Abstract
Despite accumulating evidence that bat-derived coronaviruses often require intermediate hosts to facilitate transmission to humans1, the potential role of fur animals in zoonotic coronavirus spillovers has largely been overlooked2. Here we report the isolation and characterization of a previously undescribed mink respiratory coronavirus (MRCoV) from farmed minks with pneumonia. Notably, MRCoV uses angiotensin-converting enzyme 2 (ACE2) as an entry receptor and can infect mink, bat, monkey and human cells. Cryo-electron microscopy analyses revealed that the MRCoV receptor-binding domain (RBD) binds to the same interface on ACE2 receptors as the RBD of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) despite structural differences. We identify the key determinants on the RBD of MRCoV and ACE2 that confer efficient binding. HKU5-33S, a bat coronavirus closely related to MRCoV, uses ACE2 of the bat Pipistrellus abramus for cell entry and requires only two amino acid substitutions to adapt to mink ACE2. SARS-CoV-2 protease and polymerase inhibitors potently block MRCoV infection, thereby indicating a potential therapeutic strategy. Collectively, these findings enhance our understanding of coronavirus receptor dynamics and highlight their zoonotic potential. Given the risks posed by fur farms as reservoirs for emerging pathogens, our study underscores the need for enhanced surveillance to mitigate future coronavirus outbreaks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are available in the main text or the supplementary materials. The cryo-EM maps and atomic coordinates of MRCoV RBD in complex with NvACE2 have been deposited into the Electron Microscopy Data Bank and the PDB under the accession codes EMD-60524 and 8ZWE, respectively. The MRCoV sequence has been deposited in GenBank (accession number PV174919) and the National Microbiology Data Center (accession number NMDCN0007NAP). The raw data have been deposited into the Genome Sequence Archive of the National Genomics Data Center, Beijing Institute of Genomics, Chinese Academy of Sciences/China National Center for Bioinformation under the BioProject (accession number PRJCA036572).
References
Cui, J., Li, F. & Shi, Z. L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).
Zhao, J. et al. Farmed fur animals harbour viruses with zoonotic spillover potential. Nature 634, 228–233 (2024).
Essalmani, R. et al. Distinctive roles of furin and TMPRSS2 in SARS-CoV-2 infectivity. J. Virol. 96, e0012822 (2022).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292 (2020).
Lv, Z. et al. Structural basis for neutralization of SARS-CoV-2 and SARS-CoV by a potent therapeutic antibody. Science 369, 1505–1509 (2020).
Starr, T. N. et al. ACE2 binding is an ancestral and evolvable trait of sarbecoviruses. Nature 603, 913–918 (2022).
Lu, G. et al. Molecular basis of binding between novel human coronavirus MERS-CoV and its receptor CD26. Nature 500, 227–231 (2013).
Xiong, Q. et al. Close relatives of MERS-CoV in bats use ACE2 as their functional receptors. Nature 612, 748–757 (2022).
Ma, C. B. et al. Multiple independent acquisitions of ACE2 usage in MERS-related coronaviruses. Cell 188, 1693–1710 (2025).
Peeri, N. C. et al. The SARS, MERS and novel coronavirus (COVID-19) epidemics, the newest and biggest global health threats: what lessons have we learned? Int. J. Epidemiol. 49, 717–726 (2020).
Li, W. et al. Bats are natural reservoirs of SARS-like coronaviruses. Science 310, 676–679 (2005).
Song, H. D. et al. Cross-host evolution of severe acute respiratory syndrome coronavirus in palm civet and human. Proc. Natl Acad. Sci. USA 102, 2430–2435 (2005).
Anthony, S. J. et al. Further evidence for bats as the evolutionary source of Middle East respiratory syndrome coronavirus. mBio 8, 00373-17 (2017).
Dudas, G., Carvalho, L. M., Rambaut, A. & Bedford, T. MERS-CoV spillover at the camel–human interface. eLife 7, e31257 (2018).
Li, F. Structural analysis of major species barriers between humans and palm civets for severe acute respiratory syndrome coronavirus infections. J. Virol. 82, 6984–6991 (2008).
Liu, W. J. et al. Surveillance of SARS-CoV-2 at the Huanan Seafood Market. Nature 631, 402–408 (2024).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020).
Temmam, S. et al. Bat coronaviruses related to SARS-CoV-2 and infectious for human cells. Nature 604, 330–336 (2022).
Liu, M. Q. et al. A SARS-CoV-2-related virus from Malayan pangolin causes lung infection without severe disease in human ACE2-transgenic mice. J. Virol. 97, e01719–e01722 (2023).
Zhou, H. et al. Identification of novel bat coronaviruses sheds light on the evolutionary origins of SARS-CoV-2 and related viruses. Cell 184, 4380–4391 (2021).
Wells, H. L. et al. The coronavirus recombination pathway. Cell Host Microbe 31, 874–889 (2023).
Cui, X. et al. Virus diversity, wildlife–domestic animal circulation and potential zoonotic viruses of small mammals, pangolins and zoo animals. Nat. Commun. 14, 2488 (2023).
Jahid, M. J., Bowman, A. S. & Nolting, J. M. SARS-CoV-2 outbreaks on mink farms—a review of current knowledge on virus infection, spread, spillover, and containment. Viruses 16, 81 (2024).
Oude Munnink, B. B. et al. Transmission of SARS-CoV-2 on mink farms between humans and mink and back to humans. Science 371, 172–177 (2021).
Rabalski, L. et al. Zoonotic spill-over of SARS-CoV-2: mink-adapted virus in humans. Clin. Microbiol. Infect. 28, 451.e1–451.e4 (2022).
Hammer, A. S. et al. SARS-CoV-2 transmission between mink (Neovison vison) and humans, Denmark. Emerg. Infect. Dis. 27, 547–551 (2021).
Koopmans, M. SARS-CoV-2 and the human–animal interface: outbreaks on mink farms. Lancet Infect. Dis. 21, 18–19 (2021).
Mallapaty, S. The search for animals harbouring coronavirus—and why it matters. Nature 591, 26–28 (2021).
Pickering, B. et al. Divergent SARS-CoV-2 variant emerges in white-tailed deer with deer-to-human transmission. Nat. Microbiol. 7, 2011–2024 (2022).
van Aart, A. E. et al. SARS-CoV-2 infection in cats and dogs in infected mink farms. Transbound. Emerg. Dis. 69, 3001–3007 (2022).
Xia, L., Zhang, Y., Li, Y., Li, D. & Zhou, Q. Molecular insights into cross-species spillover of coronavirus HKU5 via ACE2 receptor recognition. Preprint at bioRxiv https://doi.org/10.1101/2025.01.10.632062 (2025).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Park, Y. J. et al. Molecular basis of convergent evolution of ACE2 receptor utilization among HKU5 coronaviruses. Cell 188, 1711–1728 (2025).
Zielinska, D. F., Gnad, F., Wiśniewski, J. R. & Mann, M. Precision mapping of an in vivo N-glycoproteome reveals rigid topological and sequence constraints. Cell 141, 897–907 (2010).
Wang, Q. et al. Bat origins of MERS-CoV supported by bat coronavirus HKU4 usage of human receptor CD26. Cell Host Microbe 16, 328–337 (2014).
Han, X. et al. Structure of the S1 subunit C-terminal domain from bat-derived coronavirus HKU5 spike protein. Virology 507, 101–109 (2017).
Wu, K., Li, W., Peng, G. & Li, F. Crystal structure of NL63 respiratory coronavirus receptor-binding domain complexed with its human receptor. Proc. Natl Acad. Sci. USA 106, 19970–19974 (2009).
Zhai, X. et al. Comparison of severe acute respiratory syndrome coronavirus 2 spike protein binding to ACE2 receptors from human, pets, farm animals, and putative intermediate hosts. J. Virol. 94, e00831-20 (2020).
Hossain, M. G., Javed, A., Akter, S. & Saha, S. SARS-CoV-2 host diversity: an update of natural infections and experimental evidence. J. Microbiol. Immunol. Infect. 54, 175–181 (2021).
Chen, J. et al. A bat MERS-like coronavirus circulates in pangolins and utilizes human DPP4 and host proteases for cell entry. Cell 186, 850–863 (2023).
Chen, J. et al. Bat-infecting merbecovirus HKU5-CoV lineage 2 can use human ACE2 as a cell entry receptor. Cell 188, 1729–1742 (2025).
Lau, S. K. et al. Genetic characterization of betacoronavirus lineage C viruses in bats reveals marked sequence divergence in the spike protein of pipistrellus bat coronavirus HKU5 in Japanese pipistrelle: implications for the origin of the novel Middle East respiratory syndrome coronavirus. J. Virol. 87, 8638–8650 (2013).
Catanzaro, N. J. et al. ACE2 from Pipistrellus abramus bats is a receptor for HKU5 coronaviruses. Preprint at bioRxiv https://doi.org/10.1101/2024.03.13.584892 (2024).
Alfajaro, M. M. et al. HKU5 bat merbecoviruses use divergent mechanisms to engage bat and mink ACE2 as entry receptors. Preprint at bioRxiv https://doi.org/10.1101/2025.02.12.637862 (2025).
Ju, X. et al. A novel cell culture system modeling the SARS-CoV-2 life cycle. PLoS Pathog. 17, e1009439 (2021).
Reed, L. & Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Hygiene 27, 493–497 (1938).
Yang, Z. Estimating the pattern of nucleotide substitution. J. Mol. Evol. 39, 105–111 (1994).
Yang, Z. Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. J. Mol. Evol. 39, 306–314 (1994).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Ou, X. et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 11, 1620 (2020).
Chen, M. & Zhang, X. E. Construction and applications of SARS-CoV-2 pseudoviruses: a mini review. Int. J. Biol. Sci. 17, 1574–1580 (2021).
Sheahan, T. P. et al. An orally bioavailable broad-spectrum antiviral inhibits SARS-CoV-2 in human airway epithelial cell cultures and multiple coronaviruses in mice. Sci. Transl. Med. 12, eabb5883 (2020).
Owen, D. R. et al. An oral SARS-CoV-2 Mpro inhibitor clinical candidate for the treatment of COVID-19. Science 374, 1586–1593 (2021).
Yu, Y., Ju, X. & Ding, Q. A nucleocapsid-based transcomplementation cell culture system of SARS-CoV-2 to recapitulate the complete viral life cycle. Bio. Protoc. 11, e4257 (2021).
Li, Y. et al. Long-term effects of omicron BA.2 breakthrough infection on immunity-metabolism balance: a 6-month prospective study. Nat. Commun. 15, 2444 (2024).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Acknowledgements
We thank Q. Ding for providing the SARS-CoV-2 GFP/ΔN system; W. Zhang, J. Liang and J. Liu for technical support; and staff members of instrument platforms and/or biosafety laboratories from the Changchun Veterinary Research Institute of the Chinese Academy of Agricultural Sciences, and the Institute of Microbiology of the Chinese Academy of Science for their advice and assistance. Collaborations involving restricted materials should be coordinated with Dr. Bi. The project was supported by the National Key R&D Program of China (2025YFE0101000, 2023YFC2307500 and 2024YFC2607501), the National Natural Science Foundation of China (NSFC) Distinguished Young Scholar (32425053), the NSFC (32170158, 32100128, 32000659 and 32100128), the Major Project of Guangzhou National Laboratory (GZNL2023A01001), the National Science and Technology Infrastructure of China (National Pathogen Resource Center-NPRC-32), the Innovation Team and Talents Cultivation Program of National Administration of Traditional Chinese Medicine (ZYYCXTD-D-202208), the Natural Science Foundation of Jiangsu Province (BK20240087, BK20200545, BK20200553, BK20241578 and BK20241565), the CAMS Innovation Fund for Medical Sciences (2020-I2M-5-001), the Postdoctoral Fellowship Program of CPSF (grant numbers GZB20240314 and GZB20240316), the China Postdoctoral Science Foundation (2024M751428) and the Jiangsu Funding Program for Excellent Postdoctoral Talent (2024ZB266 and 2024ZB831). G.M.D. acknowledges support from the project KP-06-IP-Kitai/3 entitled ‘Study on broad-spectrum therapeutic drugs for zoonotic RNA viral infectious diseases’ funded by the Bulgarian National Science Fund. S.S. has moved to Fudan University and acknowledges the National Key Research and Development Program of China (grant no. 2021YFD1801101), for contributing conceptual input, particularly regarding the identification and molecular characterization of pathogens from farmed animals, and also thanks Fudan University for its support.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.S., Y.B. and S.Z. Plasmid construction: Yutong Wang, X.-x.L., Y.X. and J.W. Western blotting and immunofluorescence assays: X.-x.L., X. Zhang and X. Zhai. Protein purification: X.M., Yu Wang, W.J., Yanjun Wang, Y.X. and J.W. SPR assays: W.J. and Yanjun Wang. Structural data analyses: M.V. and W.J. PV experiments: H.J., N.W., X.M. and Yu Wang. Virus isolation and authentic virus experiments: N.W. and J.S. Validation: Yanjun Wang, J.S., X.M., Yu Wang, D.G., C.L., G.M.D. and J.C. Project administration: S.S. and Y.B. Supervision: S.S., Y.B. and S.Z. Writing original draft: N.W., H.J. and M.V. Writing, review and editing: A.M., F.A.R., G.F.G., S.Z., S.S. and Y.B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
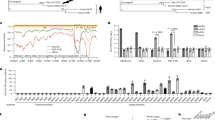
Extended Data Fig. 1 Analysis of receptor preference of MRCoV.
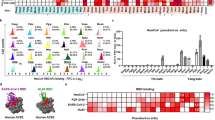
a, HEK 293T cells were transfected with plasmids expressing NvACE2, HsACE2, NvDPP4, and HsDPP4 harboring C-terminal Flag tags. At 48 h post transfection, cells were fixed and subjected to IFA using anti-Flag antibodies; nuclei were counter-stained with DAPI. Green, receptor proteins; blue, nuclei; scale bar, 500 μm. b, BHK-21 cells stably expressing NvACE2, PaACE2, or HsACE2 harboring C-terminal Flag tag were fixed and permeabilized. The expression levels of ACE2 proteins were analyzed using staining with FITC-conjugated anti-Flag antibody and flow cytometry. (a, b) The representative images from three independent experiments are shown. c, Analysis of binding of MRCoV RBD to NvACE2, PaACE2, and HsACE2 using SPR. The representative of images from three independent experiments are shown. Data are presented as the mean ± standard error of mean (s.e.m) of three independent experiments. d, Vero E6 and Huh7 cells were transfected with control siRNA (siControl) or siRNAs targeting ACE2 (#1, #2 or #3). Levels of ACE2 mRNAs in cell lysates were determined using RT-qPCR at 48 h post transfection. Data were normalized to control siRNA transfected cells (taken as 100%). Data are presented as the mean ± s.d. of three independent experiments. Statistical analysis was performed using one-way ANOVA.
Extended Data Fig. 2 Analysis of the ability of ACE2 orthologs to support the entry of MRCoV and analysis of expression of NvACE2 mutants.
a, HeLa cells were transfected with expression plasmids for Flag-tagged ACE2 from mink, Pipistrellus abramus (P.abramus), human, pangolin, Eurasian badger, monkey, pig, raccoon dog, and camel or with empty expression vector and infected with MRCoV (m.o.i. = 1). At 48 h.p.i. cells were fixed and stained for ACE2 expression and for dsRNA (a replication intermediate of the virus), nuclei were counterstained with DAPI. Red, ACE2; green, dsRNA; blue, nuclei; scale bar, 10 μm. The representative images from three independent experiments are shown. b, Replication dynamics of MRCoV in HeLa cells expressing ACE2 proteins from indicated species. Viral RNA copies in the supernatants were determined by RT-qPCR. Data are presented as the mean ± s.d. of three independent experiments. Statistical analysis was performed using one-way ANOVA. c, HEK 293 T cell stably expressing WT NvACE2 and its mutants with indicated substitutions were fixed and subjected to IFA using anti-Flag antibodies; nuclei were counter-stained with DAPI. Green, NvACE2 and its variants; blue, nuclei; scale bar, 500 μm. The representative images from three independent experiments are shown. Percentages of NvACE2 positive cells relative to DAPI positive nuclei were calculated by Image J. Data are presented as the mean ± s.d. of measurements performed for three randomly selected fields.
Extended Data Fig. 3 Resolution and electron density map of NvACE2:MRCoV RBD complex and structural comparison of RBDs of different MERS-CoV-related CoVs.
a, Fourier shell correlation (FSC) of the half cryo-EM maps, using gold standard protocol (FSC = 0.143). b, Electron density of key interacting residues in the structure of NvACE2 (green mesh) and MRCoV RBD (yellow mesh) complex. c, Comparison of the structure of MRCoV RBD with RBDs of NeoCoV, PDF-2180-CoV, BatCoV-HKU4, BatCoV-HKU5 and MERS-CoV. Figure was created with PyMOL from PDB-files 8ZWE (MRCoV), 7WPO (NeoCoV), 7WPZ (PDF-2180-CoV), 4QZV (BatCoV-HKU4), 5XGR (BatCoV-HKU5), and 4KR0 (MERS-CoV).
Extended Data Fig. 4 Determinants governing species-specific usage of ACE2.
a, Structure alignment of NvACE2 with PaACE2 (PDB: 9D32) and with HsACE2 (PDB: 6M0J). b, Partial amino acid sequence alignment of ACE2 from different mammals. White letters with red background indicate 100% conserved residues; red letters with white background indicate similar residues; black letters indicate non-conserved residues. Residues that interact with RBD are marked with asterisks. N-glycosylation sequons NXS/T in some of the sequences are marked with triangles. Panel was prepared with ESPript 3 (ref. 63). c, RBD-interacting residues which differ between NvACE2 and PaACE2 or NvACE2 and HsACE2 are shown as sticks and the substitutions are indicated. One of the unique differences between NvACE2 and HsACE2 is underlined. d, HEK 293T cells overexpressing WT PaACE2 or the indicated mutants were infected with MRCoV PVs. GFP positive-cells were counted at 48 h.p.i. Data is normalized to WT PaACE2-expressing cells and are presented as the mean ± s.d. of four independent experiments. Statistical analysis was performed using one-way ANOVA. e, Prediction of N-glycosylation at N322 residue in mink, P.abramus, human, raccoon dog, and camel WT and mutant ACE2 proteins using https://services.healthtech.dtu.dk/services/NetNGlyc-1.0/. The number represents the probability for N-glycosylation. f, HEK 293T cells expressing WT or mutant forms of human, racoon dog, or camel ACE2 were infected with MRCoV. Viral RNA copy numbers in the supernatants were determined at 48 h.p.i. Data are presented as the mean ± s.d. of three independent experiments. Statistical analysis was performed using one-way ANOVA. g, RBD:ACE2 interactions near N322 residue of NvACE2, PaACE2 and HsACE2. Images were created with PyMOL using the mutagenesis function. h, (left) Western blot analysis of WT HsACE2 without and with PNGaseF treatment. (right) Comparison of electrophoretic mobility of WT HsACE2 and its indicated mutants. The representative images from three independent experiments are shown.
Extended Data Fig. 5 Sequence alignments of the RBDs from different BetaCoVs and the RBDs of four ACE2-binding BetaCoVs.
a, Clustal-O amino acid sequence alignment of MRCoV, MERS-CoV, and BatCoV-HKU4 RBDs. The residues that interact with DPP4 are highlighted in cyan for BatCoV-HKU4 and in green for MERS-CoV. For MRCoV, residues identical to the receptor interacting residues of BatCoV-HKU4 or MERS-CoV are highlighted with the corresponding color. Residue D543 (MRCoV numbering), identical to receptor-interacting residues of both BatCoV-HKU4 and MERS-CoV RBDs, is marked in pink. b, RBD-sequences of SARS-CoV-2 (PDB: 6M0J), MRCoV (PDB: 8ZWE), PDF-2180-CoV (PDB: 7WPZ) and NeoCoV (PDB: 7WPO) were aligned using Clustal-O. Residues interacting with ACE2 are highlighted in yellow. c, Clustal-O alignment of sequences of RBDs of MRCoV, HKU5-33S and BatCoV-HKU5. Amino acids in RBD of MRCoV interacting with NvACE2 are highlighted in green and their counterparts, that are different in HKU5-33S and BatCoV-HKU5, are highlighted in pink. Circles indicate residues that form an ionic bond and squares indicate residues that form hydrogen-bonds in the complex of MRCoV RBD with NvACE2. N-glycosylation sequons are highlighted in grey. Conserved residues are colored red. Completely conserved residues are shown in white on a red background. a-c, Panels were prepared with ESPript 3 (ref. 63). d, Homology model of HKU5-33S RBD obtained with https://swissmodel.expasy.org/ using the structure of the MRCoV RBD as template. The quality of the model is high (Global Model Quality Estimate score: 0.90; QMEAND score: 0.84 ± 0.06) as indicated by the red color, except the loops which are predicted with lower confidence. e, Structural alignment of the MRCoV RBD structure (green) and the modeled HKU5-33S RBD (blue). Residues in MRCoV interacting with NvACE2 are shown as sticks colored according to atom type (carbon green, oxygen red, nitrogen blue). Interacting residues that differ in HKU5-33S are shown with carbons in white, and the substitution is indicated.
Extended Data Fig. 6 Efficacy of different antiviral treatments against MRCoV infection.
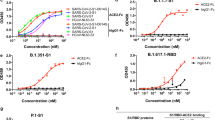
a, Dose-dependent inhibition of MRCoV and SARS-CoV-2 GFP/ΔN by SARS-CoV-2 specific antisera (n = 5). Data from one of three independent experiments are shown. b, Antiviral activities of small molecule SARS-CoV-2 inhibitors NHC (β-D-N4-hydroxycytidine) and PF-07321332 (nirmatrelvir) against MRCoV and SARS-CoV-2 GFP/ΔN. Data are presented as the mean ± s.d. of three independent experiments. (a,b), MRCoV infection was measured by determination of viral RNA copies in the cell culture supernatants using RT-qPCR; efficiency of SARS-CoV-2 GFP/ΔN infection was quantified using CQ1 confocal imaging cell counter.
Supplementary information
Supplementary Information (download DOCX )
Supplementary Figs. 1–3 and Supplementary Table 1.
Supplementary Table 2 (download XLSX )
Virus and receptor gene information.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, N., Ji, W., Jiao, H. et al. A MERS-CoV-like mink coronavirus uses ACE2 as an entry receptor. Nature 642, 739–746 (2025). https://doi.org/10.1038/s41586-025-09007-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09007-w
This article is cited by
-
Structural and catalytic diversity of coronavirus proofreading exoribonuclease
Nature Communications (2025)
-
Targeting ACE2 with a camelid antibody inhibits SARS-CoV-2 binding and has protective effects in vivo
Nature Communications (2025)