Abstract
To grow at distant sites, metastatic cells must overcome major challenges posed by the unique cellular and metabolic composition of secondary organs1. Pancreatic ductal adenocarcinoma (PDAC) is an aggressive disease that metastasizes to the liver and lungs. Despite evidence of metabolic reprogramming away from the primary site, the key drivers that dictate the ability of PDAC cells to colonize the liver or lungs and survive there remain undefined. Here we identified PCSK9 as predictive of liver versus lung colonization by integrating metastatic tropism data of human PDAC cell lines2, in vivo metastasis modelling in mice and gene expression correlation analysis. PCSK9 negatively regulates low density lipoprotein (LDL)-cholesterol import and, accordingly, PCSK9-low PDAC cells preferentially colonize LDL-rich liver tissue. LDL-cholesterol taken up by liver-avid PCSK9-low cells supports activation of pro-growth mTORC1 activation at the lysosome, and through conversion into the signalling oxysterol, 24(S)-hydroxycholesterol, reprogrammes the microenvironment to release nutrients from neighbouring hepatocytes. Conversely, PCSK9-high, lung-avid PDAC cells rely on transcriptional upregulation of the distal cholesterol synthesis pathway to generate intermediates—7-dehydrocholesterol and 7-dehydrodesmosterol—with protective action against ferroptosis, a vulnerability in the oxygen-rich microenvironment of the lung. Increasing the amount of PCSK9 redirected liver-avid cells to the lung whereas ablating PCSK9 drove lung-avid cells to the liver, thereby establishing PCSK9 as necessary and sufficient for secondary organ site preference. Our studies reveal PCSK9-driven differential utilization of the distal cholesterol synthesis pathway as a key and potentially actionable driver of metastatic growth in PDAC.
This is a preview of subscription content, access via your institution
Access options





Similar content being viewed by others
Data availability
Metabolomics data including 7-DHC and 7-DHD quantification were deposited on the Metabolomics Workbench platform at http://dx.doi.org/10.21228/M8RV70 (project ID: PR002405)70. Single-cell RNA-sequencing data are available from ref. 27. Additional resources from the Cancer Research Training Program (CRTP) can be found at https://crtp.ccr.cancer.gov/.
References
Gerstberger, S., Jiang, Q. & Ganesh, K. Metastasis. Cell 186, 1564–1579 (2023).
Jin, X. et al. A metastasis map of human cancer cell lines. Nature 588, 331–336 (2020).
Massague, J. & Obenauf, A. C. Metastatic colonization by circulating tumour cells. Nature 529, 298–306 (2016).
Elia, I. & Haigis, M. C. Metabolites and the tumour microenvironment: from cellular mechanisms to systemic metabolism. Nat. Metab. 3, 21–32 (2021).
Liu, K. H. et al. Lung metastases in patients with stage IV pancreatic cancer: prevalence, risk factors, and survival impact. J. Clin. Med. 8, 1402 (2019).
Deeb, A., Haque, S. U. & Olowokure, O. Pulmonary metastases in pancreatic cancer, is there a survival influence? J. Gastrointest. Oncol. 6, E48–E51 (2015).
Downs-Canner, S. et al. The indolent nature of pulmonary metastases from ductal adenocarcinoma of the pancreas. J. Surg. Oncol. 112, 80–85 (2015).
Yasukawa, M., Kawaguchi, T., Kawai, N., Tojo, T. & Taniguchi, S. Surgical treatment for pulmonary metastasis of pancreatic ductal adenocarcinoma: study of 12 cases. Anticancer Res. 37, 5573–5576 (2017).
Link, J. M. et al. Ongoing replication stress tolerance and clonal T cell responses distinguish liver and lung recurrence and outcomes in pancreatic cancer. Nat. Cancer 6, 123–144 (2025).
Cheung, E. C. et al. Dynamic ROS control by TIGAR regulates the initiation and progression of pancreatic cancer. Cancer Cell 37, 168–182.e164 (2020).
Maddipati, R. et al. MYC levels regulate metastatic heterogeneity in pancreatic adenocarcinoma. Cancer Discov. 12, 542–561 (2022).
Mucciolo, G. et al. EGFR-activated myofibroblasts promote metastasis of pancreatic cancer. Cancer Cell 42, 101–118.e111 (2024).
Pommier, A. et al. Unresolved endoplasmic reticulum stress engenders immune-resistant, latent pancreatic cancer metastases. Science 360, eaao4908 (2018).
Raghavan, S. et al. Microenvironment drives cell state, plasticity, and drug response in pancreatic cancer. Cell 184, 6119–6137.e6126 (2021).
Tsanov, K. M. et al. Metastatic site influences driver gene function in pancreatic cancer. Preprint at bioRxiv https://doi.org/10.1101/2024.03.17.585402 (2024).
Wang, X. et al. Identification of a subset of immunosuppressive P2RX1-negative neutrophils in pancreatic cancer liver metastasis. Nat. Commun. 12, 174 (2021).
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016).
Cancer Genome Atlas Research Network. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell 32, 185–203.e13 (2017).
Collisson, E. A. et al. Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nat. Med. 17, 500–503 (2011).
Connor, A. A. et al. Integration of genomic and transcriptional features in pancreatic cancer reveals increased cell cycle progression in metastases. Cancer Cell 35, 267–282.e267 (2019).
Moffitt, R. A. et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat. Genet. 47, 1168–1178 (2015).
O’Kane, G. M. et al. GATA6 expression distinguishes classical and basal-like subtypes in advanced pancreatic cancer. Clin. Cancer Res. 26, 4901–4910 (2020).
Aung, K. L. et al. Genomics-driven precision medicine for advanced pancreatic cancer: early results from the COMPASS trial. Clin. Cancer Res. 24, 1344–1354 (2018).
Link, J. M. et al. Tumor-infiltrating leukocyte phenotypes distinguish outcomes in related patients with pancreatic adenocarcinoma. JCO Precis. Oncol. https://doi.org/10.1200/PO.20.00287 (2021).
Boj, S. F. et al. Organoid models of human and mouse ductal pancreatic cancer. Cell 160, 324–338 (2015).
Kruger, S. et al. Isolated pulmonary metastases define a favorable subgroup in metastatic pancreatic cancer. Pancreatology 16, 593–598 (2016).
Zhang, S. et al. Single cell transcriptomic analyses implicate an immunosuppressive tumor microenvironment in pancreatic cancer liver metastasis. Nat. Commun. 14, 5123 (2023).
Brown, M. S. & Goldstein, J. L. Cholesterol feedback: from Schoenheimer’s bottle to Scap’s MELADL. J. Lipid Res. 50, S15–S27 (2009).
Brown, M. S., Radhakrishnan, A. & Goldstein, J. L. Retrospective on cholesterol homeostasis: the central role of Scap. Annu. Rev. Biochem. 87, 783–807 (2018).
Ishida, C. T. et al. SREBP-dependent regulation of lipid homeostasis is required for progression and growth of pancreatic ductal adenocarcinoma. Cancer Res. Commun. 4, 2539–2552 (2024).
Kotowski, I. K. et al. A spectrum of PCSK9 alleles contributes to plasma levels of low-density lipoprotein cholesterol. Am. J. Hum. Genet. 78, 410–422 (2006).
Sarkar, S. K. et al. A transient amphipathic helix in the prodomain of PCSK9 facilitates binding to low-density lipoprotein particles. J. Biol. Chem. 295, 2285–2298 (2020).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9–Niemann-Pick C1 signaling complex. Science 355, 1306–1311 (2017).
Shin, H. R. et al. Lysosomal GPCR-like protein LYCHOS signals cholesterol sufficiency to mTORC1. Science 377, 1290–1298 (2022).
Nelson, E. R. et al. 27-Hydroxycholesterol links hypercholesterolemia and breast cancer pathophysiology. Science 342, 1094–1098 (2013).
Courtney, R. & Landreth, G. E. LXR regulation of brain cholesterol: from development to disease. Trends Endocrinol. Metab. 27, 404–414 (2016).
Wang, B. & Tontonoz, P. Liver X receptors in lipid signalling and membrane homeostasis. Nat. Rev. Endocrinol. 14, 452–463 (2018).
Dixon, S. J. & Olzmann, J. A. The cell biology of ferroptosis. Nat. Rev. Mol. Cell Biol. 25, 424–442 (2024).
Garcia-Bermudez, J. et al. Squalene accumulation in cholesterol auxotrophic lymphomas prevents oxidative cell death. Nature 567, 118–122 (2019).
Freitas, F. P. et al. 7-Dehydrocholesterol is an endogenous suppressor of ferroptosis. Nature 626, 401–410 (2024).
Li, Y. et al. 7-Dehydrocholesterol dictates ferroptosis sensitivity. Nature 626, 411–418 (2024).
Yamada, N. et al. Inhibition of 7-dehydrocholesterol reductase prevents hepatic ferroptosis under an active state of sterol synthesis. Nat. Commun. 15, 2195 (2024).
Mullen, P. J., Yu, R., Longo, J., Archer, M. C. & Penn, L. Z. The interplay between cell signalling and the mevalonate pathway in cancer. Nat. Rev. Cancer 16, 718–731 (2016).
Riscal, R., Skuli, N. & Simon, M. C. Even cancer cells watch their cholesterol! Mol. Cell 76, 220–231 (2019).
Snaebjornsson, M. T., Janaki-Raman, S. & Schulze, A. Greasing the wheels of the cancer machine: the role of lipid metabolism in cancer. Cell Metab. 31, 62–76 (2020).
Vasseur, S. & Guillaumond, F. Lipids in cancer: a global view of the contribution of lipid pathways to metastatic formation and treatment resistance. Oncogenesis 11, 46 (2022).
Ye, J. & DeBose-Boyd, R. A. Regulation of cholesterol and fatty acid synthesis. Cold Spring Harb. Perspect. Biol. 3, a004754 (2011).
Mahboobnia, K. et al. PCSK9 and cancer: rethinking the link. Biomed. Pharmacother. 140, 111758 (2021).
Kietzmann, T. Metabolic zonation of the liver: the oxygen gradient revisited. Redox Biol. 11, 622–630 (2017).
Daemen, A. et al. Metabolite profiling stratifies pancreatic ductal adenocarcinomas into subtypes with distinct sensitivities to metabolic inhibitors. Proc. Natl Acad. Sci. USA 112, E4410–E4417 (2015).
Gabitova-Cornell, L. et al. Cholesterol pathway inhibition induces TGF-β signaling to promote basal differentiation in pancreatic cancer. Cancer Cell 38, 567–583.e511 (2020).
Karasinska, J. M. et al. Altered gene expression along the glycolysis–cholesterol synthesis axis is associated with outcome in pancreatic cancer. Clin. Cancer Res. 26, 135–146 (2020).
Guillaumond, F. et al. Cholesterol uptake disruption, in association with chemotherapy, is a promising combined metabolic therapy for pancreatic adenocarcinoma. Proc. Natl Acad. Sci. USA 112, 2473–2478 (2015).
Jacobs, E. J., Newton, C. C., Thun, M. J. & Gapstur, S. M. Long-term use of cholesterol-lowering drugs and cancer incidence in a large United States cohort. Cancer Res. 71, 1763–1771 (2011).
Nielsen, S. F., Nordestgaard, B. G. & Bojesen, S. E. Statin use and reduced cancer-related mortality. N. Engl. J. Med. 367, 1792–1802 (2012).
Hillis, A. L. et al. Targeting cholesterol biosynthesis with statins synergizes with AKT inhibitors in triple-negative breast cancer. Cancer Res. 84, 3250–3266 (2024).
Zhou, Q., Jiao, Z., Liu, Y., Devreotes, P. N. & Zhang, Z. The effects of statins in patients with advanced-stage cancers - a systematic review and meta-analysis. Front. Oncol. 13, 1234713 (2023).
Mei, W. et al. A commonly inherited human PCSK9 germline variant drives breast cancer metastasis via LRP1 receptor. Cell 188, 371–389.e28 (2025).
Liang, X. et al. Rapid and highly efficient mammalian cell engineering via Cas9 protein transfection. J. Biotechnol. 208, 44–53 (2015).
Susaki, E. A. et al. Advanced CUBIC protocols for whole-brain and whole-body clearing and imaging. Nat. Protoc. 10, 1709–1727 (2015).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Charni-Natan, M. & Goldstein, I. Protocol for primary mouse hepatocyte isolation. STAR Protoc. 1, 100086 (2020).
Guenther, M. et al. The impact of adjuvant therapy on outcome in UICC stage I pancreatic cancer. Int. J. Cancer 151, 914–919 (2022).
Adams, C. R. et al. Transcriptional control of subtype switching ensures adaptation and growth of pancreatic cancer. eLife 8, e45313 (2019).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 e324 (2019).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Zhang, L. et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell 181, 442–459.e429 (2020).
Leary, J. R. et al. Sub-cluster identification through semi-supervised optimization of rare-cell silhouettes (SCISSORS) in single-cell RNA-sequencing. Bioinformatics 39, btad449 (2023).
Rademaker, G. Summary of project PR002405. Metabolomics Workbench http://dx.doi.org/10.21228/M8RV70 (2025).
Acknowledgements
This work was supported by a BAEF (Belgian American Education Foundation) fellowship, Fonds Léon Fredericq, Rotary and Wallonie Bruxelles International grants (G.R.); a National Science Foundation Graduate Research Fellowship (NSF2034836) (G.A.H.); NHLBI R35 HL161241 and the Nina Ireland Program for Lung Heath (M.R.L.); National Cancer Institute R01CA260205 (R.Z.); National Cancer Institute R01CA240603 and R01CA260249, the Ed Marra Passion to Win Fund, the Weston Havens Fund and the AACR-MPM Transformative Cancer Research Award (R.M.P.); National Cancer Institute R01CA199064 (J.J.Y.), U01CA274298, P50CA257911 and U24CA211000 (X.L.P. and J.J.Y.); 7R01CA260860 and 7R01CA256969 (E.A.C.); Sky Foundation fellowship (L.M.P.); and National Institute of Diabetes and Digestive and Kidney Diseases R01DK131227 and P30DK026743 (B.W.). The authors thank the UCSF Parnassus Flow CoLab, RRID:SCR_018206 for assistance with flow cytometry and the UCSF Liver Center, supported by P30DK026743, for assistance with hepatocyte isolation.
Author information
Authors and Affiliations
Contributions
G.R. conceived the project, performed most experiments and wrote the manuscript. G.A.H. conducted tail vein and intracardiac injections and assisted with experiments. Y.S. performed the light-sheet imaging experiment. S.D. performed immunoblotting and RT–qPCR analyses. L.M.-P., S.F.K., S.B., S.O., M.G., V.H. and M.H. accrued and provided the matched PDAC primary patient specimens. A.L.L. cloned the PCSK9 plasmids and variants. X.L.P. and C.L. performed analysis of single-cell RNA-sequencing data. L.Q. performed the intrasplenic injections. M.A.L. performed flow cytometry and data analysis. K.W.W. and G.E.K. contributed PDAC specimens and performed blinded analysis. B.W., E.A.C., M.R.L., J.J.Y. and R.Z. supervised the research and provided intellectual feedback. R.M.P. conceived the project, supervised the research, wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Liver- and Lung-avid cell lines exhibit distinct features.
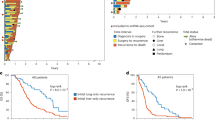
a, b. Petal plots depicting metastatic potential (length) and penetrance (width) for C1-Liver lines (a) and C2-Lung lines (b), extracted from the MetMap 500 analysis. Br, Brain; Bo, Bone; Ki, Kidney; Li, Liver; Lu, Lung. c. Representative H&E images showing tumor growth (red outline) in the liver (top) and lung (bottom) 3 weeks post intra-cardiac injection of the indicated PDAC cell lines. Scale bars: 500 µm. d. Quantification of liver/lung colonization ratio of C1-Liver and C2-lung lines for the experiment in c. (n = 6 mice per/cluster). Statistical significance was calculated using a two-tailed t-test. e. Flow cytometry-based measurement of relative eGFP or mCherry enrichment in the liver or lungs. mCherry positive cells were more abundant in the liver while eGFP positive cells were more abundant in the lungs (related to Fig. 1l–n). Statistical significance was calculated using a two-tailed t-test. Error bars depict standard deviation. (n = 3 tissues per condition). f. Gene set enrichment analysis (GSEA) plots showing relative enrichment of the PDAC classical gene signature derived from Adams et al. and Moffit et al. in cluster 1 (C1-Liver) and cluster 2 (C2-Lung) cell lines. Statistical significance was calculated using a permutation test with FDR adjustment. g. Scatter plot showing correlation between liver/lung metastatic potential ratio (see Supplementary Table 2) and classical score from Adams et al. and Moffit et al. Each dot represents a PDAC cell line (n = 25). Linear regression followed by Pearson correlation. Error margins (grey) depict standard error of the mean. h. Representative immunofluorescence images of GATA6 (green), Pan-Cytokeratin (CK19) (red), DAPI (blue) in liver (top) and lung (bottom) patient PDAC metastases. Note the nuclear localization of GATA6 in lung metastases. Scale bars: 20 µm. i. Quantification of the percentage nuclear GATA6 in liver (56 fields from n = 3 samples) and lung (45 fields from n = 3 samples) metastases sections. Statistical significance was calculated using a two tail T-test. j. Representative H&E images of human liver and lung PDAC metastases. Scale bar: 20 µm. k. Bar plot depicting quantification of the differentiation status of PDAC liver and lung metastases shown in j, (Liver metastases n = 45; Lung metastases n = 10). Statistical significance was calculated using a two-tailed t-test. l. Representative immunofluorescence images of ZEB1 (green), Pan-Cytokeratin (CK19) (red), DAPI (blue) in liver and lung patient PDAC metastatic lesions. Note the nuclear localization of ZEB1 in the liver metastases. Scale bars: 20 µm. m. Quantification of the percentage nuclear ZEB1 in liver (55 fields from n = 5 samples) and lung (40 fields from n = 5 samples) metastases sections. Statistical significance was calculated using a two tail T-test. For flow cytometry source data, see Supplementary Fig. 2.
Extended Data Fig. 2 Basal/classical PDAC subtypes do not impact metastatic organ preference.
a. Immunoblot of the indicated proteins following CRISPR mediated knockout of GATA6 in HPAC cells. b. Heatmap depicting normalized mRNA expression of the indicated genes, measured by qRT-PCR in HPAC cells with or without GATA6 knockout. c. In vitro proliferation rate of the cells shown in a. (n = 3 replicates per group). d. Representative H&E images of lung (top), and liver (bottom) 4 weeks post tail-vein or intrasplenic injection, respectively, of HPAC cells following GATA6 knockout. Scale Bar: 1 mm. e. Percentage tumour area in the lungs from samples shown in d. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. sgNT: n = 5; sgGATA6_1: n = 4; sgGATA6_2: n = 5. f. Percentage tumour area in the liver from samples shown in d. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. sgNT: n = 5; sgGATA6_1: n = 3; sgGATA6_2: n = 4. All error bars represent standard deviation of the mean. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 High PCSK9 expression is correlated with classical PDAC subtype and lung colonization.
a. Gene expression correlation of PCSK9 relative to the indicated classical (CDH1, S100P, FOXQ1) and basal (VIM, ZEB1) genes across all PDAC cells lines available on DepMap. Statistically significant correlations are indicated in red, and non-significant correlations are indicated in black. Gene rank for the indicated genes are shown in each plot. b. Scatter plot showing expression correlation between PCSK9 and individual classical (CDH1, GATA6, S100P, FOXQ1) and basal (ZEB1, VIM) genes (data extracted from n = 50 cell lines available in DepMap). Linear regression followed by Pearson correlation. Error margins (grey) depict standard error of the mean. c. ELISA based quantification of secreted PCSK9 protein from conditioned media collected from C1-Liver (n = 5) and C2-Lung (n = 5) cell lines. Dots represent n = 3 biological replicates per cell line. Statistical significance was calculated using a two tail T-test. d. Representative flow cytometry-based detection of plasma membrane LDLR in C1-Liver (n = 3) and C2-Lung (n = 3) cell lines. e. Quantification of the mean fluorescent intensity of plasma membrane LDLR from the cell lines in d. (n = 3 biological replicates per cell line). Statistical significance was calculated using a two tail T-test. f. Immunoblot of the indicated proteins in mouse derived KPC cell lines. g. Representative H&E images of tumor burden in the liver (top) and lung (bottom) 2 weeks post intrasplenic or tail-vein injection, respectively, of the indicated KPC cell lines. (n = 3 mice/cell line). Scale Bar: 500 µm. h. Petal plots depicting metastatic potential (length) and penetrance (colour) for the indicated cell lines based on data generated in g. Li, Liver; Lu, Lung. i. UMAP plot of single cell sequencing from n = 3 primary PDAC lesions depicting two clusters representing the classical (blue) and basal (yellow) subtypes in ductal cells. j. UMAP plot showing enrichment of PCSK9 mRNA expression in the classical PDAC subtype as depicted in i. k. Quantification of PCSK9 mRNA expression in ductal cells corresponding to basal and classical PDAC subtypes. Statistical significance was calculated using a two-tailed t-test. l. UMAP plot of single cell sequencing from n = 3 primary PDAC and n = 4 matched liver metastases. Colors represent different cell types. m. PCSK9 mRNA expression in ductal cells from primary PDAC and liver metastases. Statistical significance was calculated using a two-tailed t-test. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 4 Liver-avid cells uptake LDL cholesterol.
a,b. Flow cytometry plots of plasma membrane Dil-LDL in KP4 and HPAC cell lines (a). The dotted line depicts the negative control staining. Graph depicts quantification of data (b). Data points represent the average of 3 technical replicates from n = 3 biological replicates. Error bars represent standard deviation. c,d. Representative immunofluorescence images (c) and quantification (d) of Dil-LDL uptake (red) in C1-Liver and C2-Lung lines. Cells are co-stained for LAMP2 (green) and DAPI (blue). Scale bars: 10 µm. (n = 10 fields/cell line). e,f. Representative immunofluorescence images (e) and quantification (f) of Dil-LDL uptake (red) in four mouse KPC derived cell lines. Note that KPC1 and KPC2 take up exogenous LDL while KPC3 and KPC4 do not. Cells are co-stained for LAMP1 (green) and DAPI (blue). Scale bars: 10 µm. (n = 10 fields/cell line). g-i. Representative immunofluorescence images depicting Filipin (green) in C1-Liver (g), C2-Lung (h) lines and quantification (i). Cells are co-stained for Lysotracker (red) and DAPI (blue). Scale bars: 10 µm. Data points represent the average of at least 7 fields from n = 5 cell lines/group. j. Immunoblot for the indicated proteins in KPC derived cells isolated from isogenic primary tumor and liver metastases. k,l. Representative immunofluorescence images (k) and quantification (l) of Dil-LDL uptake (red) in the KPC lines in j. Cells are co-stained for LAMP1 (green) and DAPI (blue). Scale bars: 10 µm. (n = 10 fields/cell line). m,n. Representative immunofluorescence images (m) and quantification (n) of Filipin (green) in the KPC lines in j and k. Cells are co-stained for Lysotracker (red) and DAPI (blue). (n = 10 fields/cell line). Scale bars: 10 µm. Statistical significance was calculated using a two-tailed t-test for data shown in b, d, i, l, and n. For gel source data, see Supplementary Fig. 1. For flow cytometry source data, see Supplementary Fig. 2.
Extended Data Fig. 5 Liver-avid cells rely on LDL-cholesterol uptake for growth.
a. Growth curves of KPC lines cultured in DMEM with FBS or LPDS. (n = 3 replicates per condition). b. Immunoblot of the indicated proteins in C1-Liver lines following treatment with DMEM containing FBS or LPDS for 72 h. c. Heatmap depicting the expression levels of mevalonate pathway genes in C1-Liver and C2-Lung lines. Each row represents a specific gene and columns represent biological replicates for each cell line. Expression levels are expressed as z-scores. d. Immunoblot of the indicated proteins in C2-Lung lines following treatment with DMEM containing FBS or LPDS for 72 h. e,f. Relative cell confluency of MiaPaCa2 (e) and KP4 (f) cells cultured in the indicated media supplemented with cholesterol, LDL or oleic acid. Statistical significance is calculated using a one-way ANOVA with Bonferroni correction. (n = 9 biological replicates per conditions). g,h. Immunoblots of the indicated proteins in KP4 cells (g) and additional cell lines (h) following CRISPR mediated knockout of LDLR. i. Representative images of colony forming assays of C1-Liver lines and C2-Lung lines following CRISPR mediated knockout of LDLR. j. Quantification of colony forming number from images in i. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. (n = 9 biological replicates/condition). k. Representative H&E images showing tumor growth (red outline) in the liver 4 weeks post intra-splenic injection of KP4 cells expressing control sgRNA or LDLR targeting sgRNA. Scale bars: 1 mm. l. Quantification of tumor area from data in g. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. sgNT: n = 5 mice; sgLDLR_1: n = 4 mice; sgLDLR_2: n = 5 mice. All error bars represent standard deviation of the mean. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 6 Lung-avid cells rely on cholesterol biosynthesis.
a. Immunoblot of the indicated proteins in C1-Liver and C2-Lung lines. b. Quantitative real-time PCR measurement of the indicated genes in C1-Liver (n = 5) and C2-Lung (n = 5) lines expressed as z-score. Each point represents the average of n = 3 biological replicates for each line. Statistical significance was calculated using a two-tailed t-test. c. Heatmap depicting mRNA expression of the indicated genes measured by qRT-PCR displayed as z-score. (n = 3 biological replicates per line). d. Immunoblot for SREBP2 in the indicated cell lines. e. Representative immunohistochemistry images of SREBP2 in liver (left) and lung (right) metastases. Scare bar = 100 μm. f. Quantification of SREBP2 staining intensity from n = 10 liver and n = 9 lung metastases depicted in e. Statistical significance was calculated using a two tailed t-test. g. Expression levels of cholesterol biosynthesis genes in single cell RNA sequencing data from primary PDAC (n = 3) and the matched liver metastases (n = 4). Statistical significance was calculated using a two-tailed t-test. h. DepMap CRISPR dependency ranking of top genes associated with PCSK9 high status across PDAC. Cholesterol biosynthesis genes are indicated. Red dots indicate significant genes and black dots indicate non-significant genes. i. Reactions of the cholesterol biosynthesis pathway with key enzymes highlighted in green. j. Immunoblot for the indicated proteins following CRISPR mediated knockout of the indicated genes (related to data presented in fig. 3c,d). k. Representative images of colony forming assays of C1-Liver and C2-Lung lines upon CRISPR-mediated knockout of the indicated genes. All error bars represent standard deviation of the mean. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 7 Altering PCSK9-LDLR status leads to a switch in cholesterol metabolism and metastatic organ preference.
a. Representative immunofluorescence images of DiL-LDL (red) uptake in KP4 cells or following expression of the indicated PCSK9 variants. Cells are co-stained for LAMP2 (green) and DAPI (blue) Scale bar: 10 µm. b. Quantification of DiL-LDL fluorescence intensity from each condition depicted in a. (n = 10 fields/condition). Statistical significance was calculated using a one-way ANOVA with LSD correction. c. Representative immunofluorescence images of DiL-LDL (red) uptake in control HPAC cells (sgNT) or following knockout of PCSK9 (sgPCSK9). Scale bar: 10 µm. d. Quantification of DiL-LDL fluorescence intensity from each condition depicted in c. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. (n = 10 fields/condition). e. Measurement of percentage confluency of HPAC cells transfected with sgNT or sgPCSK9. (n = 3 replicates per condition). f. Immunoblot of the indicated proteins in HPAC cells following ectopic expression of LDLR. g. Representative immunofluorescence images of DiL-LDL (red) uptake in control HPAC cells following ectopic expression of LDLR. Scale bar: 5 µm. h. Quantification of DiL-LDL fluorescence intensity from each condition depicted in g. Control: n = 14 fields/condition, LDLR: n = 10 fields/condition. Statistical significance was calculated using a two-tailed t-test. i. Measurement of percentage confluency of control (eGFP) HPAC cells and following expression of LDLR. (n = 3 replicates per condition). j. Representative H&E images of tumor burden (red outline) in the liver 4 weeks post intra-splenic injection of HPAC cells with or without ectopic LDLR expression. Scale bars: 2 mm. k. Quantification of percent liver invasion from n = 5 mice/condition from data depicted in j. Statistical significance was calculated using a two-tailed t-test. l. Immunoblot of the indicated proteins in KPC3 cells (HY16936) following CRISPR mediated knockout of Pcsk9. m. Representative immunofluorescence images of DiL-LDL (red) uptake in control KPC3 cells (sgNT) or following knockout of Pcsk9 (sgPcsk9). Scale bar: 10 µm. n. Quantification of DiL-LDL fluorescence intensity from each condition depicted in m. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. (n = 10 fields/condition). o. Heatmap showing expression levels of the indicated cholesterol biosynthesis genes measured by qRT-PCR and represented as z scores, following CRISPR mediated knockout of Pcsk9 in KPC3 cells. p. Measurement of percentage confluency of KPC3 cells transfected with sgNT or sgPcsk9. (n = 3 replicates per condition). q. Representative H&E images showing tumor burden (red outline) in the lungs (top) and liver (bottom) 2 weeks post tail-vein or intra-splenic injection, respectively, of KPC3 cells transfected with sgNT or sgPcsk9. Scale bars: 1 mm. r,s. Quantification of percentage tumor burden in the lung (r) and liver (s) from the experiment in q. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. (n = 5 mice/group). t. Image of tumors excised from the pancreas 2 weeks post orthotopic transplantation of KPC3 cells expressing sgNT or sgPcsk9. u. Quantification of tumor weight in each condition depicted in t. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. (n = 5 tumors/condition). All error bars represent standard deviation of the mean. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 8 PCSK9 expression level is organ dependent.
a. Representative immunofluorescence images depicting CK19 (red) and Pcsk9 (green) staining in mouse MT23 derived tumors 2 weeks post orthotopic transplant into the pancreas, intra-splenic injection to seed the liver and tail-vein injection to seed the lungs of C57Bl/6 mice. Scale bars: 50 µm. b. Quantification of Pcsk9 staining intensity in CK19 positive ductal structures from tumours grown in the pancreas (n = 3 mice; 12 fields), liver (n = 3 mice; 10 fields) and lung (n = 3 mice; 11 fields). Statistical significance was calculated using a one-way ANOVA with Bonferroni correction.
Extended Data Fig. 9 Liver-avid PDAC cells utilize cholesterol to activate mTORC1 and condition the liver microenvironment.
a. Representative immunofluorescence images (left) and quantification (right) of Filipin staining in C1-Liver (MiaPaCa2) and C2-Lung (HPAF-II) cell lines growth in DMEM with FBS or LPDS for 72 h. Statistical significance was calculated using a one-way ANOVA followed by pairwise t-tests with Bonferroni correction. Error bars depict standard deviation. n = 10 fields for MiaPaCa2 (DMEM and LPDS) and HPAF-II (DMEM) and n = 6 fields for HPAF-II LPDS. b. Immunoblot showing 4EBP1 and P70S6K phosphorylation and total protein levels in HPAC PCSK9 KO cells treated with LPDS for 72 h. c. Representative immunofluorescence images of p-S6 (Ser240/244) in HPAC PCSK9 KO liver and lung tumors. Scale bar: 20 µm. Graph at right shows quantification of p-S6 (Ser240/244) staining intensity in n = 10 PDAC liver metastases and n = 9 PDAC lung metastases. Each data point represents the average of at least 4 fields/sample. Statistical significance was calculated using a two tail T-test. d. Analysis of RPPA shows that mTOR and mTOR targets are anti-correlated with PCSK9 levels. e. Normalized mRNA expression measured by quantitative real-time PCR of the indicated transcripts in C1-Liver (n = 5) and C2-Lung (n = 5) lines. Dots represent the average of 3 biological replicates per cell line. Statistical significance was calculated using a two-tailed t-test. Line within the box plot represents the median. f,g. Scatter plots showing anti-correlation between PCSK9 and CYP46A1 (f) and no correlation with other cholesterol hydroxylases (g). Data derived from TCGA PAAD dataset. Linear regression followed by Pearson correlation. Error bars depict standard error of the mean. (n = 183 PDAC specimens). h. Total intracellular cholesterol in mouse hepatocytes following treatment with KP4 or HPAC CM for 24 h, normalized to total cell number. (n = 3 biological replicates). Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. Error bars depict standard deviation. i. Sequencing reads showing successful deletion of CYP46A1 using 2 independent sgRNAs in KP4 cells. j. Heatmap depicting the expression levels of LXR target genes in mouse hepatocytes as measured by qRT-PCR, following treatment for 24 h with CM isolated from KP4 parental and CYP46A1 knockout cells or treated with 24-hydroxycholesterol (24-HC; 25 µM). Each row represents a specific gene. Each column represents a biological replicate per condition. Expression levels are expressed as z-scores, with positive z-scores indicating higher expression (red) and negative z-scores indicating lower expression (blue). k. Total intracellular cholesterol in mouse hepatocytes following treatment with CM isolated from KP4 parental or CYP46A1 KO CM, normalized to cell number. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. Error bars depict standard deviation. (n = 3 biological replicates/group). l. Representative H&E images showing tumor burden in the liver (top) and lungs (bottom) following intra-splenic or tail-vein injection, respectively, of KP4 cells following CRISPR mediated knockout of CYP46A1. (n = 5 mice/group). Scale Bar: 500 µm. m,n. Percentage tumour area in the liver (m) and lungs (n) from samples shown in l. Statistical significance was calculated using one-way ANOVA with Bonferroni correction. Error bars depict standard deviation. o. Measurement of KP4 growth rate in vitro following CYP46A1 knockout. Error bars depict standard deviation. (n = 3 replicates per group).
Extended Data Fig. 10 Lung-avid PDAC cells utilize cholesterol synthesis to generate sterols that protect against ferroptosis.
a. Scatter plots showing TCGA PAAD derived expression correlation between PCSK9 and individual cholesterol biosynthesis pathway genes. Linear regression followed by Pearson correlation. Error bars depict standard error of the mean. (n = 183 PDAC specimens). b. Average ML210 IC50 measurement in C1-Liver (n = 5) and C2-Lung (n = 5) lines. Statistical significance was calculated using a two-tailed t-test. Error bars depict standard deviation. (n = 5 lines per condition). c. SYTOXgreen images (left) and quantification (right) of HPAC cell death (green fluorescence) following knock-out of PCSK9 and treatment with RSL3 for 24 h. Statistical significance was calculated using a one-way ANOVA with Bonferroni correction. Error bars depict standard deviation. (n = 3 biological replicates/group). d. Sequencing reads showing successful CRISPR mediated deletion of SC5D in HPAC cells. e. Measurement of HPAC cell growth following knockout of SC5D and treatment with increasing doses of RSL3 (left) and ML210 (right) for 96 h. Error bars depict standard error of the mean. (n = 3 biological replicates/group). f. Representative H&E images showing tumor growth (red outline) in the lungs 4 weeks post tail-vein injection of HPAC cells following CRISPR mediated knockout of SC5D. Scale bars: 1 mm. g. Quantification of percentage tumor burden from the experiment in f. Statistical significance is calculated using a one-way ANOVA with Bonferroni correction. Error bars depict standard deviation. (n = 5 mice/group). h. SYTOXgreen images and quantification of KP4 cell death (green fluorescence) following expression of a control vector or PCSK9D374Y and treatment with RSL3 or ML210 for 24 h. Statistical significance was calculated using a two-tailed t-test. Error bars depict standard deviation. (n = 3 biological replicates/group). Scale bar: 200 µm. i. Sequencing reads showing successful CRISPR mediated deletion of SC5D in KP4 cells. j. Measurement of KP4 cell growth following knockout of SC5D and treatment with increasing doses of RSL3 (left) or ML210 (right) for 96 h. Error bars depict standard error of the mean. (n = 3 biological replicates per group). k. Cancer Therapeutics Response Portal (CTRP) analysis of inhibitor sensitivity associated with PCSK9 low status across all cancers. (n = 860 cell lines). Inhibitors of GPX4 are indicated. Box represents mean with interquartile multiplier of 0.5.
Supplementary information
Supplementary Figures (download PDF )
Supplementary Figs. 1 and 2
Supplementary Video 1 (download MP4 )
PDAC cell colonization of the liver. Representative light-sheet microscopy video of whole liver 2 weeks post-injection, showing aSMA (white), HPAC-eGFP (green) and KP4-mCherry (red).
Supplementary Video 2 (download MP4 )
PDAC cell colonization of the lungs. Representative light-sheet microscopy video of whole lung 2 weeks post-injection, showing aSMA (white), HPAC-eGFP (green) and KP4-mCherry (red).
Supplementary Tables (download ZIP )
Supplementary Tables 1–9
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rademaker, G., Hernandez, G.A., Seo, Y. et al. PCSK9 drives sterol-dependent metastatic organ choice in pancreatic cancer. Nature 643, 1381–1390 (2025). https://doi.org/10.1038/s41586-025-09017-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09017-8
This article is cited by
-
Ferroptosis in cancer toward molecular insights and clinical translation in pancreatic cancer
Molecular Cancer (2026)
-
Ferroptosis and metastasis: molecular checkpoints, microenvironmental dynamics, and therapeutic opportunities
Molecular Cancer (2026)
-
Targeting ferroptosis and cuproptosis in gastrointestinal cancers: molecular mechanisms, metabolic vulnerabilities, and therapeutic interventions
Molecular Biomedicine (2025)
-
Hyperlipidemia drives tumor growth in a mouse model of obesity-accelerated breast cancer growth
Cancer & Metabolism (2025)
-
A marker for sterol-dependent metastatic tropism
Nature Structural & Molecular Biology (2025)