Abstract
Microtubules are hollow cylindrical cytoskeletal polymers of laterally associated protofilaments that contain head-to-tail aligned ɑ/β-tubulin heterodimers. Although the exposed microtubule exterior is readily accessible to proteins, the mechanism governing the accessibility of the confined microtubule lumen to luminal particles remains unknown. Here we show that certain tubulin family proteins (isotypes) facilitate luminal accessibility because of the mechanical properties and lateral interactions that they confer to the microtubules. We characterized the microtubules reconstituted from defined compositions of Caenorhabditis elegans tubulin isotypes. These tubulin isotypes form microtubules with comparable protofilament numbers but different luminal accessibility. We further revealed the role of tubulin isotypes in regulating the strength of inter-protofilament lateral interactions, which determines luminal accessibility through the mechanosensitivity of reversible protofilament separation. Deformation of the microtubule lattice, which generates stresses exceeding the strength of the lateral interactions, creates gaps between adjacent protofilaments, enhancing the accessibility of the lumen. Together, our findings uncovered the tubulin isotype-dependent mechanical plasticity that confers force sensitivity to the microtubule lattice and modulates the energy barrier for luminal proteins to access the lumen.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data generated in this study are available from the corresponding authors upon request.
Code availability
The codes for quantifying ATAT-2 dissociation rate on curved microtubules are available via GitHub at https://github.com/JacksonYe61/code-for-measure-time-laps-intensity-of-curved-microtubule-in-Fiji.
References
Nogales, E., Whittaker, M., Milligan, R. A. & Downing, K. H. High-resolution model of the microtubule. Cell 96, 79–88 (1999).
Bodakuntla, S., Jijumon, A. S., Villablanca, C., Gonzalez-Billault, C. & Janke, C. Microtubule-associated proteins: structuring the cytoskeleton. Trends Cell Biol. 29, 804–819 (2019).
Akhmanova, A. & Kapitein, L. C. Mechanisms of microtubule organization in differentiated animal cells. Nat. Rev. Mol. Cell Biol. 23, 541–558 (2022).
Ichikawa, M. & Bui, K. H. Microtubule inner proteins: a meshwork of luminal proteins stabilizing the doublet microtubule. Bioessays 40, 1700209 (2018).
Ludueña, R. F. & Banerjee, A. in The Role of Microtubules in Cell Biology, Neurobiology, and Oncology Cancer Drug Discovery and Development (ed. Fojo, T.) Ch. 6 (Humana, 2008).
Dodd, D. O. et al. Ciliopathy patient variants reveal organelle-specific functions for TUBB4B in axonemal microtubules. Science 384, eadf5489 (2024).
Strassel, C. et al. An essential role for ɑ4A-tubulin in platelet biogenesis. Life Sci. Alliance https://doi.org/10.26508/lsa.201900309 (2019).
Hoff, K. J., Neumann, A. J. & Moore, J. K. The molecular biology of tubulinopathies: understanding the impact of variants on tubulin structure and microtubule regulation. Front. Cell. Neurosci. 16, 1023267 (2022).
Burton, P. R. Luminal material in microtubules of frog olfactory axons: structure and distribution. J. Cell Biol. 99, 520–528 (1984).
Tsuji, C. & Dodding, M. P. Lumenal components of cytoplasmic microtubules. Biochem. Soc. Trans. 50, 1953–1962 (2022).
Bounoutas, A., O’Hagan, R. & Chalfie, M. The multipurpose 15-protofilament microtubules in C. elegans have specific roles in mechanosensation. Curr. Biol. 19, 1362–1367 (2009).
Chalfie, M. & Thomson, J. N. Structural and functional diversity in the neuronal microtubules of Caenorhabditis elegans. J. Cell Biol. 93, 15–23 (1982).
Fukushige, T. et al. MEC-12, an alpha-tubulin required for touch sensitivity in C. elegans. J. Cell Sci. 112, 395–403 (1999).
Cueva, J. G., Hsin, J., Huang, K. C. & Goodman, M. B. Posttranslational acetylation of ɑ-tubulin constrains protofilament number in native microtubules. Curr. Biol. 22, 1066–1074 (2012).
Topalidou, I. et al. Genetically separable functions of the MEC-17 tubulin acetyltransferase affect microtubule organization. Curr. Biol. 22, 1057–1065 (2012).
Morley, S. J. et al. Acetylated tubulin is essential for touch sensation in mice. eLife https://doi.org/10.7554/eLife.20813 (2016).
Savage, C. et al. mec-7 is a beta-tubulin gene required for the production of 15-protofilament microtubules in Caenorhabditis elegans. Genes Dev. 3, 870–881 (1989).
Ward, I. M. & Sweeney, J. Mechanical Properties of Solid Polymers (Wiley, 2013).
Akella, J. S. et al. MEC-17 is an ɑ-tubulin acetyltransferase. Nature 467, 218–222 (2010).
Shida, T., Cueva, J. G., Xu, Z., Goodman, M. B. & Nachury, M. V. The major ɑ-tubulin K40 acetyltransferase ɑTAT1 promotes rapid ciliogenesis and efficient mechanosensation. Proc. Natl Acad. Sci. USA 107, 21517–21522 (2010).
Coombes, C. et al. Mechanism of microtubule lumen entry for the α-tubulin acetyltransferase enzyme αTAT1. Proc. Natl Acad. Sci. USA 113, E7176–E7184 (2016).
Szyk, A. et al. Molecular basis for age-dependent microtubule acetylation by tubulin acetyltransferase. Cell 157, 1405–1415 (2014).
Odde, D. Diffusion inside microtubules. Eur. Biophys. J. 27, 514–520 (1998).
Xu, Z. et al. Microtubules acquire resistance from mechanical breakage through intralumenal acetylation. Science 356, 328–332 (2017).
Webster, D. R. & Borisy, G. G. Microtubules are acetylated in domains that turn over slowly. J. Cell Sci. 92, 57–65 (1989).
Seetharaman, S. et al. Microtubules tune mechanosensitive cell responses. Nat. Mater. 21, 366–377 (2022).
Chretien, D. & Wade, R. H. New data on the microtubule surface lattice. Biol. Cell 71, 161–174 (1991).
Desai, A. & Mitchison, T. J. Microtubule polymerization dynamics. Annu. Rev. Cell Dev. Biol. 13, 83–117 (1997).
Ti, S. C., Alushin, G. M. & Kapoor, T. M. Human β-tubulin isotypes can regulate microtubule protofilament number and stability. Dev. Cell 47, 175–190.e175 (2018).
Luo, J. et al. Tubulin acetyltransferases access and modify the microtubule luminal K40 residue through anchors in taxane-binding pockets. Nat. Struct. Mol. Biol. 32, 358–368 (2025).
Vemu, A. et al. Severing enzymes amplify microtubule arrays through lattice GTP-tubulin incorporation. Science https://doi.org/10.1126/science.aau1504 (2018).
Roll-Mecak, A. & McNally, F. J. Microtubule-severing enzymes. Curr. Opin. Cell Biol. 22, 96–103 (2010).
Fu, G. et al. Integrated regulation of tubulin tyrosination and microtubule stability by human ɑ-tubulin isotypes. Cell Rep. 42, 112653 (2023).
Memet, E. et al. Microtubules soften due to cross-sectional flattening. eLife https://doi.org/10.7554/eLife.34695 (2018).
Hou, X., Cao, S., Rong, Q., Zheng, W. & Li, G. Effects of steel fiber and strain rate on the dynamic compressive stress-strain relationship in reactive powder concrete. Constr. Build. Mater. 170, 570–581 (2018).
Miyoshi, T. et al. Enhancement of energy absorption in a closed-cell aluminum by the modification of cellular structures. Scr. Mater. 41, 1055–1060 (1999).
Brazier, L. G. On the flexure of thin cylindrical shells and other ‘thin’ sections. Proc. R. Soc. A 116, 11 (1927).
Gittes, F., Mickey, B., Nettleton, J. & Howard, J. Flexural rigidity of microtubules and actin filaments measured from thermal fluctuations in shape. J. Cell Biol. 120, 923–934 (1993).
Needleman, D. J. et al. Synchrotron X-ray diffraction study of microtubules buckling and bundling under osmotic stress: a probe of interprotofilament interactions. Phys. Rev. Lett. 93, 198104 (2004).
Kasas, S. et al. Mechanical properties of microtubules explored using the finite elements method. ChemPhysChem 5, 252–257 (2004).
Kis, A. et al. Nanomechanics of microtubules. Phys. Rev. Lett. 89, 248101 (2002).
Pampaloni, F. & Florin, E. L. Microtubule architecture: inspiration for novel carbon nanotube-based biomimetic materials. Trends Biotechnol. 26, 302–310 (2008).
Pampaloni, F. et al. Thermal fluctuations of grafted microtubules provide evidence of a length-dependent persistence length. Proc. Natl Acad. Sci. USA 103, 10248–10253 (2006).
Taute, K. M., Pampaloni, F., Frey, E. & Florin, E. L. Microtubule dynamics depart from the wormlike chain model. Phys. Rev. Lett. 100, 028102 (2008).
Kuo, Y. W., Mahamdeh, M., Tuna, Y. & Howard, J. The force required to remove tubulin from the microtubule lattice by pulling on its ɑ-tubulin C-terminal tail. Nat. Commun. 13, 3651 (2022).
Ward, A. et al. Solid friction between soft filaments. Nat. Mater. 14, 583–588 (2015).
Hawkins, T., Mirigian, M., Selcuk Yasar, M. & Ross, J. L. Mechanics of microtubules. J. Biomech. 43, 23–30 (2010).
Kononova, O. et al. Tubulin bond energies and microtubule biomechanics determined from nanoindentation in silico. J. Am. Chem. Soc. 136, 17036–17045 (2014).
Jiang, N., Bailey, M. E., Burke, J., Ross, J. L. & Dima, R. I. Modeling the effects of lattice defects on microtubule breaking and healing. Cytoskeleton 74, 3–17 (2017).
Friedmann, D. R., Aguilar, A., Fan, J., Nachury, M. V. & Marmorstein, R. Structure of the ɑ-tubulin acetyltransferase, ɑTAT1, and implications for tubulin-specific acetylation. Proc. Natl Acad. Sci. USA 109, 19655–19660 (2012).
Taschner, M., Vetter, M. & Lorentzen, E. Atomic resolution structure of human ɑ-tubulin acetyltransferase bound to acetyl-CoA. Proc. Natl Acad. Sci. USA 109, 19649–19654 (2012).
Mickey, B. & Howard, J. Rigidity of microtubules is increased by stabilizing agents. J. Cell Biol. 130, 909–917 (1995).
Lavaud, M., Salez, T., Louyer, Y., & Amarouchene, Y. Stochastic inference of surface-induced effects using Brownian motion. Phys. Rev. Res. 3, 6 (2021).
Felderhof, B. U. Effect of the wall on the velocity autocorrelation function and long-time tail of Brownian motion. J. Phys. Chem. B 109, 21406–21412 (2005).
Wang, B., Anthony, S. M., Bae, S. C. & Granick, S. Anomalous yet Brownian. Proc. Natl Acad. Sci. USA 106, 15160–15164 (2009).
Chechkin, A. V., Seno, F., Metzler, R. & Sokolov, I. M. Brownian yet non-Gaussian diffusion: from superstatistics to subordination of diffusing diffusivities. Phys. Rev. X 7, 021002 (2017).
Takahashi, K. et al. Suppression of flatness in circular tube-draw bending by applying side compression. Mater. Trans. 53, 870–874 (2012).
Bouzigues, C. I., Tabeling, P. & Bocquet, L. Nanofluidics in the Debye layer at hydrophilic and hydrophobic surfaces. Phys. Rev. Lett. 101, 114503 (2008).
Joly, L., Ybert, C. & Bocquet, L. Probing the nanohydrodynamics at liquid-solid interfaces using thermal motion. Phys. Rev. Lett. 96, 046101 (2006).
Mo, J., Simha, A. & Raizen, M. G. Brownian motion as a new probe of wettability. J. Chem. Phys. 146, 134707 (2017).
Otsuka, H., Nagasaki, Y. & Kataoka, K. Surface characterization of functionalized polylactide through the coating with heterobifunctional poly(ethylene glycol)/polylactide block copolymers. Biomacromolecules 1, 39–48 (2000).
Chretien, D., Metoz, F., Verde, F., Karsenti, E. & Wade, R. H. Lattice defects in microtubules: protofilament numbers vary within individual microtubules. J. Cell Biol. 117, 1031–1040 (1992).
Nishida, K. et al. Expression patterns and levels of all tubulin isotypes analyzed in GFP knock-in C. elegans strains. Cell Struct. Funct. 46, 51–64 (2021).
Lu, Y. M. & Zheng, C. The expression and function of tubulin isotypes in Caenorhabditis elegans. Front. Cell Dev. Biol. 10, 860065 (2022).
Wang, X. et al. Cryo-EM structure of cortical microtubules from human parasite Toxoplasma gondii identifies their microtubule inner proteins. Nat. Commun. 12, 3065 (2021).
Portran, D., Schaedel, L., Xu, Z., Thery, M. & Nachury, M. V. Tubulin acetylation protects long-lived microtubules against mechanical ageing. Nat. Cell Biol. 19, 391–398 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rice, S. et al. A structural change in the kinesin motor protein that drives motility. Nature 402, 778–784 (1999).
Smith, M. B. et al. Segmentation and tracking of cytoskeletal filaments using open active contours. Cytoskeleton 67, 693–705 (2010).
Maurer, S. P. et al. EB1 accelerates two conformational transitions important for microtubule maturation and dynamics. Curr. Biol. 24, 372–384 (2014).
Liew, K. M., Xiang, P. & Zhang, L. W. Mechanical properties and characteristics of microtubules: a review. Compos. Struct. 123, 98–108 (2015).
Ömer Civalek, Ç. D. Bending analysis of microtubules using nonlocal Euler-Bernoulli beam theory. Appl. Math. Modell. 35, 2053–2067 (2011).
COMSOL Multiphysics v.6 (COMSOL AB, 2021).
Acknowledgements
S.-C.T. acknowledges support from the Research Grants Council of Hong Kong under the General Research Fund (grant nos. 17122822 and 17118421; principal investigator S.-C.T.) and the Collaborative Research Fund (grant no. C7064-22GF; project coordinator S.-C.T.). Y.L. acknowledges support from the Research Grants Council of Hong Kong under the General Research Fund (grant no. 17210520), the Health@InnoHK programme of the Innovation and Technology Commission of the Hong Kong SAR Government and the National Natural Science Foundation of China (grant no. 12272332). We thank T. M. Kapoor (Rockefeller University) for comments on this paper; J. Guo, M. Chen and H. Zhu (Imaging and Flow Cytometry Core, Center for PanorOmic Sciences, LKS Faculty of Medicine, University of Hong Kong) for support with fluorescence microscopes; Biological Cryo-EM Center at the Hong Kong University of Science and Technology (HKUST) for sample screening, grid preparation and data collection; and Y. Zhang for technical support. A donation from the Lo Kwee Seong Foundation generously supports the HKUST Cryo-EM Center.
Author information
Authors and Affiliations
Contributions
Y.Y., Z.H., Y.L. and S.-C.T. conceptualized the study. Y.Y. and J.L. generated the protein materials used in this study. Y.Y. performed and analysed the optical tweezer experiments. Y.Y. and J.L. performed and analysed the TIRF microscopy experiments. Y.Y. performed and analysed the TIRF-based single-molecule experiments. Y.Y., Z.H., Y.L. and S.-C.T. developed the 3D finite element model of the anisotropic microtubule lattice. W.H.L. and Y.Z. collected and analysed the cryo-EM data. Z.L. and X.D.L. characterized the proteins by means of mass spectrometry. Y.L. and S.-C.T. supervised the study. Y.Y., Z.H., Y.L. and S.-C.T. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Physics thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification and characterization of recombinant proteins.
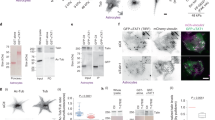
(a) Reversed-phase LC-MS spectra of purified TBA-2/TBB-2. See Luo et al. for the LC-MS spectra of purified MEC-12/MEC-730. (b) Representative Coomassie blue-stained SDS-PAGE, immunoblot analyses, and rhodamine fluorescence imaging of purified recombinant proteins. See Luo et al. for the Coomassie blue-stained SDS-PAGE, immunoblot analyses, and rhodamine fluorescence imaging of purified MEC-12/MEC-730. We performed Coomassie blue-stained SDS–PAGE analysis for every batch of protein that we purified. (c and d) Representative TIRF microscopy images showing the incorporation of FITC-labeled tubulin (green) into defects of the rhodamine-labeled TBA-2/TBB-2 (c) or MEC-12/MEC-7 (d) microtubule lattice (red) before and after the treatment of MBP-tagged human katanin P60. Scale bars: 5 µm. Similar results were observed in two independent experiments. (e) Representative cryo-EM images of GMPCPP-stabilized TBA-2/TBB-2 (top) and MEC-12/MEC-7 (bottom) microtubules showing no visible breakage or defects. Similar results were observed in 596 (TBA-2/TBB-2) and 1040 (MEC-12/MEC-7) micrographs from one cryo-EM data collection. Scale bars: 50 nm. (f and g) The FITC fluorescence intensity on per micrometer GMPCPP-stabilized TBA-2/TBB-2 microtubules (0 nM katanin: 1143 ± 374 (a.u.), n=88; 10 nM katanin: 3732 ± 1688 (a.u.), n=90; 50 nM katanin: 4780 ± 1934 (a.u.), n=78, mean ± SD are indicated) (f) and MEC-12/MEC-7 microtubules (0 nM katanin: 1515 ± 678 (a.u.), n=71; 10 nM katanin: 3077 ± 1232 (a.u.), n=73; 50 nM katanin: 5968 ± 2329 (a.u.), n=70, mean ± SD are indicated) (g) treated with different concentrations of MBP-tagged human katanin P60. The mean and SD are based on analyzed microtubules pooled from two independent experiments. The P values were calculated using a two-tailed unpaired Student’s t-test.
Extended Data Fig. 2 The distribution of ATAT-2 binding on TBA-2/TBB-2 and MEC-12/MEC-7 microtubules.
(a and b) Representative time-lapse images showing ATAT-2-GFP (green) binding to and dissociation from GMPCPP-stabilized TBA-2/TBB-2 (a) and MEC-12/MEC-7 (b) microtubules (red). The slow-dissociating GFP fluorescence signal at the microtubule ends and the shaft is indicated by white arrows and arrowheads, respectively. Scale bars: 3 µm. Similar results were observed in at least three independent experiments. (c) Heatmaps of the time-dependent decrease of the ATAT-2-GFP fluorescence intensity on a GMPCPP-stabilized TBA-2/TBB-2 or MEC-12/MEC-7 microtubule ~30 sec after perfusing the TIRF chamber with the ATAT-2-free buffer. The arrows indicate the positions of microtubule ends. Similar results were observed in at least three independent experiments. (d) The decay of the normalized GFP-tagged ATAT-2 fluorescence intensity on GMPCPP-stabilized TBA-2/TBB-2 (black, n=87) or MEC-12/MEC-7 (green, n=82) microtubules upon perfusion with the ATAT-2-free buffer. The GFP fluorescence intensity was normalized against that from the first image of the time-lapse started ~30 sec after perfusing the TIRF chamber with the ATAT-2-free buffer. The solid lines show the single exponential decay fitted to the data points (mean ± SEM) from three independent experiments. The lifetimes (τ) are indicated. (e) The ‘averaged’ normalized rhodamine fluorescence intensity at the end region of TBA-2/TBB-2 (n=72) and MEC-12/MEC-7 (n=67) microtubules from at least three independent experiments. The lengths of the microtubule end regions are indicated. (f) Schematic of the end and shaft regions of a microtubule. (g) The portions of the slow-dissociating ATAT-2-GFP on different regions of microtubules at ~30 sec after the TIRF chamber was perfused with the ATAT-2-free buffer. The mean and SD are based on analyzing TBA-2/TBB-2 (n=147) and MEC-12/MEC-7 (n=123) microtubules from at least three independent experiments. (h) The landing frequency of ATAT-2-GFP on TBA-2/TBB-2 (black bar, n=3838) and MEC-12/MEC-7 (green bar, n=2827) microtubules. Assuming a Poisson distribution, the SEM of the landing frequency was calculated as (observed frequency)0.5/ (imaging time × total microtubule length)0.5. The mean and SEM are based on data from at least three independent experiments. The P values were calculated using a two-tailed unpaired Student’s t-test. (i) The exponential decay of the numbers of detectable ATAT-2-GFP molecules (n=1130) on the coverslip surface under the imaging conditions of the single-molecule assays. The data are from two independent experiments. The measured data (black dots) were fitted with a single exponential distribution (red line) to determine the photobleaching rate (kb). (j) Distribution of the duration of ATAT-2 single molecules on GMPCPP-stabilized TBA-2/TBB-2 (black, n=3838) and MEC-12/MEC-7 (green, n=2827) microtubules from at least three independent experiments. tTBA-2/TBB-2 and tMEC-12/MEC-7 are the average duration (mean ± SD).
Extended Data Fig. 3 Microtubules can sustain cycles of compression without fatigue.
(a-d) The compression (blue) and relaxation (red) force-strain curves (a and c) and the strains with maximum energy absorption efficiency (b and d) of one GMPCPP-stabilized TBA-2/TBB-2 (a and b) or MEC-12/MEC-7 (c and d) microtubule after compression-relaxation cycles. Similar results were observed in at least three independent experiments. (e and f) Schematic of the line scan analysis (e) and the corresponding intensity profiles (f) of TBA-2/TBB-2 microtubule cross-sections (green lines) before compression and at the maximum strain in the optical trap experiments. (g) Intensity profiles of the cross-section line scans from another seven TBA-2/TBB-2 microtubules at the maximum strain. (h and i) Schematic of the line scan analysis (h) and the corresponding intensity profiles (i) of MEC-12/MEC-7 microtubule cross-sections (green lines) before compression and at the maximum strain in the optical trap experiments. (j) Intensity profiles of the cross-section line scans from another seven MEC-12/MEC-7 microtubules at the maximum strain. The background threshold (red dashed lines) was determined from the mean (µ, blue dashed lines) and standard deviation (σ) of the fluorescence intensity of a 40-pixel x 40-pixel area (red squares). We used the fluorescence intensity half-heights (gray dashed line) to determine the width of peaks from microtubule cross-section line scans. The peak width of microtubules before compression was indicated (blue double arrow). All eight GMPCPP-stabilized TBA-2/TBB-2 microtubules remain intact at the maximum strain as (i) the fluorescence intensities are larger than the background threshold (red dashed lines) and (ii) the peak widths of microtubules at the maximum strain are larger than 90% of the width before compression (blue double arrows). All eight GMPCPP-stabilized MEC-12/MEC-7 microtubules are splayed at the maximum strain as the fluorescence intensities are lower than the background threshold (red dashed lines). Scale bars: 3 µm. The data are from at least three independent experiments.
Extended Data Fig. 4 Representative tensile force-strain curves of microtubules.
(a-f) Representative tensile force-strain curves of GMPCPP-stabilized TBA-2/TBB-2 (a-c) and MEC-12/MEC-7 (d-f) microtubules with different lengths, showing the disconnection and re-connection of NeutrAvidin-coated beads on microtubules. Similar results were observed in at least three independent experiments.
Extended Data Fig. 5 The 3D finite element method-based computational model of microtubules.
(a) Rotation (or orientation change) of protofilament j will lead to non-zero displacement δi, j in its local coordinate system (upper row). The resulting moment Mi, j restraining the relative rotation between two protofilaments is effectively the same as that by a rotational spring (lower row). p is the initial distance between the long axes of two neighboring protofilaments. θ is the angle of rotation of protofilament j. (b and c) Simulation results of the cross-sectional flattening and protofilament separation of TBA-2/TBB-2 (b) and MEC-12/MEC-7 (c) microtubules. The heatmap indicates the maximum normal stress loaded on each protofilament in the microtubule lattice. (d) The largest protofilament separation distance in the simulated TBA-2/TBB-2 (black) and MEC12/MEC-7 (green) microtubules under compressive strains from 0 % to 50 % with 5% intervals. (e) The enlarged view of (d) highlights the <10 nm protofilament separation in TBA-2/TBB-2 microtubules at the examined strains.
Extended Data Fig. 6 The luminal accessibility of GMPCPP-stabilized TBA-2/TBB-2 microtubules increases with compressive strains.
Heatmaps showing the initial binding and the dissociation of GFP-tagged ATAT-2 over time on GMPCPP-stabilized TBA-2/TBB-2 microtubules experiencing 0% to ~44% compressive strain (n = 16 microtubules from at least three independent experiments).
Extended Data Fig. 7 The amount of luminal ATAT-2-GFP on GMPCPP-stabilized MEC-12/MEC-7 microtubules decreases with compressive strains.
Heatmaps showing the initial binding and the decrease of GFP-tagged ATAT-2 over time on GMPCPP-stabilized MEC-12/MEC-7 microtubules experiencing 0% to ~44% compressive strain (n = 10 microtubules from at least three independent experiments).
Extended Data Fig. 8 The luminal accessibility of GDPstateTBA-2/TBB-2 microtubules increases with compressive strains.
Heatmaps showing the initial binding and the dissociation of GFP-tagged ATAT-2 over time on GDP-state TBA-2/TBB-2 microtubules experiencing 0% to ~44% compressive strain (n = 11 microtubules from at least three independent experiments).
Extended Data Fig. 9 Examining the lattice mechanical plasticity by bending microtubules proximal to the coverslip surface.
(a) Representative kymographs showing the rapid depolymerization of GDP-state MEC-12/MEC-7 microtubules after the GTP-bound MEC-17/MEC-7 tubulin was replaced with the buffer containing 20 µM GMPCPP-bound MEC-12/MEC-7 tubulin. Microtubule seeds and dynamic microtubule extensions were labeled with 3% and 14% rhodamine-labeled tubulin, respectively. Vertical scale bars: 5 µm; horizontal scale bars: 10 s. Similar results were observed in at least three independent experiments. (b and c) Schematic of the line scan analysis (b) and the corresponding intensity profiles (c) of TBA-2/TBB-2 microtubule cross-sections (green lines) before compression and at the maximum strain in the surface bending experiments. (d and e) Schematic of the line scan analysis (d) and the corresponding intensity profiles (e) of MEC-12/MEC-7 microtubule cross-sections (green lines) before compression and at the maximum strain in the surface bending experiments. (f and g) Representative intensity profiles cross-section line scans from two GMPCPP-stabilized TBA-2/TBB-2 (f) and MEC-12/MEC-7 (g) microtubules under compression in the surface bending experiments. The threshold of the background signal (red dashed line), the fluorescence intensity half-height of a microtubule (gray dashed line), and the peak width of microtubules before compression (blue double arrow) are indicated. (h) The percentage of splayed and non-splayed GMPCPP-stabilized MEC-12/MEC-7 and TBA-2/TBB-2 microtubule extensions while being bent on the coverslip. Data are from at least three independent experiments.
Extended Data Fig. 10 The sequence-structure analysis of the interacting interface between ATAT-2 and microtubules.
(a and b) Primary sequence alignment of MEC-12 and TBA-2 (a) and MEC-7 and TBB-2 (b). The residues of MEC-12 and MEC-7 that interact with ATAT-2 are highlighted in bold font. The ATAT-2 interacting residues divergent between these tubulin isotypes are italicized and underlined. (c) The ATAT-2 interacting residues are mapped onto the luminal surface of MEC-12/MEC-7 microtubules (PDB 8Y9F). The interacting residues that are identical and divergent between MEC-12/TBA-2 and MEC-7/TBB-2 are highlighted in pink and orange, respectively. The taxane-binding pockets and the α-tubulin H1-S2 loop (the K40 loop) are indicated.
Supplementary information
Supplementary Video 1 (download AVI )
A GMPCPP-stabilized TBA-2/TBB-2 microtubule undergoes cycles of compression and relaxation. Scale bar, 5 µm.
Supplementary Video 2 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 3 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 4 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 5 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 6 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 7 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 8 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 9 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation. The cross-sectional line scans of each MEC-12/MEC-7 microtubule at the maximum strain are shown in Extended Data Fig. 3. Scale bar, 5 µm.
Supplementary Video 10 (download AVI )
A GMPCPP-stabilized TBA-2/TBB-2 microtubule under tensile forces. Scale bar, 5 µm.
Supplementary Video 11 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule under tensile forces. Scale bar, 5 µm.
Supplementary Video 12 (download AVI )
A GMPCPP-stabilized TBA-2/TBB-2 microtubule undergoes cycles of compression and relaxation when displaced ~100 nm from the coverslip surface. Scale bar, 5 µm.
Supplementary Video 13 (download AVI )
A GMPCPP-stabilized MEC-12/MEC-7 microtubule undergoes cycles of compression and relaxation when displaced ~100 nm from the coverslip surface. Scale bar, 5 µm.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ye, Y., Hao, Z., Luo, J. et al. Tubulin isotypes of C. elegans harness the mechanosensitivity of the lattice for microtubule luminal accessibility. Nat. Phys. 21, 1420–1430 (2025). https://doi.org/10.1038/s41567-025-02983-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41567-025-02983-w