Abstract
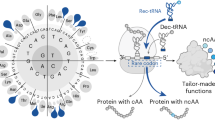
The incorporation of non-canonical amino acids (ncAAs) enables customized chemistry to tailor protein functions1,2,3. Genetic code expansion offers a general approach for ncAA encoding by reassigning stop codons as the ‘blank’ codon; however, it is not completely orthogonal to translation termination for cellular transcripts. Here, to generate more bona fide blank codons, we developed an RNA codon-expansion (RCE) strategy that introduces and decodes bioorthogonally assignable pseudouridine (Ψ) codons (ΨGA, ΨAA or ΨAG) on specified mRNA transcripts to incorporate ncAAs in mammalian cells. The RCE strategy comprises a programmable guide RNA4, an engineered decoder tRNA, and aminoacyl-tRNA synthetase. We first developed the RCE(ΨGA) system, which incorporates functional ncAAs into proteins via the ΨGA codon, demonstrating a higher translatome-wide and proteomic specificity compared with the genetic code expansion system. We further expanded our strategy to produce the RCE(ΨAA) and RCE(ΨAG) systems, with all three Ψ codon:(Ψ codon)-tRNAPyl pairs exhibiting mutual orthogonality. Moreover, we demonstrated that the RCE system cooperates compatibly with the genetic code expansion strategy for dual ncAA encoding. In sum, the RCE method utilized Ψ as a post-transcriptional ‘letter’ to encode and decode RNA codons in specific mRNA transcripts, opening a new route for genetic alphabet expansion and site-specific ncAA incorporation in eukaryotic cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All next-generation sequencing data generated for this study have been deposited in the NCBI Sequence Read Archive (SRA) under accession code PRJNA1090628. The human reference genome GRCh38 (December 2013) was downloaded from the following link: https://hgdownload.soe.ucsc.edu. The H. sapiens proteome database (UP000005640) was downloaded from https://www.uniprot.org. The custom database containing reporter sequences featuring 20 different natural amino acids at positions corresponding to the premature termination codon location is provided in Supplementary Table 1. The tRNA reference sequences were derived from GtRNAdb81 (http://gtrnadb.ucsc.edu/; accessed September 2024). ISR-related genes were collected from the GeneCards database (https://www.genecards.org/; accessed September 2024). Source data are provided with this paper.
Code availability
Custom codes are available on GitHub (https://github.com/yanxueqing621/RCE_project). These scripts include the pipelines for off-target Ψ sites identification based on PRAISE sequencing data, evaluation of potential off-target readthrough events based on Ribo-seq data, and RNA-seq data analysis.
Change history
11 August 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41586-025-09464-3
References
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Lemke, E. A. The exploding genetic code. ChemBioChem 15, 1691–1694 (2014).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Song, J. et al. CRISPR-free, programmable RNA pseudouridylation to suppress premature termination codons. Mol. Cell 83, 139–155.e139 (2023).
Neumann, H., Wang, K., Davis, L., Garcia-Alai, M. & Chin, J. W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 464, 441–444 (2010).
Orelle, C. et al. Protein synthesis by ribosomes with tethered subunits. Nature 524, 119–124 (2015).
Fried, S. D., Schmied, W. H., Uttamapinant, C. & Chin, J. W. Ribosome subunit stapling for orthogonal translation in E. coli. Angew. Chem. Int. Ed. 54, 12791–12794 (2015).
Isaacs, F. J. et al. Precise manipulation of chromosomes in vivo enables genome-wide codon replacement. Science 333, 348–353 (2011).
Lajoie, M. J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Ostrov, N. et al. Design, synthesis, and testing toward a 57-codon genome. Science 353, 819–822 (2016).
Wang, K. et al. Defining synonymous codon compression schemes by genome recoding. Nature 539, 59–64 (2016).
Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Nyerges, A. et al. Synthetic genomes unveil the effects of synonymous recoding. Preprint at bioRxiv https://doi.org/10.1101/2024.06.16.599206 (2024).
Grome, M. W. et al. Engineering a genomically recoded organism with one stop codon. Nature 639, 512–521 (2025).
Malyshev, D. A. et al. A semi-synthetic organism with an expanded genetic alphabet. Nature 509, 385–388 (2014).
Fischer, E. C. et al. New codons for efficient production of unnatural proteins in a semisynthetic organism. Nat. Chem. Biol. 16, 570–576 (2020).
Thompson, D. B. et al. The future of multiplexed eukaryotic genome engineering. ACS Chem. Biol. 13, 313–325 (2018).
Reinkemeier, C. D., Girona, G. E. & Lemke, E. A. Designer membraneless organelles enable codon reassignment of selected mRNAs in eukaryotes. Science 363, eaaw2644 (2019).
Reinkemeier, C. D. & Lemke, E. A. Dual film-like organelles enable spatial separation of orthogonal eukaryotic translation. Cell 184, 4886–4903.e4821 (2021).
Hao, M. et al. Tracking endogenous proteins based on RNA editing-mediated genetic code expansion. Nat. Chem. Biol. 20, 721–731 (2024).
Ranjan, N. & Leidel, S. A. The epitranscriptome in translation regulation: mRNA and tRNA modifications as the two sides of the same coin? FEBS Lett. 593, 1483–1493 (2019).
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2021 update. Nucleic Acids Res. 50, D231–D235 (2022).
Shi, H., Chai, P., Jia, R. & Fan, X. Novel insight into the regulatory roles of diverse RNA modifications: Re-defining the bridge between transcription and translation. Mol. Cancer 19, 78 (2020).
Karijolich, J. & Yu, Y.-T. Converting nonsense codons into sense codons by targeted pseudouridylation. Nature 474, 395–398 (2011).
Luo, N. et al. Near-cognate tRNAs increase the efficiency and precision of pseudouridine-mediated readthrough of premature termination codons. Nat. Biotechnol. 48, 114–123 (2025).
Charette, M. & Gray, M. W. Pseudouridine in RNA: what, where, how, and why. IUBMB Life 49, 341–351 (2000).
Zhang, M. et al. Quantitative profiling of pseudouridylation landscape in the human transcriptome. Nat. Chem. Biol. 19, 1185–1195 (2023).
Hofhuis, J. et al. The functional readthrough extension of malate dehydrogenase reveals a modification of the genetic code. Open Biol. 6, 160246 (2016).
Dunkelmann, D. L., Willis, J. C. W., Beattie, A. T. & Chin, J. W. Engineered triply orthogonal pyrrolysyl–tRNA synthetase/tRNA pairs enable the genetic encoding of three distinct non-canonical amino acids. Nat. Chem. 12, 535–544 (2020).
Seelam Prabhakar, P., Takyi, N. A. & Wetmore, S. D. Posttranscriptional modifications at the 37th position in the anticodon stem-loop of tRNA: structural insights from MD simulations. RNA 27, 202–220 (2021).
Nozawa, K. et al. Pyrrolysyl-tRNA synthetase–tRNAPyl structure reveals the molecular basis of orthogonality. Nature 457, 1163–1167 (2009).
Suzuki, T. et al. Crystal structures reveal an elusive functional domain of pyrrolysyl-tRNA synthetase. Nat. Chem. Biol. 13, 1261–1266 (2017).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Wang, J. et al. AAV-delivered suppressor tRNA overcomes a nonsense mutation in mice. Nature 604, 343–348 (2022).
Hollingsworth, T. J. & Gross, A. K. The severe autosomal dominant retinitis pigmentosa rhodopsin mutant Ter349Glu mislocalizes and induces rapid rod cell death. J. Biol. Chem. 288, 29047–29055 (2013).
Vidal, R. et al. A stop-codon mutation in the BRI gene associated with familial British dementia. Nature 399, 776–781 (1999).
Shibata, N. et al. Degradation of stop codon read-through mutant proteins via the ubiquitin–proteasome system causes hereditary disorders. J. Biol. Chem. 290, 28428–28437 (2015).
Namy, O., Duchateau-Nguyen, G. & Rousset, J.-P. Translational readthrough of the PDE2 stop codon modulates cAMP levels in Saccharomyces cerevisiae. Mol. Microbiol. 43, 641–652 (2002).
Capone, J. P., Sharp, P. A. & RajBhandary, U. L. Amber, ochre and opal suppressor tRNA genes derived from a human serine tRNA gene. EMBO J. 4, 213–221 (1985).
Wangen, J. R. & Green, R. Stop codon context influences genome-wide stimulation of termination codon readthrough by aminoglycosides. eLife 9, e52611 (2020).
Schueren, F. et al. Peroxisomal lactate dehydrogenase is generated by translational readthrough in mammals. eLife 3, e03640 (2014).
Shi, N. et al. Restoration of dystrophin expression in mice by suppressing a nonsense mutation through the incorporation of unnatural amino acids. Nat. Biomed. Eng. 6, 195–206 (2022).
Yeatman, T. J. A renaissance for SRC. Nat. Rev. Cancer 4, 470–480 (2004).
Zhang, G. et al. Bioorthogonal chemical activation of kinases in living systems. ACS Cent. Sci. 2, 325–331 (2016).
Sletten, E. M. & Bertozzi, C. R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. 48, 6974–6998 (2009).
Li, J. & Chen, P. R. Development and application of bond cleavage reactions in bioorthogonal chemistry. Nat. Chem. Biol. 12, 129–137 (2016).
Li, J., Jia, S. & Chen, P. R. Diels–Alder reaction-triggered bioorthogonal protein decaging in living cells. Nat. Chem. Biol. 10, 1003–1005 (2014).
Bian, Y. et al. Ultra-deep tyrosine phosphoproteomics enabled by a phosphotyrosine superbinder. Nat. Chem. Biol. 12, 959–966 (2016).
Jang, H. S., Jana, S., Blizzard, R. J., Meeuwsen, J. C. & Mehl, R. A. Access to faster eukaryotic cell labeling with encoded tetrazine amino acids. J. Am. Chem. Soc. 142, 7245–7249 (2020).
Bryson, D. I. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat. Chem. Biol. 13, 1253–1260 (2017).
Ding, W. et al. Rare codon recoding for efficient noncanonical amino acid incorporation in mammalian cells. Science 384, 1134–1142 (2024).
Liang, S.-H. & Clarke, M. F. Regulation of p53 localization. Eur. J. Biochem. 268, 2779–2783 (2001).
O’Keefe, K., Li, H. & Zhang, Y. Nucleocytoplasmic shuttling of p53 is essential for MDM2-mediated cytoplasmic degradation but not ubiquitination. Mol. Cell. Biol. 23, 6396–6405 (2003).
Gautier, A. et al. Genetically encoded photocontrol of protein localization in mammalian cells. J. Am. Chem. Soc. 132, 4086–4088 (2010).
Guangcan, S. et al. How to use open-pFind in deep proteomics data analysis?—A protocol for rigorous identification and quantitation of peptides and proteins from mass spectrometry data. Biophys. Rep. 7, 207–226 (2021).
Wilson, C., Chen, P. J., Miao, Z. & Liu, D. R. Programmable m6A modification of cellular RNAs with a Cas13-directed methyltransferase. Nat. Biotechnol. 38, 1431–1440 (2020).
Liu, X.-M., Zhou, J., Mao, Y., Ji, Q. & Qian, S.-B. Programmable RNA N6-methyladenosine editing by CRISPR–Cas9 conjugates. Nat. Chem. Biol. 15, 865–871 (2019).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Gruber, A. R., Lorenz, R., Bernhart, S. H., Neuböck, R. & Hofacker, I. L. The Vienna RNA Websuite. Nucleic Acids Res. 36, W70–W74 (2008).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2013).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in unique molecular identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Lauria, F. et al. riboWaltz: optimization of ribosome P-site positioning in ribosome profiling data. PLoS Comp. Biol. 14, e1006169 (2018).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Dong, M. et al. Sensitive, robust, and cost-effective approach for tyrosine phosphoproteome analysis. Anal. Chem. 89, 9307–9314 (2017).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41–44 (2020).
Shaw, J. J. & Green, R. Two distinct components of release factor function uncovered by nucleophile partitioning analysis. Mol. Cell 28, 458–467 (2007).
Gagnon, K. T. & Maxwell, E. S. Electrophoretic mobility shift assay for characterizing RNA–protein interaction. Methods Mol. Biol. 703, 275–291 (2011).
Case, D. A. et al. AmberTools. J. Chem. Inf. Model. 63, 6183–6191 (2023).
Eastman, P. et al. OpenMM 8: molecular dynamics simulation with machine learning potentials. J. Phys. Chem. B 128, 109–116 (2024).
McGibbon, RobertT. et al. MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 109, 1528–1532 (2015).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins Struct. Funct. Bioinformatics 78, 1950–1958 (2010).
Zgarbová, M. et al. Refinement of the Cornell et al. nucleic acids force field based on reference quantum chemical calculations of glycosidic torsion profiles. J. Chem. Theory Comput. 7, 2886–2902 (2011).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Chan, P. P. & Lowe, T. M. GtRNAdb 2.0: an expanded database of transfer RNA genes identified in complete and draft genomes. Nucleic Acids Res. 44, D184–D189 (2016).
Acknowledgements
The authors thank the National Center for Protein Sciences at Peking University in Beijing, China, for assistance with the 4150 TapeStation System (G. Li), mass spectrometry (D. Liu and Q. Zhang), Zeiss LSM 980 confocal microscope and Spin SR confocal microscope (L. Fu); G. Jia, R. Liu, F. Lin, X. Rao, C. Shao, M. Zhang and Y. Ma for discussions and materials; the Center for Quantitative Biology at Peking University for assistance with the ImageXpress Micro 4 high-content imaging system; and X. Li for help. Part of the analysis was performed on the High-Performance Computing Platform of the Center for Life Science (Peking University). We acknowledge funding from the National Natural Science Foundation of China (22137001 to P.R.C., 22337001 to C.Y., 92353000 to P.R.C. and 22425071 to C.Y.), the Ministry of Science and Technology (2023YFA1506500 and 2021YFA1302600 to P.R.C.; 2023YFC3402200 to C.Y.), Beijing Municipal Science and Technology Commission Project (Z231100002723005 to C.Y.), New Cornerstone Science Foundation (to C.Y.) through the XPLORER PRIZE, as well as New Cornerstone Science Foundation (to P.R.C.) through the New Cornerstone Investigator Program. This work was also supported by the Ministry of Agriculture and Rural Affairs of China.
Author information
Authors and Affiliations
Contributions
P.R.C., C.Y. and J.L. conceived the research. J.L. guided, designed and performed the experiments. X.Y. analysed the sequencing data. H. Wu analysed and organized the fluorescence data. Z.J. and Y.S. contributed in the screening, immunoprecipitation and proteomic experiments. X.W., R.G. and H. Wen conducted molecular dynamics simulation. Y.R. assisted the experiments in screening. C.L. performed acid-denaturing northern blots. Y.Z. and Z.J. guided the phosphoproteomic profiling. Y. M. conducted the tRNA sequencing and PRAISE sequencing experiments. All authors commented and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
C.Y. is an inventor on patents related to the RESTART technology (PCT/CN2022/095172). The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Kathrin Lang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The yields of ΨGA codons within targeted mRNAs under different dosage of gsnoRNA and PylRS-tRNAPyl.
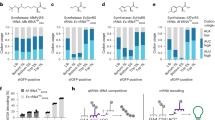
a-c, The IGV views and ΨGA codon yields within three targeted reporters: LDLR-TGA (a), AGXT-TGA (b), and Screen-TGA (c) at 0%, 25%, 50%, and 100% of the corresponding guide snoRNA. d, The IGV views and ΨGA codon yields within targeted Screen-TGA reporter under zero, 1:1, 1:4 or 1:8 stoichiometry to PylRS-tRNAPyl. Data of bar plots are presented as the mean values (n = 2 biologically independent replicates).
Extended Data Fig. 2 High encoding specificity of RCE strategy verified by the limited off-target Ψ edits as well as low off-target Ψ editing ratios across the transcriptome.
a, Complementary regions between gsnoRNA (black) and target site in SrcK295TGA-GFP reporter (blue). b, Complementary regions between gsnoRNA (black) and an off-target site in the endogenous gene (red). c, Sequence motif of the RCE-induced off-target Ψ sites. d, The on-target and off-target regions within the secondary structure of the gsnoRNA-Src predicted by RNAfold. The base-pair probabilities are indicated with color. Red and blue colors represent high and low probabilities, respectively. e,f IGV views representing the reads mapped at the off-target sites in 3’ UTRs of NM_001032283.3(e) and XM_017027173.2(f). g, Pie chart showing the distribution of RCE-induced off-target Ψ sites in the transcriptome. h, Scatterplot showing the expression level of genes containing the off-target Ψ sites by RNA-seq. Data are represented as the mean values (n = 2 biologically independent replicates). i, Scatterplot showing the translation level of genes containing the off-target Ψ sites by Ribo-seq. Data are represented as the mean values (n = 2 biologically independent replicates). The correlations between different conditions were measured via Pearson’s correlation score by defaulted parameters.
Extended Data Fig. 3 Evaluating the ncAA incorporation fidelity of the RCE system.
a, Schematic illustration of the workflow for identifying peptides with different amino acid incorporation via LC-tandem MS. b-g, The RCE system allowed site-specific incorporation of ncAAs with high fidelity on the Venus-TGA (b), Screen-TGA (d), and Src-K295*-Y527F-GFP (f) reporters. The corresponding Venus-R (c), Screen-R (e), and Src-K295R-Y527F-GFP (g) reporters were used for comparison of natural amino acid incorporation. h-j, Acid denaturing Northern Blots illustrating that the decoder tRNAs were highly charged with ncAA in all three RCE systems: RCE (ΨGA) (h), RCE (ΨAG) (i), RCE (ΨAA) (j). Each experiment was independently repeated twice with similar results. k, Diagram of the integrated stress response pathway. l, Western blotting demonstrating that the integrated stress response was not activated in the RCE systems. Tunicamycin treatment is used as a control of ISR activation. Each experiment was independently repeated twice with similar results. m, Scatterplot showing the translation level of ISR-related genes measured by Ribo-seq. Data are represented as the mean values (n = 2 biologically independent replicates).
Extended Data Fig. 4 MD simulation-based prediction supports the ΨGA preference of the screened mmtRNAPyl(UCA)-A37G variant.
a-c, Structural comparison among the ribosome-tRNA-mRNA complex from cryo-EM (PDB ID: 4JYA) (a), and all-atom MD simulation (b), along with the merged complex structures between cryo-EM and MD (c). The results show that the tRNA and mRNA align well between the structures from cryo-EM and MD simulation. d, The paired probability of the averaged trajectories demonstrating the binding between tRNA variants and UGA- or ΨGA-containing mRNAs. e-h, Centroids of the tRNA-mRNA paring structures of the wild-type mmtRNAPyl(UCA) (e,f) and the selected mmtRNAPyl(UCA)-A37G (g,h), illustrating that mmtRNAPyl(UCA)-37G forms more stable hydrogen bonds to ΨGA codon than to UGA codon.
Extended Data Fig. 5 The RCE system exhibited a high translational specificity when the reporter construct was expressed at an endogenous level.
a, Incorporated tetrazine-containing ncAA—TetBu could be labeled with TCO-biotin via bioorthogonal ligation. b, Western blotting of N-terminal HA-tagged products demonstrating that the protein product of the on-target reporter was expressed at endogenous concentration. RS/(ΨGA)-tRNATet alone was used to show the readthrough of tRNATet on the UGA codon of targeted reporter. The experiment was independently repeated twice with similar results. c,d, Western blotting (c) and quantitative analysis (d) showing the biotin signal for the GCE and RCE systems, demonstrating that the RCE system exhibited a low mis-incorporation level close to the background levels of endogenous biotinylated proteins. Data of bar plots are presented as the mean values (n = 2 biologically independent experiments). e,f, Western blotting (e) and quantitative analysis (f) comparing the off-target TetBu incorporation of the GCE(UGA), RS/(ΨGA)-tRNATet alone, and RCE(ΨGA) systems, without the targeted reporter, further demonstrating the ncAA incorporation specificity of the RCE system. Data of bar plots are presented as the mean values (n = 2 biologically independent experiments).
Extended Data Fig. 6 Proteome analysis demonstrated high translational specificity of the RCE(ΨGA) system.
a, Schematic illustration of the affinity-based enrichment method for detecting the biotinylated peptides. b,c, Volcano plots showing the ncAA-incorporated endogenous proteins induced by the RCE(ΨGA) system (b) and the GCE(UGA) system (c). Red dots represent ncAA-incorporated endogenous proteins. P-values were calculated by linear models and subsequently adjusted via the Benjamini-Hochberg method. X-axis is fold change of protein abundance in the GCE(UGA)- or RCE(ΨGA)-treated cells compared to the control cells.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Supplementary Discussion, legends for Supplementary Tables 1–7, Supplementary Figs. 1–24 and references.
Supplementary Tables (download ZIP )
Supplementary Tables 1–7; see Supplementary Information document for descriptions.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Yan, X., Wu, H. et al. RNA codon expansion via programmable pseudouridine editing and decoding. Nature 643, 1410–1420 (2025). https://doi.org/10.1038/s41586-025-09165-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09165-x
This article is cited by
-
Chemical biology of precise RNA targeting and intervention: advances and perspectives
Science China Chemistry (2026)
-
Functions and therapeutic applications of pseudouridylation
Nature Reviews Molecular Cell Biology (2025)