Abstract
Distinct hepatocyte subpopulations are spatially segregated along the portal–central axis and are critical to understanding metabolic homeostasis and injury in the liver1. Although several bioactive molecules, including ascorbate and bilirubin, have been described as having a role in directing zonal fates, zonal liver architecture has not yet been replicated in vitro2,3. Here, to evaluate hepatic zonal polarity, we developed a self-assembling zone-specific liver organoid by co-culturing ascorbate- and bilirubin-enriched hepatic progenitors derived from human induced pluripotent stem cells. We found that preconditioned hepatocyte-like cells exhibited zone-specific functions associated with the urea cycle, glutathione synthesis and glutamate synthesis. Single-nucleus RNA-sequencing analysis of these zonally patterned organoids identifies a hepatoblast differentiation trajectory that dictates periportal, interzonal and pericentral human hepatocytes. Epigenetic and transcriptomic analysis showed that zonal identity is orchestrated by ascorbate- or bilirubin-dependent binding of EP300 to TET1 or HIF1α. Transplantation of the self-assembled zonally patterned human organoids improved survival of immunodeficient rats who underwent bile duct ligation by ameliorating the hyperammonaemia and hyperbilirubinaemia. Overall, this multi-zonal organoid system serves as an in vitro human model to better recapitulate hepatic architecture relevant to liver development and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The RNA-seq, ChIP–seq and snRNA-seq data reported in this paper have been deposited to NCBI Gene Expression Omnibus (GEO) with the following accession number: GSE222654. Publicly available datasets were downloaded from the following sources: Camp et al.34 (GSE96981), Guan et al.35 (GSE154883), Harrison et al.36 (GSE245379), Hess et al.37 (GSE207889), Shinozawa et al.28 (GSE141183), Zhang et al.38 (GSE188541) and Smith et al.43 (PRJNA239635). For primary adult and fetal datasets, we downloaded H5AD data deposited in the Human Cell Atlas (https://collections.cellatlas.io/liver-development). The hg38 human reference genome is available at NCBI Genome under accession number GCF_000001405.26. Source data are provided with this paper.
Code availability
Reference codes for bioinformatics analyses can be found at https://github.com/hasanwraeth/Multi-zonal-Liver-Organoids-from-Human-Pluripotent-Stem-Cells.
Change history
22 May 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41586-025-09123-7
References
Ben-Moshe, S. & Itzkovitz, S. Spatial heterogeneity in the mammalian liver. Nat. Rev. Gastroenterol. Hepatol. 16, 395–410 (2019).
Kang, J. et al. Simultaneous deletion of the methylcytosine oxidases Tet1 and Tet3 increases transcriptome variability in early embryogenesis. Proc. Natl Acad. Sci. USA 112, E4236–E4245 (2015).
Kim, S. G. et al. Bilirubin activates transcription of HIF-1α in human proximal tubular cells cultured in the physiologic oxygen content. J. Kor. Med. Sci. 29, S146–S154 (2014).
Martini, T., Naef, F. & Tchorz, J. S. Spatiotemporal metabolic liver zonation and consequences on pathophysiology. Annu. Rev. Pathol. Mech. Dis. 18, 439–466 (2023).
Bartl, M. et al. Optimality in the zonation of ammonia detoxification in rodent liver. Arch. Toxicol. 89, 2069–2078 (2015).
Husson, A., Brasse-Lagnel, C., Fairand, A., Renouf, S. & Lavoinne, A. Argininosuccinate synthetase from the urea cycle to the citrulline–NO cycle. Eur. J. Biochem. 270, 1887–1899 (2003).
Häussinger, D. Nitrogen metabolism in liver: structural and functional organization and physiological relevance. Biochem. J 267, 281–290 (1990).
van Straten, G. et al. Aberrant expression and distribution of enzymes of the urea cycle and other ammonia metabolizing pathways in dogs with congenital portosystemic shunts. PLoS ONE 9, e100077 (2014).
McNaughton, L. et al. Distribution of nitric oxide synthase in normal and cirrhotic human liver. Proc. Natl Acad. Sci. USA 99, 17161–17166 (2002).
Baldelli, S. et al. Glutathione and nitric oxide: key team players in use and disuse of skeletal muscle. Nutrients 11, 2318 (2019).
Yu, Y. et al. A comparative analysis of liver transcriptome suggests divergent liver function among human, mouse and rat. Genomics 96, 281–289 (2010).
Jiang, C. et al. Comparative transcriptomics analyses in livers of mice, humans, and humanized mice define human-specific gene networks. Cells 9, 2566 (2020).
Cunningham, R. P. & Porat-Shliom, N. Liver zonation—revisiting old questions with new technologies. Front. Physiol. 12, 732929 (2021).
Kolbe, E. et al. Mutual zonated interactions of Wnt and Hh signaling are orchestrating the metabolism of the adult liver in mice and human. Cell Rep. 29, 4553–4567.e4557 (2019).
Wahlicht, T. et al. Controlled functional zonation of hepatocytes in vitro by engineering of Wnt signaling. ACS Synth. Biol. 9, 1638–1649 (2020).
Nejak-Bowen, K. N., Zeng, G., Tan, X., Cieply, B. & Monga, S. P. Beta-catenin regulates vitamin C biosynthesis and cell survival in murine liver. J. Biol. Chem. 284, 28115–28127 (2009).
Bacchus, H., Heiffer, M. H. & Altszuler, N. Potentiating effect of ascorbic acid on cortisone-induced gluconeogenesis. Proc. Soc. Exp. Biol. Med. 79, 648–650 (1952).
Greene, Y. J., Harwood, H. J. & Stacpoole, P. W. Ascorbic acid regulation of 3-hydroxy-3-methylglutaryl coenzyme A reductase activity and cholesterol synthesis in guinea pig liver. Biochim. Biophys. Acta 834, 134–138 (1985).
Creeden, J. F., Gordon, D. M., Stec, D. E. & Hinds, T. D. Jr. Bilirubin as a metabolic hormone: the physiological relevance of low levels. Am. J. Physiol. Endocrinol. Metab. 320, E191–E207 (2021).
Gazzin, S. et al. Bilirubin accumulation and Cyp mRNA expression in selected brain regions of jaundiced Gunn rat pups. Pediatr. Res. 71, 653–660 (2012).
Tanii, H. et al. Induction of cytochrome P450 2A6 by bilirubin in human hepatocytes. Pharmacol. Pharm. 4, 182–190 (2013).
Ma, R., Martínez-Ramírez, A. S., Borders, T. L., Gao, F. & Sosa-Pineda, B. Metabolic and non-metabolic liver zonation is established non-synchronously and requires sinusoidal Wnts. eLife 9, e46206 (2020).
Cui, J., Pan, Y. H., Zhang, Y., Jones, G. & Zhang, S. Progressive pseudogenization: vitamin C synthesis and its loss in bats. Mol. Biol. Evol. 28, 1025–1031 (2011).
Reza, H. A. et al. Synthetic augmentation of bilirubin metabolism in human pluripotent stem cell-derived liver organoids. Stem Cell Rep. 18, 2071–2083 (2023).
Horio, F., Ozaki, K., Yoshida, A., Makino, S. & Hayashi, Y. Requirement for ascorbic acid in a rat mutant unable to synthesize ascorbic acid. J. Nutr. 115, 1630–1640 (1985).
Kimura, M. et al. En masse organoid phenotyping informs metabolic-associated genetic susceptibility to NASH. Cell 185, 4216–4232.e4216 (2022).
Ouchi, R. et al. Modeling steatohepatitis in humans with pluripotent stem cell-derived organoids. Cell Metab. 30, 374–384.e376 (2019).
Shinozawa, T. et al. High-fidelity drug-induced liver injury screen using human pluripotent stem cell-derived organoids. Gastroenterology 160, 831–846.e810 (2021).
McCarty, W. J., Usta, O. B. & Yarmush, M. L. A microfabricated platform for generating physiologically-relevant hepatocyte zonation. Sci. Rep. 6, 26868 (2016).
Andrews, T. S. et al. Single-cell, single-nucleus, and spatial rna sequencing of the human liver identifies cholangiocyte and mesenchymal heterogeneity. Hepatol. Commun. 6, 821–840 (2022).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 4383 (2018).
Wesley, B. T. et al. Single-cell atlas of human liver development reveals pathways directing hepatic cell fates. Nat. Cell Biol. 24, 1487–1498 (2022).
Camp, J. G. et al. Multilineage communication regulates human liver bud development from pluripotency. Nature 546, 533–538 (2017).
Guan, Y. et al. A human multi-lineage hepatic organoid model for liver fibrosis. Nat. Commun. 12, 6138 (2021).
Harrison, S. P. et al. Scalable production of tissue-like vascularized liver organoids from human PSCs. Exp. Mol. Med. 55, 2005–2024 (2023).
Hess, A. et al. Single‐cell transcriptomics stratifies organoid models of metabolic dysfunction‐associated steatotic liver disease. EMBO J. 42, e113898 (2023).
Zhang, C. J. et al. A human liver organoid screening platform for DILI risk prediction. J. Hepatol. 78, 998–1006 (2023).
Dann, E., Henderson, N. C., Teichmann, S. A., Morgan, M. D. & Marioni, J. C. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat. Biotechnol. 40, 245–253 (2022).
Ton, M.-L. N. et al. An atlas of rabbit development as a model for single-cell comparative genomics. Nat. Cell Biol. 25, 1061–1072 (2023).
Thakur, A. et al. Hepatocyte nuclear factor 4-alpha is essential for the active epigenetic state at enhancers in mouse liver. Hepatology 70, 1360–1376 (2019).
Raisner, R. et al. Enhancer activity requires CBP/P300 bromodomain-dependent histone H3K27 acetylation. Cell Rep. 24, 1722–1729 (2018).
Smith, R. P. et al. Genome-wide discovery of drug-dependent human liver regulatory elements. PLoS Genet. 10, e1004648 (2014).
Ölander, M. et al. Hepatocyte size fractionation allows dissection of human liver zonation. J. Cell. Physiol. 236, 5885–5894 (2021).
Kang, S. W. S. et al. A spatial map of hepatic mitochondria uncovers functional heterogeneity shaped by nutrient-sensing signaling. Nat. Commun. 15, 1799 (2024).
Mariotti, V., Strazzabosco, M., Fabris, L. & Calvisi, D. F. Animal models of biliary injury and altered bile acid metabolism. Biochim. Biophys. Acta 1864, 1254–1261 (2018).
Claeys, W. et al. A mouse model of hepatic encephalopathy: bile duct ligation induces brain ammonia overload, glial cell activation and neuroinflammation. Sci. Rep. 12, 17558 (2022).
Goto, Y., Ohashi, K., Utoh, R., Yamamoto, M. & Okano, T. Hepatocyte transplantation through the hepatic vein: a new route of cell transplantation to the liver. Cell Transplant. 20, 1259–1270 (2011).
Paris, J. & Henderson, N. C. Liver zonation, revisited. Hepatology 76, 1219–1230 (2022).
Scheidecker, B. et al. Induction of in vitro metabolic zonation in primary hepatocytes requires both near-physiological oxygen concentration and flux. Front. Bioeng. Biotechnol. 8, 524 (2020).
Bartl, M. et al. Model-based optimization to explain liver zonation in nitrogen metabolism. In 55th International Scientific Colloquium 235–240 (TU Ilmenau, 2021).
Harrison, S. P. et al. Liver organoids: recent developments, limitations and potential. Front. Med. https://doi.org/10.3389/fmed.2021.574047 (2021).
Wei, Y. et al. Liver homeostasis is maintained by midlobular zone 2 hepatocytes. Science 371, eabb1625 (2021).
He, L. et al. Proliferation tracing reveals regional hepatocyte generation in liver homeostasis and repair. Science 371, eabc4346 (2021).
Li, W., Li, L. & Hui, L. Cell plasticity in liver regeneration. Trends Cell Biol. 30, 329–338 (2020).
Brosch, M. et al. Epigenomic map of human liver reveals principles of zonated morphogenic and metabolic control. Nat. Commun. 9, 4150 (2018).
Xu, W. et al. Hypoxia activates Wnt/β-catenin signaling by regulating the expression of BCL9 in human hepatocellular carcinoma. Sci. Rep. 7, 40446 (2017).
Kietzmann, T. Metabolic zonation of the liver: the oxygen gradient revisited. Redox Biol. 11, 622–630 (2017).
Iansante, V., Mitry, R. R., Filippi, C., Fitzpatrick, E. & Dhawan, A. Human hepatocyte transplantation for liver disease: current status and future perspectives. Pediatr. Res. 83, 232–240 (2018).
Reza, H. A., Okabe, R. & Takebe, T. Organoid transplant approaches for the liver. Transpl. Int. 34, 2031–2045 (2021).
Wu, H., Kirita, Y., Donnelly, E. L. & Humphreys, B. D. Advantages of single-nucleus over single-cell RNA sequencing of adult kidney: rare cell types and novel cell states revealed in fibrosis. J. Am. Soc. Nephrol. 30, 23–32 (2019).
He, L. et al. Transcriptional co-activator p300 maintains basal hepatic gluconeogenesis. J. Biol. Chem. 287, 32069–32077 (2012).
Acknowledgements
The authors thank A. Kodaka for scientific illustration materials; N. Saiki for bioinformatic support; Y.-C. Hu and C. Mayhew for Gulo-edited hiPS cell generation; all members of the Takebe, Zorn, Helmrath and Wells laboratories for their support and feedback; A. Potter for expertise in single-cell isolation and troubleshooting; and the thesis committee (J. A. Bezerra, V. Hwa, R. C. C. Quillin, J. Tchieu and J. M. Wells) for their continued support of H.A.R. and constructive criticism. We acknowledge the CCHMC Confocal Imaging Core (RRID:SCR_022628), Pathology Research Core (RRID:SCR_022637), Pluripotent Stem Cell Facility (RRID:SCR_022634), Transgenic Animal and Genome Editing Core (RRID:SCR_022642), Veterinary Services Facility, and Single Cell Genomics Core (RRID:SCR_022653) for providing gene-expression profiling and analysis. Human liver and pancreas were received from the islet cell laboratory of A. N. Balamurugan. We extend our deepest gratitude to the courageous families who generously donated their loved one’s organs and tissues for biomedical research. Thanks to Network of Hope, Louisville/Cincinnati; and Lifeline of Ohio, Columbus for supporting these special families. This work was supported by Cincinnati Children’s Research Foundation CURE grant, the Falk Transformational Award Program, New York Stem Cell Foundation, NIH Director’s New Innovator Award (DP2 DK128799-01) and R01DK135478 to T.T. This work was also supported by an NIH grant UG3/UH3 DK119982, Cincinnati Center for Autoimmune Liver Disease Fellowship Award, PHS Grant P30 DK078392 (Integrative Morphology Core and Pluripotent Stem Cell and Organoid Core) of the Digestive Disease Research Core Center in Cincinnati, Takeda Science Foundation Award, Mitsubishi Foundation Award, Cannon Foundation Award and by Japan Agency for Medical Research and Development (AMED) under grant numbers: JP24bm1223006, JP24ym0126809, JP21bm0404045, JP18fk0210037, JP18bm0704025, JP23gm1610005, JP23fk0210106, JP24gm1210012, JP24fk0210150 and JP21fk0210060, JST Moonshot: JPMJMS2022 and JPMJMS2033, JSPS KAKENHI: JP18H02800, 19K22416, 21H04822 and World Premier International Research Center Initiative (WPI) PRIMe, MEXT, Japan.
Author information
Authors and Affiliations
Contributions
H.A.R. designed and performed research, analysed data, performed the bioinformatics analyses and wrote the paper. C.S., A.A.R., K.I., S.S. and K.G. assisted in staining, ChIP, snRNA-seq and transplantation experiments and revised the paper. A.B. and J.M. designed research and revised the paper. T.T. designed research and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Jan Tchorz, Adrian Teo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Intracellular redox management enables CPS1+ hepatocyte specification in HLOs.
a) H&E histology, GLS2 IHC, and GS IHC images of liver sections are shown in panels from ODS od/od (GULO mutant) rat treated with 0.2% Ascorbic acid (AsA), ODS od/od rat treated without AsA. Scale bars indicate 100 µm. The graph shows the GLS2 or GS positive area ratio versus the hematoxylin positive area in ODS od/od (GULO mutant) rat treated with 0.2% Ascorbic acid (AsA) and ODS od/od rat treated without AsA. Data points are shown for GLS2 area at portal vein (37 portal vein sections of + AsA rats, 27 portal vein sections of - AsA rats) and GS area in 8 images of +/− AsA rat. b) Schematic for development of Z1-HLOs and doxycycline induction to induce CPS1+ hepatocyte specification (left). Brightfield and fluorescence images of mCherry expression in ascorbate depleted Dox (100 ng/ml) treated Z1-HLOs compared to HLOs with ascorbic acid depletion at D20 and control HLOs (right). c) ELISA for mGULO protein concentration in Dox treated Z1-HLOs compared to control HLOs. (n = 9 independent experiments). d) Cellular Antioxidant concentration in Dox treated Z1-HLOs compared to control HLOs. (n = 9 independent experiments). e) ROS levels in Dox treated and extracellular ascorbate induced Z1-HLOs compared to control HLOs. (n = 9 independent experiments). f) RT-qPCR of genes for Z1-HLOs. (mean ± SD and n = 9 independent experiments). g) Albumin ELISA for Z1-HLOs treated with Dox compared to control HLOs and PHH normalized by cell viability. (n = 9 independent experiments). h) Immunofluorescence images of Dox treated Z1-HLOs for CPS1, ACSS2 and CDH1 compared to control HLOs and primary liver. Scale bar indicates 200 µm. (n = 3 independent experiments). In c-e, g data are represented as boxplots where the middle line is the median, the lower and upper hinges correspond to the first and third quartiles, the upper and lower whisker extends from the hinge to the largest and smallest value respectively no further than 1.5 × IQR from the hinge (where IQR is the inter-quartile range). c-e, f, g, one-way ANOVA with multiple comparisons and Tukey’s correction.
Extended Data Fig. 2 Low dose bilirubin promotes GLUL+ hepatocyte specification in HLOs.
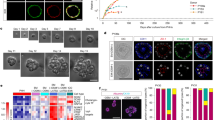
a) Schematic for low dose bilirubin treatment and Z3-HLO development to induce GLUL+ expression. b) Cell viability assay with different concentration of bilirubin to titrate dose for maximal viability. (n = 9 independent experiments). c) Brightfield image of Z3-HLOs treated with low dose bilirubin (1 mg/L) compared to control, and luminal outline using ImageJ, arrows indicate luminal projections that are similar to bile canaliculi found in human liver. Scale bar indicates 200 µm. d) Comparison of size and circularity of lumen of the control and 1 mg/L bilirubin treated Z3-HLOs. (Data is mean ± SD, n = 9 independent experiments). e) Heatmap of Zone 3 genes from RNAseq dataset for bilirubin treated Z3-HLOs compared to control. f) RT-qPCR of genes for Z3-HLOs. (mean ± SD and n = 9 independent experiments). g) CYP3A4 activity assay in response to Rifampicin in control, Z1-, Z3-HLOs, and PHH (left). CYP1A2 activity assay in response to Omeprazole in control, Z1-, Z3-HLOs, and PHH (right). (n = 9 independent experiments). h) Immunofluorescence images of Z3-HLOs for GLUL, NR3C1 and CDH1 compared to control HLOs and primary liver. Scale bar indicates 200 µm. (n = 3 independent experiments). i) Heatmap of Z1- and Z3- HLOs depicting expression of zonal genes and lack of consensus expression of markers such as ARG1 and AKR1C1. j) Caspase 3 activity assay in Z3-HLOs (left) and cell viability assay in Z3-HLOs compared to Z1-HLOs after treatment with Zone 1 toxin (right). (n = 9 independent experiments). k) Caspase 3 activity assay in Z3-HLOs (left) and cell viability assay in Z3-HLOs compared to Z1-HLOs after treatment with Zone 3 toxin (right). (n = 9 independent experiments). In b, g, j, k data are represented as boxplots where the middle line is the median, the lower and upper hinges correspond to the first and third quartiles, the upper and lower whisker extends from the hinge to the largest and smallest value respectively no further than 1.5 × IQR from the hinge (where IQR is the inter-quartile range). b,f, g,j,k one-way ANOVA with multiple comparisons and Tukey’s correction. d, Kruskal–Wallis test (left) and unpaired two-tailed Student’s t-test (right).
Extended Data Fig. 3 Bilirubin induced fusion requires close proximity and cytoskeletal signaling.
a) Brightfield images of bilirubin induced fusion in high density HLOs compared to low density and no bilirubin treatment. Scale bar indicates 200 µm. b) Comparison of mean segment length in high density HLOs compared to low density and no bilirubin treatment. (n = 9 independent experiments). c) Brightfield and live staining images (NucBlue: Blue, Cytoskeleton: White) show progression of organoid fusion after continued treatment with bilirubin (1 mg/L). Scale bar indicates 200 µm. (n = 3 independent experiments). d) Comparison of mean segment length of the HLOs from D1 to D7. (n = 9 independent experiments). e) NOTCH activity assay in bilirubin treated HLOs compared to control. (n = 9 independent experiments). f) Percentage of fused organoids after bilirubin treatment in DAPT (Notch inhibitor) and NSC (Ezrin Inhibitor, NSC668394) treated HLOs compared to control. (n = 9 independent experiments). g) Brightfield images of bilirubin induced fused HLOs compared to DAPT or NSC668394 treatment and control HLOs. Scale bar indicates 200 µm. (n = 3 independent experiments). h) CLF assay for self-assembled organoids compared to control. Scale indicates 200 µm. (n = 3 independent experiments). i) Percentage of fused organoids for each type of organoid. (n = 9 independent experiments). In e data is represented as boxplot where the middle line is the median, the lower and upper hinges correspond to the first and third quartiles, the upper and lower whisker extends from the hinge to the largest and smallest value respectively no further than 1.5 × IQR from the hinge (where IQR is the inter-quartile range). b,d, Kruskal-Wallis with multiple comparisons and Dunn-Holland-Wolfe correction. e, unpaired two-tailed Student’s t-test. i, one-way ANOVA with multiple comparisons and Tukey’s correction.
Extended Data Fig. 4 Immunostaining of mZ-HLOs compared to neonatal liver show similar features.
a) Brightfield images of fused HLOs following continued treatment with bilirubin compared to single dose treatment after 10 days. (n = 3 independent experiments). b) Immunofluorescence images (bottom) of mZ-HLOs depicting GFP, mCherry, PROX1 and A1AT. Scale bar indicates 200 µm. (n = 3 independent experiments). c) Immunofluorescence images of mZ-HLOs for pan liver markers: TUBA1A, CTNBB1, luminal marker: ZO-1 (depicting continuous lumen); Zone 1 markers: ARG1 and SLBP; Zone 2 marker TERT; Zone 3 markers: AHR, and MRP2; and Cholangiocyte marker CK7. Scale bar indicates 200 µm. Numbers on the bar indicate the percentage of fused organoids that express dual and single positivity for the indicated antigen staining. (n = 3 independent experiments). d) Immunohistochemistry images of neonatal liver sections for pan liver marker TUBA1A; Cholangiocyte marker CK7; Zone 1 markers: ARG1, SLBP, and GLS2; Zone 2 marker TERT; and Zone 3 markers: AHR, ALDH6A1 and MRP2. Scale bar indicates 200 µm. (P: Portal vein, C: Central vein). (n = 3 independent experiments).
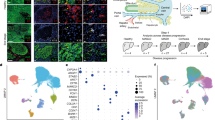
Extended Data Fig. 5 Single cell profiling of mZ-HLOs indicate the emergence of zonal like populations.
a) UMAP plot with the major populations (hepatocytes, cholangiocytes, endothelial cells, macrophages, stellate cells, and mesenchyme) of all nuclei in mZ-HLOs. b) Distinct expression profile all populations in mZ-HLOs. The size of the circle indicates the percentage of nuclei in each population expressing each gene. The color represents the average expression level for the indicated gene. c) Heatmap showing scaled mean expression of all genes in each cluster. Top 10 marker genes in each cluster have been added as labels. d) Expression of known hepatoblast and zonal hepatocyte marker genes in each population. e) Violin plot for expression of AFP (hepatoblast gene), GSS (interzonal hepatocytes), GHR (pericentral hepatocyte), and GLS2 (periportal hepatocyte).
Extended Data Fig. 6 Pseudo-spatial profiling of mZ-HLOs show similarity of zonal expression to primary liver tissue.
a) Spatial plot for TAT (zone 1), HAMP (zone 2), and CYP3A4 (zone 3) markers in 10X Xenium healthy human liver dataset (publicly available dataset). b) UMAP plot of mZ-HLO with hepatocyte populations (top) and distribution of replicate data (bottom). c) Feature plot for TAT (zone 1), HAMP (zone 2), and CYP3A4 (zone 3) markers. d) UMAP plot for zonal hepatocyte populations from primary liver (Andrews et al.30) and mZ-HLOs integrated together (top). UMAP plot depicting distribution for total hepatocyte populations from primary liver and mZ-HLOs integrated together (bottom). e) Expression of known hepatoblast and zonal hepatocyte marker genes in mZ-HLOs benchmarked against Andrews et al.30 snRNAseq dataset. f) UMAP plot for all cell types (inset: sample distribution) from primary liver datasets and mZ-HLOs integrated together. g) Feature plot for GLS2 (zone 1), HAMP (zone 2), and GLUL (zone 3) markers.
Extended Data Fig. 7 RNA velocity and pseudotime analysis in mZ-HLOs.
a) Phase portrait of AFP, GLUL, and GLS2 depicting the dynamics of the gene splicing in the nuclei with the velocity and expression of AFP, GLUL, and GLS2 in nuclei as feature plots. b) SOM (Self Organizing Map) of single-nuclei transcriptome-derived zonation profiles for mZ-HLOs based on the different populations. c) Boxplot showing the pseudotime of each nuclei population in mZ-HLOs. d) Pathway enrichment analysis examining which cellular pathways represented in the hepatoblast, pericentral, periportal, and interzonal hepatocyte populations. Circles (nodes) represent pathways, sized by the number of genes included in that pathway. Related pathways, indicated by light blue lines, are grouped into a theme (black circle) and labeled. Intra-pathway and inter-pathway relationships are shown in light blue and represent the number of genes shared between each pathway. In c data is represented as boxplot where the middle line is the median, the lower and upper hinges correspond to the first and third quartiles, the upper and lower whisker extends from the hinge to the largest and smallest value respectively no further than 1.5 × IQR from the hinge (where IQR is the inter-quartile range), while data beyond the end of the whiskers are outlying points that are plotted individually.
Extended Data Fig. 8 EP300 differentially regulates zonal genes in mZ-HLOs in conjunction to distinct transcription factors.
a) Peak density plots showing EP300 bound loci, a marker of active enhancers. Profile plot of all peaks are in the top panel. b) Genome browser view of HNF4A (pan marker) showing the EP300 ChIPseq peak. c) Genome browser view of CTNNB1 (pan marker) showing the EP300 ChIPseq peak. d) Genome browser view of SLBP (zone 1 gene) showing the EP300 ChIPseq peak. e) Genome browser view of AKR1C1 (zone 2 gene) showing the EP300 ChIPseq peak. f) Genome browser view of GHR (zone 3 gene) showing the EP300 ChIPseq peak. g) Top 10 upregulated Gene Ontology terms (Biological Process) for the gene regulated bound by EP300 in the Z1-HLOs. h) Top 10 upregulated Gene Ontology terms (Biological Process) for the gene regulated bound by EP300 in the Z3-HLOs.
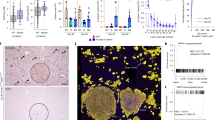
Extended Data Fig. 9 EP300 and partner transcription factors are important for zonal liver development.
a) Experimental timeline for testing role of EP300 in zonal liver development using Ad-shp300 (adenoviral vector for p300 shRNA, top). H&E stain for sections from rat liver injected with Ad-shSCR (adenoviral vector for scrambled shRNA, middle) and Ad-shp300 (adenoviral vector for p300 shRNA, bottom). Scale bar indicates 200 µm. (P: Portal vein, C: Central vein). b) ICH stain for PROX1, ARG1, and GLUL of sections from rat liver injected with Ad-shSCR (adenoviral vector for scrambled shRNA, top) and Ad-shp300 (adenoviral vector for p300 shRNA, bottom). Scale bar indicates 200 µm. (P: Portal vein, C: Central vein). c) RT-qPCR of ALB, ACSS2, ASL, CPS1, and OTC (zone 1) gene for Z1- and Z3-HLOs compared to freshly isolated PHH, H20 (20 um periportal hepatocytes) and H40 (40 um pericentral hepatocytes) (Data is mean ± SD, n = 9 independent experiments). d) RT-qPCR of ALDH1A2, ALDH6A1, HIF1A, SREBF1, and GLUL (zone 3) gene for Z1- and Z3-HLOs compared to freshly isolated PHH, H20 (20 um periportal hepatocytes) and H40 (40 um pericentral hepatocytes) (Data is mean ± SD, n = 9 independent experiments). e) EP300-TF ChIP-reChIP-PCR for H20 (20 um periportal hepatocytes) and H40 (40 um pericentral hepatocytes). (n = 3 independent experiments). f) EP300-TF ChIP-reChIP-qPCR for samples in (e). Data are mean ± SD, n = 9 independent experiments. g) Schematic for bilirubin and ascorbate mediated distinct epigenetic regulation leading to differential gene expression. c, d, one-way ANOVA with multiple comparisons and Tukey’s correction. f, unpaired two-tailed Student’s t-test.
Extended Data Fig. 10 Interzonal dependent lipid and glucose metabolism in mZ-HLOs.
a) Triglyceride assay for mZ-HLOs with and without Firsocostat treatment compared to Z1-, Z3-, control HLOs, and PHH (n = 9 independent experiments). b) Lipase activity assay for mZ-HLOs with and without Firsocostat treatment compared to Z1-, Z3-, control HLOs, and PHH (n = 9 independent experiments). c) Glucose assay for mZ-HLOs with and without FBPi treatment compared to Z1-, Z3-, control HLOs, and PHH (n = 9 independent experiments). d) Glucokinase activity assay for mZ-HLOs with and without FBPi treatment compared to Z1-, Z3-, control HLOs, and PHH (n = 9 independent experiments). In a, c data are represented as boxplots where the middle line is the median, the lower and upper hinges correspond to the first and third quartiles, the upper and lower whisker extends from the hinge to the largest and smallest value respectively no further than 1.5 × IQR from the hinge (where IQR is the inter-quartile range). a,c, one-way ANOVA with multiple comparisons and Tukey’s correction.
Extended Data Fig. 11 mZ-HLOs exhibit zone specific regenerative potential in response to toxins.
a) Experimental timeline for testing zonal regenerative potential of mZ-HLOs in response to zone specific toxins. b) Immunofluorescence images of mZ-HLOs for proliferative marker Ki-67, GLUL, ARG1 and HAMP in response to Allyl Alcohol (Zone 1 toxin, left). Immunofluorescence images of mZ-HLOs for proliferative marker Ki-67, GLUL and ARG1 in response to Acetaminophen (Zone 3 toxin). Scale bar indicates 200 µm. c) Comparison of Ki-67 + nuclei in different fluorescent regions in response to zone specific toxins. (Data is mean ± SD, n = 9 independent experiments). d) Comparison of length of different fluorescent regions in response to zone specific toxins. (Data is mean ± SD, n = 9 independent experiments). e) Immunofluorescence images of CPS1, TET1, GLUL, NR3C1, and mCherry in Z1-HLOs with Dox treatment and after Dox withdrawal and persistent bilirubin treatment at D20 and 25. Scale bar indicates 200 µm. f) RT-qPCR of ACSS2, CPS1, ALDH1A2 and GLUL gene for Z3-HLOs with Dox (Dox +) and with bilirubin after Dox withdrawal (Dox – Bilirubin +) compared to control HLOs (Data is mean ± SD, n = 9 independent experiments). c, d, unpaired two-tailed Student’s t-test. f, one-way ANOVA with multiple comparisons and Tukey’s correction.
Extended Data Fig. 12 mZ-HLOs invade into the liver parenchyma of RRG rats after transplantation.
a) Immunofluorescence images for human TUBA1A of mZ-HLOs transplanted in RRG rat liver. Scale bar indicates 200 µm. (n = 3 independent experiments). b) Immunofluorescence images for human ASGR1 of mZ-HLOs transplanted in RRG rat liver. Scale bar indicates 200 µm. (n = 3 independent experiments). c) Immunofluorescence images for GFP, mCherry, human ARG1 and GLUL of mZ-HLOs transplanted in RRG rat liver. Scale bar indicates 200 µm. (n = 3 independent experiments). d) Immunofluorescence images for human ASGR1 of Z1 and Z3-HLOs transplanted in RRG rat liver through the portal vein and inferior vena cava. Scale bar indicates 200 µm. Numbers on the bar indicate the ratio of the area of integrated organoids and total area of the liver parenchyma in view in 103 pixel2 (Data is mean ± SD, n = 9 independent experiments).
Supplementary information
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Reza, H.A., Santangelo, C., Iwasawa, K. et al. Multi-zonal liver organoids from human pluripotent stem cells. Nature 641, 1258–1267 (2025). https://doi.org/10.1038/s41586-025-08850-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-08850-1
This article is cited by
-
Advances in liver and pancreas organoids: how far we have come and where we go next
Nature Reviews Gastroenterology & Hepatology (2026)
-
Human assembloids recapitulate periportal liver tissue in vitro
Nature (2026)
-
Metabolic dysfunction-associated steatotic liver disease and steatohepatitis-associated hepatocarcinoma preclinical models
Nature Reviews Gastroenterology & Hepatology (2026)
-
Single-cell sequencing and organoids: applications in organ development and disease
Molecular Biomedicine (2025)
-
Human liver immunology: from in vitro models to new insights
Cellular & Molecular Immunology (2025)