Abstract
Although the role of cellular immunity in checkpoint immunotherapy (CPI) for cancer is well established1,2, the effect of antibody-mediated humoral immunity is comparably underexplored. Here we used rapid extracellular antigen profiling3 to map the autoantibody reactome within a cohort of 374 patients with cancer treated with CPIs and 131 healthy control participants for autoantibodies to 6,172 extracellular and secreted proteins (the ‘exoproteome’). Globally, patients with cancer treated with CPIs had diverse autoreactivities that were elevated relative to control individuals but changed minimally with treatment. Autoantibody signatures in patients treated with CPI strikingly distinguished them from healthy individuals. Although associations of specific autoantibodies with immune-related adverse events were sparse, we detected numerous individual autoantibodies that were associated with greatly altered odds ratios for response to therapy. These included autoantibodies to immunomodulatory proteins, such as cytokines, growth factors and immunoreceptors, as well as tumour surface proteins. Functional evaluation of several autoantibody responses indicated that they neutralized the activity of their target proteins, which included type I interferons (IFN-I), IL-6, OSM, TL1A, and BMPR1A and BMPR2. Modelling the effects of autoantibodies to IFN-I and TL1A in preclinical mouse tumour models resulted in enhanced CPI efficacy, consistent with their effects in patients. In conclusion, these findings indicate that autoantibodies to the exoproteome modify CPI responses and highlight therapeutically actionable pathways that can be exploited to augment immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data used to generate figures and tables in this study are included in the Source Data. The single-cell RNA sequencing data generated from this study have been deposited in the Gene Expression Omnibus under the accession number GSE294482. The reference dataset used in Extended Data Fig. 8 is under the Gene Expression Omnibus accession number GSE199563. REAP data will be made available on reasonable request from the corresponding authors, subject to restrictions related to patient privacy in accordance with institutional policies and the Health Insurance Portability and Accountability Act. Source data are provided with this paper.
Code availability
The custom code for the analysis of REAP data is available on GitHub (https://github.com/RinglabCancerREAP/Cancer_REAP).
References
Sharma, P. et al. The next decade of immune checkpoint therapy. Cancer Discov. 11, 838–857 (2021).
Kim, T. K., Vandsemb, E. N., Herbst, R. S. & Chen, L. Adaptive immune resistance at the tumour site: mechanisms and therapeutic opportunities. Nat. Rev. Drug Discov. 21, 529–540 (2022).
Wang, E. Y. et al. High-throughput identification of autoantibodies that target the human exoproteome. Cell Rep. Methods 2, 100172 (2022).
Larkin, J. et al. Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 381, 1535–1546 (2019).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e20 (2018).
Meylan, M. et al. Tertiary lymphoid structures generate and propagate anti-tumor antibody-producing plasma cells in renal cell cancer. Immunity 55, 527–541.e5 (2022).
Mazor, R. D. et al. Tumor-reactive antibodies evolve from non-binding and autoreactive precursors. Cell 185, 1208–1222.e21 (2022).
Cabrita, R. et al. Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature 577, 561–565 (2020).
Helmink, B. A. et al. B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577, 549–555 (2020).
Petitprez, F. et al. B cells are associated with survival and immunotherapy response in sarcoma. Nature 577, 556–560 (2020).
Rivera, A., Chen, C. C., Ron, N., Dougherty, J. P. & Ron, Y. Role of B cells as antigen-presenting cells in vivo revisited: antigen-specific B cells are essential for T cell expansion in lymph nodes and for systemic T cell responses to low antigen concentrations. Int. Immunol. 13, 1583–1593 (2001).
Bruno, T. C. et al. Antigen-presenting intratumoral B cells affect CD4+ TIL phenotypes in non-small cell lung cancer patients. Cancer Immunol. Res. 5, 898–907 (2017).
Cui, C. et al. Neoantigen-driven B cell and CD4 T follicular helper cell collaboration promotes anti-tumor CD8 T cell responses. Cell 184, 6101–6118.e13 (2021).
Kotera, Y., Fontenot, J. D., Pecher, G., Metzgar, R. S. & Finn, O. J. Humoral immunity against a tandem repeat epitope of human mucin MUC-1 in sera from breast, pancreatic, and colon cancer patients. Cancer Res. 54, 2856–2860 (1994).
Disis, M. L. et al. High-titer HER-2/neu protein-specific antibody can be detected in patients with early-stage breast cancer. J. Clin. Oncol. 15, 3363–3367 (1997).
Ludwig, R. J. et al. Mechanisms of autoantibody-induced pathology. Front. Immunol. 8, 603 (2017).
Bastard, P. et al. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science 370, eabd4585 (2020).
Wang, E. Y. et al. Diverse functional autoantibodies in patients with COVID-19. Nature 595, 283–288 (2021).
Britschgi, M. et al. Neuroprotective natural antibodies to assemblies of amyloidogenic peptides decrease with normal aging and advancing Alzheimer’s disease. Proc. Natl Acad. Sci. USA 106, 12145–12150 (2009).
Jaycox, J. R., Dai, Y. & Ring, A. M. Decoding the autoantibody reactome. Science 383, 705–707 (2024).
Dimitrov, J. D. & Lacroix-Desmazes, S. Noncanonical functions of antibodies. Trends Immunol. 41, 379–393 (2020).
Wang, J. et al. Fibrinogen-like protein 1 is a major immune inhibitory ligand of LAG-3. Cell 176, 334–347.e12 (2019).
Yu, X. et al. The surface protein TIGIT suppresses T cell activation by promoting the generation of mature immunoregulatory dendritic cells. Nat. Immunol. 10, 48–57 (2009).
Jaiswal, S. et al. CD47 is up-regulated on circulating hematopoietic stem cells and leukemia cells to avoid phagocytosis. Cell 138, 271–285 (2009).
Hailemichael, Y. et al. Interleukin-6 blockade abrogates immunotherapy toxicity and promotes tumor immunity. Cancer Cell 40, 509–523.e6 (2022).
Benci, J. L. et al. Tumor interferon signaling regulates a multigenic resistance program to immune checkpoint blockade. Cell 167, 1540–1554.e12 (2016).
Vitiello, G. A. & Miller, G. Targeting the interleukin-17 immune axis for cancer immunotherapy. J. Exp. Med. 217, e20190456 (2020).
Huseni, M. A. et al. CD8+ T cell-intrinsic IL-6 signaling promotes resistance to anti-PD-L1 immunotherapy. Cell Rep. Med. 4, 100878 (2023).
Salazar, V. S., Gamer, L. W. & Rosen, V. BMP signalling in skeletal development, disease and repair. Nat. Rev. Endocrinol. 12, 203–221 (2016).
Thomas, C. et al. Structural linkage between ligand discrimination and receptor activation by type I interferons. Cell 146, 621–632 (2011).
van der Wijst, M. G. P. et al. Type I interferon autoantibodies are associated with systemic immune alterations in patients with COVID-19. Sci. Transl. Med. 13, eabh2624 (2021).
Hamilton, J. A. et al. Endogenous interferon-β regulates survival and development of transitional B cells. J. Immunol. 199, 2618–2623 (2017).
Antonelli, G., Scagnolari, C., Moschella, F. & Proietti, E. Twenty-five years of type I interferon-based treatment: a critical analysis of its therapeutic use. Cytokine Growth Factor Rev. 26, 121–131 (2015).
Benci, J. L. et al. Opposing functions of interferon coordinate adaptive and innate immune responses to cancer immune checkpoint blockade. Cell 178, 933–948.e14 (2019).
Reticker-Flynn, N. E. et al. Lymph node colonization induces tumor-immune tolerance to promote distant metastasis. Cell 185, 1924–1942.e23 (2022).
Mathew, D. et al. Combined JAK inhibition and PD-1 immunotherapy for non–small cell lung cancer patients. Science 384, eadf1329 (2024).
Zak, J. et al. JAK inhibition enhances checkpoint blockade immunotherapy in patients with Hodgkin lymphoma. Science 384, eade8520 (2024).
Giles, J. R. et al. Shared and distinct biological circuits in effector, memory and exhausted CD8+ T cells revealed by temporal single-cell transcriptomics and epigenetics. Nat. Immunol. 23, 1600–1613 (2022).
Zhu, J., Petit, P.-F. & Van den Eynde, B. J. Apoptosis of tumor-infiltrating T lymphocytes: a new immune checkpoint mechanism. Cancer Immunol. Immunother. 68, 835–847 (2019).
Wang, E. C. Y. et al. DR3 regulates negative selection during thymocyte development. Mol. Cell. Biol. 21, 3451–3461 (2001).
Wack, A., Terczyńska-Dyla, E. & Hartmann, R. Guarding the frontiers: the biology of type III interferons. Nat. Immunol. 16, 802–809 (2015).
Tang, Z. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Postow, M. A., Sidlow, R. & Hellmann, M. D. Immune-related adverse events associated with immune checkpoint blockade. N. Engl. J. Med. 378, 158–168 (2018).
de Moel, E. C. et al. Autoantibody development under treatment with immune-checkpoint inhibitors. Cancer Immunol. Res. 7, 6–11 (2019).
Weber, J. S. et al. Safety profile of nivolumab (NIVO) in patients (pts) with advanced melanoma (MEL): a pooled analysis. J. Clin. Oncol. 33, 9018–9018 (2015).
Jiang, Y. et al. Identification and characterization of a novel RF-amide peptide ligand for orphan G-protein-coupled receptor SP9155. J. Biol. Chem. 278, 27652–27657 (2003).
Cohen, J. C., Boerwinkle, E., Mosley, T. H. & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Furie, R. et al. Anifrolumab, an anti-interferon-α receptor monoclonal antibody, in moderate-to-severe systemic lupus erythematosus. Arthritis Rheumatol. 69, 376–386 (2017).
Bradford, H. F. et al. Inactive disease in patients with lupus is linked to autoantibodies to type I interferons that normalize blood IFNα and B cell subsets. Cell Rep. Med. 4, 100894 (2023).
Bastard, P. et al. Autoantibodies neutralizing type I IFNs are present in ~4% of uninfected individuals over 70 years old and account for ~20% of COVID-19 deaths. Sci. Immunol. 6, eabl4340 (2021).
Michot, J. M. et al. Immune-related adverse events with immune checkpoint blockade: a comprehensive review. Eur. J. Cancer 54, 139–148 (2016).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 46, 2699 (2018).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Migone, T. S. et al. TL1A is a TNF-like ligand for DR3 and TR6/DcR3 and functions as a T cell costimulator. Immunity 16, 479–492 (2002).
Qin, K. et al. Exposure of progressive immune dysfunction by SARS-CoV-2 mRNA vaccination in patients with chronic lymphocytic leukemia: a prospective cohort study. PLoS Med. 20, e1004157 (2023).
Zhou, T. et al. IL-18BP is a secreted immune checkpoint and barrier to IL-18 immunotherapy. Nature 583, 609–614 (2020).
Acknowledgements
We thank all members of the Ring and Kluger laboratories for technical assistance and helpful discussions. Extended Data Fig. 1 was created with Cognition Studio. Figures 1a, 2a, 3b,c,e and 4i and Extended Data Figs. 5a and 6a were created with BioRender (https://biorender.com). Y.D. was supported by a Yale Gruber Fellowship. J.R.J. was supported by the Yale Medical Scientist Training Program. A.M.R. was supported by grants from the Mark Foundation for Cancer Research and the Pew Charitable Trusts, and gifts from the Anderson and Bezos Families. K.C.H. and H.M.K. were supported by grants from the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Y.D., M.A., H.M.K. and A.M.R. designed the study. Y.D. and E.Y.W. established the REAP platform. Y.D. performed the REAP screens. K.C.H., H.M.K. and S.M.C. provided the clinical resources. L.A., M.A. and L.Z. collected the clinical samples, reviewed electronic health records and curated the clinical metadata. Y.D., J.C., H.v.A., Y.K. and L.F. performed the data analysis and data visualization. Y.D., K.Q., J.R.J. and S.F. performed the in vitro and in vivo validation experiments. H.M.K. and A.M.R. supervised the study. Y.D. and A.M.R. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A.M.R., E.Y.W. and Y.D. are inventors of a patent application assigned to Yale University describing the REAP technology (patent number: WO2021189053A1). In addition, A.M.R. is the founder and a director of Seranova Bio, the commercial licensee of REAP. S.M.C. and L.F. are employees of Seranova Bio. H.v.A. holds equity in Seranova Bio. L.A., K.Q., M.A., J.R.J., J.C., L.Z., S.F., Y.K., K.C.H. and H.M.K. declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Graphical abstract.
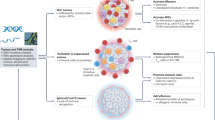
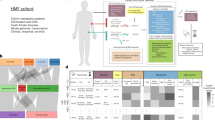
Plasma samples from CPI-treated patients and healthy donors were evaluated for autoantibodies against 6,172 distinct human extracellular antigens using REAP. REAP-derived autoantibody signatures were capable of distinguishing patients from healthy individuals and predicting response to CPI therapy. The impact of individual autoantibodies on clinical outcomes was assessed through calculation of odds ratios, highlighting treatment-enhancing and treatment-inhibiting autoantibodies. Autoantibodies of interest were biophysically validated and assessed for functional activity via ex vivo assays and preclinical models. The graphical abstract was created using Cognition Studio.
Extended Data Fig. 2 Autoantibody dynamics and associations with age and sex.
Related to Fig. 1. (a) Distribution of longitudinal sample collection over time. (b) REAP scores of PD-1 and CTLA4 from pre and post treatment plasma samples. PD−1 signal (n = 189, p = 1.51E-38). CTLA4 signal (n = 212, p = 3.58E-22). Significance was assessed by a paired two-sided Student’s t-test. (c) Overlay of distinct autoantibody reactivities from an individual patient. Red lines represent autoantibodies detected in the pre-treatment sample and grey lines represent autoantibodies detected post-treatment. (d) Scatter plot: Autoantibody REAP values in pre- and post-treatment. Each point represents a detected autoantibody. Density plot: post/pre REAP score differences for all detected reactivities. (e) For both healthy donors (controls) and cancer patients, there was a lack of correlation between age and the number of autoantibodies present, as assessed by a linear regression model (p = not significant). (f) Comparison of autoantibody numbers between males and females in both healthy donors and cancer patients (pre-treatment). Significance was assessed by a paired two-sided Student’s t-test. (g) Gender composition of healthy donors and cancer patients. (h) Linear regression models of autoantibody reactivity. Table shows coefficient estimates (standard errors) and p-values for three linear regression models predicting the number of autoantibody hits at different reactivity thresholds (>2, >4, and >6). Cancer Status coefficient represents the difference in number of autoantibodies between pre-treatment cancer patients and healthy controls adjusting for sex and log-transformed age.
Extended Data Fig. 3 Heatmap of REAP scores for immune-related proteins.
Immune-related proteins were categorized into specific groups as indicated in the figure and patients stratified according to response status.
Extended Data Fig. 4 ELISA validation.
ELISA validations for 11 additional REAP-detected autoantigens. The experiments were conducted in unicate with two technical replicates per sample. Data are presented as mean value with dot plots. Significance was assessed by a two-sided Student’s t-test.
Extended Data Fig. 5 Additional characterization of plasma and PBMC from patients with neutralizing autoantibodies against IFN-I.
(a) IFN-I signaling (STAT1 phosphorylation measured by flow cytometry) was assessed across a range of plasma dilutions for the six patients with broadly-reactive IFN-I autoantibodies for IFNA2, IFNA4, and IFNA8 as described in the Methods. Experiment was conducted in duplicate with two technical replicates. The schematic was created using BioRender (https://biorender.com). (b) Related to Fig. 3c,d. Flow cytometry gating strategies for monocytes, dendritic cells, B cells, natural killer cells, regulatory T cells, non-regulatory CD4 T cells and CD8 T cells. Antibodies used for staining were listed in the methods. (c) Percentage of cells in each cluster for IFN-I autoantibody positive patients compared to control patients. (d) Related to Fig. 3f,g. Kaplan–Meier survival curves for mice bearing B16F10 (top) or CT26 (bottom) tumors under different treatment conditions. Figures reflect pooled results from two independent experiments with numbers of animals per group indicated in the legend. Significance was assessed by the log-rank test. (e) Related to Fig. 3h,i. Tumor weight-normalized analyses of T cell subpopulations in different treatment groups. Data points are representative of individual tumors measured in unicate. Significance was assessed using a two-sided Student’s t-test. (f) Related to Fig. 3h,i. Flow cytometry gating strategies for CD8+PD1+ T cells. Antibodies used for staining were listed in the methods.
Extended Data Fig. 6 Single-cell RNA sequencing sample preparation.
(a) Study design: Mice grafted with MC38b were treated with either anti-mouse PD-1 alone or in combination with anti-mouse TL1A. The mice were euthanized 24 h following the second dose. Tumors were then harvested, digested into a single-cell suspension, sorted into targeted populations, and analyzed using 10x single-cell sequencing. The schematic was created using BioRender (https://biorender.com). (b) Flow cytometry gating strategies for sorting out CD45+CD3+ (TIL cells), CD45+CD3− (non-TIL immune cells), and CD45−CD3− (tumor and stromal cells).
Extended Data Fig. 7 Single-cell transcriptomic landscape of the TME in anti-PD1 and anti-TL1A treated tumors.
(a) t-SNE projection showing distribution of all cells colored by treatment condition (dark grey, PBS; orange, anti-PD1; blue, anti-PD1 and anti-TL1A). (b) t-SNE plot of all cells colored by expression of genes supporting cell-type assignments. (c, d, e) Bar plot showing frequency of CD4 T cells (c), NK cells (d), and myeloid cells (e) within clusters of interests in different treatment group. (f) Related to Fig. 4j,k. Flow cytometry gating strategies for CD8 T cells. Antibodies used for staining were listed in the methods.
Extended Data Fig. 8 CD8 T cells subclusters assignment strategy.
Giles et al. study38 on CD8+ T cells subtypes in chronic LCMV infection model was used as a reference to assign cell types in this study. Based on the gene expression pattern, Exh_Int, Exh_ISG and Exh_KLR are grouped as transitory exhausted T cell (Exh_Trans).
Extended Data Fig. 9 Comparison of odds ratios for autoantibodies by cancer type and cohort group.
(a, b) Comparison of odds ratios for anti-IFN-I, anti-IFN-III, and anti-BMPR1A/BMPR2 across different cancer types. Pan cancer (n = 374), melanoma (n = 148), NSCLC (n = 102), RCC (n = 58). Data are presented as odds ratio with 95% confidence intervals. (c) Analysis of TCGA transcript expression data for type-III IFN in multiple cancer types. Data are presented as mean value with dot plots. (d) Comparison of odds ratios for anti-IFN-I, and anti-BMPR1A/BMPR2 across different cohorts. Meta cohort (n = 374), Yale cohort (n = 249), MT cohort (n = 125). Data are presented as odds ratio with 95% confidence intervals.
Extended Data Fig. 10 Composition of irAEs across CPI response status.
Related to Fig. 5. This heatmap visualizes the category and severity of irAEs observed. Patients were grouped according to their CPI response status. Color indicates the severity of the specified irAE. Thyroiditis/hypothyroidism, hypophysitis, enteritis/colitis, pancreatitis, and hepatitis were the major observed irAEs in the cohort.
Supplementary information
Supplementary Table 1 (download DOCX )
Patient demographic
Supplementary Table 2 (download XLSX )
OR variable estimate
Supplemental Table 3 (download XLSX )
Frequencies of selected autoantibodies in current study and published cohorts
Supplemental Table 4 (download XLSX )
Summary of odds ratios comparing cancer patients to healthy individuals and CPI responders vs non-responders
Supplemental Table 5 (download XLSX )
Library antigen list
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dai, Y., Aizenbud, L., Qin, K. et al. Humoral determinants of checkpoint immunotherapy. Nature 644, 527–536 (2025). https://doi.org/10.1038/s41586-025-09188-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09188-4
This article is cited by
-
Beyond suppression: the paradox of JAK inhibitors as amplifiers of cancer immunotherapy
Nature Reviews Immunology (2026)
-
Heterocellular crosstalk and architecture of the pancreatic tumour microenvironment
Nature Reviews Cancer (2026)
-
Humoral IgG1 responses to tumor antigens underpin clinical outcomes in immune checkpoint blockade
Nature Medicine (2026)
-
Tertiary lymphoid structures in cancer: spatiotemporal heterogeneity, immune orchestration, and translational opportunities
Journal of Hematology & Oncology (2025)