Abstract
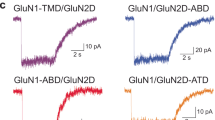
Ion-channel activity reflects a combination of open probability and unitary conductance1. Many channels display subconductance states that modulate signalling strength2,3, yet the structural mechanisms governing conductance levels remain incompletely understood. Here we report that conductance levels are controlled by the bending patterns of pore-forming transmembrane helices in the heterotetrameric neuronal channel GluN1a-2B N-methyl-D-aspartate receptor (NMDAR). Our single-particle electron cryomicroscopy (cryo-EM) analyses demonstrate that an endogenous neurosteroid and synthetic positive allosteric modulator (PAM), 24S-hydroxycholesterol (24S-HC), binds to a juxtamembrane pocket in the GluN2B subunit and stabilizes the fully open-gate conformation, where GluN1a M3 and GluN2B M3′ pore-forming helices are bent to dilate the channel pore. By contrast, EU1622-240 binds to the same GluN2B juxtamembrane pocket and a distinct juxtamembrane pocket in GluN1a to stabilize a sub-open state whereby only the GluN2B M3′ helix is bent. Consistent with the varying extents of gate opening, the single-channel recordings predominantly show full-conductance and subconductance states in the presence of 24S-HC and EU1622-240, respectively. Another class of neurosteroid, pregnenolone sulfate, engages a similar GluN2B pocket, but two molecules bind simultaneously, revealing a diverse neurosteroid recognition pattern. Our study identifies that the juxtamembrane pockets are critical structural hubs for modulating conductance levels in NMDAR.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Cryo-EM data have been deposited to the PDB and Electron Microscopy Data Bank (EMDB): non-active-state Gly, Glu, PS-bound hGluN1a-2B NMDAR (PDB: 9OOS; EMDB: EMD-70671), pre-active-state Gly, Glu, PS-bound hGluN1a-2B NMDAR (PDB: 9OOT; EMDB: EMD-70672), closed-state Gly, Glu, 24S-HC-bound hGluN1a-2B NMDAR (PDB: 9OOQ; EMDB: EMD-70669), open-state Gly, Glu, 24S-HC-bound hGluN1a-2B NMDAR (PDB: 9OOR; EMDB: EMD-70671) and sub-open-state Gly, Glu, EU1622-240-bound rGluN1a-2B NMDAR (PDB: 9OOU; EMDB: EMD-70673). Source data are provided with this paper.
References
Hille, B. Ion Channels of Excitable Membranes 3rd edn (Sinauer, 2001).
Jahr, C. E. & Stevens, C. F. Glutamate activates multiple single channel conductances in hippocampal neurons. Nature 325, 522–525 (1987).
Cull-Candy, S. G. & Usowicz, M. M. Multiple-conductance channels activated by excitatory amino acids in cerebellar neurons. Nature 325, 525–528 (1987).
Schneggenburger, R. & Ascher, P. Coupling of permeation and gating in an NMDA-channel pore mutant. Neuron 18, 167–177 (1997).
Banke, T. G. & Traynelis, S. F. Activation of NR1/NR2B NMDA receptors. Nat. Neurosci. 6, 144–152 (2003).
Popescu, G. & Auerbach, A. Modal gating of NMDA receptors and the shape of their synaptic response. Nat. Neurosci. 6, 476–483 (2003).
Bliss, T. V. P. & Collingridge, G. L. A synaptic model of memory: long-term potentiation in the hippocampus. Nature 361, 31–39 (1993).
Malenka, R. C. & Bear, M. F. LTP and LTD: an embarrassment of riches. Neuron 44, 5–21 (2004).
Hansen, K. B. et al. Structure, function, and pharmacology of glutamate receptor ion channels. Pharmacol. Rev. 73, 298–487 (2021).
Karakas, E. & Furukawa, H. Crystal structure of a heterotetrameric NMDA receptor ion channel. Science 344, 992–997 (2014).
Lee, C. H. et al. NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 511, 191–197 (2014).
Wang, J. X. & Furukawa, H. Dissecting diverse functions of NMDA receptors by structural biology. Curr. Opin. Struct. Biol. 54, 34–42 (2019).
Mony, L. & Paoletti, P. Mechanisms of NMDA receptor regulation. Curr. Opin. Neurobiol. 83, 102815 (2023).
Zhou, C. & Tajima, N. Structural insights into NMDA receptor pharmacology. Biochem. Soc. Trans. 51, 1713–1731 (2023).
Wu, E., Zhang, J., Zhang, J. & Zhu, S. Structural insights into gating mechanism and allosteric regulation of NMDA receptors. Curr. Opin. Neurobiol. 83, 102806 (2023).
Hansen, K. B. et al. Structure, function, and allosteric modulation of NMDA receptors. J. Gen. Physiol. 150, 1081–1105 (2018).
Ratner, M. H., Kumaresan, V. & Farb, D. H. Neurosteroid actions in memory and neurologic/neuropsychiatric disorders. Front. Endocrinol. 10, 169 (2019).
Hanson, J. E. et al. Therapeutic potential of N-methyl-D-aspartate receptor modulators in psychiatry. Neuropsychopharmacology 49, 51–66 (2024).
Zorumski, C. F. et al. New directions in neurosteroid therapeutics in neuropsychiatry. Neurosci. Biobehav. Rev. 172, 106119 (2025).
Hrcka Krausova, B. et al. Site of action of brain neurosteroid pregnenolone sulfate at the N-methyl-D-aspartate receptor. J. Neurosci. 40, 5922 (2020).
Perszyk, R. E. et al. Biased modulators of NMDA receptors control channel opening and ion selectivity. Nat. Chem. Biol. 16, 188–196 (2020).
Chou, T. H. et al. Molecular mechanism of ligand gating and opening of NMDA receptor. Nature 632, 209–217 (2024).
Ullman, E. Z. et al. Mechanisms of action underlying conductance-modifying positive allosteric modulators of the NMDA receptor. Mol. Pharmacol. 106, 334–353 (2024).
Paul, S. M. et al. The major brain cholesterol metabolite 24(S)-hydroxycholesterol is a potent allosteric modulator of N-methyl-D-aspartate receptors. J. Neurosci. 33, 17290–17300 (2013).
Wu, F. S., Gibbs, T. T. & Farb, D. H. Pregnenolone sulfate: a positive allosteric modulator at the N-methyl-D-aspartate receptor. Mol. Pharmacol. 40, 333–336 (1991).
Fritzemeier, R. G. et al. Thienopyrimidinone derivatives as a GluN2B/C/D biased, positive allosteric modulator of the N-methyl-d-aspartate receptor. J. Med. Chem. 68, 9303–9322 (2025).
Premkumar, L. S., Qin, F. & Auerbach, A. Subconductance States of a mutant NMDA receptor channel kinetics, calcium, and voltage dependence. J. Gen. Physiol. 109, 181–189 (1997).
Stern, P., Béhé, P., Schoepfer, R. & Colquhoun, D. Single-channel conductances of NMDA receptors expressed from cloned cDNAs: comparison with native receptors. Proc. R. Soc. Lond. B 250, 271–277 (1997).
Banke, T. G., Dravid, S. M. & Traynelis, S. F. Protons trap NR1/NR2B NMDA receptors in a nonconducting state. J. Neurosci. 25, 42–51 (2005).
Huang, Z. & Gibb, A. J. Mg2+ block properties of triheteromeric GluN1–GluN2B–GluN2D NMDA receptors on neonatal rat substantia nigra pars compacta dopaminergic neurones. J. Physiol. 592, 2059–2078 (2014).
Kang, H. et al. Structural basis for channel gating and blockade in tri-heteromeric GluN1-2B-2D NMDA receptor. Neuron https://doi.org/10.1016/j.neuron.2025.01.013 (2025).
Chou, T. H., Tajima, N., Romero-Hernandez, A. & Furukawa, H. Structural basis of functional transitions in mammalian NMDA receptors. Cell 182, 357–371 (2020).
Tajima, N. et al. Activation of NMDA receptors and the mechanism of inhibition by ifenprodil. Nature 534, 63–68 (2016).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. P. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graphics 14, 354–360 (1996).
Amin, J. B. et al. Two gates mediate NMDA receptor activity and are under subunit-specific regulation. Nat. Commun. 14, 1623 (2023).
Twomey, E. C., Yelshanskaya, M. V., Grassucci, R. A., Frank, J. & Sobolevsky, A. I. Channel opening and gating mechanism in AMPA-subtype glutamate receptors. Nature 549, 60–65 (2017).
Gangwar, S. P. et al. Kainate receptor channel opening and gating mechanism. Nature 630, 762–768 (2024).
Swanson, G. T., Kamboj, S. K. & Cull-Candy, S. G. Single-channel properties of recombinant AMPA receptors depend on RNA editing, splice variation, and subunit composition. J. Neurosci. 17, 58 (1997).
Zhang, W. et al. A transmembrane accessory subunit that modulates kainate-type glutamate receptors. Neuron 61, 385–396 (2009).
Watanabe, J., Beck, C., Kuner, T., Premkumar, L. S. & Wollmuth, L. P. DRPEER: a motif in the extracellular vestibule conferring high Ca2+ flux rates in NMDA receptor channels. J. Neurosci. 22, 10209–10216 (2002).
Perszyk, R. E. et al. Hodgkin-Huxley-Katz Prize Lecture: genetic and pharmacological control of glutamate receptor channel through a highly conserved gating motif. J. Physiol. https://doi.org/10.1113/JP278086 (2020).
Rosenmund, C., Stern-Bach, Y. & Stevens, C. F. The tetrameric structure of a glutamate receptor channel. Science 280, 1596–1599 (1998).
Yelshanskaya, M. V., Patel, D. S., Kottke, C. M., Kurnikova, M. G. & Sobolevsky, A. I. Opening of glutamate receptor channel to subconductance levels. Nature 605, 172–178 (2022).
Coombs, I. D. & Cull-Candy, S. G. Single-channel mechanisms underlying the function, diversity and plasticity of AMPA receptors. Neuropharmacology 198, 108781 (2021).
Benveniste, M. & Mayer, M. L. Kinetic analysis of antagonist action at N-methyl-D-aspartic acid receptors. Two binding sites each for glutamate and glycine. Biophys. J. 59, 560–573 (1991).
Hale, W. D., Huganir, R. L. & Twomey, E. C. Architecture, activation, and conformational plasticity in the GluA4 AMPA receptor. Preprint at bioRxiv https://doi.org/10.1101/2025.06.12.659357 (2025).
Furukawa, H., Simorowski, N. & Michalski, K. Effective production of oligomeric membrane proteins by EarlyBac-insect cell system. Methods Enzymol. 653, 3–19 (2021).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rubinstein, J. L. & Brubaker, M. A. Alignment of cryo-EM movies of individual particles by optimization of image translations. J. Struct. Biol. 192, 188–195 (2015).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Chou, T. H. et al. Structural insights into binding of therapeutic channel blockers in NMDA receptors. Nat. Struct. Mol. Biol. 29, 507–518 (2022).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Lewis, C. A. Ion-concentration dependence of the reversal potential and the single channel conductance of ion channels at the frog neuromuscular junction. J. Physiol. 286, 417–445 (1979).
Jatzke, C., Hernandez, M. & Wollmuth, L. P. Extracellular vestibule determinants of Ca2+ influx in Ca2+-permeable AMPA receptor channels. J. Physiol. 549, 439–452 (2003).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc.Bioinform. 2016, 5.6.1–5.6.37 (2016).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78, 1950–1958 (2010).
Joung, I. S. & Cheatham, T. E. III Determination of alkali and halide monovalent ion parameters for use in explicitly solvated biomolecular simulations. J. Phys. Chem. B 112, 9020–9041 (2008).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Nosé, S. A unified formulation of the constant temperature molecular dynamics methods. J. Chem. Phys. 81, 511–519 (1984).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Acknowledgements
We thank N. Simorowski and J. Zhang for technical support; R. Perszyk for helpful discussions; and D. Thomas and M. Wang for managing the cryo-EM facility and computing facility at Cold Spring Harbor Laboratory, respectively. This work was funded by the NIH (NS111745 and MH085926 to H.F., NS111619 to S.F.T.), Austin’s purpose (H.F. and S.F.T.), Robertson funds at CSHL, Doug Fox Alzheimer’s fund, Heartfelt Wing Alzheimer’s fund and the Gertrude and Louis Feil Family Trust (all to H.F.). The computational work was performed with assistance from an NIH grant (S10OD028632-01).
Author information
Authors and Affiliations
Contributions
H.K. and H.F. conceived the project. H.K. and R.S. obtained cryo-EM structures. H.K. conducted TEVC electrophysiology experiments. E.Z.U. and S.F.T. conducted single-channel recordings. M.E. conducted molecular dynamics simulations. S.P. and D.C.L. synthesized EU1622-240. H.K. and H.F. wrote the manuscript with input from all of the authors.
Corresponding author
Ethics declarations
Competing interests
S.F.T. and H.F. are members of the medical advisory boards for the CureGRIN Foundation. S.F.T. is an advisory board member for the GRIN2B Foundation; a member of the scientific advisory boards for Eumentis Therapeutics and Neurocrine; a consultant for GRIN Therapeutics and Seyltx; a co-founder of NeurOp and AgriThera; and is on the Board of Directors for NeurOp. D.C.L. is on the Board of Directors for NeurOp. Several authors (S.P., S.F.T. and D.C.L.) are co-inventors on Emory-owned IP involving NMDA receptor modulators. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM workflow for PS-bound GluN1a-2B NMDAR structures.
a, Single-particle cryo-EM analysis of the glycine, glutamate, and PS-bound GluN1a-2B NMDAR. In this dataset, non-active and pre-active states were identified. Data processing was performed on sets of movies listed in Supplementary Table 2. The white bar in the micrograph indicates 20 nm. b-c, Map quality assessment of non-active state (b) and pre-active state (c).
Extended Data Fig. 2 Cryo-EM workflow for 24S-HC-bound GluN1a-2B NMDAR structures.
a, Single-particle cryo-EM analysis of the glycine, glutamate, and 24S-HC-bound GluN1a-2B NMDAR. In this dataset, the closed and open states were identified. Data processing was performed on sets of movies listed in Supplementary Table 2. The white bar in the micrograph indicates 20 nm. b-c, Map quality assessment of closed state (b) and open state (c).
Extended Data Fig. 3 Additive potentiating effects of PS and 24S-HC.
a, Superposition of the PS (light orange) and 24S-HC (yellow) binding sites demonstrating distinct binding modes. At Site-1, the hydroxyl group at the tip of 24S-HC and the sulfate group of PS reside in a similar area surrounded by GluN2B pre-M1’ and M1’. 24S-HC binds parallel to GluN2B M1’ helix, while PS wedges into the GluN2B M2’ and M3’ helices. At Site-2, the ketone group of PS overlaps with the phospholipid acyl group (grey). b, Concentration-responses of 24S-HC in the presence of various concentrations of PS. For each PS concentration, EC50 values with 95% CI were 531 nM (([411 nM, 941 nM], 2 µM PS), 568 nM (([383 nM, 2.2 µM], 10 µM PS), 714 nM (([444 nM, 2.25 µM], 50 µM PS). The Hillslope was 1.19 for 2 µM, 1.14 for 10 µM, and 0.768 for 50 µM. Experiments were done with six or seven individual oocytes per PS concentration. Data for 0 μM PS condition were previously shown in Fig. 2d. Error bars indicate mean ± SEM.
Extended Data Fig. 4 Mechanism of 24S-HC potentiation.
a, Superposition of the 24S-HC stabilized open (deep teal) and closed (grey) structures. Binding of 24S-HC at the GluN2B juxtamembrane pocket favours movement of GluN2B pre-M1’ (dotted arrow). This pre-M1’ movement is essential to accommodate the bending of the pore-forming GluN2B M3’ helices. b, Conformational coupling of LBDs and TMDs in GluN1 (upper panel) and GluN2B (lower panel). Shown here are the superpositions of the closed (grey) and open (magenta and deep teal) states. The Cα atoms of the gating ring residues, right above the channel gate, for GluN1a (Ile664) and GluN2B (Gln662), are shown as spheres. Note the substantial positional shift of GluN2B Gln662 residues between open and closed states, contrasted with the minimal displacement of GluN1a Ile664 residues (dotted lines).
Extended Data Fig. 5 HOLE analysis of 24S-HC-bound GluN1a-2B NMDAR in closed state and EU1622-240-bound GluN1a-2B NMDAR in sub-open state, and PMF calculation of 24S-HC-bound GluN1a-2B NMDAR in fully open state.
a, In the closed state, the channel is highly constricted at both VIVI-gate and the Asn-ring. b, In the sub-open state, the channel is dilated at the VIVI-gate, with partial dilation at the Asn-ring. Arrows on the model represent the kinked region of the GluN2B M3’ helices. c, All-atom potential of mean force (PMF) calculation of the free energy for Na+ ions around the TMD region of the 24S-HC-bound fully open state. The wider pore radius at the Thr-ring in the fully open state (2.2 Å in the sub-open state, 3.0 Å in the fully open state) corresponds to a modest energy barrier (~ 1.7 kcal mol−1) that permits Na+ permeation without stable binding, whereas the Asn-ring displays favourable free energy consistent with its role as a secondary gate. PMF calculation of EU1622-240-bound GluN1a-2B was adapted from ref. 21.
Extended Data Fig. 6 Hydration pattern of the channel pore in closed, sub-open, and fully open states.
a, Cross-section of the channel pore of the closed (left), sub-open (middle), and fully open (right) states of GluN1a-2B NMDAR. Water molecules are coloured in slate spheres overlaid with cryo-EM density in grey mesh. Key residues involved in the water network are shown. b, Local resolution of water density in the channel pore.
Extended Data Fig. 7 Conformational states of the PS-bound GluN1a-2B NMDAR.
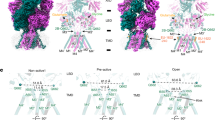
The structure of the PS-bound structures in the non-active and pre-active states viewed from the side (upper panels) and top (lower panels) of the TMDs. The pre-active state shows a greater distance between the GluN2B Gln662 residues compared to the non-active state due to the GluN1a-2B LBD dimer rotations (curved arrows). In both cases, the channel gate remains closed. The hydrophobic gate residues are shown as magenta (GluN1) and deep teal (GluN2B) spheres.
Extended Data Fig. 8 Cryo-EM workflow for EU1622-240-bound GluN1a-2B NMDAR structure.
a, Single-particle cryo-EM analysis of glycine, glutamate, and EU1622-240-bound GluN1a-2B NMDAR. The most predominant conformation of this dataset was the sub-open state. Data processing was performed on sets of movies listed in Supplementary Table 2. The white bar in the micrograph indicates 20 nm. b, Map quality assessment of the sub-open state.
Extended Data Fig. 9 EU1622-240 reduces Ca2+ permeability.
a, The representative current-voltage curves for GluN1-2B responses in the presence 100 μM glutamate and 30 μM glycine with either vehicle (left) or 24S-HC/EU1622-240 (right) in different calcium concentrations. b, The average reversal potentials for vehicle and 24S-HC/EU1622-240 were determined from current-voltage relationships in solutions with different concentrations of Ca2+. c, Mean ΔReversal potential (high Ca2+ minus low Ca2+). Statistical significance was assessed using a two-tailed t-test; the asterisk (*) indicates p = 0.0405, n.s., not significant. d, The average permeability ratio of Ca2+ to monovalent cations derived from a modified version of Lewis equation. Values are shown as mean ± S.E.M. Data are from independent measurements: vehicles (n = 6), 24S-HC (n = 13), EU1622-240 (n = 10). Statistical significance was assessed using a two-tailed t-test; the asterisk (*) indicates p = 0.0112, n.s., not significant.
Supplementary information
Supplementary Figure 1 (download DOCX )
Umbrella histogram and block analysis of (a) Cl- and (b) Na+. Umbrella sampling histograms show good window overlap along the reaction coordinate for both ions. Block analysis used 2-ns incremental increases in the data used for each per PMF calculation. Both systems converged to within thermal energy by the final block.
Supplementary Tables (download DOCX )
Supplementary Tables 1-4.
Supplementary Video 1 (download MP4 )
Conformational transitions between the closed state, sub-open state, and fully-open state of GluN1a-2B NMDAR. Global conformational changes are presented first, followed by zoomed-in views highlighting local rearrangements around the GluN1a M3 and GluN2B M3′ helices. GluN1a is coloured in magenta, and GluN2B in deep teal.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kang, H., Steigerwald, R., Ullman, E.Z. et al. Mechanism of conductance control and neurosteroid binding in NMDA receptors. Nature 648, 220–228 (2025). https://doi.org/10.1038/s41586-025-09695-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09695-4