Abstract
Legume roots secure nitrogen by forming a symbiosis with soil rhizobia but remain resistant to pathogenic bacteria1,2,3,4. How this tolerance to rhizobia is achieved without compromising plant immunity is largely unknown. Here we identify the cytoplasmic kinase MtLICK1/2, which interacts with nodulation factor receptor MtLYK3 to drive symbiotic signalling and suppress plant immunity. Rhizobial infection and nodule development are defective in Mtlick1/2, phenocopying the Mtlyk3-1 mutant. MtLICK1/2 and MtLYK3 undergo reciprocal transphosphorylation during rhizobial symbiosis. Phosphorylated MtLYK3 activates the receptor-like kinase MtDMI2 to stimulate symbiotic signalling. MtLICK1/2 is activated in the rhizobial infection area to suppress plant immunity. Thus, MtLICK1/2 and MtLYK3 together amplify symbiotic signalling and dampen host immunity to enable legume–rhizobium symbiosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are available within this article and its Supplementary Information. Original gel blots are shown in Supplementary Fig. 1. Original data points in graphs are shown in the Source Data files. These data were also uploaded on Figshare (https://figshare.com/s/4881ae6bff042fc75ef4). The M. truncatula R108 Tnt1 transposon insertion lines of MtLICK1 (NF9810) and MtLICK2 (NF4329) were obtained from the Noble Foundation Tnt1 database (https://medicago-mutant.dasnr.okstate.edu/mutant/database.php). The protein and genome source was obtained from Phytozome (doe.gov). Source data are provided with this paper.
Code availability
No customized code was generated in this study.
References
Zipfel, C. & Oldroyd, G. E. Plant signalling in symbiosis and immunity. Nature 543, 328–336 (2017).
Martin, F. M., Uroz, S. & Barker, D. G. Ancestral alliances: plant mutualistic symbioses with fungi and bacteria. Science 356, eaad4501 (2017).
van Velzen, R., Doyle, J. J. & Geurts, R. A resurrected scenario: single gain and massive loss of nitrogen-fixing nodulation. Trends Plant Sci. 24, 49–57 (2019).
Poole, P., Ramachandran, V. & Terpolilli, J. Rhizobia: from saprophytes to endosymbionts. Nat. Rev. Microbiol. 16, 291–303 (2018).
Arrighi, J. F. et al. The Medicago truncatula lysin motif-receptor-like kinase gene family includes NFP and new nodule-expressed genes. Plant Physiol. 142, 265–279 (2006).
Haney, C. H. et al. Symbiotic rhizobia bacteria trigger a change in localization and dynamics of the Medicago truncatula receptor kinase LYK3. Plant Cell 23, 2774–2787 (2011).
Limpens, E. et al. LysM domain receptor kinases regulating rhizobial nod factor-induced infection. Science 302, 630–633 (2003).
Moling, S. et al. Nod factor receptors form heteromeric complexes and are essential for intracellular infection in Medicago nodules. Plant Cell 26, 4188–4199 (2014).
Mulder, L., Lefebvre, B., Cullimore, J. & Imberty, A. LysM domains of Medicago truncatula NFP protein involved in Nod factor perception. Glycosylation state, molecular modeling and docking of chitooligosaccharides and Nod factors. Glycobiology 16, 801–809 (2006).
Smit, P. et al. Medicago LYK3, an entry receptor in rhizobial nodulation factor signaling. Plant Physiol. 145, 183–191 (2007).
Radutoiu, S. et al. Plant recognition of symbiotic bacteria requires two LysM receptor-like kinases. Nature 425, 585–592 (2003).
Madsen, E. B. et al. A receptor kinase gene of the LysM type is involved in legume perception of rhizobial signals. Nature 425, 637–640 (2003).
Indrasumunar, A. et al. Inactivation of duplicated nod factor receptor 5 (NFR5) genes in recessive loss-of-function non-nodulation mutants of allotetraploid soybean (Glycine max L. Merr.). Plant Cell Physiol. 51, 201–214 (2010).
Indrasumunar, A. et al. Nodulation factor receptor kinase 1α controls nodule organ number in soybean (Glycine max L. Merr). Plant J. 65, 39–50 (2011).
Oldroyd, G. E. Speak, friend, and enter: signalling systems that promote beneficial symbiotic associations in plants. Nat. Rev. Microbiol. 11, 252–263 (2013).
Broghammer, A. et al. Legume receptors perceive the rhizobial lipochitin oligosaccharide signal molecules by direct binding. Proc. Natl Acad. Sci. USA 109, 13859–13864 (2012).
Bozsoki, Z. et al. Ligand-recognizing motifs in plant LysM receptors are major determinants of specificity. Science 369, 663–670 (2020).
Gourion, B., Berrabah, F., Ratet, P. & Stacey, G. Rhizobium-legume symbioses: the crucial role of plant immunity. Trends Plant Sci. 20, 186–194 (2015).
Cao, Y., Halane, M. K., Gassmann, W. & Stacey, G. The role of plant innate immunity in the legume-rhizobium symbiosis. Annu. Rev. Plant Biol. 68, 535–561 (2017).
Bravo, A., York, T., Pumplin, N., Mueller, L. A. & Harrison, M. J. Genes conserved for arbuscular mycorrhizal symbiosis identified through phylogenomics. Nat. Plants 2, 15208 (2016).
Journet, E. et al. Medicago truncatula ENOD11: a novel RPRP-encoding early nodulin gene expressed during mycorrhization in arbuscule-containing cells. Mol. Plant Microbe Interact. 14, 737–748 (2001).
Marsh, J. F. et al. Medicago truncatula NIN is essential for rhizobial-independent nodule organogenesis induced by autoactive calcium/calmodulin-dependent protein kinase. Plant Physiol. 144, 324–335 (2007).
Villamor, J. G. et al. Profiling protein kinases and other ATP binding proteins in Arabidopsis using Acyl-ATP probes. Mol. Cell Proteomics 12, 2481–2496 (2013).
Madsen, E. B. et al. Autophosphorylation is essential for the in vivo function of the Lotus japonicus Nod factor receptor 1 and receptor-mediated signalling in cooperation with Nod factor receptor 5. Plant J. 65, 404–417 (2011).
Klaus-Heisen, D. et al. Structure-function similarities between a plant receptor-like kinase and the human interleukin-1 receptor-associated kinase-4. J. Biol. Chem. 286, 11202–11210 (2011).
Nakagawa, T. et al. From defense to symbiosis: limited alterations in the kinase domain of LysM receptor-like kinases are crucial for evolution of legume-rhizobium symbiosis. Plant J. 65, 169–180 (2011).
Endre, G. et al. A receptor kinase gene regulating symbiotic nodule development. Nature 417, 959–962 (2002).
Ried, M. K., Antolin-Llovera, M. & Parniske, M. Spontaneous symbiotic reprogramming of plant roots triggered by receptor-like kinases. eLife 3, e03891 (2014).
Yoshida, S. & Parniske, M. Regulation of plant symbiosis receptor kinase through serine and threonine phosphorylation. J. Biol. Chem. 280, 9203–9209 (2005).
Gust, A. A. Peptidoglycan perception in plants. PLoS Pathog. 11, e1005275 (2015).
Liang, Y. et al. Nonlegumes respond to rhizobial Nod factors by suppressing the innate immune response. Science 341, 1384–1387 (2013).
DeFalco, T. A. & Zipfel, C. Molecular mechanisms of early plant pattern-triggered immune signaling. Mol. Cell 81, 3449–3467 (2021).
Liang, X. & Zhou, J. M. Receptor-like cytoplasmic kinases: central players in plant receptor kinase-mediated signaling. Annu. Rev. Plant Biol. 69, 267–299 (2018).
Hohmann, U., Lau, K. & Hothorn, M. The structural basis of ligand perception and signal activation by receptor kinases. Annu. Rev. Plant Biol. 68, 109–137 (2017).
Ma, X., Xu, G., He, P. & Shan, L. SERKing coreceptors for receptors. Trends Plant Sci. 21, 1017–1033 (2016).
Roth, R. et al. A rice serine/threonine receptor-like kinase regulates arbuscular mycorrhizal symbiosis at the peri-arbuscular membrane. Nat. Commun. 9, 4677 (2018).
Montero, H. et al. A mycorrhiza-associated receptor-like kinase with an ancient origin in the green lineage. Proc. Natl Acad. Sci. USA 118, e2105281118 (2021).
Wong, J. et al. A Lotus japonicus cytoplasmic kinase connects Nod factor perception by the NFR5 LysM receptor to nodulation. Proc. Natl Acad. Sci. USA 116, 14339–14348 (2019).
Irving, T. B. et al. KIN3 impacts arbuscular mycorrhizal symbiosis and promotes fungal colonisation in Medicago truncatula. Plant J. 110, 513–528 (2022).
Radutoiu, S. et al. LysM domains mediate lipochitin-oligosaccharide recognition and Nfr genes extend the symbiotic host range. EMBO J. 26, 3923–3935 (2007).
Stracke, S. et al. A plant receptor-like kinase required for both bacterial and fungal symbiosis. Nature 417, 959–962 (2002).
Boisson-Dernier, A. et al. Agrobacterium rhizogenes-transformed roots of Medicago truncatula for the study of nitrogen-fixing and endomycorrhizal symbiotic associations. Mol. Plant Microbe Interact. 14, 695–700 (2001).
Gietz, R. D. & Woods, R. A. Transformation of yeast by lithium acetate single stranded carrier DNA polyethylene glycol method. Methods Enzymol. 350, 87–96 (2002).
Akamatsu, A. et al. An OsCEBiP/OsCERK1-OsRacGEF1-OsRac1 module is an essential early component of chitin-induced rice immunity. Cell Host Microbe 13, 465–476 (2013).
Padmanabhan, M. S. et al. Novel positive regulatory role for the SPL6 transcription factor in the N TIR-NB-LRR receptor-mediated plant innate immunity. PLoS Pathog. 9, e1003235 (2013).
Jia, N., Zhu, Y. & Xie, F. An efficient protocol for model legume root protoplast isolation and transformation. Front. Plant Sci. 9, 670 (2018).
Feng, F. et al. A combination of chitooligosaccharide and lipochitooligosaccharide recognition promotes arbuscular mycorrhizal associations in Medicago truncatula. Nat. Commun. 10, 5047 (2019).
Plener, L., Manfredi, P., Valls, M. & Genin, S. PrhG, a transcriptional regulator responding to growth conditions, is involved in the control of the type III secretion system regulon in Ralstonia solanacearum. J. Bacteriol. 192, 1011–1019 (2010).
Wang, Y. et al. A bacterial effector protein uncovers a plant metabolic pathway involved in tolerance to bacterial wilt disease. Mol. Plant 14, 1281–1296 (2021).
Yamchi, A. et al. Proteomics analysis of Medicago truncatula response to infection by the phytopathogenic bacterium Ralstonia solanacearum points to jasmonate and salicylate defence pathways. Cell Microbiol. 20, e12796 (2018).
Salzer, P., Corbiere, H. & Boller, T. Hydrogen peroxide accumulation in Medicago truncatula roots colonized by the arbuscular mycorrhiza-forming fungus Glomus intraradices. Planta 208, 319–325 (1999).
Thordal-Christensen, H., Zhang, Z., Wei, D. & Collinge, D. B. Subcellular localization of H2O2 in plants. H2O2 accumulation in papillae and hypersensitive response during the barley-powdery mildew interaction. Plant J. 11, 1187–1194 (1997).
You, Q. et al. An E3 ubiquitin ligase-BAG protein module controls plant innate immunity and broad-spectrum disease resistance. Cell Host Microbe 20, 758–769 (2016).
Benezech, C. et al. Medicago-Sinorhizobium-Ralstonia co-infection reveals legume nodules as pathogen confined infection sites developing weak defenses. Curr. Biol. 30, 1–8 (2019).
Gimenez-Ibanez, S., Zamarreno, A. M., Garcia-Mina, J. M. & Solano, R. An evolutionarily ancient immune system governs the interactions between Pseudomonas syringae and an early-diverging land plant lineage. Curr. Biol. 29, 2270–2281.e4 (2019).
Erbs, G. et al. Peptidoglycan and muropeptides from pathogens Agrobacterium and Xanthomonas elicit plant innate immunity: structure and activity. Chem. Biol. 15, 438–448 (2008).
Gust, A. A. et al. Bacteria-derived peptidoglycans constitute pathogen-associated molecular patterns triggering innate immunity in Arabidopsis. J. Biol. Chem. 282, 32338–32348 (2007).
Liu, B. et al. Lysin motif-containing proteins LYP4 and LYP6 play dual roles in peptidoglycan and chitin perception in rice innate immunity. Plant Cell 24, 3406–3419 (2012).
Acknowledgements
We thank Life Science Editors for editing assistance on the manuscript; and L. Zhu and W. Cai for their help with microscope pictures and picture analysis. This research was supported by the National Science Foundation (32425001, 32088102, 32000189, 31730103 and 32050081), the CAS Project for Young Scientists in Basic Research (YSBR-011), the New Cornerstone Science Foundation (NCI202248), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0630000 and XDB27040207), the National Key R&D Program of China (2024YFA0918203, 2022YFF1001800 and 2019YFA0904703) and the Science and Technology Commission of Shanghai Municipality (24N12800200). E.W. acknowledges support from the XPLORER PRIZE.
Author information
Authors and Affiliations
Contributions
E.W., D.W. and R.J. directed the research. D.W. and R.J. performed most of the experiments, including the genetic phenotype analysis, subcellular localization, Y2H, co-IP, RNA analysis, in vivo and/or in vitro phosphorylation assays, PGN purification, ROS detection, MAPK detection, western blots, R. solanacearum spot inoculations and quantification. X.S., H.G., X.T., X.L. and H.D. contributed to molecular cloning, qRT–PCR and transformation work. A.Z. performed the pathogen infection assays. S.L. and K.X. contributed to the transformation work. C.T., J.Y. and W.C. provided important suggestions and analysis to rescue this project. A.P.M.’s laboratory provided the inoculation system and Medicago cultivation support for the pathogen inoculation assays. E.W. oversaw the entire study. E.W., D.W. and R.J. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The phylogenetic tree of LICK family, MtLICK1/2 do not interact with MtNFP or MtDMI2 in Y2H.
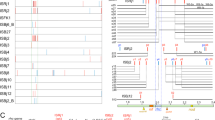
a, Phylogenetic tree of LICK family members in Medicago truncatula, Glycine max, Lotus japonicus, Oryza sativa, Arabidopsis thaliana, Populus tremula, Vitis vinifera, Nicotiana benthamiana, Brassica napus, Sisymbrium irio, and Marchantia polymorpha. The phylogenetic tree was constructed using maximum likelihood method. ECD, extracellular domain. TM, transmembrane domain. KD, kinase domain. ECD, TM and KD were predicted by TMHMM-2.0. Extracellular domain and transmembrane domain were predicted by TMHMM-2.0. b, Yeast two-hybrid (Y2H) assay of MtNFP or MtDMI2 cytoplasmic domain, MtLICK1 and MtLICK2 with MtLICK1. Interaction of BK-MtLYK3CD and AD-MtLICK1 as a positive control. Empty vector (EV) was used as a negative control. c, Yeast two-hybrid (Y2H) assay of MtNFP or MtDMI2 cytoplasmic domain, MtLICK1 and MtLICK2 with MtLICK2. Interaction of BK-MtLYK3CD and AD-MtLICK2 as a positive control. Empty vector (EV) was used as a negative control. For b and c, the experiments were repeated three times with similar results.
Extended Data Fig. 2 MtLYK3 interacts with MtLICK1/2 on the cell embrane.
a, MtLICK1/2 is localized at the plasma membrane of N. benthamiana epidermal cells. RFP-OsRAC1 and GFP-MtLICK1/2 were transiently co-expressed in N. benthamiana leaves transformed with p35S:RFP-OsRAC1 and p35S:GFP-MtLICK1/2. RFP-OsRAC1 is plasma membrane localization marker. Scale bars, 20 µm. b, MtLICK1 and MtLICK2 are co-localized with MtLYK3 at the plasma membrane of N. benthamiana epidermal cell, MtLYK3-RFP is plasma membrane localization receptor-like kinase. MtLYK3-RFP and GFP-MtLICK1/2 were transiently co-expressed in N. benthamiana leaves transformed with pLjUBQ:MtLYK3-RFP and p35S:GFP-MtLICK1/2. Scale bars, 20 µm. c, MtLYK3-Flag or MtBRI1-Flag and GFP-MtLICK1 or GFP-MtLICK2 were transiently co-expressed in N. benthamiana leaves and Co-IP assay was performed with a-Flag. MtBRI1-Flag and GFP-MtLICK1/2, GFP-MtLICK1/2 and Flag or GFP and MtLYK3-Flag were transiently co-expressed in N. benthamiana leaves serve as control. MtBRI1 was used as a negative plasma membrane protein control. d, MtLICK1/2-Flag or Flag were expressed in Medicago stable transgenic plants (Mtlyk3 mutant transformed with pMtLYK3:gMtLYK3-GFP) hairy roots transformed with pMtLICK1:MtLICK1WT-Flag, pMtLICK2:MtLICK2WT-Flag, and pLjUBQ:Flag and Co-IP assay was performed with a-GFP antibodies. Flag was expressed in Medicago stable transgenic plants (Mtlyk3 mutant transformed with pMtLYK3:gMtLYK3-GFP) and MtLICK1/2-Flag were expressed in wild-type plants (A17) by hairy root transformation serve as control. e, Detection of protein-protein interactions in Medicago (A17) root protoplast by BiFC. MtLYK3-Yn (N-terminal 174 amino acids of YFP) and MtLICK1/2-Yc (C-terminal 66 amino acids of YFP) were transiently co-expressed in Medicago (A17) root protoplasts transformed with p35S:MtLYK3-Yn and p35S:MtLICK1/2-Yc. Protein interactions were not detected in the other two combinations: MtLYK3-Yn and Yc empty vector (EV) as well as Yn empty vector (EV) and MtLICK1/2-Yc. Scale bars, 10 µm. Data are representative of three independent experiments. f,g, qRT-PCR was used to detect mRNA levels of MtLICK1 and MtLICK2 in wild-type (WT) R108 1-week roots at 6-hour post inoculation (hpi) with Nod factor (NF) (3 biological repetitions per group). MtEF-1 was used as the reference gene. Data are mean ± SEM. Each biological replicate is plotted as black dot and triangle. Statistical analysis was performed using Two-sided Student’s t-test compared to Mock. h,i, qRT-PCR was used to detect mRNA levels of MtLICK1 and MtLICK2 in R108 roots at 1 hpi, 6 hpi, 1 dpi, 14 dpi and 21dpi with S. meliloti 1021 (3 biological repetitions per group). MtEF-1 was used as the reference gene. Data are mean ± SEM. Each biological replicate is plotted as black dot and triangle. Statistical analysis was performed using Two-sided Student’s t-test compared to Mock. j, Expression patterns of MtLICK1/2 and MtLYK3 in root hairs were determined by GUS reporter analysis. The roots of wild-type plants were transformed with pMtLICK1/2: GUS or pMtLYK3: GUS and GUS activity detected by treating with X-Gluc at 37 °C for 8 h (see Methods). Scale bars, 100 µm. k, Schematic representation of MtLICK1 genomic sequence and identification of Tnt1 insertion mutant of MtLICK1. Exons are shown as red boxes. Localization of Tnt1 insertion is represented as green lines with a green triangle. l, Schematic representation of MtLICK2 genomic sequence and identification of Tnt1 insertion mutant of MtLICK2. Exons are shown as red boxes. Localization of Tnt1 insertion is represented as green lines with a green triangle. For c, f-i and k, l the experiments were repeated three times with similar results. For d, the experiments were repeated two times with similar results.
Extended Data Fig. 3 MtLICK1 and MtLICK2 are all involved in rhizobial symbiosis.
a, qRT-PCR was used to detect mRNA levels of MtLICK1 or MtLICK2 in WT (R108) and Mtlick1 or Mtlick2 roots (n = 3 technical replicates per group). MtEF-1 was used as the reference gene. Data are mean ± SEM. Each technical replicate is plotted as black dot and triangle. Statistical analysis was performed using Two-sided Student’s t-test compared to WT (R108). b, Number of nodules and bumps at 21 dpi with S. meliloti 1021 in Mtlick1 or Mtlick2 hairy roots transformed with empty vector (EV), pMtLICK1:genomicMtLICK1 or pMtLICK2:genomicMtLICK2. WT (R108)/EV serves as a positive control, Mtlick1/EV or Mtlick2/EV as a negative control. Data are mean ± SEM. (WT/EV, n = 12; Mtlick1/EV, n = 21; Mtlick1/gMtLICK1, n = 23; Mtlick2/EV, n = 19; Mtlick2/gMtLICK2, n = 22). Each plant analyzed is plotted as a black dot, square, triangle, cross and circle respectively). Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlick1/EV, Mtlick2/EV or WT/EV. c, RNA interference of MtLICK1 in Mtlick2 mutant plants. Nodules and bumps number of Mtlick2 mutant hairy roots transformed with RNAi empty vector (Mtlick2/EV) or MtLICK1 RNAi (Mtlick2/MtLICK1 RNAi) and transformed plants at 21 dpi with S. meliloti 1021. Mtlick2/EV serves as control. EV, empty vector. (EV, n = 13; RNAi, n = 13). Data are mean ± SEM. Each plant analyzed is plotted as black dot (EV) and triangle (RNAi). Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlick2/EV. d, qRT-PCR was used to detect mRNA levels of MtLICK1 in Mtlick2/EV and Mtlick2/MtLICK1 RNAi roots in transformed plants at 21 dpi with S. meliloti 1021. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological repetitions analyzed is plotted as black dot and triangle respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlick2/EV. e, Representative images of nodules in wild-type (WT) R108 and Mtlick1/2 seedlings grown on vermiculite at 21 dpi with S. meliloti 1021. Scale bars, 1 mm. f-j, Number of nodules and bumps at 21 dpi with S. meliloti 1021 in Mtlick1/2 hairy roots transformed with empty vector (EV), pMtLICK1:genomicMtLICK1 or pMtLICK2:genomicMtLICK2. WT (R108)/EV serves as a positive control, Mtlick1/2/EV as a negative control. Data are mean ± SEM. (WT/EV, n = 26; Mtlick1/2/EV, n = 13; Mtlick1/2/gMtLICK1, n = 26) (WT/EV, n = 26; Mtlick1/2/EV, n = 29; Mtlick1/2/gMtLICK2, n = 24). Each plant analyzed is plotted as a black dot, triangle and cross respectively). Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlick1/2/EV or WT/EV. In h,j, qRT-PCR was used to detect mRNA levels of MtLICK1 or MtLICK2 in transformed plants at 21 dpi with S. meliloti 1021. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological repetitions analyzed is plotted as black dot and triangle respectively. In g-j, statistical analysis was performed using Two-sided Student’s t-test compared to WT/EV. In f, representative images of nodule in transformed plants at 21 dpi with S. meliloti 1021. Scale bars, 5 mm. k, Representative images of nodule in transformed plants at 21 dpi with S. meliloti 1021 in Mtlick1/2 hairy roots transformed with pLjUBQ:GFP-MtLICK1/2. Scale bars, 1 mm. l-n, Overexpression of MtLICK1 and MtLICK2 in wild-type (WT) A17 plants. Number of nodules and bumps at 21 dpi with S. meliloti 1021 in A17 plants hairy roots transformed with empty vector (EV), pLjUBQ:MtLICK1 and pLjUBQ:MtLICK2. WT/EV used as negative control. WT/MtLICK1, n = 43; WT/MtLICK2, n = 28; EV, n = 38; each plant analyzed is plotted as black dot, triangle and cross respectively. Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to WT/EV. In m,n, qRT-PCR was used to detect mRNA levels of MtLICK1 or MtLICK2 in transformed plants at 21 dpi with S. meliloti 1021. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological repetitions analyzed is plotted as black dot and triangle respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to WT/EV. o,p, qRT-PCR was used to detect mRNA levels of symbiosis marker genes in WT (R108), Mtlick1, Mtlick2 and Mtlick1/2 at 14 dpi with S. meliloti 1021. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological repetitions analyzed is plotted as black dot and triangle respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to WT Sm. Sm, S. meliloti 1021. q,r, qRT-PCR was used to detect mRNA levels of symbiosis marker genes (MtENOD11 and MtNIN) in WT (A17), Mtlyk3-1 (Mthcl-1), WT (R108) and Mtlick1/2 7 d roots at 1 h, 3 h, and 24 h post Nod factor treatment. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological repetitions analyzed is plotted as black dot and triangle respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to WT (A17) or WT (R108) treated with Nod factor (NF). s, qRT-PCR was used to detect mRNA levels of MtLICK3 in transformed plants Mtlick1/2/MtLICK3 RNAi roots. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological repetitions analyzed is plotted as black dot and triangle respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to WT (R108)/ EV or Mtlick1/2/EV. All the experiments above were repeated three times with similar results.
Extended Data Fig. 4 MtLYK3 regulates MtLICK1/2 kinase activity by phosphorylation.
a, Schematic representation of MtLICK1 and MtLICK2, including the location of phosphorylation sites and ATP binding site in the kinase domain. b, Auto- and trans-phosphorylation activities of the indicated affinity purified recombinant MtLICK1WT/MtLICK2WT and mutants. Loading gels (bottom) were stained with CBB. The histograms below show a statistical analysis of the relative signal intensity based on three repeated in vitro phosphorylation assay data. Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to MtLICK1WT or MtLICK2WT. Asterisks indicate significant difference. c, Number of nodules and bumps at 21 dpi with S. meliloti 1021 in Mtlick1/2 hairy roots transformed with pLjUBQ: MtLICK1WT (LICK1), empty vector (EV), pLjUBQ: MtLICK1S60A (S60A), pLjUBQ: MtLICK1S60D (S60D), pLjUBQ: MtLICK1S203A (S203 A), pLjUBQ: MtLICK1S203D (S203D), pLjUBQ: MtLICK1T206A (T206A) and, pLjUBQ: MtLICK1T206D (T206D). Mtlick1/2/MtLICK1WT served as positive control, Mtlick1/2/EV served as negative control. (WT, n = 17; EV, n = 17; S60A, n = 13; S60D, n = 18; S201A, n = 12; S201D, n = 12; T206A, n = 13; T206D, n = 14; each plant analyzed is plotted as black dot, black triangle, cross, hollow triangle, square, rhombus, hollow circle and hollow square respectively.) Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlick1/2/pLjUBQ: MtLICK1WT. d, Number of nodules and bumps at 21 dpi with S. meliloti 1021 in Mtlick1/2 hairy roots transformed with pLjUBQ:MtLICK2WT (LICK2), empty vector (EV), pLjUBQ: MtLICK2S55A (S55A), pLjUBQ: MtLICK2S55D (S55D), pLjUBQ: MtLICK2S198A (S198A), pLjUBQ: MtLICK2S198D (S198A), pLjUBQ: MtLICK2T201A (T201A) and, pLjUBQ: MtLICK2T201D (T201D). Mtlick1/2/MtLICK2WT served as positive control, Mtlick1/2/EV served as negative control. (WT, n = 47; EV, n = 27; S55A, n = 41; S55D, n = 33; T201A, n = 19; T201D, n = 10; each plant analyzed is plotted as black dot, black triangle, cross, hollow triangle, square, rhombus, hollow circle and hollow square respectively.) Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlick1/2/pLjUBQ: MtLICK2WT. e,f, qRT-PCR was used to detect mRNA levels of MtLICK1WT/3A/3D or MtLICK2WT/3A/3D in Mtlick1/2 hairy roots transformed with pLjUBQ:MtLICK1/2WT, pLjUBQ:MtLICK1/23A and pLjUBQ:MtLICK1/23D at 21 dpi with S. meliloti 1021. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological repetitions analyzed is plotted as black dot and triangle respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to WT/EV. g, Auto- and trans-phosphorylation activities of the indicated affinity purified recombinant MtLYK3WT/MtLYK3G334E (MtHCL-1). Loading control gel (bottom) stained with CBB. All the experiments above were repeated three times with similar results.
Extended Data Fig. 5 Phosphorylation of MtLICK1/2 by MtLYK3 regulates the interaction between MtLICK1/2 and MtLYK3.
a,b, MtLICK1/2-Flag or Flag was expressed in Medicago stable transgenic plants (MtLYK3-GFP, Mtlyk3 mutant transformed with pMtLYK3:gMtLYK3-GFP) hairy roots transformed with pLjUBQ:MtLICK1/2-Flag or pLjUBQ:Flag by hairy root transformation. After Nod factor treatment as indicated time (0 min, 15 min and 60 min), Co-IP assay was performed with a-GFP. Flag was expressed in Medicago stable transgenic plants (pMtLYK3:gMtLYK3-GFP) and MtLICK1/2-Flag were expressed in wild-type A17 roots by hairy root transformation serve as control. The histograms right show a statistical analysis of the relative signal intensity based on three repeated Co-IP assay data. Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to 0 min Nod factor treatment. c, Yeast two-hybrid (Y2H) assay of MtLYK3 cytoplasmic domain with wild-type and mutant MtLICK1/2. Interaction of BK-MtLYK3CD and AD-MtLICK1/2 as the positive control. Empty vector (EV) was used as a negative control. For a and b, the experiments were repeated three times with similar results. For c, the experiments were repeated two times with similar results.
Extended Data Fig. 6 Phosphorylated MtLICK1/2 are less efficient in forming a complex with MtLYK3.
a,b, Western blot detection of MtLICK1WT and mutants in yeast. Western blot membrane was immunoblotted with a-HA and a-Myc to indicate the amount of MtLICK1 or MtLYK3CD respectively. c,d, Western blot detection of MtLICK2WT and mutants in yeast. Western blot membrane was immunoblotted with a-HA and a-Myc to indicate the amount of MtLICK2 or MtLYK3CD respectively. e,f, MtLICK1/2WT or 3D -Flag or Flag was expressed in Medicago stable transgenic plants (pMtLYK3:gMtLYK3-GFP) hairy roots transformed with pLjUBQ:MtLICK1/2WT or 3D or pLjUBQ:Flag by hairy root transformation. Co-IP assay was performed with a-GFP. Flag was expressed in Medicago stable transgenic plants (pMtLYK3:gMtLYK3-GFP) by hairy root transformation serves as control. The histograms right show a statistical analysis of the relative signal intensity based on three repeated Co-IP assay data. Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to MtLICK1WT or MtLICK2WT. g,h, MtLYK3-HA and GFP-MtLICK1/2WT or GFP-MtLICK1/23D-Flag were transiently co-expressed in N. benthamiana leaves transformed with pLjUBQ:MtLYK3-HA, p35S:GFP-MtLICK1/2WT and p35S:GFP-MtLICK1/23D-Flag. Co-IP assay was performed with a-GFP. GFP and HA or GFP and MtLYK3-HA were transiently co-expressed in N. benthamiana leaves serve as control. The histograms right show a statistical analysis of the relative signal intensity based on three repeated Co-IP assay data. Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to MtLICK1WT or MtLICK2WT. All the experiments were repeated three times with similar results.
Extended Data Fig. 7 Phosphorylation of MtLYK3 is essential for MtLYK3 kinase activity and symbiotic signaling.
a, Kinase activity of affinity purified recombinant MtLYK3CD and MtLICK1. Loading control gel (bottom) stained with CBB. b, Trans-phosphorylation activity of immunoprecipitated MtLYK3-Flag from N. benthamiana leaves transformd with pLjUBQ:MtLYK3-Flag (expressing MtLYK3-Flag) or transformd with pLjUBQ:MtLYK3-Flag and p35S:GFP-MtLICK1/2 (co-expressing MtLYK3-Flag and GFP-MtLICK1/2) against the universal substrate myelin basic protein (MyBP). Immunoprecipitation was performed with a-Flag agarose beads. The same quantity was blotted with a-Flag and a-GFP to indicate the amount of MtLYK3-Flag or GFP-MtLICK1/2 respectively. Loading control gel (bottom) stained with CBB to indicate the amount of MyBP. The histograms right show a statistical analysis of the relative signal intensity based on three repeated in vitro phosphorylation assay data. Data are mean ± SEM. c, Alignment of the amino acid sequences around the activation loop (AL) and YAQ regions of the LysM receptor kinases. The locations of the aEF helix, aF helix and aEF/aF loop in MtLYK3 and AtCERK1 are indicated above and below the alignments respectively. Consensus amino acid residues for all sequences are indicated by letter. The activation segment, defined as the region between two conserved tripeptide motifs (DFG-PE). d, Kinase activity of affinity purified recombinant MtLYK3CDWT, MtLYK3CDY483F and MtLYK3CDK349E against a MyBP substrate. MyBP, Myelin Basic Protein. Loading control gel (bottom) stained with CBB. e, Detection of auto-phosphorylation on affinity-purified recombinant MtLYK3 cytoplasmic domain (MtLYK3CD) following an in vitro kinase assay with cold ATP. Three membranes were immunoblotted with a-pS (Serine), a-pT (Threonine) and a-pY (Tyrosine) antibodies respectively. Loading control gel (bottom) stained with CBB. f, qRT-PCR was used to detect mRNA levels of symbiosis marker genes in Mtlyk3-1 (Mthcl-1) hairy roots transformed with pLjUBQ:MtLYK3WT, empty vector (EV), and pLjUBQ:MtLYK3Y483F at 21 dpi; each replicate is plotted as black dot, triangle and cross respectively. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlyk3-1/pLjUBQ:MtLYK3WT. g, MtLYK3 protein accumulation in Mtlyk3-1 (Mthcl-1) hairy roots transformed with pLjUBQ:MtLYK3WT (WT), pLjUBQ:MtLYK3Y483F (Y483F) and empty vector (EV). Western blot with a-Flag antibodies. Loading control gel (bottom) stained with CBB. h, MtLYK3 protein accumulation in N. benthamiana leaves transformed with pLjUBQ:MtLYK3WT (WT), pLjUBQ:MtLYK3Y483F (Y483F) and empty vector (EV). Western blot with a-Flag antibodies. Loading control gel (bottom) stained with CBB. i, qRT-PCR was used to detect mRNA levels of MtLYK3WT-Flag and MtLYK3Y483F-Flag in Mtlyk3-1 (Mthcl-1) hairy roots transformed with pLjUBQ:MtLYK3WT (WT), pLjUBQ:MtLYK3Y483F (Y483F) and empty vector (EV). MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological replicate is plotted as black dot, triangle and cross respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlyk3-1/MtLYK3WT. All the experiments above were repeated three times with similar results.
Extended Data Fig. 8 Phosphorylation of MtLYK3Y488 is required for rhizobial symbiosis.
a, Detection of Y488 phosphorylation using a-pY488 specific antibodies on affinity-purified recombinant MtLYK3 cytoplasmic domain (MtLYK3CD) following an in vitro kinase assay with cold ATP. Membrane was immunoblotted with a-pY488 antibodies. Loading control gel (bottom) stained with CBB. b, MtLYK3 protein accumulation in Mtlyk3-1 (Mthcl-1) hairy roots transformed with pLjUBQ:MtLYK3WT (WT), pLjUBQ:MtLYK3Y488F (Y488F) and empty vector (EV). Western blot with a-Flag antibodies. Loading control gel (bottom) stained with CBB. c, MtLYK3 protein accumulation in N. benthamiana leaves transformed with pLjUBQ:MtLYK3WT (WT), pLjUBQ:MtLYK3Y488F (Y488F) and empty vector (EV). Western blot with a-Flag antibodies. The other gel with same loading quantity was stained with CBB for loading control. d, qRT-PCR was used to detect mRNA levels of MtLYK3WT-Flag and MtLYK3Y488F-Flag in Mtlyk3-1 (Mthcl-1) hairy roots transformed with pLjUBQ:MtLYK3WT (WT), empty vector (EV) and pLjUBQ:MtLYK3Y488F (Y488F). MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological replicate is plotted as black dot, triangle and triangle cross respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlyk3-1/MtLYK3WT. e,f,g, qRT-PCR was used to detect mRNA levels of symbiosis markers in Mtlyk3-1 (Mthcl-1) hairy roots transformed with pLjUBQ:MtLYK3WT (WT), empty vector (EV) and pLjUBQ:MtLYK3Y488F (Y488F) at 7 dpi e, 14 dpi f, and 21 dpi g; each biological replicate is plotted as black dot, triangle and cross respectively. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological replicate is plotted as black dot, square and triangle. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtlyk3-1/pLjUBQ:MtLYK3WT. h, Phosphorylation activity of the indicated affinity purified recombinant MtLYK3CDWT, MtLYK3CDY488F and MtLYK3CDK349E against an MBP-MtDMI2K624E substrate. Loading control gel (bottom) stained with CBB. The histograms right show a statistical analysis of the relative signal intensity based on three repeated in vitro phosphorylation assay data. Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to MtLYK3CDWT. i, Schematic representation of the phosphorylation sites of MtDMI2 mediated by MtLYK3 through LC-MS. JMD, juxtamembrane domain. CTT, C-terminal tail. j, Auto- and trans-phosphorylation activities of the indicated affinity purified recombinant MtDMI2WT/MtDMI213A against a MyBP substrate. MyBP, Myelin Basic Protein. Loading control gel (bottom) stained with CBB. The histograms right show a statistical analysis of the relative signal intensity based on three repeated in vitro phosphorylation assay data. Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to MtDMI2WT. k,l, Number of nodules and bumps at 21 dpi with S. meliloti 1021 in Mtdmi2-1 hairy roots transformed with pLjUBQ:MtDMI2WT (WT), empty vector (EV), and pLjUBQ:MtDMI213A (13A). Mtdmi2-1/EV served as negative control. (WT, n = 34; EV, n = 51; 13A, n = 32; each plant analyzed is plotted as black dot, triangle, and cross respectively). Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtdmi2-1/pLjUBQ: MtDMI2WT or Mtdmi2-1/EV. In l, representative images of nodules in transformed plants at 21 dpi with S. meliloti 1021. DsRed protein expressed from the construct was used as a selection marker for transformations. Scale bars, 5 mm. m, qRT-PCR was used to detect mRNA levels of symbiosis markers in Mtdmi2-1 hairy roots transformed with pLjUBQ:MtDMI2WT (WT), empty vector (EV) and pLjUBQ:MtDMI213A (13A) at 21 dpi. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Each biological replicate is plotted as black dot, square and triangle. Statistical analysis was performed using Two-sided Student’s t-test compared to Mtdmi2-1/pLjUBQ:MtDMI2WT. All the experiments above were repeated three times with similar results.
Extended Data Fig. 9 MtLICK1 interact with MtRBOHs to regulate plant immunity.
a,b,c, qRT-PCR was used to detect mRNA levels of immunity marker genes in transformed plants WT (R108) and Mtlick1/2 at 2 days, post S. meliloti 1021 inoculation. MtEF-1 was used as the reference gene. (3 biological repetitions per group). d-i, qRT-PCR was used to detect mRNA levels of immunity marker genes (MtPR4, MtPR10 and MtWRKY) in transformed plants Mtlick1/2/MtLICK1WT, Mtlick1/2/MtLICK13A, Mtlick1/2/MtLICK13D, Mtlick1/2/MtLICK2WT, Mtlick1/2/MtLICK23A, and Mtlick1/2/MtLICK23D at 3 days, post S. meliloti 1021 inoculation. MtEF-1 was used as the reference gene. (3 biological repetitions per group). Data are mean ± SEM. Statistical analysis was performed using one-way ANOVA (Tukey test). Different letters indicate significant difference (P < 0.05). j, ROS production following treatment of wild-type (WT) R108 and Mtlick1/2 root segments with 0.4 mg/mL peptidoglycan (PGN) over 40 min. WT-Mock, n = 17; Mtlick1/2-Mock, n = 20; WT-PGN, n = 21; Mtlick1/2-PGN, n = 18). Data are mean ± SEM. Each plant analyzed is plotted as dots and triangle respectively. Statistical analysis was performed using Two-sided Student’s t-test compared to WT-PGN. RLU, relative luminescence unit. k, ROS production following treatment of wild-type (WT) R108 and Mtlick1/2 root segments with 0.4 mg/mL PGN derived from S. meliloti 1021(WT(R108) Mock, n = 21; WT(R108) PGN, n = 20; Mtlick1/2 Mock, n = 19; Mtlick1/2 PGN, n = 17). Data are mean ± SEM. Each plant analyzed is plotted as dots and triangle respectively. RLU, relative luminescence unit. l,m,n, R. solanacearum soil-drenching inoculation assays in Medicago truncatula. In l, the results are presented as disease progression, showing the average wilting symptoms on a scale from 0 to 4. Healthy plants (score 0), leaf curling (score 1), wilting petioles (score 2) and leaf chlorosis (score 3) and whole plant collapse (score 4). (WT (R108), n = 36; Mtlick1, n = 36; Mtlick2, n = 36; Mtlick1/2, n = 34). m, shows the survival analysis of data in l, Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to wild-type plants (R108). n, Phenotype of plant survival 14 days post-inoculation with R. solanacearum. Scale bar, 2 cm. o, DAB staining of 7-day Medicago roots pretreated with R. solanacearum for 3 h. Scale bars, 0.5 mm. The below histograms show a statistical analysis of the relative staining intensity based on three repeated DAB staining assay data. The relative staining intensity was detected by using Image J software (WT (R108), n = 11; Mtlick1, n = 11; Mtlick2, n = 11; Mtlick1/2, n = 11; each plant analyzed is plotted as black dot, triangle, box and cross respectively). Data are mean ± SEM. Statistical analysis was performed using Two-sided Student’s t-test compared to WT (R108). p, GFP-MtLICK1/2 and MtRBOHB/C-HA were transiently co-expressed in N. benthamiana leaves and Co-IP assay was performed with a-GFP antibodies. GFP-MtLICK1/2 and HA or MtLTI6b-GFP (low temperature–induced protein 6B, PM marker) and MtRBOHB/C-HA or GFP and MtRBOHB/C-HA were transiently co-expressed in N. benthamiana leaves serve as control. q, Number of infection events at 7 dpi with S. meliloti 1021-LacZ (Sm) in wild type (R108) plants with H2O2 treatment. The plants, without H2O2 treatment, were used as control. (0 mM, n = 20; 1 mM, n = 20; 4 mM, n = 20). Data are mean ± SEM. Each plant analyzed is plotted as dots, triangles and circles respectively. Statistical analysis was performed using one-way ANOVA (Tukey test). Different letters indicate significant difference (P < 0.05). r, Number of nodules and bumps at 14 dpi with S. meliloti 1021 (Sm) in wild type (R108) plants with H2O2 treatment. The plants, without H2O2 treatment, were used as control. (0 mM, n = 20; 1 mM, n = 20; 4 mM, n = 20). Data are mean ± SEM. Each plant analyzed is plotted as dots, circles, triangles, crosses and boxes respectively. Statistical analysis was performed using Two-sided Student’s t-test. For a-p, the experiments were repeated three times with similar results. For q and r, the experiments were repeated two times with similar results.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figure 1 and Supplementary Tables 1–5.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, D., Jin, R., Shi, X. et al. A kinase mediator of rhizobial symbiosis and immunity in Medicago. Nature 643, 768–775 (2025). https://doi.org/10.1038/s41586-025-09057-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09057-0
This article is cited by
-
Transcriptome-guided discovery of novel plant-associated genes in a rhizosphere Pseudomonas
Microbiome (2025)
-
MtLICK1/2: gatekeepers of symbiosis and immunity in Medicago truncatula
Science China Life Sciences (2025)