Abstract
The multifaceted dysfunction of tumour-infiltrating T cells, including exhaustion and mitochondrial dysfunction, remains a major obstacle in cancer immunotherapy1,2,3,4,5,6. Transcriptomic and epigenomic regulation of T cell dysfunction have been extensively studied7,8,9, but the role of proteostasis in regulating these obstacles remains less defined. Here we combined computational analyses of atlases of T cell exhaustion and mitochondrial fitness with performed targeted in vivo CRISPR screens, which identified the E3 ubiquitin ligase KLHL6 as a dual-negative regulator of both T cell exhaustion and mitochondrial dysfunction. Mechanistically, KLHL6 expression promoted TOX poly-ubiquitination and subsequent proteasomal degradation, thereby attenuating the transition of progenitor exhausted T cells towards terminal exhaustion. Simultaneously, KLHL6 maintained mitochondrial fitness by constraining the excessive mitochondrial fission that occurs during chronic T cell receptor stimulation by means of post-translational regulation of the PGAM5–Drp1 axis. However, KLHL6 is naturally downregulated by T cell receptor ligation, mitigating its potentially beneficial ubiquitin ligase activities during exposure to chronic stimulation. Enforcing KLHL6 expression in T cells markedly improved efficacy and long-term persistence against tumours and during viral infections in vivo. These findings uncover KLHL6 as a multifunctional, clinically actionable target for cancer immunotherapy, and highlight the potential of modulating proteostasis and ubiquitin modification to improve immunotherapy.
Similar content being viewed by others
Main
Durable responses to T-cell-based cancer immunotherapies remain limited in a substantial fraction of patients10,11. A major barrier to efficacy is T cell dysfunction within the tumour microenvironment (TME), driven in part by exhaustion, mitochondrial dysfunction and various immunosuppressive factors12,13. Despite extensive transcriptomic and epigenomic insights into T cell dysfunction7,8,9, our understanding of how post-translational modifications affect proteostasis to contribute to regulating this multifaceted process is still limited. Ubiquitin ligases have important roles in controlling protein degradation and homeostasis14, thereby modulating diverse biological events, including T cell development, activation and trafficking15,16,17. The multifunctional nature of these ligases, which can target diverse substrates for degradation, offers unique opportunities to address many obstacles of anti-tumour T cell responses simultaneously. However, the fundamental roles of the diverse universe of ubiquitin ligases, particularly in anti-tumour T cell biology, remain incompletely understood, limiting potential modulation of ubiquitin ligase activity as a therapeutic strategy in cancer immunotherapy. In this study, we integrated computational analyses with targeted in vivo CRISPR screens, and identified the E3 ubiquitin ligase KLHL6 as a dual-negative regulator of T cell exhaustion and mitochondrial dysfunction during chronic antigen stimulation. Enforced expression of KLHL6 in adoptively transferred T cells improved anti-tumour and anti-viral efficacy. The findings highlight the crucial role of ubiquitin modifications in dictating T cell fate and function, and implicate KLHL6 as a promising clinically actionable multifunctional target to enhance cancer immunotherapy.
CRISPR screen for E3 ligases in T cells
To elucidate general mechanisms of T cell dysfunction, we analysed 136 bulk RNA sequencing (RNA-seq) samples of CD8+ T cells from eight independent infection and cancer studies to define the exhaustion transcriptomic landscape (Fig. 1a, Extended Data Fig. 1a and Supplementary Table 1). Information-theoretic analyses distilled transcriptional variation across thousands of genes into two core modules (modules 1 and 2), each comprising opposing gene sets (Extended Data Fig. 1a and Supplementary Table 2). These two modules recapitulated the global transcriptome of CD8+ T cells under chronic (tumour and/or infection) and acute (infection) conditions (Extended Data Fig. 1a), capturing the principal programs underlying exhaustion. Module 1 positively enriched genes were low in naive and memory T cells but high in early and late exhausted T (Tex) cells, whereas negatively enriched genes showed the inverse trend. Module 2 positively enriched genes were selectively elevated in late exhausted cells, and negatively enriched genes were higher in naive and early exhausted states (Extended Data Fig. 1a and Supplementary Table 2). Gene set enrichment analysis (GSEA) revealed functional distinctions: module 1 genes associated with chronic stimulation, exhaustion and differentiation from naive and/or memory states, whereas module 2 genes related to proliferation, self-renewal and progenitor exhausted T (Tpex) cell features (Extended Data Fig. 1b,c and Supplementary Table 3). Projecting time-course datasets of antigen-specific CD8+ T cells (acute, Listeria or chronic, LCMV-Clone 13/tumour) onto a two-module map confirmed module 1 effectively separated acute from chronic antigen exposure, and module 2 tracked early-to-late exhaustion progression8,18 (Fig. 1b,c). Projecting independent time-series transcriptomes of CD8+ T cells in acute and chronic LCMV infection19 onto the same map revealed a coherent temporal trajectory of CD8+ T cell differentiation, with early samples (day 6–8) from both infections clustering together, consistent with early precursor T cells adapting to both acute and chronic infection20,21 (Extended Data Fig. 1d,e). These findings support the biological relevance of the two gene modules in capturing dynamic trajectories of CD8+ T cell differentiation across several physiological contexts.
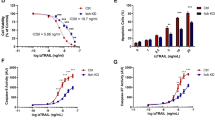
a, Schematic illustration of workflow for computational analysis of RNA-seq atlases of T cell exhaustion. b,c, Samples from two published datasets (GSE89307, GSE86881) were projected onto a two-dimensional map defined by computationally derived gene modules. b, Antigen-specific TCR-transgenic T cells collected across acute Listeria infection and tumour progression at matched time points following adoptive transfer. c, Antigen-specific naive and chronically exhausted T cells isolated during late-stage LCMV infection. d, GSEA enrichment of proteostasis-associated pathways in modules 1 and 2. e, Experimental schematic of CRISPR screen for E3 ligases that regulate T cell exhaustion and mitochondrial function. f,g, Rank plots (left) of gene-level enrichment scores in exhausted versus non-exhausted T cells (f) and in T cell populations with dysfunctional versus functional mitochondria (g). The CRISPR enrichment scores (log2 fold change of PD-1+TIM-3+ versus PD-1−TIM-3− or (MTDR/MTG)lo versus (MTDR/MTG)hi) were determined by comparing the indicated subsets for target genes from Cas9+sgRNA (mCherry)+ cells isolated from tumours on day 7 after ACT. The x axis shows targeted genes; the y axis shows the CRISPR enrichment score of each targeted gene; the dot colour represents false discovery rate (FDR). Distribution of several top-hit sgRNAs (right). Axis represents log2 fold change (FC). The histogram shows distribution of all sgRNAs. Red bars represent targeted sgRNAs, grey bars represent all other sgRNAs. h,i, Representative plots (left) and quantification (right) of the proportions of PD-1+TIM-3+ (h, n = 7 mice) and (MTDR/MTG)lo (i, n = 6 mice) TILs in sgCtrl-transduced or sgKlhl6-transduced Cas9+ OT-I T cells from B16-OVA tumours obtained on day 7 after ACT. Diagram in a created in Biorender. Li, G. (2025) https://BioRender.com/d5c767f. Diagram in e created in BioRender. Li, G. (2025) https://BioRender.com/dw0yfsp. Data are presented as mean ± s.e.m. Statistical analyses were determined by two-way ANOVA with Tukey’s multiple-comparisons test (h,i). *P < 0.05, ***P < 0.001 and ****P < 0.0001.
Notably, we observed significant enrichment of proteostasis-related pathways in modules 1 and 2, particularly the ubiquitin–proteasome system (UPS) (Fig. 1d and Supplementary Table 3). Given the limited understanding of proteostasis in T cell dysfunction, we focused on the UPS pathway. Hierarchical clustering of E3 ligase-related genes from OT-I T cells isolated from B16-OVA tumours identified two patterns, with one upregulated in single positive (SP, PD-1+TIM-3−) and double-positive (DP, PD-1+TIM-3+) tumour-infiltrating T cells (TILs) and the other downregulated (Extended Data Fig. 1f–h), underscoring potential roles of E3 ligase-mediated proteostasis in shaping exhausted T cell differentiation. Given the link between mitochondrial dysfunction and exhaustion under TME metabolic stress3,4,5,6, we analysed mitochondrial-associated genes and pathways in human and mouse TILs6,22, again revealing strong associations between UPS and mitochondrial function (Extended Data Fig. 1i,j). To prioritize functional candidates, we selected 78 E3 ligase-related genes negatively enriched in both exhaustion modules and 133 E3 ligase candidates positively associated with mitochondrial function for two parallel in vivo CRISPR screens targeting exhaustion and mitochondrial fitness (Extended Data Fig. 1k,l and Supplementary Table 4).
Using an established in vivo CRISPR-screening system23, single-guide RNA (sgRNA)-transduced Cas9+ OT-I T cells were transferred into B16-OVA tumour-bearing mice (Fig. 1e). Seven days after adoptive cell transfer (ACT), PD-1+TIM-3+ versus PD-1−TIM-3− cells (exhaustion screen) and (MitoTracker Deep Red (MTDR, mitochondrial membrane potential)/MitoTracker Green (MTG, mitochondrial mass))lo versus (MTDR/MTG)hi cells (mitochondrial dysfunctional versus functional screen) were isolated for sgRNA enrichment (Fig. 1e and Supplementary Table 5). Exhaustion screen identified positive and negative regulators including Klhl6, Traf6, Pcgf1, Trim45 and Rspry1 (Fig. 1f), with Traf6 known to promote CD8+ T cell memory and anti-tumour responses24,25 but underexplored in exhaustion. The mitochondrial screen revealed several established regulators such as Rnf34, Mkrn1 and Rnf7 (refs. 26,27,28), not previously studied in T cells, and new regulators such as Klhl6, Trim8 and Rspry1 (Fig. 1g). Integrating both screens, KLHL6 emerged as the top-ranked E3 ligase regulating both T cell exhaustion and mitochondrial fitness (Fig. 1f,g). In vivo validation with two Klhl6-targeting sgRNAs confirmed Klhl6 deletion reduced OT-I cell accumulation in the TME and increased frequencies of exhausted (PD-1+TIM-3+ or LAG-3+TIM-3+) and mitochondrially depolarized (MTDR/MTG)lo CD8+ TILs (Fig. 1h,i and Extended Data Fig. 2a–e). Further validation confirmed Pcgf1 or Traf6 deletion increased exhausted PD-1+TIM-3+CD8+ TILs, whereas Rspry1 knockout (KO) reduced exhaustion, consistent with screening results (Extended Data Fig. 2f). Phenotypic assessments of mitochondrial regulators aligned with screen results (Extended Data Fig. 2g). These findings strongly implicate KLHL6 as a critical regulator of both T cell exhaustion and mitochondrial function.
KLHL6 deficiency impairs T cell function
The ubiquitinase KLHL6, a Cullin3-RING E3 complex component, is known to regulate B cell differentiation29,30. To investigate its role in T cell immunity, we generated KLHL6-deficient (Klhl6−/−) mice and evaluated immunological characteristics (Extended Data Fig. 2h,i). As reported in ref. 30, KLHL6 deficiency impaired mature B cell formation but had no great effect on T cell development, peripheral homeostasis or activation, and only modestly reduced proliferation in vitro (Extended Data Fig. 2j–o). To assess the impact on anti-tumour T cell responses in a mouse B16-OVA melanoma model (Extended Data Fig. 2p), we crossed Klhl6−/− mice with OT-I mice (Klhl6−/− OT-I mice). By day 14, Klhl6−/− OT-I cells showed weaker tumour control than wild-type (WT) cells (Fig. 2a), with reduced accumulation in tumour, draining lymph node (dLN) and spleen (Fig. 2b and Extended Data Fig. 2q). Klhl6−/− CD8+ TILs showed increased PD-1+TIM-3+ and reduced PD-1−TIM-3− and PD-1+TIM-3− populations (Fig. 2c), elevated PD-1, TIM-3, LAG-3 and TOX expression (Extended Data Fig. 2r,s), and diminished tumour necrosis factor (TNF) and IFNγ production (Fig. 2d). Mice receiving Klhl6−/− OT-I cells had poorer survival (Extended Data Fig. 2t). To minimize inter-tumour variability, equal WT and Klhl6−/− OT-I T cells were cotransferred into the same tumour-bearing hosts (Fig. 2e). Klhl6−/− cells were consistently significantly decreased in tumour, dLN and spleen (Fig. 2f), and showed elevated PD-1, TIM-3 and TOX expression within tumours (Fig. 2g–i). KLHL6 deletion induced broad transcriptional changes (Extended Data Fig. 3a), including upregulation of exhaustion-related genes and downregulation of stemness-associated genes (Fig. 2j and Extended Data Fig. 3b). Gene Ontology analysis showed enriched oxidative stress-induced senescence, transforming growth factor-β (TGFβ) signalling and G2 cell-cycle arrest pathways in KLHL6-deficient cells (Extended Data Fig. 3c). GSEA revealed reduced memory and effector signatures but increased apoptosis and cell-cycle arrest signatures in Klhl6−/−CD8+ TILs (Extended Data Fig. 3d,e). Collectively, KLHL6 loss drove CD8+ T cells towards exhaustion and dysfunction.
a,b, CD45.2+ C57BL/6N mice were subcutaneously injected with B16-OVA melanoma cells and, 9 days later, treated by ACT with 3 × 106 CD45.1/2+ Klhl6+/+ (WT) or Klhl6−/− (KO) OT-I T cells (n = 14 mice). Tumour weights (a) and the numbers of transferred OT-I T cells (b) were assessed at day 14 post-ACT. c, Percentages of PD-1−TIM-3−, PD-1+TIM-3− and PD-1+TIM-3+ subsets among CD45.1/2+ TILs (n = 14 mice). d, Cytokine production (TNF and IFNγ) in transferred CD45.1/2+ TILs was determined (n = 14 mice). e–i, For cotransfer experiments, CD45.1/2+ WT and CD45.1+ KO OT-I T cells were mixed at a 1:1 ratio and then adoptively transferred into tumour-bearing CD45.2+ mice 9 days after tumour inoculation, and the mice were euthanized for analysis at day 14 after ACT (n = 7 mice). Experimental design (e), the proportions of WT and KO OT-I cells in tumour, dLN and spleen (f), the percentages of PD-1+TIM-3+ population (g) and the expression levels of PD-1 and TIM-3 (h) and TOX (i) in cotransferred WT and KO TILs. j, Heat map of exhaustion-associated genes in WT and KO TILs at day 14 post-ACT (n = 3 independent samples). k, GSEA plots of signatures of mitochondrial OXPHOS and mitochondrial respiratory chain complex assembly in WT versus KO TILs (n = 3 independent samples). GSEA was performed using a one-sided, permutation-based modified K–S test with adjustments for multiple comparisons. l–n, Cotransferred WT and KO TILs were sorted for Seahorse assays on day 14 post-ACT. OCR (l), SRC (m) and mitochondrial ATP production (n) were measured (n = 8 independent tests; 20 mice). o,p, The frequencies of (MTDR/MTG)lo subsets in transferred WT and KO OT-I TILs (o) and their distribution among exhausted (PD-1+TIM-3− and PD-1+TIM-3+) and non-exhausted (PD-1−TIM-3−) populations (p) at day 14 after ACT (n = 8 mice). Diagram in e created in BioRender. Li, G. (2025) https://BioRender.com/5se3g09. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (a,b,f–i,m–o) or two-way ANOVA with Sidak’s multiple-comparisons test (c,d,l,p). *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001; NS, not significant. FCCP, carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone; MFI, mean fluorescence intensity; NES, normalized enrichment score; R&A, rotenone and antimycin A.
GSEA also showed reduced OXPHOS and respiratory chain activity in Klhl6−/− T cells (Fig. 2k and Extended Data Fig. 3f). MTG and tetramethylrhodamine ethyl ester (TMRE) (membrane potential) staining revealed a markedly reduced TMRE/MTG ratio, indicating impaired mitochondrial activity (Extended Data Fig. 3g). Metabolic analyses demonstrated reduced oxygen consumption rate (OCR), spare respiratory capacity (SRC) and ATP production but elevated glycolytic proton efflux rate (glycoPER) in Klhl6−/− T cells (Extended Data Fig. 3h–k). Similarly, KLHL6-deficient TILs showed diminished mitochondrial function, increased glycoPER and a higher frequency of (MTDR/MTG)lo depolarized mitochondria compared with WT cells6,31 (Fig. 2l–o and Extended Data Fig. 3l–n). Exhausted TIL subsets (PD-1+TIM-3−, PD-1+TIM-3+) harboured more depolarized mitochondria than non-exhausted (PD-1−TIM-3−), and KLHL6 deficiency further exacerbated depolarization in exhausted TILs (Fig. 2p), underscoring a positive regulatory role maintaining mitochondrial fitness.
KLHL6 expression profiling of OT-I subpopulations from spleen and tumour (PD-1−TIM-3−, PD-1+TIM-3− and PD-1+TIM-3+) revealed Klhl6 expression highest in splenic T cells followed by PD-1−TIM-3− cells compared with the other tumour-reactive subsets (Extended Data Fig. 4a). T cell receptor (TCR) signalling downregulated KLHL6 protein expression in vitro (Extended Data Fig. 4b,c). Time-series analysis showed transient decline-recovery of KLHL6 expression following TCR stimulation but sustained repression on repeated stimulation (Extended Data Fig. 4d,e). Transcriptomic analysis of CD8+ T cells in acute and chronic infections or tumours8,19, confirmed persistent KLHL6 downregulation in chronic but transient reduction in acute contexts (Extended Data Fig. 4f,g). These findings were validated with an ex vivo time-course analysis of KLHL6 messenger RNA (mRNA) and protein levels during acute (Armstrong) and chronic (Clone 13) LCMV infections in virus-specific CD8+ P14 T cells (Extended Data Fig. 4h–j). Collectively, chronic TCR signalling resulted in sustained repression of KLHL6 expression, contributing to CD8+ T cell exhaustion and metabolic insufficiency.
KLHL6 limits Tpex terminal differentiation
As chronic TCR stimulation downregulates KLHL6 and promotes T cell exhaustion in the TME, we asked whether constitutive KLHL6 overexpression (KLHL6-OE) driven by a retroviral promoter could mitigate exhaustion, restore mitochondrial function and enhance T cell anti-tumour effects in a B16-OVA model (Extended Data Fig. 5a). Mice receiving KLHL6-OE OT-I T cells showed greater accumulation of transferred cells in tumours, spleens and dLNs at day 14 after ACT (Fig. 3a,b and Extended Data Fig. 5b). KLHL6-OE TILs showed increased proliferation, reduced apoptosis and elevated Bcl-2/Bim ratios (Extended Data Fig. 5c–f), consistent with enhanced intra-tumoural accumulation. The proportion and absolute number of PD-1−TIM-3− cells within KLHL6-OE OT-I TILs increased, whereas percentage of PD-1+TIM-3+ cells decreased despite an increased absolute count, suggesting enhanced survival and/or replenishment of exhausted cells (Extended Data Fig. 5g,h). Intra-tumoural KLHL6-OE TILs had reduced TOX, increased TCF-1 and enhanced cytokine production (Fig. 3c and Extended Data Fig. 5i,j). In spleen and dLN, KLHL6-OE expanded central memory-like (TCM-like: CD62L+CD44+) CD8+ T cells (Extended Data Fig. 5k). Tumour-infiltrating exhausted T cells are heterogeneous, including Tpex (TCF-1+TIM-3− or Ly108+TIM-3−) and terminally exhausted (Texterm: TCF-1−TIM-3+ or Ly108−TIM-3+) subsets9,32. Notably, KLHL6 overexpression increased both percentage and absolute number of Tpex (Ly108+TIM-3−) cells, while reducing percentage (but not number) of Texterm (Ly108−TIM-3+) cells, opposite to the pattern in KLHL6-deficient cells (Fig. 3d,e and Extended Data Fig. 5l,m). KLHL6-OE Tpex and Texterm cells showed lower levels of TOX and PD-1 compared with controls (Extended Data Fig. 5n,o), supporting a direct role for KLHL6 in restraining T cell exhaustion. KLHL6-OE TILs also showed enhanced mitochondrial fitness, with increased OCR, SRC and ATP production, reduced glycoPER and fewer (MTDR/MTG)lo depolarized mitochondria (Extended Data Fig. 5p–t), consistent with in vitro findings (Extended Data Fig. 5u–y). Correspondingly, tumour-bearing mice receiving KLHL6-OE T cells showed prolonged survival (Fig. 3f).
a–e, In total 3 × 106 Control or KLHL6-OE OT-I T cells were transferred into tumour-bearing CD45.2+ mice 9 days after tumour inoculation, and the mice were euthanized for analysis at day 14 after ACT (n = 7 mice). Percentages (a) and numbers (b) of transferred CD45.1+ OT-I TILs, cytokine production (TNF and IFNγ) (c), and frequencies (d) and numbers (e) of Tpex (Ly108+TIM-3−) and Texterm (Ly108−TIM-3+) subsets in Control and KLHL6-OE OT-I TILs were determined. f, Survival curves of tumour-bearing mice receiving Control or KLHL6-OE CD45.1+ OT-I T cells (n = 10 mice). Mice with tumour volumes greater than 1,500 mm3 were euthanized and this was defined as death. g, Transferred Control and KLHL6-OE OT-I TILs at day 14 post-ACT were subjected to scRNA-seq analysis. UMAP embedding showing clusters of all transferred CD8+ OT-I TILs (left) and their relative proportions (right); cluster annotations: 0, proliferation; 1, proinflammation; 2, exhaustion; 3, progenitor and 4, early activation. h, UMAP density plots comparing Control and KLHL6-OE TILs. i, Violin plots of Tpex and Texterm gene signatures; the boxplot spans from the first to the third quartile of the distribution, with the median positioned in the centre and whiskers representing the minimum and maximum values, excluding outliers. Values plotted represent cells from a single replicate. j, Dot plots showing expression of signature genes, with both colour and size indicating effect size. k–m, CD45.1+ Control or KLHL6-OE OT-I Tpex cells were transferred into CD45.2+ congenic mice that had been subcutaneously implanted with B16-OVA cells 2 days earlier, and the mice were euthanized for analysis on days 8 and 16 post-ACT. k, Experimental design. l,m, Representative plots (l) and percentages (m) of Ly108+TIM-3−, Ly108+TIM-3+ and Ly108−TIM-3+ subsets among transferred cells before and after transfer (n = 7 mice). Diagram in k created in BioRender. Li, G. (2025) https://BioRender.com/rxkc0bf. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (a,b), two-way ANOVA with Sidak’s multiple-comparisons test (c–e), two-way ANOVA with Tukey’s multiple-comparisons test (m) and log-rank (Mantel–Cox) test (f). *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
We next performed single-cell RNA-seq (scRNA-seq) on KLHL6-OE and control OT-I TILs at day 14 post-ACT. Uniform manifold approximation and projection (UMAP) projection resolved five subpopulations with distinct programs (Fig. 3g and Extended Data Fig. 6a,b), with cells from both groups present across the map but at differing densities (Fig. 3h). KLHL6-OE enriched for cluster 3 (Tpex) and reduced cluster 2 (Texterm) (Fig. 3g,h), concomitant with upregulation of Tpex signature and stem-like genes, and downregulation of Texterm signature and exhaustion-related genes (Fig. 3i,j and Extended Data Fig. 6c,d). Pathway enrichment analysis in KLHL6-OE TILs showed increased 4-1BB, IL-2 and TCR signalling, reduced replicative senescence and potassium channel activity, and enhanced mitochondrial programs including biogenesis, OXPHOS and ATP synthesis (Extended Data Fig. 6e).
To further elucidate the role of KLHL6 in the Tpex-to-Texterm transition, we isolated control or KLHL6-OE Tpex cells from tumours, transferred them into congenic mice implanted with B16-OVA tumours 2 days earlier, and tracked differentiation trajectory at days 8 and 16 post-transfer33 (Fig. 3k). KLHL6-OE restrained but did not completely block transition of Tpex to Texterm, increased TIL accumulation and improved tumour control compared with controls (Fig. 3l,m and Extended Data Fig. 6f,g). In a long-term cotransfer assay (days 21/28), KLHL6-OE OT-I cells persisted at higher proportions and numbers, expressed lower TIM-3 and higher Ki-67, showed greater polyfunctionality (IFNγ+TNF+IL-2+) and maintained elevated TCM or stem-like fractions (Extended Data Fig. 6h–n), suggestive of more durable anti-tumour immunity. Finally, in a human HepG2-NY-ESO xenograft model treated with NY-ESO-specific 1G4 TCR-T cells34 (Extended Data Fig. 6o), KLHL6-OE TCR-T cells again showed enhanced tumour control, greater accumulation in TME and blood, reduced LAG-3, TIM-3 and TOX expression, elevated cytokine production and enhanced splenic TCM formation (Extended Data Fig. 6p–t). Overall, enforced KLHL6 expression preserved Tpex self-renewal, restrained terminal exhaustion and improved durable anti-tumour responses in mouse and human models.
KLHL6 resists exhaustion in chronic infection
KLHL6 function during chronic viral infection was investigated using Klhl6−/− P14 mice that express a transgenic TCR for the LCMV DbGP33–41 epitope. Naive Klhl6−/− and WT P14 CD8+ T cells were mixed 1:1 and transferred into congenic recipients, which were then infected with acute LCMV-Arm or chronic LCMV-CL13. KLHL6 deficiency minimally affected early T cell activation in either setting (Extended Data Fig. 7a). However, during chronic LCMV-CL13 infection, Klhl6−/− T cells showed significant numerical reductions on days 8 and 28 post-infection (p.i.) (Extended Data Fig. 7b,c). At day 8 post-chronic infection, P14 cells typically differentiate into either terminal ‘effector-like’ cells (TCF-1−GzmB+ or Ly108−TIM-3+) or Tex precursors (Texprec: TCF-1+GzmB− or Ly108+TIM-3−)23,35. Compared with WT, Klhl6−/− cells produced proportionally more Texprec and fewer terminal effector-like cells, although absolute numbers of both subsets were decreased (Extended Data Fig. 7d,e). By day 8 p.i., KLHL6 deletion already impaired cytokine production, increased mitochondrial damage and enhanced apoptosis (Extended Data Fig. 7f–h). At day 28 p.i., KLHL6-deficient P14 cells showed elevated expression of exhaustion markers (for example, TIM-3 and TOX) and reduced TCF-1 (Extended Data Fig. 7i). KLHL6 deletion resulted in a decreased proportion of Ly108+CX3CR1− Tpex cells and an increased frequency of Ly108−CX3CR1− Texterm cells, whereas the Ly108−CX3CR1+ Texint population was only modestly affected (Extended Data Fig. 7j). Moreover, all KLHL6-deficient subsets showed decreased absolute numbers and lower Bcl-2/Bim ratios (Extended Data Fig. 7j,k). Similar skewing was observed using Ly108 and CD69 gating to define Tpex1, Tpex2, Texint and Texterm subsets36 (Extended Data Fig. 7l). Furthermore, KLHL6 deletion increased TOX expression in all four subsets, and markedly impaired P14 effector function and viral control (Extended Data Fig. 7m–o).
We ectopically expressed KLHL6 in WT P14 cells through retroviral transduction to assess protective effects (Extended Data Fig. 8a). KLHL6-OE P14 cells preferentially differentiated into effector-like cells rather than Texprec during chronic LCMV-CL13 infection, with increased total numbers and subset abundance on day 8 p.i. (Extended Data Fig. 8b–d). KLHL6-OE reduced apoptosis, enhanced cytokine production, decreased PD-1 and TIM-3 expression, and increased TCF-1 levels at day 21 p.i. (Extended Data Fig. 8e–g). Furthermore, enforced KLHL6 expression delayed differentiation of Tpex to Texterm (Extended Data Fig. 8h). All KLHL6-OE subsets showed numerical advantages and reduced TOX expression compared with respective controls (Extended Data Fig. 8i,j). Similar trends were observed using Ly108 and CX3CR1 markers (Extended Data Fig. 8k,l). Adoptive transfer of KLHL6-OE P14 T cells significantly improved viral control at day 15 p.i. (Extended Data Fig. 8m). Together, these findings demonstrate KLHL6 restrained terminal exhaustion, promoted exhausted T cell survival and enhanced protective immunity during chronic viral infection.
KLHL6 mediates TOX ubiquitination
To investigate how KLHL6 regulates anti-tumour responses and exhaustion, we performed E3-substrate tagging by ubiquitin biotinylation (E-STUB) with mass spectrometry37 and identified 82 putative substrates (log2 fold change greater than 1) (Fig. 4a, Extended Data Fig. 9a and Supplementary Table 7). TOX, a key transcription factor for T cell exhaustion7,38,39, was significantly enriched (Fig. 4a and Extended Data Fig. 9b,c). Reciprocal co-immunoprecipitation (Co-IP) assays confirmed KLHL6–TOX interactions in primary human T cells, Jurkat cells and EL4 cells (Fig. 4b and Extended Data Fig. 9d). Truncation analysis revealed strong binding between the TOX carboxy-terminal domain (amino acids 330–526) and KLHL6 (Extended Data Fig. 9e,f). Ectopic KLHL6 expression reduced TOX protein levels in a dose-dependent manner (Extended Data Fig. 9g). A cycloheximide chase assay to visualize protein degradation kinetics confirmed KLHL6 shortened the half-life of endogenous TOX (Fig. 4c). KLHL6 deficiency or overexpression significantly regulated TOX protein levels in both stimulated and unstimulated T cells (Fig. 4d and Extended Data Fig. 9h). KLHL6-mediated reduction of TOX was largely abrogated by the proteasomal inhibitor MG132 (Fig. 4e), suggesting KLHL6 promotes TOX degradation by means of the proteasome. Ubiquitination analysis demonstrated KLHL6 overexpression promoted poly-ubiquitination of TOX, whereas KLHL6 deficiency reduced it (Fig. 4f,g and Extended Data Fig. 9i,j). Moreover, TCR stimulation markedly reduced TOX ubiquitination in both mouse and human primary T cells, coinciding with reduced KLHL6 expression following TCR engagement (Extended Data Fig. 9k).
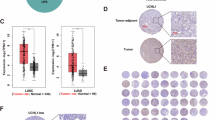
a, E-STUB assay and label-free mass spectrometry were used to identify KLHL6-proximal ubiquitylated substrates. Fold changes in protein abundance between KLHL6-BirA and Empty-BirA groups were calculated by two-sided moderated t-test (limma). b, Interaction between endogenous KLHL6 and TOX in human T cells. c, Cycloheximide (CHX) chase assay showing TOX degradation in Jurkat cells transduced with empty or KLHL6-Myc vectors (n = 3 independent samples). d, TOX levels in WT or Klhl6−/− OT-I T cells with or without anti-CD3 restimulation (n = 3 independent samples). e, Quantification of endogenous TOX protein in Jurkat cells transduced with an empty vector or KLHL6-Myc, with or without MG132 treatment (10 μM, 6 h; n = 3 independent samples). f,g, Ubiquitination of endogenous TOX in KLHL6-overexpressing human T cells (f) and in mouse Klhl6−/− CD8+ T cells (g). h, Violin plots showing the TOX and TCF-1 signatures in transferred Control and KLHL6-OE TILs from scRNA-seq data in Fig. 3g. The boxplot spans from the first to the third quartile of the distribution, with the median positioned in the centre. Whiskers represent the minimum and maximum values, excluding outliers. Values plotted represent cells from a single replicate. i–k, OT-I CD8+ T cells from CD45.1+ WT or KO donor mice were transduced with shCtrl or shTox retrovirus and adoptively transferred (4 × 106) into B16-OVA tumour-bearing mice. The mice were euthanized at day 14 for analysis after adoptive transfer (n = 8 mice). Percentages (i) and absolute numbers (j) of Texterm (Ly108−TIM-3+) and Tpex (Ly108+TIM-3−) subsets, and tumour weights (k) were assessed. l, Correlation of KLHL6 expression with Tpex, Texterm and TOX signatures in human CD8+ TILs from human pan-cancer scRNA-seq data. For immunoblot source data, see Supplementary Fig. 1. Illustration in l created in BioRender. Li, G. (2025) https://BioRender.com/p3754eu. Data in b,f,g are representative of three independent experiments. Data are presented as mean ± s.e.m. Statistical analyses were determined by two-way ANOVA with Tukey’s multiple-comparisons test (c–e,i–k). *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001; NS, not significant. IP, immunoprecipitation.
KLHL6 is known to catalyse Lys48-linked poly-ubiquitination of substrates for proteasomal degradation29,40,41. Indeed, mutation of Lys48 but not Lys63 on ubiquitin greatly diminished KLHL6-induced TOX poly-ubiquitination (Extended Data Fig. 9l). We mapped by mutagenesis potential KLHL6-targeted lysines in TOX and identified four conserved residues (Lys245, Lys246, Lys248 and Lys323) as the primary sites for KLHL6-induced degradation (Extended Data Fig. 9m). Mutation of all four sites (4KR) rendered TOX nearly unresponsive to KLHL6-mediated ubiquitination and degradation, without affecting their interaction (Extended Data Fig. 9n–p). A cycloheximide chase assay confirmed TOX(4KR) showed a markedly prolonged half-life compared with TOX(WT) (Extended Data Fig. 9q), emphasizing the importance of these lysine residues in KLHL6-mediated proteasomal degradation.
KLHL6 loss causes TOX-dependent exhaustion
TOX is a master regulator for initiating and promoting T cell exhaustion7,36,38,39,42. In CD8+ TILs from B16-OVA tumours, TOX expression correlated positively with TIM-3 and negatively with TCF-1 (Extended Data Fig. 10a). TOXhi cells contained more Texterm and fewer Tpex populations compared with TOXlo cells (Extended Data Fig. 10b). Consistent with KLHL6 promoting TOX degradation, scRNA-seq of KLHL6-OE TILs showed reduced TOX and enhanced TCF-1 signatures (Fig. 4h). To determine whether KLHL6 regulates exhaustion through TOX, we examined differentiation of Klhl6−/−OT-I TILs transduced with a short-hairpin RNA (shRNA) targeting Tox. This knockdown (KD) approach was preferred over sgRNA-mediated KO because complete TOX deficiency impairs survival in Tex cells7,38. TOX KD in KLHL6-deficient TILs markedly decreased the proportion of Ly108−TIM-3+ Texterm cells, and favoured accumulation of the Ly108+TIM-3− Tpex subpopulation and total TIL numbers compared with KLHL6-deficient cells without Tox KD (Fig. 4i,j and Extended Data Fig. 10c), consistent with TOX reduction mitigating the Texterm skewing caused by KLHL6 loss. Although TOX levels in KLHL6-deficient TILs with Tox KD were restored to levels similar to WT T cells (Extended Data Fig. 10d), tumour control was only partially rescued (Fig. 4k), suggesting more downstream targets may contribute to KLHL6-mediated anti-tumour effects. Analysis of pan-cancer scRNA-seq data from TILs of 316 patients across various cancer types22 revealed that KLHL6lo CD8+ TILs were enriched with exhaustion-associated genes and TOX and Texterm signatures, but had lower expression of stem-like genes and Tpex signature compared with KLHL6hi cells (Fig. 4l and Extended Data Fig. 10e). Collectively, these findings suggest KLHL6-mediated regulation of TOX protein levels controls the transition of self-renewing Tpex cells to Texterm cells within tumours.
To assess potential functional consequences of KLHL6-resistant TOX, TOX(WT) or TOX(4KR) was overexpressed in CD8+ T cells through retroviral transduction at comparable infection efficiencies, followed by 5-day in vitro expansion (Extended Data Fig. 10f). TOX(WT) overexpression induced exhaustion features, including PD-1 and LAG-3 upregulation and reduced cytokine production7 (Extended Data Fig. 10g–i). Notably, TOX(4KR) cells showed a small but significant difference compared with the TOX(WT) group, possibly because low endogenous KLHL6 expression cannot rapidly amplify expression differences between WT and mutant TOX, as evidenced by higher TOX levels and increased inhibitory receptor expression and cytokine suppression (Extended Data Fig. 10g–i). Transcriptomic analysis revealed TOX(WT) overexpression upregulated exhaustion genes, whereas TOX(4KR) further amplified this profile, including Pdcd1, Havcr2, Nr4a3, Nfkb2, Hif1a and Bhlhe41 (Extended Data Fig. 10j). Furthermore, TOX(4KR) cells showed reduced effector gene expression relative to TOX(WT) cells (Extended Data Fig. 10k). In the B16-OVA model, TOX(4KR)-overexpressing OT-I TILs showed on day 14 post-ACT higher TOX levels, increased inhibitory receptor expression, and significantly fewer TNF+IFNγ+ effector cells compared with TOX(WT) cells (Extended Data Fig. 10l–o). These results indicate resistance to KLHL6-mediated degradation stabilized TOX and exacerbated T cell exhaustion, further supporting the mechanistic role of KLHL6 as a critical regulator of T cell anti-tumour function.
KLHL6–PGAM5 axis sustains T cell immunity
Mitochondrial capacity is a key contributor to T cell exhaustion and anti-tumour immunity3,5,6. Transmission electron microscopy demonstrated KLHL6-deficient T cells contain smaller mitochondria with shortened cristae than WT cells (Fig. 5a). Mitochondrial morphology reflects the balance between fission and fusion, and KLHL6 loss shifted this equilibrium towards excessive fission, as evidenced by decreased expression of fusion mediators, including mitofusin 2 (Mfn2) and optic atrophy 1 (Opa1), and elevated levels of the fission factor dynamin-related protein 1 (Drp1) (Extended Data Fig. 11a). Reduced phosphorylation of Drp1 at Ser637 (p-Drp1S637) indicated preferential increase of the active form, which promotes mitochondrial fragmentation (Extended Data Fig. 11a). Moreover, KLHL6-deficient T cells showed increased p62 and a slight decrease of LC3-phosphatidylethanolamine conjugate (LC3-II) (Extended Data Fig. 11b), suggesting impaired mitophagy and defective mitochondrial quality control43. Pharmacological inhibition of mitochondrial fission in WT and KLHL6-deficient T cells enhanced anti-tumour activity, with prolonged survival of recipient mice44 (Extended Data Fig. 11c). Together, these results demonstrate KLHL6 deficiency drives excessive mitochondrial fragmentation that may impair anti-tumour activity.
a, Representative mitochondrial morphology, individual mitochondrial area (n = 218) and total crista length per mitochondrion (n = 60 cells) in WT and KO OT-I T cells. b–f, CD8+ WT and KLHL6 KO OT-I T cells transduced with shPgam5 (shP5) or shCtrl retrovirus were cultured for 6 days in vitro. b, Immunoblots of indicated proteins. c–f, OCR (c,d), SRC (e) and mitochondrial ATP production (f) were assessed (n = 9 tests). g–i, CD45.1/2+ WT and CD45.1+ KO OT-I T cells transduced with either GFP-shCtrl or Thy1.1-shPgam5 (shP5) were mixed equally (1:1:1:1) and cotransferred into CD45.2+ mice bearing B16-OVA tumours. Mice were analysed on day 14 post-ACT (n = 7 mice). Percentages of TNF+IFNγ+CD8+ TILs (g), frequencies (h) and cell numbers (i) of Tpex (Ly108+TIM-3−) and Texterm (Ly108−TIM-3+) subsets were assessed. j,k, Tumour volume (j) and survival curves (k) of tumour-bearing mice following separate transfer of 5 × 106 indicated cells were recorded (n = 10 mice). Mice with tumour volumes greater than 1,500 mm3 were euthanized and this was defined as death. l,m, 4 × 106 WT + Empty, WT + PGC1α, KO + Empty or KO + PGC1α OT-I T cells were adoptively transferred into B16-OVA tumour-bearing mice. Mice were analysed at day 14 post-ACT for the frequencies and numbers of Tpex and Texterm cells (l), and cytokine production (m) (n = 6 mice). n,o, CD45.1+ WT and KO OT-I T cells were transduced with either shCtrl, shPgam5 (shP5), shTox or a combination of shPgam5 and shTox (shP5 + shTox) retrovirus. A total of 4 × 106 cells from each group were transferred into CD45.2+ B16-OVA tumour-bearing mice. Tpex and Texterm subsets (n) and cytokine production (o) in CD8+ TILs were assessed on day 14 post-ACT (n = 6 mice). For immunoblot source data, see Supplementary Fig. 1. Experiments in a,b were repeated three times. Data in n,o are representative of two independent experiments. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (a), two-way ANOVA with Tukey’s multiple-comparisons test (d–j,l–o), and log-rank (Mantel–Cox) test (k). *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001; NS, not significant. Scale bar, 1 μm.
To elucidate how KLHL6 modulates mitochondrial dynamics, we further analysed E-STUB data and identified phosphoglycerate mutase 5 (PGAM5), a mitochondrial Ser/Thr phosphatase that regulates mitochondrial homeostasis45,46,47, as a potential target of KLHL6 (Fig. 4a and Extended Data Fig. 9c). Experiments confirmed KLHL6 interacts with PGAM5 and promotes its UPS degradation, whereas KLHL6 deficiency in OT-I T cells elevated PGAM5 abundance (Extended Data Fig. 11d–g). On mitochondrial stress, PGAM5 recruits and dephosphorylates Drp1 at Ser637 to activate GTPase function and drive mitochondrial fission45. PGAM5 KD in KLHL6-deficient cells restored fusion-related proteins (Mfn2, Opa1) and p-Drp1S637, improved mitochondrial activity (TMRE/MTG ratio), alleviated fragmentation and increased OCR, SRC and ATP production (Fig. 5b–f and Extended Data Fig. 11h,i). PGAM5 KD modestly enhanced mitochondrial metrics in WT cells as well. In an in vivo cotransfer assay (Extended Data Fig. 11j), PGAM5 KD rescued much of the numerical defect observed in KLHL6-deficient cells, enhanced cytokine production, increased Tpex cells and reduced Texterm frequency and expression of exhaustion markers (Fig. 5g–i and Extended Data Fig. 11k,l). PGAM5 KD in WT TILs modestly enhanced cytokine production and had limited effects on Tpex frequency (Fig. 5g,h), suggesting divergent mitochondrial requirements in WT versus KLHL6-deficient T cells. PGAM5 KD also increased TCM (CD62L+CD44+) numbers in dLN and spleen (Extended Data Fig. 11m). Mice receiving PGAM5 KD T cells, either WT (WT + shP5) or KLHL6-deficient (KO + shP5), showed improved tumour control and prolonged survival compared with respective controls (Fig. 5j,k). Similar mitochondrial and anti-tumour improvements were achieved with the PGAM5 inhibitor LFHP-1c48 (Extended Data Fig. 11n–y). These results demonstrate PGAM5 contributed substantially to the mitochondrial and functional defects observed in KLHL6-deficient T cells.
To independently validate that impaired mitochondrial function contributes to KLHL6-deficient T cell dysfunction, we tested whether enhancing mitochondrial function through the ectopic expression of PGC1α, a master regulator of mitochondrial biogenesis and oxidative phosphorylation (OXPHOS)49, could rescue the functional defects of KLHL6-deficient T cells. Enforced PGC1α expression is reported to improve metabolism and restore effector function in Tex cells31,50. PGC1α overexpression in KLHL6-deficient T cells did enhance mitochondrial function, with increased basal and maximal OCR, SRC and ATP production (Extended Data Fig. 12a,b). In the B16-OVA tumour model, PGC1α overexpression improved tumour control and increased intra-tumoural accumulation of transferred KLHL6-deficient T cells (Extended Data Fig. 12c,d). Furthermore, enforced PGC1α expression reduced mitochondrial depolarization, restricted terminal exhaustion, promoted formation of Tpex cells and enhanced cytokine production in KLHL6-deficient TILs (Fig. 5l,m and Extended Data Fig. 12e). These results support the conclusion that mitochondrial insufficiency is a driver of KLHL6-deficient T cell dysfunction.
Dual targeting boosts anti-tumour immunity
Previous results showed KLHL6 supports T cell anti-tumour function primarily by limiting TOX-driven exhaustion differentiation and PGAM5-mediated mitochondrial dysfunction. To test whether dual targeting of TOX and PGAM5 could additively improve KLHL6-deficient T cell function, we simultaneously knocked down both genes in KLHL6-deficient T cells and evaluated performance in vivo (Extended Data Fig. 12f). KD of TOX or PGAM5 alone partially restored anti-tumour immunity, with improved tumour control, enhanced TIL accumulation, reduced terminal exhaustion and increased cytokine production (Fig. 5n,o and Extended Data Fig. 12g–i). Dual KD resulted in near-complete rescue, with phenotypic and functional features largely restored to WT levels (Fig. 5n,o and Extended Data Fig. 12g–i). These findings further confirm both TOX and PGAM5 as key functional substrates of KLHL6 regulating T cell exhaustion and anti-tumour immunity, highlighting that KLHL6 coordinates transcriptional and metabolic pathways rather than acting through a singular substrate.
To explore the relationship between these two downstream pathways, we overexpressed TOX in CD8+ T cells in vitro to test direct effects on mitochondrial function. TOX-OE did not affect mitochondrial depolarization with or without TCR restimulation (Extended Data Fig. 12j), and only modestly increased mitochondrial reactive oxygen species (ROS) levels (Extended Data Fig. 12k). Seahorse analysis revealed no changes in mitochondrial OCR, SRC or ATP production, although activation-induced glycolysis (glycoPER) was reduced on TCR stimulation, suggesting TOX does not directly regulate mitochondrial function (Extended Data Fig. 12l–n). In addition, TOX KD failed to rescue mitochondrial defects in KLHL6-deficient cells (Extended Data Fig. 12o,p). These findings support KLHL6 maintains mitochondrial fitness independently of TOX, primarily through PGAM5.
Discussion
Ubiquitin modification controls intracellular protein degradation, translocation and function, thereby regulating cellular homeostasis. Dysregulation of ubiquitination is implicated in human malignancies51. Beyond tumour-intrinsic effects, ubiquitin modification shapes host defence and modulates the TME by influencing innate and adaptive immunity52,53. Targeting UPS pathways, including E3 ligases such as Cbl-b, Fbxo38, Mdm2 and Stub1, has enhanced T cell immunotherapy efficacy15,17,54,55. However, there remains limited understanding of ubiquitin-dependent regulation of T cell dysfunction within the TME. Our integrative computational analysis-guided CRISPR screen identified the E3 ligase KLHL6 as a previously unrecognized multifunctional negative modulator of T cell dysfunction that counteracts two main obstacles: exhaustion and mitochondrial dysfunction.
KLHL6 regulated Lys48-linked ubiquitination and proteasomal degradation of TOX, which governs the transition from Tpex cells towards terminal differentiation. This post-translational control complements transcriptional and epigenetic mechanisms of TOX regulation. Ectopic KLHL6-OE reduced but did not abolish intracellular TOX, enhancing tumour control and prolonging T cell persistence without the detrimental effects of total TOX loss. Various factors, including chronic TCR stimulation3,4,6, PD-1 signalling6 and metabolic stress5,56 within the TME can trigger mitochondrial insufficiency and reinforce T cell dysfunction. We found KLHL6 preserves mitochondrial fitness through the PGAM5–Drp1 axis. Loss of KLHL6 drove pathological mitochondrial fission, probably contributing to the hyperaccumulation of fragmented mitochondria observed in TILs during chronic antigen stimulation6. Targeting PGAM5-mediated mitochondrial fission partially restored anti-tumour function in KLHL6-deficient T cells, although complete inhibition or PGAM5 deletion can cause excessive mitochondrial fusion and senescence57. Dual targeting of TOX and PGAM5 additively enhanced KLHL6-deficient T cell anti-tumour responses, highlighting functional independence and synergy. We speculate these two separate pathways converge to further drive differentiation to exhaustion and intracellular stress, as suggested by the phenotypes of KLHL6-deficient TILs in which only one of the pathways has been partially repaired.
Recently, proteolysis-targeting chimeras and molecular glues leveraging E3-substrate interactions showed potential to enhance immune checkpoint blockade and/or ACT by degrading targets such as PD-L1 in tumour cells or HPK1 in T cells58,59. However, as regulation of T cell dysfunction in the TME is multifaceted, single-targeted approaches may prove inadequate. Our findings demonstrated KLHL6 acts as a dual-negative regulator of T cell exhaustion and mitochondrial dysfunction, providing a rationale for developing proteolysis-targeting chimeras that simultaneously target many KLHL6 substrates, including TOX and PGAM5, to overcome obstacles to T-cell-based immunotherapy. Besides immunomodulatory effects in T cells, KLHL6 also functions as a tumour suppressor by promoting degradation of CDK2, Roquin2 and NOTCH2, thereby impeding tumour progression29,40,41. Thus, developing specific small-molecule agents that enhance KLHL6 expression or activity may offer further benefits by directly suppressing tumour growth while revitalizing T cell function. Furthermore, exploring other E3 ligases, deubiquitinases and/or their manipulation through proteolysis-targeting chimera-based or modifier-based strategies may broaden therapeutic opportunities in cancer immunotherapy.
In summary, our study provides an in-depth view of how KLHL6-mediated ubiquitination orchestrates T cell anti-tumour immunity by means of dual regulation of Tex lineage differentiation and mitochondrial dynamics. By elucidating molecular characteristics of T cell exhaustion, this work offers new insights into fundamental immunology and concrete avenues for advancing cancer immunotherapy.
Methods
Mice
Male and female mice were used for the study. CD45.1+ OT-I or P14 TCR-transgenic mice were housed together. CD45.2+ male and female C57BL/6N or C57BL/6JNifdc mice aged 6–8 weeks were purchased from Vital River as recipients. Female NCG (NOD/ShiLtJGpt-Prkdcem26Cd52Il2rgem26Cd22/Gpt) mice aged 6–8 weeks were purchased from GemPharmatech. Rosa26-Cas9 mice were provided as a gift from the W. Sheng laboratory at the University of Zhejiang. We crossed Rosa26-Cas9 mice with OT-I transgenic mice to generate Cas9+ OT-I mice for CRISPR–Cas9 screening in tumour antigen-specific CD8+ T cells. Four-week-old Klhl6+/− mice were purchased from Cyagen. The Klhl6+/− mice were crossed with OT-I, P14 or C57BL/6N mice to generate Klhl6−/− OT-I/P14 mice or Klhl6−/− mice for subsequent experiments. All mice were kept in a specific-pathogen-free facility, and all animal experiments were performed with the approval of the Institutional Animal Care and Use Committee of Suzhou Institute of Systems Medicine (ISM-IACUC-0151-R and ISM-IACUC-20240098). Mice were housed in standard conditions, with 12 h/12 h light/dark cycles, a controlled temperature of 22–24 °C and humidity of 60%, with unrestricted food and water availability, and were examined daily. All mice were used at 6–16 weeks old. All tumour burdens did not exceed the permission of the Institutional Animal Care and Use Committee of Suzhou Institute of Systems Medicine. Age-matched and sex-matched mice were assigned randomly to experimental and control groups.
Cell lines
Human embryonic kidney 293T (HEK293T) cells were purchased from the American Type Culture Collection (ATCC, CRL-3216) and maintained in DMEM (Gibco, C11995500BT) supplemented with 10% fetal bovine serum (FBS) (Gibco, 16000044) and 1% penicillin–streptomycin (P/S) (Gibco, 15140122). The mouse melanoma cell line B16 was transduced to express OVA257-264 antigen (a gift from Bo Huang laboratory) and maintained in DMEM with 10% FBS and 1% P/S. HepG2 cells (ATCC, HB-8065) were transduced to express human NY-ESO antigen (HepG2-ESO) and cultured in DMEM with 10% FBS and 1% P/S. Jurkat (ATCC, TIB-152) and EL4 (ATCC, TIB-39) cell lines were cultured within the complete Roswell Park Memorial Institute (RPMI)-1640 medium supplemented with 10% FBS, 1% P/S, 1% GlutaMAX (Gibco, 35050061), 10 mM HEPES (Gibco, 15630130), 1% non-essential amino acids (Gibco, 11140076), 1 mM sodium pyruvate (Gibco, 11360070) and 50 μM β-mercaptoethanol (Sigma, M6250). HEK293T, Jurkat, HepG2 and EL4 cells were pre-authenticated by ATCC by short tandem repeat (STR) sequencing. B16-OVA cells were frequently monitored based on their morphological features but have not been authenticated by STR. All cell lines were routinely tested for mycoplasma contamination.
Plasmids
Mouse Klhl6, Tox and Ppargc1a genes were amplified from the complementary DNA (cDNA) library of mice OT-I T cells, and human KLHL6 and TOX genes were amplified from human peripheral blood mononuclear cells (PBMCs). Retroviral plasmid (MSGV-Thy1.1-Klhl6, MSGV-Thy1.1-Ppargc1a and MSGV-Thy1.1-Vector) and packaging vector (pCL-Eco) plasmid were used to produce retroviruses in HEK293T cells using 293 Transfection Reagent (Mirus, MIR 2700), which were then transduced into OT-I CD8+ T cells. The MESV-shCtrl-GFP (Addgene, 85587) was used for Tox or Pgam5 KD. Primer sequences used for Tox and Pgam5 KD can be found in Supplementary Table 6. The retroviral plasmids (MSGV-NGFR-KLHL6, MSGV-Thy1.1-1G4 TCR and MSGV-NGFR-Vector) and packaging vectors (pHIT60 and RD114) were used to produce retroviruses, which were used to transduce PBMCs and Jurkat cells. Lentivirus vectors (pCCLc-MND-Thy1.1-Klhl6 and pCCLc-MND-Thy1.1-Vector) and packaging vector (PA2X and VSV-G) plasmids were used for lentivirus production in HEK293T cells using Liposomal Transfection Reagent for transduction into EL4 cell line. For transient expression experiments in HEK293T cells, the vector plasmid pcDNA4/TO or pFLAG-CMV-4 was used according to the experimental need.
Primary mouse T cell isolation, viral transduction and culture
Naive OT-I T lymphocytes were isolated from the spleens and peripheral lymph nodes of male and female OT-I mice (6–8 weeks). Spleens and peripheral lymph nodes were collected, and mashed through a 70-μm filter, and red blood cells were lysed using red blood cell lysis buffer (BioLegend, 420301) followed by washing with 1× phosphate-buffered saline (PBS). CD8+ OT-I T cells were purified using a CD8+ Naive T cell isolation kit (BioLegend, 480043) according to the manufacturer’s instructions. Primary mouse T cells were counted and then resuspended in RPMI-1640 supplemented with 10% FBS, 1% sodium pyruvate, 1% non-essential amino acids, 10 mM HEPES, 1% GlutaMAX, 1% P/S, 50 μM β-mercaptoethanol and mouse IL-2 (20 U ml−1, Peprotech, 212-12). Then, the resuspended CD8+ OT-I T cells were seeded at a concentration of 1 million cells per ml on 24-well plates with overnight-bound anti-mouse CD3 (2 μg ml−1, BioLegend, 100359) and anti-mouse CD28 (1 μg ml−1, BioLegend, 102121) antibodies. Cells were activated in 24-well plates for 48 h and then transferred out of the activation plates and passaged to new plates every 2 days with a concentration of 1 million cells per ml. For drug treatment experiments, DMSO (Sigma, D2650), 2 μM LFHP-1c (MCE, HY-139598) or 10 μM Mdivi-1 (Selleck, S7162) and 20 μM M1 (Selleck, S3375) were added to cultures daily starting on day 3 after T cell activation. In viral transduction, 7.5 × 105 OT-I cells were transduced with unconcentrated retroviral supernatant after 24 h of activation in 24-well plates coated with RetroNectin reagent (15 μg ml−1, Takara, T100B). Following centrifugation at 2,500 rpm for 90 min at 30 °C, T cells were cultured in the incubator for 24 h. The transduction was repeated 24 h later and then returned to fresh medium for culture. Drug-treated or retrovirus-transduced OT-I cells were sorted by flow cytometry and then adoptively transferred into recipient mice that were inoculated with B16-OVA tumour cells before transfer.
Human T cell isolation, viral transduction and culture
Human PBMCs from healthy donors were purchased from Sailybio and isolated using Lymphoprep (Cytiva, 17144003) according to the manufacturer’s protocol. Isolated PBMCs were cultured in RPMI-1640 medium supplemented with 5% Human Serum AB (Gemini, 100-512), 1% GlutaMAX, 1% non-essential amino acids, 1% P/S, 1 mM sodium pyruvate, 10 mM HEPES and 50 μM β-mercaptoethanol in the presence of human IL-2 (100 U ml−1, Peprotech, 200-02). PBMCs were activated by anti-human CD3 (1 μg ml−1, BioLegend, 317347) and anti-human CD28 (1 μg ml−1, BioLegend, 302943) monoclonal antibodies for 2 days and then underwent viral transduction. In brief, 1 × 106 PBMCs were transferred to a new 24-well plate and dually transduced by 1G4 TCR-specific and KLHL6-specific retroviral supernatant in the presence of 10 μg ml−1 polybrene (Sigma, TR-1003-G). Following centrifugation at 2,500 rpm for 90 min at 30 °C, PBMCs were cultured in the incubator for 24 h with fresh medium and then underwent repeated transduction. The transduced PBMCs were adoptively transferred into female NCG mice that were inoculated with HepG2-ESO tumour cells before transfer.
B16 tumour model and ACT immunotherapy
To investigate the anti-tumour activity of T cells in vivo, 2 × 105 B16-OVA melanoma cells were subcutaneously injected into female C57BL/6N mice. Nine days after tumour implantation, each tumour-bearing mouse was intravenously injected with the required number of CD8+ OT-I T cells from female OT-I mice, which had been expanded for 6 days according to different experimental designs. Tumour-bearing mice received 5 Gy of sublethal irradiation for lymphodepletion 1 day before ACT. For the analysis of tumour growth and mice survival, tumour volume was measured every 2 days and calculated as length (mm) × width (mm) × width (mm) × 0.5. Mice with tumour volumes greater than 1,500 mm3 were euthanized and defined as dead for survival analysis. For the analysis of functional phenotype, mice were euthanized and tissues from tumours, spleens and lymph nodes were collected at days 7, 14, 21 or 28 post-ACT, depending on different experimental designs. For the CellTrace Violet labelling assay, equal numbers of CellTrace Violet-labelled control and KLHL6-OE OT-I T cells were cotransferred into recipient tumour-bearing mice, and TILs were analysed by flow cytometry on day 4 post-ACT. The tumours were digested by Type II collagenase (Worthington Biochemical, LS004176) and processed with Percoll (Cytiva, 17089109). Adoptively transferred OT-I T cells were isolated from tumours, spleens, and lymph nodes, and cell numbers were counted. Isolated T cells were washed and resuspended in ice-cold PBS with 2% FBS in the presence of specific antibodies for the determination of their proportion and functional phenotype through flow cytometry.
In vivo Tpex transfer assay
Female C57BL/6N (CD45.2+) mice were subcutaneously implanted with 2 × 105 B16-OVA cells on day 0. On day 9, each tumour-bearing mouse was intravenously injected with 3 × 106 control or KLHL6-OE CD45.1+ OT-I T cells. Then 14 days after ACT, Tpex (Ly108+TIM-3−) TILs were sorted from tumours by flow cytometry. After sorting, the cells were centrifuged and resuspended in PBS. A total of 5 × 104 Tpex cells were transferred through tail vein injection into female C57BL/6N (CD45.2+) mice that had been subcutaneously implanted with 3 × 105 B16-OVA cells 2 days before. Tumour sizes were measured on day 8 after ACT and every 2 days thereafter. TILs were isolated at days 8 and 16 for phenotypic analysis as previously described in ref. 33.
NCG mouse model and 1G4 TCR-T cell therapy
Female NCG mice were subcutaneously implanted with 4 × 106 HepG2-ESO cells. Subsequently, 1G4 TCR-T cells transduced with or without KLHL6, respectively, were expanded for 12 days in vitro and adoptively transferred into the tumour-bearing mice (6 million cells per mouse) when tumour volumes reached 80 mm3. Mice were euthanized on day 16 after ACT, and the tumours were collected for weighing. For the in vivo phenotyping, the blood, tumours and spleens were collected. The spleens and blood were mashed and/or lysed with red blood cell lysis buffer for 5 min on ice. To isolate T cells from the tumour, the tumours were digested by Type II collagenase and processed with Percoll. Then, the isolated T cells were stained with antibody cocktails and analysed by flow cytometry.
LCMV infection and adoptive T cell transfer
CD45.2+ C57BL/6 recipient mice were intraperitoneally infected with 2 × 105 plaque-forming units (PFU) of LCMV-Armstrong or intravenously injected through the tail vein with 2 × 106 PFU of LCMV-Clone 13. One day before infection, mice received adoptive transfers of 5 × 104 (for Armstrong) or 5 × 103 (for Clone 13) P14 CD8+ T cells. Phenotypic analyses were performed at various time points p.i. according to the experimental design20. Naive WT CD8+ T cells and Klhl6−/− (KO) CD8+ T cells for transfer were isolated from P14 mice using a naive CD8+ T cell isolation kit and adoptively transferred into recipient mice. For retroviral transduction, naive P14 CD8+ T cells were activated for 24 h and then transduced with MSGV-Thy1.1-Klhl6 (KLHL6-OE) or MSGV-Thy1.1-Vector (Control) retrovirus. The following day, transduced CD8+ T cells were sorted, resuspended in cold 1× PBS and adoptively transferred into recipient mice, followed by LCMV infection 1 day later.
LCMV viral RNA quantification
CD45.2+ C57BL/6 recipient mice were intravenously injected with 2 × 106 PFU of LCMV-Clone 13. One day before infection, mice were adoptively transferred with 1 × 104 P14 CD8+ T cells. Liver and lung samples were collected on day 15 p.i., and viral load was quantified using a quantitative PCR (qPCR)-based assay, as previously described60. In brief, total RNA was extracted using the Qiagen RNA isolation kit and subsequently subjected to reverse transcription with the Reverse Transcription Kit (Vazyme, R323-01). cDNA was then used as template for qPCR with 2× SYBR Green qPCR Master Mix (Bimake, b21203). Primers for LCMV GP and hypoxanthine-guanine phosphoribosyltransferase (HPRT) are listed in Supplementary Table 6.
Surprisal analysis
We analysed harmonized bulk RNA-seq datasets comprising 136 samples from 8 previously published studies8,18,33,61,62,63,64,65 including gene expression profiles from CD8+ memory and effector T cells, TILs and chimeric antigen receptor T cells, as well as endogenous Tex cells exposed to chronic antigen stimulation, with or without immune checkpoint inhibition. A complete list of datasets is provided in Supplementary Table 1. Despite thousands of genes that could all be changing across various studies and conditions, we proposed that many genes are coordinately changing together as a group (or gene module), which reflects the fundamental biology of T cell exhaustion programs. Surprisal analysis has been well-documented in deconvoluting the change of thousands of genes into the change of only a couple of gene modules and one unchanged gene expression baseline66,67. The unchanged gene expression baseline reflected the biological processes that are conserved across conditions and time points. The gene module reflected the deviation from the global stable state.
We used surprisal analysis66,67, an information-theoretical analysis technique that integrates principles of thermodynamics and maximal entropy, here to simplify the transcriptome changes into two main gene modules and one unchanged gene expression baseline, which when added together, accurately capture the global transcriptomic profiles of the raw data. Briefly, the logarithm of the measured level of a transcript i at a specific study a sample b, \(\mathrm{ln}{X}_{i}(a\_b)\), is expressed as a sum of a log-transformed gene expression baseline, term \(\mathrm{ln}{X}_{i}^{0}\), and several gene modules \({\lambda }_{j}(a\_b)\times {G}_{{ij}}\), representing deviations from the common expression baseline. Each deviation term is a product of a study-sample-dependent module score\({\lambda }_{j}(a\_b)\), and the study-sample-independent module-specific contribution score Gij of the gene i. Gene i that shows large positive or negative contribution to a module j (high positive or negative Gij value) represents a gene that is functionally positively or negatively correlated with the module j. In other words, the biological function of module j could be inferred by functional enrichment analysis of genes with positive and negative Gij values. The study-sample-dependent module scores of the top modules (in this case, modules 1 and 2) should be able to illustrate the global transcriptome similarities or dissimilarities. To calculate these gene modules, we first computed the singular value decomposition of the matrix \({\rm{ln}}X(a\_b)\). As described previously67, the singular value decomposition factored this matrix in a way that determined the two sets of parameters that are required in the surprisal analysis: the Lagrange multipliers (\({\lambda }_{j}(a\_b)\)) for all gene modules at a given sample and for all samples in all studies, as well as the module-specific contribution scores (Gij for all transcripts i at each gene module j. Further enrichment analysis of the functions associated with each module was performed based on the module-specific contribution scores of the genes associated with that module. These two dominant gene modules (modules 1 and 2) each consist of two gene sets showing opposite expression trends across all samples (M+/−) (Supplementary Table 2). M1+ genes were low in naive and memory T cells but elevated in both early and late exhausted T cells from tumours, as well as in exhausted T cells from chronic infections, whereas M1− genes showed the inverse trend. M2+ genes were selectively enriched in late exhausted cells across the tumour and chronic infection settings, and M2− genes were more highly expressed in naive and early exhausted states.
GSEA of the gene modules
GSEA was performed using MSigDB (v.7.5.1) pathways and custom gene sets derived from the existing literature33,68. Genes were ranked by surprisal analysis scores and analysed separately for association with modules 1 and 2 using the R package clusterProfiler (v.4.12.0)69. Ties (zero scores) were excluded. Enrichment scores were normalized by use of permutation tests, and P values were derived accordingly. Custom gene sets consisted of the top 400 differentially expressed genes (Mann–Whitney U-test) after removing housekeeping, ribosomal and mitochondrial genes. The full GSEA results are provided in Supplementary Table 3.
Mitochondrial function analysis from public databases
To investigate genes associated with mitochondrial function in T cells, we analysed roughly 400,000 TILs from 316 patients across 21 cancer types22, correlating gene expression with pathway activity for hallmark_oxidative_phosphorylation in the Molecular Signature Database. This analysis identified 286 E3 ligases positively associated with mitochondrial function. To account for patient variability in the TIL atlas, we also analysed a ‘cleaner’ mouse RNA-seq dataset within the context of adoptive T cell therapy, in which tumour-specific T cells were sorted into two subsets: (MTDR/MTG)hi functional mitochondria and (MTDR/MTG)lo dysfunctional mitochondria6. Differential pathway enrichment analysis confirmed that ubiquitin-related pathways are significantly associated with mitochondrial function. Through this, we identified 191 E3 ligases positively linked to mitochondrial function. The 133 E3 ligases identified as overlapping between human and mouse analyses (Supplementary Table 4) were selected for in vivo CRISPR screening.
CRISPR–Cas9 screens using the retroviral E3-related library
Retroviral sgRNA vector and sgRNA cloning
In this study, CRISPR–Cas9 sgRNA was expressed using pSL21-Thy1.1 or pSL21-mCherry (Addgene, 164410)23. sgRNAs were generated by annealing two DNA oligos and then ligated into the pSL21-Thy1.1 or pSL21-mCherry vector after digestion with BbsI.
E3-related library construction
The pSL21-mCherry vector was used for the construction of sgRNA library. A computational-guided sgRNA library targeting 78 exhaustion-related E3 ligase genes and 133 mitochondrial-related E3 ligase genes were selected and synthesized. The guide RNA (gRNA) sequences were designed according to previously published data and using the gRNA-design tool (GenScript)70. The library associated with exhaustion differentiation comprised a total of 400 gRNAs, including 10 non-targeting controls and 390 unique sgRNAs, with 5 gRNAs designed for each targeting gene. Another library related to mitochondrial function included a total of 671 gRNAs, including 17 non-targeting controls and 654 unique sgRNAs, with 3–5 gRNAs designed for each targeting gene. All sgRNA oligos, including both positive and negative control sgRNAs, were synthesized by SYNBIO Technologies and pooled in equal molarity. The pooled sgRNA oligos were subsequently amplified through PCR and cloned into BbsI-digested pSL21-mCherry vector using Gibson Assembly Kit (NEB, E5510S). The product of Gibson Assembly reaction was then introduced into TG1 Electrocompetent Cells (Biosearch Technologies, 60502) by means of electroporation and cultured overnight on solid Luria-Bertani agar plates (24 × 24-cm culture plate). The total number of colonies across all plates was counted, exceeding 50× representation, and the plasmids were purified using the EndoFree Plasmid Maxi Kit (CWBIO, CW2104M). To verify the identity and relative representation of sgRNAs in the pooled plasmids, a deep-sequencing analysis was performed by a NovaSeq 6000 (PE150) instrument. We confirmed that 100% of the designed sgRNAs were cloned into the vector and the final library is diverse with a Gini index of 0.05.
In vivo screening
The in vivo screening approach was conducted following established protocols from previous studies23,71. Briefly, a retrovirus pool containing sgRNAs was generated by cotransfecting the specific library plasmids and a packaging vector (pCL-Eco) in HEK293T cells. After 48 h of transfection, the viral supernatant was collected and stored at −80 °C. Naive Cas9+ OT-I T cells were isolated from spleens and activated using anti-CD3 and anti-CD28 antibodies. At 24 h after activation, Cas9+ OT-I T cells were transduced with the retrovirus library, and the transduction process was repeated after 24 h. The transduction efficiency was assessed based on the fluorescence intensity of mCherry, and it reached roughly 40% by the end of transduction. Following viral transduction, the cells were washed and cultured in the medium supplemented with mouse IL-2 for 4 days to allow for expansion and gene editing. mCherry-positive cells were sorted by flow cytometry. Roughly 2 × 105 (400 gRNAs library) or 3.5 × 105 (671 gRNAs library) transduced Cas9+ OT-I T cells were saved as ‘day 0 input’ (around 500× cell coverage per sgRNA). Subsequently, transduced Cas9+ OT-I T cells (3 × 106) were transferred into Cas9+ hosts bearing B16-OVA melanoma tumours. At day 7 after ACT, non-exhausted T cells (PD-1−TIM-3−) and exhausted T cells (PD-1+TIM-3+) or (MTDR/MTG)hi and (MTDR/MTG)lo cells were sorted using flow cytometry and frozen at −80 °C until genomic DNA extraction. A minimum of 2 × 105 or 3.5 × 105 Cas9+ OT-I T cells per sample were collected for further analysis.
sgRNA library sequencing
Genomic DNA was extracted by using the PureLink Genomic DNA Mini Kit (Invitrogen, K182001) according to the manufacturer’s instructions. Two rounds of PCR were performed by using the PrimeSTAR HS DNA Polymerase (Takara, R045B) to amplify the sgRNAs and attach Illumina adaptors and indexes to barcode each sample. The primer sequences used to amplify sgRNAs for the PCR are as follows: next-generation sequencing forward (F), AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCTGTATTTCGATTTCTTGGCTTTATATATCTTGT; next-generation sequencing reverse (R), CAAGCAGAAGACGGCATACGAGATATTGGCGTGACTGG AGTTCAGACGTGTGCTCTTCCGATCTGACTAGCCTTATTTAAACTTGCTATGC. Different index sequences were added to distinguish between experimental groups. After each PCR reaction, the PCR products were purified using the AMPure XP beads (Beckman A63881). The library sequencing was performed using the Illumina NovaSeq 6000 (PE150) platform (Novogene).
CRISPR screen data processing and analysis
For data analysis, single-end reads were trimmed and quality filtered using the MAGeCK-VISPR package (v.0.5.5)72 and run using Python (v.3.7.4) and matched against sgRNA sequences from the sgRNA library. Read counts for sgRNAs were normalized by control guides when possible, and when not through median counts values. log2 fold changes were calculated with a 1 × 10−2 pseudo-count to account for zero-count genes and avoid infinite values; these fold changes were used as enrichment differences between DP (PD-1+TIM-3+) cell samples and those of DN (PD-1−TIM-3−) cell samples. The same analyses were also performed between (MTDR/MTG)lo versus (MTDR/MTG)hi cells. Gene-targeting sgRNAs consistently showed enrichment or depletion, whereas non-targeting controls were tightly centred around zero, indicating minimal selection bias. Gene retrieval was 100% across all targets in both screens; sgRNA retrieval was 100% in the exhaustion screen and 99.1% in the mitochondrial fitness screen. The log2 fold-change values for each gene and sgRNA from the CRISPR screens are compiled in Supplementary Table 5.
Experimental workflow in RNA-seq
For the transcriptional profiling of Tex cells, we established a B16-OVA melanoma model to analyse tumour antigen-specific CD8+ T cell exhaustion. Briefly, OT-I CD8+ T cells were activated in vitro using anti-CD3 and anti-CD28 antibodies. On day 9 following the implantation of B16-OVA tumours, 3 × 106 OT-I CD8+ T cells were adoptively transferred to each tumour-bearing mouse. On day 14 after ACT, cells were sorted from tumours and spleens using flow cytometry. PD-1 and TIM-3 were used to label different subsets of exhausted TILs: the PD-1−TIM-3− population, PD-1+TIM-3− population and PD-1+TIM-3+ population. For the transcriptional analysis of adoptively transferred WT and Klhl6−/− CD8+ OT-I T cells in the tumours, Klhl6−/− CD8+ OT-I T cells (CD45.1+) and WT CD8+ OT-I T cells (CD45.1/2+) were mixed in a 1:1 ratio and adoptively transferred into the same B16-OVA tumour-bearing mice. After 14 days, the transferred CD45.1+ and CD45.1/2+ CD8+ T cells were sorted from tumours using flow cytometry and prepared for RNA extraction. Total RNA from the isolated transferred OT-I T cells was extracted using the RNeasy Micro Kit (Qiagen, 74004) following the manufacturer’s instructions and stored at −80 °C for RNA-seq. RNA integrity was assessed using the Agilent 2100 Bioanalyzer (Agilent). Subsequently, the libraries were prepared using the TruSeq RNA sample prep kit (Illumina, FC-122-1001). These libraries were then subjected to sequencing on an Illumina NovaSeq 6000 (PE150) platform, generating roughly 40 million paired-end reads (Novogene).
RNA-seq data processing and analysis
The raw read counts were extracted and then normalized by their library size factors and read and gene lengths using edgeR (v.3.36.0)73, which was then used to calculate differential genes. Detailed information on trimming, alignment and quantification can be found as previously reported74 and further details are available at https://github.com/danielgchen/FH_bulk-RNA-seq_pipeline. In brief, data were trimmed using cutadapt (v.2.9)75, quality checked before and after trimming using FastQC (v.0.11.9), and then mapped and quantified using STAR (v.2.7.7a)76. The pathway enrichment analysis of differentially expressed genes was conducted using clusterProfiler (v.4.12.0)77. GSEA was performed with GSEA (v.4.1.0)68; log2 fold changes were calculated with a 1 × 10−2 pseudo-count to account for zero-count genes and avoid infinite values.
Experimental workflow in scRNA-seq
Activated CD8+ OT-I T cells were transduced with either Vector (MSGV-Thy1.1-Vector) or Klhl6 (MSGV-Thy1.1-Klhl6). Then, these transduced cells were adoptively transferred into B16-OVA tumour-bearing mice at a concentration of 3 × 106 cells per mouse. At day 14 after ACT, OT-I T cells were sorted from tumour samples using flow cytometry. Subsequently, these sorted single cells were encapsulated into droplets, loaded into Chromium microfluidic chips with 30 (v.3) chemistry, and barcoded using a 10× Chromium Controller (10X Genomics). The RNAs from these barcoded cells were subsequently reverse-transcribed, and sequencing libraries were prepared using reagents from a Chromium Single Cell 3′ (v3) reagent kit (10X Genomics), according to the manufacturer’s instructions. Library quantification was performed using the Qubit 3.0 Fluorometer (ThermoFisher Scientific), and library quality was assessed using the 2100 Bioanalyzer with the High Sensitivity DNA kit (Agilent). The NovaSeq 6000 platform (Illumina) was used for sequencing the libraries in 50-base pair paired-end mode.
scRNA-seq data processing and analysis
scRNA-seq analysis pipeline closely follows previously reported methods78,79. Briefly, Droplet-based sequencing data were aligned and quantified by use of Cell Ranger Single-Cell Software Suite (v.7.1.0, 10X Genomics) using refdata-gex-mm10-2020-A as a reference. Cells from each sample were first filtered for cells with 500 or more genes and 1,000 or more counts, then filtered based on (1) fewer than 50,000 counts per cell (library size); (2) fewer than 7,000 detected genes per cell and (3) proportion of mitochondrial gene counts (mitochondrial gene unique molecular identifiers (UMIs)/total UMIs) less than 5%. Doublets were identified through clustering; low-quality, low-count cells were also removed. After quality control-based filtering, Scanpy80 was used to normalize cells by means of counts per million normalization (UMI count per cell was set to 106) and log1p transformation (natural log of counts per million plus one). Principal component analysis was performed using variable genes. Leiden clustering and UMAP plots were generated based on selected principal component analysis dimensions. Normalized data are shown in the form of UMAP colour-coding or violin plots. Embedding density was used for density plots and calculated using scanpy.tl.embedding_density, which is a wrapper for the gaussian density algorithm under scipy. TOX signature was defined by taking the genes differentially upregulated, defined as a false discovery rate less than 0.05 and log2 fold change greater than or equal to 1, on Tox-overexpressed T cells in a tumour model38. Stemness and terminal exhaustion signature were defined as the following Lef1, Tcf7, Aqp3, Ccr7, Sell, Il7r, Gzmk, Dusp1, Dusp2, Fos and Junb for stemness and Pdcd1, Ctla4, Cd200r1, Cd244a, Havcr2, Lag3 and Tigit for terminal exhaustion; genes were derived from literature. Published datasets on T cell exhaustion were obtained from studies related to chronic infection and human tumour-infiltrating CD8+ T cells22,33. The public T cell exhaustion data in chronic infection was processed from raw by filtering for n counts between 2,000 and 10,000, n genes between 1,000 and 3,000, and less than 5% mitochondrial reads; data were then normalized according to the aforementioned methods and projection algorithms.
qPCR with reverse transcription
Total RNAs from cells were extracted using Trizol reagent (Takara, 9109) or the RNeasy Micro Kit, according to the manufacturer’s instructions. The extracted RNAs were reverse-transcribed into cDNA using HiScript Reverse Transcriptase (Vazyme, R323-01). Quantitative real-time PCR was performed using the ABI prism 7500 real-time PCR System (ThermoFisher) and 2× SYBR Green qPCR Master Mix (Bimake, b21203), following the respective manufacturer’s protocols. The data are presented as the fold change in gene expression normalized to an internal reference gene (B2M). The relative expression of mRNA was calculated using the 2−ΔΔCT method. Primer sequences used for qPCR can be found in Supplementary Table 6.
Flow cytometry and sorting
T cells were stained using fluorescent antibodies and subsequently analysed by flow cytometry. To prepare for staining, T cells were collected and washed once with fluorescence-activated cell sorting (FACS) buffer (PBS with 2% FBS). For surface protein staining, cells were stained with fluorescently conjugated antibodies and Live/Dead Fixable Dead Cell Stain Kit (Invitrogen, 65-0866-18) in FACS buffer, then fixed with 2% paraformaldehyde (Casmart) for 30 min at 4 °C. For transcription factor staining, cells were prestained with Live/Dead Fixable Dead Cell Stain Kit and fluorescently conjugated antibodies in FACS buffer to detect surface markers. The cells were then fixed for 30 min at 4 °C using FOXP3/transcription factor fixation buffer (Invitrogen, 00-5523-00) and stained with transcription factor antibodies in permeabilization buffer (Invitrogen, 00-8333-56). For detection of intracellular cytokines, cells were stimulated with phorbol myristate acetate in the presence of Brefeldin A (BFA) (BioLegend, 423304) for 4.5 h. Then, the prestained cells were fixed and stained with cytokines antibodies in the permeabilization buffer. After staining, cells were resuspended in FACS buffer for flow cytometric analysis. Flow cytometry data were collected using BD LSR Fortessa and BD FACSDiva (v.8.0.2), and analysed with FlowJo (v.10.4) software. Cell sorting was performed using BD FACS Aria III and BD FACSDiva (v.8.0.2). A list of antibodies and their dilutions used can be found in Supplementary Table 8.
Transmission electron microscopy
The indicated OT-I T cells were activated in each well of a 24-well plate with mouse anti-CD3 and anti-CD28 antibodies for 3 days and cultured in RPMI-1640 medium for another 3 days. Subsequently, 1 × 106 OT-I T cells were gathered and fixed in a precooled fixation buffer (2.5% glutaraldehyde, 0.1 M phosphate buffer, pH 7.4) overnight at 4 °C. After 3 washes with PBS, cells were postfixed in 1% osmium tetroxide in PBS for 2 h, dehydrated and embedded in Spurr’s resin following standard procedures. Ultrathin sections were stained with uranyl acetate and lead citrate. Mitochondrial morphology was visualized using Hitachi HT-7800 transmission electron microscopy (v.01.20) and an AMT-XR81DIR camera. For quantitation of mitochondrial cross-sectional area and total crista length, images of each cell profile containing four or five mitochondria from a single thin section for the indicated samples were analysed. Cross-sectional area and total crista length per mitochondrion were calculated using the lasso tool in Image J (v.1.8.0) software.
Seahorse analysis
To investigate metabolic characteristics, we used a Seahorse XF24 analyser (Agilent) to measure both OCR and glycoPER in in vitro-expanded T cells and TILs sorted from tumours at day 14 after ACT, according to different experimental designs. Before analysis, these cells were pretreated with the non-buffered XF medium (RPMI-1640 supplemented with 10 mM glucose, 1 mM sodium pyruvate and 2 mM glutamine). Subsequently, the cells were seeded at a density of 1.3 × 105 cells per well in an XF24 cell culture microplate and incubated in a non-CO2 environment for 1 h at 37 °C. To optimize cell adhesion, the plates underwent a 5 min spin at room temperature at 100g with zero brake. Measurements of OCR and glycoPER were conducted under both basal conditions and in response to specific compounds, such as 1.25 μM oligomycin (Oligo), 50 mM 2-deoxy-d-glucose, 1.5 μM carbonyl cyanide-p-trifluoromethoxy-phenylhydrazone, 0.5 μM rotenone and antimycin A (R&A). The SRC was calculated by subtracting basal OCR from maximum OCR. The OCR coupled with mitochondrial ATP production (coupled OCR) is defined as the OCR reduction after the injection of oligomycin, which inhibits ATP synthase. OCR and glycoPER were analysed by Seahorse wave software (Seahorse, Agilent Technologies, v.2.6).
Mitochondrial mass and membrane potential analysis
Mitochondrial mass and membrane potential were assessed using MTG (Invitrogen, M7514) and either TMRE (Invitrogen, T669) or MTDR (Invitrogen, M46753). In brief, cells were stained with 250 nM MTG and 50 nM TMRE or MTDR, and incubated at 37 °C (5% CO2) for 30 min. Following incubation, cells were washed three times with FACS buffer and subsequently subjected to surface marker staining for further flow cytometric analysis.
Western blotting
For protein expression analysis, cells were gathered, washed with cold PBS, and then lysed in 1% SDS (Sangon, 151-21-3) for 30 min on ice. The protein samples were denatured at 95 °C for 15 min and stored at −20 °C. Protein samples were separated on SDS–PAGE gels and then transferred onto methanol-activated polyvinylidene fluoride membranes (Millipore, IPVH00005). Membranes were blocked with 5% nonfat milk in PBS containing Tween-20 (0.1%) for 1 h and then incubated overnight at 4 °C with the respective primary antibodies. The following day, membranes were incubated with the corresponding HRP-coupled secondary antibodies for 2 h at room temperature, followed by signal development using ECL Western Blotting substrate (Tanon, 180-5001) and the Chemidoc automated detection system (Bio-Rad). Data analysis was performed using Image J (v.1.8.0) software. The antibodies and their dilutions used can be found in Supplementary Table 8.
E-STUB for mass spectrometry
As previously described in ref. 37, Jurkat cells were cultured in DMEM supplemented with 10% dialysed FBS. Three biological replicates were prepared for each treatment condition. Jurkat cells were transduced with retroviruses packaged in HEK293T cells using MSGV-NGFR-KLHL6-BirA and MSGV-Thy1.1-BAP-Ub plasmids to co-express KLHL6-BirA and BAP-tagged ubiquitin. Cells transduced with retroviruses packaged from MSGV-NGFR-Empty-BirA and MSGV-Thy1.1-BAP-Ub plasmids served as controls. Then 72 h post-transduction, cells were pretreated with proteasome inhibitor MG132 to promote accumulation of ubiquitylated substrates and then pulsed with 50 μM biotin for 30 min. Following biotin labelling, cells were washed once with ice-cold PBS and lysed on ice using E-STUB RIPA buffer (RIPA buffer supplemented with EDTA-free protease inhibitor cocktail, Pierce Universal Nuclease, 10 mM N-ethylmaleimide, 1 mM EGTA and 1.5 mM MgCl2). Lysates were collected into microcentrifuge tubes, rotated at 4 °C for at least 1 h and clarified by centrifugation at 12,000g for 15 min at 4 °C. Total protein lysates were incubated with 50 μl of resuspended and prewashed streptavidin beads overnight at 4 °C with rotation. The following day, beads were washed 5 times with Wash Buffer (PBS containing 0.05% Tween-20), then incubated with 30 μl of 0.1% SDS at 95 °C for 5 min in a metal bath. Samples were sent to Shanghai Omicsspace Biotech for mass spectrometry analysis. Significant changes between the relative protein abundance of the experimental samples to the control samples were assessed by two-sided moderated t-test as implemented in the limma package (v.3.54.2)81 (Supplementary Table 7).
Co-IP and ubiquitination assays
For the Co-IP assay, cells were transfected with the plasmids according to the experimental designs outlined in the figures. After 36–48 h, the cells were gathered and lysed in Co-IP lysis buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, 1 mM EGTA, 1% NP-40, 2.5 mM sodium pyrophosphate and 1 mM Na3VO4, for 1 h at 4 °C. The cell lysates were centrifuged at 12,000g for 10 min at 4 °C to remove cell debris. The supernatants were collected and incubated with anti-Flag or anti-Myc beads that had been precleaned with Co-IP buffer. These mixtures were then rotated overnight at 4 °C. On the following day, the beads were subjected to five washes with Co-IP lysis buffer and then resuspended in 1× loading buffer. Subsequently, the samples were denatured at 95 °C for 15 min and stored at −20 °C. To assess the ubiquitination of TOX and PGAM5, TOX proteins (both endogenous and exogenous TOX proteins) and endogenous PGAM5 proteins were immunoprecipitated from cell lysates using anti-Flag beads or TOX/PGAM5 antibody-coated beads, depending on the different experimental conditions. In brief, cells were collected and lysed using ultrasonic cracking, and then denatured at 95 °C for 15 min. Cell lysates were incubated in Co-IP lysis buffer with protease inhibitors (Roche) and rotated for 1 h at 4 °C. Subsequently, the cell lysates were centrifuged at 12,000g for 10 min at 4 °C to collect cell supernatant. The supernatant was incubated with the respective antibody–beads complex and rotated overnight at 4 °C. Afterwards, the beads were washed 5 times with Co-IP buffer and denatured by heating at 95 °C for 15 min. These samples were separated by SDS–PAGE, transferred to polyvinylidene fluoride membranes and then subjected to western blotting using the designated primary and secondary antibodies.
Statistical analysis
Statistical analyses were performed using GraphPad Prism (v.8.0). No statistical methods were used to predetermine sample size. Data collection and analysis were conducted without blinding to the experimental conditions. A two-tailed Student’s t-test was used to compare the two groups. For multiple comparisons, a two-way analysis of variance (ANOVA) with Tukey’s or Sidak’s multiple-comparisons test was applied. The log-rank (Mantel–Cox) test was performed to compare mouse survival curves. Data are presented as mean ± standard error of the mean (s.e.m.). The numbers of mice used per experiment and the number of experimental repeats are indicated in the figure legends. P < 0.05 was considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All underlying data are available upon request. All scRNA-seq data used in this study are publicly available and can be accessed from the Sequence Read Archive under accession numbers SRR28131283–SRR28131284. The bulk RNA-seq data can be accessed from the Sequence Read Archive under accession numbers SRR28117257–SRR28117267 and SRR34547767–SRR34547775. The links to datasets obtained from publicly available sources are provided in Supplementary Table 1. Source data are provided with this paper.
Code availability
This paper does not report original code. Any further information required to reanalyse the data reported in this study is available from the corresponding author upon request.
References
Albelda, S. M. CAR T cell therapy for patients with solid tumours: key lessons to learn and unlearn. Nat. Rev. Clin. Oncol. 21, 47–66 (2024).
Baessler, A. & Vignali, D. A. A. T cell exhaustion. Annu. Rev. Immunol. 42, 179–206 (2024).
Scharping, N. E. et al. Mitochondrial stress induced by continuous stimulation under hypoxia rapidly drives T cell exhaustion. Nat. Immunol. 22, 205–215 (2021).
Vardhana, S. A. et al. Impaired mitochondrial oxidative phosphorylation limits the self-renewal of T cells exposed to persistent antigen. Nat. Immunol. 21, 1022–1033 (2020).
Wu, H. et al. Mitochondrial dysfunction promotes the transition of precursor to terminally exhausted T cells through HIF-1alpha-mediated glycolytic reprogramming. Nat. Commun. 14, 6858 (2023).
Yu, Y. R. et al. Disturbed mitochondrial dynamics in CD8(+) TILs reinforce T cell exhaustion. Nat. Immunol. 21, 1540–1551 (2020).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8(+) T cell exhaustion. Nature 571, 211–218 (2019).
Philip, M. et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545, 452–456 (2017).
Zhou, P. et al. Single-cell CRISPR screens in vivo map T cell fate regulomes in cancer. Nature 624, 154–163 (2023).
June, C. H., O’Connor, R. S., Kawalekar, O. U., Ghassemi, S. & Milone, M. C. CAR T cell immunotherapy for human cancer. Science 359, 1361–1365 (2018).
Sharma, P. et al. Immune checkpoint therapy-current perspectives and future directions. Cell 186, 1652–1669 (2023).
Chow, A., Perica, K., Klebanoff, C. A. & Wolchok, J. D. Clinical implications of T cell exhaustion for cancer immunotherapy. Nat. Rev. Clin. Oncol. 19, 775–790 (2022).
Peng, J. J., Wang, L., Li, Z., Ku, C. L. & Ho, P. C. Metabolic challenges and interventions in CAR T cell therapy. Sci. Immunol. 8, eabq3016 (2023).
Dale, B. et al. Advancing targeted protein degradation for cancer therapy. Nat. Rev. Cancer 21, 638–654 (2021).
Stromnes, I. M. et al. Abrogating Cbl-b in effector CD8(+) T cells improves the efficacy of adoptive therapy of leukemia in mice. J. Clin. Invest. 120, 3722–3734 (2010).
Zhao, B. et al. Notch and the pre-TCR coordinate thymocyte proliferation by induction of the SCF subunits Fbxl1 and Fbxl12. Nat. Immunol. 20, 1381–1392 (2019).
LaFleur, M. W. et al. A STUB1-CHIC2 complex inhibits CD8(+) T cells to restrain tumor immunity. Nat. Immunol. 26, 1476–1487 (2025).
Pauken, K. E. et al. Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. Science 354, 1160–1165 (2016).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 37, 1130–1144 (2012).
McManus, D. T. et al. An early precursor CD8(+) T cell that adapts to acute or chronic viral infection. Nature 640, 772–781 (2025).
Chu, T. et al. Precursors of exhausted T cells are pre-emptively formed in acute infection. Nature 640, 782–792 (2025).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, abe6474 (2021).
Chen, Z. et al. In vivo CD8(+) T cell CRISPR screening reveals control by Fli1 in infection and cancer. Cell 184, 1262–1280 (2021).
Pearce, E. L. et al. Enhancing CD8 T-cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
Yu, J. et al. The OX40-TRAF6 axis promotes CTLA-4 degradation to augment antitumor CD8(+) T-cell immunity. Cell Mol. Immunol. 20, 1445–1456 (2023).
He, X. et al. RNF34 functions in immunity and selective mitophagy by targeting MAVS for autophagic degradation. EMBO J. 38, e100978 (2019).
Han, H., Chae, S., Hwang, D. & Song, J. Attenuating MKRN1 E3 ligase-mediated AMPKalpha suppression increases tolerance against metabolic stresses in mice. Cell Stress 2, 325–328 (2018).
Wang, W., Liu, J., Li, J. & Su, H. A PRKN-independent mechanism regulating cardiac mitochondrial quality control. Autophagy 21, 254–256 (2025).
Choi, J. et al. Loss of KLHL6 promotes diffuse large B-cell lymphoma growth and survival by stabilizing the mRNA decay factor roquin2. Nat. Cell Biol. 20, 586–596 (2018).
Bertocci, B. et al. Klhl6 deficiency impairs transitional B cell survival and differentiation. J. Immunol. 199, 2408–2420 (2017).
Bengsch, B. et al. Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8(+) T cell exhaustion. Immunity 45, 358–373 (2016).
Lan, X. et al. Antitumor progenitor exhausted CD8(+) T cells are sustained by TCR engagement. Nat. Immunol. 25, 1046–1058 (2024).
Miller, B. C. et al. Subsets of exhausted CD8(+) T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Robbins, P. F. et al. Tumor regression in patients with metastatic synovial cell sarcoma and melanoma using genetically engineered lymphocytes reactive with NY-ESO-1. J. Clin. Oncol. 29, 917–924 (2011).
Beltra, J. C. et al. Stat5 opposes the transcription factor Tox and rewires exhausted CD8(+) T cells toward durable effector-like states during chronic antigen exposure. Immunity 56, 2699–2718 (2023).
Beltra, J. C. et al. Developmental relationships of four exhausted CD8(+) T cell subsets reveals underlying transcriptional and epigenetic landscape control mechanisms. Immunity 52, 825–841 (2020).
Huang, H. T. et al. Ubiquitin-specific proximity labeling for the identification of E3 ligase substrates. Nat. Chem. Biol. 20, 1227–1236 (2024).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Ying, M. et al. Ubiquitin-dependent degradation of CDK2 drives the therapeutic differentiation of AML by targeting PRDX2. Blood 131, 2698–2711 (2018).
Zhou, N. et al. DLBCL-associated NOTCH2 mutations escape ubiquitin-dependent degradation and promote chemoresistance. Blood 142, 973–988 (2023).
Huang, Y. J. et al. Continuous expression of TOX safeguards exhausted CD8 T cell epigenetic fate. Sci. Immunol. 10, eado3032 (2025).
Liu, L. et al. Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat. Cell Biol. 14, 177–185 (2012).
Buck, M. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016).
Wang, Z., Jiang, H., Chen, S., Du, F. & Wang, X. The mitochondrial phosphatase PGAM5 functions at the convergence point of multiple necrotic death pathways. Cell 148, 228–243 (2012).
Ma, K. et al. Dynamic PGAM5 multimers dephosphorylate BCL-xL or FUNDC1 to regulate mitochondrial and cellular fate. Cell Death Differ. 27, 1036–1051 (2020).
Chen, G. et al. A regulatory signaling loop comprising the PGAM5 phosphatase and CK2 controls receptor-mediated mitophagy. Mol. Cell 54, 362–377 (2014).
Gao, C. et al. A novel PGAM5 inhibitor LFHP-1c protects blood–brain barrier integrity in ischemic stroke. Acta Pharm. Sin. B 11, 1867–1884 (2021).
Lin, J., Handschin, C. & Spiegelman, B. M. Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab. 1, 361–370 (2005).
Scharping, N. E. et al. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity 45, 374–388 (2016).
Senft, D., Qi, J. & Ronai, Z. A. Ubiquitin ligases in oncogenic transformation and cancer therapy. Nat. Rev. Cancer 18, 69–88 (2018).
Deng, L., Meng, T., Chen, L., Wei, W. & Wang, P. The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct. Target. Ther. 5, 11 (2020).
Hoffmann, A., Cheng, G. & Baltimore, D. NF-κB: master regulator of cellular responses in health and disease. Immun. Inflamm. 1, 2 (2025).
Zhou, J. et al. The ubiquitin ligase MDM2 sustains STAT5 stability to control T cell-mediated antitumor immunity. Nat. Immunol. 22, 460–470 (2021).
Meng, X. et al. FBXO38 mediates PD-1 ubiquitination and regulates anti-tumour immunity of T cells. Nature 564, 130–135 (2018).
Franco, F., Jaccard, A., Romero, P., Yu, Y. R. & Ho, P. C. Metabolic and epigenetic regulation of T-cell exhaustion. Nat. Metab. 2, 1001–1012 (2020).
Yu, B. et al. Mitochondrial phosphatase PGAM5 modulates cellular senescence by regulating mitochondrial dynamics. Nat. Commun. 11, 2549 (2020).
Wang, Y. et al. In vitro and in vivo degradation of programmed cell death ligand 1 (PD-L1) by a proteolysis targeting chimera (PROTAC). Bioorg. Chem. 111, 104833 (2021).
Si, J. et al. Hematopoietic progenitor kinase1 (HPK1) mediates T cell dysfunction and is a druggable target for T cell-based immunotherapies. Cancer Cell 38, 551–566 (2020).
Zhao, X. et al. The transcriptional cofactor Tle3 reciprocally controls effector and central memory CD8(+ ) T cell fates. Nat. Immunol. 25, 294–306 (2024).
Scott-Browne, J. P. et al. Dynamic changes in chromatin accessibility occur in CD8(+) T cells responding to viral infection. Immunity 45, 1327–1340 (2016).
Man, K. et al. Transcription factor IRF4 promotes CD8(+) T cell exhaustion and limits the development of memory-like T cells during chronic infection. Immunity 47, 1129–1141 (2017).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8(+) T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Mognol, G. P. et al. Exhaustion-associated regulatory regions in CD8(+) tumor-infiltrating T cells. Proc. Natl Acad. Sci. USA 114, E2776–E2785 (2017).
Chen, J. et al. NR4A transcription factors limit CAR T cell function in solid tumours. Nature 567, 530–534 (2019).
Zadran, S., Arumugam, R., Herschman, H., Phelps, M. E. & Levine, R. D. Surprisal analysis characterizes the free energy time course of cancer cells undergoing epithelial-to-mesenchymal transition. Proc. Natl Acad. Sci. USA 111, 13235–13240 (2014).
Remacle, F., Kravchenko-Balasha, N., Levitzki, A. & Levine, R. D. Information-theoretic analysis of phenotype changes in early stages of carcinogenesis. Proc. Natl Acad. Sci. USA 107, 10324–10329 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Chen, S. et al. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell 160, 1246–1260 (2015).
Wei, J. et al. Targeting REGNASE-1 programs long-lived effector T cells for cancer therapy. Nature 576, 471–476 (2019).
Li, W. et al. Quality control, modeling, and visualization of CRISPR screens with MAGeCK-VISPR. Genome Biol. 16, 281 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Su, Y. et al. Multi-omic single-cell snapshots reveal multiple independent trajectories to drug tolerance in a melanoma cell line. Nat. Commun. 11, 2345 (2020).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Wu, T. et al. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Su, Y. et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell 185, 881–895 (2022).
Su, Y. et al. Multi-omics resolves a sharp disease-state shift between mild and moderate COVID-19. Cell 183, 1479–1495 (2020).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Acknowledgements
We thank W. Sheng (Zhejiang University), Q. Shan (Suzhou Institute of Systems Medicine) and L. Ye (Third Military Medical University) for providing experimental materials and technical support; and L. Yao and C. Wu (Xiamen University) for help with electron microscopy. The diagrams in this study were created using BioRender (https://www.biorender.com). This work was supported by the National Natural Science Foundation of China (grant nos 32525028 and 32270994 to G.L., 32571074 and 32300764 to H.C.), the Noncommunicable Chronic Diseases-National Science and Technology Major Project (grant no. 2024ZD0520600), the Basic Research Program of Jiangsu (grant nos BK20250003 to G.L., BK20230280 to H.C.), the Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (grant nos 2021-RC310-014 and 2024-JKCS-15 to G.L.), the CAMS Innovation Fund for Medical Sciences (CIFMS) (grant nos 2021-I2M-1-047, 2021-I2M-1-061, 2022-I2M-2-004, 2023-I2M-2-010, 2023-I2M-QJ-019, 2024-I2M-ZD-009 and 2025-I2M-GCC-006 to G.L., 2025-I2M-TS-15 to H.C.). We thank the Suzhou Municipal Key Laboratory (SZS2023005) and the NCTIB Fund for R&D Platform for Cell and Gene Therapy. P.D.G. was supported as an investigator by Parker Institute for Cancer Immunotherapy. Y.S. was supported by the Damon Runyon Quantitative Biology Fellowship from the Damon Runyon Cancer Research Foundation (grant no. DRQ-13-22), the Mahan Fellowship at Herbold Computational Biology Program of Fred Hutchinson Cancer Research Center, the Translational Data Science Integrated Research Center New Collaboration Award and Integrated Research Center at Fred Hutchinson Cancer Research Center, Pilot Award of Immunotherapy Integrated Research Center.
Author information
Authors and Affiliations
Contributions
H.C. and G.L. conceived of the project and designed experiments. H.C., Y.S. and X.P. performed most experiments and interpreted data. H.C., Y.S., R.G. and D.G.C. performed all bioinformatics analyses. H.C., Y.X., E.X., X.P. and J.D. performed the western blotting and LCMV infection. X.P., X.D. and E.X. designed and constructed most of the plasmids. H.C., G.L., P.D.G., Y.S. and X.P. wrote and edited the paper. G.L. and P.D.G. supervised the project and provided overall direction.
Corresponding authors
Ethics declarations
Competing interests
P.D.G. is a founder of and has received funding from Juno Therapeutics; is a founder and scientific advisory board member of, has equity in and has received research support from Affini-T Therapeutics; is an scientific advisory board member of and has equity in Immunoscape, Elpiscience, Earli, Metagenomi, Nextech and Catalio; and has received research support from Lonza. R.G. has received consulting income from Takeda, Sanofi and declares ownership in Ozette Technologies and stock options in Modulus Therapeutics. G.L., H.C. and X.P. have a pending patent application filed related to this work. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Russell Jones, Robert Manguso and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Visualization of transcriptome deconvolution into 2 gene modules.
a, Application of surprisal analysis to the RNA-seq data atlases related to T cell exhaustion. Samples from two representative T cell exhaustion studies (GSE89307 and GSE86881) were shown antigen-specific CD8+ T cells from acute Listeria infection and tumour progression (right), and from chronic LCMV infection (left). In the acute/tumour model, TCR-transgenic naïve T cells were transferred prior to pathogen or tumour induction, with effector/memory and early/late exhausted states sampled across defined time points. In the chronic model, naïve and late exhausted T cells were isolated from spleens following persistent LCMV infection. The combination of the gene expression baseline and the first two gene modules accurately recapitulated the experimentally measured transcriptome profiles. Heatmaps display genes positively or negatively associated with Module 1 or Module 2 (red = high, blue = low). Sample annotations follow standard naming conventions. b,c, GSEA enrichment of pathways based on genes positively or negatively associated in Module 1 (M1, b) and Module 2 (M2, c). GSEA uses a one-sided, permutation-based modified K–S test with adjustments for multiple comparisons. d,e, Module scores for M1 and M2 were calculated using a time-series T cell RNA-seq dataset of acute and chronic LCMV infection (GSE41867) (d, n = 4 independent samples), and projected onto a two-dimensional map with the x- and y-axes representing the respective module scores (e). f, Heat map showing expression of E3-related genes in adoptively transferred T cells from spleen, and in double negative (DN, PD-1−TIM-3−), single positive (SP, PD-1+TIM-3−), and double positive (DP, PD-1+TIM-3+) cells from B16-OVA tumours at day 14 after ACT (n = 3 independent samples). g, Percentages for E3 ligase-related genes predicted to be correlated (orange) or anti-correlated (blue) with exhaustion are shown. h, Volcano plot showing differential expression of E3-related genes between PD-1−TIM-3− and PD-1+TIM-3+ TILs. The x-axis represents the log2transformed fold change (FC) values, and the y-axis represents the negative log10 of adjusted P values (n = 3 independent samples). i, GSEA of ubiquitin-associated pathways in 400,000 human TILs with functional versus dysfunctional mitochondria (defined by the activity of mitochondrial complex I) across 21 cancer types. j, GSEA of ubiquitin-associated pathways in tumour-specific TILs with (MTDR/MTG)hi versus (MTDR/MTG)lo mitochondria from a mouse ACT model. k,l, Venn diagrams showing 78 E3 ligase genes negatively linked to T cell exhaustion, identified by overlapping differentially expressed genes from two gene modules (M1 and M2) with a curated E3 ligase list (k), and 133 E3 ligase genes positively associated with mitochondrial function, shared between mouse and human T cells (l).
Extended Data Fig. 2 KLHL6 deletion drives T cell exhaustion, but does not impact thymocyte development or peripheral T cell homeostasis.
a, Sanger sequencing of the Klhl6 locus in Cas9+sgRNA+ OT-I T cells transduced with sgCtrl, sgKlhl6-1, or sgKlhl6-2 in vitro. Primer sequences are listed in Supplementary Table 6. b,c, Mice were subcutaneously injected with 2×105 B16-OVA melanoma cells, followed by transfer of 3×106 Cas9+CD8+ OT-I T cells expressing control sgRNA or sgKlhl6 (two targets) on day 9. Tumours were collected and analyzed 7 days later. Experimental design (b). The number of transferred Cas9+ OT-I T cells in the tumour (c, n = 7 mice). d,e, Representative plots (left) and proportions (right) of LAG-3+TIM-3+ TILs (d), and cell numbers of indicated subsets (e) among sgCtrl- or sgKlhl6-transduced Cas9+CD8+ OT-I TILs (n = 7 mice). f,g, Proportions of PD-1+TIM-3+ (f) and (MTDR/MTG)lo (g) TILs among sgCtrl- or the indicated sgRNA-transduced Cas9+CD8+ OT-I T cells from B16-OVA tumours (n = 6 mice). h, Schematic of CRISPR/Cas9-mediated generation of Klhl6−/− mice. Guide RNA sequences are provided in Supplementary Table 6. i, Genotyping results for Klhl6+/+ (WT) or Klhl6−/− (KO) alleles. j,k, Percentages of thymocyte subsets: CD4−CD8− double-negative (DN), CD4+CD8+ double-positive (DP), CD4+ single-positive (CD4SP) and CD8+ single-positive (CD8SP) (j), and CD44+ single-positive (DN1), CD44+CD25+ double-positive (DN2), CD25+ single-positive (DN3) and CD44−CD25− double-negative (DN4) subpopulations (k) in DN cells of WT and KO mice (n = 6 mice). l, Numbers of CD19+, CD4+, and CD8+ cells in spleens of WT and KO mice (n = 6 mice). m, Percentages of naïve (CD44loCD62Lhi), central memory (CD44hiCD62Lhi; CM) and effector/effector memory (CD44hiCD62Llo, EFF/EM) CD8+ and CD4+ T cells in spleens of WT and KO mice (n = 6 mice). n,o, Proliferation of naïve WT and KO CD8+ T cells following anti-CD3/CD28 activation measured by CFSE dilution (n), and expression of activation markers (CD44, CD69 and CD25) at days 1-2 in vitro (o). p, Experimental design related to Fig. 2a–d. q-s, Percentages of transferred CD45.1/2+ OT-I T cells in dLN and spleen (q), and quantification of PD-1, TIM-3, LAG-3, and TOX expression OT-I TILs (r,s) at day 14 post-ACT (n = 14 mice). t, Survival curves of tumour-bearing mice after transfer of 5×106 CD45.1/2+ WT and KO OT-I T cells (n = 13 mice). Mice with tumour volumes >1500 mm3 were euthanized and defined as death. Diagram in b created in BioRender. Li, G. (2025) https://BioRender.com/aicmpdn. Diagram in p created in BioRender. Li, G. (2025) https://BioRender.com/0feebot. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (q-s), two-way ANOVA with Tukey’s multiple-comparisons test (c-g), two-way ANOVA with Sidak’s multiple-comparisons test (j-m), and Log-rank (Mantel-Cox) test (t). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 3 KLHL6 deficiency impairs mitochondrial fitness.
a,b, Heat maps of differentially expressed genes (a) and stemness-associated genes (b) between transferred WT and KO CD8+ T cells isolated from B16-OVA tumour-bearing mice at day 14 after ACT (n = 3 independent samples). c, Gene ontology analysis of differentially expressed genes in WT versus KO CD8+ TILs. Red and blue indicate molecular functions enriched in WT and KO cells, respectively. d,e, GSEA plots of memory and effector signatures (d) or cell cycle arrest and apoptosis signatures (e) for WT versus KO TILs. NES, normalized enrichment score. GSEA uses a one-sided, permutation-based modified K–S test with adjustments for multiple comparisons. f, Distinct mitochondrial biological processes in WT versus KO TILs at day 14 after ACT. g, WT and KO OT-I T cells were activated with anti-CD3/CD28 for 6 days. Mitochondrial mass and potential were measured using MTG (250 nM) and TMRE (50 nM), respectively. Relative fold changes of mitochondrial mass, membrane potential and TMRE/MTG ratio were calculated (n = 3 independent samples). h-k, Seahorse assays of OCR (h,i), SRC and mitochondrial ATP production (j), and glycoPER (k) in WT and KO OT-I T cells at day 6 after activation in vitro (n = 10 tests). l, Experimental design. Equal numbers (2×106 each) of CD45.1/2+ WT and CD45.1+ KO OT-I T cells were cotransferred into CD45.2+ mice bearing B16-OVA tumour (day 9 post-inoculation) and analyzed at day 14 after ACT. m, Analysis of glycoPER in WT and KO OT-I TILs from B16-OVA tumours at day 14 after ACT (n = 8 independent tests; 20 mice). n, Gating strategy for identifying (MTDR/MTG)hi and (MTDR/MTG)lo populations among cotransferred WT and KO OT-I TILs. Diagram in l created in BioRender. Li, G. (2025) https://BioRender.com/5se3g09. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (g,j) and two-way ANOVA with Sidak’s multiple-comparisons test (i,m). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001.
Extended Data Fig. 4 TCR stimulation downregulates KLHL6 level.
a, Relative Klhl6 mRNA levels in adoptively transferred OT-I T cells from spleens and PD-1−TIM-3−, PD-1+TIM-3− or PD-1+TIM-3+ CD8+ TILs from B16-OVA tumours at day 14 after ACT (n = 3 independent samples). b,c, Schematic of OT-I T cell restimulation in vitro. OT-I T cells were isolated from spleens, activated with anti-CD3/CD28, and cultured in mIL-2-containing media for 4 days. Cells were restimulated on day 4 with anti-CD3 (2 μg/mL, 24 h), OVA (1 μg/mL, 24 h), or PMA (50 ng/mL, 12 h) and compared to the un-restimulated Controls (b). Immunoblot and quantification of KLHL6 protein level in treated cells. Actin was used as a loading control (c, n = 3 independent samples). d,e, OT-I T cells were activated and expanded in the media containing mIL-2 for 4 days, then subjected to either repeated anti-CD3 restimulation every 2 days until day 9 (three restim) or a single restimulation on day 4 followed by resting in mIL-2 media until day 9 (one restim) (d). Western blot and quantification of KLHL6 expression were performed (e, n = 3 independent samples). f, Transcriptional expression of Tcf7, Pdcd1, and Klhl6 in P14 T cells during acute and chronic LCMV infection (GSE41867) (n = 4 independent samples). The boxplot spans from the first to the third quartile of the distribution, with the median positioned in the center. Whiskers represent the minimum and maximum values, excluding outliers. g, Klhl6 mRNA expression in antigen-specific T cells over time during acute infection and tumour progression (GSE89307) (n = 2-3 independent samples). h-j, Naïve CD45.1+P14+CD8+ T cells were adoptively transferred into CD45.2+ recipients (5×103 cells/mouse for LCMV-Clone 13 or 5×104 cells/mouse for LCMV-Armstrong). One day later, mice were infected with either virus. CD45.1+P14+ T cells were sorted from the spleen at the indicated time points post-infection for analysis. Experimental design (h). mRNA (i, n = 3 independent samples) and protein (j) levels of KLHL6 in CD45.1+P14+ T cells were analyzed at indicated time points. Diagram in b created in BioRender. Li, G. (2025) https://BioRender.com/9zoxyuh. Diagram in d created in BioRender. Li, G. (2025) https://BioRender.com/5se3g09. Diagram in h created in BioRender. Li, G. (2025) https://BioRender.com/l0bgjp0. Data are presented as mean ± s.e.m. Statistical analyses were determined by two-way ANOVA with Tukey’s multiple-comparisons test (a,c,e). **P < 0.01, ***P < 0.001, and ****P < 0.0001.
Extended Data Fig. 5 Enforced expression of KLHL6 restricts T cell exhaustion and promotes mitochondrial function.
a, Experimental design, related to Fig. 3a–e. b, Percentages of transferred Control and KLHL6-OE CD45.1+ OT-I T cells detected in the spleen and dLN on day 14 post-transfer (n = 7 mice). c, CellTrace Violet (CTV)-labeled CD45.1/2+ Control and CD45.1+ KLHL6-OE OT-I T cells were cotransferred into B16-OVA tumour-bearing recipient mice. TILs were analyzed on day 4 post-ACT (n = 5 mice). d-f, Proliferation (Ki-67hi) (d), apoptosis (Annexin V+) (e), and expression of apoptotic molecules (Bcl-2, Bcl-XL, and Bim) (f) in transferred Control and KLHL6-OE TILs at day 14 post-ACT (n = 6 mice). g,h, Percentages (g) and cell numbers (h) of PD-1−TIM-3−, PD-1+TIM-3− and PD-1+TIM-3+ populations in transferred CD45.1+CD8+ TILs (n = 7 mice). i,j, TOX (i) and TCF-1 (j) expression in CD8+PD-1+ TILs (n = 7 mice). k, Percentages of CD44+CD62L+ cells in spleen and dLN (n = 7 mice). l,m, Frequencies (l) and numbers (m) of Tpex (Ly108+TIM-3−) and Texterm (Ly108−TIM-3+) subsets in WT and KO TILs (n = 14 mice), related to Extended Data Fig. 2p. n,o, TOX (n) and PD-1 (o) expression in the Tpex and Texterm subsets at day 14 post-ACT (n = 7 mice). p-t, CD45.1/2+ KLHL6-OE and CD45.1+ Control OT-I T cells (2 × 106 each) were cotransferred into CD45.2+ mice bearing B16-OVA tumour. At day 14 post-ACT, TILs were isolated for metabolic analysis, including OCR (p), SRC (q), mitochondrial ATP production (r), glycoPER (s), and the percentage of (MTDR/MTG)lo cells (t) (Seahorse: n = 6 independent tests, 20 mice; (MTDR/MTG)lo subset: n = 8 mice). u-y, CD8+ OT-I T cells transduced with KLHL6-OE or Control retrovirus were analyzed on day 6 for MTG and TMRE staining (u, n = 3 independent samples), as well as OCR (v,w), SRC (x), and mitochondrial ATP production (y) (n = 10 tests). Diagram in a created in BioRender. Li, G. (2025) https://BioRender.com/aicmpdn. Data are presented as mean ± s.e.m. Statistical analyses were performed by unpaired two-tailed Student’s t-test (b-f,i-k,q,r,t,u,x,y) or two-way ANOVA with Sidak’s multiple-comparisons (g,h,l-p,s,w). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 6 Enforced KLHL6 expression enhances mouse and human T cell anti-tumour immunity.
a-d, Violin (a), dot (b), and UMAP (c,d) plots showing effector, exhaustion, and stem-like gene signatures, together with representative gene expression patterns and Texterm (upper) and Tpex (lower) profiles from scRNA-seq in Fig. 3g. The boxplot spans from the first to the third quartile of the distribution, with the median positioned in the center. Whiskers represent the minimum and maximum values, excluding outliers. Values plotted represent cells from a single replicate. e, Gene ontology analysis of differentially expressed genes between Control and KLHL6-OE TILs from scRNA-seq data. f,g, Experimental design as in Fig. 3k. Numbers of transferred Control and KLHL6-OE OT-I T cells in B16-OVA tumours on days 8 and 16 post-ACT (f) and tumour growth curves in B16-OVA-bearing mice receiving PBS, Control, or KLHL6-OE Tpex cells (g) (n = 7 mice). h-n, CD45.2+ mice were subcutaneously injected with B16-OVA melanoma cells. After 9 days, CD45.1+ Control and CD45.1/2+ KLHL6-OE OT-I T cells were mixed at a 1:1 ratio up to 5 million total cells and adoptively transferred into the tumour-bearing mice. Mice were sacrificed at days 21/28 post-transfer (n = 7 mice). Experimental design (h). Percentages and numbers of transferred T cells in tumour, dLN, spleen, and blood were measured (i-k). MFI of PD-1, TIM-3, and Ki-67 (l), cytokine production (m), and percentages of TCM (CD44+CD62L+) and TSCM (CD44−CD62L+CD95+) subsets (n) were determined at day 28. Polyfunctional subset indicates simultaneous expression of IL-2, IFNγ, and TNFα. o-t, NCG mice were subcutaneously implanted with HepG2-ESO cells. After 12 days, 6×106 activated 1G4 TCR-T cells transduced with KLHL6-OE or Control retrovirus were adoptively transferred. Mice were analyzed at day 16 post-transfer (n = 8 mice). Experimental design (o). Tumour weights (p), numbers of TCR-T cells in tumour and blood (q), MFI of LAG-3, TIM-3 and TOX (r) and cytokine production (TNFα and IFNγ) (s) in TCR-TILs, and frequencies of TCM (CD62L+CD45RA−) and Tem (CD62L−CD45RA−) subsets from spleens (t) were evaluated. Diagram in h created in BioRender. Li, G. (2025) https://BioRender.com/5se3g09. Diagram in o created in BioRender. Li, G. (2025) https://BioRender.com/5se3g09. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (l-n,p-r), two-way ANOVA with Sidak’s multiple-comparisons test (f,i-k,s,t), or two-way ANOVA with Tukey’s multiple-comparisons test (g). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 7 KLHL6 loss promotes Texterm cell differentiation during chronic infection.
a, Naïve Klhl6+/+ (WT) and Klhl6−/− (KO) CD8+ P14 T cells were mixed at a 1:1 ratio (5×103 for LCMV-CL13 or 5×104 for LCMV-Arm) and transferred into CD45.2+ recipients, followed by intravenous LCMV infection one day later. Activation markers in transferred WT and KO P14 T cells from spleen were assessed at days 4 and 6 post-infection (p.i.). b,c, Naïve CD45.1/2+ WT and CD45.1+ KO P14 cells were cotransferred (2.5×103 each) into CD45.2+ mice, followed by LCMV-CL13 infection one day later. Mice were sacrificed on days 8 and 28 p.i. (n = 6 mice). Experimental design (b), and percentages (left) and absolute numbers (right) of WT and KO P14 cells in spleen (c). d,e, Frequencies and numbers of “effector-like” (TCF-1−GzmB+ or Ly108−TIM-3+) and Texprec (TCF-1+GzmB− or Ly108+TIM-3−) subsets in transferred WT and KO P14 T cells from spleens at day 8 p.i. (n = 6 mice). f-h, Percentages of TNFα+IFNγ+ cells (f), (MTDR/MTG)lo subsets (g), and apoptotic cells (Annexin V+PI+; Annexin V+PI−) (h) in transferred WT and KO P14 cells from spleens at day 8 p.i. of CL13-infected mice (n = 6 mice). i, Quantification of TIM-3, PD-1, TOX and GzmB expression and the percentage of TCF-1+ cells at day 28 p.i. (n = 6 mice). j,k, Frequencies and numbers of Tpex (Ly108+CX3CR1−), Texint (Ly108−CX3CR1+) and Texterm (Ly108−CX3CR1−) subsets (j), and Bcl-2/Bim expression in these subsets (k) at day 28 p.i. (n = 6 mice). l-n, Frequencies and numbers of Tpex1 (Ly108+CD69+), Tpex2 (Ly108+CD69−), Texint (Ly108−CD69−), and Texterm (Ly108−CD69+) subsets (l), TOX expression in these subsets (m), and cytokine production (TNFα and IFNγ) after PMA/BFA stimulation (n) in transferred WT and KO P14 cells at day 28 p.i. (n = 6 mice). o, Mice receiving PBS, WT P14, or KO P14 cells were infected with LCMV-CL13 one day after transfer, and LCMV viral loads in liver and lungs were measured on day 15 p.i., normalized to the PBS group. LCMV titers were assessed by qPCR relative to HPRT (PBS, n = 6 mice; WT and KO, n = 8 mice). Diagram in b created in BioRender, Li. G (2025) https://BioRender.com/l0bgjp0. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (c-n), or two-way ANOVA with Tukey’s multiple-comparisons test (o). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 8 Enforced KLHL6 expression restrains exhaustion and boosts anti-viral T cell responses.
a, Experimental design. Activated CD45.1+ P14 T cells transduced with KLHL6-OE or Control retrovirus were separately transferred into CD45.2+ recipients (5×103 cells/recipient), followed by LCMV-CL13 infection one day later. Mice were sacrificed at days 8 and 21 p.i. b,c, Representative plots and frequencies of “effector-like” (TCF-1−GzmB+ or Ly108−TIM-3+) and Texprec (TCF-1+GzmB− or Ly108+TIM-3−) subsets in transferred Control and KLHL6-OE P14 T cells from spleens at day 8 p.i. (n = 6 mice). d, Numbers of TCF-1+GzmB−, TCF-1−GzmB+ and total P14 T cells from spleen at day 8 p.i. (n = 6 mice). e, Quantification of Annexin V+ P14 cells at day 8 p.i. (n = 6 mice). f,g, Cytokine production (TNFα and IFNγ) (f) and relative MFI of TIM-3, PD-1 and TCF-1 (g) in KLHL6-OE versus Control P14 T cells from spleens at day 21 p.i. (n = 6 mice). h-j, Frequencies (h) and absolute numbers (i) of Tpex1, Tpex2, Texint and Texterm subsets, and TOX expression in these subsets (j) (n = 6 mice). k,l, Frequencies (k) and numbers (l) of Tpex (Ly108+CX3CR1−), Texint (Ly108−CX3CR1+), and Texterm (Ly108−CX3CR1−) subsets in spleens at day 21 p.i. (n = 6 mice). m, Mice receiving PBS, Control, and KLHL6-OE P14 T cells were infected with LCMV-CL13 one day after adoptive transfer, and LCMV viral loads in liver and lungs were measured on day 15 p.i., normalized to the PBS group. LCMV titers were assessed by qPCR relative to HPRT (PBS, n = 5 mice; Control and KLHL6-OE, n = 8 mice). Diagram in a created in BioRender. Li, G. (2025) https://BioRender.com/l0bgjp0. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (b-l), or two-way ANOVA with Tukey’s multiple-comparisons test (m). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 9 KLHL6 mediates Lys48-linked poly-ubiquitination and degradation of TOX.
a, Schematic of the E-STUB system. KLHL6 was fused to the biotin ligase BirA and co-expressed with BAP-tagged ubiquitin in cells, enabling biotinylation of substrates proximal to BirA-KLHL6 in a ubiquitin-specific manner. b, Rank-ordered plot of MS results showing streptavidin-enriched proteins from an E-STUB assay in Jurkat cells expressing KLHL6-BirA versus those expressing Empty-BirA. c, Jurkat cells co-expressing BAP-tagged ubiquitin with either KLHL6-BirA or Empty-BirA were treated with or without biotin (50 μM) for 15 min. Biotinylated proteins were then enriched using streptavidin beads and detected by immunoblotting. Braces indicate bands related to the target proteins. d, Interaction between KLHL6-Myc and endogenous TOX in Jurkat and EL4 cells. e,f, Schematic of full-length TOX protein (WT) and truncated mutants (ΔC1, ΔC2, and ΔN) (e), and requirement of the C-terminal domain for KLHL6 interaction (f). g, Western blot of Flag-TOX degradation in HEK293T cells transfected with empty vector or different dosages (0.5, 1, and 1.5 μg) of KLHL6-Myc. h, Endogenous TOX levels in human T cells transduced with KLHL6-Myc and cultured with or without anti-CD3 stimulation (n = 3 independent samples). i, Ubiquitination of exogenous TOX in HEK293T cells cotransfected with HA-Ub and KLHL6-Myc. j, Ubiquitination of endogenous TOX in Jurkat cells with KLHL6 overexpression. k, Ubiquitination analysis of endogenous TOX in activated mouse and human T cells upon anti-CD3 restimulation. l, Measurement of TOX ubiquitination with mutant Ub (K48R, K63R) in HEK293T cells. m, Identification of the key TOX sites responsible for KLHL6-mediated degradation. n-p, Ubiquitination (n), interaction (o) and degradation (p, n = 3 independent samples) of a quadruple mutation (4KR) of TOX in HEK293T cells cotransfected with KLHL6-Myc. A triple mutant (K245, K246, and K248) was defined as ‘3KR’, and the quadruple mutant (K245, K246, K248, and K323) defined as ‘4KR’. q, Evaluation of degradation of the TOX quadruple mutant (4KR) in HEK293T cells treated with CHX (50 µg/mL) for the indicated times (n = 3 independent samples). Diagram in a created in BioRender. Li, G. (2025) https://BioRender.com/va0cyy2. Data in (c,d,f,g,i-o) are representative of two independent experiments. Data are presented as mean ± s.e.m. Statistical analyses were determined by unpaired two-tailed Student’s t-test (p,q), or two-way ANOVA with Tukey’s multiple-comparisons test (h). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 10 A TOX mutant resistant to KLHL6-mediated degradation reinforces the exhaustion phenotype in T cells.
a, Gating strategy, histogram plots, and quantification of TIM-3 and TCF-1 levels in TOXloPD-1lo, TOXloPD-1hi, and TOXhiPD-1hi OT-I TIL populations from B16-OVA tumours at day 14 post-transfer (n = 7 mice). b, Percentages of Tpex (Ly108+TIM-3−) and Texterm (Ly108−TIM-3+) populations in TOXlo and TOXhi TILs at day 14 (n = 7 mice). c,d, Numbers of CD45.1+ OT-I TILs (c) and TOX expression levels (d) in WT+shCtrl, KO+shCtrl, and KO+shTox groups at day 14 after ACT (n = 8 mice). e, Correlation between KLHL6 expression and the indicated genes in human TILs from melanoma, renal cancer, breast cancer, and nasopharyngeal cancer, using human pan-cancer scRNA-seq data. CD8+ TILs were stratified into KLHL6hi and KLHL6lo groups. Selected genes were quantified and visualized with red indicating highly expressed and blue indicating lower expressed. f-k, Naïve CD8+ OT-I T cells were activated with anti-CD3/CD28 for 24 h, and transduced with TOX(WT) (TOXWT-OE), TOX(4KR) (TOX4KR-OE) or Empty vector (Control). Cells were cultured for 5 days in vitro before subsequent analyses. Experimental design (f), percentages of PD-1+LAG-3+ populations (g), TOX, PD-1, and LAG-3 levels (h), cytokine production (i), heatmap of differentially expressed genes (j), and GSEA for exhaustion and effector signatures (k) (n = 3 independent samples). GSEA uses a one-sided, permutation-based modified K–S test with adjustments for multiple comparisons. l-o, 3×106 Control, TOX(WT), or TOX(4KR) OT-I T cells were adoptively transferred into B16-OVA tumour-bearing mice. Mice were sacrificed on day 14 after ACT. TOX expression (l), percentages of PD-1−TIM-3−, PD-1+TIM-3− and PD-1+TIM-3+ populations (m), MFI of LAG-3, TIM-3, and PD-1 (n), and cytokine production (o) in CD45.1+CD8+ TILs were assessed (n = 6 mice). Diagram in e created in BioRender. Li, G. (2025) https://BioRender.com/p3754eu. Diagram in f created in BioRender. Li, G. (2025) https://BioRender.com/ahcb92h. Data are presented as mean ± s.e.m. Statistical analyses were performed by two-way ANOVA with Sidak’s multiple-comparisons test (b) or with Tukey’s multiple-comparisons test (a,c,d,g-i,l-o). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 11 KLHL6 modulates mitochondrial fitness and anti-tumour T cell responses via ubiquitinating PGAM5.
a, Immunoblot of mitochondrial fusion/fission proteins in WT or KLHL6 KO OT-I T cells activated by anti-CD3/CD28 for indicated times (n = 3 independent samples). b, Immunoblot of p62 and LC3 (upper band: LC3-I; lower band: LC3-II) in WT and KO OT-I T cells at day 5 post-activation. c, CD45.2+ B16-OVA-bearing mice received 4×106 WT or KO CD45.1+ OT-I cells pretreated with DMSO or M1 (20 μM) + Mdivi-1 (10 μM) (MM) for 3 days. Survival of mice was monitored (n = 8 mice). d, Interaction between introduced KLHL6-Myc and endogenous PGAM5 in Jurkat cells. e, Ubiquitination of endogenous PGAM5 in Jurkat cells transduced with KLHL6-Myc. f,g, Immunoblot analysis of endogenous PGAM5 in Jurkat cells expressing an empty vector or KLHL6-Myc with or without MG132 treatment (f) and in WT and KO T cells collected at different time points after activation (g). h,i, WT and KLHL6 KO OT-I T cells transduced with shPgam5 (shP5) or shCtrl retrovirus were cultured in vitro for 5 days. TMRE/MTG ratio (h, n = 5 independent samples) and mitochondrial morphology and area (i; scale bar, 1 μm; n = 60 cells) were analyzed. j-m, CD45.1/2+ WT and CD45.1+ KLHL6 KO OT-I T cells transduced with GFP-shCtrl or Thy1.1-shPgam5 (shP5) were mixed at a 1:1:1:1 ratio and cotransferred into CD45.2+ B16-OVA tumour-bearing mice. Mice were sacrificed for analysis on day 14 after ACT. Experimental design (j), CD8+ TIL numbers (k), PD-1 and TIM-3 expression in TILs (l), and numbers of CD44+CD62L+ cells in dLN and spleen from the four groups (m) (n = 7 mice). n-r, WT and KO OT-I T cells were activated and cultured for 3 days in vitro, then treated with DMSO or the PGAM5 inhibitor LFHP-1c (P5i, 2 μM) for another 3 days before analysis. Immunoblotting of Mfn2, Opa1, Drp1, p-Drp1S637, PGAM5, and Actin (n). OCR, SRC, mitochondrial ATP production (o-q, n = 10 tests), and TMRE/MTG ratio (r, n = 3 independent samples) were assessed. s-y, CD45.2+ mice bearing B16-OVA tumours received 4×106 activated WT or KO CD45.1+ OT-I cells pretreated with DMSO or P5i. The mice were sacrificed for analysis at day 14 after ACT. Schematic of the experiment (s), tumour weights (t), total numbers (u), cytokine production (v), PD-1 and TIM-3 levels (w), and cell numbers of indicated subsets (x) of OT-I TILs were evaluated (n = 6 mice); percentages of TCM populations in dLN and spleen (y, n = 6 mice). Diagram in j created in BioRender. Li, G. (2025) https://BioRender.com/md3c1bz. Diagram in s created in BioRender. Li, G. (2025) https://BioRender.com/ap5vq6h. Data in (b,d-g,n) are representative of three independent experiments. Data are presented as mean ± s.e.m. Statistical analyses were determined by two-way ANOVA with Tukey’s multiple-comparisons test (a,h,i,k-m,o-r,t-y) and Log-rank (Mantel-Cox) test (c). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Extended Data Fig. 12 TOX and mitochondrial fitness are key mediators of KLHL6-driven anti-tumour immune responses.
a,b, CD8+ WT and KLHL6 KO T cells transduced with either Empty vector (Empty) or PGC1α-overexpression (PGC1α) plasmids were cultured for 6 days. OCR (a, n = 10 tests) and SRC and mitochondrial ATP production (b, n = 10 tests) were measured. c-e, Tumour weights (c), absolute numbers of transferred CD8+ TILs (d), and proportion of damaged mitochondria in transferred CD8+ TILs (e) were assessed in the indicated groups (n = 6 mice), related to Fig. 5l,m. f-i, Experimental design related to Fig. 5n,o (f), tumour weights (g), absolute numbers of total (h) and Tpex (Ly108+TIM-3−) and Texterm (Ly108−TIM-3+) subsets (i) of transferred CD8+ TILs among the five groups at day 14 after ACT (n = 6 mice). j-n, CD8+ OT-I T cells transduced with either TOX-overexpressing (TOX-OE) or Empty vector (Control) retrovirus were cultured for 5 days in vitro, and then restimulated with or without CD3 antibody for 24 h before analysis. Representative plots and quantification of mitochondrial depolarization (TMRE/MTG)lo (j) and MitoSOX level (k) were assessed (n = 3 independent samples). OCR (l), SRC (left) and mitochondrial ATP production (right) (m), and glycoPER (n) were measured (n = 10 tests). o,p, Activated WT and KO CD8+ OT-I T cells were transduced with either shCtrl or shTox retrovirus at 24 h post-activation and cultured for an additional 5 days prior to analysis. OCR (o), SRC (left) and mitochondrial ATP production (right) (p) were measured (n = 8 tests). Diagram in f created in BioRender. Li, G. (2025) https://BioRender.com/md3c1bz. Data in (f-i) are representative of two independent experiments. Data are presented as mean ± s.e.m. Statistical analyses were determined by two-way ANOVA with Tukey’s multiple-comparisons test (a-e,g-p). *P < 0.05, **P < 0.01, ***P < 0.001, and ****P < 0.0001; ns, not significant.
Supplementary information
Supplementary Fig. 1 (download PDF )
Raw western blot images.
Supplementary Tables 1–8 (download ZIP )
Supplementary Table 1: Datasets used for module 1 and module 2 analysis. Supplementary Table 2: The gene contribution score to modules 1 and 2. Supplementary Table 3: GSEA and custom analysis for modules 1 and 2. Supplementary Table 4: Selection of E3-associated genes for CRISPR screening. Supplementary Table 5: Statistical analysis of in vivo CRISPR screens. Supplementary Table 6: The primers used in this study. Supplementary Table 7: Mass spectrometry results of E-STUB. Supplementary Table 8: The antibodies used in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheng, H., Su, Y., Pan, X. et al. The ubiquitin ligase KLHL6 drives resistance to CD8+ T cell dysfunction. Nature 651, 451–461 (2026). https://doi.org/10.1038/s41586-025-09926-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09926-8
This article is cited by
-
Chronic TCR signaling-driven suppression of the FOXO1-KLHL6 axis promotes T cell exhaustion
Immunity & Inflammation (2026)