Abstract
The centriole is an evolutionarily conserved microtubule-bearing organelle with a striking 9-fold symmetrical architecture that is crucial for fundamental cellular processes in eukaryotes, including polarity, signalling and motility. The last two decades have witnessed important progress in uncovering the molecular architecture and assembly principles governing centriole biogenesis, which are discussed in this Review, with a focus on human cells. Recently developed advanced microscopy approaches have increased our understanding of the mechanisms governing centriole biogenesis, from initiating the assembly process to forming a full-fledged organelle. Some two billion years after its emergence during evolution, and more than 100 years after its identification by pioneering scientists, these advances make the centriole ripe for another era of discoveries.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
van Beneden, E. & Neyt, A. Nouvelle recherches sur la fécondation et la division mitosique chez l’ascaride mégalocéphale. Bull. Acad. R. Belg. 14, 215–295 (1887).
Boveri, T. Ueber den antheil des spermatozoon an der theilung des eies. Sitzungsber. Ges. Morph. Physiol. München 3, 151–164 (1887).
Boveri, T. Zellen-Studien: Heft 4, Ueber Die Natur Der Centrosomen (Fischer, 1900).
Carvalho-Santos, Z. et al. Stepwise evolution of the centriole-assembly pathway. J. Cell Sci. 123, 1414–1426 (2010).
Hodges, M. E., Scheumann, N., Wickstead, B., Langdale, J. A. & Gull, K. Reconstructing the evolutionary history of the centriole from protein components. J. Cell Sci. 123, 1407–1413 (2010).
Bornens, M. & Azimzadeh, J. Origin and evolution of the centrosome. Adv. Exp. Med. Biol. 607, 119–129 (2007).
Zhao, H., Khan, Z. & Westlake, C. J. Ciliogenesis membrane dynamics and organization. Semin. Cell Dev. Biol. 133, 20–31 (2023).
Kumar, D. & Reiter, J. How the centriole builds its cilium: of mothers, daughters, and the acquisition of appendages. Curr. Opin. Struct. Biol. 66, 41–48 (2021).
Hilgendorf, K. I., Myers, B. R. & Reiter, J. F. Emerging mechanistic understanding of cilia function in cellular signalling. Nat. Rev. Mol. Cell Biol. 25, 555–573 (2024).
Vasquez-Limeta, A. & Loncarek, J. Human centrosome organization and function in interphase and mitosis. Semin. Cell Dev. Biol. 117, 30–41 (2021).
Bornens, M. The centrosome in cells and organisms. Science 335, 422–426 (2012).
Wu, J. & Akhmanova, A. Microtubule-organizing centers. Annu. Rev. Cell Dev. Biol. 33, 51–75 (2017).
Heald, R., Tournebize, R., Habermann, A., Karsenti, E. & Hyman, A. Spindle assembly in Xenopus egg extracts: respective roles of centrosomes and microtubule self-organization. J. Cell Biol. 138, 615–628 (1997).
Pintard, L. & Bowerman, B. Mitotic cell division in Caenorhabditis elegans. Genetics 211, 35–73 (2019).
Dutcher, S. K. & O’Toole, E. T. The basal bodies of Chlamydomonas reinhardtii. Cilia 5, 18 (2016).
Lattao, R., Kovács, L. & Glover, D. M. The centrioles, centrosomes, basal bodies, and cilia of Drosophila melanogaster. Genetics 206, 33–53 (2017).
Bayless, B. A., Galati, D. F. & Pearson, C. G. Tetrahymena basal bodies. Cilia 5, 1 (2015).
Prosser, S. L. & Pelletier, L. Centriolar satellite biogenesis and function in vertebrate cells. J. Cell Sci. 133, jcs239566 (2020).
Dang, H. & Schiebel, E. Emerging roles of centrosome cohesion. Open. Biol. 12, 220229 (2022).
Regolini, M. The centrosome as a geometry organizer. Results Probl. Cell Differ. 67, 253–276 (2019).
Bernhard, W. & De Harven, E. Presence in certain mammalian cells of an organoid probably of centriole nature; electron microscopy. C. R. Hebd. Seances Acad. Sci. 242, 288–290 (1956).
Bernhard, W. & De Harven, E. Electron microscopic study of the ultrastructure of centrioles in vertebra. Z. Zellforsch. Mikrosk. Anat. 45, 378–398 (1956).
Satir, P. Chirality of the cytoskeleton in the origins of cellular asymmetry. Philos. Trans. R. Soc. Lond. B Biol. Sci. 371, 20150408 (2016).
Guichard, P. et al. Native architecture of the centriole proximal region reveals features underlying its 9-fold radial symmetry. Curr. Biol. 23, 1620–1628 (2013).
Li, S., Fernandez, J. J., Marshall, W. F. & Agard, D. A. Three-dimensional structure of basal body triplet revealed by electron cryo-tomography. EMBO J. 31, 552–562 (2012).
Guichard, P., Chretien, D., Marco, S. & Tassin, A. M. Procentriole assembly revealed by cryo-electron tomography. EMBO J. 29, 1565–1572 (2010).
Klena, N. et al. Architecture of the centriole cartwheel-containing region revealed by cryo-electron tomography. EMBO J. 39, e106246 (2020).
Greenan, G. A., Keszthelyi, B., Vale, R. D. & Agard, D. A. Insights into centriole geometry revealed by cryotomography of doublet and triplet centrioles. eLife 7, e36851 (2018).
Nazarov, S. et al. Novel features of centriole polarity and cartwheel stacking revealed by cryo-tomography. EMBO J. 39, e106249 (2020).
Hirono, M. Cartwheel assembly. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, rstb.2013.0458 (2014).
Guichard, P., Hamel, V. & Gönczy, P. The rise of the cartwheel: seeding the centriole organelle. Bioessays 40, e1700241 (2018).
Nigg, E. A. & Stearns, T. The centrosome cycle: centriole biogenesis, duplication and inherent asymmetries. Nat. Cell Biol. 13, 1154–1160 (2011).
Firat-Karalar, E. N. & Stearns, T. The centriole duplication cycle. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130460 (2014).
Kochanski, R. S. & Borisy, G. G. Mode of centriole duplication and distribution. J. Cell Biol. 110, 1599–1605 (1990). This work demonstrates, through microinjection of labelled tubulin followed by detection of newly incorporated protein, that centriole duplication is conservative.
Kuriyama, R., Dasgupta, S. & Borisy, G. G. Independence of centriole formation and initiation of DNA synthesis in Chinese hamster ovary cells. Cell Motil. Cytoskeleton 6, 355–362 (1986).
Manandhar, G., Schatten, H. & Sutovsky, P. Centrosome reduction during gametogenesis and its significance. Biol. Reprod. 72, 2–13 (2005).
Delattre, M. & Gönczy, P. The arithmetic of centrosome biogenesis. J. Cell Sci. 117, 1619–1630 (2004).
Mahjoub, M. R., Nanjundappa, R. & Harvey, M. N. Development of a multiciliated cell. Curr. Opin. Cell Biol. 77, 102–105 (2022).
Lyu, Q., Li, Q., Zhou, J. & Zhao, H. Formation and function of multiciliated cells. J. Cell Biol. 223, e202307150 (2024).
Al Jord, A. et al. Centriole amplification by mother and daughter centrioles differs in multiciliated cells. Nature 516, 104–107 (2014).
Choksi, S. P. et al. An alternative cell cycle coordinates multiciliated cell differentiation. Nature 630, 214–221 (2024).
Weier, A.-K. et al. Multiple centrosomes enhance migration and immune cell effector functions of mature dendritic cells. J. Cell Biol. 221, e202107134 (2022).
LeGuennec, M., Klena, N., Aeschlimann, G., Hamel, V. & Guichard, P. Overview of the centriole architecture. Curr. Opin. Struct. Biol. 66, 58–65 (2021).
Jakobsen, L. et al. Novel asymmetrically localizing components of human centrosomes identified by complementary proteomics methods. EMBO J. 30, 1520–1535 (2011).
Andersen, S. S. Molecular characteristics of the centrosome. Int. Rev. Cytol. 187, 51–109 (1999). This work is the initial protein correlation profiling proteomics study identifying the compendium of proteins in human centrosomes.
Kilburn, C. L. et al. New Tetrahymena basal body protein components identify basal body domain structure. J. Cell Biol. 178, 905–912 (2007).
Keller, L. C., Romijn, E. P., Zamora, I., Yates, J. R. & Marshall, W. F. Proteomic analysis of isolated Chlamydomonas centrioles reveals orthologs of ciliary-disease genes. Curr. Biol. 15, 1090–1098 (2005).
Hamel, V. et al. Identification of Chlamydomonas central core centriolar proteins reveals a role for human WDR90 in ciliogenesis. Curr. Biol. 27, 2486–2498 (2017).
Bauer, M., Cubizolles, F., Schmidt, A. & Nigg, E. A. Quantitative analysis of human centrosome architecture by targeted proteomics and fluorescence imaging. EMBO J. 35, 2152–2166 (2016).
Gupta, G. D. et al. A dynamic protein interaction landscape of the human centrosome–cilium interface. Cell 163, 1484–1499 (2015).
Sugioka, K. et al. Centriolar SAS-7 acts upstream of SPD-2 to regulate centriole assembly and pericentriolar material formation. eLife 6, e20353 (2017).
Pelletier, L. et al. The Caenorhabditis elegans centrosomal protein SPD-2 is required for both pericentriolar material recruitment and centriole duplication. Curr. Biol. 14, 863–873 (2004).
Kemp, C. A., Kopish, K. R., Zipperlen, P., Ahringer, J. & O’Connell, K. F. Centrosome maturation and duplication in C. elegans require the coiled-coil protein SPD-2. Dev. Cell 6, 511–523 (2004).
O’Connell, K. F. et al. The C. elegans zyg-1 gene encodes a regulator of centrosome duplication with distinct maternal and paternal roles in the embryo. Cell 105, 547–558 (2001).
Dammermann, A. et al. Centriole assembly requires both centriolar and pericentriolar material proteins. Dev. Cell 7, 815–829 (2004).
Leidel, S., Delattre, M., Cerutti, L., Baumer, K. & Gönczy, P. SAS-6 defines a protein family required for centrosome duplication in C. elegans and in human cells. Nat. Cell Biol. 7, 115–125 (2005).
Delattre, M. et al. Centriolar SAS-5 is required for centrosome duplication in C. elegans. Nat. Cell Biol. 6, 656–664 (2004).
Leidel, S. & Gönczy, P. SAS-4 is essential for centrosome duplication in C elegans and is recruited to daughter centrioles once per cell cycle. Dev. Cell 4, 431–439 (2003).
Kirkham, M., Müller-Reichert, T., Oegema, K., Grill, S. & Hyman, A. A. SAS-4 is a C. elegans centriolar protein that controls centrosome size. Cell 112, 575–587 (2003).
Banterle, N. & Gönczy, P. Centriole biogenesis: from identifying the characters to understanding the plot. Annu. Rev. Cell Dev. Biol. 33, 23–49 (2017).
Goshima, G. et al. Genes required for mitotic spindle assembly in Drosophila S2 cells. Science 316, 417–421 (2007).
Balestra, F. R., Strnad, P., Fluckiger, I. & Gönczy, P. Discovering regulators of centriole biogenesis through siRNA-based functional genomics in human cells. Dev. Cell 25, 555–571 (2013).
Stemm-Wolf, A. J., O’Toole, E. T., Sheridan, R. M., Morgan, J. T. & Pearson, C. G. The SON RNA splicing factor is required for intracellular trafficking structures that promote centriole assembly and ciliogenesis. Mol. Biol. Cell 32, ar4 (2021).
Lau, L., Lee, Y. L., Sahl, S. J., Stearns, T. & Moerner, W. E. STED microscopy with optimized labeling density reveals 9-fold arrangement of a centriole protein. Biophys. J. 102, 2926–2935 (2012).
Sonnen, K. F., Schermelleh, L., Leonhardt, H. & Nigg, E. A. 3D-structured illumination microscopy provides novel insight into architecture of human centrosomes. Biol. Open. 1, 965–976 (2012).
Sillibourne, J. E. et al. Assessing the localization of centrosomal proteins by PALM/STORM nanoscopy. Cytoskeleton 68, 619–627 (2011).
Laporte, M. H. et al. Time-series reconstruction of the molecular architecture of human centriole assembly. Cell 187, 2158–2174 (2024). This work deploys ultrastructure expansion microscopy to uncover the precise distribution of 21 proteins in the centriole architecture.
Sir, J. H. et al. A primary microcephaly protein complex forms a ring around parental centrioles. Nat. Genet. 43, 1147–1153 (2011).
Lee, K. S., Park, J.-E., Il Ahn, J., Wei, Z. & Zhang, L. A self-assembled cylindrical platform for Plk4-induced centriole biogenesis. Open. Biol. 10, 200102 (2020).
Momotani, K., Khromov, A. S., Miyake, T., Stukenberg, P. T. & Somlyo, A. V. Cep57, a multidomain protein with unique microtubule and centrosomal localization domains. Biochem. J. 412, 265–273 (2008).
Wei, Z. et al. Requirement of the Cep57–Cep63 interaction for proper Cep152 recruitment and centriole duplication. Mol. Cell Biol. 40, e00535–19 (2020).
Lukinavicius, G. et al. Selective chemical crosslinking reveals a Cep57–Cep63–Cep152 centrosomal complex. Curr. Biol. 23, 265–270 (2013).
Kim, T.-S. et al. Molecular architecture of a cylindrical self-assembly at human centrosomes. Nat. Commun. 10, 1151 (2019).
Ito, K. K. et al. Cep57 and Cep57L1 maintain centriole engagement in interphase to ensure centriole duplication cycle. J. Cell Biol. 220, e202005153 (2021).
Zhao, H. et al. Cep57 and Cep57l1 function redundantly to recruit the Cep63–Cep152 complex for centriole biogenesis. J. Cell Sci. 133, jcs241836 (2020).
Zhao, H. et al. The Cep63 paralogue Deup1 enables massive de novo centriole biogenesis for vertebrate multiciliogenesis. Nat. Cell Biol. 15, 1434–1444 (2013).
Arquint, C. & Nigg, E. A. The PLK4–STIL–SAS-6 module at the core of centriole duplication. Biochem. Soc. Trans. 44, 1253–1263 (2016).
Chi, W., Wang, G., Xin, G., Jiang, Q. & Zhang, C. PLK4-phosphorylated NEDD1 facilitates cartwheel assembly and centriole biogenesis initiations. J. Cell Biol. 220, e202002151 (2021).
Haren, L. et al. NEDD1-dependent recruitment of the γ-tubulin ring complex to the centrosome is necessary for centriole duplication and spindle assembly. J. Cell Biol. 172, 505–515 (2006).
Zitouni, S. et al. CDK1 prevents unscheduled PLK4–STIL complex assembly in centriole biogenesis. Curr. Biol. 26, 1127–1137 (2016).
Kim, T.-S. et al. Hierarchical recruitment of Plk4 and regulation of centriole biogenesis by two centrosomal scaffolds, Cep192 and Cep152. Proc. Natl Acad. Sci. USA 110, E4849–E4857 (2013).
Sonnen, K. F., Gabryjonczyk, A. M., Anselm, E., Stierhof, Y. D. & Nigg, E. A. Human Cep192 and Cep152 cooperate in Plk4 recruitment and centriole duplication. J. Cell Sci. 126, 3223–3233 (2013).
Park, S.-Y. et al. Molecular basis for unidirectional scaffold switching of human Plk4 in centriole biogenesis. Nat. Struct. Mol. Biol. 21, 696–703 (2014).
Sullenberger, C., Kong, D., Avazpour, P., Luvsanjav, D. & Loncarek, J. Centrosomal organization of Cep152 provides flexibility in Plk4 and procentriole positioning. J. Cell Biol. 222, e202301092 (2023).
Gürkaşlar, H. K. & Hoffmann, I. Binding of CEP152 to PLK4 stimulates kinase activity to promote centriole assembly. Mol. Biol. Cell https://doi.org/10.1091/mbc.E24-12-0581 (2025).
Scott, P., Curinha, A., Gliech, C. & Holland, A. J. PLK4 self-phosphorylation drives the selection of a single site for procentriole assembly. J. Cell Biol. 222, e202301069 (2023).
Klebba, J. E. et al. Polo-like kinase 4 autodestructs by generating its Slimb-binding phosphodegron. Curr. Biol. 23, 2255–2261 (2013).
Cunha-Ferreira, I. et al. The SCF/Slimb ubiquitin ligase limits centrosome amplification through degradation of SAK/PLK4. Curr. Biol. 19, 43–49 (2009).
Rogers, G. C., Rusan, N. M., Roberts, D. M., Peifer, M. & Rogers, S. L. The SCF Slimb ubiquitin ligase regulates Plk4/Sak levels to block centriole reduplication. J. Cell Biol. 184, 225–239 (2009).
Holland, A. J., Lan, W., Niessen, S., Hoover, H. & Cleveland, D. W. Polo-like kinase 4 kinase activity limits centrosome overduplication by autoregulating its own stability. J. Cell Biol. 188, 191–198 (2010).
Cunha-Ferreira, I. et al. Regulation of autophosphorylation controls PLK4 self-destruction and centriole number. Curr. Biol. 23, 2245–2254 (2013).
Guderian, G., Westendorf, J., Uldschmid, A. & Nigg, E. A. Plk4 trans-autophosphorylation regulates centriole number by controlling βTrCP-mediated degradation. J. Cell Sci. 123, 2163–2169 (2010).
Wong, Y. L. et al. Reversible centriole depletion with an inhibitor of Polo-like kinase 4. Science 348, 1155–1160 (2015). This work reports the development of the selective and potent small molecule PLK4 inhibitor centrinone, which enables seamless inhibition of centriole formation.
Montenegro Gouveia, S. et al. PLK4 is a microtubule-associated protein that self-assembles promoting de novo MTOC formation. J. Cell. Sci. 132, jcs.219501 (2018).
Park, J.-E. et al. Phase separation of Polo-like kinase 4 by autoactivation and clustering drives centriole biogenesis. Nat. Commun. 10, 4959 (2019).
Yamamoto, S. & Kitagawa, D. Self-organization of Plk4 regulates symmetry breaking in centriole duplication. Nat. Commun. 10, 1810 (2019).
Hatch, E. M., Kulukian, A., Holland, A. J., Cleveland, D. W. & Stearns, T. Cep152 interacts with Plk4 and is required for centriole duplication. J. Cell Biol. 191, 721–729 (2010).
Moyer, T. C., Clutario, K. M., Lambrus, B. G., Daggubati, V. & Holland, A. J. Binding of STIL to Plk4 activates kinase activity to promote centriole assembly. J. Cell Biol. 209, 863–878 (2015).
Ohta, M. et al. Direct interaction of Plk4 with STIL ensures formation of a single procentriole per parental centriole. Nat. Commun. 5, 5267 (2014).
Wilmott, Z. M., Goriely, A. & Raff, J. W. A simple turing reaction-diffusion model explains how PLK4 breaks symmetry during centriole duplication and assembly. PLoS Biol. 21, e3002391 (2023). This work considers symmetry breaking at the onset of centriole assembly mathematically as a classic Turing reaction–diffusion model.
Takao, D., Yamamoto, S. & Kitagawa, D. A theory of centriole duplication based on self-organized spatial pattern formation. J. Cell Biol. 218, 3537–3547 (2019).
Leda, M., Holland, A. J. & Goryachev, A. B. Autoamplification and competition drive symmetry breaking: initiation of centriole duplication by the PLK4–STIL network. iScience 8, 222–235 (2018).
Cunningham, N. H. J., Bouhlel, I. B. & Conduit, P. T. Daughter centrioles assemble preferentially towards the nuclear envelope in Drosophila syncytial embryos. Open. Biol. 12, 210343 (2022).
van Breugel, M. et al. Structures of SAS-6 suggest its organization in centrioles. Science 331, 1196–1199 (2011).
Kitagawa, D. et al. Structural basis of the 9-fold symmetry of centrioles. Cell 144, 364–375 (2011). Together with van Breugel et al. (2011), this study reports a biophysical and structural analysis of SAS-6, leading to the proposal that SAS-6 self-assembly dictates the centriole 9-fold symmetry.
Banterle, N. et al. Kinetic and structural roles for the surface in guiding SAS-6 self-assembly to direct centriole architecture. Nat. Commun. 12, 6180 (2021).
Fong, C. S., Kim, M., Yang, T. T., Liao, J.-C. & Tsou, M.-F. B. SAS-6 assembly templated by the lumen of cartwheel-less centrioles precedes centriole duplication. Dev. Cell 30, 238–245 (2014).
Guichard, P. et al. Cartwheel architecture of trichonympha basal body. Science 337, 553 (2012).
Aydogan, M. G. et al. A homeostatic clock sets daughter centriole size in flies. J. Cell Biol. 217, 1233–1248 (2018).
Guichard, P. et al. Cell-free reconstitution reveals centriole cartwheel assembly mechanisms. Nat. Commun. 8, 14813 (2017).
Nakazawa, Y., Hiraki, M., Kamiya, R. & Hirono, M. SAS-6 is a cartwheel protein that establishes the 9-fold symmetry of the centriole. Curr. Biol. 17, 2169–2174 (2007).
Hiraki, M., Nakazawa, Y., Kamiya, R. & Hirono, M. Bld10p constitutes the cartwheel-spoke tip and stabilizes the 9-fold symmetry of the centriole. Curr. Biol. 17, 1778–1783 (2007).
Hilbert, M. et al. SAS-6 engineering reveals interdependence between cartwheel and microtubules in determining centriole architecture. Nat. Cell Biol. 18, 393–403 (2016).
Rodrigues-Martins, A. et al. DSAS-6 organizes a tube-like centriole precursor, and its absence suggests modularity in centriole assembly. Curr. Biol. 17, 1465–1472 (2007).
Kovacs, L. et al. Gorab is a Golgi protein required for structure and duplication of Drosophila centrioles. Nat. Genet. 50, 1021–1031 (2018).
Kovacs, L., Fatalska, A. & Glover, D. M. Targeting Drosophila Sas6 to mitochondria reveals its high affinity for Gorab. Biol. Open. 11, bio059545 (2022).
Grzonka, M. & Bazzi, H. Mouse SAS-6 is required for centriole formation in embryos and integrity in embryonic stem cells. eLife 13, e94694 (2024).
Huang, F. et al. Cartwheel disassembly regulated by CDK1–cyclin B kinase allows human centriole disengagement and licensing. J. Biol. Chem. 298, 102658 (2022).
Strnad, P. et al. Regulated HsSAS-6 levels ensure formation of a single procentriole per centriole during the centrosome duplication cycle. Dev. Cell 13, 203–213 (2007).
Yoshiba, S. et al. HsSAS-6-dependent cartwheel assembly ensures stabilization of centriole intermediates. J. Cell Sci. 132, jcs217521 (2019).
Guichard, P., Laporte, M. H. & Hamel, V. The centriolar tubulin code. Semin. Cell Dev. Biol. 137, 16–25 (2023).
Desai, A. & Mitchison, T. J. Microtubule polymerization dynamics. Annu. Rev. Cell dev. Biol. 13, 83–117 (1997).
Kuriyama, R. & Borisy, G. G. Centriole cycle in Chinese hamster ovary cells as determined by whole-mount electron microscopy. J. Cell Biol. 91, 814–821 (1981).
Moyer, T. C. & Holland, A. J. PLK4 promotes centriole duplication by phosphorylating STIL to link the procentriole cartwheel to the microtubule wall. eLife 8, e46054 (2019).
Sharma, A. et al. Centriolar CPAP/SAS-4 imparts slow processive microtubule growth. Dev. Cell 37, 362–376 (2016).
Spektor, A., Tsang, W. Y., Khoo, D. & Dynlacht, B. D. Cep97 and CP110 suppress a cilia assembly program. Cell 130, 678–690 (2007).
Iyer, S. S. et al. Centriolar cap proteins CP110 and CPAP control slow elongation of microtubule plus ends. J. Cell Biol. 224, e202406061 (2025).
Lin, Y.-N. et al. CEP120 interacts with CPAP and positively regulates centriole elongation. J. Cell Biol. 202, 211–219 (2013).
Sharma, A., Gerard, S. F., Olieric, N. & Steinmetz, M. O. Cep120 promotes microtubule formation through a unique tubulin binding C2 domain. J. Struct. Biol. 203, 62–70 (2018).
Comartin, D. et al. CEP120 and SPICE1 cooperate with CPAP in centriole elongation. Curr. Biol. 23, 1360–1366 (2013).
Archinti, M., Lacasa, C., Teixido-Travesa, N. & Luders, J. SPICE—a previously uncharacterized protein required for centriole duplication and mitotic chromosome congression. J. Cell Sci. 123, 3039–3046 (2010).
Tang, C. J., Fu, R. H., Wu, K. S., Hsu, W. B. & Tang, T. K. CPAP is a cell-cycle regulated protein that controls centriole length. Nat. Cell Biol. 11, 825–831 (2009).
Kohlmaier, G. et al. Overly long centrioles and defective cell division upon excess of the SAS-4-related protein CPAP. Curr. Biol. 19, 1012–1018 (2009).
Mahjoub, M. R., Xie, Z. & Stearns, T. Cep120 is asymmetrically localized to the daughter centriole and is essential for centriole assembly. J. Cell Biol. 191, 331–346 (2010).
Schmidt, T. I. et al. Control of centriole length by CPAP and CP110. Curr. Biol. 19, 1005–1011 (2009).
Schmidt-Cernohorska, M. et al. Flagellar microtubule doublet assembly in vitro reveals a regulatory role of tubulin C-terminal tails. Science 363, 285–288 (2019).
Serrano, L., de la Torre, J., Maccioni, R. B. & Avila, J. Involvement of the carboxyl-terminal domain of tubulin in the regulation of its assembly. Proc. Natl Acad. Sci. USA 81, 5989–5993 (1984).
Takeda, Y. et al. Molecular basis promoting centriole triplet microtubule assembly. Nat. Commun. 15, 2216 (2024).
Curinha, A. et al. Centriole structural integrity defects are a crucial feature of hydrolethalus syndrome. J. Cell Biol. 224, e202403022 (2025).
Wang, J. T., Kong, D., Hoerner, C. R., Loncarek, J. & Stearns, T. Centriole triplet microtubules are required for stable centriole formation and inheritance in human cells. eLife 6, e29061 (2017).
Dutcher, S. K., Morrissette, N. S., Preble, A. M., Rackley, C. & Stanga, J. ε-Tubulin is an essential component of the centriole. Mol. Biol. Cell 13, 3859–3869 (2002).
Dutcher, S. K. & Trabuco, E. C. The UNI3 gene is required for assembly of basal bodies of Chlamydomonas and encodes δ-tubulin, a new member of the tubulin superfamily. Mol. Biol. Cell 9, 1293–1308 (1998).
Pudlowski, R. et al. A δ-tubulin/ε-tubulin/Ted protein complex is required for centriole architecture. eL ife 13, RP98704 (2025).
Breslow, D. K. et al. A CRISPR-based screen for Hedgehog signaling provides insights into ciliary function and ciliopathies. Nat. Genet. 50, 460–471 (2018).
Bournonville, L., Laporte, M. H., Borgers, S., Guichard, P. & Hamel, V. The A–C linker controls centriole structural integrity and duplication. Nat. Commun. 16, 6836 (2025).
Sala, C. et al. An interaction network of inner centriole proteins organised by POC1A–POC1B heterodimer crosslinks ensures centriolar integrity. Nat. Commun. 15, 9857 (2024).
Le Guennec, M. et al. A helical inner scaffold provides a structural basis for centriole cohesion. Sci. Adv. 6, eaaz4137 (2020).
Steib, E. et al. WDR90 is a centriolar microtubule wall protein important for centriole architecture integrity. eLife 9, e57205 (2020).
Azimzadeh, J. et al. hPOC5 is a centrin-binding protein required for assembly of full-length centrioles. J. Cell Biol. 185, 101–114 (2009).
Van de Mark, D., Kong, D., Loncarek, J. & Stearns, T. MDM1 is a microtubule-binding protein that negatively regulates centriole duplication. Mol. Biol. Cell 26, 3788–3802 (2015).
Zach, F. et al. The retinitis pigmentosa 28 protein FAM161A is a novel ciliary protein involved in intermolecular protein interaction and microtubule association. Hum. Mol. Genet. 21, 4573–4586 (2012).
Sydor, A. M. et al. PPP1R35 is a novel centrosomal protein that regulates centriole length in concert with the microcephaly protein RTTN. eLife 7, e37846 (2018).
Chen, H.-Y. et al. Human microcephaly protein RTTN interacts with STIL and is required to build full-length centrioles. Nat. Commun. 8, 247 (2017).
Baum, P., Furlong, C. & Byers, B. Yeast gene required for spindle pole body duplication: homology of its product with Ca2+-binding proteins. Proc. Natl Acad. Sci. USA 83, 5512–5516 (1986).
Kilmartin, J. V. Sfi1p has conserved centrin-binding sites and an essential function in budding yeast spindle pole body duplication. J. Cell Biol. 162, 1211–1121 (2003).
Laporte, M. H. et al. Human SFI1 and Centrin form a complex critical for centriole architecture and ciliogenesis. EMBO J. 41, e112107 (2022).
Dantas, T. J., Wang, Y., Lalor, P., Dockery, P. & Morrison, C. G. Defective nucleotide excision repair with normal centrosome structures and functions in the absence of all vertebrate centrins. J. Cell Biol. 193, 307–318 (2011).
Gaudin, N. et al. Evolutionary conservation of centriole rotational asymmetry in the human centrosome. eLife 11, e72382 (2022).
Silflow, C. D. et al. The Vfl1 Protein in Chlamydomonas localizes in a rotationally asymmetric pattern at the distal ends of the basal bodies. J. Cell Biol. 153, 63–74 (2001).
Geimer, S. & Melkonian, M. The ultrastructure of the Chlamydomonas reinhardtii basal apparatus: identification of an early marker of radial asymmetry inherent in the basal body. J. Cell Sci. 117, 2663–2674 (2004).
Paintrand, M., Moudjou, M., Delacroix, H. & Bornens, M. Centrosome organization and centriole architecture: their sensitivity to divalent cations. J. Struct. Biol. 108, 107–128 (1992).
Chong, W. M. et al. Super-resolution microscopy reveals coupling between mammalian centriole subdistal appendages and distal appendages. eLife 9, e53580 (2020).
Ishikawa, H., Kubo, A., Tsukita, S. & Tsukita, S. Odf2-deficient mother centrioles lack distal/subdistal appendages and the ability to generate primary cilia. Nat. Cell Biol. 7, 517–524 (2005).
Bowler, M. et al. High-resolution characterization of centriole distal appendage morphology and dynamics by correlative STORM and electron microscopy. Nat. Commun. 10, 993 (2019).
Yang, T. T. et al. Super-resolution architecture of mammalian centriole distal appendages reveals distinct blade and matrix functional components. Nat. Commun. 9, 2023 (2018).
Chang, T.-J. B., Hsu, J. C.-C. & Yang, T. T. Single-molecule localization microscopy reveals the ultrastructural constitution of distal appendages in expanded mammalian centrioles. Nat. Commun. 14, 1688 (2023).
Rosa, E., Silva, I. et al. Molecular mechanisms underlying the role of the centriolar CEP164–TTBK2 complex in ciliopathies. Structure 30, 114–128.e9 (2022).
Čajánek, L. & Nigg, E. A. Cep164 triggers ciliogenesis by recruiting Tau tubulin kinase 2 to the mother centriole. Proc. Natl Acad. Sci. USA 111, E2841–E2850 (2014).
Oda, T., Chiba, S., Nagai, T. & Mizuno, K. Binding to Cep164, but not EB1, is essential for centriolar localization of TTBK2 and its function in ciliogenesis. Genes. Cell 19, 927–940 (2014).
Wong, C. & Stearns, T. Centrosome number is controlled by a centrosome-intrinsic block to reduplication. Nat. Cell Biol. 5, 539–544 (2003). By fusing cells in different phases of the cell cycle, this work reveals that centrioles in G2 possess a centrosome-intrinsic block to reduplication.
Loncarek, J., Hergert, P., Magidson, V. & Khodjakov, A. Control of daughter centriole formation by the pericentriolar material. Nat. Cell Biol. 10, 322–328 (2008).
Uhlmann, F. Separase regulation during mitosis. Biochem. Soc. Symp. 2003, 243–251 (2003).
Tsou, M. F. & Stearns, T. Mechanism limiting centrosome duplication to once per cell cycle. Nature 442, 947–951 (2006).
Tsou, M. F. et al. Polo kinase and separase regulate the mitotic licensing of centriole duplication in human cells. Dev. Cell 17, 344–354 (2009).
Cabral, G., Sans, S. S., Cowan, C. R. & Dammermann, A. Multiple mechanisms contribute to centriole separation in C. elegans. Curr. Biol. 23, 1380–1387 (2013).
Loncarek, J., Hergert, P. & Khodjakov, A. Centriole reduplication during prolonged interphase requires procentriole maturation governed by Plk1. Curr. Biol. 20, 1277–1282 (2010).
Schockel, L., Mockel, M., Mayer, B., Boos, D. & Stemmann, O. Cleavage of cohesin rings coordinates the separation of centrioles and chromatids. Nat. Cell Biol. 13, 966–972 (2011).
Agircan, F. G. & Schiebel, E. Sensors at centrosomes reveal determinants of local separase activity. PLoS Genet. 10, e1004672 (2014).
Mohr, L. et al. An alternatively spliced bifunctional localization signal reprograms human Shugoshin 1 to protect centrosomal instead of centromeric cohesin. Cell Rep. 12, 2156–2168 (2015).
Wang, X. et al. sSgo1, a major splice variant of Sgo1, functions in centriole cohesion where it is regulated by Plk1. Dev. Cell 14, 331–341 (2008).
Mennella, V. et al. Subdiffraction-resolution fluorescence microscopy reveals a domain of the centrosome critical for pericentriolar material organization. Nat. Cell Biol. 14, 1159–1168 (2012).
Lee, K. & Rhee, K. Separase-dependent cleavage of pericentrin B is necessary and sufficient for centriole disengagement during mitosis. Cell Cycle 11, 2476–2485 (2012).
Matsuo, K. et al. Kendrin is a novel substrate for separase involved in the licensing of centriole duplication. Curr. Biol. 22, 915–921 (2012).
Kim, J., Lee, K. & Rhee, K. PLK1 regulation of PCNT cleavage ensures fidelity of centriole separation during mitotic exit. Nat. Commun. 6, 10076 (2015).
Buchman, J. J. et al. Cdk5rap2 interacts with pericentrin to maintain the neural progenitor pool in the developing neocortex. Neuron 66, 386–402 (2010).
Pagan, J. K. et al. Degradation of Cep68 and PCNT cleavage mediate Cep215 removal from the PCM to allow centriole separation, disengagement and licensing. Nat. Cell Biol. 17, 31–43 (2015).
Barrera, J. A. et al. CDK5RAP2 regulates centriole engagement and cohesion in mice. Dev. Cell 18, 913–926 (2010).
Ito, K. K. et al. Multimodal mechanisms of human centriole engagement and disengagement. EMBO J. 44, 1294–1321 (2025).
Watanabe, K., Takao, D., Ito, K. K., Takahashi, M. & Kitagawa, D. The Cep57–pericentrin module organizes PCM expansion and centriole engagement. Nat. Commun. 10, 931 (2019).
Kim, J., Kim, J. & Rhee, K. PCNT is critical for the association and conversion of centrioles to centrosomes during mitosis. J. Cell Sci. 132, jcs225789 (2019).
Wang, W. J., Soni, R. K., Uryu, K. & Tsou, M. F. The conversion of centrioles to centrosomes: essential coupling of duplication with segregation. J. Cell Biol. 193, 727–739 (2011). This work identifies PLK1-mediated conversion of centrioles to centrosomes as an essential step for MTOC function and procentriole formation.
Tsuchiya, Y., Yoshiba, S., Gupta, A., Watanabe, K. & Kitagawa, D. Cep295 is a conserved scaffold protein required for generation of a bona fide mother centriole. Nat. Commun. 7, 12567 (2016).
Izquierdo, D., Wang, W.-J., Uryu, K. & Tsou, M.-F. B. Stabilization of cartwheel-less centrioles for duplication requires CEP295-mediated centriole-to-centrosome conversion. Cell Rep. 8, 957–965 (2014).
Fong, C. S., Ozaki, K. & Tsou, M.-F. B. PPP1R35 ensures centriole homeostasis by promoting centriole-to-centrosome conversion. Mol. Biol. Cell 29, 2801–2808 (2018).
Atorino, E. S., Hata, S., Funaya, C., Neuner, A. & Schiebel, E. CEP44 ensures the formation of bona fide centriole wall, a requirement for the centriole-to-centrosome conversion. Nat. Commun. 11, 903 (2020).
O’Neill, A. C. et al. Spatial centrosome proteome of human neural cells uncovers disease-relevant heterogeneity. Science 376, eabf9088 (2022).
Carden, S. et al. Proteomic profiling of centrosomes across multiple mammalian cell and tissue types by an affinity capture method. Dev. Cell 58, 2393–2410.e9 (2023). This work develops an efficient affinity capture method that enables the seamless purification of centrioles from various human cell lines.
Wolf, N., Hirsh, D. & McIntosh, J. R. Spermatogenesis in males of the free-living nematode, Caenorhabditis elegans. J. Ultrastruct. Res. 63, 155–169 (1978).
Tollervey, F., Rios, M. U., Zagoriy, E., Woodruff, J. B. & Mahamid, J. Molecular architectures of centrosomes in C. elegans embryos visualized by cryo-electron tomography. Dev. Cell 60, 885–900.e5 (2025).
Pelletier, L., O’Toole, E., Schwager, A., Hyman, A. A. & Muller-Reichert, T. Centriole assembly in Caenorhabditis elegans. Nature 444, 619–623 (2006).
Pierron, M. et al. Centriole elimination during Caenorhabditis elegans oogenesis initiates with loss of the central tube protein SAS-1. EMBO J. 42, e115076 (2023).
Gibbons, I. R. & Grimstone, A. V. On flagellar structure in certain flagellates. J. Biophys. Biochem. Cytol. 7, 697–716 (1960).
Bezler, A. et al. Atypical and distinct microtubule radial symmetries in the centriole and the axoneme of Lecudina tuzetae. Mol. Biol. Cell 33, ar75 (2022).
Schrevel, J. & Besse, C. A functional flagella with a 6 + 0 pattern. J. Cell Biol. 66, 492–507 (1975).
Phillips, D. M. Giant centriole formation in sciara. J. Cell Biol. 33, 73–92 (1967).
Gomes Pereira, S. et al. The 3D architecture and molecular foundations of de novo centriole assembly via bicentrioles. Curr. Biol. 31, 4340–4353.e7 (2021).
Rout, M. P. & Sali, A. Principles for integrative structural biology studies. Cell 177, 1384–1403 (2019).
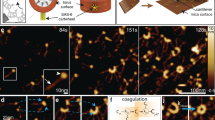
Nievergelt, A. P., Banterle, N., Andany, S. H., Gönczy, P. & Fantner, G. E. High-speed photothermal off-resonance atomic force microscopy reveals assembly routes of centriolar scaffold protein SAS-6. Nat. Nanotechnol. 13, 696–701 (2018).
Martins, A. R., Machado, P., Callaini, G. & Bettencourt-Dias, M. Microscopy methods for the study of centriole biogenesis and function in Drosophila. Methods Cell Biol. 97, 223–242 (2010).
Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 355, 606–612 (2017).
An, H.-L., Kuo, H.-C. & Tang, T. K. Modeling human primary microcephaly with hiPSC-derived brain organoids carrying CPAP-E1235V disease-associated mutant protein. Front. Cell Dev. Biol. 10, 830432 (2022).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Gabriel, E. et al. CPAP promotes timely cilium disassembly to maintain neural progenitor pool. EMBO J. 35, 803–819 (2016).
Fritz-Laylin, L. K. et al. Rapid centriole assembly in Naegleria reveals conserved roles for both de novo and mentored assembly. Cytoskeleton 73, 109–116 (2016).
Graser, S., Stierhof, Y.-D. & Nigg, E. A. Cep68 and Cep215 (Cdk5rap2) are required for centrosome cohesion. J. Cell Sci. 120, 4321–4331 (2007).
Lovera, M. & Lüders, J. The ciliary impact of nonciliary gene mutations. Trends Cell Biol. 31, 876–887 (2021).
Reiter, J. F. & Leroux, M. R. Genes and molecular pathways underpinning ciliopathies. Nat. Rev. Mol. Cell Biol. 18, 533–547 (2017).
Martin, C.-A. et al. Mutations in PLK4, encoding a master regulator of centriole biogenesis, cause microcephaly, growth failure and retinopathy. Nat. Genet. 46, 1283–1292 (2014).
Khan, M. A. et al. A missense mutation in the PISA domain of HsSAS-6 causes autosomal recessive primary microcephaly in a large consanguineous Pakistani family. Hum. Mol. Genet. 23, 5940–5949 (2014).
Kumar, A., Girimaji, S. C., Duvvari, M. R. & Blanton, S. H. Mutations in STIL, encoding a pericentriolar and centrosomal protein, cause primary microcephaly. Am. J. Hum. Genet. 84, 286–290 (2009).
LoMastro, G. M. et al. PLK4 drives centriole amplification and apical surface area expansion in multiciliated cells. eLife 11, e80643 (2022).
Ferrante, M. I. et al. Identification of the gene for oral–facial–digital type I syndrome. Am. J. Hum. Genet. 68, 569–576 (2001).
Thauvin-Robinet, C. et al. The oral–facial–digital syndrome gene C2CD3 encodes a positive regulator of centriole elongation. Nat. Genet. 46, 905–911 (2014).
Joseph, N. et al. Disease-associated mutations in CEP120 destabilize the protein and impair ciliogenesis. Cell Rep. 23, 2805–2818 (2018).
Gönczy, P. Centrosomes and cancer: revisiting a long-standing relationship. Nat. Rev. Cancer 15, 639–652 (2015).
Godinho, S. A. & Basto, R. Centrosomes and cancer: balancing tumor-promoting and inhibitory roles. Trends Cell Biol. 35, 515–526 (2025).
Boveri, T. Über mehrpolige mitosen als mittel zur analyse des zellkerns. Verhandl. Phys.-Med. Ges. Würzburg 35, 67–90 (1902).
Levine, M. S. et al. Centrosome amplification is sufficient to promote spontaneous tumorigenesis in mammals. Dev. Cell 40, 313–322.e5 (2017).
Rosario, C. O. et al. A novel role for Plk4 in regulating cell spreading and motility. Oncogene 34, 3441–3451 (2015).
Rosario, C. O. et al. Plk4 is required for cytokinesis and maintenance of chromosomal stability. Proc. Natl Acad. Sci. USA 107, 6888–6893 (2010).
Ganem, N. J., Godinho, S. A. & Pellman, D. A mechanism linking extra centrosomes to chromosomal instability. Nature 460, 278–282 (2009).
Arnandis, T. et al. Oxidative stress in cells with extra centrosomes drives non-cell-autonomous invasion. Dev. Cell 47, 409–424.e9 (2018).
Marteil, G. et al. Over-elongation of centrioles in cancer promotes centriole amplification and chromosome missegregation. Nat. Commun. 9, 1258 (2018).
Meitinger, F. et al. Control of cell proliferation by memories of mitosis. Science 383, 1441–1448 (2024).
Fulcher, L. J., Sobajima, T., Batley, C., Gibbs-Seymour, I. & Barr, F. A. MDM2 functions as a timer reporting the length of mitosis. Nat. Cell Biol. 27, 262–272 (2025).
Kiermaier, E., Stötzel, I., Schapfl, M. A. & Villunger, A. Amplified centrosomes — more than just a threat. EMBO Rep. 25, 4153–4167 (2024).
Yeow, Z. Y. et al. Targeting TRIM37-driven centrosome dysfunction in 17q23-amplified breast cancer. Nature 585, 447–452 (2020).
Meitinger, F. et al. TRIM37 controls cancer-specific vulnerability to PLK4 inhibition. Nature 585, 440–446 (2020).
Musacchio, A. On the role of phase separation in the biogenesis of membraneless compartments. EMBO J. 41, e109952 (2022).
Woodruff, J. B. The material state of centrosomes: lattice, liquid, or gel? Curr. Opin. Struct. Biol. 66, 139–147 (2021).
Raff, J. W. Phase separation and the centrosome: a fait accompli? Trends Cell Biol. 29, 612–622 (2019).
Woodruff, J. B. et al. The centrosome is a selective condensate that nucleates microtubules by concentrating tubulin. Cell 169, 1066–1077.e10 (2017).
Yamamoto, S., Yabuki, R. & Kitagawa, D. Biophysical and biochemical properties of Deup1 self-assemblies: a potential driver for deuterosome formation during multiciliogenesis. Biol. Open. 10, bio056432 (2021).
Yeh, H.-W. et al. Cep57 regulates human centrosomes through multivalent interactions. Proc. Natl Acad. Sci. USA 121, e2305260121 (2024).
Ahn, J. I. et al. Phase separation of the Cep63•Cep152 complex underlies the formation of dynamic supramolecular self-assemblies at human centrosomes. Cell Cycle 19, 3437–3457 (2020).
Ryniawec, J. M. et al. Polo-like kinase 4 homodimerization and condensate formation regulate its own protein levels but are not required for centriole assembly. Mol. Biol. Cell 34, ar80 (2023).
Qi, F. et al. Dynamic SAS-6 phosphorylation aids centrosome duplication and elimination in C. elegans oogenesis. EMBO Rep. https://doi.org/10.1038/s44319-025-00485-7 (2025).
Voß, Y., Klaus, S., Lichti, N. P., Ganter, M. & Guizetti, J. Malaria parasite centrins can assemble by Ca2+-inducible condensation. PLoS Pathog. 19, e1011899 (2023).
Kalbfuss, N. & Gönczy, P. Towards understanding centriole elimination. Open. Biol. 13, 230222 (2023).
Werner, S., Pimenta-Marques, A. & Bettencourt-Dias, M. Maintaining centrosomes and cilia. J. Cell Sci. 130, 3789–3800 (2017).
Takumi, K. & Kitagawa, D. Experimental and natural induction of de novo centriole formation. Front. Cell Dev. Biol. 10, 861864 (2022).
Nabais, C., Pereira, S. G. & Bettencourt-Dias, M. Noncanonical biogenesis of centrioles and basal bodies. Cold Spring Harb. Symp. Quant. Biol. 82, 123–135 (2017).
Woolley, D. M. & Fawcett, D. W. The degeneration and disappearance of the centrioles during the development of the rat spermatozoon. Anat. Rec. 177, 289–301 (1973).
Courtois, A., Schuh, M., Ellenberg, J. & Hiiragi, T. The transition from meiotic to mitotic spindle assembly is gradual during early mammalian development. J. Cell Biol. 198, 357–370 (2012).
Hepler, P. K. The blepharoplast of Marsilea: its de novo formation and spindle association. J. Cell Sci. 21, 361–390 (1976).
Klink, V. P. & Wolniak, S. M. Centrin is necessary for the formation of the motile apparatus in spermatids of Marsilea. Mol. Biol. Cell 12, 761–776 (2001).
Fritz-Laylin, L. K. & Fulton, C. Naegleria: a classic model for de novo basal body assembly. Cilia 5, 10 (2016).
Fritz-Laylin, L. K., Assaf, Z. J., Chen, S. & Cande, W. Z. Naegleria gruberi de novo basal body assembly occurs via stepwise incorporation of conserved proteins. Eukaryot. Cell 9, 860–865 (2010).
Peel, N., Stevens, N. R., Basto, R. & Raff, J. W. Overexpressing centriole-replication proteins in vivo induces centriole overduplication and de novo formation. Curr. Biol. 17, 834–843 (2007).
Rodrigues-Martins, A., Riparbelli, M., Callaini, G., Glover, D. M. & Bettencourt-Dias, M. Revisiting the role of the mother centriole in centriole biogenesis. Science 316, 1046–1050 (2007).
Nabais, C. et al. Plk4 triggers autonomous de novo centriole biogenesis and maturation. J. Cell Biol. 220, e202008090 (2021).
Khodjakov, A. et al. De novo formation of centrosomes in vertebrate cells arrested during S phase. J. Cell Biol. 158, 1171–1181 (2002).
Wang, W. J. et al. De novo centriole formation in human cells is error-prone and does not require SAS-6 self-assembly. eLife 4, e10586 (2015).
Sala, R., Farrell, K. C. & Stearns, T. Growth disadvantage associated with centrosome amplification drives population-level centriole number homeostasis. Mol. Biol. Cell 31, 2646–2656 (2020).
Acknowledgements
The author is grateful to G. N. Hatzopoulos for help preparing most figure panels, and thanks F. Douma for Fig. 1b as well as P. Keeling for generating a Creative Commons version of the figure for Supplementary Box 1 that could be adapted here. Moreover, G. N. Hatzopoulos, F. Douma and S. Aich are acknowledged for their useful comments on the manuscript. The author apologizes to those scientists whose work could not be mentioned due to space limitations or ignorance. Work on centriole assembly in the author’s laboratory during writing of this review was supported by the European Research Council (AdG 588437) and Swiss Cancer Research (KFS-6018-02-2024-R).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Peer review
Peer review information
Nature Reviews Molecular Cell Biology thanks Mónica Bettencourt-Dias, Fanny Gergely and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Glossary
- APC/C
-
(anaphase-promoting complex, also known as the cyclosome). A large protein complex with E3 ubiquitin ligase activity that poly-ubiquitinates target cell cycle proteins, thus marking them for degradation by the 26S proteasome.
- Axoneme
-
A microtubule-based structure in cilia and flagella that exhibits a 9-fold radial symmetry of microtubule doublets stemming from the likewise symmetrical centriolar microtubule doublets.
- Cryo electron tomography
-
(cryo-ET). Electron microscopy approach in which the specimen is preserved in the native state by freezing at cryogenic temperature, and then imaged through a tilt series, followed by specimen reconstruction.
- Dynamic instability
-
Refers to the ensemble dynamic behaviour of microtubules: once nucleated, microtubules switch between growth and shrinkage phases at their plus end, with catastrophe and rescue events between the two phases.
- G1/S transition
-
A cell cycle transition between the G1 phase that follows mitosis and the S phase, when DNA replication takes place; the G1/S transition generally commits the cell to completing the cell cycle.
- Pericentriolar material
-
(PCM). A region surrounding centrioles from where most microtubules are nucleated in many animal cells.
- Ultrastructural expansion microscopy
-
(U-ExM). A method in which the specimen is augmented isotropically using a swellable hydrogel, thereby enlarging the effective resolution correspondingly.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gönczy, P. Critical constituents and assembly principles of centriole biogenesis in human cells. Nat Rev Mol Cell Biol 27, 260–277 (2026). https://doi.org/10.1038/s41580-025-00921-5
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41580-025-00921-5