Abstract
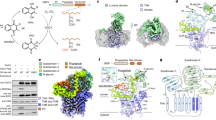
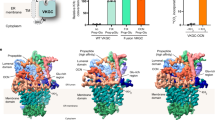
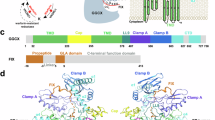
The γ-carboxylation of glutamate residues enables Ca2+-mediated membrane assembly of protein complexes that support broad physiological functions, including haemostasis, calcium homeostasis, immune response and endocrine regulation1,2,3,4. Modulating γ-carboxylation levels provides prevalent treatments for haemorrhagic and thromboembolic diseases5. This unique post-translational modification requires vitamin K hydroquinone (KH2) to drive highly demanding reactions6 catalysed by the membrane-integrated γ-carboxylase (VKGC). Here, to decipher the underlying mechanisms, we determined cryo-electron microscopy structures of human VKGC in unbound form, with KH2 and four haemostatic and non-haemostatic proteins possessing propeptides and glutamate-rich domains in different carboxylation states. VKGC recognizes substrate proteins through knob-and-hole interactions with propeptides, thereby bringing tethered glutamate-containing segments for processive carboxylation within a large chamber that provides steric control. Propeptide binding also triggers a global conformational change to signal VKGC activation. Through sequential deprotonation and KH2 epoxidation, VKGC generates a free hydroxide ion as an exceptionally strong base that is required to deprotonate the γ-carbon of glutamate for CO2 addition. The diffusion of this superbase—protected and guided by a sealed hydrophobic tunnel—elegantly resolves the challenge of coupling KH2 epoxidation to γ-carboxylation across the membrane interface. These structural insights and extensive functional experiments advance membrane enzymology and propel the development of treatments for γ-carboxylation disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM maps have been deposited into the Electron Microscopy Data Bank under accession numbers EMD-44939 (apo VKGC), EMD-44941 (VKGC–TMG2), EMD-44940 (VKGC–TMG2 with KH2), EMD-44935 (VKGC–FIX with KH2), EMD-44936 (VKGC–FX with KH2), EMD-44937 (VKGC–PC with KH2) and EMD-44942 (partially carboxylated VKGC–FIX with KH2). The coordinates have been deposited at the PDB under accession numbers 9BVO (apo VKGC), 9BVQ (VKGC–TMG2), 9BVP (VKGC–TMG2 with KH2), 9BVK (VKGC–FIX with KH2), 9BVL (VKGC–FX with KH2), 9BVM (VKGC–PC with KH2) and 9BVR (partially carboxylated VKGC–FIX with KH2). The MS data have been deposited in ProteomeXchange under identifier PXD052939. Source data are provided with this paper.
References
Furie, B. & Furie, B. C. Molecular basis of vitamin K-dependent gamma-carboxylation. Blood 75, 1753–1762 (1990).
Dahlback, B. Vitamin K-dependent protein S: beyond the protein C pathway. Semin. Thromb. Hemost. 44, 176–184 (2018).
Murshed, M., Schinke, T., McKee, M. D. & Karsenty, G. Extracellular matrix mineralization is regulated locally; different roles of two gla-containing proteins. J. Cell Biol. 165, 625–630 (2004).
Lacombe, J. et al. Vitamin K-dependent carboxylation regulates Ca2+ flux and adaptation to metabolic stress in beta cells. Cell Rep. 42, 112500 (2023).
Sadler, J. E. K is for koagulation. Nature 427, 5–6 (2004).
Dowd, P., Hershline, R., Ham, S. W. & Naganathan, S. Vitamin K and energy transduction: a base strength amplification mechanism. Science 269, 1684–1691 (1995).
Mosnier, L. O., Zlokovic, B. V. & Griffin, J. H. The cytoprotective protein C pathway. Blood 109, 3161–3172 (2007).
Ma, H. et al. Vitamin K2-dependent GGCX and MGP are required for homeostatic calcium regulation of sperm maturation. iScience 14, 210–225 (2019).
Bandyopadhyay, P. K. Vitamin K-dependent gamma-glutamylcarboxylation: an ancient posttranslational modification. Vitam. Horm. 78, 157–184 (2008).
Xu, Z., Spencer, H. J., Harris, V. A. & Perkins, S. J. An updated interactive database for 1692 genetic variants in coagulation factor IX provides detailed insights into hemophilia B. J. Thromb. Haemost. 21, 1164–1176 (2023).
Cao, Q. et al. Molecular basis of inherited protein C deficiency results from genetic variations in the signal peptide and propeptide regions. J. Thromb. Haemost. 21, 3124–3137 (2023).
Oldenburg, J. et al. Missense mutations at ALA-10 in the factor IX propeptide: an insignificant variant in normal life but a decisive cause of bleeding during oral anticoagulant therapy. Br. J. Haematol. 98, 240–244 (1997).
Palla, R., Peyvandi, F. & Shapiro, A. D. Rare bleeding disorders: diagnosis and treatment. Blood 125, 2052–2061 (2015).
Castiglione, V. et al. Evaluation of inactive Matrix-Gla-protein (MGP) as a biomarker for incident and recurrent kidney stones. J. Nephrol. 33, 101–107 (2020).
Rost, S. et al. Compound heterozygous mutations in the γ-glutamyl carboxylase gene cause combined deficiency of all vitamin K‐dependent blood coagulation factors. Br. J. Haematol. 126, 546–549 (2004).
De Vilder, E. Y., Debacker, J. & Vanakker, O. M. GGCX-associated phenotypes: an overview in search of genotype-phenotype correlations. Int. J. Mol. Sci. 18, 240 (2017).
Morris, D. P., Stevens, R. D., Wright, D. J. & Stafford, D. W. Processive post-translational modification. Vitamin K-dependent carboxylation of a peptide substrate. J. Biol. Chem. 270, 30491–30498 (1995).
Stenina, O., Pudota, B. N., McNally, B. A., Hommema, E. L. & Berkner, K. L. Tethered processivity of the vitamin K-dependent carboxylase: factor IX is efficiently modified in a mechanism which distinguishes Gla’s from Glu’s and which accounts for comprehensive carboxylation in vivo. Biochemistry 40, 10301–10309 (2001).
Higgins-Gruber, S. L. et al. Effect of vitamin K-dependent protein precursor propeptide, vitamin K hydroquinone, and glutamate substrate binding on the structure and function of γ-glutamyl carboxylase. J. Biol. Chem. 285, 31502–31508 (2010).
Suttie, J. W. Vitamin K-dependent carboxylase. Annu. Rev. Biochem. 54, 459–477 (1985).
Sugiura, I., Furie, B., Walsh, C. T. & Furie, B. C. Propeptide and glutamate-containing substrates bound to the vitamin K-dependent carboxylase convert its vitamin K epoxidase function from an inactive to an active state. Proc. Natl Acad. Sci. USA 94, 9069–9074 (1997).
Kulman, J. D., Harris, J. E., Xie, L. & Davie, E. W. Identification of two novel transmembrane gamma-carboxyglutamic acid proteins expressed broadly in fetal and adult tissues. Proc. Natl Acad. Sci. USA 98, 1370–1375 (2001).
Tie, J., Wu, S. M., Jin, D., Nicchitta, C. V. & Stafford, D. W. A topological study of the human gamma-glutamyl carboxylase. Blood 96, 973–978 (2000).
Sugiura, I., Furie, B., Walsh, C. T. & Furie, B. C. Profactor IX propeptide and glutamate substrate binding sites on the vitamin K-dependent carboxylase identified by site-directed mutagenesis. J. Biol. Chem. 271, 17837–17844 (1996).
Lomize, A. L., Todd, S. C. & Pogozheva, I. D. Spatial arrangement of proteins in planar and curved membranes by PPM 3.0. Protein Sci. 31, 209–220 (2022).
Wu, S. M., Morris, D. P. & Stafford, D. W. Identification and purification to near homogeneity of the vitamin K-dependent carboxylase. Proc. Natl Acad. Sci. USA 88, 2236–2240 (1991).
Watzka, M. et al. Bleeding and non-bleeding phenotypes in patients with GGCX gene mutations. Thromb. Res. 134, 856–865 (2014).
Hao, Z. et al. γ-Glutamyl carboxylase mutations differentially affect the biological function of vitamin K-dependent proteins. Blood 137, 533–543 (2021).
Ghosh, S. et al. GGCX mutations show different responses to vitamin K thereby determining the severity of the hemorrhagic phenotype in VKCFD1 patients. J. Thromb. Haemost. 19, 1412–1424 (2021).
Ghosh, S. et al. GGCX variants leading to biallelic deficiency to gamma-carboxylate GRP cause skin laxity in VKCFD1 patients. Hum. Mutat. 43, 42–55 (2022).
Tie, J. K. et al. Determination of disulfide bond assignment of human vitamin K-dependent gamma-glutamyl carboxylase by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J. Biol. Chem. 278, 45468–45475 (2003).
Tie, J.-K., Zheng, M.-Y., Pope, R. M., Straight, D. L. & Stafford, D. W. Identification of the N-linked glycosylation sites of vitamin K-dependent carboxylase and effect of glycosylation on carboxylase function. Biochemistry 45, 14755–14763 (2006).
Stanley, T. B., Jin, D.-Y., Lin, P.-J. & Stafford, D. W. The propeptides of the vitamin K-dependent proteins possess different affinities for the vitamin K-dependent carboxylase. J. Biol. Chem. 274, 16940–16944 (1999).
Jorgensen, M. J. et al. Recognition site directing vitamin K-dependent gamma-carboxylation resides on the propeptide of factor IX. Cell 48, 185–191 (1987).
Hao, Z., Jin, D. Y., Stafford, D. W. & Tie, J. K. Vitamin K-dependent carboxylation of coagulation factors: insights from a cell-based functional study. Haematologica 105, 2164–2173 (2020).
Pan, L. C. & Price, P. A. The propeptide of rat bone gamma-carboxyglutamic acid protein shares homology with other vitamin K-dependent protein precursors. Proc. Natl Acad. Sci. USA 82, 6109–6113 (1985).
Stanley, T. B., Humphries, J., High, K. A. & Stafford, D. W. Amino acids responsible for reduced affinities of vitamin K-dependent propeptides for the carboxylase. Biochemistry 38, 15681–15687 (1999).
Belvini, D. et al. Molecular genotyping of the Italian cohort of patients with hemophilia B. Haematologica 90, 635–642 (2005).
Herrmann, F. H. et al. Factor VII deficiency: clinical manifestation of 717 subjects from Europe and Latin America with mutations in the factor 7 gene. Haemophilia 15, 267–280 (2009).
Johnsen, J. M. et al. Novel approach to genetic analysis and results in 3000 hemophilia patients enrolled in the My Life, Our Future initiative. Blood Adv. 1, 824–834 (2017).
Pezeshkpoor, B. et al. Variants in FIX propeptide associated with vitamin K antagonist hypersensitivity: functional analysis and additional data confirming the common founder mutations. Ann. Hematol. 97, 1061–1069 (2018).
Mutucumarana, V. P., Acher, F., Straight, D. L., Jin, D.-Y. & Stafford, D. W. A conserved region of human vitamin K-dependent carboxylase between residues 393 and 404 is important for its interaction with the glutamate substrate. J. Biol. Chem. 278, 46488–46493 (2003).
Rishavy, M. A. & Berkner, K. L. Insight into the coupling mechanism of the vitamin K-dependent carboxylase: mutation of histidine 160 disrupts glutamic acid carbanion formation and efficient coupling of vitamin K epoxidation to glutamic acid carboxylation. Biochemistry 47, 9836–9846 (2008).
Mutucumarana, V. P. et al. Expression and characterization of the naturally occurring mutation L394R in human γ-glutamyl carboxylase. J. Biol. Chem. 275, 32572–32577 (2000).
Zacchi, L. F. et al. Coagulation factor IX analysis in bioreactor cell culture supernatant predicts quality of the purified product. Commun. Biol. 4, 390 (2021).
Kaufman, R. J., Wasley, L. C., Furie, B. C., Furie, B. & Shoemaker, C. B. Expression, purification, and characterization of recombinant gamma-carboxylated factor IX synthesized in Chinese hamster ovary cells. J. Biol. Chem. 261, 9622–9628 (1986).
Presnell, S. R., Tripathy, A., Lentz, B. R., Jin, D. Y. & Stafford, D. W. A novel fluorescence assay to study propeptide interaction with gamma-glutamyl carboxylase. Biochemistry 40, 11723–11733 (2001).
Li, Q. et al. Mutations in the GGCX and ABCC6 genes in a family with pseudoxanthoma elasticum-like phenotypes. J. Invest. Dermatol. 129, 553–563 (2009).
Rishavy, M. A. et al. Brønsted analysis reveals Lys218 as the carboxylase active site base that deprotonates vitamin K hydroquinone to initiate vitamin K-dependent protein carboxylation. Biochemistry 45, 13239–13248 (2006).
Dowd, P., Ham, S. W. & Hershline, R. Role of oxygen in the vitamin K-dependent carboxylation reaction: incorporation of a second atom of oxygen 18 from molecular oxygen-18O2 into vitamin K oxide during carboxylase activity. J. Am. Chem. Soc. 114, 7613–7617 (1992).
Parker, C. H. et al. A conformational investigation of propeptide binding to the integral membrane protein γ-glutamyl carboxylase using nanodisc hydrogen exchange mass spectrometry. Biochemistry 53, 1511–1520 (2014).
Soute, B. A. et al. Congenital deficiency of all vitamin K-dependent blood coagulation factors due to a defective vitamin K-dependent carboxylase in Devon Rex cats. Thromb. Haemost. 68, 521–525 (1992).
Furie, B. C. et al. The gamma-carboxylation recognition site is sufficient to direct vitamin K-dependent carboxylation on an adjacent glutamate-rich region of thrombin in a propeptide-thrombin chimera. J. Biol. Chem. 272, 28258–28262 (1997).
Knobloch, J. E. & Suttie, J. W. Vitamin K-dependent carboxylase. Control of enzyme activity by the “propeptide” region of factor X. J. Biol. Chem. 262, 15334–15337 (1987).
Davis, C. H., Deerfield, D. II., Wymore, T., Stafford, D. W. & Pedersen, L. G. A quantum chemical study of the mechanism of action of vitamin K carboxylase (VKC) III. Intermediates and transition states. J. Mol. Graph. Model. 26, 409–414 (2007).
Bierbaumer, S. et al. Enzymatic conversion of CO2: from natural to artificial utilization. Chem. Rev. 123, 5702–5754 (2023).
Larson, A. E., Friedman, P. A. & Suttie, J. W. Vitamin K-dependent carboxylase. Stoichiometry of carboxylation and vitamin K 2,3-epoxide formation. J. Biol. Chem. 256, 11032–11035 (1981).
Rishavy, M. A. & Berkner, K. L. Vitamin K oxygenation, glutamate carboxylation, and processivity: defining the three critical facets of catalysis by the vitamin K-dependent carboxylase. Adv. Nutr. 3, 135–148 (2012).
Suttie, J. W. Mechanism of action of vitamin K: demonstration of a liver precursor of prothrombin. Science 179, 192–194 (1973).
Nelsestuen, G. L. & Suttie, J. W. Mode of action of vitamin K. Calcium binding properties of bovine prothrombin. Biochemistry 11, 4961–4964 (1972).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Chu, R. et al. Redesign of a four-helix bundle protein by phage display coupled with proteolysis and structural characterization by NMR and X-ray crystallography. J. Mol. Biol. 323, 253–262 (2002).
Mukherjee, S. et al. Synthetic antibodies against BRIL as universal fiducial marks for single-particle cryoEM structure determination of membrane proteins. Nat. Commun. 11, 1598 (2020).
Tsutsumi, N. et al. Structure of human Frizzled5 by fiducial-assisted cryo-EM supports a heterodimeric mechanism of canonical Wnt signaling. eLife 9, e58464 (2020).
Li, C. et al. FastCloning: a highly simplified, purification-free, sequence- and ligation-independent PCR cloning method. BMC Biotechnol. 11, 92 (2011).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rubinstein, J. L. & Brubaker, M. A. Alignment of cryo-EM movies of individual particles by optimization of image translations. J. Struct. Biol. 192, 188–195 (2015).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. D. Features and development of Coot. Acta Crystallogr. D. 66, 486–501 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Freedman, S. J., Furie, B. C., Furie, B. & Baleja, J. D. Structure of the calcium ion-bound gamma-carboxyglutamic acid-rich domain of factor IX. Biochemistry 34, 12126–12137 (1995).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Pravda, L. et al. MOLEonline: a web-based tool for analyzing channels, tunnels and pores (2018 update). Nucleic Acids Res. 46, W368–W373 (2018).
Van Zundert, G. C. P. et al. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J. Mol. Biol. 428, 720–725 (2016).
Lacombe, J., Rishavy, M. A., Berkner, K. L. & Ferron, M. VKOR paralog VKORC1L1 supports vitamin K-dependent protein carboxylation in vivo. JCI Insight 3, e96501 (2018).
Gao, W. et al. Characterization of missense mutations in the signal peptide and propeptide of FIX in hemophilia B by a cell-based assay. Blood Adv. 4, 3659–3667 (2020).
Li, H. et al. The interactions of dopamine and oxidative damage in the striatum of patients with neurodegenerative diseases. J. Neurochem. 152, 235–251 (2020).
Wisniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Meier, F. et al. Parallel accumulation-serial fragmentation (PASEF): multiplying sequencing speed and sensitivity by synchronized scans in a trapped ion mobility device. J. Proteome Res. 14, 5378–5387 (2015).
Meier, F. et al. Online parallel accumulation-serial fragmentation (PASEF) with a novel trapped ion mobility mass spectrometer. Mol. Cell. Proteomics 17, 2534–2545 (2018).
Matyash, V., Liebisch, G., Kurzchalia, T. V., Shevchenko, A. & Schwudke, D. Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J. Lipid Res. 49, 1137–1146 (2008).
Hsu, F.-F. & Turk, J. Electrospray ionization with low-energy collisionally activated dissociation tandem mass spectrometry of glycerophospholipids: mechanisms of fragmentation and structural characterization. J. Chromatogr. B 877, 2673–2695 (2009).
Berkner, K. L. & Pudota, B. N. Vitamin K-dependent carboxylation of the carboxylase. Proc. Natl Acad. Sci. USA 95, 466–471 (1998).
Ramstrom, M. & Sandberg, H. Characterization of gamma-carboxylated tryptic peptides by collision-induced dissociation and electron transfer dissociation mass spectrometry. Eur. J. Mass Spectrom. 17, 497–506 (2011).
Kelleher, N. L. et al. Localization of labile posttranslational modifications by electron capture dissociation: the case of gamma-carboxyglutamic acid. Anal. Chem. 71, 4250–4253 (1999).
Huang, M., Furie, B. C. & Furie, B. Crystal structure of the calcium-stabilized human factor IX Gla domain bound to a conformation-specific anti-factor IX antibody. J. Biol. Chem. 279, 14338–14346 (2004).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Ohkubo, Y. Z. & Tajkhorshid, E. Distinct structural and adhesive roles of Ca2+ in membrane binding of blood coagulation factors. Structure 16, 72–81 (2008).
Lin, P.-J. et al. The putative vitamin K-dependent gamma-glutamyl carboxylase internal propeptide appears to be the propeptide binding site. Biochemistry 277, 28584–28591 (2002).
Acknowledgements
We thank J.-K. Tie and M. Ferron for discussions and for providing the GGCX−/− reporter cell line and the anti-Gla antibody, respectively. W.L. is supported by the National Heart, Lung, and Blood Institute (R01 HL121718), the Vagelos Endowed Chair, the Established Investigator Award and Collaborative Sciences Award from American Heart Association, Children’s Discovery Institute (MCII 2020-854), the National Institute of Allergy and Infectious Diseases (R01 AI158500) and the Forefront of Science Award from W. M. Keck Foundation. W.L. and M.L.G. are supported by National Institute of General Medical Sciences (R01 GM131008). B.A.G. is supported by the National Institute of Neurological Disorders and Stroke (R01 NS111997), the Eunice Kennedy Shriver National Institute of Child Health & Human Development (R01 HD106051) and the National Science Foundation (CHE 2127882). Z.L. is supported by a BMB Research Seed Grant (PJ000027587) and National Institute on Aging (P30 AG066444). B.L. is supported by the Hormel Institute, University of Minnesota.
Author information
Authors and Affiliations
Contributions
Q.C. performed cryo-EM, functional and MS experiments. M.S. provided primary support to Q.C., and H.C., G.S., S.L. and J.X. assisted. B.L. and W.L. determined the structures. A.A., Z.L., F.-F.H., M.C. and J.S. performed MS analyses with the support of M.L.G. and B.A.G. W.L. conceived and directed the project, and wrote the manuscript with input from Q.C., A.M.K. and all of the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Darrel Stafford and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The fusion strategy improves the structural resolution and protein expression while preserving VKGC catalytic activity.
a, 2D classification of wild-type VKGC and TMG2-BRIL-VKGC fusion protein. The particles of wild-type VKGC were extracted at box size of 320 pixels (1.1 Å/pixel), and TMG2-BRIL-VKGC at box size of 320 pixels (0.885 Å/pixel). The 2D classifications for both wild-type and fusion proteins show high-resolution features in many regions. b, Comparison of cryo-EM density maps of wild-type VKGC (left) and TMG2-BRIL-VKGC (right), coloured by domains (as in Fig. 1h). For wild-type VKGC, the 3D reconstruction yielded only a 4.4 Å resolution map with part of the VKGC extramembrane region disordered. c, Expression levels of TMG2-VKGC with and without BRIL insertion, assessed by Western blot using an anti-VKGC antibody. The original blots are shown in Supplementary Fig. 2. UN, untransfected. This experiment was performed once. d, FSEC profile comparison of TMG2-VKGC with and without BRIL insertion. These proteins are tagged with a C-terminal eGFP, expressed in HEK 293T cells, and extracted in DDM. e, The elution profile of size-exclusion chromatography during the protein purification of TMG2-BRIL-VKGC. f, SDS-PAGE of the peak fractions (17-22 corresponding to 16-18 mL) from size-exclusion chromatography. M, protein marker. The original gel is shown in Supplementary Fig. 3. This experiment was performed once. g, Immunoblots showing γ-carboxylation of TMG2-BRIL-VKGC in VKGC-knockout (k/o) cells treated with vitamin K (K, reduced to KH2 in cells). Carboxylation is not observed in untreated or warfarin (W) treated cells. The anti-Gla antibody recognizes wild-type (WT) VKGC due to self-carboxylation, and vitamin K treatment induces an electrophoresis-mobility shift85. This shift is not evident in the fusion protein, likely due to its larger size (with an eGFP tag) or hindrance of self-carboxylation by the fusion. For gel source data, see Supplementary Fig. 4. This experiment was performed once. h, Self-incorporation of 14CO2 into VKGC fusion proteins (20 pmol). The autoradiographs (left) were quantified to determine the 14CO2 incorporation levels (right). The radioactivity counting (mean ± s.d. from n = 3 biological replicates) did not include the upper bands, which represent slight aggregation of protein samples in the gel lanes. The TMG2 Prop-Glu fusion (i.e., TMG2-BRIL-VKGC) was used in the control without KH2. The original autoradiographs are shown in Supplementary Fig. 6. i, Competition of free FIX Prop against TMG2 Prop-Glu and VKGC fusion protein (1 µM). Fluorescein-labelled proFIX18 was used, and the protein-bound fraction was captured by crosslinking. Wild-type VKGC and BSA served as controls. The original gels are shown in Supplementary Fig. 7. This experiment was performed once. j, Epoxidase activities of wild-type VKGC protein (1 µM) is similar with 1:1 and 1:10 ratios of FIX Prop-Glu. k, Ammonium sulfate has little effect on the activity of the TMG2 Prop-Glu and VKGC fusion protein. l, Ammonium sulfate substantially increases the epoxidase activity with the FLEEL substrate. Reactions were conducted according to Sugiura, et al.21, with 0.01 µM wild-type VKGC, 64 µM proFIX18 (Prop) or proFIX28 (Prop-Glu), 3.6 mM FLEEL (sGlu), 222 µM KH2, 8 mM DTT, and a 30 min incubation. m, Higher DTT concentration slightly increases the epoxidase activity in presence of proFIX18 and FLEEL. In our conditions, ~2 mM DTT was introduced with KH2 addition, whereas Sugiura, et al.21 used an additional 8 mM DTT. Other conditions in this experiment are the same as in l. n, The relative activities of different substrates remain similar with different protein concentrations of wild-type VKGC. For comparison, the condition without ammonium sulfate and with 2 µM proFIX18 and 5 mM FLEEL was used. Data in j-n are mean ± s.d. from n = 3 biological replicates.
Extended Data Fig. 2 Quality of cryo-EM maps.
From Left to right, Local resolution illustrations of the cryo-EM maps, angular distribution plots, half-map FSC curves, and model-to-map FSCs. Respective representations are shown for fusion constructs of VKGC with TMG2 Prop-Glu (a), VKGC with TMG2 Prop-Glu and KH2 (b), VKGC with FIX Prop-Glu and KH2 (c), VKGC with FX Prop-Glu and KH2 (d), VKGC with PC Prop-Glu and KH2 (e), and VKGC with partially carboxylated FIX Prop-Glu and KH2 (f), and wild-type VKGC in unbound state (g).
Extended Data Fig. 3 Cryo-EM data processing.
The dataset and density maps of VKGC with Prop-Glu of TMG2 and KH2 are shown as a representative. Other datasets are similarly processed and quality of density maps are similar. a, A representative raw cryo-EM image. This experiment was performed once with thousands of raw images collected. b, Representative 2D classes. c, The data processing procedure. Two rounds of 2D classification and two rounds of heterogenous refinements generated a 3.3 Å final map. d, Density maps of VKGC individual regions. e, Density maps of KH2 and two bound phosphatidylcholine (PhosC) molecules. f, Density maps of TMG2 propeptide and Glu-rich regions.
Extended Data Fig. 4 The large VKGC chamber may permit the binding of the entire Glu-rich region and the folding of Gla-rich region into helices.
a, Model of the 64 AA Prop-Glu region of FIX in an extended conformation within the large chamber of VKGC. The modelling used HADDOCK76, with basic residues in VKGC and Glu residues in FIX Glu region selected for direct interactions. The modelled part of Glu-rich region is coloured in purple, and the propeptide and Glu-rich region observed in cryo-EM structure in orange and red, respectively. b, View of the FIX Prop-Glu model with VKGC shown in electrostatic surface. The lower part of the large chamber is positively charged, encircled by a belt of basic residues that potentially facilitate the binding of negatively charged Glu or Gla residues24,58 during processive γ-carboxylation. c, The cryo-EM model of partially carboxylated FIX in front (left) and side (right) views with the ER membrane modelled by PPM analysis25. The double arrow indicates proximity of the Gla-rich region to the membrane opening of VKGC. d, MS quantification of γ-carboxylation levels of individual Glu residues in partially carboxylated FIX fused with VKGC. Data are mean ± s.d. from n = 3 biological replicates. Glu/Gla residues detectable by ETD fragmentation were shown; ETD was used to optimally preserve γ-carboxylation86,87. The relative carboxylation levels are consistent with HCD fragmentation, but the absolute levels in HCD are relatively lower due to partial loss of this labile post-translational modification. e, MS-MS spectra of γ-carboxylated peptides (used in d) with the ETD fragmentation mode. f, Conformational change of the FIX Glu-rich region from the uncarboxylated state (propeptide-tethered, VKGC bound, and without Ca2+; Fig. 4a–e), the partially carboxylated state (propeptide-tethered, VKGC bound, and with Ca2+, as in c), to the carboxylated, functional state (in absence of propeptide and VKGC, but with Ca2+; PDB 1NL0)88. The uncarboxylated and partially carboxylated structures are superimposed, and the fully carboxylated structure is shown in a similar N to C direction. g, Density maps of the propeptide (left) and partially carboxylated Gla-rich region (right, unsharpened map) of FIX.
Extended Data Fig. 5 Structural location of VKCFD1 mutations.
a-b, Location of VKCFD1 mutations (red spheres) on the VKGC structure, in side (a) and top (b) views. c, Cluster of VKCFD1 mutations (in bold letters) near a propeptide-binding pocket (site III in Fig. 3). d, Cluster of VKCFD1 mutations near the c-Glu binding site. e, VKCFD1 mutations at the KH2 binding site. S300F and F299S are found in the dysfunctional allele of VKCFD1 patients (Supplementary Table 1) and exhibit no γ-carboxylation activities (Fig. 5f and Extended Data Fig. 8c)28. Another VKCFD1 mutation, S284P, may act as a helix breaker in RH7, altering several residues that interact with KH2 (Fig. 5d). f, VKCFD1 mutations located at the α/β domain interface interacting with other structural components. These mutations potentially interfere with the allosteric motion or propeptide binding of VKGC.
Extended Data Fig. 6 Binding interactions between VKGC and propeptides of VKDPs.
a, Binding of the VKDP propeptides (Prop) to the α/β and jellyroll domains. b, Polar interactions at the interface between the α/β and jellyroll domains. c, Sequence alignment of propeptides from human VKDPs and VKGC-binding affinities of these propeptides19. PS, protein S. PZ, protein Z. OCN, osteocalcin. MGP, matrix Gla protein. GRP, Gla-rich protein or unique cartilage matrix-associated protein. The MGP propeptide does not contain the recognition motif for furin cleavage and is not removed in mature protein. Uncommon residues in VKDPs at the three key sites are indicated by red letter, with those known to affect propeptide binding in bold. The sequence logo89 above depicts the residue conservation level at each position. n.g., negligible. d, Cellular γ-carboxylation activities of VKGC mutants at the propeptide binding sites I, II and III. FIX-Gla-PC is used as the γ-carboxylation reporter. Data are mean ± s.d. from n = 3 biological replicates.
Extended Data Fig. 7 Glu-rich regions of VKDPs.
a, VKGC binding interactions with the 5-AA Glu-containing segments from FIX, FX, TMG2, and PC. b, Electrostatic surface showing the binding grooves for individual side chains in the 5-AA segments. These interactions are highly adaptive to different Glu-containing sequences. c, Sequence alignment of the Glu-rich regions from human VKDPs. OCN, GRP, and MGP are shown separately due to their low sequence homology to other VKDPs in the Gla domain. The dots in MGP indicate position of the propeptide that is flanked by Glu-rich sequences. For clarity, the non-consensus Glu residues in OCN, MGP, and GRP, and those at the C-terminal (after +33) of other VKDPs are shown in red letters, and those known to be carboxylated in bold. The sequence alignments were generated by Clustal W v2.1. The grey and light grey shading denote highly and moderately conserved residues, respectively. The sequence logos89 above alignments depict the residue conservation level at each position. Secondary structures (ω-loop and helices h1, h2, and h3) and disulfide bond (S-S) in the Gla domain are indicated below. Inset, Structural location of the Gla residues, Ca2+ ions, and hydrophobic residues (in ω-loop). The crystal structure of human FIX Gla domain (PDB 1NL0)88 is shown with membrane (grey shading) insertion90.
Extended Data Fig. 8 Comparisons of VKGC-TMG2 fusion in bound and unbound states with KH2 or with propeptide.
a, Overall structures with and without bound KH2. b, The c-Glu and KH2 binding sites in the two states. The KH2-bound structure (coloured) and unbound structure (grey) share essentially the same conformation. c, Cellular γ-carboxylation activities of VKGC mutants at the KH2 binding pocket. FIX-Gla-PC is used as the γ-carboxylation reporter. Data are mean ± s.d. from n = 3 or 6 biological replicates. d, Structure of wild-type VKGC in unbound state (coloured as in Fig. 1h). The jellyroll and α/β domains are partially dissociated, creating a gap in between. e, Propeptide binding induces rotation of the jellyroll domain and bent helix (arrows), and an adaptive change in the α/β domain. The unbound state (coloured) and propeptide-bound state (grey) are superimposed by the membrane domain. f, Key residues in the bent helix and L11 that impact propeptide binding. Left, Front view of the propeptide binding site. V502, S510, and W512 are located at the back of α/β and jellyroll domains. Mutations of these residues have been shown to affect propeptide binding91. Right, A 180° rotated view of the back, showing the three residues (in grey) participating in key interactions with surrounding residues (coloured by domains) that stabilize the two domains for propeptide binding. Green lines indicate hydrogen bonds. g-i. Backbone conformational changes between the propeptide bound and unbound structures. The structures of unbound, wild-type VKGC (coloured by domains) and VKGC fused with TMG2 Prop-Glu (red) are superimposed by membrane domain. Changes in the jellyroll domain (g), HL9 and L11 (h), and HL5-6 (i) are discernible from 4.4 Å density map of the unbound VKGC. The red arrows indicate representative backbone changes in the superimposed Prop-Glu bound structure that are out of the map densities of the unbound structure.
Extended Data Fig. 9 Sequence alignment of VKGC homologues.
The alignment is between VKGC homologues from distinct eukaryotic species (Homo, Homo sapiens. Mus, Mus musculus. Xenopus, Xenopus laevis. Takifugu, Takifugu rubripes. Drosophila, Drosophila melanogaster. Conus, Conus textile. Ciona, Ciona intestinalis) and bacterial species (Lepto, Leptospira interrogans. Strep, Streptomyces paromomycinus). The sequence alignments were generated by Clustal W v2.1, and extra sequences in certain homologues are omitted (represented by dots) for simplicity. Key catalytic residues are coloured in red. Below the aligned sequences, the asterisks indicate conserved residues, and single and double dots represent varying levels of sequence similarity. Brown spheres above the alignment indicate VKCFD1 mutations. Residues participating in the bindings of propeptide (P), 5-AA Glu-containing segment (G), and KH2 (K), or forming the hydrophobic tunnel (T) are indicated in coloured letters above the alignment. The protein folding topology (Cyto: cytoplasmic side, Lum: ER lumenal side) and secondary structures (as in Fig. 2b. TM, transmembrane helices. RH, reentrant helices) are denoted further above.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–2 and Supplementary Figs. 1–7.
Supplementary Data 1 (download PDF )
Validation reports of deposited coordinates and maps.
Supplementary Video 1 (download MP4 )
Propeptide binding induces conformational changes that trigger KH2 epoxidation and facilitate binding of the Glu-rich region. The video shows structural changes of VKGC alternating between the unbound state and FIX Prop–Glu-bound state, superimposed by the membrane domain. Propeptide binding between the jellyroll and α/β domains induces the rotation of the jellyroll domain. This rotation causes the associated bent helix to move, in turn altering the conformations of HL5–6 and HL9. These HL regions form part of the KH2-binding pocket, with HL9 also serving as the binding site for the five-amino-acid Glu-containing segment. Thus, propeptide binding reshapes these binding sites to facilitate KH2 epoxidation and the binding of Glu-containing segments.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cao, Q., Ammerman, A., Saimi, M. et al. Molecular basis of vitamin-K-driven γ-carboxylation at the membrane interface. Nature 639, 816–824 (2025). https://doi.org/10.1038/s41586-025-08648-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-08648-1
This article is cited by
-
Molecular basis of vitamin K-dependent protein γ-glutamyl carboxylation
Cell Research (2025)
-
Structural insights into the vitamin K-dependent γ-carboxylation of osteocalcin
Cell Research (2025)
-
Structural insight into bicarbonate-mediated carboxylation by human vitamin K-dependent carboxylase
Nature Communications (2025)
-
Osteocalcin has many tricks to get γ-carboxylated
Cell Research (2025)