Abstract
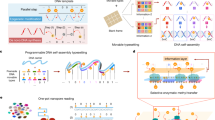
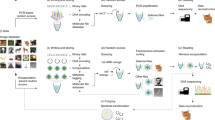
Any modern information system is expected to feature a set of primordial features and functions: a substrate stably carrying data; the ability to repeatedly write, read, erase, reload and compute on specific data from that substrate; and the overall ability to execute such functions in a seamless and programmable manner. For nascent molecular information technologies, proof-of-principle realization of this set of primordial capabilities would advance the vision for their continued development. Here we present a DNA-based store and compute engine that captures these primordial capabilities. This system comprises multiple image files encoded into DNA and adsorbed onto ~50-μm-diameter, highly porous, hierarchically branched, colloidal substrate particles comprised of naturally abundant cellulose acetate. Their surface areas are over 200 cm2 mg−1 with binding capacities of over 1012 DNA oligos mg−1, 10 TB mg−1 or 104 TB cm−3. This ‘dendricolloid’ stably holds DNA files better than bare DNA with an extrapolated ability to be repeatedly lyophilized and rehydrated over 170 times compared with 60 times, respectively. Accelerated ageing studies project half-lives of ~6,000 and 2 million years at 4 °C and −18 °C, respectively. The data can also be erased and replaced, and non-destructive file access is achieved through transcribing from distinct synthetic promoters. The resultant RNA molecules can be directly read via nanopore sequencing and can also be enzymatically computed to solve simplified 3 × 3 chess and sudoku problems. Our study establishes a feasible route for utilizing the high information density and parallel computational advantages of nucleic acids.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw sequencing data, sequences of DNA oligos and source data for all plots are available at https://doi.org/10.5281/zenodo.12169723 (ref. 62) and https://doi.org/10.5281/zenodo.12192541 (ref. 63) and https://github.com/keung-lab/Lin-et-al-2024.git, https://github.com/dna-storage/framed/tree/sdc_nature_submission and https://github.com/dna-storage/framed/releases. All other data are available upon reasonable request. Source data are provided with this paper.
Code availability
The software algorithms we developed to perform the reported analyses are available at https://doi.org/10.5281/zenodo.12169723 (ref. 62), https://github.com/dna-storage/framed/tree/sdc_nature_submission and https://github.com/dna-storage/framed/releases under a permissive open source license with instructions for installation. We implemented code in Python using many standard open-source packages, including biopython, primer3, numpy, scipy, pandas and others. These dependences are documented in the form of a Python requirements.txt file that guides installation of additional dependent software packages.
References
Ceze, L., Nivala, J. & Strauss, K. Molecular digital data storage using DNA. Nat. Rev. Genet. 20, 456–466 (2019).
Copeland, B. J. in The Stanford Encyclopedia of Philosophy Winter 2020 edn (ed. Zalta, E. N.) (Stanford Univ., 2020).
Ceruzzi, P. E. A history of modern computing. Choice Rev. Online 36, 36-4531–36-4531 (1999).
Goldman, N. et al. Towards practical, high-capacity, low-maintenance information storage in synthesized DNA. Nature 494, 77–80 (2013).
Church, G. M., Gao, Y. & Kosuri, S. Next-generation digital information storage in DNA. Science 337, 1628 (2012).
Grass, R. N., Heckel, R., Puddu, M., Paunescu, D. & Stark, W. J. Robust chemical preservation of digital information on DNA in silica with error-correcting codes. Angew. Chem. Int. Ed. 54, 2552–2555 (2015).
Blawat, M. et al. Forward error correction for DNA data storage. Procedia Comput. Sci. 80, 1011–1022.
Erlich, Y. & Zielinski, D. DNA Fountain enables a robust and efficient storage architecture. Science 355, 950–954 (2017).
Lee, H. H., Kalhor, R., Goela, N., Bolot, J. & Church, G. M. Terminator-free template-independent enzymatic DNA synthesis for digital information storage. Nat. Commun. 10, 2383 (2019).
Palluk, S. et al. De novo DNA synthesis using polymerase-nucleotide conjugates. Nat. Biotechnol. 36, 645–650 (2018).
Lopez, R. et al. DNA assembly for nanopore data storage readout. Nat. Commun. 10, 2933 (2019).
Mao, C., LaBean, T. H., Reif, J. H. & Seeman, N. C. Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407, 493–496 (2000).
Adleman, L. M. Molecular computation of solutions to combinatorial problems. Science 266, 1021–1024 (1994).
Organick, L. et al. Random access in large-scale DNA data storage. Nat. Biotechnol. 36, 242–248 (2018).
Tabatabaei Yazdi, S. M. H., Yuan, Y., Ma, J., Zhao, H. & Milenkovic, O. A rewritable, random-access DNA-based storage system. Sci. Rep. 5, 1–10 (2015).
Yazdi, S. M. H. T., Gabrys, R. & Milenkovic, O. Portable and error-free DNA-based data storage. Sci. Rep. https://doi.org/10.1038/s41598-017-05188-1 (2017).
Bornholt, J. et al. A DNA-based archival storage system. In Proc. 21st International Conference on Architectural Support for Programming Languages and Operating Systems—ASPLOS ’16 (eds Conte, T. & Zhou, Y.) 637–649 (ACM Press, 2016).
Bögels, B. W. A. et al. DNA storage in thermoresponsive microcapsules for repeated random multiplexed data access. Nat. Nanotechnol. 18, 912–921 (2023).
Benenson, Y. et al. Programmable and autonomous computing machine made of biomolecules. Nature 414, 430–434 (2001).
Bell, N. A. W. & Keyser, U. F. Digitally encoded DNA nanostructures for multiplexed, single-molecule protein sensing with nanopores. Nat. Nanotechnol. 11, 645–651 (2016).
Dickinson, G. D. et al. An alternative approach to nucleic acid memory. Nat. Commun. 12, 2371 (2021).
Chen, K. et al. Digital data storage using DNA nanostructures and solid-state nanopores. Nano Lett. 19, 1210–1215 (2019).
Chen, K., Zhu, J., Bošković, F. & Keyser, U. F. Nanopore-based DNA hard drives for rewritable and secure data storage. Nano Lett. 20, 3754–3760 (2020).
Zhang, Y. et al. DNA origami cryptography for secure communication. Nat. Commun. 10, 5469 (2019).
Numajiri, K., Kimura, M., Kuzuya, A. & Komiyama, M. Stepwise and reversible nanopatterning of proteins on a DNA origami scaffold. Chem. Commun. 46, 5127 (2010).
Roh, S., Williams, A. H., Bang, R. S., Stoyanov, S. D. & Velev, O. D. Soft dendritic microparticles with unusual adhesion and structuring properties. Nat. Mater. 18, 1315–1320 (2019).
Williams, A. H. et al. Printable homocomposite hydrogels with synergistically reinforced molecular-colloidal networks. Nat. Commun. 12, 2834 (2021).
Bergenstråhle, M., Wohlert, J., Himmel, M. E. & Brady, J. W. Simulation studies of the insolubility of cellulose. Carbohydr. Res. 345, 2060–2066 (2010).
Lindman, B., Medronho, B., Alves, L., Norgren, M. & Nordenskiöld, L. Hydrophobic interactions control the self-assembly of DNA and cellulose. Q. Rev. Biophys. 54, e3 (2021).
Bang, R. S., Roh, S., Williams, A. H., Stoyanov, S. D. & Velev, O. D. Fluid flow templating of polymeric soft matter with diverse morphologies. Adv. Mater. 35, 2211438 (2023).
Ali, M. E. et al. in Reference Module in Materials Science and Materials Engineering (Elsevier, 2016); https://doi.org/10.1016/B978-0-12-803581-8.04075-3
Paul, A. & Bhattacharya, S. Chemistry and biology of DNA-binding small molecules. Curr. Sci. 102, 212–231 (2012).
Koch, J. et al. A DNA-of-things storage architecture to create materials with embedded memory. Nat. Biotechnol. 38, 39–43 (2020).
Lin, K. N., Grandhi, T. S. P., Goklany, S. & Rege, K. Chemotherapeutic drug‐conjugated microbeads demonstrate preferential binding to methylated plasmid DNA. Biotechnol. J. 13, 1700701 (2018).
Stetefeld, J., McKenna, S. A. & Patel, T. R. Dynamic light scattering: a practical guide and applications in biomedical sciences. Biophys. Rev. 8, 409–427 (2016).
Lin, K. N., Volkel, K., Tuck, J. M. & Keung, A. J. Dynamic and scalable DNA-based information storage. Nat. Commun. 11, 2981 (2020).
Fisher, S. et al. A scalable, fully automated process for construction of sequence-ready human exome targeted capture libraries. Genome Biol. 12, R1 (2011).
DeAngelis, M. M., Wang, D. G. & Hawkins, T. L. Solid-phase reversible immobilization for the isolation of PCR products. Nucleic Acids Res. 23, 4742–4743 (1995).
Chen, Y.-J. et al. Quantifying molecular bias in DNA data storage. Nat. Commun. 11, 3264 (2020).
Matange, K., Tuck, J. M. & Keung, A. J. DNA stability: a central design consideration for DNA data storage systems. Nat. Commun. 12, 1358 (2021).
Lauková, L., Konečná, B., Janovičová, Ľ., Vlková, B. & Celec, P. Deoxyribonucleases and their applications in biomedicine. Biomolecules 10, 1036 (2020).
Robinson, P. K. Enzymes: principles and biotechnological applications. Essays Biochem 59, 1–41 (2015).
Loenen, W. A. M., Dryden, D. T. F., Raleigh, E. A., Wilson, G. G. & Murray, N. E. Highlights of the DNA cutters: a short history of the restriction enzymes. Nucleic Acids Res. 42, 3–19 (2014).
Allemand, J. F., Bensimon, D., Jullien, L., Bensimon, A. & Croquette, V. pH-dependent specific binding and combing of DNA. Biophys. J. 73, 2064–2070 (1997).
Vandeventer, P. E. et al. Multiphasic DNA adsorption to silica surfaces under varying buffer, pH, and ionic strength conditions. J. Phys. Chem. B 116, 5661–5670 (2012).
Cai, J. & Zhang, L. Rapid dissolution of cellulose in LiOH/urea and NaOH/urea aqueous solutions. Macromol. Biosci. 5, 539–548 (2005).
Jiménez-Ángeles, F. & Firoozabadi, A. Hydrophobic hydration and the effect of NaCl salt in the adsorption of hydrocarbons and surfactants on clathrate hydrates. ACS Cent. Sci. 4, 820–831 (2018).
Workman, R. E. et al. Nanopore native RNA sequencing of a human poly(A) transcriptome. Nat. Methods 16, 1297–1305 (2019).
Soneson, C. et al. A comprehensive examination of nanopore native RNA sequencing for characterization of complex transcriptomes. Nat. Commun. 10, 3359 (2019).
Smith, M. A. et al. Molecular barcoding of native RNAs using nanopore sequencing and deep learning. Genome Res. 30, 1345–1353 (2020).
Qiu, M. et al. RNA nanotechnology for computer design and in vivo computation. Philos. Trans. R Soc. A 371, 20120310 (2013).
Faulhammer, D., Cukras, A. R., Lipton, R. J. & Landweber, L. F. Molecular computation: RNA solutions to chess problems. Proc. Natl Acad. Sci. USA 97, 1385–1389 (2000).
Takahashi, C. N., Nguyen, B. H., Strauss, K. & Ceze, L. Demonstration of end-to-end automation of DNA data storage. Sci. Rep. 9, 4998 (2019).
Newman, S. et al. High density DNA data storage library via dehydration with digital microfluidic retrieval. Nat. Commun. 10, 1706 (2019).
Luo, Y. et al. Integrated microfluidic DNA storage platform with automated sample handling and physical data partitioning. Anal. Chem. 94, 13153–13162 (2022).
Gerasimova, Y. V. & Kolpashchikov, D. M. Towards a DNA nanoprocessor: reusable tile‐integrated DNA circuits. Angew. Chem. 128, 10400–10403 (2016).
Guz, N. et al. Bioelectronic interface connecting reversible logic gates based on enzyme and DNA reactions. ChemPhysChem 17, 2247–2255 (2016).
Polak, R. E. & Keung, A. J. A molecular assessment of the practical potential of DNA-based computation. Curr. Opin. Biotechnol. 81, 102940 (2023).
Yang, S. et al. DNA as a universal chemical substrate for computing and data storage. Nat. Rev. Chem. 8, 179–194 (2024).
Cherry, K. M. & Qian, L. Scaling up molecular pattern recognition with DNA-based winner-take-all neural networks. Nature 559, 370–376 (2018).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Keung Lab. keung-lab/Lin-et-al-2024: v1.0.1. Zenodo https://doi.org/10.5281/zenodo.12169723 (2024).
Lin, K. & Keung, A. FASTQ data for: a primordial DNA store and compute engine. Zenodo https://doi.org/10.5281/zenodo.12192541 (2024).
Acknowledgements
We thank K. J. Tomek, M. Lee and K. Matange for helpful discussions and R. M. Kelly for use of their vacuum concentrator. We thank the Biomanufacturing Training and Education Center (BTEC) at NCSU for the use of their lyophilizer device. We also thank S. Mukherjee for providing training on the use of the Zetasizer Nano ZSP for measuring the zeta-potential of the SDC–DNA samples. This work was supported by the National Science Foundation (ECCS-2027655 and CSR-1901324). K.N.L. was supported by a Department of Education Graduate Assistance in Areas of Need fellowship, P200A160061. R.E.P. was supported by T32GM133366. Some artwork was created with BioRender.
Author information
Authors and Affiliations
Contributions
K.N.L., O.D.V. and A.J.K. conceived the study. K.N.L. planned and performed the wet-lab experiments with guidance from O.D.V. and A.J.K. K.N.L. and K.V. designed the three-file oligo library for SDC. K.V. performed file encoding and decoding simulation with guidance from J.M.T. P.W.H. performed nanopore RNA sequencing with guidance from W.T. K.N.L and R.E.P. planned and performed the simulation for computation library design. C.C. fabricated SDC materials made from cellulose acetate, cellulose and agarose with guidance from O.D.V. K.N.L. and A.S.C. designed the microfluidic system with guidance from A.S.M. K.N.L. performed zeta-potential measurements. K.V. and K.N.L. processed the next-generation sequencing and nanopore sequencing data. K.N.L. and A.J.K. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare the following competing interests: A.J.K. and J.M.T. are cofounders of DNAli Data Technologies that has potential interest in translating and commercializing DNA-based information systems. A.J.K., K.V., J.M.T. and K.N.L. are inventors of patent WO 2020/096679 which has been licensed to DNAli Data Technologies and from which some of this work is derived. W.T. has two patents (8,748,091 and 8,394,584), licensed to Oxford Nanopore Technologies (ONT), which were used for direct RNA sequencing in this work. W.T. has received travel funds to speak at symposia organized by ONT. The other authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Tom de Greef, Casey Platnich and Victor Zhirnov for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Adsorbing DNA onto SDC can lead to stable RNA generation.
A) Morphology of SDC with (left) and without (right) magnetic nanoparticles. Images were taken using a Nikon Ts2 Inverted microscope. All samples were imaged using the same 10× objective. Samples were imaged using the same microscope settings, and adjusted identically for quantification purposes. B) Equivalent masses of magnetic nanoparticles alone (MNP) or caSDC infused with MNP (MNP-caSDC) were incubated with DNA (dsDNA), washed, and subjected to IVT. An RNA gel indicates no RNA generated from the magnetic nanoparticles. C) Representative strand distribution density (left), skewness (middle), and kurtosis (right) plots for experimental samples of File1. These samples included direct sequencing of the File1 DNA obtained from the DNA synthesis provider (original File1), cDNA obtained after IVT of File1 DNA adsorbed to caSDC (IVT of SDC-File1), and cDNA obtained after IVT of unbound File1 DNA (IVT of File1). D) Representative strand distribution density (left), skewness (middle), and kurtosis (right) plots for experimental samples of File2. These samples included direct sequencing of the File2 DNA obtained from the DNA synthesis provider (original File2), cDNA obtained after IVT of File2 DNA adsorbed to caSDC (‘IVT of SDC-File2’), and cDNA obtained after IVT of unbound File2 DNA (IVT of File2). Plotted values represent the arithmetic mean, and error bars represent the standard deviation of three independent IVT reactions. Statistics was calculated using One-Way ANOVA with Tukey–Kramer post-hoc for panel C and D. a p = 1.77×10−4, b p = 2.92×10−3, c p = 1.67×10−4, d p = 3.01×10−2, e p = 6.12×10−1, f p = 9.51×10−4, g p = 3.76×10−3, h p = 3.18×10−2, i p = 4.28×10−2, j p = 1.54×10−5, k p = 1.63×10−2, l p = 1.29×10−2.
Extended Data Fig. 2 Complex files adsorbed on SDCs can be repeatedly accessed with robust and stable performance.
A) Skewness (left) and kurtosis (right) plots for strand distribution density of cDNA obtained after each sequential round of IVT to access File1 DNA adsorbed to caSDC. The results demonstrated a consistent strand distribution for sequential rounds of accessing File1 DNA adsorbed to the SDCs. B) Percent error for each DNA sequence position in the cDNA obtained after IVT of File1 DNA adsorbed to caSDC after each sequential round of IVT, and cDNA obtained after IVT of unbound File1 DNA (IVT of File1). The error rate was calculated by dividing the number of errors of a given type occurring at a nucleotide position by the total number of reads for that sequence (Method). C) Number of sequencing reads for strands in cDNA obtained after each sequential round of IVT to access File1 DNA adsorbed to caSDC. D) Percent error for each DNA sequence position in the cDNA obtained from lyophilized File3-SDC after IVT of the first lyophilization (1st round), and after the 3rd and 5th rounds of lyophilization, as obtained by Illumina sequencing. E) Percent error for each DNA sequence position in the cDNA obtained from IVT after 0, 24 and 48 hours of incubation of lyophilized or solubilized File3-SDC at 65 ˚C. The error rate was calculated by dividing the number of errors of a given type occurring at a nucleotide position by the total number of reads for that sequence. F) 300 ng DNA was input into identical IVT reactions either in a microfluidic system placed in an incubator at 37 ˚C or in a PCR tube placed in a PCR machine held at 37 ˚C for overnight. Plotted values represent the arithmetic mean, and error bars represent the standard deviation of three independent IVT reactions. Statistics was calculated using One-Way ANOVA with Tukey–Kramer post-hoc. Comparisons are relative to the first experimental condition in panel C. a p = 3.93×10−1, b p = 6.76×10−1, c p = 8.12×10−1, d p = 3.81×10−1, e p = 3.38×10−1, f p = 1.62×10−1, g p = 1.10×10−2, h p = 6.50×10−3, i p = 8.42×10−3.
Extended Data Fig. 3 Complex files adsorbed onto SDCs can be specifically erased, and new information can be reloaded onto SDCs.
A) Number of sequencing reads for strands in cDNA obtained from IVT of File DNA adsorbed to caSDC after processed with restriction digestion as file deletion. The values were measured by Illumina sequencing. B) Percent of unique strands of each file found in cDNA after File3 was specifically deleted from the three-file database as measured by Illumina sequencing. The deletion was executed with 1 µL or 5 µL of restriction enzyme. Values were plotted as a percentage of the total unique strands. C) The fraction of all sequencing reads for a targeted file, obtained from cDNA after IVT of File DNA adsorbed to caSDC after reloading. Annotation of each operation is listed in Extended Data Table 5. FileX->FileY indicates FileX was deleted and FileY was then loaded, with IVT of FileY performed, measured, and plotted. Values were measured by Illumina sequencing and plotted as a percentage of the total sequencing reads. D) Skewness (left) and kurtosis (right) plots for strand distribution density of cDNA obtained after IVT of unmodified File DNA (‘Unmod.’), erasing treated File DNA adsorbed to the SDCs (File1+DNaseI) and reloaded new File DNA after each operation on the SDCs. Values were measured by Illumina sequencing. ‘→’ denotes removing current File DNA on SDCs and reloading with new file information. Annotation of each operation is listed in Extended Data Table 5. E) Percent error for each DNA sequence position in the cDNA obtained after incubating File3-SDC under various buffer conditions, followed by IVT. F) Percent error for each DNA sequence position in the cDNA obtained after IVT of unmodified File DNA (Unmod.) and reloaded File DNA. The error rate was calculated by dividing the number of errors of a given type occurring at a nucleotide position by the total number of reads for that sequence. Values were measured by Illumina sequencing and plotted after normalizing to its number of sequencing reads found in untreated File DNA prior to IVT. Plotted values represent the arithmetic mean, and error bars represent the standard deviation of three independent IVT reactions.
Extended Data Fig. 4 Complex file DNA adsorbed on SDCs can be directly sequenced after IVT using Oxford nanopore sequencing.
A) Violin plots of the strand distributions for experimental samples of File2. These samples include direct sequencing of the File2 DNA obtained from the DNA synthesis provider (Original File2), RNA and cDNA obtained after IVT of File2 DNA adsorbed to caSDC (IVT of SDC-File2), RNA and cDNA obtained after unbound File2 DNA (IVT of File2). B) Violin plots of the strand distributions for experimental samples of File3. These samples include direct sequencing of the File3 DNA obtained from the DNA synthesis provider (Original File3), RNA and cDNA obtained after IVT of File3 DNA adsorbed to caSDC (IVT of SDC-File3), RNA and cDNA obtained after unbound File3 DNA (IVT of File3). C,D (C) Skewness and (D) kurtosis plots for strand distribution density of RNA obtained after IVT of unbound File DNA, File DNA adsorbed to caSDC, and of DNA obtained after direct sequencing of File DNA from synthesis provider. Plotted RNA samples were processed with Oxford nanopore sequencing (ONT) and DNA samples were processed with Illumina Sequencing (Illumina). Each plotted value represents the arithmetic mean, and error bars represent the standard deviation of three independent IVT reactions. Statistics was calculated using One-Way ANOVA with Tukey–Kramer post-hoc for panel A and B. a p = 2.71×10−3, b p = 1.56×10−10, c p = 3.58×10−4, d p = 1.59×10−2, e p = 8.14×10−1, f p = 1.52×10−10, g p = 1.50×10−10, h p = 1.55×10−10.
Extended Data Fig. 5 Implementation of addressable in-storage computation.
A) Schematic of computation rules for Puzzle1. The payload of each oligo in the DNA library is divided into nine sections, with each section representing the a specific position on the puzzle board. Each position is composed of a specific 20 nt DNA sequence (bit sequence). Combination of these oligos completes the full starting configuration for Puzzle1. In computation, short DNA oligos are used as nucleic acid operators to hybridize to specific puzzle RNA strands which contain information violates puzzle rules. This process triggers endonuclease activities and leaves behind RNA strands representing correct puzzle solutions. These remaining strands are purified and retained for downstream processes and NGS analysis. B) Schematic of correct solutions for Puzzle1, Puzzle2, and Puzzle3, as well as the intentionally incorrect solution for Puzzle3.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–5, Methods and References.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download ZIP )
Unprocessed images.
Source Data Fig. 2 (download ZIP )
Unprocessed images.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download ZIP )
Unprocessed images.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, K.N., Volkel, K., Cao, C. et al. A primordial DNA store and compute engine. Nat. Nanotechnol. 19, 1654–1664 (2024). https://doi.org/10.1038/s41565-024-01771-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41565-024-01771-6
This article is cited by
-
Advances and challenges in non-canonical nucleic acids data storage
Nature Communications (2026)
-
Functions and applications of enzymes in nucleic acid nanotechnology
Journal of Nanobiotechnology (2025)
-
Synthetic nucleic-acid droplets: a bioprogramming platform for designer microliquids
Polymer Journal (2025)
-
The coming wave of confluent biosynthetic, bioinformational and bioengineering technologies
Nature Communications (2025)
-
Deciphering the molecular mechanisms of FET fusion oncoprotein–DNA hollow co-condensates
Nature Communications (2025)