Abstract
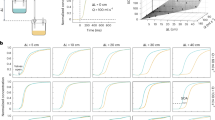
Sensorimotor feedback is a fundamental mechanism of active sensing. In olfaction, the primary motor behaviour is the sniff. Thus, in active olfactory sensing, we would expect the dynamics of the sniff to change according to detailed odour characteristics. Furthermore, percept-related modulations of sniffing behaviour should correspond to neural activity in participating brain regions. Here we analysed a high-precision functional MRI dataset including more than 4,300 sniffs per participant taken of 160 odours during ~18 hours of scanning, to probe the relationship between odour-induced sniff modulations and perceptual features at a high level of granularity. We found that fine-grained perceptual odour information—and even odour identity—can be decoded from sniffing dynamics, and that olfactory brain regions, particularly the amygdala, are involved in percept-driven modulation of sniffing behaviour. Thus, olfactory cortical areas participate in real-time modulations of sniffs according to perceptual properties of the odour at a high level of granularity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Analyses presented here used a previously published database30,36. Additional behavioural data for group-level analyses are available via GitHub at https://github.com/viveksgr/SFP.
Code availability
All analyses were performed in MATLAB R2023b. Code for preprocessing and reproducing the results presented in this article is available via GitHub at https://github.com/viveksgr/SFP.
References
Kepecs, A., Uchida, N. & Mainen, Z. F. The sniff as a unit of olfactory processing. Chem. Senses 31, 167–179 (2006).
Wachowiak, M. All in a sniff: olfaction as a model for active sensing. Neuron 71, 962–973 (2011).
Mainland, J. & Sobel, N. The sniff is part of the olfactory percept. Chem. Senses 31, 181–196 (2006).
Sobel, N. et al. Sniffing and smelling: separate subsystems in the human olfactory cortex. Nature 392, 282–286 (1998).
Laing, D. G. Natural sniffing gives optimum odour perception for humans. Perception 12, 99–117 (1983).
Cury, K. M. & Uchida, N. Robust odor coding via inhalation-coupled transient activity in the mammalian olfactory bulb. Neuron 68, 570–585 (2010).
Verhagen, J. V., Wesson, D. W., Netoff, T. I., White, J. A. & Wachowiak, M. Sniffing controls an adaptive filter of sensory input to the olfactory bulb. Nat. Neurosci. 10, 631–639 (2007).
Kareken, D. A. et al. Olfactory system activation from sniffing: effects in piriform and orbitofrontal cortex. NeuroImage 22, 456–465 (2004).
Wachowiak, M. & Cohen, L. B. Representation of odorants by receptor neuron input to the mouse olfactory bulb. Neuron 32, 723–735 (2001).
Miura, K., Mainen, Z. F. & Uchida, N. Odor representations in olfactory cortex: distributed rate coding and decorrelated population activity. Neuron 74, 1087–1098 (2012).
Shusterman, R., Smear, M. C., Koulakov, A. A. & Rinberg, D. Precise olfactory responses tile the sniff cycle. Nat. Neurosci. 14, 1039–1044 (2011).
Jordan, R., Fukunaga, I., Kollo, M. & Schaefer, A. T. Active sampling state dynamically enhances olfactory bulb odor representation. Neuron 98, 1214–1228 (2018).
Johnson, B. N., Mainland, J. D. & Sobel, N. Rapid olfactory processing implicates subcortical control of an olfactomotor system. J. Neurophysiol. 90, 1084–1094 (2003).
Prescott, J., Burns, J. & Frank, R. A. Influence of odor hedonics, food-relatedness, and motivational state on human sniffing. Chemosens. Percept. 3, 85–90 (2010).
Ferdenzi, C., Fournel, A., Thévenet, M., Coppin, G. & Bensafi, M. Viewing olfactory affective responses through the sniff prism: effect of perceptual dimensions and age on olfactomotor responses to odors. Front. Psychol. 6, 1776 (2015).
Johnson, B. N., Russell, C., Khan, R. M. & Sobel, N. A comparison of methods for sniff measurement concurrent with olfactory tasks in humans. Chem. Senses 31, 795–806 (2006).
de los Cobos Pallares, F. P., Bautista, T. G., Stanić, D., Egger, V. & Dutschmann, M. Brainstem-mediated sniffing and respiratory modulation during odor stimulation. Respir. Physiol. Neurobiol. 233, 17–24 (2016).
Dlouhy, B. J. et al. Breathing inhibited when seizures spread to the amygdala and upon amygdala stimulation. J. Neurosci. 35, 10281–10289 (2015).
Masaoka, Y. & Homma, I. in Post-genomic Perspectives in Modeling and Control of Breathing (eds Champagnat, J. et al.) 9–14 (Springer, 2004).
Nobis, W. P. et al. Amygdala-stimulation-induced apnea is attention and nasal-breathing dependent. Ann. Neurol. 83, 460–471 (2018).
Nobis, W. P. et al. The effect of seizure spread to the amygdala on respiration and onset of ictal central apnea. J. Neurosurg. 132, 1313–1323 (2019).
Yang, C. F., Kim, E. J., Callaway, E. M. & Feldman, J. L. Monosynaptic projections to excitatory and inhibitory preBötzinger complex neurons. Front. Neuroanat. 14, 58 (2020).
Trevizan-Baú, P. et al. Forebrain projection neurons target functionally diverse respiratory control areas in the midbrain, pons, and medulla oblongata. J. Comp. Neurol. 529, 2243–2264 (2021).
Rhone, A. E. et al. A human amygdala site that inhibits respiration and elicits apnea in pediatric epilepsy. JCI Insight 5, e134852 (2020).
Lacuey, N., Zonjy, B., Londono, L. & Lhatoo, S. D. Amygdala and hippocampus are symptomatogenic zones for central apneic seizures. Neurology 88, 701–705 (2017).
Howard, J. D., Plailly, J., Grueschow, M., Haynes, J.-D. & Gottfried, J. A. Odor quality coding and categorization in human posterior piriform cortex. Nat. Neurosci. 12, 932–938 (2009).
Stettler, D. D. & Axel, R. Representations of odor in the piriform cortex. Neuron 63, 854–864 (2009).
Roland, B., Deneux, T., Franks, K. M., Bathellier, B. & Fleischmann, A. Odor identity coding by distributed ensembles of neurons in the mouse olfactory cortex. eLife 6, e26337 (2017).
Pashkovski, S. L. et al. Structure and flexibility in cortical representations of odour space. Nature 583, 253–258 (2020).
Sagar, V., Shanahan, L. K., Zelano, C. M., Gottfried, J. A. & Kahnt, T. High-precision mapping reveals the structure of odor coding in the human brain. Nat. Neurosci. 26, 1595–1602 (2023).
Schoonover, C. E., Ohashi, S. N., Axel, R. & Fink, A. J. Representational drift in primary olfactory cortex. Nature 594, 541–546 (2021).
Gonzalez, J., Torterolo, P., Bolding, K. A. & Tort, A. B. L. Communication subspace dynamics of the canonical olfactory pathway. iScience 27, 12 (2024).
Kay, L. M. et al. Olfactory oscillations: the what, how and what for. Trends Neurosci. 32, 207–214 (2009).
Rosero, M. A. & Aylwin, M. L. Sniffing shapes the dynamics of olfactory bulb gamma oscillations in awake behaving rats. Eur. J. Neurosci. 34, 787–799 (2011).
Stern, M., Bolding, K. A., Abbott, L. & Franks, K. M. A transformation from temporal to ensemble coding in a model of piriform cortex. eLife 7, e34831 (2018).
Sagar, V., Shanahan, L. K., Zelano, C. M., Gottfried, J. A. & Kahnt, T. Neural Encoding Models of Olfaction (NEMO) dataset. Zenodo https://doi.org/10.5281/zenodo.7636722 (2023).
Noto, T., Zhou, G., Schuele, S., Templer, J. & Zelano, C. Automated analysis of breathing waveforms using BreathMetrics: a respiratory signal processing toolbox. Chem. Senses 43, 583–597 (2018).
Keller, A. & Vosshall, L. B. Olfactory perception of chemically diverse molecules. BMC Neurosci. 17, 1–17 (2016).
Keller, A. et al. Predicting human olfactory perception from chemical features of odor molecules. Science 355, 820–826 (2017).
Secundo, L. et al. Individual olfactory perception reveals meaningful nonolfactory genetic information. Proc. Natl Acad. Sci. USA 112, 8750–8755 (2015).
Bensafi, M., Pouliot, S. & Sobel, N. Odorant-specific patterns of sniffing during imagery distinguish ‘bad’ and ‘good’ olfactory imagers. Chem. Senses 30, 521–529 (2005).
Moberly, A. H. et al. Olfactory inputs modulate respiration-related rhythmic activity in the prefrontal cortex and freezing behavior. Nat. Commun. 9, 1528 (2018).
Echevarria-Cooper, S. L. et al. Mapping the microstructure and striae of the human olfactory tract with diffusion MRI. J. Neurosci. 42, 58–68 (2021).
Zhou, G., Lane, G., Cooper, S. L., Kahnt, T. & Zelano, C. Characterizing functional pathways of the human olfactory system. eLife 8, e47177 (2019).
Zelano, C. et al. Nasal respiration entrains human limbic oscillations and modulates cognitive function. J. Neurosci. 36, 12448–12467 (2016).
Smear, M., Shusterman, R., O’Connor, R., Bozza, T. & Rinberg, D. Perception of sniff phase in mouse olfaction. Nature 479, 397–400 (2011).
Rojas-Líbano, D., Frederick, D. E., Egaña, J. I. & Kay, L. M. The olfactory bulb theta rhythm follows all frequencies of diaphragmatic respiration in the freely behaving rat. Front. Behav. Neurosci. 8, 214 (2014).
Fontanini, A. & Bower, J. M. Slow-waves in the olfactory system: an olfactory perspective on cortical rhythms. Trends Neurosci. 29, 429–437 (2006).
Adrian, E. The electrical activity of the mammalian olfactory bulb. Electroencephalogr. Clin. Neurophysiol. 2, 377–388 (1950).
Liversedge, S. P. & Findlay, J. M. Saccadic eye movements and cognition. Trends Cogn. Sci. 4, 6–14 (2000).
Hoffman, J. E. & Subramaniam, B. The role of visual attention in saccadic eye movements. Percept. Psychophys. 57, 787–795 (1995).
Ahissar, E. & Arieli, A. Figuring space by time. Neuron 32, 185–201 (2001).
Leszczynski, M. & Schroeder, C. E. The role of neuronal oscillations in visual active sensing. Front. Integr. Neurosci. 13, 32 (2019).
Rucci, M., Iovin, R., Poletti, M. & Santini, F. Miniature eye movements enhance fine spatial detail. Nature 447, 852–855 (2007).
Ko, H.-K., Poletti, M. & Rucci, M. Microsaccades precisely relocate gaze in a high visual acuity task. Nat. Neurosci. 13, 1549–1553 (2010).
Szwed, M., Bagdasarian, K. & Ahissar, E. Encoding of vibrissal active touch. Neuron 40, 621–630 (2003).
Ganguly, K. & Kleinfeld, D. Goal-directed whisking increases phase-locking between vibrissa movement and electrical activity in primary sensory cortex in rat. Proc. Natl Acad. Sci. USA 101, 12348–12353 (2004).
Whitton, J. P., Hancock, K. E. & Polley, D. B. Immersive audiomotor game play enhances neural and perceptual salience of weak signals in noise. Proc. Natl Acad. Sci. USA 111, E2606–E2615 (2014).
Kehl, M. S. et al. Single-neuron representations of odours in the human brain. Nature 634, 626–634 (2024).
Hernandez, D. E. et al. Fast updating feedback from piriform cortex to the olfactory bulb relays multimodal identity and reward contingency signals during rule-reversal. Nat. Commun. 16, 937 (2025).
Cain, W. S. & Murphy, C. L. Interaction between chemoreceptive modalities of odour and irritation. Nature 284, 255–257 (1980).
Brand, G. Olfactory/trigeminal interactions in nasal chemoreception. Neurosci. Biobehav. Rev. 30, 908–917 (2006).
Frasnelli, J., Charbonneau, G., Collignon, O. & Lepore, F. Odor localization and sniffing. Chem. Senses 34, 139–144 (2009).
Rehan, M. et al. Predicting odor pleasantness from odorant structure: pleasantness as a reflection of the physical world. J. Neurosci. 27, 10015–10023 (2007).
Zarzo, M. Psychologic dimensions in the perception of everyday odors: pleasantness and edibility. J. Sens. Stud. 23, 354–376 (2008).
Yeshurun, Y. & Sobel, N. An odor is not worth a thousand words: from multidimensional odors to unidimensional odor objects. Ann. Rev. Psychol. 61, 219–241 (2010).
Noto, T., Zhou, G., Yang, Q., Lane, G. & Zelano, C. Human primary olfactory amygdala subregions form distinct functional networks, suggesting distinct olfactory functions. Front. Syst. Neurosci. 15, 752320 (2021).
Scalia, F. & Winans, S. S. The differential projections of the olfactory bulb and accessory olfactory bulb in mammals. J. Comp. Neurol. 161, 31–55 (1975).
Carmichael, S. T., Clugnet, M. C. & Price, J. L. Central olfactory connections in the macaque monkey. J. Comp. Neurol. 346, 403–434 (1994).
Yang, Q. et al. The human olfactory amygdala: anatomical connections between the olfactory bulb and amygdala subregions. Imaging Neurosci. 3, imag_a_00571 (2025).
Sosulski, D. L., Bloom, M. L., Cutforth, T., Axel, R. & Datta, S. R. Distinct representations of olfactory information in different cortical centres. Nature 472, 213–216 (2011).
Root, C. M., Denny, C. A., Hen, R. & Axel, R. The participation of cortical amygdala in innate, odour-driven behaviour. Nature 515, 269–273 (2014).
Short, S. M. & Wachowiak, M. Temporal dynamics of inhalation-linked activity across defined subpopulations of mouse olfactory bulb neurons imaged in vivo. eNeuro 6, ENEURO.0189-19.2019 (2019).
Sobel, N. et al. Odorant-induced and sniff-induced activation in the cerebellum of the human. J. Neurosci. 18, 8990–9001 (1998).
Friston, K. J., Williams, S., Howard, R., Frackowiak, R. S. & Turner, R. Movement-related effects in fMRI time-series. Magn. Reson. Med. 35, 346–355 (1996).
Prince, J. S. et al. Improving the accuracy of single-trial fMRI response estimates using GLMsingle. eLife 11, e77599 (2022).
Allen, E. J. et al. A massive 7T fMRI dataset to bridge cognitive neuroscience and artificial intelligence. Nat. Neurosci. 25, 116–126 (2022).
Acknowledgements
This work was supported by the NIDCD, NIMH and NINDS of the National Institutes of Health under award numbers R01DC015426 (to C.Z.), R01DC018539 (to C.Z), T32MH067564 (to V.S.) and T32NS047987 (to V.S.). This research was supported in part by the Intramural Research Program of the National Institutes of Health (NIH) (ZIA DA000642 to T.K.). The contributions of the NIH author (T.K.) are considered works of the United States Government. The findings and conclusions presented in this paper are those of the authors and do not necessarily reflect the views of the NIH or the US Department of Health and Human Services. This research was supported in part through the computational resources and staff contributions provided for the Quest high performance computing facility at Northwestern University, which is jointly supported by the Office of the Provost, the Office for Research and Northwestern University Information Technology. We thank Rockefeller University and Sage Bionetworks–DREAM for the DREAM database of perceptual ratings.
Author information
Authors and Affiliations
Contributions
V.S., G.L., T.K. and C.Z. devised the experiment and conceptualized the analyses. A.S., Q.Y. and N.A. collected additional experimental data. V.S., A.S. and G.Z. performed the analyses. V.S., G.L., T.K. and C.Z. prepared and revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Human Behaviour thanks Thomas Hummel, Aharon Ravia and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Decoding accuracy based on raw sniff traces.
a. Decoding accuracy (percentage of trials reported correctly) of a multiclass SVM model that predicts (out of sample) odor identity based on the raw sniff traces at each trial. Decoding was significant for all subjects (mean subject: classification performance = 2.57%, P = 0.000, one-tailed-weighted z-test; all subjects: P = 0.000; one-tailed z-test; N = 4560 (S1), N = 4320 (S2, S3) trials). Data are presented as mean values and error bars indicate 95% C.I. b. Distribution of correlation of perceptual ratings (except intensity and pleasantness) between the odor predicted by the classifier in Fig. 2c. and the correct odor, on trials in which the decoder made a wrong prediction. The perceptual ratings of the odor incorrectly predicted by model were significantly correlated to that of correct odor (S1: rSVM = 0.097, P = 0.000, 4480 trials, two-tailed Wilcoxon signed rank test; S2: rSVM = 0.320, P = 0.000, 4132 trials, two-tailed Wilcoxon signed rank test; S3: rSVM = 0.309, P = 0.000, 4229 trials, two-tailed Wilcoxon signed rank test). Further, this correlation was significant after accounting for mean correlation (rmean) of perceptual ratings among the odors in S2 (rSVM-rmean = 0.013, P = 0.006, one-tailed permutation test) and S3 (rSVM-rmean = 0.010, P = 0.002, one-tailed permutation test) but not S1 (rSVM-rmean = −0.003,P = 0.480, one-tailed permutation test). The center lines denote the median, the box limits denote the upper and lower quartiles, the whiskers are 1.5× interquartile range and points are outliers. c. Pattern correlation analysis after regressing out intensity and pleasantness, showing cross-trial correlation among pairs of trials of the same or different odors (rsame = 0.083; rdiff = 0.013; for mean subject, P = 0.000 for each subject; N = 4560 trials in S1 and N = 4320 trials in S2 and S3; two-tailed Wilcoxon rank sum test). Data are presented as mean values and error bars indicate 95% C.I.
Extended Data Fig. 2 Subject-wise sniff traces.
a. Subject-wise sniff traces for all trials used in the study. Trials are sorted in the temporal order of their presentation. b. Feature-based representation of sniff trials along 24 features. The first 12 features were extracted from the BreathMetrics toolbox and the remaining were derived features described in the Methods.
Extended Data Fig. 3 Multilinguistic regression models based on group subject data.
Performance of multilinguistic regression models for each descriptor, predicting discretized ratings (low, medium, high) of a perceptual descriptor on a trial, given the sniffing parameters when using average ratings across subjects (left) and ratings from the DREAM dataset (right). Model performance was significant for several descriptors (chance correct = 33.34%, P < 0.05, for 14/21 descriptors in all subjects for group average, and 7/21 descriptors for ratings from the DREAM dataset, one-tailed binomial test (FDR corrected), N = 4560 trials in S1 and N = 4320 trials in S2 and S3). Data are presented as mean values and error bars indicate 95% C.I.
Extended Data Fig. 4 Odor decoding accuracy from sniffs acquired from additional subjects.
Decoding accuracy on additional behavioral data from 3 subjects. Percentage of trials reported correctly of a multiclass SVM model that predicts (out of sample) odor identity for 10 odors based on the features of the sniff at each trial (200 trials/subject). Together with the 3 original subjects, the combined dataset of 6 subjects shows significant group-level decoding (P = 0.031, N = 6, two-tailed Wilcoxon signed rank test), suggesting generalizability of the effect. Data are presented as mean values and error bars indicate 95% C.I.
Extended Data Fig. 5 SVM decoding accuracy after accounting for odors with high accuracy.
The left three panels show histograms of mean accuracy (number of trials with significant prediction/total number of trials) per odor for 3 subjects. The right panel shows decoding accuracy (percentage of trials reported correctly) of a multiclass SVM model that predicts (out of sample) odor identity based on the features of the sniff at each trial, after removing odors that show high prediction accuracy (> 0.2) as seen in the left panels. Decoding was significant for all subjects (mean subject: classification performance = 2.86%, P = 0.000, one tailed z-test). Data are presented as mean values and error bars indicate 95% C.I.
Extended Data Fig. 6 Subject-wise temporal RSA after accounting for intensity and pleasantness.
a. The temporal profile of the perceptual similarity regression coefficients (standardized beta values) (orange) overlaid on the sniff flow temporal profile (blue) averaged across subjects when perceptual similarity was based on all descriptors except intensity. Solid lines indicate the time-points when the regression coefficient was significant (P < 0.05, subject-wise one-tailed t-test, panel b for at least 2/3 subjects) and dashed lines denote otherwise. b. Temporal perceptual representational similarity (without intensity) in individual subjects. Solid lines indicate the time-points where the regression coefficient was significant (P < 0.05, FDR corrected across time-bins, one-tailed t-test, N = all unique pairs of 4560 (S1), 4320 (S2, S3) trials: 10394520 (S1) and 9329040 (S2, S3))), and dashed lines denote otherwise. c. Perceptual representational similarity (without intensity) in a. plotted as a function of sniff flow rate. d. Perceptual similarity (without intensity) plotted as a function of sniff flow for individual subjects. e. The temporal profile of the perceptual similarity regression coefficients (standardized beta values, orange) overlaid on the sniff flow temporal profile (blue) averaged across subjects when perceptual similarity was based on all descriptors except pleasantness. Solid lines indicate the time-points when the regression coefficient was significant (P < 0.05, subject-wise one-tailed t-test, panel f for at least 2/3 subjects)) and dashed lines denote otherwise. f. Temporal perceptual representational similarity (without pleasantness) for individual subjects. Solid lines indicate the time-points when the regression coefficient was significant (P < 0.05, FDR corrected across time-bins, one-tailed t-test, N = all unique pairs of 4560 (S1), 4320 (S2, S3) trials: 10394520 (S1) and 9329040 (S2, S3))) and dashed lines denote otherwise. g. Perceptual representational similarity (without pleasantness) in e. plotted as a function of sniff flow rate. h. Perceptual similarity (without pleasantness) plotted as a function of sniff flow for individual subjects. All subject-wise results were corrected for multiple comparisons across time-bins by an FDR correction (Benjamini-Hochberg procedure).
Extended Data Fig. 7 Temporal RSA for subsets of odors and different perceptual descriptors.
a. Time series RSA for different subsets of trials, to demonstrate range of variability of information peaks. Rows indicate time-series representational similarities (regression coefficients) for distinct bootstrap samples while columns indicate separate time-bins. b. Time series RSA for different perceptual descriptors. Rows indicate time-series representational similarities (regression coefficients) for distinct descriptors while columns indicate separate time-bins.
Extended Data Fig. 8 Subject-wise and control analyses for neural RSA.
a. Searchlight analyses to determine where representations of sniff shape, perceptual and random clustering of sniffs were significant (P < 0.05, t-test, FDR corrected for all conditions and subjects, except S3, perceptual cluster; P < 0.01 two-tailed t-test). Raw sniff information (in blue) was pervasive throughout the olfactory regions. A subset of these regions captured aspects of sniff information modulated by perceptual features (red for voxels overlapping with raw sniff voxels, otherwise, green). b. Regression coefficients of correlation of sniff-shape/perceptual/random clustered sniff-similarity matrices with correspondingly clustered neural data after orthogonalization of sniff and perceptual clustering. The clustering was orthogonalized using the orthogonalization algorithm described in Methods until the adjusted random index (ARI) between the clustering was insignificant (ARI before orthogonalization: S1: 0.003; S2: 0.009; S3: 0.004, P = 0.000, one-tailed permutation test; ARI after orthogonalization: S1: 0.0004; S2: 0.0008; S3: 0.0005, P > 0.1, one-tailed permutation test). Regression coefficients of raw sniffing parameters based on sniff shape were significantly higher than the random control (P = 0.000, all areas, one-tailed t-test, N = 10394520 (S1) and 9329040 (S2, S3)). Similarly, regression coefficients of perceptual clustering were significantly higher than the random control (P = 0.000, all areas, one-tailed t-test, N = 10394520 (S1) and 9329040 (S2, S3)). c. Regression coefficients of correlation of sniff-shape/perceptual/random clustered sniff-similarity matrices with correspondingly clustered neural data based on perceptual features other than intensity and pleasantness. Regression coefficients of raw sniffing parameters based on sniff shape were significantly higher than the random control (P=0.000, all areas, one-tailed t-test, N = 10394520 (S1) and 9329040 (S2, S3)). Similarly, regression coefficients of perceptual clustering were significantly higher than the random control (P=0.000, all areas, one-tailed t-test, N = 10394520 (S1) and 9329040 (S2, S3)). Data are presented as mean values and error bars indicate s.e.m. across subjects in all panels.
Extended Data Fig. 9 Searchlight-based clustered RSA for individual subjects with control analyses accounting for intensity and pleasantness.
a. Searchlight-based clustered RSA for individual subjects, with clustering performed on individual perceptual features. The brain maps show number of perceptual features for which the clustered RSA was significant (P < 0.05, one-tailed t-test, N = 10394520 (S1) and 9329040 (S2, S3)). b. Number of perceptual features (except intensity and pleasantness) for which clustered-RSA was significant across regions (averaged across all odor-responsive voxels at threshold P = 0.05, one-tailed t-test). Number of significantly modulating features were highest in the AMY (F4, 10132 = 189.788; P = 0.000, repeated measures ANOVA; P = 0.000 for post-hoc comparison between AMY and other anatomical regions, except PC; P = 0.009, pairwise t-test, FDR corrected). Data are presented as mean values and error bars indicate s.e.m. across subjects in all panels.
Extended Data Fig. 10 Similarity between measuring sniffs from breathing belt and spirometer.
a. Pearson’s r of raw sniff trace obtained from spirometer and derivative of breathing belt for 12 runs. Correlation is across a large number of samples (>106) and is highly significant (P = 0.000, mean r = 0.768, N = 1209200 samples for the shortest run, two-tailed t-test) b. Pearson’s r of sniff features extracted from the BreathMetrics toolbox for spirometer trace and derivative of breathing belt for all runs. Chance r is shown in black line and the correlation is significant in all features (all P < 0.001, one-tailed t-test, except exhaleTimeToTrough: P = 0006; inhaleDurations: P = 0.004 and exhaleDurations: P = 0.027, N = 141 (mean number of sniffs in each run)). For all panels data are presented as mean values and error bars indicate s.e.m. across runs.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sagar, V., Sheriff, A., Yang, Q. et al. The human brain modulates sniffs according to fine-grained perceptual features of odours. Nat Hum Behav 10, 137–147 (2026). https://doi.org/10.1038/s41562-025-02327-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41562-025-02327-x