Abstract
Animal–microorganism symbioses are omnipresent, with both partners often gaining benefits as mutualists. A single mutation in the carbon catabolite repression system in Escherichia coli enables mutualism with the stinkbug Plautia stali. Here we find that this mutation is not present in natural symbioses. Given that the carbon catabolite repression pathway affects the expression of >500 downstream genes, we investigated their role in mutualisms. We find that disruption of a single gene, tnaA, encoding tryptophanase makes E. coli mutualistic to P. stali, resulting in the accumulation of tryptophan and the reduction of toxic indole. A survey of wild populations of P. stali and other stinkbug species revealed that their typical microbial symbionts, Pantoea, consistently lack the tnaA gene. Some Pantoea species such as Pantoea ananatis retain the tnaA gene and cannot establish symbiosis with P. stali, but tnaA-disrupted P. ananatis partially restored the symbiotic capability. When a natural Pantoea mutualist of P. stali was transformed with a functional tna operon, its symbiotic capability reduced significantly. Our finding suggests that tryptophanase disruption may have facilitated the evolution of gut bacterial mutualists in insects.
Similar content being viewed by others
Main
Microbial symbioses prevail in biological kingdoms, in which diverse relationships encompass parasitism, commensalism and mutualism1,2. Among them, the most intimate symbioses are found among mutualistic ones, in which the host and the symbiont constitute an integrated biological entity and suffer disadvantages without the partnership3,4. Originally, such microbial symbionts must have had no relationship with their host organisms, plausibly having existed as environmental microorganisms. It is of fundamental evolutionary interest how ordinary free-living microorganisms have become indispensable mutualists. How many and what mutations are required for the evolution of mutualism? How quickly does the evolution of mutualism proceed? To address these questions, experimental evolutionary approaches may provide valuable insights5,6,7,8,9,10,11,12.
Recently, an experimental evolutionary system consisting of an insect, Plautia stali, as host and a bacterium, Escherichia coli, as symbiont was established, which brought about unprecedented insight into an early stage of the evolution of mutualistic symbiosis13,14. The stinkbug P. stali possesses a midgut symbiotic organ full of a specific bacterial symbiont of the genus Pantoea, which is essential for growth and survival of the host insect15,16,17,18. The famous model bacterium E. coli is a component of the mammalian gut microbiome and has no previous relationship with the insect19,20. However, when symbiont-deprived newborn nymphs of P. stali were experimentally inoculated and maintained with a hypermutating E. coli strain, multiple evolutionary lines showed a significantly improved adult emergence rate and body colour within several months to a year, indicating rapid and recurrent evolution of ‘mutualistic’ E. coli13. Analysis of the independently evolved mutualistic E. coli lines revealed that single loss-of-function mutations on cyaA and crp genes that convergently disrupt the carbon catabolite repression (CCR) global transcriptional regulator system, which is involved in bacterial metabolic switching in response to nutritional stresses21,22, are responsible for the mutualistic host phenotypes. These results revealed that elaborate mutualistic symbiosis can evolve very easily and rapidly through a single gene mutation13.
The finding that only a single gene disruption is sufficient for making E. coli an insect mutualist is certainly striking, but it should be noted that disruption of the CCR pathway globally affects the expression levels of over 500 downstream genes encoded on the E. coli genome23. Hence, we expected that the causative genes directly responsible for the improved host phenotypes must be identified among the downstream E. coli genes under CCR regulation, although the nature of such genes was totally unknown. Here we report the identification of a bacterial enzyme gene under CCR regulation whose disruption underpins the evolution of insect–bacterium mutualism not only in the laboratory but also in nature.
Results
No CCR disruption in natural symbionts of P. stali
In Japanese populations of P. stali, six Pantoea-allied symbiotic bacteria, Pantoea spp. A, B, C, D, E and F (abbreviated as Sym A and Sym B for uncultivable ones, and Sym C, Sym D, Sym E and Sym F for cultivable ones), are present15. While all the symbiotic bacteria can support normal growth and reproduction of P. stali, genome sequencing revealed that their genomes consistently retain the intact CCR pathway genes cyaA and crp (Supplementary Table 1a). These observations strongly suggested that the CCR disruption, which was observed in the laboratory evolution of mutualism with hypermutating E. coli13, is not involved in the evolution of mutualistic symbionts in natural populations of P. stali.
Survey of downstream E. coli genes affected by CCR disruption
Considering that disruption of the CCR global transcriptional regulator system generally affects expression levels of hundreds of genes encoded on bacterial genomes23, it seemed plausible that some genes downstream of the CCR pathway may actually be responsible for the mutualistic phenotypes of the CCR-disruptive E. coli mutants. In this context, we focused on 58 CCR-regulated genes that were identified to be commonly down- or upregulated in two independent mutualistic E. coli evolutionary lines, CmL05 and GmL07, identified in our previous study13. These genes consisted of a diverse array of functional genes such as transporter genes for non-glucose sugars, carbohydrate metabolism genes, quorum sensing genes, extracellular matrix production genes, transcription factor genes and others (Extended Data Fig. 1).
Elevated tryptophan levels after the evolution of mutualism as well as CCR disruption in E. coli
To gain insight into metabolic aspects of the mutualistic evolutionary and mutant E. coli lines, we conducted comparative transcriptomic and metabolomic analyses of the E. coli-infected P. stali. A promising clue came from quantitative analysis of free amino acids in P. stali infected with the CCR-disruptive E. coli mutants ΔcyaA and Δcrp, in which a specific essential amino acid, tryptophan, showed over ten times higher levels in the haemolymph and symbiotic organ compared with the insects infected with the wild-type control E. coli strain ΔintS (Fig. 1a,b and Extended Data Fig. 2). Of the 58 candidate genes, 2 genes were related to tryptophan: tnaA encoding tryptophanase and tnaB encoding a component of tryptophan transporter (Fig. 1c,d and Extended Data Fig. 1). We obtained deletion mutants of these genes, ΔtnaA and ΔtnaB, and inoculated the E. coli mutants into P. stali. Tryptophan levels in the haemolymph and symbiotic organ were significantly elevated in the ΔtnaA-infected insects but not in the ΔtnaB-infected insects (Fig. 1a,b and Extended Data Fig. 2). The elevated tryptophan levels in the ΔtnaA-infected insects were comparable to (1) those in insects infected with the CCR-disruptive mutants ΔcyaA and Δcrp, (2) those in insects infected with the mutualistic evolved E. coli strain CmL05 (ref. 13) and (3) those in normal symbiotic insects with the natural Pantoea symbiont Sym A (Fig. 1a,b, and Extended Data Figs. 2 and 3a,b). A tryptophanase assay confirmed that not only ΔtnaA but also ΔcyaA, Δcrp and CmL05 lost the tryptophanase activity whereas ΔintS and ΔtnaB were tryptophanase positive (Extended Data Fig. 3c–e).
a,b, Effects of knockout mutants of E. coli, ΔcyaA, Δcrp, ΔtnaA and ΔtnaB, on tryptophan levels in haemolymph (a) and the symbiotic organ (b). c,d, Transcriptomic data on the downregulation of the tnaA gene (c) and tnaB gene (d) after the evolution of mutualism in the evolutionary E. coli lines CmL05 and GmL07 (Extended Data Fig. 1). e,f, Effects of knockout mutants of E. coli, ΔcyaA, Δcrp, ΔtnaA and ΔtnaB, on adult emergence rates (e) and body colour (f). Note that Sym A, the natural symbiont of P. stali15, comprises a mutualistic positive control, whereas ΔintS E. coli represents a non-beneficial negative control13. For the box plots, centre lines, limits and dots show medians, first and third quartiles, and data points, respectively. Different alphabetical letters (a, b, c and d) indicate statistically significant differences (pairwise Wilcoxon rank–sum test with Hommel’s correction: P < 0.05, two sided). Biological replicate numbers are indicated on the graphs. g, External appearance of the adult insects infected with Sym A, ΔcyaA, Δcrp, ΔtnaA, ΔtnaB and ΔintS obtained in the study. CPM, count per million; Trp, tryptophan.
Improved host phenotypes induced by tryptophanase disruption in E. coli
The ΔtnaA-infected insects showed significantly higher adult emergence rates and remarkably greenish body colour compared with the ΔintS-infected control insects, which were comparable to those of the ΔcyaA- and Δcrp-infected insects, and also comparable to those of the normal symbiotic insects with the natural Pantoea symbiont Sym A (Fig. 1e–g and Extended Data Fig. 4a,b). However, the ΔtnaA-infected insects showed no significant improvement in body size compared with the ΔintS-infected control insects, which was also the case for the ΔcyaA- and Δcrp-infected insects (Extended Data Fig. 4c,d). When a functional tnaA gene was introduced into the ΔtnaA E. coli strain (Supplementary Fig. 1a), inoculation of the recombinant ΔtnaA::tnaA E. coli strain resulted in reduced haemolymphal tryptophan and attenuation of the improved host performance (Extended Data Fig. 5). When the tnaA gene in the ΔcyaA E. coli strain was constitutively expressed by introduction of an ectopic promoter sequence (Supplementary Fig. 1b), inoculation of the recombinant ΔcyaA Pconst-tnaA E. coli strain also resulted in lower haemolymphal tryptophan and cancellation of the improved host performance (Extended Data Fig. 5). These results revealed that (1) disruption of the tryptophanase gene in E. coli, which is under CCR regulation, significantly improves the survival and body colour of infected P. stali, (2) the improved host performance due to infection with CCR-disruptive mutants, ΔcyaA and Δcrp, is attributable to downregulation of the CCR downstream target gene tnaA and (3) tryptophanase disruption is a pivotal mechanism that underpins the evolution of P. stali–E. coli mutualism.
Why host performance improves through tryptophanase disruption in E. coli
Extended Data Fig. 6 summarizes the results showing the molecular mechanisms involved in the laboratory evolution of P. stali–E. coli mutualism. Before the evolution of mutualism, bacterial CCR operates, tryptophanase is expressed, tryptophan is broken down and the host suffers poor performance (Extended Data Fig. 6a). After the evolution of mutualism, bacterial CCR is disrupted, tryptophanase is suppressed, tryptophan accumulates and the host shows good performance (Extended Data Fig. 6b). Why does tryptophanase disruption in symbiotic E. coli result in improved performance of the host P. stali? Considering that tryptophanase converts tryptophan into indole, pyruvate and ammonium24, we conceived two plausible hypotheses, which are not necessarily mutually exclusive. The first hypothesis is that tryptophanase disruption suppresses toxic indole production and thereby improves host fitness, on the grounds that perturbation of tryptophan metabolism mediated by gut microbiota towards the indole pathway tends to be linked to pathology and disease25. The second hypothesis is that tryptophanase disruption results in the accumulation of the essential amino acid tryptophan and thereby contributes to host fitness, given that symbiont-mediated provisioning of tryptophan is important for diverse plant-sucking insects26.
Effects of indole and tryptophan feeding
To test these hypotheses, we administered different concentrations of indole and tryptophan to P. stali nymphs infected with the tryptophanase-deficient ΔtnaA E. coli and those infected with the control ΔintS E. coli via drinking water. As indole doses were elevated, adult emergence rates declined in both the ΔtnaA-infected insects and the ΔintS-infected insects, with the level of decline less conspicuous in the ΔtnaA-infected insects than in the ΔintS-infected insects (Fig. 2a,b). These results favoured the notion that indole accumulation is detrimental to the growth and survival of P. stali. As tryptophan doses were elevated, adult emergence rates were not affected in the ΔtnaA-infected insects (Fig. 2c) but were suppressed in the ΔintS-infected insects (Fig. 2d). These results seemed unexpected at a glance considering that tryptophan is an essential amino acid. However, it should be noted that the laboratory insects were provided with highly nutritious food (raw peanuts); tryptophan feeding may thus lead to excessive tryptophan intake, and the tryptophanase-positive ΔintS E. coli may convert the excess tryptophan into toxic indole. Quantification of tryptophan and indole in haemolymph samples of these experimental insects revealed that (1) the ΔtnaA-infected insects showed little haemolymphal indole; (2) by contrast, the ΔintS-infected insects showed significantly higher levels of haemolymphal indole; (3) in the ΔintS-infected insects, indole and tryptophan feeding tended to result in elevated levels of haemolymphal indole; and (4) in the ΔintS-infected insects, tryptophan levels were consistently low (Fig. 2e,f). These results accounted for the observations that not only indole feeding but also tryptophan feeding resulted in negative fitness consequences preferentially in the ΔintS-infected insects (Fig. 2a–d). Metabolomic analysis confirmed that higher haemolymphal tryptophan levels and lower haemolymphal indole levels were observed in the insects infected with the CCR-deficient mutant ΔcyaA and the evolved E. coli strain CmL05G13, compared with the ΔintS-infected insects (Extended Data Fig. 7a–c). It was also shown that some tryptophan-derived metabolites, such as 5-hydroxytryptamine, kynurenine, 3-hydroxykynurenine, indole-3-acetic acid and indole-3-carboxylic acid, tended to show higher haemolymphal levels in the insects infected with the CCR-deficient mutant and evolutionary E. coli strains, ΔcyaA and CmL05G13, compared with the ΔintS-infected insects (Extended Data Fig. 7d–k).
a,b, Effects of indole feeding via drinking water on growth and survival of P. stali infected with tryptophanase-disrupted ΔtnaA E. coli (a) and control ΔintS E. coli (b). c,d, Effects of tryptophan feeding via drinking water on growth and survival of P. stali infected with tryptophanase-disrupted ΔtnaA E. coli (c) and control ΔintS E. coli (d). e,f, Effects of indole and tryptophan feeding via drinking water on haemolymphal indole levels (e) and tryptophan levels (f) of P. stali infected with tryptophanase-disrupted ΔtnaA E. coli and control ΔintS E. coli. For box plots, centre lines, limits and dots show medians, first and third quartiles, and data points, respectively. Different alphabetical letters (a, b, c and d) indicate statistically significant differences (pairwise Wilcoxon rank–sum test with Hommel’s correction: P < 0.05, two sided). Biological replicate numbers are indicated on the graphs.
Effects of tryptophan overproduction by E. coli
In addition to the feeding experiments, we examined the effects of upregulated tryptophan production by genetically manipulated E. coli. When a tryptophan-overproducing E. coli mutant, ΔtrpR, which is disruptive of the trp operon repressor trpR27, was inoculated into P. stali, the ΔtrpR-infected insects showed significantly improved adult emergence rates and body colour compared with the control ΔintS-infected insects, whereas the levels of improvement were not comparable to those of the ΔtnaA-infected insects (Extended Data Fig. 8a,b). Notably, despite the tryptophan overproduction by ΔtrpR E. coli, the ΔtrpR-infected insects did not show elevated tryptophan levels in haemolymph (Extended Data Fig. 8c,d). It seems plausible, although speculative, that the tryptophan production by ΔtrpR E. coli is at such a level that the host insects promptly use up the limited essential amino acid for their growth and development. These results corroborated the notion that E. coli-derived tryptophan contributes to the improvement of host fitness.
Absence of the tryptophanase gene in natural symbiotic bacteria of stinkbugs
Given that tryptophanase disruption makes E. coli mutualistic to P. stali in the laboratory, it is of interest whether natural symbiotic bacteria of stinkbugs retain the tnaA gene or not. First, we inspected six genomes of natural Pantoea-allied symbionts Sym A, B, C, D, E and F of P. stali, in which no tnaA gene was found (Fig. 3 and Supplementary Table 1a). Next, we determined seven genomes of Pantoea-allied Sym C isolated from seven additional Ryukyu Island populations of P. stali, from which no tnaA gene was detected (Fig. 3 and Supplementary Table 1a). Next, we determined five genomes of Pantoea-allied Sym C of other stinkbugs collected at Ryukyu Islands, namely, three local isolates from Axiagastus rosmarus, one isolate from Lampromicra miyakona and one isolate from Scutellera amethystina, all of which encoded no tnaA gene (Fig. 3 and Supplementary Table 1a). Finally, using an inoculation and screening procedure with symbiont-free newborn nymphs of P. stali15, we screened and isolated environmental bacteria capable of supporting growth of P. stali from soil samples collected at five Ryukyu Islands, namely six isolates from Ishigaki Island, two isolates from Okinawa Island, two isolates from Miyako Island, three isolates from Yonaguni Island and four isolates from Tokunoshima Island. All the environmental bacterial isolates potentially symbiotic to P. stali were phylogenetically placed in the genus Pantoea and devoid of the tnaA gene in their genomes (Fig. 3 and Supplementary Table 1a). Enzymatic assay confirmed that the natural symbiotic bacteria of P. stali as well as the environmental bacterial isolates potentially symbiotic to P. stali consistently lack tryptophanase activity (Extended Data Fig. 9a–d). Inoculation of these bacterial isolates into symbiont-free newborn nymphs of P. stali verified that they can support growth and survival of the host stinkbugs, with the stinkbug-derived isolates tending to induce better host performance than the soil-derived isolates (Extended Data Fig. 10). These results suggested that the lack of the tnaA gene may be related to the ability of Pantoea-allied bacteria to establish symbiosis with P. stali.
The maximum likelihood phylogeny is inferred from amino acid sequences of 106 concatenated essential single-core genes (35,647 aligned amino acid sites). Statistical support values for each clade are shown at the node in the order of maximum likelihood and Bayesian analyses. Collection localities are shown in the map of the mainland and Ryukyu Islands of Japan. The colours of the bacterial taxon labels and the squares on the map correspond to the symbiont categories depicted at the bottom left (Supplementary Table 1a). The presence or absence of the tnaA gene is shown beside the taxon labels.
Pantoea ananatis with the tryptophanase gene are incapable of symbiosis with P. stali
Of 105 Pantoea genomes retrieved from the GenBank database, 78 genomes lacked the tnaA gene while 27 genomes retained the tnaA gene, with the majority, 19 genomes, affiliated to P. ananatis (Supplementary Table 1b). We obtained 4 strains of P. ananatis from culture collections, which were all confirmed to be tnaA and tryptophanase positive (Extended Data Fig. 9a–e). When they were inoculated into symbiont-free newborn nymphs of P. stali, few adult insects emerged (Extended Data Fig. 9f), indicating that the tnaA-carrying P. ananatis strains are incapable of establishing symbiosis with P. stali.
Tryptophanase disruption improved the symbiotic performance of P. ananatis
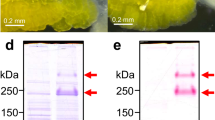
Is the inability of P. ananatis to establish symbiosis with P. stali relevant to the tryptophanase gene on the bacterial genome? We generated a knockout mutant of P. ananatis JCM6986 by homologous recombination targeting the tnaA gene (Fig. 4a). PCR detection confirmed deletion of the tnaA gene (Fig. 4b), and an enzymatic assay verified the loss of tryptophanase activity in the mutant (Fig. 4c). When the ΔtnaA mutant of P. ananatis was inoculated into symbiont-free newborn nymphs of P. stali, the nymphal survival and adult emergence rate significantly improved (Fig. 4d,e), although the adult emergence rate was only less than 10% on average (Fig. 4e). These results indicated that the non-symbiotic Pantoea strain becomes, although partially, mutualistic to P. stali by loss-of-function mutation of the tryptophanase gene.
a, Knockout scheme of the tnaA gene by homologous recombination. b, PCR check of homologous recombination. c, Enzymatic assay of tryptophanase disruption. d, Effects on the survival curve. Line points, limits and dots show means, standard deviations and data points, respectively. Statistical analysis was conducted on the 36th day data using Welch’s two-sample test (P = 3.73 × 10−4, two sided). e, Effects on the adult emergence rate. For box plots, centre lines, limits and dots show medians, first and third quartiles, and data points, respectively. Statistical analysis was conducted using a Wilcoxon rank–sum test (P = 0.00041, two sided). Biological replicate numbers are indicated on the graphs. FRT, flippase recognition target; NC, negative control; WT, wild-type P. ananatis.
Natural Pantoea symbiont reduced symbiotic performance when transformed with functional tryptophanase
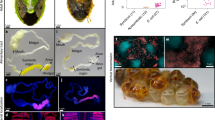
Finally, we artificially introduced a functional tna operon of P. ananatis into the genome of Sym F, a natural symbiont of P. stali that is cultivable and able to support host growth and survival15, by homologous recombination targeting the presumably non-functional transposon-related gene intB (Fig. 5a and Supplementary Fig. 2). The intB::tnaAB symbiont transformant showed significant tryptophanase activity (Fig. 5b), verifying that the introduced tna operon is functioning in the transformed symbiont strain. When symbiont-free newborn nymphs of P. stali were inoculated with the tryptophanase-producing intB::tnaAB symbiont, their adult emergence rate and body colour were negatively affected compared with those inoculated with the control ΔintB symbiont strain (Fig. 5c–f). In these insects, infection with the tryptophanase-producing intB::tnaAB symbiont resulted in lower tryptophan levels and higher indole levels than infection with the control ΔintB symbiont strain (Fig. 5g,h). These results indicated that the absence of the tryptophanase gene may contribute to the mutualistic properties of the natural symbiont of P. stali.
a, Introduction scheme of the tna operon by homologous recombination. b, Enzymatic assay of the expression and functioning of the introduced tna operon. c, Effects on the adult emergence rate. d, Effects on adult body colour. e, Effects on adult female body size. f, Effects on adult male body size. g, Effects on tryptophan levels in haemolymph. h, Effects on indole levels in haemolymph. For box plots, centre lines, limits and dots show medians, first and third quartiles, and data points, respectively. Statistical analysis was conducted by two-sided Wilcoxon rank–sum test (P values are shown on the graphs). Biological replicate numbers are indicated on the graphs.
Discussion
Our previous study showed that a single gene mutation, ΔcyaA or Δcrp, disrupting the bacterial CCR pathway makes E. coli mutualistic to P. stali13, which led to the notion that elaborate mutualistic symbiosis can evolve more easily and rapidly than conventionally envisioned. On account of the diverse phenotypic changes observed with the mutualistic E. coli strains and mutants13, we expected that multiple genes downstream of the CCR pathway may be involved in the evolution of P. stali–E. coli mutualism. Unexpectedly, however, we found that a single enzyme gene, tnaA, encoding tryptophanase, which is under CCR regulation, is the major effect gene whose disruption is sufficient for establishing P. stali–E. coli mutualism. This finding further corroborates the notion that elaborate mutualistic symbiosis can evolve easily and rapidly by a single gene mutation.
Plausibly, tryptophanase disruption contributes to host fitness via reduction of toxic indole and via accumulation of the potentially limited essential amino acid tryptophan. It should be noted that diverse stinkbugs rely on their gut symbiotic bacteria to provide essential amino acids and vitamins28,29,30,31, and the E. coli genome encodes the genes needed for synthesis of all these nutrients32. In plant-sucking aphids, the essential bacterial symbiont Buchnera encodes and amplifies synthetic genes for tryptophan on a plasmid33,34. Detailed physiological studies, for example those using a nutritionally defined artificial diet developed for aphids35,36, are needed for further understanding of the insect–bacterium nutritional interactions and interdependency.
Our genomic and functional investigations revealed that the loss of the tryptophanase gene is not only underpinning the laboratory evolution of P. stali–E. coli mutualism but, plausibly, also involved in the evolution of bacterial mutualists of the genus Pantoea that have recurrently occurred in natural populations of P. stali and other stinkbugs15,37,38,39. Of course, a variety of genetic changes of both partners must have contributed to the establishment and maintenance of the stinkbug–bacterium mutualistic symbioses in nature, and only a part of which may be attributable to the loss of the tnaA gene of the symbiont side. On account of the consistent absence of the tnaA gene among the diverse stinkbug symbiont genomes, which encompass cultivable ones with large genome sizes to uncultivable ones with reduced genome sizes (Supplementary Table 1a,c), we hypothesize that, although speculative, tryptophanase disruption may have facilitated the evolution of stinkbug–bacterium mutualism. Tryptophanase-deficient environmental Pantoea strains may have predisposed the establishment of symbiosis with stinkbugs. Alternatively, tryptophanase disruption may tend to occur at an early stage of the stinkbug–bacterium symbiosis in the course of symbiont genome degeneration. In either case, it seems plausible that tryptophanase disruption acts as a pivotal mutation of the symbiont side that facilitates, canalizes and stabilizes the relationship towards mutualism.
By contrast, while loss-of-function mutations of cyaA and crp, which disrupt CCR regulation, were identified as responsible for the evolution of P. stali–E. coli mutualism in the laboratory13, most of the Pantoea-allied natural symbiotic bacteria associated with P. stali and other stinkbugs, particularly those whose genomes are not so reduced, retain the cyaA and crp genes in their genomes (Supplementary Table 1a,c). These observations suggest that, in nature, disruption of the CCR pathway is generally not involved in the evolution of gut bacterial mutualists that are indispensable for the plant-sucking stinkbugs40,41,42. Considering that CCR regulation is important for bacterial adaptation to fluctuating environments by switching the main carbon source from a depleted one to an abundant one21,22, it is conceivable, although speculative, that CCR disruption can evolve under stable environments such as laboratory conditions, but it may be generally detrimental for bacteria that are thriving under fluctuating natural environments. These observations provide an important lesson that symbiotic evolution in the laboratory does not necessarily reflect symbiotic evolution in nature.
Among diverse life forms, the tnaA gene is found in various Gram-negative bacteria, whereas it is less common in Gram-positive bacteria, archaea and eukaryotes43. In particular, tnaA is most commonly detected in Gammaproteobacteria, to which E. coli, Pantoea spp. and many insect symbionts belong44. How widely tnaA disruption is relevant to the evolution of mutualism in diverse host–microorganism symbiotic associations is currently elusive and deserves future studies.
Using the model system for the experimental evolution of symbiosis between P. stali and E. coli, we have shown that even a single bacterial gene mutation can facilitate the evolution of mutualism. In the real world, however, the processes and mechanisms of the evolution of mutualism must be much more complex, entailing multiple genes and mutations of both the host and symbiont. Integrative approaches to the evolution of mutualism conducted in this study, in which experimental evolution in the laboratory and the natural diversity of symbiosis are jointly investigated, will lead to a deeper understanding of how elaborate mutualistic symbioses have been established and maintained.
Methods
Insect samples, bacterial strains and primers used in this study
Stinkbug samples and their symbiotic bacteria used in this study are listed in Supplementary Table 1a. Genome data of Pantoea isolates and stinkbug symbionts were retrieved from DNA databases (Supplementary Table 1b,c). P. ananatis isolates JCM6986, JCM14682 and JCM15056 were obtained from the Japan Collection of Microorganisms, while strain AJ13355 was provided by Ajinomoto. E. coli strains and mutants used in this study are listed in Supplementary Table 1d. The PCR primers used in this study are listed in Supplementary Table 1e.
Insect rearing, symbiont sterilization and bacterial inoculation
For most experiments, a laboratory strain of P. stali was used. The insects were reared in clean plastic or paper containers and fed with sterilized peanuts and sterilized water supplemented with 0.05% ascorbic acid in climate chambers at 25 ± 1 °C under a long day regime of 16 h light and 8 h dark as described45. To prepare symbiont-deprived newborn nymphs, collected egg masses were soaked in 4% formaldehyde for 20 min, kept twice in sterilized water for 10 min each, air-dried in a clean bench and placed in sterile plastic Petri dishes with cotton balls. The Petri dishes were kept in an incubator at 25 °C, where symbiont-free newborn nymphs emerged. Bacteria were cultured in liquid LB medium and diluted to OD600 = 0.1. The diluted culture medium (around 1.5 ml) was applied to cotton balls in each Petri dish, through which the symbiont-free newborn nymphs orally acquired the bacterial suspension. After sucking bacteria-containing water, the first instar nymphs moulted to second instar within 4–5 days without feeding, to which several pieces of sterilized peanuts and a 1.5-ml tube of sterilized water containing 0.05% ascorbic acid were introduced. Then, 3–4 days later, the mature second instar nymphs were transferred to a new rearing cage consisting of a paper container, a plastic lid with a large hole for ventilation, draining mesh for preventing insect escape, a 25-ml bottle of sterilized water containing 0.05% ascorbic acid and sterilized peanuts. This rearing cage system, which was renewed every week, was devised to stably maintain the insect colonies in good condition for an extended period. The emerged adult insects were sexed, counted, kept in a refrigerator overnight and image scanned from their dorsal side using a scanner (EPSON GT-X980) 6 weeks after egg collection. On the basis of the scanned images, the body colour and body size of the insects were measured using the image analysing software Natsumushi v.1.10 (ref. 46).
Analysis of amino acids, tryptophan-derived metabolites and indole
Each haemolymph sample was collected using a glass capillary (1 µl, Drummond) from the neck of an ice-anaesthetized adult insect, suspended in 100 µl of 80% (v/v) methanol and stored at –80 °C until use. Each symbiotic midgut sample dissected from an adult insect was homogenized in 100 µl of 80% methanol and stored under the same conditions. After homoarginine, homophenylalanine, [15N]-tryptophan and 6-hydroxyindole were added as internal standards, each sample was centrifuged, and an aliquot of the supernatant was subjected to liquid chromatography and mass spectrometry analysis of amino acids, tryptophan-derived metabolites and indole. The detection and quantification of these metabolites were performed using a liquid chromatography and mass spectrometry system (Waters, ACQUITY UPLC H-class and Xevo G2-XS qTOF) with an electrospray ionization source. Amino acid composition was measured after propyl-chloroformate derivatization as previously described18,31. For measurement of tryptophan and related compounds, the sample aliquots were concentrated under N2 flow and resuspended in 0.05% (v/v) formic acid. Then, they were separated on a column (Waters, BEH C18, 1.7 µm, 2 mm × 100 mm) with a gradient elution of 0.05% formic acid and methanol. Each compound was selectively measured at a positive multiple reaction monitoring mode. As indole is not efficiently ionized under electrospray ionization, we derivatized it with p-dimethylaminocinnamaldehyde as described47. The derivatized indole was separated on the same analytical column and quantified at a positive multiple reaction monitoring mode.
Feeding experiments with tryptophan and indole
L-tryptophan (FUJIFILM Wako Pure Chemical Corporation) and indole (Tokyo Chemical Industry) were dissolved and serially diluted in sterilized water containing 0.05% ascorbic acid. For tryptophan, concentrations of 1.14 × 100, 10−1, 10−2 and 10−3 mg ml−1 were prepared. For indole, concentrations of 0.5 × 100, 10−1, 10−2 and 10−3 mg ml−1 were prepared. The experimental insects were reared with sterilized peanuts and supplemented water as described, but the supplemented water was renewed every 3 days or 4 days to minimize the deterioration of the supplemented reagents. The emerged adult insects were counted and subjected to measurements of haemolymphal tryptophan and indole levels 6 weeks after egg collection.
Tryptophanase activity assay
A qualitative assessment of tryptophanase activity was conducted essentially as described48. Each bacterial strain was cultured in 3 ml of LB or M9-based liquid medium at 25 °C with shaking at 180 rpm for 24 h or 48 h. Then, 100 μl of Kovács indole reagent (Sigma-Aldrich) was added to the bacterial culture, and a reddish colour indicated the presence of indole (Extended Data Fig. 3c). A quantitative assessment of tryptophanase activity during bacterial growth was conducted essentially as described49. Each bacterial strain was cultured in LB or M9 liquid medium at 25 °C with shaking at 180 rpm overnight, diluted with LB liquid medium to OD600 = 0.1 and dispensed to 24 test tubes as 1-ml aliquots. The test tubes were incubated at 25 °C with shaking at 180 rpm, from which 3 samples were taken every hour and subjected to measurement of OD600. Then, the samples were centrifuged at room temperature at 10,000 rpm for 3 min, and the supernatants were subjected to indole quantification using an Indole Assay Kit (MAK326, Sigma-Aldrich).
Complementation of the tnaA gene in the ΔtnaA E. coli strain
The kanamycin resistance gene (KmR) was deleted from the E. coli tnaA::KmR strain using flippase (FLP) recombinase-mediated excision (Supplementary Table 1d). To restore tnaA function, a tnaA–Tn5 fusion construct was generated. The tnaA coding region was amplified by PCR using the primers BW25113_tnaA_rescue_F and BW25113_tnaA_rescue_R, while the KmR gene was amplified using the primers BW25113_tnaA_rescue_nptII_F and BW25113_tnaA_rescue_nptII_R (Supplementary Table 1e). The assembled tnaA–KmR cassette was re-amplified using the primers BW25113_tnaA_rescue_F and BW25113_tnaA_rescue_nptII_R (Supplementary Table 1e), and introduced into the BW25113 ΔtnaA strain via electroporation using λ Red recombination. The resulting strain was designated as BW25113 ΔtnaA::tnaA (Supplementary Table 1d). Successful integration of the construct was confirmed by PCR using the primers BW25113_tnaA-nptII_check_F and BW25113_tnaA-nptII_check_R (Supplementary Table 1e).
Construction of the ΔcyaA P_const–tnaA E. coli strain
To decouple tnaA expression from cAMP-CRP regulation, the native promoter and 5′ untranslated region of the tna operon were replaced by a synthetic constitutive promoter J23119. This promoter lacks the CRP-binding site, allowing transcription independent of intracellular cAMP levels and glucose concentration (Supplementary Fig. 1b). The kanamycin resistance gene was deleted from the cyaA::KmR E. coli strain using FLP recombinase-mediated excision (Supplementary Table 1d). The kanamycin resistance cassette containing the J23119 promoter was provided by N. Obana (University of Tsukuba) and amplified by PCR using the primers Pconst-tnaA_nptII-J23119_F and Pconst-tnaA_nptII-J23119_R (Supplementary Table 1e). The assembled fragment was introduced into the BW25113 ΔcyaA E. coli strain via electroporation using λ Red recombination. The final strain, BW25113 ΔcyaA Pconst-tnaA, was verified by PCR using the primers Pconst-tnaA_check_F and Pconst-tnaA_check_R (Supplementary Table 1e).
Knockout of the tnaA gene of P. ananatis
The presence of the tnaA gene in P. ananatis isolates was confirmed by PCR using the specific primers PA_tnaA_125F and PA_tnaA_1357R (Supplementary Table 1e). To knock out the tnaA gene in P. ananatis strain JCM6986, the primers PA_JCM6986_ΔtnaA_F and PA_JCM6986_ΔtnaA_R (Supplementary Table 1e) were used to amplify by PCR a kanamycin resistance gene (KmR) region from the E. coli ∆intS mutant. The PCR product was purified using a QIAquick PCR Purification Kit (Qiagen). Transformation of P. ananatis JCM6986 with the plasmid pRed/ET (Gene Bridges) was performed using a Gene Pulser/MicroPulser (Bio-Rad Laboratories). Subsequently, the tnaA gene in the P. ananatis genome was replaced by the KmR cassette50. The successful insertion of the KmR was confirmed by the acquisition of kanamycin resistance in P. ananatis, and the loss of the pRed/ET plasmid was verified by the absence of tetracycline resistance. The deletion of the tnaA gene was further confirmed by specific PCR amplification using the primers PA_JCM6986_tnaA_check_F and PA_JCM6986_tnaA_check_R (Supplementary Table 1e). The resultant strain was designated as P. ananatis JCM6986 ΔtnaA (Supplementary Table 1d).
Transformation and expression of the tna operon in Pantoea sp. F
Given that deletion of the tnaC gene has been reported to result in constitutive expression of tnaAB genes51, we first disrupted the tnaC gene in P. ananatis JCM6986 by inserting a KmR region amplified by PCR using the primers PA_JCM6986_ΔtnaC_F and PA_JCM6986_ΔtnaC-RUT_R from the E. coli ∆intS mutant (Supplementary Table 1e) into the tnaC locus (Supplementary Fig. 2). The successful deletion of tnaC was verified by specific PCR amplification with the primers PA_JCM6986_tnaC_check_F and PA_JCM6986_tnaC-RUT_check_R (Supplementary Table 1e). Next, the mutated tna operon from the resultant strain P. ananatis JCM6986 tnaC-RUT::KmR (Supplementary Fig. 2) was amplified using the primers SymF_intB/JCM6986_tnaAB_F and SymF_intB/JCM6986_tnaAB_R (Supplementary Table 1e). The intB gene of Pantoea sp. Plst-Sym F (Supplementary Table 1a) was then replaced by the mutated tna operon from the PCR fragment, resulting in the construction of the Plst-Sym F intB::tna operon tnaC-RUT::KmR (Supplementary Fig. 2). Finally, the KmR cassette was excised using FLP recombinase expressed from the plasmid pFLP3. The loss of pFLP3 was facilitated by the sacB-based suicide gene system52, thereby creating the final strain, Plst-Sym F intB::tnaAB (Supplementary Table 1d and Supplementary Fig. 2). The successful insertion of tnaAB was verified by specific PCR amplification with the primers SymF_intB_check_F and SymF_intB_check_R (Supplementary Table 1e). As a control strain in the infection experiments, the strain Plst-Sym F ΔintB (Supplementary Table 1d) was constructed by the same protocol for the strain Plst-Sym F intB::tnaAB except that the primers SymF_ΔintB_F and SymF_ΔintB_R (Supplementary Table 1e) were used to obtain a KmR region. The successful deletion of intB was verified by specific PCR amplification with the primers SymF_intB_check_F and SymF_intB_check_R (Supplementary Table 1e).
Genome sequencing and analysis
DNA samples of the uncultivable symbionts A-Plst-Sym A and B-Plst-Sym B (Supplementary Table 1a) were extracted from the symbiotic organs dissected from adult insects of P. stali. DNA samples of the cultivable bacteria were extracted from overnight cultures in LB liquid medium at 25 °C. The extraction of DNA from the materials was conducted using DNeasy Blood and Tissue Kits (Qiagen). The DNA samples were subjected to library preparation and sequencing using either the PacBio RSII or PacBio Sequel sequencing system (Pacific Biosciences of California). For the symbionts Sym A, C-Lami-ISGK-165 and C-Scam-OKNW-431 (Supplementary Table 1a), the DNA samples were additionally sequenced using the MinION system in combination with Ligation Sequencing Kit V14 and R10.4.1 flow cells (Oxford Nanopore Technologies). Base calling of Nanopore long reads was conducted using Guppy version 6.5.7 + ca6d6af (Oxford Nanopore Technologies) with minimap version 2.24-r1122 (ref. 53). De novo assembly was performed using Flye v.2.9.1 with default settings and an estimated genome size of 5.0 Mb (ref. 54). When libraries were sequenced with the PacBio RSII or Sequel system, raw PacBio reads were mapped to the draft assemblies via BLASR v.5.3.3 (ref. 55). The assembled sequences were then polished using the original PacBio reads with Arrow v.2.3.2 (ref. 56). The chromosomal genomes of A-Plst-SymA, C-Lami-ISGK-165 and C-Scam-OKNW-431 were not assembled into one circular contig owing to the presence of long segmental repeats among different chromosomal regions. For these three strains, de novo assemblies of the ONT reads with Flye were used for gap closing. The draft assemblies were polished with two rounds of medaka (v.1.4.1; https://github.com/nanoporetech/medaka). To reveal assembly errors, original sequencing reads were mapped to completing circular genomes using BWA v.0.7.17 (ref. 57). Possible sequence errors were identified using these mapped data via bam-readcont v.1.0.1 (ref. 58) and then manually inspected using IGV v.2.16.2 (ref. 59). Genome annotation was performed using DFAST v.1.2.20 (ref. 60).
Molecular phylogenetic analysis
The genome sequences of the Pantoea symbionts, environmental Pantoea strains and allied bacteria were annotated using DFAST v.1.2.20. Then, the proteome sets were analysed with publicly available ones using bcgTree v.1.2.0 (ref. 61), which automatically extracted 107 essential single-copy core genes from amino acid sequences of the whole genome data. Ambiguously aligned regions were trimmed by using Gblocks v.0.9.1b with manual inspection62. In total, 106 gene alignments (1 gene, rpmH, was excluded owing to having missing data) were concatenated and a partitioning file was generated to mark the boundaries of each gene. Corresponding amino acid substitution models were estimated by the Bayesian information criteria using ModelTest-NG v.0.2.0 (ref. 63). The maximum likelihood analysis was conducted using RAxML-NG v0.9.0 (ref. 64). Bootstrap values were obtained with 1,000 resamples. Bayesian inference analysis was conducted using MrBayes v.3.2.7a (ref. 65) with 10 million generations of Markov chain Monte Carlo runs with sampling every 100 generations. The first 25% of the samples were discarded as burn-in, and the remaining trees were used to calculate posterior probabilities. The stationarity of the runs was assessed using Tracer v.1.7.2 (ref. 66).
Transcriptomic analyses
Total RNA was extracted from homogenates of the symbiotic organ using RNAiso (Takara Bio) in combination with the RNeasy Mini Kit (Qiagen). Ribosomal RNAs of both insect and bacterial origin were removed from the total RNA using the Ribo‑Zero Gold rRNA Removal Kit (Epidemiology; Illumina). The rRNA‑depleted RNA samples were used to construct paired‑end sequencing libraries with either the SureSelect Strand‑Specific RNA Library Prep Kit (Agilent Technologies) or the TruSeq RNA Library Prep Kit v2 (Illumina). Libraries were sequenced on an Illumina HiSeq 3000 or HiSeq X platform.
Raw sequencing reads were quality trimmed and mapped to the E. coli BW25113 reference genome (accession number NZ_CP009273), and gene‑level read counts were obtained using CLC Genomics Workbench v10.0 (Qiagen). Normalization of read counts and differential gene expression analyses were performed using edgeR v3.32.1 (ref. 67).
Statistics and reproducibility
We statistically compared the effects of experimental treatments on the host phenotypes, including adult emergence rates, colour hues, and indole and tryptophan contents, using Wilcoxon rank–sum test on account of their non-Gaussian distributions. For multiple comparisons, P values were adjusted using Hommel’s method. Exact P values were provided in source data files. All statistical analyses were conducted using R version 4.4.0 (ref. 68). The number of replicates for each experiment is indicated in the respective figures. No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during the experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All genome sequencing data produced in this study were deposited in the DNA Data Bank of Japan (DDBJ) Sequence Read Archive under accession numbers AP035891–AP036010 (Supplementary Table 1a). The data have been deposited with links to BioProject accession number PRJDB17425 in the DDBJ BioProject database. The transcriptomics data are available via DDBJ under BioProject accession number PRJDB5544. Source data are provided with this paper.
References
Douglas, A. E. The Symbiotic Habit (Princeton Univ. Press, 2010).
McFall-Ngai, M. et al. Animals in a bacterial world, a new imperative for the life sciences. Proc. Natl Acad. Sci. USA 110, 3229–3236 (2013).
Leigh, E. J. The evolution of mutualism. J. Evol. Biol. 23, 2507–2508 (2010).
Bronstein, J. L. Mutualism (Oxford Univ. Press, 2015).
Hoang, K. L., Moran, L. T. & Gerardo, N. M. Experimental evolution as an underutilized tool for studying beneficial animal–microbe interactions. Front. Microbiol. 7, 1444 (2016).
King, K. C. et al. Rapid evolution of microbe-mediated protection against pathogens in a worm host. ISME J. 10, 1915–1924 (2016).
Pankey, M. S. et al. Host-selected mutations converging on a global regulator drive an adaptive leap towards symbiosis in bacteria. eLife 6, e24414 (2017).
Tso, G. H. W. et al. Experimental evolution of a fungal pathogen into a gut symbiont. Science 362, 589–595 (2018).
Robinson, C. D. et al. Experimental bacterial adaptation to the zebrafish gut reveals a primary role for immigration. PLoS Biol. 16, e2006893 (2018).
Mehta, A. P. et al. Engineering yeast endosymbionts as a step toward the evolution of mitochondria. Proc. Natl Acad. Sci. USA 115, 11769–11801 (2018).
Drew, G. C., Stevens, E. J. & King, K. C. Microbial evolution and transitions along the parasite–mutualist continuum. Nat. Rev. Microbiol. 19, 623–638 (2021).
Obeng, N. et al. Bacterial c-di-GMP has a key role in establishing host–microbe symbiosis. Nat. Microbiol. 8, 1809–1819 (2023).
Koga, R. et al. Single mutation makes Escherichia coli an insect mutualist. Nat. Microbiol. 7, 1141–1150 (2022).
Kaltenpoth, M. Fast track to mutualism. Nat. Microbiol. 7, 1104–1105 (2022).
Hosokawa, T. et al. Obligate bacterial mutualists evolving from environmental bacteria in natural insect populations. Nat. Microbiol. 1, 15011 (2016).
Oishi, S., Moriyama, M., Koga, R. & Fukatsu, T. Morphogenesis and development of midgut symbiotic organ of the stinkbug Plautia stali (Hemiptera: Pentatomidae). Zool. Lett. 5, 16 (2019).
Oishi, S., Harumoto, T., Okamoto-Furuta, K., Moriyama, M. & Fukatsu, T. Mechanisms underpinning morphogenesis of a symbiotic organ specialized for hosting an indispensable microbial symbiont in stinkbugs. mBio 14, e00522-23 (2023).
Oishi, S., Moriyama, M., Mizutani, M., Futahashi, R. & Fukatsu, T. Regulation and remodeling of microbial symbiosis in insect metamorphosis. Proc. Natl Acad. Sci. USA 120, e2304879120 (2022).
Walk, S. T. et al. Cryptic lineages of the genus Escherichia. Appl. Environ. Microbiol. 75, 6534–6544 (2009).
Tenaillon, O., Skurnik, D., Picard, B. & Denamur, E. The population genetics of commensal Escherichia coli. Nat. Rev. Microbiol. 8, 207–217 (2010).
Deutscher, J., Francke, C. & Postma, P. W. How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol. Mol. Biol. Rev. 70, 939–1031 (2006).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 6, 613–624 (2008).
Santos-Zavaleta, A. et al. RegulonDB v 10.5: tackling challenges to unify classic and high throughput knowledge of gene regulation in E. coli K-12. Nucleic Acids Res. 47, D212–D220 (2019).
Happold, F. C. in Advances in Enzymology and Related Areas of Molecular Biology Vol. 10 (ed. Nord, F. F.) 51–81 (Wiley, 1950).
Agus, A., Planchais, J. & Sokol, H. Gut microbiota regulation of tryptophan metabolism in health and disease. Cell Host Microbe 23, 716–724 (2018).
McCutcheon, J. P. & Moran, N. A. Functional convergence in reduced genomes of bacterial symbionts spanning 200 My of evolution. Genome Biol. Evol. 2, 708–718 (2010).
Gunsalus, J. P. & Yanofsky, C. Nucleotide sequence and expression of Escherichia coli trpR, the structural gene for the trp aporepressor. Proc. Natl Acad. Sci. USA 77, 7117–7121 (1980).
Nikoh, N., Hosokawa, T., Oshima, K., Hattori, M. & Fukatsu, T. Reductive evolution of bacterial genome in insect gut environment. Genome Biol. Evol. 3, 702–714 (2011).
Kaiwa, N. et al. Symbiont-supplemented maternal investment underpinning host’s ecological adaptation. Curr. Biol. 24, 2465–2470 (2014).
Salem, H. et al. Vitamin supplementation by gut symbionts ensures metabolic homeostasis in an insect host. Proc. R. Soc. B 281, 20141838 (2014).
Moriyama, M. & Fukatsu, T. Host’s demand for essential amino acids is compensated by an extracellular bacterial symbiont in a hemipteran insect model. Front. Physiol. 13, 1028409 (2022).
Blattner, E. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1462 (1997).
Lay, C. Y., Baumann, L. & Baumann, P. Amplification of trpEG: adaptation of Buchnera aphidicola to an endosymbiotic association with aphids. Proc. Natl Acad. Sci. USA 91, 3819–3823 (1994).
Baumann, P. Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu. Rev. Microbiol. 59, 155–189 (2005).
Mittler, T. E. & Dadd, R. H. Artificial feeding and rearing of the aphid, Myzus persicae (Sulzer), on a completely defined synthetic diet. Nature 195, 404 (1962).
Shibao, H., Kutsukake, M., Lee, J. M. & Fukatsu, T. Maintenance of soldier-producing aphids on an artificial diet. J. Insect Physiol. 48, 495–505 (2002).
Hosokawa, T., Matsuura, Y., Kikuchi, Y. & Fukatsu, T. Recurrent evolution of gut symbiotic bacteria in pentatomid stinkbugs. Zool. Lett. 2, 24 (2016).
Otero-Bravo, A., Goffredi, S. & Sabree, Z. L. Cladogenesis and genomic streamlining in extracellular endosymbionts of tropical stink bugs. Genome Biol. Evol. 10, 680–693 (2018).
Otero-Bravo, A. & Sabree, Z. L. Multiple concurrent and convergent stages of genome reduction in bacterial symbionts across a stink bug family. Sci. Rep. 11, 7731 (2021).
Salem, H., Florez, L., Gerardo, N. & Kaltenpoth, M. An out-of-body experience: the extracellular dimension for the transmission of mutualistic bacteria in insects. Proc. R. Soc. B 282, 20142957 (2015).
Hosokawa, T. & Fukatsu, T. Relevance of microbial symbiosis to insect behavior. Curr. Opin. Insect Sci. 39, 91–100 (2020).
Sugiyama, R., Moriyama, M., Koga, R. & Fukatsu, T. Host range of naturally and artificially evolved symbiotic bacteria for a specific host insect. mBio 15, e01342-24 (2024).
Boya, B. R., Kumar, P., Lee, J. H. & Lee, J. Diversity of the tryptophanase gene and its evolutionary implications in living organisms. Microorganisms 9, 2126 (2021).
McCutcheon, J. P. & Moran, N. A. Extreme genome reduction in symbiotic bacteria. Nat. Rev. Microbiol. 10, 13–26 (2012).
Nishide, Y. et al. Aseptic rearing procedure for the stinkbug Plautia stali (Hemiptera: Pentatomidae) by sterilizing food-derived bacterial contaminants. Appl. Entomol. Zool. 52, 407–415 (2017).
Tanahashi, M. & Fukatsu, T. Natsumushi: image measuring software for entomological studies. Entomol. Sci. 21, 347–360 (2018).
Porubsky, P. R., Scott, E. E. & Williams, T. D. p-Dimethylaminocinnamaldehyde derivatization for colorimetric detection and HPLC–UV/vis–MS/MS identification of indoles. Arch. Biochem. Biophys. 475, 14–17 (2008).
MacWilliams, M. P. Indole Test Protocol (American Society for Microbiology, 2009); https://asm.org/protocols/indole-test-protocol
Li, G. & Young, K. D. A cAMP-independent carbohydrate-driven mechanism inhibits tnaA expression and TnaA enzyme activity in Escherichia coli. Microbiology 160, 2079–2088 (2014).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
van der Stel, A. X. et al. Structural basis for the tryptophan sensitivity of TnaC-mediated ribosome stalling. Nat. Commun. 12, 5340 (2021).
Choi, K. H. et al. A Tn 7-based broad-range bacterial cloning and expression system. Nat. Methods 2, 443–448 (2005).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. Assembly of long error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Chaisson, M. J. & Tesler, G. Mapping single molecule sequencing reads using basic local alignment with successive refinement (BLASR): application and theory. BMC Bioinformatics 13, 238 (2012).
Chin, C.-S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569 (2013).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Khanna, A. et al. Bam-readcount—rapid generation of basepair-resolution sequence metrics. J. Open Source Softw. 7, 3722 (2022).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Tanizawa, Y., Fujisawa, T. & Nakamura, Y. DFAST: a flexible prokaryotic genome annotation pipeline for faster genome publication. Bioinformatics 34, 1037–1039 (2018).
Ankenbrand, M. J. & Keller, A. bcgTree: automatized phylogenetic tree building from bacterial core genomes. Genome 59, 783–791 (2016).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Darriba, D. et al. ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol. 37, 291–294 (2020).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455 (2019).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 67, 901–904 (2018).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. EdgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2024).
Acknowledgements
We thank N. Obana (University of Tsukuba) for providing the J23119 promoter-containing cassette, D. Haraguchi (Okinawa Prefectural Plant Protection Center) and Y. Kikuchi (National Institute of Advanced Industrial Science and Technology) for providing insect samples, and T. Tanaka, T. Matsushita, T. Hachikawa, N. Shimizu, S. Toyoda, M. Taguchi, M. Kobayashi and N. Yamane for technical assistance. This study was supported by the Japan Science and Technology Agency ERATO grant number JPMJER1902 (T.F. and R.K.) and the Japan Society for the Promotion of Science (JSPS) KAKENHI grant numbers JP25221107 (T.F. and R.K.), JP17H06388 (T.F. and S.S.), JP22128001 (T.F. and S.S.) and JP22128007 (T.F.).
Author information
Authors and Affiliations
Contributions
Y.W., M.M., R.K. and T.F. conceived the project and designed the experiments. Y.W. conducted most of the experimental works including bacterial infection, insect rearing, fitness evaluation, E. coli and Pantoea mutant generation and so on. M.M. performed metabolomic analyses of amino acids, indole and its derivatives, and so on. R.K. conducted genomic and phylogenetic analyses of diverse stinkbug symbionts and soil-derived bacterial isolates. T.H. performed a field survey to collect natural insects and soil samples, from which Pantoea strains symbiotic to P. stali were isolated. K.O. and H.T. conducted inoculation experiments of insect-derived and soil-derived Pantoea isolates into P. stali and evaluated fitness consequences. S.S. and N.N. supported genomic and phylogenetic analyses. T.F. wrote the paper with input from all authors. All authors approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Jun-Bo Luan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genes commonly down- or up-regulated in independent mutualistic E. coli evolutionary lines,CmL05 and GmL07, and also down- or up-regulated by the CCR disruption in E. coli.
(a) Venn diagram showing the numbers of commonly down-regulated genes. (b-h) Functional categories and expression levels of the commonly down-regulated genes. (b) Transporters. (c) Carbohydrate metabolism. (d) Amino acid metabolism. (e) Lipid metabolism. (f) Quorum sensing. (g) Transcriptional regulators. (h) Others. (i) Venn diagram showing the numbers of commonly up-regulated genes. (j-k) Functional categories and expression levels of the commonly up-regulated genes. (j) Nucleotide metabolism. (k) Transporter. The inset figure at the bottom right represents the explanations of the elements in the plots. For box plots, center lines, limits, and dots show medians, first and third quartiles, and data points. Biological replicate numbers and exact FDR q-values are provided in the source data file.
Extended Data Fig. 2 Levels of free amino acids in P. stali infected with a natural symbiont Pantoea sp. A (Sym A), mutant E. coli strains (∆cyaA, ∆crp, ∆tnaA and ∆tnaB), and a wild-type E. coli strain (∆intS).
(a) Free amino acids in hemolymph. (b) Free amino acids in symbiotic organ. Note that tryptophan levels were drastically higher in the insects infected with Sym A, ∆cyaA, ∆crp and ∆tnaA than those infected with ∆tnaB and ∆intS. Bars, limits, and dots show means, standard deviations, and data points. Biological replicate numbers are indicated on the graphs.
Extended Data Fig. 3 Tryptophan production and tryptophanase activity of E. coli mutant strains.
(a) Levels of free amino acids in hemolymph of P. stali infected with a mutant E. coli strain (∆cyaA), an evolutionary E. coli strain (CmL05G13), and a wild-type E. coli strain (∆intS). Bars, limits, and dots show means, standard deviations, and data points. (b) Comparison of the tryptophan levels. For box plots, center lines, limits, and dots show medians, first and third quartiles, and data points. Note that ∆cyaA and CmL05G13 are both CCR-deficient. Different alphabetical letters (a, b) indicate statistically significant differences (pairwise Wilcoxon rank-sum test with Hommel’s correction: P < 0.05, two-sided). The exact P-values are provided in the source data file. (c) Tryptophanase assay of the E. coli control strain ΔintS, the E. coli mutant strains ΔtnaA, ΔtnaB, ΔcyaA and Δcrp, and the evolutionary E. coli strain CmL05G13. Red color indicates tryptophanase activity. (d, e) Cell growth dynamics (d) and indole production (e) of ΔintS and CmL05G13. Bacterial cell culture was conducted at 25 °C in LB liquid medium with shaking. Line points, limits, and dots show means, standard deviations, and data points. Biological replicate numbers are indicated on the graphs.
Extended Data Fig. 4 Effects of a natural symbiont Pantoea sp. A (Sym A), mutant E. coli strains (∆cyaA, ∆crp, ∆tnaA and ∆tnaB), and a wild-type E. coli strain (∆intS) on phenotypes of P. stali.
(a) Adult female body color. (b) Adult male body color. (c) Adult female body size. (d) Adult male body size. For box plots, center lines, limits, and dots show medians, first and third quartiles, and data points. Different alphabetical letters (a, b, c) indicate statistically significant differences (pairwise Wilcoxon rank-sum test with Hommel’s correction: P < 0.05, two-sided). The exact P-values are provided in the source data file. Biological replicate numbers are indicated on the graphs.
Extended Data Fig. 5 Rescue phenotypes of P. stali infected with E. coli mutants affecting tryptophanase expression.
(a-c) Effects of E. coli mutants ΔcyaA Pconst-tnaA (ΔcyaA strain whose tnaA is driven to express constitutively) and ΔtnaA::tnaA (ΔtnaA strain which is complemented with functional tnaA) in comparison with the mutualistic E. coli strains ΔcyaA and ΔtnaA and the wildtype control E. coli strain ΔintS. (a) Adult emergence rate. (b) Adult body color. (c) Tryptophan level in hemolymph. For box plots, center lines, limits, and dots show medians, first and third quartiles, and data points. Different alphabetical letters (a–d) indicate statistically significant differences (pairwise Wilcoxon rank-sum test with Hommel’s correction: P < 0.05, two-sided). The exact P-values are provided in the source data file. (d) Enzymatic assay of tryptophanase activity of the E. coli strains. (e) Levels of free amino acids in hemolymph of P. stali infected with the E. coli mutants. Bars, limits, and dots show means, standard deviations, and data points. Biological replicate numbers are indicated on the graphs.
Extended Data Fig. 6 Molecular mechanisms underlying the evolution of P. stali-E. coli mutualism uncovered in this study.
(a) Before the evolution of mutualism. (b) After the evolution of mutualism.
Extended Data Fig. 7 Titers of tryptophan-derived metabolites in hemolymph of P. stali infected with different E. coli strains.
(a) A schematic overview of tryptophan metabolism. Compounds quantified by LC-MS are highlighted in bold. Red, blue and black represent metabolites that increased, decreased and unchanged by infection with the host performance-improving E. coli mutants (CmL05G13, ΔcyaA and ΔtnaA), respectively. (b-k) Titers of tryptophan and its derived metabolites in the host hemolymph. (b) Tryptophan (Trp). (c) Indole. (d) 5-Hydroxytryptophan (5HTrp). (e) Hydroxyindoleacetate (HIAA). (f) Kynurenine (Kyn). (g) 3-Hydroxykynurenine (3HK). (h) Xanthurenic acid (XA). (i) Indole-3-acetic acid (IAA). (j) Indole-3-carboxylic acid (ICA). (k) Indole-3-ethanol (IEt). Other abbreviations: 3HAA, 3-Hydroxyanthranilic acid; CA, Cinnabarinic acid; 5HT, 5-Hydroxytryptamine; IAM, Indole-3-acetamide; IAAId, Indole-3-acetaldehyde; IGA, Indole-3-glyoxicacid. For box plots, center lines, limits, and dots show medians, first and third quartiles, and data points. Different alphabetical letters (a-d) indicate statistically significant differences (pairwise Wilcoxon rank-sum test with Hommel’s correction: P < 0.05, two-sided). The exact P-values are provided in the source data file.
Extended Data Fig. 8 Effects of a tryptophan overproducing mutant E. coli strain ∆trpR on phenotypes of P. stali in comparison with the mutualistic E. coli strain ∆tnaA and the wild-type control E. coli strain ∆intS.
(a) Adult emergence rate. (b) Adult body color. (c) Tryptophan level in hemolymph. For box plots, center lines, limits, and dots show medians, first and third quartiles, and data points. Different alphabetical letters (a, b, c) indicate statistically significant differences (pairwise Wilcoxon rank-sum test with Hommel’s correction: P < 0.05, two-sided). The exact P-values are provided in the source data file. (d) Levels of free amino acids in hemolymph. Bars, limits, and dots show means, standard deviations, and data points. Biological replicate numbers are indicated on the graphs.
Extended Data Fig. 9 Microbial and symbiotic properties of P. ananatis isolates.
(a) In vitro evaluation of tryptophan synthesis by natural Pantoea symbionts of P. stali and environmental P. ananatis isolates. All bacterial strains proliferated in M9 minimal medium containing glucose as the sole carbon source, indicating that they are all capable of synthesizing tryptophan. Line points, limits, and dots show means, standard deviations, and data points. Biological replicate numbers are indicated on the graphs. (b-d) Enzymatic assay of tryptophanase activity and indole production by the bacterial strains cultured in M9-based media at 25 °C, shaking at 180 rpm, for 48 h. (b) In M9 medium, neither tryptophanase activity nor indole production was detected from the P. ananatis isolates with functional tnaA gene, probably because of low bacterial density and metabolic activity in the minimal medium. (c, d) When either 0.1% tryptophan or 1% Tryptone was added, tryptophanase activity and consequent indole production (red) were detected from the P. ananatis isolates but not from the natural Pantoea symbionts. Note that tnaA-disrupted P. ananatis strain (JCM6986 ΔtnaA) was incapable of indole production. (e) PCR detection of tnaA gene in the P. ananatis isolates. (f) Adult emergence rates of P. stali infected with the P. ananatis isolates. For box plots, center lines, limits, and dots show medians, first and third quartiles, and data points. Different alphabetical letters (a, b, c) indicate statistically significant differences (pairwise Wilcoxon rank-sum test with Hommel’s correction: P < 0.05, two-sided). Biological replicate numbers are indicated on the graphs. The exact P-values are provided in the source data file.
Extended Data Fig. 10 Adult emergence rates of P. stali infected with original Pantoea symbionts, heterospecific Pantoea symbionts from different stinkbug species, and potential Pantoea symbionts isolated from environmental soil samples.
Bars, limits and dots show means, standard deviations and data points. Biological replicate numbers are shown in parentheses.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2.
Supplementary Table (download XLSX )
Supplementary Table 1a–e.
Source data
Source Data Fig. 1 (download XLSX )
Numerical and statistical source data.
Source Data Fig. 2 (download XLSX )
Numerical and statistical source data.
Source Data Fig. 3 (download TXT )
Multiple alignments formatted for MrBayes and RAxML-NG and substitution model for RAxML-NG.
Source Data Fig. 3 (download XLSX )
Taxonomy labels for the tree drawing.
Source Data Fig. 4 (download XLSX )
Numerical source data.
Source Data Fig. 4 (download JPEG )
Unprocessed gel image of Fig. 4b.
Source Data Fig. 5 (download XLSX )
Numerical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Numerical and statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Numerical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Numerical and statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Numerical and statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Numerical and statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Numerical and statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Numerical and statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Numerical and statistical source data.
Source Data Extended Data Fig. 9 (download JPEG )
Unprocessed gel image of Fig. 9e.
Source Data Extended Data Fig. 10 (download XLSX )
Numerical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Moriyama, M., Koga, R. et al. Tryptophanase disruption promotes insect–bacterium mutualism. Nat Microbiol 11, 759–769 (2026). https://doi.org/10.1038/s41564-026-02264-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-026-02264-z