Abstract
Somatically acquired mutations in the E1 ubiquitin-activating enzyme UBA1 within haematopoietic stem and progenitor cells (HSPCs) were recently identified as the cause of the adult-onset autoinflammatory syndrome VEXAS (vacuoles, E1 enzyme, X linked, autoinflammatory, somatic)1. UBA1 mutations in VEXAS lead to clonal expansion within the HSPC and myeloid, but not lymphoid, compartments. Despite its severity and prevalence, the mechanisms by which UBA1 mutations cause multiorgan autoinflammation and haematological disease are unclear. Here we use somatic gene-editing approaches to model VEXAS-associated UBA1 mutations in primary macrophages and HSPCs. Uba1-mutant mouse macrophages exposed to inflammatory stimuli underwent aberrant apoptotic and necroptotic cell death mediated by caspase-8 (CASP8) and RIPK3–MLKL, respectively. Accordingly, in mice challenged with TNF or LPS, the UBA1 inhibitor TAK-243 exacerbated inflammation in a RIPK3–CASP8-dependent manner. By contrast, Uba1 mutation in HSPCs induced an unfolded protein response and myeloid bias independently of RIPK3–CASP8. Mechanistically, aberrant cell death of Uba1-mutant macrophages coincided with a kinetic defect in Lys63/Met1 (that is, linear) polyubiquitylation of inflammatory signalling complexes. Collectively, our results link VEXAS pathogenesis to the pathogenesis of rarer monogenic autoinflammatory syndromes, highlight specific ubiquitin-associated defects stemming from an apical mutation in the ubiquitylation cascade and support therapeutic targeting of the inflammatory cell death axis in VEXAS.
Similar content being viewed by others
Main
Since the recent discovery of VEXAS syndrome1, hundreds of patients with VEXAS have been identified, with prevalence estimates reaching around 1 in 4,269 male individuals aged over 50 years2,3. VEXAS is part of a disease paradigm in which age-associated somatic mutations in the haematopoietic lineage drive clonal expansion and are associated with haematological malignancy and inflammatory conditions4,5. However, in VEXAS, inflammatory manifestations are overt and severe, involving the skin, lungs, vasculature and musculoskeletal system6.
In most patients, mutations affect Met41 in UBA1, substituting it with leucine, valine or threonine. Alternatively, mutations in the exon 3 splice acceptor site cause aberrant splicing that results in skipping of Met411,7,8,9,10. Normally, translation of UBA1 yields two isoforms that arise from translation initiation at Met1 (UBA1a) or Met41 (UBA1b). Met41 mutations in VEXAS thereby diminish UBA1b translation and lead to aberrant translation initiation at Met67 to generate an enzymatically impaired isoform, UBA1c1,8. However, the molecular mechanisms through which somatic Met41 mutations or ‘Met41 skipping’ splice mutations in UBA1 lead to the clinical sequelae of VEXAS are unclear.
UBA1 is the dominant E1 ubiquitin-activating enzyme in cells. Although rare, multiple other genetic mutations affecting ubiquitin-related proteins cause autoinflammatory syndromes and share some clinical features with VEXAS. These include genetic mutations in components of the linear ubiquitin chain assembly complex (LUBAC), the linear deubiquitylase OTULIN, the ubiquitylation substrate RIPK1 and the ubiquitin-binding protein A2011,12,13,14,15,16,17,18,19,20,21,22,23. These mutations variably affect NF-κB signal transduction and inflammatory cell death. Although UBA1 has a more apical role in ubiquitylation than these proteins, we sought to evaluate the effects of VEXAS-associated Uba1 mutations on inflammatory signalling and cell death.
Base editing Uba1 to model VEXAS
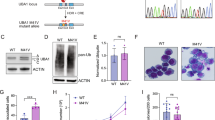
We first developed a somatic gene-editing approach to model endogenous Uba1 M41T mutation. To do so, we electroporated bone-marrow-derived macrophages (BMDMs) or ex vivo expanded HSPCs with mRNA encoding an ABE8e base editor coupled to a SpRY SpCas9 variant (ABE8e-SpRY) plus either a single guide RNA (sgRNA) capable of creating an A(T)G>A(C)G (p.M41T) mutation in Uba1 (hereafter sgM41T) or a control sgRNA24,25,26 (sgGFP; Fig. 1a). Male primary cells were used as UBA1 is X-chromosome linked and VEXAS predominantly affects male individuals. Within 72 h of electroporation and without any selection conditions, this approach yielded high levels (around 85%) of the desired M41T edit in BMDMs and HSPCs27, precisely mimicking the M41T mutation seen in VEXAS (Fig. 1b); we term these cells Uba1M41T/y cells. Moreover, a bystander T>C mutation within the intron upstream of exon 3 in Uba1 was detected in around 9.6% of sequenced Uba1 alleles, but this mutation spares the exon 3 splice acceptor motif (Extended Data Fig. 1a,b).
a, Schematic of the somatic gene-editing approach in BMDMs and HSPCs to generate Uba1M41T/y cells. The diagram was created using BioRender. b, Sanger sequencing of the genomic region surrounding Met41 in Uba1 BMDMs 5 days after electroporation with either sgM41T (Uba1M41T/y) or sgGFP (Uba1+/y) plus mRNA encoding ABE8e-SpRY. c, Immunoblot analysis of the indicated BMDMs 9 days after electroporation. d, The viability of Uba1+/y (n = 6), Uba1M41T/y (n = 6) and Uba1KO/y (n = 3) BMDMs 9 days after electroporation. e,f, Immunoblot analysis of BMDMs of the indicated genotypes treated with TNF (e) or LPS (f) for the indicated lengths of time. g, IL-6 secretion by Uba1+/y or Uba1M41T/y BMDMs treated with the indicated stimuli for 24 h. n = 3 biological replicates per stimulus. h, Immunoblot analysis of BMDMs of the indicated genotypes treated with TNF for the indicated lengths of time. All assays in this figure were performed 9 days after electroporation. Results are representative of 2 or 3 independent experiments. For d and g, data are mean ± s.e.m.; the dots represent biological replicates. P values were calculated using one-way analysis of variance (ANOVA) with Bonferroni’s multiple-comparison test (d) and unpaired two-sided t-tests (g); ****P < 0.0001.
Immunoblotting of Uba1M41T/y BMDMs confirmed a decrease in UBA1b, as expected due to impairment of translation initiation at Met41. Moreover, a faster-migrating UBA1 protein isoform was detected on SDS–PAGE, analogous to the UBA1c isoform observed in cells of patients with VEXAS (Fig. 1c). Residual production of UBA1b in M41T BMDMs may occur due to non-canonical translation initiation at the mutant M41T codon, as previously suggested, and/or residual UBA1b expression from non-edited cells8. To examine off-target editing, we evaluated 12 of the highest-likelihood off-target editing sites, all of which fell outside of annotated protein-coding genes. No off-target edit exceeded a variant allele frequency of 0.47% (Extended Data Fig. 1c). These results establish our cellular systems as robust and faithful models of the genomic and protein-level changes caused by M41T mutation of Uba1 in primary murine macrophages.
In parallel, we CRISPR-deleted Uba1 by electroporating BMDMs with recombinant Cas9 plus control or Uba1-targeting sgRNAs (Uba1KO/y). Notably, while Uba1M41T/y BMDMs exhibited only modestly decreased global ubiquitylation and preserved cell viability, Uba1KO/y BMDMs displayed a pronounced decrease in cellular ubiquitylation and spontaneous cell death (Fig. 1d and Extended Data Fig. 1d,e). These findings suggest that the M41T mutant of Uba1 is not a complete loss-of-function allele.
Signalling in Uba1-mutant cells
Given the pronounced inflammatory phenotype in VEXAS, we first evaluated the effects of UBA1(M41T) expression on innate immune signalling downstream of TNF and toll-like receptor TLR4. Uba1M41T/y or Uba1+/y BMDMs were stimulated with a time-course of TNF or bacterial lipopolysaccharide (LPS; a TLR4 agonist; Fig. 1e,f). Both TNF and LPS elicited broadly similar signalling profiles by immunoblotting. Notably, we observed modestly perturbed NF-κB signalling in Uba1M41T/y BMDMs, including dampened phosphorylation of IKKα/β and dampened or delayed phosphorylation of p105, although phosphorylation and degradation of IκBα remained largely unaffected (Fig. 1e,f). Phosphorylation of p38 and JNK also appeared unaffected. Consistent with these subtle effects, secretion of the NF-κB-dependent cytokine IL-6 was slightly decreased in Uba1M41T/y BMDMs stimulated with LPS, Pam3CSK4 (TLR1–TLR2 heterodimer agonist), R837 (TLR7 agonist) or CpG-B DNA (TLR9 agonist) (Fig. 1g). We conclude that expression of UBA1(M41T) in BMDMs led to modest reductions in TNF- and LPS-induced signalling and TLR-dependent cytokine production, which did not provide a facile explanation for the pathological autoinflammatory features of VEXAS.
Uba1 and inflammatory cell death
In contrast to the modest signalling changes described above, TNF induced heightened autophosphorylation of RIPK1 in Uba1M41T/y BMDMs, which was abolished in Uba1M41T/yRipk1D138N/D138N BMDMs carrying a kinase-inactivating point mutation in RIPK1 (Fig. 1h). Given the role of RIPK1 in cell death, we tested the susceptibility of Uba1M41T/y BMDMs to TNF-induced cell death. Indeed, treatment with TNF alone induced cell death in a fraction of Uba1M41T/y BMDMs, whereas no death was observed in Uba1+/y BMDMs (Fig. 2a and Extended Data Fig. 2a). To further examine TNF-induced cell death, we co-treated cells with TNF and low doses of the inhibitor of apoptosis (IAP) antagonist compound birinapant (hereafter, IAPi). IAPi promotes TNF-induced cell death by depleting the E3 ligases cIAP1 and cIAP228,29,30,31. TNF plus low dose IAPi killed nearly all Uba1M41T/y BMDMs, while largely sparing control BMDMs (Fig. 2a and Extended Data Fig. 2a). We next tested additional TNF-related cell death stimuli, including co-treating BMDMs with TNF plus a low dose of emricasan (a pan-caspase inhibitor), which induces necroptosis, or TNF plus IFNγ. Both these co-treatments led to significantly more death among Uba1M41T/y BMDMs compared with controls (Fig. 2a and Extended Data Fig. 2a), suggesting that VEXAS-associated M41T mutation in Uba1 aberrantly enhances multiple forms of TNF-induced cell death.
a, Cell death was measured over 24 h among BMDMs of the indicated genotypes treated with the indicated stimuli 9 days after electroporation, quantified as the percentage of YOYO-1+ cells among total cells, averaged across 3 images taken per well. Data are mean ± s.e.m. of triplicate wells per genotype and condition. P values were calculated using one-way ANOVA with Tukey’s multiple-comparison test. The timepoints used for statistical tests were as follows: t = 24 h (TNF, TNF + emricasan (Emr), TNF + IFNγ, LPS, LPS + IFNγ); t = 16 h (TNF + IAPi); t = 7 h (LPS + emricasan). b, Crispresso227 analysis of amplicon sequencingof the genomic region surrounding exon 3 splice (spl.) acceptor site of Uba1 (DNA) and of a Uba1 reverse transcription–PCR (RT–PCR) product generated from RNA, both isolated from BMDMs 5 days after electroporation with Cas9 ribonucleoprotein. The deletion frequency is shown in black at each nucleotide (nt) position with the reference sequence and gene structure depicted above. Bottom left, the relative frequency of reads containing deletions of the indicated lengths. Values are expressed as the percentage of all deletion-containing reads. Middle, schematic of the most commonly detected deletion events. Right, immunoblots of Uba1∆Spl/y BMDMs 8 days after electroporation. c, Cell death of BMDMs of the indicated genotypes over 24 h after indicated treatments 9 days after electroporation, quantified as the percentage YOYO-1+ cells among total cells. Data are mean ± s.e.m. of triplicate wells, with three images taken per well for quantification. d,e, Immunoblots of BMDMs treated with TNF plus emricasan (TE) (d) or either TNF or LPS alone or in combination with IAPi, IFNγ or emricasan (e) as indicated for 6 h. The results are representative of 2–3 independent experiments. Cl., cleaved; Unstim., unstimulated.
In addition to TNFR1, TLR4 has been linked to the necroptotic and apoptotic cell death machinery through the adaptor TRIF32,33. We therefore stimulated BMDMs with LPS either alone or in combination with IFNγ or emricasan. All of these stimuli elicited exacerbated cell death among Uba1M41T/y BMDMs (Fig. 2a and Extended Data Fig. 2a). Importantly, LPS-induced death of Uba1M41T/y BMDMs was not solely due to autocrine or paracrine TNF induced by LPS stimulation because LPS, LPS plus IFNγ and LPS plus emricasan killed Tnf−/−Uba1M41T/y BMDMs more robustly than Tnf−/−Uba1+/y BMDMs (Extended Data Fig. 2b). Given the hyperactivation of RIPK1 induced by TNF stimulation in Uba1M41T/y BMDMs (Fig. 1h), we tested the requirement for RIPK1 kinase activity in mediating the cell death phenotype of Uba1M41T/y BMDMs. Consistent with a contributory role for RIPK1 kinase activity, the RIPK1 kinase inhibitor necrostatin-1s (Nec-1s) or genetic inactivation of RIPK1 (Ripk1D138N/D138N) partially suppressed the sensitization to cell death of Uba1M41T/y BMDMs induced by TNF- and LPS-containing stimuli to a varying degree across individual stimuli (Extended Data Fig. 2c,d). Thus, expression of UBA1(M41T) in macrophages exacerbates both RIPK1-kinase-dependent and kinase-independent inflammatory cell death.
Base editing of BMDMs to produce two recurrent clonal haematopoiesis-associated mutations Tet2H1380R or Dnmt3aM852V did not exacerbate cell death after exposure to either LPS, TNF plus emricasan or TNF plus IAPi34,35,36 (Extended Data Fig. 2e). Thus, the increased propensity of at least two distinct immune receptors (TNFR1 and TLR4) to kill BMDMs was specific to cells containing the Uba1M41T/y mutation and not a general feature of cells prone to aberrant clonal expansion in the haematopoietic compartment. As an additional set of controls, inflammatory cell death of Uba1M41T/y BMDMs was increased compared with that in Uba1+/y BMDMs generated using either a non-targeting sgRNA (sgGFP) or sgRNAs targeting unrelated genomic loci (sgTrac or sgRosa26) (Extended Data Fig. 2f).
We also performed base editing of endogenous UBA1 to generate the M41T mutation in human macrophages differentiated from peripheral blood monocytes from healthy male donors. Like mouse BMDMs, human UBA1M41T/y monocyte-derived macrophages exhibited increased cell death in response to a similar set of TNF- and LPS-containing stimuli (Extended Data Fig. 3a). Moreover, base-edited UBA1M41T human THP-1 monocytic cells also displayed increased cell death elicited with TNF- and LPS-containing stimuli compared with UBA1-wild-type (WT) cells electroporated with a control sgRNA targeting the AAVS1 locus (Extended Data Fig. 3b,c). Lentiviral expression of UBA1(WT) in UBA1M41T THP-1 cells reduced cell death induced by TNF or TNF plus IAPi to levels similar to those observed in control cells, confirming that these phenotypes arose due to on-target UBA1 editing (Extended Data Fig. 3d). Thus, aberrant inflammatory cell death programs characterize both mouse and human myeloid cells carrying the VEXAS-associated M41T mutation in UBA1.
To determine whether VEXAS-associated UBA1 M41V and splice-site mutations similarly exacerbate inflammatory cell death, we first generated Uba1M41V/y BMDMs through electroporation of a distinct sgRNA (sgM41V) and ABE8e-NG base-editor pair (Extended Data Fig. 3e). Like Uba1M41T/y BMDMs, Uba1M41V/y BMDMs showed decreased UBA1b levels and expression of UBA1c, modestly decreased total ubiquitin and significantly increased cell death after treatment with TNF- and LPS-containing stimuli (Extended Data Fig. 3f,g). To model VEXAS-associated splice-site mutations that recurrently occur in the splice acceptor site upstream of exon 3, we targeted this region with an sgRNA complexed with recombinant Cas9 protein. As expected, Cas9-mediated targeting of this site in male BMDMs (Uba1∆Spl/y) led to heterogenous insertions and deletions at this splice acceptor site at the genomic level (Fig. 2b). Notably, amplicon sequencing of Uba1 mRNA surrounding this region revealed a recurrent in-frame 12-nucleotide-skipping event spanning Met41 and identical to that observed in patients with VEXAS (Fig. 2b)7. Other mRNA reads in these cells included skipping exon 3 altogether (Fig. 2b). Immunoblotting confirmed decreased UBA1b expression with concomitant expression of a presumed UBA1c isoform in Uba1∆Spl/y BMDMs alongside a modest decrease in total ubiquitin (Fig. 2b). Like Uba1M41T/y and Uba1M41V/y BMDMs, Uba1∆Spl/y BMDMs also showed increased inflammatory forms of cell death (Fig. 2c). Thus, we conclude that dysregulated inflammatory cell death is a prominent feature across human and mouse cells and multiple VEXAS-associated UBA1 mutations.
TNF and LPS can promote CASP8-mediated apoptosis or RIPK3–MLKL-mediated necroptosis when CASP8 is ablated37,38,39,40. We therefore assessed the relative contributions of apoptosis versus necroptosis by editing Uba1 M41T in WT, Mlkl−/− and Mlkl−/−Casp8−/− BMDMs. Exacerbated cell death in response to emricasan plus either TNF or LPS was fully abrogated in Uba1M41T/yMlkl−/− BMDMs compared with that in Uba1M41T/yMlkl+/+ BMDMs. Cell death induced by TNF plus IAPi, TNF plus IFNγ, LPS or LPS plus IFNγ were partially diminished by deletion of Mlkl and fully abrogated by co-deleting Mlkl together with Casp8 in Uba1M41T/y BMDMs, implicating components of necroptosis and apoptosis involved in the cell death response (Fig. 2a). Confirming these findings, partial amelioration of the cell death phenotype was observed in Uba1M41T/y Ripk3−/− BMDMs compared with that in Uba1M41T/y BMDMs, respectively, while more complete abrogation of exacerbated death of Uba1M41T/y BMDMs was achieved after combined deletion of Ripk3 and Casp8 (Extended Data Fig. 4a). Consistent with exacerbated apoptosis and necroptosis, we observed accelerated phosphorylation of RIPK3 and MLKL, markers of necroptosis, in Uba1M41T/y BMDMs after emricasan co-treatment with either TNF or LPS (Fig. 2d,e) and increased cleavage of caspases 3, 7 and 8, markers of apoptosis, after treatment with TNF plus IAPi, TNF plus IFNγ, LPS or LPS plus IFNγ (Fig. 2e). Ripk3 deletion in Uba1∆Spl/y BMDMs minimally ameliorated cell death induced by TNF, TNF plus IAPi, TNF plus IFNγ, LPS or LPS plus IFNγ, which was largely CASP8 dependent (Fig. 2c). By contrast, Ripk3 deletion blocked cell death of Uba1∆Spl/y BMDMs that were treated with either TNF or LPS in combination with emricasan (Fig. 2c). These differences between Uba1∆Spl/y and Uba1M41T/y BMDMs could reflect variable inflammation-induced cell death mechanisms among different VEXAS-associated Uba1 mutations.
We next evaluated whether the UBA1 inhibitor TAK-243 could similarly induce inflammatory cell death in BMDMs. Treatment of BMDMs with 1 μM of TAK-243 killed WT, Ripk3−/− and Ripk3−/−Casp8−/− BMDMs, although Ripk3−/−Casp8−/− BMDMs died at a slower rate (Extended Data Fig. 4b). This suggests that high concentrations of TAK-243 can trigger cell death that is independent of RIPK3-dependent necroptosis and is accelerated by, but does not require, CASP8. Similarly, co-deletion of Mlkl and Casp8 did not substantially restore viable cell yields of Uba1KO/y BMDMs to normal (Extended Data Fig. 4c,d). By contrast, 100 nM TAK-243 treatment did not kill BMDMs unless they were co-treated with TNF- or LPS-containing stimuli (Extended Data Fig. 4e). Moreover, similar to our observations with Uba1M41T/y and Uba1∆Spl/y BMDMs, TAK-243-mediated exacerbation of cell death induced by TNF- and LPS-containing stimuli was mediated largely through RIPK3 and CASP8 (Extended Data Fig. 4e).
On the basis of these results, we tested whether partial UBA1 inhibition in vivo using TAK-243 could exacerbate inflammatory challenges with systemic TNF or LPS. Intraperitoneal (i.p.) injection of low dose (3 mg per kg) TAK-243 in WT mice did not elicit a detectable systemic inflammatory response as measured by changes in core body temperature. By contrast, co-injection of TAK-243 plus intravenous TNF led to severe hypothermia and mortality of WT mice compared with injection of TNF alone (Fig. 3a). This exacerbation could be substantially blocked by RIPK1 inhibition with Nec-1s or by genetic inactivation of RIPK1 kinase activity, both of which provided a survival benefit (Fig. 3a,b). Ripk3−/− mice also showed partial protection from TAK-243-mediated exacerbation of systemic TNF injection, whereas Ripk3−/−Casp8−/− mice were protected to an even greater degree (Extended Data Fig. 4f). To evaluate whether TAK-243 could also exacerbate the acute sepsis model of i.p. LPS injection, we generated bone-marrow chimeras in which WT mice were reconstituted with bone-marrow cells from WT, Ripk1D138N/D138N, Ripk3−/− or Ripk3−/−Casp8−/− mice. In WT control chimeras, the systemic inflammatory response to i.p. injection of LPS was markedly exacerbated by co-treatment with 3 mg per kg TAK-243 (Fig. 3c). Exacerbation of LPS-mediated inflammation by TAK-243 was attenuated in Ripk3−/− and even further in Ripk3−/−Casp8−/− chimeric mice (Fig. 3c). Thus, TAK-243 augments LPS-mediated inflammation in a manner that depends on RIPK3–CASP8 within haematopoietic cells. By contrast, Ripk1D138N/D138N chimeras were not discernibly protected (Extended Data Fig. 4g), possibly owing to RIPK1 kinase-independent cell death induced by LPS (Extended Data Fig. 2c,d). Although a contribution of off-target effects of TAK-243 cannot be excluded, these findings suggest that UBA1 impairment exacerbates inflammation through the RIPK3–CASP8 axis in vivo.
a, Rectal temperatures and survival of C57BL/6 mice after i.v. injection of TNF (200 μg per kg), pretreated 2 h before with either vehicle or TAK-243 (3 mg per kg i.p.), and 30 min before with either PBS or Nec-1s (6 mg per kg, i.v.). The mean rectal temperatures are shown for n = 10–15 mice per condition. b, Rectal temperatures and survival of Ripk1+/+ (n = 10) and Ripk1D138N/D138N (n = 8) mice after i.v. injection of TNF (200 μg per kg), pretreated 2 h before with TAK-243 (3 mg per kg, i.p.). c, Rectal temperatures and survival of bone-marrow chimeras of the indicated genotypes (n = 6–7 mice per cohort) after i.p. injection of LPS (5 mg per kg), pretreated 2 h before with vehicle or TAK-243 (3 mg per kg, i.p.). Experiments were performed 5 weeks after 10 Gy irradiation and bone marrow transplantation. P values comparing survival curves in a–c were calculated using the log-rank (Mantel–Cox) test; *P < 0.05, **P < 0.01, ***P < 0.001. For a–c, data are mean ± s.e.m.
Uba1, inflammasomes and UPR signalling
Activation of the inflammasome has been suggested to occur in VEXAS based on studies of patient samples41. To evaluate inflammasome signalling, Pam3CSK4-primed Uba1M41T/y BMDMs were exposed to agonists of NLRP3 (nigericin or ATP), NLRC4 (flagellin or Rod-Tox) or AIM2 (poly(dA:dT)). LDH release induced by all of these inflammasome stimuli were similar among Uba1M41T/y and Uba1+/y BMDMs (Extended Data Fig. 5a), suggesting comparable induction of pyroptosis. By contrast, production of IL-1β was slightly diminished among Uba1M41T/y BMDMs (Extended Data Fig. 5b), which may be due to dampened NF-κB signalling during Pam3CSK4 priming (Fig. 1g). Consistent with these findings, Casp1−/−Casp11−/−Uba1M41T/y BMDMs were still killed at elevated levels compared with Casp1−/−Casp11−/−Uba1+/y cells in response to TNF- and LPS-containing stimuli (Extended Data Fig. 5c). These findings suggest that inflammasome activation in patients with VEXAS may be a secondary rather than primary effect stemming from UBA1 mutation in myeloid cells.
We next assessed autophagy and the unfolded protein response (UPR), which have previously been observed in UBA1-mutant monocytes in VEXAS1,42. Uba1M41T/y mouse BMDMs did not show elevated baseline or bafilomycin-induced expression of p62, GABARAP or LC3 (Extended Data Fig. 5d). These cells also did not display altered levels of PERK phosphorylation, XBP1-s, ATF4, BiP or CHOP either at baseline or after induction of endoplasmic reticulum stress with tunicamycin (Extended Data Fig. 5e). By contrast, UBA1M41T THP-1 monocytes did show evidence of an elevated UPR based on increased expression of ATF4 (Extended Data Fig. 3b). Inhibition of PERK using AMG’44 (ref. 43), which blocked tunicamycin-induced CHOP expression and PERK autophosphorylation (Extended Data Fig. 5f), modestly increased cell death of Uba1M41T/y BMDMs elicited by TNF- and LPS-containing stimuli (Extended Data Fig. 5g). Thus, PERK-dependent signalling, which was previously implicated in VEXAS, does not drive the cell death phenotype observed here, but its inhibition may further heighten inflammatory cell death of Uba1M41T/y cells. Treatment of Uba1M41T/y cells with TNF plus IAPi also did not induce UPR markers ATF4, BiP or XBP1-s, despite a partial reduction in UBA1 levels in these cells after treatment (Extended Data Fig. 5h). Thus, the enhanced cell death phenotype induced by UBA1 mutation is observed among cells that either do not (mouse BMDMs) or do (human THP-1 monocytes) show baseline UPR activation1 (Fig. 2a and Extended Data Figs. 3b,c and 5e).
Uba1 mutation and myeloid bias
UBA1 mutations in VEXAS originate in HSPCs. To evaluate these effects in human HSPCs (hHSPCs), we isolated CD34+ HSPCs from human cord blood (Extended Data Fig. 6a). As expected, base editing of these cells to generate UBA1M41T/y hHSPCs led to pronounced vacuoles and signs of an UPR as determined by immunoblotting of ATF4 and BiP (Fig. 4a and Extended Data Fig. 6b,c). We next transplanted control UBA1+/y hHSPCs or UBA1M41T/y hHSPCs into immunodeficient NOD.Cg-KitW-41JTyr+PrkdcscidIl2rgtm1Wjl/ThomJ (NBSGW) mice. As expected and consistent with other reports44, engraftment of UBA1+/y hHSPCs progressively increased over time, characterized by predominant production of CD19+ lymphocytes. By contrast, UBA1M41T/y hHSPCs failed to expand over time and generated almost exclusively myeloid progeny (Fig. 4b,c, Extended Data Fig. 6d and Supplementary Fig. 2), paralleling myeloid-biased haematopoiesis and progressive bone-marrow failure observed in patients with VEXAS.
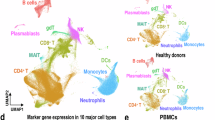
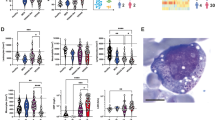
a, Sanger sequencing of genomic region surrounding Met41 in human UBA1 among CD34+ HSPCs collected 3 days after electroporation with either sgAAVS1 (UBA1+/y; top) or sgM41T (UBA1M41T/y; middle) plus mRNA encoding ABE8e-NGG. Bottom, representative Sanger sequencing of human CD33+ cells sorted from NBSGW recipient mice 4 weeks after transplantation of UBA1M41T/y HSPCs. Pre-Tx, pre-transplant. b,c, The percentage of human CD45+ cells within the total CD45+ cell population of bone marrow (b) and the percentage of CD19+ or CD33+ cells among the human CD45+ population (c) in NBSGW mice transplanted with HSPCs of the indicated genotypes (n = 10 mice per genotype) at 4 or 8 weeks after transplantation. Data are mean ± s.e.m.; each dot represents the results from a single mouse. For b and c, P values were calculated using unpaired two-sided t-tests. Right, representative flow cytometry plot gated on live human CD45+ cells in the bone marrow of NBSGW mice at 4 weeks after transplantation. The mean ± s.e.m. percentage is displayed adjacent to each gate. d, Schematic of the scRNA-seq experimental approach. e,f, Reduced dimensionality projection and joint clustering of cells collected 5 and 8 days after electroporation across all genotypes. Colouring highlights major cell types and clusters (e) or genotype or timepoint (f), as indicated. g, The percentage of cells within the macrophage-like cluster for the indicated genotypes and timepoints. Statistical analysis was performed using two-sided Fisher’s exact tests. h, Gene set module scores for the indicated pathways, stratified by genotype. Statistical analysis was performed using two-sided Mann–Whitney U-tests. The box plots show the median (centre line), first and third quartiles (box limits) and 1.5× interquartile range (whiskers). For g and h, ****P < 2.2 × 10−16. i, The frequency of cells within each annotated cell type, stratified by genotype and timepoint as indicated. In d–i, three biological replicates per genotype were pooled and then analysed for each timepoint.
To examine the underlying mechanisms in mouse HSPCs, we generated Uba1M41T/y HSPCs and noted spontaneous acquisition of myeloid markers in cell culture including CD11b, F4/80 and Gr-1 despite culture conditions that normally promote preservation of a stem cell state45 (Extended Data Fig. 7a–d). Moreover, expression of UBA1(M41T) decreased the fraction of Lineage (Lin)−KIT+ cells and Lin− SCA-1+KIT+CD150+ HSPCs (Extended Data Fig. 7b and Supplementary Fig. 3). Morphologically, we noted numerous macrophage-like cells within Uba1M41T/y HSPC cultures (Extended Data Fig. 7d) and detected signs of an UPR in Uba1M41T/y HSPCs as revealed by elevated levels of ATF4, CHOP and XBP1-s (Extended Data Fig. 7e). Like BMDMs, Uba1M41T/y HSPCs also underwent increased levels of cell death in response to the same LPS- and TNF-containing stimuli (Extended Data Fig. 7f) and showed decreased proliferation and viability over time in culture (Extended Data Fig. 7g). Notably, TNF- or LPS-induced cell death of Uba1M41T/y BMDMs led to increased release of damage-associated molecular patterns (DAMPs), including IL-1β, IL-18 and IL-1α, of which IL-1β or IL-1α treatment was sufficient to induce myeloid differentiation of HSPCs (Extended Data Fig. 7h–j), consistent with previous reports46. These findings raise the possibility that UBA1 mutation may promote myeloid bias both cell intrinsically and indirectly through DAMPs released through increased myeloid cell death.
Whereas the inflammatory cell death of Uba1M41T/y macrophages was mediated by the RIPK1–RIPK3–CASP8 axis (Fig. 2a and Extended Data Fig. 4a), Uba1 M41T editing in HSPCs led to increased myeloid bias even among cells deficient in Ripk3 or co-deficient in both Ripk3 and Casp8 (Extended Data Fig. 8a). Moreover, Uba1M41T/y, Uba1M41T/yRipk3−/−, Uba1M41T/yRipk3−/−Casp8−/− and Uba1M41T/yRipk1D138N/D138N HSPCs all failed to engraft into lethally irradiated mice after transplantation (Extended Data Fig. 8b–e). This engraftment failure occurred despite Uba1M41T/y HSPCs displaying no detectable bone marrow homing or CXCR4 expression defects (Extended Data Fig. 8f). The inability of Uba1M41T/y HSPCs to successfully engraft in the transplant context was cell-intrinsic, as lethally irradiated mice transplanted with a 1:1 mix of CD45.1+ Uba1+/y and CD45.2+ Uba1M41T/y HSPCs also displayed preferential engraftment by CD45.1+ Uba1+/y cells at the expense of CD45.2+ Uba1M41T/y cells (Extended Data Fig. 8g,h and Supplementary Fig. 4). Single-cell RNA-sequencing (scRNA-seq) analysis of Uba1M41T/y HSPCs collected 5- and 8-days after electroporation confirmed that Uba1 mutation causes spontaneous differentiation towards a macrophage-like state (Fig. 4d–i and Extended Data Fig. 9a), which was substantiated by analysis of Met41 within sequencing reads mapping to the 5′ end of the Uba1 transcript (Extended Data Fig. 9b). Supporting the validity of our model, Uba1-mutant HSPCs activated the UPR, suppressed translation and shared an overlapping gene expression signature with that observed in patients with VEXAS42 (Fig. 4h and Extended Data Fig. 9c). Myeloid differentiation of Uba1M41T/y progressed over time in culture and induced bystander differentiation of Uba1+/y cells by day 8 (Extended Data Fig. 9b–d), whereas the UPR was maximally engaged in Uba1M41T/y HSPCs 5 days after electroporation and subsequently abated (Extended Data Fig. 9c). Importantly, both myeloid differentiation and a VEXAS gene expression signature remained evident among Uba1M41T/y HSPCs on Ripk3−/− or Ripk3−/−Casp8−/− genetic backgrounds (Fig. 4h,i and Extended Data Fig. 9c,e). Thus, we conclude that VEXAS-associated UBA1 mutation induces myeloid bias and impairs the ability of HSPCs to undergo emergency haematopoiesis independently of RIPK3–CASP8-mediated cell death.
Uba1 and ubiquitin-dependent signalling
We next sought to decipher the molecular defect(s) in TNF signalling associated with cell death of Uba1M41T/y macrophages. TNF triggers recruitment of RIPK1 and TRADD to TNFR1; recruitment of TRAF2, cIAP1 and cIAP2 through TRADD then leads to modification of the TNFR1-associated signalling complex with Lys63-linked polyubiquitin30,47,48. In turn, Lys63-linked polyubiquitin chains provide docking sites for HOIP, a component of LUBAC that mediates linear polyubiquitylation of the TNFR1-associated signalling complex49,50,51. Recruitment of the canonical IKKα/β complex through its linear polyubiquitin-binding subunit NEMO (also known as IKKγ) subsequently enables productive NF-κB signalling52. This signalling complex, known as complex I53, depends on rapidly inducible and complex pathways of ubiquitylation that we hypothesized may be perturbed in VEXAS, leading to pathological cell death.
We first stimulated Uba1+/y and Uba1M41T/y macrophages with Flag-tagged TNF for 5 min and immunoprecipitated TNF-induced complex I. This revealed similar TNF-induced TRADD polyubiquitylation and recruitment to TNFR1, but the overall polyubiquitylation patterns on RIPK1 and TNFR1 in Uba1M41T/y macrophages were markedly perturbed (Fig. 5a and Extended Data Fig. 10a,e). Specifically, linear polyubiquitylation within complex I was greatly decreased in Uba1M41T/y macrophages (Fig. 5a and Extended Data Fig. 10a,e). Consistent with impaired linear polyubiquitylation, recruitment of NEMO and IKKβ to complex I was reduced in Uba1M41T/y macrophages, as were downstream NF-κB signalling responses, as previously noted (Figs. 1e,f and 5a). Notably, kinetic analysis of TNF-induced complex I revealed that RIPK1 ubiquitylation and linear ubiquitylation normalized in Uba1M41T/y BMDMs at later timepoints after TNF stimulation, before signalling complex disassembly (Extended Data Fig. 10a). These findings suggest that Uba1 mutation leads to a kinetic ubiquitylation defect localized to TNF-induced complex I, modestly impairing NF-κB signalling and concomitant with the inflammatory cell death propensity of Uba1-mutant macrophages.
a, Immunoblots of Flag immunoprecipitates and input lysates from Uba1+/y or Uba1M41T/y BMDMs treated with Flag–TNF as indicated. b, Immunoblots of denaturing Lys63-polyubiquitin immunoprecipitates and input lysates from Uba1+/y or Uba1M41T/y BMDMs treated for the indicated number of minutes (m). c, Immunoblots of untreated or linear-specific (OTULIN) deubiquitylase-treated Flag immunoprecipitates and input lysates from Uba1+/y or Uba1M41T/y BMDMs treated with Flag–TNF for 5 min. d, Immunoblots of pull-downs using linear polyubiquitin-specific TUBEs and input lysates from LPS-treated BMDMs of the indicated genotypes. e, Proposed model of a feedforward cycle of aberrant HSPC myeloid differentiation and autoinflammation driven by a RIPK1–RIPK3–CASP8 death axis in UBA1-mutant myeloid cells. The diagram was created using BioRender. Experiments in a–d were performed 9 days after electroporation and are representative of three (a) or two (b–d) independent experiments.
To further analyse the nature of this ubiquitylation defect, we stimulated control and Uba1M41T/y BMDMs with TNF and enriched Lys48-, Lys63- or linear-polyubiquitin chains using tandem-ubiquitin-binding entities (TUBEs) specific for these ubiquitin linkage types. Lys48-TUBE pull-downs revealed comparable global as well as RIPK1-associated Lys48-linked ubiquitin among TNF-stimulated Uba1+/y and Uba1M41T/y BMDMs (Extended Data Fig. 10b). By contrast, the ubiquitylation pattern of RIPK1 enriched using Lys63- or linear-TUBEs appeared to be altered at 5 min after TNF stimulation in Uba1M41T/y BMDMs before appearing to nearly normalize at later timepoints (Extended Data Fig. 10c,d). Given that these pull-downs are performed under non-denaturing conditions, we also isolated proteins linked covalently to Lys63-polyubiquitin chains by using a Lys63-polyubiquitin linkage-specific antibody in the presence of 3 M urea to disrupt non-covalent protein–protein associations54. Immunoblotting of these pull-downs from Uba1+/y and Uba1M41T/y macrophages stimulated with TNF revealed that Lys63-polyubiquitylated RIPK1 migrated slightly faster on SDS–PAGE in Uba1M41T/y macrophages (Fig. 5b), suggesting moderately impaired Lys63 polyubiquitin chain formation on RIPK1.
To determine the contribution of each polyubiquitin chain linkage type to the altered ubiquitylation pattern on RIPK1, we treated complex I immunoprecipitated from Uba1+/y and Uba1M41T/y macrophages with Lys63- and/or linear-specific deubiquitylases (AMSH, OTULIN or the pan-deubiquitylase USP21). AMSH cleaved a large fraction of ubiquitylated RIPK1 in Uba1+/y BMDMs, whereas its effect was less pronounced on ubiquitylated RIPK1 in Uba1M41T/y macrophages, indicative of a partial deficit in Lys63 polyubiquitylation of RIPK1 (Extended Data Fig. 10e). OTULIN treatment also cleaved the highest-molecular-mass species of ubiquitylated RIPK1 in Uba1+/y but not Uba1M41T/y BMDMs (Fig. 5c and Extended Data Fig. 10e), consistent with a combined impairment in both Lys63- and linear polyubiquitylation of RIPK1. Moreover, TNFR1 was differentially ubiquitylated in Uba1+/y and Uba1M41T/y macrophages and this was at least partly attributable to differences in its linear polyubiquitylation, as OTULIN treatment cleaved the slower-migrating TNFR1 bands in Uba1+/y but not Uba1M41T/y macrophages (Extended Data Fig. 10e). Recruitment of the LUBAC components HOIP and SHARPIN to complex I was normal and only slightly impaired, respectively (Fig. 5a and Extended Data Fig. 10a). To determine whether this molecular defect extends beyond TNF, we also isolated LPS-induced signalling complexes using linear polyubiquitin-specific TUBEs, which revealed dampened total and IRAK1-associated linear polyubiquitin within Uba1M41T/y relative to Uba1+/y macrophages (Fig. 5d). These results imply that the VEXAS-associated M41T mutation in Uba1 may provoke inflammatory forms of cell death by compromising the normal choreography of polyubiquitylation events that culminate in linear polyubiquitylation of inflammatory signalling complexes.
Discussion
Here, to distinguish between proximal and secondary events in the pathological inflammatory responses elicited by UBA1 mutation, we used base editing and Cas9 to model endogenous Met41 or splice-site mutations that collectively mimic the majority of VEXAS-associated UBA1 mutations. We found that VEXAS-associated Uba1 mutations in macrophages precipitated a combination of exacerbated CASP8-mediated apoptosis and MLKL-mediated necroptosis that coincided with defective linear polyubiquitylation during TNF and LPS signalling. During TNF signalling, this linear polyubiquitylation defect was preceded by a partial impairment in Lys63 polyubiquitylation of RIPK1. This observation links the pathogenesis of autoinflammation in VEXAS with that of rare inflammatory diseases caused by germline mutations in the linear ubiquitylation machinery11,12,13,14,15,18.
VEXAS occurs due to somatic mutation of UBA1 within HSPCs, in which M41T mutation prompted myeloid-biased differentiation by mechanism(s) operating independently of the RIPK1–RIPK3–CASP8 axis. Indeed, transcriptional signatures of myeloid differentiation and the UPR—previously noted in cells from patients with VEXAS1,42,55—were detected in Uba1-mutant HSPCs even after Ripk3 and Casp8 co-deletion. Thus, independent molecular mechanisms emanating from impaired UBA1 function may cause myeloid-biased clonal expansion of UBA1-mutant HSPCs and the autoinflammation mediated by their mutant myeloid progeny.
Macrophages across all UBA1 mutations examined were highly prone to dying after exposure to TNF- or LPS-containing stimuli. These inflammatory forms of cell death, which release DAMPs, may further heighten myeloid-biased haematopoiesis in a feedforward cycle46,56 (Fig. 5e), although this model awaits testing. Notably, Uba1-mutant HSPCs appear similarly prone to inflammatory cell death (Extended Data Fig. 7f), although their evident clonal expansion over years to decades in humans suggests that these cells can overcome this propensity to eventually become clonally dominant44.
Our data suggest that a dysregulated RIPK1–RIPK3–CASP8 axis may have a key role in the autoinflammatory pathology of VEXAS, nominating this pathway as a therapeutic target. Thus, aberrant cell death emerges as a far more common contributor to genetic inflammatory disease than previously appreciated.
Methods
Mice
The Memorial Sloan Kettering Cancer Center Institutional animal care and use committee responsible for ethical compliance approved all animal protocols. Mlkl−/− (ref. 57), Ripk3−/− (ref. 54), Casp8−/− (ref. 58), Tnf−/− (ref. 59), Tnfrsf1a−/− (ref. 60) and Ripk1D138N/D138N (ref. 58) mice have been described previously. C57BL6 WT mice were purchased from Jackson Laboratories and Charles Rivers laboratories. CD45.1+ C57BL6/J, NOD.Cg-KitW-41JTyr+PrkdcscidIl2rgtm1Wjl/ThomJ (NBSGW) and Casp1−/−Casp11−/− mice were purchased from Jackson Laboratories. All mouse strains were maintained in the Sloan Kettering Institute animal facility under specific-pathogen-free conditions in accordance with institutional guidelines and ethical regulations. Mice were housed under a 12 h–12 h light–dark cycle between 08:00 and 20:00 in a temperature-controlled room (22 ± 1 °C, 40–60% humidity), with free access to water and food.
Cell culture
Primary BMDMs were differentiated from male mouse bone marrow for 5–7 days on non-treated plates in Dulbecco’s modified Eagle medium (DMEM) high glucose containing 20 mM HEPES, 50 μM 2-mercaptoethanol, 2 mM L-glutamine, 1 mM sodium pyruvate, 100 U ml−1 penicillin, 100 μg ml−1 streptomycin, 10% FCS and 10% supernatants from NIH 3T3 cells expressing M-CSF.
Mouse HSPCs were sorted and ex vivo expanded based on a previous report45. In brief, bone marrow was obtained from mice (male) by mortar and pestle followed by ACK lysis and KIT immunomagnetic enrichment using Miltenyi anti-mouse CD117 microbeads, and then around 1,000-2,500 lineage−KIT+SCA-1+CD150+ cells were sorted directly into fibronectin-coated 96-well plates with F12 medium supplemented with 1× insulin-transferrin-selenium-ethanolamine (IST-X), penicillin–streptomycin, 10 mM HEPES, 1 mg ml−1 87% polyvinyl alcohol (PVA), 100 ng ml−1 TPO and 10 ng ml−1 SCF. Cells were expanded over the course of 7–10 days before electroporation. The medium was changed every 2–3 days by replacing more than 90% of the medium within the well.
For preparation of human monocytes, male donor buffy coats were purchased from the New York Blood Center. Peripheral blood mononuclear cells were isolated using anti-human CD14 magnetic beads (Miltenyi). Cells were seeded onto non-treated plates in RPMI high glucose supplemented with 10% FBS (Gibco), 2 mM GlutaMax, 100 U ml−1 penicillin, 100 μg ml−1 streptomycin and human M-CSF 100 ng ml−1 (Peprotech). Plated monocytes were cultured for 5 days until adherent before electroporation, which was performed using the sgRNA 5′-TGGCCATTCCCTAGGAATAG-3′, co-electroporated with mRNA encoding ABE8e-SpRY. Cells were then replated onto tissue-culture-treated 96-well plates for Incucyte analysis.
For preparation of human CD34+ HSPCs, fresh male human cord blood units were used as a source. We acknowledge the Abraham J. & Phyllis Katz Cord Blood Foundation (doing business as Cleveland Cord Blood Center) and volunteers from communities in Cleveland, Atlanta and San Francisco, USA, as the source of the cord blood materials used in this study, which were obtained under an MTA. Cord blood units were deidentified and coded before use and obtained through signed informed consent for use in research studies. CD34+ cells were isolated using the EasySep Human CD34 Positive Selection Kit II (StemCell Technologies, 17896). Cells were cultured at a concentration of 0.5–1 × 106 cells per ml in CD34-HSPC medium (StemSpan SFEM II, 1× CC100 (StemCell Technologies, 2690), 50 ng ml−1 TPO, 2 mM L-glutamine, 100 μg ml−1 streptomycin) for 48 h before electroporation.
THP-1 cells were newly purchased from ATCC (TIB-202) and underwent standard quality-control specifications including short-tandem repeat profiling for authentication and mycoplasma testing. Cells were cultured in THP-1 medium (RPMI, 10% FBS, 100 U ml−1 penicillin, 100 μg ml−1 streptomycin, 10 mM HEPES, 1 mM sodium pyruvate, 2mM L-glutamine, 50 μM 2-mercaptoethanol) at 0.4–2 × 106 cells per ml.
Wright–Giemsa staining of gene-edited HSPCs was performed 6–8 days after electroporation using Hema-3 stain (Thermo Fisher Scientific, 23-123869). Around 50,000 cells were centrifuged onto frosted microscope slides using Cytofunnels and a Cytospin centrifuge (Shandon) at 600 rpm for 5 min. Vacuoles were manually quantified across 6–7 high-power fields using the ×20 objective on the Zeiss AxioImager microscope, an ORCA/ER CCD camera (Hamamatsu Photonics) and ZEN 3.3 software (Zeiss).
CRISPR–Cas9 and base-editor electroporation
CRISPR-mediated deletion of Uba1 in mouse BMDMs was performed by Cas9 ribonucleoprotein electroporation as previously described61. In brief, BMDMs were collected and extensively washed with PBS, and then 5 × 106 BMDMs were electroporated with recombinant Cas9 protein v.3 (IDT) complexed with Uba1-specific sgRNAs (seed sequences: 5′-ATGACCAAGGAACTACCCAG-3′; 5′-CATTGGTAAAAATCGAGCGG-3′; 5′-CGACTTGCTGAACTCAACAG-3′) in nucleofector solution P3 (Lonza) using the Lonza 4D-nucleofector system (4D-Nucleofector Core Unit, Lonza, AAF-1003B; 4D-Nucleofector X Unit, AAF-1003X) with previously optimized conditions. The following GFP-targeting sgRNA sequences were used as controls: 5′-GGTGGTGCAGATGAACTTCA-3′ and 5′-GGCATCGACTTCAAGGAGGA-3′. sgRNAs targeting the Rosa26 locus (5′-ACTCCAGTCTTTCTAGAAGA-3′) as well as the Trac locus (5′-CACTCACCTGAACTGGGGTA-3′) were used as additional controls. To generate Uba1∆Spl/y BMDMs, the sgRNA sequence 5′-CGTTCTTCGCCATTCCCTAG-3′ targeting the splice acceptor of exon 3 was used. For human experiments, an additional control sgRNA targeting the human AAVS1 locus was used.
For Uba1 base editing in mouse BMDMs and HSPCs, the above sgRNA sequences targeting Rosa26, GFP and/or Trac were used as controls. To engineer the M41T mutation, ABE8e-SpRY was co-electroporated with the following sgRNA 5′-CGCCATTCCCTAGAGACAGA-3′. To engineer the M41V mutation, ABE8e-NG was co-electroporated with the sgRNA 5′-GGAATGGCGAAGAACGGCAG-3′. To engineer the M41T mutation in human cells (for example, THP-1), cells were co-electroporated with mRNA-encoding ABE8e-NGG and the sgRNA 5′-TGGCCATTCCCTAGGAATAG-3′ or sgAAVS1 control (5′-AGAGCTAGCACAGACTAGAG-3′).
Cells were washed twice with PBS and then electroporated using the Neon NxT Electroporation System (NEON1S, Thermo Fisher Scientific) and corresponding 100 μl kit (N10096, Thermo Fisher Scientific) or 10 μl kit (1096, Thermo Fisher Scientific). BMDMs were electroporated using 1,400 V; 20 ms; 2 pulses. HSPCs were electroporated at 1,700 V; 20 ms; 1 pulse. THP-1 cells were electroporated using 1,300 V; 20 ms; 2 pulses. Electroporations of BMDMs were performed using 5 × 106 BMDMs per 100 μl electroporation, 10 μg of base-editor encoding mRNA and 400 pmol of sgRNA. Electroporations of THP-1 cells and mouse HSPCs was performed on 2.5–3.5 × 105 cells per 10 μl electroporation using 1.5 μg of base-editor encoding mRNA and 40 pmol of sgRNA. Cells were immediately plated after electroporation into non-antibiotic-containing medium, which was exchanged after 24 h for antibiotic containing medium. Electroporated BMDMs were maintained in the medium described above on non-treated plates and replated the day before stimulation onto tissue culture-treated plates. For human CD34+ HSPC gene editing, electroporation was performed using the Lonza 4D-nucleofector system using the EO-100 setting (around 250,000 cells per electroporation). Cells were replated in CD34-HSPC medium and cultured for 1–2 days before transplantation.
Mouse BMDMs, human THP-1 monocytes, human monocyte-derived macrophages and mouse HSPCs were treated as indicated with LPS (from Escherichia coli K12, Invivogen; 100 ng ml−1), R837 (2 μg ml−1, Invivogen), Pam3CSK4 (1 μg ml−1, Invivogen), CpG-B (ODN 1826, 5 μM, Invivogen), TNF (100 ng ml−1, Peprotech), emricasan (2 or 4 μM, Selleck Chemicals), birinapant (500 nM, 1 μM or 2 μM, Selleck Chemicals), necrostatin-1s (30 μM, Cell Signaling Technology), human IFNγ (100 ng ml−1, Peprotech), mouse IFNγ (100 ng ml−1, Peprotech), human IFNγ (100 ng ml−1, Cell Signaling Technology), AMG’44 (10 μM, Sigma-Aldrich), tunicamycin (5 μg ml−1, Millipore) or bafilomycin A1 (50 nM, Cayman Chemicals). IFNγ was given as a 2 h pretreatment before stimulation. For cytokine secretion assays, BMDMs were stimulated with TLR agonists for 24 h in 96-well tissue culture plates containing 40,000–100,000 cells per well. IL-6 was measured using the IL-6 Mouse ProcartaPlex Simplex Kit (EPX01A-20603-901, Thermo Fisher Scientific) and ProcartaPlex Mouse Basic Kit (EPX010-20440-901; Thermo Fisher Scientific) on a Luminex xMAP INTELLIFLEX System (APX2020, Thermo Fisher Scientific) according to the manufacturer’s protocol. Mouse HSPCs were also treated with recombinant IL-1α (25 ng ml−1, Thermo Fisher Scientific, 211-11A-10UG) or IL-1β (25 ng ml−1, Thermo Fisher Scientific, 211-11B-10UG) for 96 h before analysis for differentiation markers.
Bone marrow transplantation
Lin−KIT+SCA-1+CD150+ HSPCs were sorted from male 6–10-week-old mice of various genotypes and cultured ex vivo in PVA-based medium described above for 7–10 days before electroporation. Electroporated cells were genotyped after 2–3 days to confirm editing. Around 200,000 cells were subsequently transplanted 3 days after electroporation into cohorts of lethally irradiated (10 Gy) recipient male mice. After transplantation, mice were serially bled or underwent bone marrow aspiration to monitor engraftment. Genomic DNA was isolated and Uba1 genotyping was performed by PCR and Sanger sequencing. The presence of the Uba1 M41T genotype was quantified using the EditR software.
For LPS sepsis experiments, bone-marrow chimeras were generated by transplanting around 1 × 106 whole bone marrow cells from Ripk3+/+Casp8+/+, Ripk3−/−Casp8+/+ or Ripk3−/−Casp8−/− mice into 6–10-week-old recipient C57BL6/J mice that were lethally irradiated with 10 Gy. Experiments were then performed ≥5 weeks after transplantation.
For human CD34 transplantation, 100,000–200,000 cells were transplanted 1–2 days after electroporation through tail-vein injection into 6-10-week-old recipient NBSGW mice. Engraftment was monitored by bone-marrow aspiration at serial timepoints after transplantation, beginning at 4 weeks. In brief, the mice were anaesthetized with isoflurane and provided meloxicam for analgesia, and the leg was disinfected with three sets of alternating scrubs with betadine/povidine iodine solution and alcohol pads. A wetted 0.1 ml Tuberculin Syringe and needle (0.5 cm3, 27.5 G needle) was inserted into the shaft of the needle. The needle plunger was pulled back gently to aspirate roughly 5 μl of bone marrow. Successful aspiration was confirmed visually by a string of bone marrow cells followed by the appearance of blood in the top of the needle in the base of the syringe. After completion of the procedure, the mice were observed until they could maintain sternal recumbency.
Plasmids and base editor cloning
A pUC19-derived in vitro transcription (IVT)-compatible backbone generated by Gibson cloning included (1) a T7 promoter modified to be compatible with the CleanCap co-transcriptional mRNA capping strategy; (2) a human alpha-globin derived 5′ UTR; (3) a AES-mtRNR1 fusion 3′ UTR; (4) a P2A-mScarlet reporter; (5) a 120 nucleotide poly(A) sequence with an intervening G nucleotide to minimize recombination; and (6) type II restriction site directly downstream of the poly(A) to create a clean poly(A)-terminated linear template62,63. ABE8e-SpRY, ABE8e-NG,and ABE8e-NGG were subsequently cloned into this backbone by Gibson cloning.
In vitro mRNA transcription
The IVT base-editor template was linearized with BbsI and purified through phenol–chloroform extraction and ethanol precipitation. The IVT reaction was performed on the linearized template for 2 h at 37 °C using N1-methylpseudouridine-5′-triphosphate and CleanCap AG co-transcriptional capping (TriLink). The DNA template was then digested away with TURBO DNase (Thermo Fisher Scientific). mRNA was then purified using the PureLink RNA mini columns (Thermo Fisher Scientific). RNA was quantified and quality control was performed by running the RNA on a glyoxal gel to confirm the appropriate size and lack of significant RNA degradation (Thermo Fisher Scientific, NorthernMax). Alternatively, mRNA encoding ABE8e-SpRY was purchased from TriLink.
Flow cytometry
Flow cytometry was performed as previously described45. In brief, after preparing single-cell suspensions, cells were stained in PBS containing 2% heat-inactivated FBS and 1 mM EDTA at 4 °C. The following fluorophore-conjugated anti-mouse antibodies were used: CD11b-BUV395 (M1/70, BD Biosciences, 563553), CD19-BUV737 (1D3, eBioscience, 367-0193-82), CD117-BV650 (ACK2, BioLegend, 135125), CD150-BV785 (TC15-12F12.2, BioLegend, 115937), CD48-FITC (HM48-1, eBioscience, 11-0481-85), SCA-1-PE-Cy7 (D7, BioLegend, 108114), F4/80-APC (BM8, BioLegend, 123116). Lineage cocktail includes (all linked to APC-Cy7): CD3 (17A2, BioLegend, 100222), CD19 (6D5, BioLegend, 115530), Ter-119 (TER119, BioLegend, 116223), Gr1 (RB6-8C5, BioLegend, 108424), FCeR1a (MAR-1, BioLegend, 134325), NK1.1 (PK136, BioLegend, 108724), CD11b (M1/70, BioLegend, 101226), CD16/32-FITC (101305), KIT-PE (BioLegend, 135105), TPOr (primary antibody, Thermo Fisher Scientific, BS-11311R, donkey anti-rabbit IgG (H+L) highly cross-adsorbed secondary antibody, Alexa Fluor Plus 647 (Thermo Fisher Scientific, A32795), CD45-BUV395 (BD Biosciences, 564279). The following anti-human antibodies were used: CD45-FITC (304038), CD19-PE-Cy5 (BioLegend, 302209), CD33-BUV805 (BD Biosciences, 749018). Antibodies were used for flow cytometry at a 1:200 dilution. Samples were acquired on a Cytek Aurora (5-laser) or Attune NxT (Thermo Fisher Scientific) and analysed using FlowJo software v.9.9.6 or v.10.10.
Lentivirus production and infection
The Uba1a WT cDNA sequence was cloned into a lentiviral backbone upstream of a P2A-mNeonGreen sequence. mNeonGreen only (empty) and Uba1a-P2A-mNeonGreen lentivirus were generated in HEK293T cells by transfection with packaging plasmids pMD2.G and psPax2 using Lipofectamine 3000 reagent (Thermo Fisher Scientific). Lentivirus was collected at 24 and 48 h after transfection, concentrated by ultracentrifugation and spinfected onto THP-1 cells 800g for 60 min. mNeonGreen+ cells were sorted 96 h after infection and expanded before base editing of endogenous UBA1.
Western blotting
For western blotting, cells were lysed in 20 mM Tris HCl pH 7.4, 135 mM NaCl, 1.5 mM MgCl2, 1 mM EGTA, 1% (v/v) Triton X-100, 10% (v/v) glycerol and Halt protease and phosphatase inhibitor cocktail, EDTA-free (Thermo Fisher Scientific, 78447). Insoluble material was removed by centrifugation at 4 °C before addition of LDS sample buffer followed by incubation at 70 °C for at least 10 min. Alternatively, for analysis of cell death programs by western blot, cells were lysed in 20 mM Tris HCl pH 7.4, 135 mM NaCl, 1% (w/v) SDS, Halt protease and phosphatase inhibitor cocktail and benzonase endonuclease (Thermo Fisher Scientific, 70-746-3). Cells were lysed by sonicating for 10 min before addition of LDS sample buffer followed by incubation at 70 °C for at least 10 min. The following primary antibodies were used for western blotting: UBA1a/b (Cell Signaling Technologies, 4891; and Proteintech, 67198-1-Ig), β-actin (Cell Signaling Technologies, 13E5, 5125), RIPK1 (Cell Signaling Technologies, D94C12, 3493), phospho-p105 Ser932 (Cell Signaling Technologies, 18E6, 4806), phospho-IκBα Ser 32 (Cell Signaling Technologies, 14D4, 2859), IκBα (Cell Signaling Technologies, 9242), phospho-IKKα/β (Cell Signaling Technologies, 16A6, 2697), phospho-p38 MAPK Thr180/Tyr182 (Cell Signaling Technologies, 9211), p38 MAPK (D13E1), XP rabbit monoclonal antibody (Cell Signaling Technologies, 8690), phospho-SAPK/JNK (Thr183/Tyr185) (Cell Signaling Technologies, 81E11, 4668), ubiquitin (Cell Signaling Technologies, E6K4Y, 20326), GAPDH (Cell Signaling Technologies, D16H11, 8884), cleaved CASP8 Asp387 (Cell Signaling Technologies, D5B2, 8592), cleaved CASP3 Asp175 (Cell Signaling Technologies, 5A1E, 9664), cleaved CASP7 Asp198 (Cell Signaling Technologies, 9491), RIPK3 (Cell Signaling Technologies, D8J3L, 15828), phospho-RIPK1 Ser166/Thr169 (Genentech, GEN175-DP-A1), linear ubiquitin (Genentech, 1F11/3F5/Y102L), TRADD (Genentech, GN-21-3), HOIP (Genentech, 11D6H2G5), phospho-RIPK3 Thr231/Ser232 (Genentech, GEN-135-35-9), IKKγ/NEMO (Abcam, ab178872), IRAK1 (Cell Signaling Technologies, D51G7, 4504S), CASP8 (Enzo, 1G12, ALX-804-447), CASP3 (Cell Signaling Technologies, 9662), CASP7 (Cell Signaling Technologies, 9492), LC3B (Cell Signaling Technologies, 27755), ATF4 (Cell Signaling Technologies, 11815), CHOP (Cell Signaling Technologies, 5554), XBP-1s (Cell Signaling Technologies, 82914), PERK (Cell Signaling Technologies, 3192), BiP (Cell Signaling Technologies, 3177), p62 (Abnova, H00008878-M01), GABARAP (Abcam, ab109364), TNFR1 (Cell Signaling Technologies, 13377S), SHARPIN (Proteintech, 14626-1-AP), IKKβ (Cell Signaling Technologies, 8943S), K48 ubiquitin (Genentech), Lys63 polyubiquitin (Abcam, ab271929), anti-MLKL (phospho S345) antibody (Abcam, (EPR9515(2)), ab196436) and MLKL (Sigma-Aldrich, 6A2852). Western blot source data are provided in Supplementary Fig. 1.
Immunoprecipitation and deubiquitinating enzyme reactions
For the K48 polyubiquitin TUBE (tandem-repeated ubiquitin-binding entity) experiment, 7.5 × 106 BMDMs were plated the day before onto 15 cm tissue-culture-treated plates. The next day, cells were treated with 20 μM MG132 for 1 h before stimulation with 100 ng ml−1 recombinant mouse TNF (Peprotech) for the indicated timepoints. Plates were washed with prechilled PBS and collected by cell scraping. Cells were centrifuged at 400g for 5 min at 4 °C. Cell pellets were then resuspended in 400 µl Triton lysis buffer (20 mM Tris HCl pH 7.4, 135 mM NaCl, 1.5 mM MgCl2, 1 mM EGTA, 1% (v/v) Triton X-100, 10% (v/v) glycerol) containing complete protease and phosphatase inhibitor cocktail, 10 mM N-ethylmaleimide (Thermo Fisher Scientific), 20 μM MG132 (MedChemExpress, HY-13259), 20 μM PR-619 (Lifesensors, SI1619) and 5 mM 1,10-phenanthroline, monohydrate (Life Sensors, SI1649) and lysed for 15 min on ice followed by centrifugation at 20,000g for 10 min. 7.5% of input was set aside, and 25 µl of pre-washed magnetic K48 ubiquitin TUBE beads (Lifesensors, UM407M) were added to the remaining lysate. Beads were incubated by rotation with lysate for 4 h at 4 °C followed by washing three times with lysis buffer. Immunoprecipitates were then eluted in 2× loading dye at room temperature for 15 min followed by immunoblotting.
For TNF–Flag immunoprecipitation (IP), 7.5 × 106 BMDMs were plated the day before onto 15 cm tissue-culture-treated plates and the cells were stimulated for the indicated lengths of time with 250 ng ml−1 of recombinant Flag–TNF (Enzo Life Sciences). Cells were then washed with cold PBS, scraped and centrifuged at 400g for 5 min at 4 °C. Cell pellets were then resuspended in 400 μl Triton lysis buffer (20 mM Tris HCl pH 7.4, 135 mM NaCl, 1.5 mM MgCl2, 1 mM EGTA, 1% (v/v) Triton X-100, 10% (v/v) glycerol and complete protease and phosphatase inhibitor cocktail, 10 mM) containing 10 mM N-ethylmaleimide and lysed for 10 min on ice before removal of insoluble fraction by centrifugation at 20,000g for 10 min. Then 100 μl of prewashed anti-Flag M2 magnetic beads (Millipore Sigma) was added to each sample and rotated at 4 °C for 4 h. The beads were then washed four times with lysis buffer and immunoprecipitates were eluted with 2× loading dye, heated at 70 °C for 5 min and then analysed by immunoblotting.
To perform a deubiquitinating enzyme reaction on the complex I immunoprecipitate, beads were resuspended in 50 mM Tris HCl pH 7.5 buffer with 50 mM NaCl and 5 mM dithiothreitol (Thermo Fisher Scientific). They were treated with AMSH (2 μM), OTULIN (1 μM) and USP21 (1 μM) at 37 °C in the presence of 100 U of λPPase (NEB) and MnCl2 (1 mM). After 1 h incubation, the samples were denatured at 100 °C for 10 min and were further treated with 500 units of PNGase F (NEB) for 1 h at 37 °C. The reaction was stopped by adding 4× LDS loading dye.
For denaturing Lys63-polyubiquitin-specific immunoprecipitations, 7.5 × 106 BMDMs were plated the day before onto 15 cm tissue-culture-treated plates and stimulated with 100 ng ml−1 recombinant murine TNF (Peprotech) for 0 or 5 min. The plates were washed with cold PBS and collected by cell scraping. Cells were centrifuged at 400g for 5 min, and cell pellets were directly lysed for 15 min at room temperature in denaturing lysis buffer: 6 M urea, 135 mM NaCl, 50 mM Tris pH 7.5, 1.5 mM MgCl2, 1 mM EGTA, 1% Triton X-100, 10 mM 1× protease/phosphatase inhibitor (Thermo Fisher Scientific) and 10 mM N-ethylmaleimide. Lysates were cleared by centrifugation at 20,000g for 10 min. 6.25% input was collected, and the remaining lysate was diluted to 3 M urea and then precleared with 20 μl of protein A/G agarose beads (Pierce), rotating at 4 °C for 30 min. Beads were removed and 5 μg of Lys63-polyubiquitin-specific antibody (Genentech, Apu3.A8) was added to each lysate and incubated by rotation at 4 °C overnight. The next morning, 20 μl of washed protein A/G beads was added to each sample and incubated by rotation for 3 h. The samples were then transferred to new Eppendorf tubes, washed four times with 1 ml of urea lysis buffer (diluted to 3 M) and eluted with 25 μl of 2× SDS at 70 °C for 10 min.
For the linear- and Lys63-specific polyubiquitin TUBE experiments, 7.5 × 106 BMDMs were plated the day before onto 15 cm tissue-culture-treated plates and stimulated with 100 ng ml−1 recombinant mouse TNF (Peprotech) for the indicated timepoints. Plates were washed with prechilled PBS and collected by cell scraping. Cells were centrifuged at 400g for 5 min at 4 °C. The cell pellets were then resuspended in 400 µl Triton lysis buffer (20 mM Tris HCl pH 7.4, 135 mM NaCl, 1.5 mM MgCl2, 1 mM EGTA, 1% (v/v) Triton X-100, 10% (v/v) glycerol) containing complete protease and phosphatase inhibitor cocktail and 10 mM N-ethylmaleimide and lysed for 15 min on ice followed by centrifugation at 20,000g for 10 min. 7.5% of input was set aside, and 25 µl of prewashed magnetic linear-polyubiquitin-specific (Lifesensors, UM406M) or Lys63-polyubiquitin-specific (Lifesensors, UM404M) TUBE beads were added to the remaining lysate. The beads were incubated by rotation with lysate for 4 h at 4 °C followed by washing three times with lysis buffer. The immunoprecipitates were then eluted in 2× loading dye at room temperature for 15 min followed by immunoblotting.
Cell death analyses
At 8 days after electroporation, BMDMs were seeded at 40,000 cells per well in 96-well plates and treated the next day with the stimuli indicated in the figures in BMDM medium containing 500 nM YOYO-1 iodide (Invitrogen, Y3601). The IncuCyte SX5 system (Sartorius) acquired three images per well every hour for 24–48 h at 37 °C and 5% CO2. YOYO-1+ cells (dead cells) and total cell counts were quantified from resultant images by the IncuCyte analysis software. Percentages of dead cells for each well at each timepoint were calculated by dividing the YOYO-1+ cell count by total cell counts, averaging the values derived from the three images and multiplying by 100.
On/off-target base-editing analysis
Genomic DNA was isolated from BMDMs, human monocyte-derived macrophages or HSPCs 5–9 days after electroporation using the DNeasy Blood & Tissue Kit (Qiagen) according to the manufacturer’s instructions. PCR was then performed using Q5 polymerase (NEB) for 30 cycles to amplify an approximately 250 bp region surrounding the target mutation in Uba1 (Extended Data Table 1) or the set of highest-likelihood off-target sites as determined by CHOPCHOP64. Uba1-specific amplicons were subjected to Sanger sequencing by Azenta. For a quantitative measure of gene editing through next-generation sequencing, overhangs were included onto locus-specific primers (PCR1) to prime a second PCR step (PCR2) that appends illumina sequencing adapters onto the 5′ and 3′ ends, as well as i5 and i7 indices. Amplicons were size-selected using 1× (v/v) SPRIselect beads (Beckman Coulter), pooled and processed for high-throughput sequencing. Amplicon deep sequencing data were analysed using CRISPResso2 (v.2.2.14)27 in batch mode to quantify base-editing outcomes across on-target and off-target loci. Analyses were executed with the following parameters: --quantification_window_center -17, --quantification_window_size 10, --min_average_read_quality 30, --base_editor_output, --max_paired_end_reads_overlap 200, and --trim_sequences. The quantification window (±10 bp centred 17 bp upstream of the PAM) was defined to encompass the canonical base-editing window and adjacent nucleotides. For each amplicon, CRISPResso2 reported the frequency of modified reads—defined as reads containing nucleotide substitutions, insertions or deletions within this window—relative to the total aligned reads. For estimation of editing frequency by Sanger sequencing, EditR software was used.
Uba1 ∆Spl/y splice-site analysis
Genomic DNA was isolated from BMDMs 5 days after electroporation and PCR was performed using primers that amplified an approximately 250 bp region surrounding the target site (Extended Data Table 1). To assess downstream effects on splicing, RNA was simultaneously isolated from BMDMs using the Qiagen RNeasy Plus mini kit (74134). cDNA was generated using AffinityScript cDNA synthesis kit (Agilent, 600559), and PCR was performed using forward and reverse primers targeting exon 1 and exon 5 of the mature Uba1 transcript (Extended Data Table 1). PCR (DNA) and RT–PCR (RNA) products were purified and sent for PlasmidSaurus Premium PCR sequencing. Fastq files were then analysed using CRISPResso227 to identify the frequency of different deletion lengths. Quantification window was set to 30, centre of quantification window was set to −3. The deletion histogram output of CRISPResso2 was used to quantify the relative frequency of different deletion events within the Uba1 RT–PCR product.
Cell cycle analysis
Cell cycle profiling was performed using the Click-iT EdU Alexa Fluor 647 Flow Cytometry Assay Kit (Invitrogen). Briefly, mouse HSPCs in culture were pulsed for 2 h with 10 µM 5-ethynyl-2′-deoxyuridine. Approximately 1 × 105 cells were then washed, fixed, permeabilized and subjected to a Click-iT reaction (according to the manufacturer’s protocol) for covalent addition of an Alexa Fluor 647 azide. Cells were co-stained with PI, and the percentage of G1-, S- and G2/M-phase cells was measured by flow cytometry analysis.
LPS and TNF in vivo experiments
Two hours before injection of TNF or LPS, 6–10-week-old mice were treated with either vehicle or TAK-243 (3 mg per kg, i.p.). Mice were then treated with recombinant LPS (E. coli O55:B5, L2880-10MG, i.p., 5 mg per kg) or TNF (200 μg per kg, i.v., Thermo Fisher Scientific, 315-01A-1MG). Rectal temperatures were taken before injection of TNF or LPS (0 h) and then at 3 h, 6 h, 24 h, 48 h and up to 72 h. Mice with a temperature of ≤25 °C or with significant morbidity were euthanized. For Nec-1s rescue studies, Nec-1s (6 mg per kg, Cell Signaling Technology, 17802) was administered 15–30 min before TNF i.v. injection. Male mice were used for all experiments except for Extended Data Fig. 4f, in which a mix of male and female mice were used, evenly distributed by gender across genotypes.
Inflammasome assay
BMDMs were primed with 1 µg ml−1 Pam3CSK4 (Invivogen, tlrl-pms) for 5 h in 96-well tissue culture plates containing 40,000–100,000 cells per well. Primed cells were stimulated as indicated with 10 μg ml−1 nigericin (Invivogen, tlrl-nig) for 1 h, 5 mM ATP (Sigma-Aldrich, A2383) for 4 h, 0.5 μg ml−1 flagellin (Invivogen, tlrl-pafla) with FugeneHD (Promega, E2311), 1 μg ml−1 LFn-Rod (Invivogen, tlrl-rod) with 1 μg ml−1 Anthrax protective antigen (Millipore, 176905) or 2 μg ml−1 PolydA:dT (Invivogen, tlrl-patn) with Lipofectamine2000 (Invitrogen) overnight. Lumit IL-1β Human/Mouse Immunoassay (Promega) measured IL-1β in culture supernatants and CytoTox 96 Non-Radioactive Cytotoxicity Assay (Promega) measured LDH release. These data were acquired using Varioskan LUX (Thermo Fisher Scientific).
DAMP measurement
For cytokine secretion assays, BMDMs were stimulated as indicated with 100 ng ml−1 recombinant mouse TNF (Peprotech), LPS (100 ng ml−1) and IFNγ (100 ng ml−1, Peprotech) for 24 h in 96-well tissue culture plates containing 40,000–100,000 cells per well. IL-1α, IL-1β and IL-18 were measured in culture supernatants using the Mouse ProcartaPlex Simplex Kit (PPX-03-MXPRNM6; Thermo Fisher Scientific) on the Luminex xMAP INTELLIFLEX System (APX2020, Thermo Fisher Scientific) according to the manufacturer’s protocol.
scRNA-seq
Ripk3+/+Casp8+/+, Ripk3−/−Casp8+/+ and Ripk3−/−Casp8−/− HSPCs were isolated and cultured for 8 days before electroporation in triplicate with sgRosa26 or sgM41T sgRNAs along with ABE8e-SpRY mRNA. Cells from each of the corresponding six genotypes were collected on either day 5 or day 8 after electroporation. Cells were blocked with TruStain FcX (anti-mouse CD16/32) antibody (BioLegend, 101319) for 10 min and then stained per genotype with TotalSeq-B hashtag antibodies (BioLegend, B0301-B0306) for 30 min at 4 °C. Cells were then washed four times, counted and pooled before submission for 3′ scRNA-seq. Single-cell suspensions were stained with Trypan blue and the Countess II Automated Cell Counter (Thermo Fisher Scientific) was used to assess both cell number and viability. After quality control, the samples were loaded onto Chromium GEM-X Single Cell 3′ v4 (10x Genomics, 1000690) system and GEM generation, cDNA synthesis, cDNA amplification and library preparation of 31,000–38,000 cells was performed using the Chromium GEM-X Single Cell 3′ Kit v4 (10x Genomics, 1000691) according to the manufacturer’s protocol. cDNA amplification included 11 cycles, and 10 µl of the material was used to prepare sequencing libraries with 10 cycles of PCR. Indexed libraries were pooled and sequenced on a NovaSeq X in a PE28/88 run using the NovaSeq X 25B Reagent Kit (Illumina). An average of 23 thousand paired reads was generated per cell. Amplification products generated using the methods described above included both cDNA and feature barcodes tagged with cell barcodes and unique molecular identifiers. Smaller feature barcode fragments were separated from longer amplified cDNA using a 0.6× cleanup using aMPure XP beads (Beckman Coulter, A63882). Libraries were constructed using the 3′ Feature Barcode Kit (10x Genomics, 1000276) according to the manufacturer’s protocol with ten cycles of PCR. Indexed libraries were pooled and sequenced on the NovaSeq X system in a PE28/88 run using the NovaSeq X 25B Reagent Kit (Illumina). An average of 224 million paired reads was generated per sample.
Bioinformatic analysis (scRNA-seq analysis, splicing analysis)
Raw sequencing data were processed using the CellRanger v.8.0.1 pipeline using the default parameters. Cell hashing data were quantified and demultiplexed using HTODemux in Seurat. Pseudobulk and single-cell genotyping of the consequence of the Uba1 base edit were performed using the pileup functionality in Rsamtools (as part of Bioconductor)65. All downstream gene expression analyses were performed using the Seurat v.5 ecosystem66. Specifically, cell filtering, dimensionality reduction and gene expression visualization were performed using the default Seurat parameters with two exceptions. First, to regress cell cycle, we used the CellCycleScoring function and used the annotated phase per cell as a covariate that was regressed out of the harmony principal component correction67. Second, we used a resolution parameter of 0.4 in the FindClusters function to yield archetypal cell type clusters and to mitigate over-clustering. Gene set analyses were performed using the AddModuleScore function in Seurat with the default parameters, using the same gene sets nominated from analyses of human patients with VEXAS42 after converting human genes to murine genes. We developed a gene module of 88 genes that were significantly upregulated or downregulated comparing VEXAS mutant to WT cells to compute the Ganesan VEXAS module by filtering the HSPC differentially expressed genes such that genes with a log[FC] > 2 and an adjusted P < 0.01 were used as the input for gene module scoring per cell42. Statistical analyses between the gene edits were computed using nonparametric two-sided Mann–Whitney U-tests.
Statistics and reproducibility
Data were analysed using Microsoft Excel and GraphPad Prism v.10.4.2 for statistical analysis. Methods for P value determination are included in the relevant figure legends. Unless otherwise specified in the figure legends, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Depicted error bars generally represent s.e.m. Figure legends indicate the statistical tests used. Investigators were not blinded to allocation during experiments and outcome assessment. Samples were grouped according to genotype or treatment, and therefore not randomized. Animals were age-matched in all experiments. No sample size calculations were performed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
High-throughput sequencing data have been uploaded to the Gene Expression Omnibus (GEO: GSE299813). Source data are provided with this paper.
Code availability
Custom code to reproduce scRNA-seq analyses in this manuscript is available at GitHub (https://github.com/clareaulab/vexas-scrna).
References
Beck, D. B. et al. Somatic mutations in UBA1 and severe adult-onset autoinflammatory disease. N. Engl. J. Med. 383, 2628–2638 (2020).
Beck, D. B. et al. Estimated prevalence and clinical manifestations of UBA1 variants associated with VEXAS syndrome in a clinical population. JAMA 329, 318–324 (2023).
Georgin-Lavialle, S. et al. Further characterization of clinical and laboratory features in VEXAS syndrome: large-scale analysis of a multicentre case series of 116 French patients. Br. J. Dermatol. 186, 564–574 (2022).
Oganesyan, A., Hakobyan, Y., Terrier, B., Georgin-Lavialle, S. & Mekinian, A. Looking beyond VEXAS: coexistence of undifferentiated systemic autoinflammatory disease and myelodysplastic syndrome. Semin. Hematol. 58, 247–253 (2021).
Obiorah, I. E. et al. Benign and malignant hematologic manifestations in patients with VEXAS syndrome due to somatic mutations in UBA1. Blood Adv. 5, 3203–3215 (2021).
Grayson, P. C., Patel, B. A. & Young, N. S. VEXAS syndrome. Blood 137, 3591–3594 (2021).
Bourbon, E. et al. Therapeutic options in VEXAS syndrome: insights from a retrospective series. Blood 137, 3682–3684 (2021).
Ferrada, M. A. et al. Translation of cytoplasmic UBA1 contributes to VEXAS syndrome pathogenesis. Blood 140, 1496–1506 (2022).
Templé, M. et al. Atypical splice-site mutations causing VEXAS syndrome. Rheumatology 60, e435–e437 (2021).
Poulter, J. A. et al. Novel somatic mutations in UBA1 as a cause of VEXAS syndrome. Blood 137, 3676–3681 (2021).
Damgaard, R. B. et al. The deubiquitinase OTULIN is an essential negative regulator of inflammation and autoimmunity. Cell 166, 1215–1230 (2016).
Boisson, B. et al. Immunodeficiency, autoinflammation and amylopectinosis in humans with inherited HOIL-1 and LUBAC deficiency. Nat. Immunol. 13, 1178–1186 (2012).
Heger, K. et al. OTULIN limits cell death and inflammation by deubiquitinating LUBAC. Nature 559, 120–124 (2018).
Boisson, B. et al. Human HOIP and LUBAC deficiency underlies autoinflammation, immunodeficiency, amylopectinosis, and lymphangiectasia. J. Exp. Med. 212, 939–951 (2015).
Zhou, Q. et al. Biallelic hypomorphic mutations in a linear deubiquitinase define otulipenia, an early-onset autoinflammatory disease. Proc. Natl Acad. Sci. USA 113, 10127–10132 (2016).
Cuchet-Lourenço, D. et al. Biallelic RIPK1 mutations in humans cause severe immunodeficiency, arthritis, and intestinal inflammation. Science 361, 810–813 (2018).
Zhou, Q. et al. Loss-of-function mutations in TNFAIP3 leading to A20 haploinsufficiency cause an early-onset autoinflammatory disease. Nat. Genet. 48, 67–73 (2016).
Oda, H. et al. Biallelic human SHARPIN loss of function induces autoinflammation and immunodeficiency. Nat. Immunol. 25, 764–777 (2024).
Lalaoui, N. et al. Mutations that prevent caspase cleavage of RIPK1 cause autoinflammatory disease. Nature 577, 103–108 (2020).
Tao, P. et al. A dominant autoinflammatory disease caused by non-cleavable variants of RIPK1. Nature 577, 109–114 (2020).
Peltzer, N. et al. HOIP deficiency causes embryonic lethality by aberrant TNFR1-mediated endothelial cell death. Cell Rep. 9, 153–165 (2014).
Rickard, J. A. et al. TNFR1-dependent cell death drives inflammation in Sharpin-deficient mice. eLife 3, e03464 (2014).
Gerlach, B. et al. Linear ubiquitination prevents inflammation and regulates immune signalling. Nature 471, 591–596 (2011).
Walton, R. T., Christie, K. A., Whittaker, M. N. & Kleinstiver, B. P. Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science 368, 290–296 (2020).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Varfolomeev, E. et al. IAP antagonists induce autoubiquitination of c-IAPs, NF-κB activation, and TNFα-dependent apoptosis. Cell 131, 669–681 (2007).
Vince, J. E. et al. IAP antagonists target cIAP1 to induce TNFα-dependent apoptosis. Cell 131, 682–693 (2007).
Bertrand, M. J. M. et al. cIAP1 and cIAP2 facilitate cancer cell survival by functioning as E3 ligases that promote RIP1 ubiquitination. Mol. Cell 30, 689–700 (2008).
Dondelinger, Y. et al. RIPK3 contributes to TNFR1-mediated RIPK1 kinase-dependent apoptosis in conditions of cIAP1/2 depletion or TAK1 kinase inhibition. Cell Death Differ. 20, 1381–1392 (2013).
Meylan, E. et al. RIP1 is an essential mediator of Toll-like receptor 3-induced NF-κB activation. Nat. Immunol. 5, 503–507 (2004).
He, S., Liang, Y., Shao, F. & Wang, X. Toll-like receptors activate programmed necrosis in macrophages through a receptor-interacting kinase-3-mediated pathway. Proc. Natl Acad. Sci. USA 108, 20054–20059 (2011).
Tazi, Y. et al. Unified classification and risk-stratification in acute myeloid leukemia. Nat. Commun. 13, 4622 (2022).
Bolton, K. L. et al. Cancer therapy shapes the fitness landscape of clonal hematopoiesis. Nat. Genet. 52, 1219–1226 (2020).
Bernard, E. et al. Molecular international prognostic scoring system for myelodysplastic syndromes. NEJM Evid. 1, EVIDoa2200008 (2022).
Kaiser, W. J. et al. RIP3 mediates the embryonic lethality of caspase-8-deficient mice. Nature 471, 368–372 (2011).
Oberst, A. et al. Catalytic activity of the caspase-8-FLIP(L) complex inhibits RIPK3-dependent necrosis. Nature 471, 363–367 (2011).
Alvarez-Diaz, S. et al. The pseudokinase MLKL and the kinase RIPK3 have distinct roles in autoimmune disease caused by loss of death-receptor-induced apoptosis. Immunity 45, 513–526 (2016).
Kaiser, W. J. et al. Toll-like receptor 3-mediated necrosis via TRIF, RIP3, and MLKL. J. Biol. Chem. 288, 31268–31279 (2013).
Kosmider, O. et al. VEXAS syndrome is characterized by inflammasome activation and monocyte dysregulation. Nat. Commun. 15, 910 (2024).
Ganesan, S. et al. Single-cell genotype-phenotype mapping identifies therapeutic vulnerabilities in VEXAS syndrome. Preprint at bioRxiv https://doi.org/10.1101/2024.05.19.594376 (2024).
Smith, A. L. et al. Discovery of 1H-pyrazol-3(2H)-ones as potent and selective inhibitors of protein kinase R-like endoplasmic reticulum kinase (PERK). J. Med. Chem. 58, 1426–1441 (2015).
Molteni, R. et al. Mechanisms of hematopoietic clonal dominance in VEXAS syndrome. Nat. Med. 31, 1911–1924 (2025).
Wilkinson, A. C. et al. Long-term ex vivo haematopoietic-stem-cell expansion allows nonconditioned transplantation. Nature 571, 117–121 (2019).
Pietras, E. M. et al. Chronic interleukin-1 exposure drives haematopoietic stem cells towards precocious myeloid differentiation at the expense of self-renewal. Nat. Cell Biol. 18, 607–618 (2016).
Shu, H. B., Takeuchi, M. & Goeddel, D. V. The tumor necrosis factor receptor 2 signal transducers TRAF2 and c-IAP1 are components of the tumor necrosis factor receptor 1 signaling complex. Proc. Natl Acad. Sci. USA 93, 13973–13978 (1996).
Ermolaeva, M. A. et al. Function of TRADD in tumor necrosis factor receptor 1 signaling and in TRIF-dependent inflammatory responses. Nat. Immunol. 9, 1037–1046 (2008).
Emmerich, C. H. et al. Lys63/Met1-hybrid ubiquitin chains are commonly formed during the activation of innate immune signalling. Biochem. Biophys. Res. Commun. 474, 452–461 (2016).
Emmerich, C. H. et al. Activation of the canonical IKK complex by K63/M1-linked hybrid ubiquitin chains. Proc. Natl Acad. Sci. USA 110, 15247–15252 (2013).
Haas, T. L. et al. Recruitment of the linear ubiquitin chain assembly complex stabilizes the TNF-R1 signaling complex and is required for TNF-mediated gene induction. Mol. Cell 36, 831–844 (2009).
Rahighi, S. et al. Specific recognition of linear ubiquitin chains by NEMO is important for NF-kappaB activation. Cell 136, 1098–1109 (2009).
Micheau, O. & Tschopp, J. Induction of TNF receptor I-mediated apoptosis via two sequential signaling complexes. Cell 114, 181–190 (2003).
Newton, K., Sun, X. & Dixit, V. M. Kinase RIP3 is dispensable for normal NF-kappa Bs, signaling by the B-cell and T-cell receptors, tumor necrosis factor receptor 1, and Toll-like receptors 2 and 4. Mol. Cell. Biol. 24, 1464–1469 (2004).
Wu, Z. et al. Early activation of inflammatory pathways in UBA1-mutated hematopoietic stem and progenitor cells in VEXAS. Cell Rep. Med. 4, 101160 (2023).
Avagyan, S. et al. Resistance to inflammation underlies enhanced fitness in clonal hematopoiesis. Science 374, 768–772 (2021).
Murphy, J. M. et al. The pseudokinase MLKL mediates necroptosis via a molecular switch mechanism. Immunity 39, 443–453 (2013).
Newton, K. et al. Activity of protein kinase RIPK3 determines whether cells die by necroptosis or apoptosis. Science 343, 1357–1360 (2014).
Pasparakis, M., Alexopoulou, L., Episkopou, V. & Kollias, G. Immune and inflammatory responses in TNF alpha-deficient mice: a critical requirement for TNF alpha in the formation of primary B cell follicles, follicular dendritic cell networks and germinal centers, and in the maturation of the humoral immune response. J. Exp. Med. 184, 1397–1411 (1996).
Peschon, J. J. et al. TNF receptor-deficient mice reveal divergent roles for p55 and p75 in several models of inflammation. J. Immunol. 160, 943–952 (1998).
Freund, E. C. et al. Efficient gene knockout in primary human and murine myeloid cells by non-viral delivery of CRISPR-Cas9. J. Exp. Med. 217, e20191692 (2020).
Trepotec, Z., Geiger, J., Plank, C., Aneja, M. K. & Rudolph, C. Segmented poly(A) tails significantly reduce recombination of plasmid DNA without affecting mRNA translation efficiency or half-life. RNA 25, 507–518 (2019).
Orlandini von Niessen, A. G. et al. Improving mRNA-based therapeutic gene delivery by expression-augmenting 3’ UTRs identified by cellular library screening. Mol. Ther. 27, 824–836 (2019).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Acknowledgements
We thank M. Glickman, A. Rudensky and O. Abdel-Wahab for advice and discussions; L. Ivashkiv and M. Huang for assistance with human monocyte culture; E. Vidal Rodriguez and F. Munoz for animal husbandry; and J. Pruneda for advice and reagents related to deubiquitylase enzyme assays. We acknowledge the use of the Integrated Genomics Operation Core, funded by the NCI Cancer Center Support Grant (CCSG, P30 CA08748), Cycle for Survival and the Marie-Josée and Henry R. Kravis Center for Molecular Oncology. S.W.L. is the Geoffrey Beene Chair of Cancer Biology and a Howard Hughes Medical Institute Investigator. Some schematics were created using BioRender and exported using a publication license. Funding was provided by a Burroughs Wellcome Fund Career Award for Medical Scientists (A.D.G.), Cycle for Survival Equinox Innovation Award (A.D.G.), the Ludwig Center at Memorial Sloan Kettering Cancer Center (A.D.G. and T. Das), a Josie Robertson Investigator Award (A.D.G.), NIH grant R01 AI187249 (A.D.G.), the Edward P. Evans Foundation Young Investigator Award (V.K.N.), the NCI K12 CA184746 Paul Calabresi Career Development Award (V.K.N.), the NCI Clinical Scholars Biomedical Research Training Program T32 CA009512 (V.K.N.), the American Society of Clinical Oncology Young Investigator Award (V.K.N.), NIH grant 1F31HL182193-01 (R.J.L.), the Center for Experimental Immuno-Oncology Scholars Program of MSK (Y.K.), the Anna Maria and Stephen Kellen Pre-Doctoral Fellowship in Cancer Immunology at MSK (J.K.M) and the NCI AML SPORE grant (P50 CA254838) (S.W.L., V.K.N. and A.D.G.).
Author information
Authors and Affiliations
Contributions
A.D.G. and V.K.N conceived, designed and planned the study. V.K.N., T. Das, T. Devine, J.K.M., L.J.W, P.M., R.J.L., K.C., M.T., Y.K. and A.E.C. performed experiments under the supervision of A.D.G.; V.K.N. and T. Devine were also supervised by S.W.L.; K.C. and M.T. were also supervised by E.d.S.; C.A.L. and V.K.N. performed bioinformatic analysis. A.D.G., V.K.N. and T. Das wrote the paper with edits and input from all of the other authors.
Corresponding author
Ethics declarations
Competing interests
A.D.G. is a consultant for Genentech. While not directly related to this manuscript, S.W.L. declares outside consultancy and equity for Oric Pharmaceuticals, Blueprint Medicines, Mirimus, Senecea Therapeutics, Faeth Therapeutics and PMV Pharmaceuticals; and outside consultancy (no equity) for Fate Therapeutics. C.A.L. is a consultant to Cartography Biosciences. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Peter van Galen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of base edited Uba1M41T/y cells.
a, CRISPResso227 analysis of Uba1 amplicon next-generation sequencing in BMDMs 5 days after electroporation with ABE8e-SpRY mRNA and sgM41T. Editing frequency is depicted at intended target nucleotide (M41T: ATG > ACG) as well as at a bystander edit position (T > C) immediately upstream of the Uba1 exon 3 splice acceptor site. b, CRISPResso2 analysis of Uba1 amplicon sequencing at allelic resolution27. c, Frequency of modified reads at 12 of the highest likelihood sgM41T off-target sites predicted by CHOPCHOP64. Sites were assessed by amplicon sequencing using primers flanking each off-target site. Amplicons were analysed by next-generation sequencing (n = 1-4 biological replicates analysed per genomic locus). d, Immunoblots of BMDMs at 9 or 12 days after electroporation for the indicated genotypes. e, Percent M41T editing frequency in BMDMs at days 5 and 9 post-electroporation (n = 4 biological replicates per condition). Bars in c represent mean values; dots represent biological replicates; error bars represent standard deviation (c). Statistical test in e: paired two-sided student’s t-test.
Extended Data Fig. 2 Uba1M41T/y BMDMs are sensitized to inflammatory cell death that is partially rescuable by RIPK1 kinase inhibition.
a, Representative images of YOYO-1+ (green) Uba1+/y or Uba1M41T/y BMDMs 24 h after treatment with indicated stimuli. Scale bar in images in a represents 100 μm. b-d, Cell death measured over 24 h after treatment with indicated stimuli in BMDMs of indicated genotypes, with or without treatment with RIPK1 kinase inhibitor Nec-1s (c), quantified as percent YOYO-1+ cells among total cells. e, Cell death of BMDMs electroporated with ABE8e-SpRY and indicated sgRNAs designed to generate the indicated mutations in Uba1, Tet2 or Dnmt3a. Representative sanger sequencing of targeted mutations in Uba1, Tet2 and Dnmt3a is depicted on the right. f, Cell death of Uba1M41T/y (sgM41T) or Uba1+/y BMDMs generated via electroporation with 3 different control sgRNAs (sgGFP, sgRosa26, sgTrac) and treated with indicated stimuli, quantified as percent YOYO-1+ cells among total cells. All experiments were performed 9 days after electroporation. Lines in b-f represent mean values of triplicate wells, with 3 images averaged per well for quantification; error bands represent SEM; results are representative of 2 or 3 independent experiments.
Extended Data Fig. 3 Human UBA1M41T myeloid cells and mouse Uba1M41V/y macrophages are prone to inflammatory cell death.
a, Sanger sequencing of UBA1 M41T edit in human monocyte-derived macrophages (left). Cell death measured over 48 h in human monocyte-derived macrophages of indicated genotypes treated with indicated stimuli (right) 9 days after electroporation. Death was quantified as percent YOYO-1+ cells among total cells. b, Immunoblots of THP-1 monocytes 5 days after electroporation with Abe8e-NGG and sgM41T. c, Cell death measured of human THP-1 monocytes of indicated genotypes treated with indicated stimuli 5 days after electroporation, quantified as percent YOYO-1+ cells among total cells. d, Cell death measured over 48 h in human THP-1 monocytes of indicated genotypes expressing a lentiviral vector encoding mNeonGreen alone (‘Empty’) or mNeonGreen linked via P2A sequence to an sgRNA-resistant wild-type form of Uba1 (Uba1WT-oe). Cells were treated as indicated 5 days after electroporation. Cell death was quantified as in c. P values calculated at 48 h after treatment using one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test. e, Representative sanger sequencing of Uba1 M41V edit in BMDMs 9 days after electroporation with ABE8e-NG base editor mRNA and sgM41V or control sgRNA. f, Immunoblots of indicated BMDMs. g, Cell death measured over 24-48 h in BMDMs of indicated genotypes treated with indicated stimuli 9 days after electroporation, quantified as percent YOYO-1+ cells among total cells. Lines in a, c, d, g represent mean values of triplicate wells, with 3 images taken per well for quantification; error bands represent SEM. Experiments are representative of 2 (a, d, g) or 3 (c) independent experiments.
Extended Data Fig. 4 Uba1 M41T mutation or partial inhibition of UBA1 with low-dose TAK-243 causes RIPK3 and Caspase-8 dependent inflammatory cell death.
a, Cell death measured over 48 h after treatment with indicated stimuli in BMDMs of indicated genotypes 9 days after electroporation, quantified as percent YOYO-1+ cells among total cells. b, Cell death analysis after treatment with indicated stimuli in BMDMs of indicated genotypes, in presence or absence of 1 μM TAK-243, quantified as percent YOYO-1+ cells among total cells. c, Immunoblots of BMDMs of indicated genotypes 9 days after electroporation. d, Viable BMDM cell yields 9 days after electroporation relative to Uba1+/y (sgRosa26 electroporated) controls of matched Mlkl-Casp8 genotypes. Bars represent mean; error bars represent SEM. e, Cell death analysis of BMDMs of indicated genotypes after treatment with indicated stimuli in presence or absence of 100 nM TAK-243, quantified as percent YOYO-1+ cells among total cells. Lines in a, b, e represent mean values of triplicate wells, with 3 images taken per well for quantification; error bands represent SEM; results are representative of 2 independent experiments. f, g, Rectal temperatures and survival of C57BL/6 mice of indicated genotypes after i.v. injection of TNF (200 μg/kg) (f) or i.p. injection of LPS (5 mg/kg) (g), pre-treated 2 h prior with either vehicle or TAK-243 (3 mg/kg i.p.). Lines represent mean rectal temperatures for the indicated number of mice per condition. In g, experiments were performed 5 weeks after 10 Gy irradiation and bone marrow transplantation. P values comparing survival curves in (f-g) were calculated using the Log-rank (Mantel-Cox) test. Error bars in rectal temperature curves in (f-g) indicate SEM.
Extended Data Fig. 5 UPR and inflammasome signalling and inflammatory cell death of Uba1M41T/y BMDMs.
a, b LDH (a) and IL-1\({\rm{\beta }}\) (b) release from Pam3CSK4-primed BMDMs of indicated genotypes treated with indicated agonists of NLRP3, NLRC4 and AIM2 inflammasomes. Treatment was performed 9 days after electroporation. Bars in a, b represent mean values; error bars represent SEM. Each dot represents a biological replicate (n = 3). P values calculated using unpaired, two-sided t-test. c, Cell death measured over 48 h in BMDMs of indicated genotypes treated with indicated stimuli 9 days after electroporation. d-f, Immunoblots of BMDMs of indicated genotypes treated with Bafilomycin A1 (‘Baf’, 50 nM), Tunicamycin (‘Tm’, 5 μg/mL), and/or PERK inhibitor AMG’44 (10 μM) for indicated lengths of time. g, Cell death measured over 48 h in BMDMs of indicated genotypes and treated with indicated stimuli in presence or absence of AMG’44 (10 μM). h, Immunoblots of BMDMs of indicated genotypes treated with TNF alone or in combination with Birinapant (IAPi) for indicated lengths of time or Tunicamycin for 6 h. All assays were performed 9 days after electroporation. Lines in c, g represent mean values of triplicate wells, with 3 images taken per well for quantification; error bands represent SEM. Results are representative of 2 (c, h) or 3 (a, b, d, e, f, g) independent experiments.
Extended Data Fig. 6 Human CD34+ HSPC-derived VEXAS model recapitulates features of human disease.
a, Schema for engineering UBA1M41T/y HSPCs and characterizing their effect on hematopoiesis via transplantation into NBSGW recipient mice. Created in BioRender. Gitlin, A. (2025) https://BioRender.com/2vd7lid. b, Wright-Giemsa stain of human HSPCs 6 days after electroporation and quantification of vacuolated cells (red arrows) per high power field (HPF) (n = 6-7 HPF per genotype, counted manually). c, Immunoblots of human HSPCs of indicated genotypes 6 days after electroporation. d, Crispresso2-based27 quantification of % M41T alleles detected by amplicon sequencing of a region surrounding Met41 in UBA1 from genomic DNA of human CD45+ cells sorted from bone marrow of NBSGW mice 4 weeks after transplantation with HSPCs of the indicated genotypes (n = 3 mice per genotype). Each dot represents genotyping quantification from an individual mouse. Bars in b and d represent mean; error bars represent SEM. P value in b was calculated using unpaired, two-sided t-test. Results in b and c are representative of 2 independent experiments.
Extended Data Fig. 7 Effects of Uba1 mutation on HSPCs and DAMPs.
a, Immunoblots of Uba1+/y or Uba1M41T/y HSPCs 5 days after electroporation. b, Frequency of indicated cell populations (L: Lineage−, S: Sca-1+, K: c-KIT+, Gr-1: Ly-6G and/or Ly-6C) among HSPCs 6 days after electroporation with ABE8e-SpRY plus sgGFP (Uba1+/y) or sgM41T (Uba1M41T/y) (n = 5 biological replicates per genotype). ****P < 0.0001. c, Representative flow cytometry plot of CD11b and F4/80 expression 6 days after electroporation of Uba1+/y or Uba1M41T/y HSPCs, % of cells in each quadrant is indicated. d, Wright-Giemsa stain of Uba1+/y or Uba1M41T/y HSPCs 8 days after electroporation. e, Immunoblots of Uba1+/y or Uba1M41T/y HSPCs 3 days after electroporation. f, Cell death measured over 24 h in HSPCs of indicated genotypes treated with indicated stimuli 4 days after electroporation. Lines represent mean values of triplicate wells, with 3 images taken per well for quantification; error bands represent SEM. g, Analysis of cell cycle (% of cells in S2, G, or M phase) by EdU incorporation vs Propidium iodide stain, viability, and M41T frequency (via Sanger sequencing) at the indicated timepoints after electroporation of HSPCs of the indicated genotypes (n = 4 biological replicates). h, IL-1α (n = 3), IL-1β (n = 3) and IL-18 (n = 6) release by Uba1+/y or Uba1M41T/y BMDMs treated with indicated stimuli for 24 h. i, j Representative flow cytometry plot (i) and quantification (j) of myeloid differentiation of HSPCs 96 h after treatment with IL-1α or IL-1β (n = 2 per condition). Dots in b, g, h, j represent biological replicates; bars represent mean values; error bars represent SEM. Results are representative of 3 (a-d) or 2 (e-j) independent experiments. P value for % M41T graph (right) in panel g was calculated using paired two-sided t-test. Otherwise, P values in b, g, h were calculated using unpaired, two-sided t-test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 8 Myeloid bias and transplant failure of Uba1M41T/y HSPCs is independent of RIPK1-RIPK3-Caspase-8 axis.
a, Plot depicting percentage of CD11b+ cells among ex vivo cultured WT (n = 5), Ripk3−/− (n = 3-4), and Ripk3−/− Casp8−/− (n = 2) HSPCs 6 days after electroporation. Each dot represents a biological replicate. b. Schema for murine HSPC transplantation experiments with indicated genotypes. Created in BioRender. Gitlin, A. (2025) https://BioRender.com/4r2l9ho. c, Plot depicts percent of Uba1-M41T edited cells among pre-transplant (pre-tx) Uba1M41T/y HSPCs and post-transplant (post-tx) CD11b-enriched cells from pooled peripheral blood isolated at 3- and 6-weeks post transplantation. Genotyping of n = 5 pooled mice at each timepoint is represented as a single dot. d, Genotyping of Uba1 Met41 from peripheral blood or bone marrow of transplanted mice at indicated timepoints. Each dot represents results from an individual mouse. e, Flow cytometric analysis of the peripheral blood of reconstituted mice at 18-weeks, confirming donor (CD45.2) engraftment. Each value was obtained from a single transplanted mouse (n = 3-5 per genotype). f, Left, CXCR4 mean fluorescence intensity (MFI) in mouse HSPCs of indicated genotypes, 5 days after electroporation (n = 2 biological replicates). Right, percent donor (CD45.2) cells identified within bone marrow isolated 24 h after transplantation into lethally irradiated CD45.1+ recipients (n = 5 mice per genotype). g, Schema for mixed bone marrow chimera experimental design. Created in BioRender. Gitlin, A. (2025) https://BioRender.com/svu0zql. h, Percent CD45.2+ (of total CD45 positive) cells measured by flow cytometry of the peripheral blood of mixed bone marrow chimeras of indicated genotypes at indicated timepoints, gated on subpopulations of hematopoietic cells as indicated (n = 5 mice per genotype). Bars in panels represent mean values; error bars represent SEM; dots represent biological replicates. P values calculated using unpaired two-sided t-test. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 9 Single-cell RNA-seq analysis of Uba1M41T/y HSPCs.
a, Reduced dimensionality projection and joint clustering of HSPCs from all genotypes and timepoints. Expression of representative genes from each major annotated cell type is presented. b, Annotation of cells by M41T status, as determined, where possible, directly from sequencing reads mapping to the 5’ end of Uba1. c, Gene set module scores, stratified by genotype and timepoint post electroporation. d, Percent M41T editing in ex vivo HSPC cultures quantified directly from scRNA-seq reads. e, Reduced dimensionality projection and joint clustering of HSPCs separated and coloured by indicated genotypes and timepoints. Statistical test: two-sided Mann Whitney Test. **P < 0.01, ***P < 2.2e-16. Boxplots: centre line, median; box limits, first and third quartiles; whiskers, 1.5× interquartile range. In panel d, dots represent mean values and error bars represent SEM. 3 biological replicates per genotype were pooled and then analysed for each timepoint.
Extended Data Fig. 10 Temporal and linkage-specific analysis of polyubiquitin chains in Uba1M41T/y BMDMs.
a, Immunoblots of Flag-immunoprecipitates and input lysates from Uba1+/y or Uba1M41T/y BMDMs treated with Flag-TNF for indicated lengths of time. b, Immunoblots of pulldowns using K48-polyubiquitin-specific TUBEs and input lysates from MG132-treated BMDMs of indicated genotypes stimulated with TNF for indicated lengths of time. c, Immunoblots of pulldowns using Lys63-polyubiquitin-specific TUBEs and input lysates from BMDMs of indicated genotypes treated with TNF for indicated length of time. d, Immunoblots of pulldowns using linear-polyubiquitin-specific TUBEs and input lysates from BMDMs of indicated genotypes treated with TNF for indicated length of time. e, Immunoblots of Flag-immunoprecipitates treated with Lys63-specific (AMSH), linear-specific (OTULIN), or pan (USP21) deubiquitylases and input lysates from Uba1+/y or Uba1M41T/y BMDMs treated with Flag-TNF for 5 min. All assays were performed 9 days after electroporation. Results are representative of 2-3 independent experiments.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–4. Supplementary Fig. 1: uncropped western blots. Supplementary Fig. 2: the gating strategy for human CD34 engraftment analysis. Supplementary Fig. 3: the gating strategy for assessment of mouse HSPC differentiation. Supplementary Fig. 4: the gating strategy to characterize haematopoietic reconstitution in transplanted mice.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Narendra, V.K., Das, T., Wierciszewski, L.J. et al. Independent mechanisms of inflammation and myeloid bias in VEXAS syndrome. Nature 649, 1273–1281 (2026). https://doi.org/10.1038/s41586-025-09815-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09815-0
This article is cited by
-
Unravelling the cellular mechanisms of VEXAS syndrome
Nature Reviews Rheumatology (2026)