Abstract
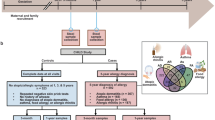
Early-life microbial exposures shape immune development and allergy risk. Food allergen sensitization, reflected by the presence of food allergen-specific immunoglobulin E (IgE), is an early indication of impaired immune tolerance. Here we show that early-life transmission of aromatic lactate-producing bifidobacteria strains in 147 children followed from birth to 5 years of age, facilitated by vaginal delivery, exposure to older siblings and exclusive breastfeeding for the first 2 months, led to increased levels of aromatic lactates in the infant gut. This microbiota–metabolite signature was inversely associated with the development of food allergen-specific IgE until 5 years and atopic dermatitis at 2 years. The observed effect was mediated by 4-hydroxy-phenyllactate, which inhibited IgE, but not IgG, production in ex vivo human immune cell cultures. Together, these findings define an early-life microbiota–metabolite–immune axis linking microbial transmission and feeding practices with reduced allergic sensitization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
High-quality, non-host shotgun metagenomic sequencing data from the ALADDIN cohort are deposited in the Sequence Read Archive (SRA) under the BioProject PRJEB84944 and PRJEB102491 at the European Nucleotide Archive (ENA). The sequencing reads from PAPS are available at ENA under accession number PRJEB51055. The sequencing reads from the BIS cohort are available at ENA under accession number PRJNA576314. The gene catalogue and MAGs are available via Zenodo at https://doi.org/10.5281/zenodo.17810079 (ref. 65). Data on individual lifestyles, metabolomics and faecal aromatic lactic acid concentrations are pseudonymized (coded) personal data, protected and are not available due to data privacy laws. Access to controlled data can requested by contacting johan.alm@ki.se. Requests will require submission and approval of a written data use agreement that ensures compliance with all applicable legal and ethical regulations. The agreement specifies that data may only be used for the purposes outlined in the approved request and may not be shared with third parties. The expected timeframe for granting access depends on completion of institutional and legal procedures, but we aim to respond to initial requests within 4 weeks. Source data are provided with this paper.
Code availability
The SNV pipeline for generating phylogenetic trees is available via GitHub at https://github.com/pennyneve/SNVcalling.
References
Ege, M. J. et al. Prenatal exposure to a farm environment modifies atopic sensitization at birth. J. Allergy Clin. Immunol. 122, 407–412 (2008).
Renz, H. & Skevaki, C. Early life microbial exposures and allergy risks: opportunities for prevention. Nat. Rev. Immunol. 21, 177–191 (2021).
Gray, L. E. K. et al. Deserters on the atopic march: risk factors, immune profile and clinical outcomes of food sensitized-tolerant infants. Allergy 75, 1404–1413 (2020).
Prescott, S. L. et al. A global survey of changing patterns of food allergy burden in children. World Allergy Org. J. 6, 21 (2013).
Schoos, A. M. et al. Sensitization trajectories in childhood revealed by using a cluster analysis. J. Allergy Clin. Immunol. 140, 1693–1699 (2017).
Torow, N. & Hornef, M. W. The neonatal window of opportunity: setting the stage for life-long host-microbial interaction and immune homeostasis. J. Immunol. 198, 557–563 (2017).
Stenius, F. et al. Lifestyle factors and sensitization in children—the ALADDIN birth cohort. Allergy 66, 1330–1338 (2011).
Jelding-Dannemand, E., Malby Schoos, A. M. & Bisgaard, H. Breast-feeding does not protect against allergic sensitization in early childhood and allergy-associated disease at age 7 years. J. Allergy Clin. Immunol. 136, 1302–1308 (2015).
Strachan, D. P. Hay-fever, hygiene, and household size. Brit. Med. J. 299, 1259–1260 (1989).
Ownby, D. R., Johnson, C. C. & Peterson, E. L. Exposure to dogs and cats in the first year of life and risk of allergic sensitization at 6 to 7 years of age. JAMA 288, 963–972 (2002).
Depner, M. et al. Maturation of the gut microbiome during the first year of life contributes to the protective farm effect on childhood asthma. Nat. Med. 26, 1766–1775 (2020).
Stein, M. M. et al. Innate immunity and asthma risk in Amish and Hutterite farm children. New Engl. J. Med. 375, 411–421 (2016).
Eggesbø, M., Botten, G., Stigum, H., Nafstad, P. & Magnus, P. Is delivery by cesarean section a risk factor for food allergy?. J. Allergy Clin. Immuno. 112, 420–426 (2003).
Miettinen, R., Hermansson, H., Merikukka, M., Gissler, M. & Isolauri, E. Mode of delivery—impact on risk of noncommunicable diseases. J. Allergy Clin. Immunol. 136, 1398–1399 (2015).
Shao, Y. et al. Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth. Nature 574, 117–121 (2019).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 852 (2015).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Galazzo, G. et al. Development of the microbiota and associations with birth mode, diet, and atopic disorders in a longitudinal analysis of stool samples, collected from infancy through early childhood. Gastroenterology 158, 1584–1596 (2020).
Stokholm, J. et al. Delivery mode and gut microbial changes correlate with an increased risk of childhood asthma. Sci. Transl. Med. 12, eaax9929 (2020).
van Nimwegen, F. A. et al. Mode and place of delivery, gastrointestinal microbiota, and their influence on asthma and atopy. J. Allergy Clin. Immunol. 128, 948–955 (2011).
Henrick, B. M. et al. Bifidobacteria-mediated immune system imprinting early in life. Cell 184, 3884–3898.e11 (2021).
Feehley, T. et al. Healthy infants harbor intestinal bacteria that protect against food allergy. Nat. Med. 25, 448–453 (2019).
Chatzigiannidou, I. et al. Temporal dynamics and microbial interactions shaping the gut resistome in early infancy. Nat. Commun. 16, 8139 (2025).
Laursen, M. F. et al. Bifidobacterium species associated with breastfeeding produce aromatic lactic acids in the infant gut. Nat. Microbiol. https://doi.org/10.1038/s41564-021-00970-4 (2021).
Meng, D. et al. Indole-3-lactic acid, a metabolite of tryptophan, secreted by Bifidobacterium longum subspecies infantis is anti-inflammatory in the immature intestine. Pediatr. Res. 88, 209–217 (2020).
Ehrlich, A. M. et al. Indole-3-lactic acid associated with Bifidobacterium-dominated microbiota significantly decreases inflammation in intestinal epithelial cells. BMC Microbiol. 20, 357 (2020).
Holmes, I., Harris, K. & Quince, C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS ONE 7, e30126 (2012).
Roßberg, S. et al. Orally applied bacterial lysate in infants at risk for atopy does not prevent atopic dermatitis, allergic rhinitis, asthma or allergic sensitization at school age: follow-up of a randomized trial. Allergy 75, 2020–2025 (2020).
Melén, E. et al. Male sex is strongly associated with IgE-sensitisation to airborne but not food allergens: results up to age 24 years from the BAMSE birth cohort. Clin. Translat. Allergy 10, 15 (2020).
Vuillermin, P. J. et al. Maternal carriage of Prevotella during pregnancy associates with protection against food allergy in the offspring. Nat. Commun. 11, 1452 (2020).
Douwes, J. et al. Farm exposure in utero may protect against asthma, hay fever and eczema. Eur. Respir. J. 32, 603–611 (2008).
Uekert, S. J. et al. Sex-related differences in immune development and the expression of atopy in early childhood. J. Allergy Clin. Immunol. 118, 1375–1381 (2006).
Sakanaka, M. et al. Varied pathways of infant gut-associated bifidobacterium to assimilate human milk oligosaccharides: prevalence of the gene set and its correlation with bifidobacteria-rich microbiota formation. Nutrients https://doi.org/10.3390/nu12010071 (2019).
Dubois, L. et al. Paternal and induced gut-microbiota seeding complement maternal transmission in shaping neonatal gut colonisation. Cell Host Microbe https://doi.org/10.1016/j.chom.2024.05.004 (2024).
McGuire, M. K. et al. What’s normal? Oligosaccharide concentrations and profiles in milk produced by healthy women vary geographically. Am. J. Clin. Nutr. 105, 1086–1100 (2017).
Palframan, R. J., Gibson, G. R. & Rastall, R. A. Carbohydrate preferences of Bifidobacterium species isolated from the human gut. Curr. Iss. Intest. Microbiol. 4, 71–75 (2003).
DunnGalvin, A. et al. Highly accurate prediction of food challenge outcome using routinely available clinical data. J. Allergy Clin. Immunol. 127, 633–639 (2011).
Hickman, B. et al. Gut microbiota wellbeing index predicts overall health in a cohort of 1000 infants. Nat. Commun. 15, 8323 (2024).
Christensen, E. M. et al. The developing airway and gut microbiota in early life is influenced by age of older siblings. Microbiome 10, 106 (2022).
Koplin, J. J. et al. Environmental and demographic risk factors for egg allergy in a population-based study of infants. Allergy 67, 1415–1422 (2012).
Elizur, A., Rachel-Jossefi, S., Rachmiel, M., Eisenberg, E. & Katz, Y. Consumption of cow’s milk formula in the nursery and the development of milk allergy. Clin. Transl. Allergy 14, e12352 (2024).
Peters, A. et al. Metabolites of lactic acid bacteria present in fermented foods are highly potent agonists of human hydroxycarboxylic acid receptor 3. PLoS Genet. 15, e1008145 (2019).
Uhlén, M. et al. A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science 366, eaax9198 (2019).
Gutiérrez-Vázquez, C. & Quintana, F. J. Regulation of the immune response by the aryl hydrocarbon receptor. Immunity 48, 19–33 (2018).
Taft, D. H. et al. Bifidobacterial dominance in the early infant gut microbiome is associated with improved vaccine responses and reduced intestinal inflammation. Microbiome 6, 157 (2018).
Casaburi, G. et al. Metagenomic insights of the infant microbiome community structure and function across multiple sites in the United States. Sci. Rep. 11, 1472 (2021).
Lewis, Z. T. et al. Maternal fucosyltransferase 2 status affects the gut bifidobacterial communities of breastfed infants. Microbiome 3, 13 (2015).
Huda, M. N. et al. Stool microbiota and vaccine responses of infants. Pediatrics 134, e362–e372 (2014).
Vogel, K. et al. Bifidobacteria shape antimicrobial T-helper cell responses during infancy and adulthood. Nat. Commun. 14, 5943 (2023).
Hanifin, J. M. & Rajka, G. Diagnostic features of atopic dermatitis. Acta Dermato-Venereol. 60, 44–47 (1980).
Severity scoring of atopic dermatitis: the SCORAD index. Consensus Report of the European Task Force on Atopic Dermatitis. Dermatology https://doi.org/10.1159/000247298 (1993).
Wettermark, B. et al. The new Swedish Prescribed Drug Register–opportunities for pharmacoepidemiological research and experience from the first six months. Pharmacoepidemiol. Drug Saf. 16, 726–735 (2007).
Ballardini, N., Nilsson, C., Nilsson, M. & Lilja, G. ImmunoCAP Phadiatop Infant–a new blood test for detecting IgE sensitisation in children at 2 years of age. Allergy 61, 337–343 (2006).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Lau, S. et al. Oral application of bacterial lysate in infancy decreases the risk of atopic dermatitis in children with 1 atopic parent in a randomized, placebo-controlled trial. J. Allergy Clin. Immunol. 129, 1040–1047 (2012).
Kechin, A., Boyarskikh, U., Kel, A. & Filipenko, M. cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J. Comput. Biol. 24, 1138–1143 (2017).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Callahan, B. J., McMurdie, P. J. & Holmes, S. P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 11, 2639–2643 (2017).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Glöckner, F. O. et al. 25 years of serving the community with ribosomal RNA gene reference databases and tools. J. Biotechnol. 261, 169–176 (2017).
Eriksen, C. et al. IgG and IgM cooperate in coating of intestinal bacteria in IgA deficiency. Nat. Commun. 14, 8124 (2023).
Tingley, D., Yamamoto, T., Hirose, K., Keele, L. & Imai, K. mediation: R package for causal mediation analysis. J. Stat. Softw. 59, 1–38 (2014).
Myers, P. Gene catalogue and MAGs from the ALADDIN birth cohort. Zenodo https://doi.org/10.5281/zenodo.17810079 (2025).
Acknowledgements
We acknowledge the families participating in the ALADDIN study for their trust and contribution and the ALADDIN team for their involvement in this work, especially nurse and coordinator M. Eriksson, medical doctor F. Stenius, biomedical analyst C. Wallén and researcher C. Johansson. We also acknowledge the families participating in the PAPS study. We thank the families participating in BIS and acknowledge the contribution of the fieldwork team and the BIS Investigator Group: A.-L. Ponsonby, D. Burgner, J. Carlin, S. Ranganthan, R. Saffery, M. Tang, F. Collier, L. Gray, A. Loughman, and T. Mansel. We also thank L. B. Rosholm at DTU Bioengineering for technical support. We thank the following funding bodies for support to ALADDIN: the Swedish Research Council (2012-3011), the Swedish Research Council for Working Life and Social Research (2006-1630), the regional agreement on medical training and clinical research (ALF) between Stockholm County Council and the Karolinska Institutet and the Karolinska University Hospital, the Swedish Asthma and Allergy Research Association, the Swedish Cancer and Allergy Fund, the Ekhaga Foundation, the Frimurare Barnhuset Foundation in Stockholm and the Hesselman Foundation, all to J.A. Thermo Fisher Scientific, Uppsala, Sweden, provided the study with reagents. Support to PAPS by the German Research Foundation (grant no. DFG Ha 2162/4–1), Symbiopharm Herborn, German Ministry of Nutrition and Agriculture and Joint Programming Initiative A healthy diet for a healthy life (JPI HDHL) Joint Action Intestinal Microbiomics (50–52905–98–599), German Research Foundation (DFG) LA 2473/2-1 via the Clinical Research Group (KFO 339 Food Allergy and Tolerance Food@ LA 2473/2-1) and the Bundesanstalt für Landwirtschaft und Ernährung Förderkennzeichen 2815ERA05E (all S.L.). Support by The Independent Research Fund Denmark (grant no. 0171-00006B to H.R.M.), Innovation Fund Denmark (grant no. 4203-00005B to S.B.) and FII institute for the FII institute Research Chair at DTU in Immune-based Prediction of Disease from 2021 to 2023 to S.B.
Author information
Authors and Affiliations
Contributions
P.N.M. and R.K.D. were responsible for bioinformatics and data analysis and for drafting the manuscript together with S.B. A.S. and J.A. were responsible for collection of the ALADDIN samples, S.L. and E.H. for collection of the PAPS samples and P.V. for collection of the BIS samples. A.M. analysed the HMOs in mother’s milk, and M.A.H. provided expert help in its interpretation. J.M.M., I.C., P.L.J. and C.E. supported the data analysis. N.V.B, M.O’H., P.V. and J.P. were responsible for 16S rRNA gene amplicon sequencing data and analysis. R.K.D. prepared samples for faecal water metabolomics and implemented the Ig ELISA analysis. M.P. ran metabolomics samples. H.M.R. was responsible for metabolomics analysis of aromatic lactates. M.F.L., D.A., M.I.B. and T.R.L. were responsible for qPCR analysis. N.L.N. and K.K. were responsible for shotgun metagenomic sequencing of ALADDIN faecal samples. H.B.N. supervised the bioinformatics analysis. A.H.T., R.K.D., E.M.S. and S.B. implemented and performed cellular IgE assays. A.S., J.A., K.K. and S.B. were responsible for the study concept and design. All authors were responsible for providing comments to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.S. was a member in the Joint Steering Committee for the Human Translational Microbiome Program (CTMR) at Karolinska Institutet together with Ferring Pharmaceuticals, Switzerland (2016–2023). R.K.D., P.N.M., C.E., A.H.T. and S.B. have a patent (patent no. WO2025158027-A1, 2025, International/PCT (WIPO)) covering findings in Fig. 3 and Extended Data Fig. 3. The other authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Mahesh Desai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Abundance of bifidobacteria in the first 5 years of life.
Relative abundance of B. bifidum, B. breve, B. infantis, B. longum, B. dentium, B. adolescentis, B. catenulatum, and B. pseudocatenulatum at eight time points from 0–60 months in each of 56 children with longitudinal sampling from the ALADDIN birth cohort.
Extended Data Fig. 2 Early life abundance of aldh+ bifidobacteria associate with later reduced food allergen–specific IgE in ALADDIN and in the replication cohort PAPS when using 4 time points for faecal abundance annotation between 0 and 6 months and V3 16S rRNA amplicon sequencing but not with reduced food allergen-specific skin prick test in the replication cohort BIS when based on abundance at 2 time points (1 and 6 months) and V4 16S rRNA amplicon sequencing.
a, Maximum relative abundance of aldh+ bifidobacteria across 4 time points (3–6 d, 3w, 2 m, 6 m) in children with or without food allergen–specific IgE (cutoff: 0.35 kUA/L) at 6, until 12, and until 24 months of age in the ALADDIN cohort. b, Maximum relative abundances of aldh+ and aldh− bifidobacteria across 4 time points (w5, 13, 21, 31) in children with or without food allergen–specific IgE (cutoff: 0.35 kUA/L) until 36 months of age in the PAPS cohort. Bacteria were identified using amplicon sequencing of the V3 region of the 16S rRNA gene, and aldh+ and aldh− bifidobacteria were distinguished using two positions that discriminate them in the V3 region. c, Spearman’s rank correlation coefficients (SCC) between the maximum relative abundances of aldh+ bifidobacteria from 0–6 months of life and the maximum concentrations of food allergen–specific IgE in children with food allergen–specific IgE ≥ 0.1 kUA/L at 6 months (n = 43), until 12 months (n = 69), until 24 months (n = 92), and until 60 months (n = 118) in the ALADDIN cohort. The line represents the linear regression line; grey area represents 95% CI around the regression estimate. d, Maximum relative abundances of aldh+ and aldh− bifidobacteria in the BIS cohort between 1 and 6 months of age versus a positive food allergen skin prick test at 12 months of age. Bacteria were identified using amplicon sequencing of the V4 region of the 16S rRNA gene. There is only one discriminative position in the V4 region of the 16S rRNA gene that can be used to distinguish aldh+ from aldh− bifidobacteria, which makes their identification less valid as compared to V3 region identification. a–c, Statistics based on two-sided Wilcoxon rank-sum test. For box plots, the horizontal lines indicate the median; box boundaries indicate the interquartile range; whiskers represent values within 1.5× the interquartile range of the first and third quartiles.
Extended Data Fig. 3 Faecal levels of ILA and PLA do not mediate the association between aldh+ bifidobacteria and food allergen–specific IgE until 5 years of age and do not link to later diminished allergic disease in children.
a, Faecal levels of ILA and PLA in 2 months old children with or without food allergen–specific IgE (cutoff ≥ 0.35 kUA/L) until 5 years of age. b, Forest plot displaying a causal mediation analysis quantifying the contribution of 2 months faecal PLA levels to the association of max 0–6 m aldh+ faecal relative abundance with food allergen–specific IgE until 5 years of age. Points represent the effect estimate and horizontal lines the 95% CI around each effect estimate. Significance was tested by bootstrapping with 1,000 resamples. The estimate corresponds to the change in log odds of food allergen–specific IgE until 5 years of age per one-unit increase. c, Total effect and mediation model, including tests of significant associations preceding the mediation analysis. Association between max 0–6 m faecal aldh+ relative abundance and food allergen–specific IgE until 5 years of age (path c) and 2 m faecal 4-OH-PLA or PLA levels and food allergen–specific IgE (path b) were tested with a generalized linear model. The 2 m faecal 4-OH-PLA or PLA levels were predicted by max 0–6 m faecal aldh+ relative abundance (path a) using a linear model. Path c´ corresponds to the direct effect in the mediation analysis. d, Faecal levels of ILA and PLA in 2 months old children that are diagnosed with atopic dermatitis, as defined by SCORAD, at 2 years of age (yes/no). (a, d) Numbers below box plots represent the number of samples with yes or no for the given indication. Horizontal lines indicate the median; box boundaries indicate the interquartile range; whiskers represent values within 1.5× the interquartile range of the first and third quartiles; points outside whiskers show outliers. Statistics based on two-sided Welch t-test.
Extended Data Fig. 4 Correlations between individual Bifidobacterium species and faecal levels of aromatic lactates at 2 or 6 months of age.
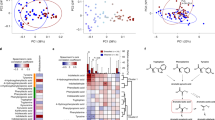
Spearman’s rank correlation coefficients (SCC) between faecal levels of aromatic lactates at 2 or 6 months of age and individual aldh+ and aldh− bifidobacteria when detected (n = 141 at 2 months, n = 54 at 6 months). The Holm-Bonferroni method was used to adjust for multiple testing at each time point, with asterisks indicating *Padj < 0.05, **Padj < 0.01, ***Padj < 0.001.
Extended Data Fig. 5 Addition of IL-4 to CD40L-primed blood-derived immune cells preferentially increases IgE production, and addition of physiological doses of 4-OH-PLA does not alter cellular viability.
a, b, e, IgE and IgG1-4 production in human blood-derived immune cells stimulated ex vivo for 10 days with 50 ng/mL HA-tagged CD40L, 100 ng/mL cross-linking anti-HA antibody, and 50 ng/mL rhIL-4 (activation stimuli) to induce IgE production. a, Fold change in concentrations of IgE and IgG1-4 at day 10 upon addition of activation stimuli at day 0, 4, and 7 as compared to cells provided with the CD40L-stimuli without IL-4. b, IgG1-4 production after 10 days stimulation with vehicle with/without 5 or 50 μM 4-OH-PLA as compared to vehicle only. Data is displayed as mean ± SEM of replicates from 6 healthy individuals. c, Gating strategy for determining the percentage of Day 10 viable blood-derived immune cells using Live/Dead Fixable Violet stain. d, Day 10 viability of blood-derived immune cells from three donors subjected to activation stimuli in a suspension with or without 50 μM 4-OH-PLA. e, IgE and IgG1-4 production after 10 days stimulation with vehicle with/without 5 or 50 μM ILA as compared to vehicle only. Data is displayed as mean ± SEM of replicates from 6 healthy donors. Statistics: b, e Two-sided, paired t-test, with P ≥ 0.05 considered non-significant, and e, P (5 µM) = 0.0044 and P (50 µM) = 0.00067.
Extended Data Fig. 6 Influence of early-life environmental factors on the abundance of aldh+ and aldh− bifidobacteria from 0–6 months of age in ALADDIN.
Relative abundances of aldh+ bifidobacteria and aldh− bifidobacteria at 3–6 days, 3 weeks, 2 months, and 6 months in vaginally delivered children, a, from anthroposophic vs. non-anthroposophic families, b, girls vs. boys, or c, with or without child or maternal pets/farm animal exposure in utero and until 2 months of age. d, Prevalence of summed aldh+ bifidobacteria with relative abundance above 2% in children at 12 months (n = 56), 18 months (n = 55), 24 months (n = 56), and 60 months (n = 55), and in mothers during third trimester of pregnancy (M-preg 3tr) and at 2 months postpartum (M-2m pp, both n = 56). For box plots, the horizontal lines indicate the median; box boundaries indicate the interquartile range; whiskers represent values within 1.5× the interquartile range of the first and third quartiles; dots represent individual data points. Statistics: Two-sided Wilcoxon rank-sum test. FDR cutoffs (Q): not significant (ns) ≥ 0.05.
Extended Data Fig. 7 Influence of early-life environmental factors on the abundance of aldh+ and aldh− bifidobacteria from 1–6 months of age in children enrolled in the PAPS cohort.
Relative abundances of aldh+ bifidobacteria and aldh− bifidobacteria at 5, 13, 21, and 31 weeks in vaginally vs. caesarean section–delivered children (a), and in vaginally delivered children with or without older siblings (b). Relative abundance of aldh+ bifidobacteria at 5, 13, 21, and 31 weeks in vaginally delivered breastfed children with or without formula supplementation before 2 months of age (c), and in caesarean section–delivered children with or without formula supplementation before 2 months of age (d). For box plots, the horizontal line indicates the median; box boundaries indicate the interquartile range; whiskers represent values within 1.5× the interquartile range of the first and third quartiles; dots represent individual data points. Statistics: Two-sided Wilcoxon rank-sum test. FDR cutoffs (Q): ** < 0.01, *** < 0.001, and not significant (ns) ≥ 0.05.
Extended Data Fig. 8 Influence of early-life environmental factors on the faecal levels of the aromatic lactate 4-OH-PLA in children enrolled in ALADDIN.
Levels of 4-OH-PLA at 2 and 6 months of age in vaginally and caesarean section-delivered children (a), in vaginally delivered children with or without older siblings (b), and in vaginally delivered breastfed children with or without formula supplementation before 2 months of age (c). For box plots, the horizontal line indicates the median; box boundaries indicate the interquartile range; whiskers represent values within 1.5× the interquartile range of the first and third quartiles; points outside whiskers show outliers. Statistics based on two-sided Welch t-test.
Extended Data Fig. 9 Detection of maternal carriage of B. infantis and B. breve using qPCR.
a, Data points showing detection of B. infantis and B. breve in mothers prior to delivery and at 2 months postpartum by qPCR. Detection limit was 300 gene copies per gram faeces. b, Odds ratio of co-presence of B. infantis or B. breve in mothers or children. Pre-delivery / 3–6 days corresponds to the mother sampled prior to delivery (n = 56) and child sampled at 3–6 days after birth (only vaginally delivered children included, n = 43). 2 months / 0–2 months corresponds to the mother sampled at 2 months postpartum (n = 56) and child within 0–2 months (n = 56). Since B. infantis was not detected in mothers prior to delivery, only the Odds ratio for co-presence at 2 months is shown. Points indicate odds ratio estimate; lines indicate 95% CI around each odds ratio estimate.
Extended Data Fig. 10 Maternal secretion and prevalence of Bifidobacterium spp. in infants.
Prevalence of Bifidobacterium spp. in breastfed children with a secretor (n = 47) or non-secretor mother (n = 23) was compared using samples from 3–6 days (n = 29/17), 3 weeks (n = 28/19), 2 months (n = 46/23), and 6 months (n = 44/22) from children with secretor/non-secretor mothers, respectively. a, Prevalence of individual Bifidobacterium species in breastfed children of secretor versus non-secretor mothers. Odds ratio estimates (log2) with 95% CI around each odds ratio estimate are shown. Points are coloured according to the direction of association: pink indicates higher prevalence in children with secretor mothers, green indicates higher prevalence in children with non-secretor mother, and grey indicates non-significant co-associations (P ≥ 0.05). A relative abundance of 1% was used as cutoff. b, Heatmap of significant co-occurrence of Bifidobacterium species pairs in breastfed children of secretor versus non-secretor mothers. Each cell represents a species pair; colours indicate the odds ratio estimate (log2) for co-existence: positive values mean that the pair is more likely to co-exist in children with secretor mothers, while negative values mean that co-existence is more likely in children with non-secretor mothers. Only pairs with significant associations (P < 0.05) are shown. Species pairs were considered co-existing when both species had a relative abundance > 1% in a sample. Statistics: Two-sided Fisher’s exact test for count data.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–8, Methods and References.
Supplementary Table 1 (download XLSX )
Taxa identification.
Source data
Source Data Figs. 1, 3 and 4 (download XLSX )
Source data for Figs. 1a–c,f, 3c and 4d.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Myers, P.N., Dehli, R.K., Mie, A. et al. Early-life colonization by aromatic-lactate-producing bifidobacteria lowers the risk of allergic sensitization. Nat Microbiol 11, 429–441 (2026). https://doi.org/10.1038/s41564-025-02244-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02244-9