Abstract
Extreme weather events are occurring more often, resulting in increasingly frequent mass mortality events for plants and animals. Identifying why individuals die during these events and their long-term consequences for populations can enable a mechanistic understanding of species’ vulnerability to global change. Here we report on early-arriving purple martins (Progne subis)—a migratory songbird—that were killed at >50% of their breeding sites across two US states during a severe winter storm event in 2021. Victims exhibited substantial allelic differences from individuals sampled before and after the storm event. The surviving population suffered delayed breeding, reproductive failure and, in 2022, late breeding-ground arrival. Phenological trait values returned to the mean by 2024, yet the population may be unlikely to recover demographically until at least 2027. Purple martins are markedly declining in the region and signatures of past events suggest that frequent mass mortality events may be challenging their resiliency.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw genomic sequences are deposited on NCBI (no. PRJNA1241750). Arrival and nest data are citizen science databases managed by the PMCA. Arrivals can be obtained through the data portal access (https://www.purplemartin.org/research/8/scout-arrival-study/). Access to nest data can be obtained by submitting a request to the PMCA and agreeing to data use policies. Source data are provided with this paper.
Code availability
Scripts for all analyses are available via GitHub at https://github.com/Mstager/Purple_Martin_scripts.git.
References
Lande, R. Risks of population extinction from demographic and environmental stochasticity and random catastrophes. Am. Nat. 142, 911–927 (1993).
Reed, D. H., O’Grady, J. J., Ballou, J. D. & Frankham, R. The frequency and severity of catastrophic die-offs in vertebrates. Anim. Conserv. 6, 109–114 (2003).
Ameca y Juárez, E. I., Mace, G. M., Cowlishaw, G. & Pettorelli, N. Natural population die-offs: causes and consequences for terrestrial mammals. Trends Ecol. Evol. 27, 272–277 (2012).
Ostfeld, R. S. & Keesing, F. Pulsed resources and community dynamics of consumers in terrestrial ecosystems. Trends Ecol. Evol. 15, 232–237 (2000).
Thébault, E. & Fontaine, C. Stability of ecological communities and the architecture of mutualistic and trophic networks. Science 329, 853–856 (2010).
Renner, H. M. et al. Catastrophic and persistent loss of common murres after a marine heatwave. Science 386, 1272–1276 (2024).
Campbell-Staton, S. C. et al. Winter storms drive rapid phenotypic, regulatory, and genomic shifts in the green anole lizard. Science 357, 495–498 (2017).
Fey, S. B. et al. Recent shifts in the occurrence, cause, and magnitude of animal mass mortality events. Proc. Natl Acad. Sci. USA 112, 1083–1088 (2015).
Cohen, J. et al. Recent Arctic amplification and extreme mid-latitude weather. Nat. Geosci. 7, 627–637 (2014).
Donihue, C. M. et al. Hurricane-induced selection on the morphology of an island lizard. Nature 560, 88–91 (2018).
Brown, C. R. & Brown, M. B. Intense natural selection on body size and wing and tail asymmetry in cliff swallows during severe weather. Evolution 52, 1461–1475 (1998).
Bumpus, H. C. Eleventh lecture. The elimination of the unfit as illustrated by the introduced sparrow, Passer domesticus. (A fourth contribution to the study of variation.). In Biological Lectures: Woods Hole Marine Biological Laboratory 209–225 (Ginn & Co., 1898).
Brown, C. R., O’Brien, V. A., Wagnon, G. S., Moore, A. T. & Sherry, R. Nonrandom weather-related mortality in a purple martin (Progne subis) roost. Southw. Naturalist 62, 210–212 (2017).
Brown, M. B. & Brown, C. R. Intense natural selection on morphology of cliff swallows (Petrochelidon pyrrhonota) a decade later: did the population move between adaptive peaks?. Auk 128, 69–77 (2011).
Piatt, J. F. et al. Extreme mortality and reproductive failure of common murres resulting from the northeast Pacific marine heatwave of 2014–2016. PLoS ONE 15, e0226087 (2020).
Aikens, E. O., Merkle, J. A., Xu, W. & Sawyer, H. Pronghorn movements and mortality during extreme weather highlight the critical importance of connectivity. Curr. Biol. 35, 1927–1934 (2025).
Senner, N. R. et al. When Siberia came to the Netherlands: the response of continental black-tailed godwits to a rare spring weather event. J. Anim. Ecol. 84, 1164–1176 (2015).
Monthly Synoptic Discussion for February 2021. NOAA National Centers for Environmental Information https://www.ncei.noaa.gov/access/monitoring/monthly-report/synoptic/202102 (2021).
Rolland, V., Balenger, S. L., Grindstaff, J. L. & Siefferman, L. Natural selection after severe winter favors larger and duller bluebirds. Am. Nat. 204, 561–573 (2024).
Walsh, J. et al. Chapter 2: Our Changing Climate. Climate Change Impacts in the United States: The Third National Climate Assessment (2014); https://www.nrc.gov/docs/ML1821/ML18215A328.pdf
Thornton, P. E. et al. Daymet: daily surface weather data on a 1-km grid for North America, version 3. ORNL DAAC https://doi.org/10.3334/ORNLDAAC/1328 (2016).
Neufeld, L. R. et al. Breeding latitude is associated with the timing of nesting and migration around the annual calendar among Purple Martin (Progne subis) populations. J. Ornithol. 162, 1009–1024 (2021).
Winkler, D. W., Luo, M. K. & Rakhimberdiev, E. Temperature effects on food supply and chick mortality in tree swallows (Tachycineta bicolor). Oecologia 173, 129–138 (2013).
Williams, C. Studies in the effect of weather conditions on the activity and abundance of insect populations. Philos. Trans. R. Soc. Lond. B https://doi.org/10.1098/rstb.1961.0011 (1961).
Taylor, L. R. Analysis of the effect of temperature on insects in flight. J. Anim. Ecol. 32, 99–117 (1963).
Peng, R. K., Fletcher, C. R. & Sutton, S. L. The effect of microclimate on flying dipterans. Int. J. Biometeorol. 36, 69–76 (1992).
Briers, R. A., Cariss, H. M. & Gee, J. H. R. Flight activity of adult stoneflies in relation to weather. Ecol. Entomol. 28, 31–40 (2003).
Newton, I. Weather-related mass-mortality events in migrants. Ibis 149, 453–467 (2007).
Guglielmo, C. G., McGuire, L. P., Gerson, A. R. & Seewagen, C. L. Simple, rapid, and non-invasive measurement of fat, lean, and total water masses of live birds using quantitative magnetic resonance. J. Ornithol. 152, 75–85 (2011).
Boyle, W. A., Winkler, D. W. & Guglielmo, C. G. Rapid loss of fat but not lean mass prior to chick provisioning supports the flight efficiency hypothesis in tree swallows. Funct. Ecol. 26, 895–903 (2012).
Kendeigh, S. C. Effect of air temperature on the rate of energy metabolism in the English sparrow. J. Exp. Zool. 96, 1–16 (1944).
Both, C., Bijlsma, R. G. & Ouwehand, J. Repeatability in spring arrival dates in pied flycatchers varies among years and sexes. Ardea 104, 3–21 (2016).
Stutchbury, B. J. M. et al. Tracking long-distance songbird migration by using geolocators. Science 323, 896–896 (2009).
Fraser, K. C., Shave, A., de Greef, E., Siegrist, J. & Garroway, C. J. Individual variability in migration timing can explain long-term, population-level advances in a songbird. Front. Ecol. Evol. 7, (2019).
Brown, C. R. & Bomberger Brown, M. Weather-mediated natural selection on arrival time in cliff swallows (Petrochelidon pyrrhonota). Behav. Ecol. Sociobiol. 47, 339–345 (2000).
Kokko, H., Gunnarsson, T. G., Morrell, L. J. & Gill, J. A. Why do female migratory birds arrive later than males?. J. Anim. Ecol. 75, 1293–1303 (2006).
Enbody, E. D. et al. Community-wide genome sequencing reveals 30 years of Darwin’s finch evolution. Science 381, eadf6218 (2023).
Lynch, M., Wei, W., Ye, Z. & Pfrender, M. The genome-wide signature of short-term temporal selection. Proc. Natl Acad. Sci. USA 121, e2307107121 (2024).
De Greef, E., Suh, A., Thorstensen, M. J., Delmore, K. E. & Fraser, K. C. Genomic architecture of migration timing in a long-distance migratory songbird. Sci. Rep. 13, 2437 (2023).
Machado, H. E. et al. Broad geographic sampling reveals the shared basis and environmental correlates of seasonal adaptation in Drosophila. eLife 10, e67577 (2021).
Singh, D., Swarup, V., Le, H. & Kumar, V. Transcriptional signatures in liver reveal metabolic adaptations to seasons in migratory blackheaded buntings. Front. Physiol. 9, 1568 (2018).
Srikanth, K., Kwon, A., Lee, E. & Chung, H. Characterization of genes and pathways that respond to heat stress in Holstein calves through transcriptome analysis. Cell Stress Chaperones 22, 29 (2016).
Singh, A. K. et al. Genomewide expression analysis of the heat stress response in dermal fibroblasts of Tharparkar (zebu) and Karan-Fries (zebu × taurine) cattle. Cell Stress Chaperones 25, 327–344 (2020).
Yu, J., Lou, Y. & Zhao, A. Transcriptome analysis of follicles reveals the importance of autophagy and hormones in regulating broodiness of Zhedong white goose. Sci. Rep. 6, 36877 (2016).
Aboura, A. et al. Array comparative genomic hybridization profiling analysis reveals deoxyribonucleic acid copy number variations associated with premature ovarian failure. J. Clin. Endocrinol. Metab. 94, 4540–4546 (2009).
Morton, E. S. & Derrickson, K. C. The biological significance of age-specific return schedules in breeding purple martins. Condor 92, 1040 (1990).
Bull, J. C. et al. Climate causes shifts in grey seal phenology by modifying age structure. Proc. Biol. Sci. 288, 20212284 (2021).
Taff, C. C. & Shipley, J. R. Inconsistent shifts in warming and temperature variability are linked to reduced avian fitness. Nat. Commun. 14, 7400 (2023).
Truchelut, R. E. et al. Earlier onset of North Atlantic hurricane season with warming oceans. Nat. Commun. 13, 4646 (2022).
Senner, N. R., Stager, M. & Cheviron, Z. A. Spatial and temporal heterogeneity in climate change limits species’ dispersal capabilities and adaptive potential. Ecography 41, 1428–1440 (2018).
Youngflesh, C. et al. Demographic consequences of phenological asynchrony for North American songbirds. Proc. Natl Acad. Sci. USA 120, e2221961120 (2023).
Fink, D. et al. eBird Status and Trends. https://doi.org/10.2173/WZTW8903 (Cornell Lab of Ornithology, 2023).
Shaw, R. E. et al. Global meta-analysis shows action is needed to halt genetic diversity loss. Nature https://doi.org/10.1038/s41586-024-08458-x (2025).
Brown, C. R., Airola, D. A. & Tarof, S. Purple Martin (Progne subis), version 2.0. Birds of the World https://doi.org/10.2173/bow.purmar.02 (2021).
Nebel, S., Mills, A., McCracken, J. & Taylor, P. Declines of aerial insectivores in North America follow a geographic gradient. Avian Conserv. Ecol. 5, (2010).
Rosenberg, K. V. et al. Decline of the North American avifauna. Science 366, 120–124 (2019).
Sauer, J. R., Link, W. A. & Hines, J. E. The North American breeding bird survey, analysis results 1966–2019. Data release 1051. US Geological Survey (USGS) https://doi.org/10.5066/P96A7675 (2020).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2025).
Climate at a Glance. Divisional Mapping. National Centers for Environmental Information (NCEI) (accessed 10 March 2025); https://www.ncei.noaa.gov/access/monitoring/climate-at-a-glance/divisional/mapping
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Miles, A. et al. cggh/scikit-allel: v1.3.13. Zenodo https://doi.org/10.5281/zenodo.13772087 (2024).
Thomas, P. D. et al. PANTHER: making genome-scale phylogenetics accessible to all. Protein Sci. 31, 8–22 (2022).
Czech, L., Spence, J. P. & Expósito-Alonso, M. grenedalf: population genetic statistics for the next generation of pool sequencing. Bioinformatics 40, btae508 (2024).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Pinheiro, J. et al. nlme: Linear and Nonlinear Mixed Effects Models. R package version 3.1-168 https://cran.r-project.org/web/packages/nlme/index.html (2024).
Stutchbury, B. J. M., Hill, J. R., III, Kramer, P. M., Rush, S. A. & Tarof, S. A. Sex and age-specific annual survival in a neotropical migratory songbird, the purple martin (Progne subis). Auk 126, 278–287 (2009).
Stoffel, M. A., Nakagawa, S. & Schielzeth, H. rptR: repeatability estimation and variance decomposition by generalized linear mixed-effects models. Methods Ecol. Evol. 8, 1639–1644 (2017).
Legendre, S. Projecting populations. In Population Ecology in Practice (eds Murray, D. L. & Sandercock, B. K.) 193–214 (2020).
Brown, C. R. Clutch size and reproductive success of adult and subadult purple martins. Southw. Naturalist 23, 597–603 (1978).
Wagner, R. H., Schug, M. D. & Morton, E. S. Condition-dependent control of paternity by female purple martins: implications for coloniality. Behav. Ecol. Sociobiol. 38, 379–389 (1996).
Finlay, J. C. Breeding biology of purple martins at the northern limit of their range. Wilson Bull. 83, (1971).
de Greef, E., Brashear, W., Delmore, K. E. & Fraser, K. C. Population structure, patterns of natal dispersal and demographic history in a declining aerial insectivore, the purple martin Progne subis. J. Avian Biol. 2022, e02929 (2022).
Acknowledgements
We thank the PMCA landlords; the citizen scientists that contributed to the data collection; S. Cardiff, N. Mason and the LSU Museum of Natural Sciences; M. Harvey and P. Lavretsky at the UT El Paso Biodiversity Collection; UMass students J. Applegate and S. Thakur; A. Gerson for use of the QMR; J. Casey for hospitality; E. de Greef for sharing the purple martin genome annotation; and UMass BaMPhEE, W. Tong, W. Burnside and the Stager and Senner lab groups for comments on a previous version of this manuscript. This work was funded by grants from the PMCA (to A.M.F. and M.S.), as well as funding from the University of Central Florida (to A.M.F.) and the University of Massachusetts Amherst (to M.S.).
Author information
Authors and Affiliations
Contributions
M.S. and A.M.F. designed the study. J.S. and R.K.A. helped to coordinate collection of the carcasses by the museum. D.L.D. received carcasses at the museum and prepared them as specimens. A.M.F., J.S. and R.K.A. conducted field sampling. J.K.G. provided permits for field sampling. K.D. performed morphological and body composition assays. A.M.F. and M.S. performed laboratory work. P.M.B. and R.R.F. conducted genomic analyses. M.S. and N.R.S. designed and executed all other analyses and drafted the manuscript. All authors contributed feedback on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Eric Ameca, Shane Campbell-Staton and Virginie Rolland for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Body composition of purple martin carcasses.
a) Lean and (b) fat masses (as a proportion of total body mass) for n = 281 carcasses. Dashed line indicates mean value.
Extended Data Fig. 2 Early arrival dates for 10 U.S. South Central states.
Gray dots represent individual reports. Larger black circles indicate the mean arrival date for the year with + indicating the first and third quartiles. Vertical line denotes the mean arrival date for the period of 1998-2021 with light gray shading to indicate 2 standard deviations from the mean. Red shaded horizontal bar highlights 2022, the year following the storm event. Black arrows indicate years in which mean arrival date \(\ge\)2 standard deviations from the long-term mean. Sample sizes are as follows: nTexas = 9589, nLouisiana = 3190, nMississippi = 1337, nAlabama = 2314, nGeorgia = 1773, nFlorida = 3569, nArkansas = 1544, and nOklahoma = 1946.
Extended Data Fig. 3 Nest success across the Louisiana breeding season.
Relationship between nest success (that is, whether or not a nest had at least one young fledge) and first egg date from n = 1095 nests (open circles) in Louisiana from 1998-2024. Line represents the results of a logistic regression (\(\beta\) = −0.03 \(\pm \,\)0.005, P < 0.01) with gray shading to indicate 95% confidence interval.
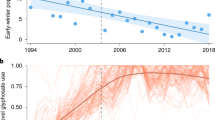
Extended Data Fig. 4 Louisiana arrival and temperature trends.
Long-term trends in first arrival dates (blue dots; n = 3190 arrivals) and June temperatures (black dots; n = 27 years) in Louisiana from 1998-2024. Blue line represents a linear mixed-effect regression (\(\beta\) = 0.15 \(\pm \,\)0.07, p = 0.03) with shaded gray indicating 95% confidence interval; only sites with >10 years of data were included, and site was incorporated as a random effect. Red line represents the long-term trend in mean June temperatures in Louisiana (1998-2024) resulting from a linear regression (\(\beta\) = 0.04 \(\pm \,\)0.01, P = 0.01); dotted lines indicate 95% confidence interval.
Extended Data Fig. 5 Principal components analysis on genetic variation across the genome using 922,941 SNPs.
Each point represents one of 61 individuals color-coded by sampling site. Symbols indicate sample cohort: individuals that died during the storm (x) or individuals sampled in May 2021 after the storm (dots).
Extended Data Fig. 6 Providing perspective on latitudinal correlations through random subsampling of SNP dataset.
Histogram depicting the number of SNPs that significantly changed with latitude in each of 100, randomly subsampled groups composed of 2,624 non-Fst-outlier SNPs. Blue dotted line indicates the number of Fst outlier SNPs whose allele frequency corresponded with latitude.
Supplementary information
Source data
Source Data Fig. 1 (download XLSX )
(Sheet 1) Localities for 30 sampling sites of purple martin carcasses. Latitudes and longitudes have been rounded to the nearest 100th to maintain privacy of homeowners. (Sheet 2) Origin and morphological data for individuals sampled in May 2021. Latitudes and longitudes have been rounded to the nearest 100th to maintain homeowner privacy. Sex (male (M) or female (F)) and age (subadult (SY) or adult (ASY)) were determined by plumage.
Source Data Fig. 3 (download XLSX )
Individual information for 66 samples for which whole-genome sequencing was performed. Collection date and collection locale with corresponding latitudes and longitudes (rounded to the nearest 100th to maintain homeowner privacy) are listed. Cohort (survivor or victim), age (ASY (after second year), adult; SY (second year), subadult) and sex (M, male; F, female) are indicated. BioSample accession no. corresponds to raw sequence archived on the NCBI’s Sequence Read Archive.
Source Data Extended Data Fig. 1 (download XLSX )
Individual data for 292 carcasses, including LSU MNS tissue number, collection date, location, latitude and longitude; sex (determined by plumage); morphological measurements; and body composition (fat and lean masses).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Stager, M., Benham, P.M., Senner, N.R. et al. Storm-induced mass mortality results in both immediate and long-term consequences for a migratory songbird. Nat Ecol Evol (2026). https://doi.org/10.1038/s41559-026-03005-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41559-026-03005-5