Abstract
Most neurodevelopmental disorders with single gene diagnoses act via haploinsufficiency, in which only one of the two gene copies is functional1. SCN2A haploinsufficiency is one of the most frequent causes of neurodevelopmental disorder, often presenting with autism spectrum disorder, intellectual disability and, in a subset of children, refractory epilepsy2. Here, using SCN2A haploinsufficiency as a proof-of-concept, we show that upregulation of the existing functional gene copy through CRISPR activation (CRISPRa) can rescue neurological-associated phenotypes in Scn2a haploinsufficient mice. We first show that restoring Scn2a expression in adolescent heterozygous Scn2a conditional knock-in mice rescues electrophysiological deficits associated with Scn2a haploinsufficiency (Scn2a+/−). Next, using an adeno-associated virus CRISPRa-based treatment in adolescent mice, we show that we can correct intrinsic and synaptic deficits in neocortical pyramidal cells, a major cell type that contributes to neurodevelopmental disorders and seizure aetiology in SCN2A haploinsufficiency. Furthermore, we find that systemic delivery of CRISPRa protects Scn2a+/− mice against chemoconvulsant-induced seizures. Finally, we also show that adeno-associated virus CRISPRa treatment rescues excitability in SCN2A haploinsufficient human stem-cell-derived neurons. Our results showcase the potential of this therapeutic approach to rescue SCN2A haploinsufficiency and demonstrates that rescue even at adolescent stages can ameliorate neurodevelopmental phenotypes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All RNA-seq data are available at the NCBI Gene Expression Omnibus as Bioproject GSE302897. Requests for materials can be directed to N.A. (CRT-based approaches) or K.J.B. (mouse models). Source data are provided with this paper.
Code availability
No custom code was generated for this work.
Change history
23 February 2026
In the version of the article initially published, in the “CRISPRa in vitro optimization” section of the Methods, the text “rAAV vectors were generated using similar plasmids and cloning methods as described previously” originally cited ref. 1 but has now been corrected to ref. 17 in the HTML and PDF versions of the article.
08 December 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41586-025-09995-9
References
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Sanders, S. J. et al. Progress in understanding and treating SCN2A-mediated disorders. Trends Neurosci. https://doi.org/10.1016/j.tins.2018.03.011 (2018).
Maenner, M. J. et al. Prevalence of autism spectrum disorder among children aged 8 years – Autism and Developmental Disabilities Monitoring Network, 11 sites, United States, 2016. MMWR Surveill. Summ. https://doi.org/10.15585/MMWR.SS6904A1 (2020).
Kaplanis, J. et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 586, 757–762 (2020).
Fu, J. M. et al. Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nat. Genet. 54, 1320–1331 (2022).
Epi25 Collaborative Exome sequencing of 20,979 individuals with epilepsy reveals shared and distinct ultra-rare genetic risk across disorder subtypes. Nat. Neurosci. 27, 1864–1879 (2024).
Wang, D., Tai, P. W. L. & Gao, G. Adeno-associated virus vector as a platform for gene therapy delivery. Nat. Rev. Drug Discov. 18, 358–378 (2019).
Wu, Z., Yang, H. & Colosi, P. Effect of genome size on AAV vector packaging. Mol. Ther. 18, 80–86 (2010).
Howe, K. L. et al. Ensembl 2021. Nucleic Acids Res. 49, D884–D891 (2021).
Matharu, N. & Ahituv, N. Modulating gene regulation to treat genetic disorders. Nat. Rev. Drug Discov. 19, 757–775 (2020).
Werling, D. M. et al. Whole-genome and RNA sequencing reveal variation and transcriptomic coordination in the developing human prefrontal cortex. Cell Rep. 31, 107489 (2020).
Jenkins, P. M. & Bender, K. J. Axon initial segment structure and function in health and disease. Physiol. Rev. 105, 765–801 (2025).
Spratt, P. W. E. et al. The autism-associated gene Scn2a contributes to dendritic excitability and synaptic function in the prefrontal cortex. Neuron 103, 673–685 (2019).
Nelson, A. D. et al. Physical and functional convergence of the autism risk genes Scn2a and Ank2 in neocortical pyramidal cell dendrites. Neuron 112, 1133–1149 (2024).
Lu, C. et al. Overexpression of NEUROG2 and NEUROG1 in human embryonic stem cells produces a network of excitatory and inhibitory neurons. FASEB J. 33, 5287–5299 (2019).
Li, T. et al. Action potential initiation in neocortical inhibitory interneurons. PLoS Biol. https://doi.org/10.1371/journal.pbio.1001944 (2014).
Matharu, N. et al. CRISPR-mediated activation of a promoter or enhancer rescues obesity caused by haploinsufficiency. Science 363, eaau0629 (2019).
Grimm, D. et al. In vitro and in vivo gene therapy vector evolution via multispecies interbreeding and retargeting of adeno-associated viruses. J. Virol. 82, 5887–5911 (2008).
Bae, S., Park, J. & Kim, J.-S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Wang, W. et al. PTPN14 is required for the density-dependent control of YAP1. Genes Dev. 26, 1959–1971 (2012).
Correia, J. C. et al. Zfp697 is an RNA-binding protein that regulates skeletal muscle inflammation and remodeling. Proc. Natl Acad. Sci. USA 121, e2319724121 (2024).
Busse, D. C. et al. Interferon-induced protein 44 and interferon-induced protein 44-like restrict replication of respiratory syncytial virus. J. Virol. 94, e00297–20 (2020).
Baum, M. L. et al. CSMD1 regulates brain complement activity and circuit development. Brain Behav. Immun. 119, 317–332 (2024).
Spratt, P. W. E. et al. Paradoxical hyperexcitability from NaV1.2 sodium channel loss in neocortical pyramidal cells. Cell Rep. 36, 109483 (2021).
Chung, J. H., Larsen, A. R., Chen, E. & Bunz, F. A PTCH1 homolog transcriptionally activated by p53 suppresses hedgehog signaling. J. Biol. Chem. 289, 33020–33031 (2014).
Liang, L. et al. Developmental dynamics of voltage-gated sodium channel isoform expression in the human and mouse brain. Genome Med. 13, 135 (2021).
Yuan, Y. et al. Antisense oligonucleotides restore excitability, GABA signalling and sodium current density in a Dravet syndrome model. Brain 147, 1231–1246 (2024).
Zhang, J. et al. Severe deficiency of the voltage-gated sodium channel NaV1.2 elevates neuronal excitability in adult mice. Cell Rep. 36, 109495 (2021).
Miyamoto, H. et al. Impaired cortico-striatal excitatory transmission triggers epilepsy. Nat. Commun. 10, 1917 (2019).
Reynolds, C., King, M. D. & Gorman, K. M. The phenotypic spectrum of SCN2A-related epilepsy. Eur. J. Paediatr. Neurol. 24, 117–122 (2020).
Colasante, G. et al. dCas9-based Scn1a gene activation restores inhibitory interneuron excitability and attenuates seizures in Dravet syndrome mice. Mol. Ther. 28, 235–253 (2020).
Colasante, G. et al. In vivo CRISPRa decreases seizures and rescues cognitive deficits in a rodent model of epilepsy. Brain 143, 891–905 (2020).
Yamagata, T. et al. CRISPR/dCas9-based Scn1a gene activation in inhibitory neurons ameliorates epileptic and behavioral phenotypes of Dravet syndrome model mice. Neurobiol. Dis. 141, 104954 (2020).
Chang, H.-C. et al. rAAV-CRISPRa therapy corrects Rai1 haploinsufficiency and rescues selective disease features in Smith–Magenis syndrome mice. J. Biol. Chem. 299, 102728 (2023).
Wang, G. et al. Multiplexed activation of endogenous genes by CRISPRa elicits potent antitumor immunity. Nat. Immunol. 20, 1494–1505 (2019).
Liao, H.-K. et al. In vivo target gene activation via CRISPR/Cas9-mediated trans-epigenetic modulation. Cell 171, 1495–1507 (2017).
Böhm, S. et al. A gene therapy for inherited blindness using dCas9-VPR-mediated transcriptional activation. Sci. Adv. 6, eaba5614 (2020).
Kemaladewi, D. U. et al. A mutation-independent approach for muscular dystrophy via upregulation of a modifier gene. Nature 572, 125–130 (2019).
Mich, J. K. et al. Interneuron-specific dual-AAV SCN1A gene replacement corrects epileptic phenotypes in mouse models of Dravet syndrome. Sci. Transl. Med. 17, eadn5603 (2025).
Waszkielewicz, A. M. et al. Ion channels as drug targets in central nervous system disorders. Curr. Med. Chem. 20, 1241–1285 (2013).
Johnson, J. P. et al. NBI-921352, a first-in-class, NaV1.6 selective, sodium channel inhibitor that prevents seizures in Scn8a gain-of-function mice, and wild-type mice and rats. eLife 11, e72468 (2022).
Ferdosi, S. R. et al. Multifunctional CRISPR-Cas9 with engineered immunosilenced human T cell epitopes. Nat. Commun. 10, 1842 (2019).
Mehta, A. & Merkel, O. M. Immunogenicity of Cas9 protein. J. Pharm. Sci. 109, 62–67 (2020).
Gaj, T., Sirk, S. J., Shui, S.-L. & Liu, J. Genome-editing technologies: principles and applications. Cold Spring Harb. Perspect. Biol. 8, a023754 (2016).
Levy, G. & Barak, B. Postnatal therapeutic approaches in genetic neurodevelopmental disorders. Neural Regen. Res. 16, 414–422 (2021).
Markati, T., Duis, J. & Servais, L. Therapies in preclinical and clinical development for Angelman syndrome. Expert Opin. Investig. Drugs 30, 709–720 (2021).
Silva-Santos, S. et al. Ube3a reinstatement identifies distinct developmental windows in a murine Angelman syndrome model. J. Clin. Invest. 125, 2069–2076 (2015).
Wolter, J. M. et al. Cas9 gene therapy for Angelman syndrome traps Ube3a-ATS long non-coding RNA. Nature 587, 281–284 (2020).
Berg, A. T. et al. Expanded clinical phenotype spectrum correlates with variant function in SCN2A-related disorders. Brain 147, 2761–2774 (2024).
Eaton, M. et al. Generation and basic characterization of a gene-trap knockout mouse model of Scn2a with a substantial reduction of voltage-gated sodium channel Nav 1.2 expression. Genes Brain Behav. 20, e12725 (2021).
Tatsukawa, T. et al. Scn2a haploinsufficient mice display a spectrum of phenotypes affecting anxiety, sociability, memory flexibility and ampakine CX516 rescues their hyperactivity. Mol. Autism 10, 15 (2019).
Shin, W. et al. Scn2a haploinsufficiency in mice suppresses hippocampal neuronal excitability, excitatory synaptic drive, and long-term potentiation, and spatial learning and memory. Front. Mol. Neurosci. 12, 145 (2019).
Léna, I. & Mantegazza, M. NaV1.2 haploinsufficiency in Scn2a knock-out mice causes an autistic-like phenotype attenuated with age. Sci. Rep. 9, 12886 (2019).
Schamiloglu, S., Wu, H., Zhou, M., Kwan, A. C. & Bender, K. J. Dynamic foraging behavior performance is not affected by Scn2a haploinsufficiency. eNeuro 10, ENEURO.0367-23.2023 (2023).
Goertsen, D. et al. AAV capsid variants with brain-wide transgene expression and decreased liver targeting after intravenous delivery in mouse and marmoset. Nat. Neurosci. 25, 106–115 (2022).
Keiser, M. S. et al. Toxicity after AAV delivery of RNAi expression constructs into nonhuman primate brain. Nat. Med. 27, 1982–1989 (2021).
Hordeaux, J. et al. The GPI-linked protein LY6A drives AAV-PHP.B transport across the blood-brain barrier. Mol. Ther. 27, 912–921 (2019).
Yao, Y. et al. Variants of the adeno-associated virus serotype 9 with enhanced penetration of the blood-brain barrier in rodents and primates. Nat. Biomed. Eng. 6, 1257–1271 (2022).
Blesa, J. et al. BBB opening with focused ultrasound in nonhuman primates and Parkinson’s disease patients: targeted AAV vector delivery and PET imaging. Sci. Adv. 9, eadf4888 (2023).
Ewels, P. A. et al. The nf-core framework for community-curated bioinformatics pipelines. Nat. Biotechnol. 38, 276–278 (2020).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Fang, Z., Liu, X. & Peltz, G. GSEApy: a comprehensive package for performing gene set enrichment analysis in Python. Bioinformatics 39, btac757 (2023).
Planells-Cases, R. et al. Neuronal death and perinatal lethality in voltage-gated sodium channel αII-deficient mice. Biophys. J. 78, 2878–2891 (2000).
Van Erum, J., Van Dam, D. & De Deyn, P. P. PTZ-induced seizures in mice require a revised Racine scale. Epilepsy Behav. 95, 51–55 (2019).
Markram, H. et al. Reconstruction and simulation of neocortical microcircuitry. Cell 163, 456–492 (2015).
Ben-Shalom, R. et al. Opposing effects on NaV1.2 function underlie differences between SCN2A variants observed in individuals with autism spectrum disorder or infantile seizures. Biol. Psychiatry 82, 224–232 (2017).
Ruden, J. B., Dixit, M., Zepeda, J. C., Grueter, B. A. & Dugan, L. L. Robust expression of functional NMDA receptors in human induced pluripotent stem cell-derived neuronal cultures using an accelerated protocol. Front. Mol. Neurosci. 14, 777049 (2021).
Acknowledgements
We thank the Bender and Ahituv laboratory members for comments and discussions on this manuscript, and members of the FamilieSCN2A Foundation who provided the core motivation and inspiration for this work. We acknowledge the UCSF Parnassus Flow CoLab (RRID:SCR_018206) supported in part by NIH grant no. P30 DK063720 and by the NIH S10 Instrumentation Grant no. S10 1S10OD021822-01. This work was supported by grants from SFARI (grant no. 629287: K.J.B., N.A.; grant no. 513133: K.J.B.), the Broad Institute Target Practice Initiative (K.J.B.), the Autism Science Foundation (S.T.), the Weill Neurohub Investigator Program (K.J.B., N.A.), an NSERC Predoctoral Fellowship (P.W.E.S.), the Ford Foundation Dissertation Fellowship (S.S.H.), the Weill Foundation Graduate Student Fellowship (S.S.H.), an NSF Graduate Fellowship (S.T.) and the NIH (grant no. R01 MH125978: K.J.B.; grant no. R01 MH136475: K.J.B., N.A.; grant nos. F32 MH125536 and K99 MH135209: A.D.N.; grant nos. R01 NS078118 and R01 NS121287: J.T.P.; grant no. R01 MH115045, R01 NS108874 and R01 MH118298: J.Q.P.; grant no. T32 GM007449: S.S.H.; grant no. MH126960: P.M.J.; grant no. R01 MH129751: S.J.S.), EMBO (ALTF 585-2021: C.A.), the Medical Research Council Centre of Research Excellence in Therapeutic Genomics (MR/Z504725/1: S.J.S., N.A.), the UCSF Bakar Aging Research Institute postdoctoral fellowship (C.A.) and the Bettencourt Schueller foundation (C.A.).
Author information
Authors and Affiliations
Contributions
Conceptualization: K.J.B., S.J.S., N.A. Methodology: S.T., A.D.N., P.W.E.S., H.K., J.Z., N.M., J.Q.P., J.T.P., P.M.J., R.B.-S., N.A., K.J.B. Software: H.K., J.Z., R.B.-S. Formal analysis: S.T., A.D.N., P.W.E.S., V.B., B.K., L.M., P.M.J., H.K., J.Z., K.J.B. Investigation: S.T., A.D.N., P.W.E.S., E.C.H., X.Z., H.K., Z.L., C.A., V.B., K.Y., J.Z., S.S.H., A.S., C.M.K., C.L., S.E.T., S.S., Y.C.L., K.J.B. Writing—original draft: S.T., A.D.N., N.A., K.J.B. Writing—review and editing: all authors. Visualization: S.T., A.D.N., P.W.E.S., H.K., J.Z., K.J.B. Supervision: N.A., K.J.B. Funding acquisition: S.T., A.D.N., P.W.E.S., S.S.H., J.Q.P., J.T.P., S.J.S., N.A., K.J.B.
Corresponding authors
Ethics declarations
Competing interests
N.M. is the cofounder and former board member and CSO of Regel Therapeutics, N.A. is the cofounder of Regel Therapeutics and both N.A. and K.J.B. are on the scientific advisory board of Regel Therapeutics. P.W.E.S. is a Program Director at Regel Therapeutics. N.M. and N.A. are the inventors on patent ‘Gene therapy for haploinsufficiency’ WO2018148256A9. N.A., K.J.B. and S.J.S. receive funding from BioMarin Pharmaceutical Incorporated. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Elvir Becirovic, James Trimmer and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Excitatory pyramidal neurons in the mPFC are GFP+ in Cre-negative Scn2a+/KI animals.
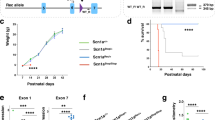
a. Coronal brain sections from P60 Scn2a+/KI mouse (Cre-) immunostained with anti-GFP and anti-parvalbumin (PV). b. Zoom of area highlighted by dashed box in panel a, with GFP and Parvalbumin channels separated then merged at bottom. Parv+ somata are circled in the GFP panel. c. Further zoom of region in layer 5b in panel b. Parv+ somata are circled as in panel c. d. Quantification of mean fluorescence intensify of GFP in PV-negative cells, PV-positive cells, and neuropil (area without somata as a measure of background fluorescence). Data are from 2 mice. Circles are mean GFP intensity values in ROIs of Parv- somata, Parv+ somata, or neighboring neuropil in single optical sections. Box plots are medians, quartiles, and 100% tails. n = 67 Parv- and 37 Parv+ cells analyzed; Parv- vs. Parv + : ****p < 0.0001. Parv- vs. neuropil: ****p < 0.0001. Holm-Šídák multiple comparisons test.
Extended Data Fig. 2 In vitro optimization of CRISPRa constructs in mouse Neuroblastoma-2A cells.
a. Fold change of Scn2a expression in Neuro-2a cells transfected with plasmids containing sgRNAs targeting the promoter of mouse Scn2a compared to a no-sgRNA VP64 control. Blue bars represent plasmids with largest increase in Scn2a expression. Circles are replicates, overlaid on mean ± SD. b. Fold change of Scn2a transduced with rAAV-DJ virus in Neuro-2a cells. Circles are replicates, overlaid on mean ± SD. c. Total animal weight at time of 8 mg/kg 4-AP administration (animals aged 69–119 days). Circles are animals. Box plots are medians, quartiles and 100% tails. d. RT-qPCR analysis of dCas9 and mCherry mRNA within the mPFC of tail vein injected Scn2a+/+ + CRISPRa (light gray) and Scn2a+/− + CRISPRa (purple) versus uninjected controls Scn2a+/+ (dark gray) or Scn2a+/− (cyan). Injected animals with at least a 10-fold increase in expression levels of both dCas9 and mCherry to the average Scn2a+/+ uninjected controls were included in EEG datasets in Fig. 3.
Extended Data Fig. 3 Scn2a-CRISPRa off-target analysis in cell lines.
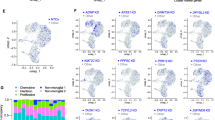
a. RNA-seq expression levels of Scn2a TAD-domain genes from Scn2a-rAAV-CRISPRa treated Neuro-2a cells compared to VP64-only. Circles are individual replicates, box plots are medians, quartiles and 100% tails. p-values noted above data. Wald test. b. RNA-seq expression levels of sodium channels from Scn2a-rAAV-CRISPRa treated Neuro-2a cells compared to VP64-only. Circles are individual replicates, box plots are medians, quartiles and 100% tails. p-values noted above data. Wald test. c. Volcano plot representing Log2 fold-change in expression levels for each gene. Wald test. P-values noted above each comparison. Grey dots represent no significant DEGs for CRISPRa, purple dots signify Scn2a and nearby genes, and orange dots denote upregulated genes. d. Gene ontology (GO) analysis showing enrichment of molecular functions. e. Gene ontology (GO) analysis showing enrichment of biological processes. f. Analysis of sgRNA sequence off-targeting using Cas-OFFinder.
Extended Data Fig. 4 Intrinsic electrophysiology of Scn2a+/KI and CRISPRa treated neurons.
a. AP threshold from P57-85 Scn2a-rAAV-empty vector Scn2a+/+ (light gray) and Scn2a+/− (magenta) neurons, Scn2a-rAAV-CRISPRa treated Scn2a+/+ (dark gray) and Scn2a+/− (purple) neurons, and Scn2a+/KI Cre- (green) and Scn2a+/KI Cre+ (gray) neurons. Threshold of the first AP evoked by a near-rheobase current. Scn2a+/+ + empty: n = 17 cells; Scn2a+/− + empty: n = 27 cells; Scn2a+/+ + CRISPRa: n = 24 cells; Scn2a+/− + CRISPRa: n = 19 cells; Scn2a+/KI Cre-: n = 29 cells; Scn2a+/KI Cre + : n = 21 cells. No significant differences. Holm-Šídák multiple comparisons test. Circles are individual cells, box plots are medians, quartiles and 100% tails. b. Input resistance (MΩ). Scn2a+/+ + empty: n = 17 cells; Scn2a+/− + empty: n = 27 cells; Scn2a+/+ + CRISPRa: n = 17 cells; Scn2a+/− + CRISPRa: n = 11 cells; Scn2a+/KI Cre-: n = 26 cells; Scn2a+/KI Cre + : n = 19 cells. No significant differences. Holm-Šídák multiple comparisons test. Circles are individual cells, box plots are medians, quartiles and 100% tails. c. Rheobase current (pA) to generate first spike. Scn2a+/+ + empty: n = 16 cells; Scn2a+/− + empty: n = 27 cells; Scn2a+/+ + CRISPRa: n = 17 cells; Scn2a+/− + CRISPRa: n = 12 cells; Scn2a+/KI Cre-: n = 25 cells; Scn2a+/KI Cre + : n = 19 cells. No significant differences. Holm-Šídák multiple comparisons test. Circles are individual cells, box plots are medians, quartiles and 100% tails. d. APs per 300 ms stimulation epoch for each current amplitude. Left: Quantification of firing rate slope of data on Right. Scn2a+/+ + empty: n = 16 cells; Scn2a+/− + empty: n = 27 cells; Scn2a+/+ + CRISPRa: n = 17 cells; Scn2a+/− + CRISPRa: n = 12 cells; Scn2a+/KI Cre-: n = 25 cells; Scn2a+/KI Cre + : n = 19 cells. No significant differences. Holm-Šídák multiple comparisons test. Circles are individual cells, box plots are medians, quartiles and 100% tails.Right: Number of APs versus current amplitude injected. Circles and bars are mean ± SEM at each stimulus intensity.
Extended Data Fig. 5 Scn2a-CRISPRa off-target analysis in the mouse neocortex.
a. RNA-seq expression levels of Scn2a TAD-domain genes from Scn2a-rAAV-CRISPRa treated Neuro-2a cells compared to VP64-only. Circles are individual mice, box plots are medians, quartiles and 100% tails. b. RNA-seq expression levels of sodium channels from Scn2a-rAAV-CRISPRa treated Neuro-2a cells compared to VP64-only. Circles are individual mice, box plots are medians, quartiles and 100% tails. c. Volcano plot representing Log2 fold-change in expression levels for each gene for WT vs Scn2a+/− “Het”. Wald test. P-values noted above each comparison. Grey dots represent no significant DEGs for CRISPRa, purple dots signify Scn2a and nearby genes, and orange dots denote upregulated genes. d. Same as c, but for CRISPRa-treated Scn2a+/− “CRISPRa” vs WT. e. Same as c, but for CRISPRa-treated Scn2a+/− “CRISPRa” vs Scn2a+/− “Het”. f. Analysis of sgRNA sequence off-targeting using Cas-OFFinder. g. Data representation of NeuN-positive neuronal nuclei isolated from cortical tissue for FACS sorting. Representative FACS plots visualized with FlowJo V10 with percentage of parent gates for each population. Singlets were selected based on FSC-H versus FSC-A. h. From singlets, DAPI-positive events were gated to identify nuclei. i. NeuN-positive neuronal nuclei were selected using an anti-NeuN antibody conjugated to Alexa Fluor 488.
Extended Data Fig. 6 CRISPRa expression persists through 16 months post-systemic injection.
Peak AP dV/dt (top) at 30-, 56-, 107-, and 210-days following tail-vein injection of Scn2a-rAAV-CRISPRa-PHP.eB in Scn2a+/−mice (purple). Circles represent individual neurons. Lines are linear regression. qPCR of mCherry (middle) or dCas9 (bottom) mRNA from neocortical samples collected from tail-vein or retro-orbital Scn2a-rAAV-CRISPRa-PHP.eB injected Scn2a+/+ (gray) or Scn2a+/− mice (purple) across time. Circles represent single mice. Lines are linear regression.
Extended Data Fig. 7 Behavioral and EEG response to increasing doses of 4-AP in Scn2a+/− mice.
a. Example PFC EEGs from animals receiving 8 and 15 mg/kg 4-AP. 4-AP administered at onset of all recordings. Dashed box denotes a tonic clonic seizure and subsequent mortality occurring with 15 mg/kg dosing in Scn2a+/− animal. EEG data within boxed area is shown at higher resolution on right. b. Survival curves for all WT and Scn2a+/− animals for 6, 8 and 15 mg/kg 4-AP. WT: black, Scn2a+/−: cyan.
Extended Data Fig. 8 In vitro optimization of CRISPRa constructs in human SH-SY5Ycells.
a. Fold change of SCN2A expression in SH-SY5Y cells transfected with plasmids containing sgRNAs targeting the promoter of human SCN2A compared to a no sgRNA VP64 control. b. Fold change of SCN2A transduced with rAAV-DJ virus in human SH-SY5Y cells. c. SCN2A mRNA expression from SCN2A+/+ (black), SCN2A+/− (cyan), and SCN2A-rAAV-CRISPRa treated SCN2A+/− (purple) hESC-derived neurons normalized to wild type average. SCN2A+/+ vs. SCN2A+/−: *p = 0.03. Holm-Šídák multiple comparisons test. n = 3 replicates for all conditions. circles are individual replicates. Bars are mean ± SD.
Extended Data Fig. 9 Axon initial segment structural plasticity occurs in SCN2A+/− neurons and is rescued by CRISPRa.
a. Representative images of SCN2A+/+ (black) and SCN2A+/− (cyan) human stem-cell-derived neurons immunostained with antibodies against ankyrin-G (green) and MAP2 (magenta). Arrowheads denote start and end points used to quantify AIS length. b. Quantification of AIS length. SCN2A+/+: n = 56 cells, 3 dishes. SCN2A+/−: n = 56 cells, 3 dishes. ****p < 0.0001. Mann-Whitney test. c. Representative images of SCN2A+/− neurons expressing Scn2a-rAAV-CRISPRa-mCherry (purple) and mCherry-negative internal SCN2A+/− controls (cyan). Immunostaining against ankyrin-G and MAP2. d. Quantification of AIS length. SCN2A+/−: n = 122 cells, 3 dishes. SCN2A+/− + CRISPRa: n = 23 cells, 3 dishes. ****p < 0.0001. Mann-Whitney test. Circles are individual AIS calculations. Box plots are medians, quartiles, and 100% tails.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tamura, S., Nelson, A.D., Spratt, P.W.E. et al. CRISPR activation for SCN2A-related neurodevelopmental disorders. Nature 646, 983–991 (2025). https://doi.org/10.1038/s41586-025-09522-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09522-w
This article is cited by
-
Effect of Alzheimer’s disease medications on neurocognitive outcomes in children and adolescents with autism spectrum disorder and low IQ: a scoping review
Translational Psychiatry (2025)
-
CRISPR activation tackles neurodevelopmental disorders
Nature Reviews Drug Discovery (2025)