Abstract
Cereblon (CRBN) is the target of thalidomide derivatives1 that achieve therapeutic efficacy against some haematologic neoplasias2,3,4 by recruiting neosubstrates for degradation5,6,7. Despite the intense investigation of orthosteric thalidomide derivatives, little is known about alternate binding sites on CRBN. Here we report an evolutionarily conserved cryptic allosteric binding site on CRBN. Small-molecule SB-405483 binds the allosteric site to cooperatively enhance the binding of orthosteric ligands and alter their neosubstrate degradation profiles. A survey of over 100 orthosteric ligands and their degradation targets reveals trends in the classes of compounds and neosubstrates in which degradation outcomes are enhanced or inhibited by SB-405483. Structural investigations provide a mechanistic basis for the effects of the allosteric ligand by shifting the conformational distribution of CRBNopen to a novel CRBNint and increasing the CRBNclosed state. The discovery of a cryptic allosteric binding site on CRBN that alters the functional effects of orthosteric ligands opens new directions with broad implications for improving the selectivity and efficacy of CRBN therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data generated or analysed during this study are available in the main text or Supplementary Information. Uncropped, full western blot images and gels are provided in Supplementary Figs. 1–6. Proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD052094 and PXD067391. Proteomics raw data were searched against SwissProt Human (Homo sapiens) protein database (21 February 2019; 20,355 total entries). Cryo-EM maps and associated atomic models were deposited to the Electron Microscopy Data Bank (EMDB) and the Protein Data Bank (PDB), respectively, with the following accession numbers: CRBNclosed/DDB1 bound to lenalidomide and SB-405483: PDB 9SFM. CRBNopen/DDB1ΔBPB: EMD-70778 (composite map), EMD-70776 (consensus map) and EMD-70777 (focus map). CRBNint/DDB1ΔBPB: EMD-70781 (consensus map). CRBNclosed/DDB1ΔBPB: EMD-70784 (composite map), EMD-70782 (consensus map) and EMD-70783 (focus map). CRBNCK1α/DDB1ΔBPB bound to lenalidomide and SB-405483: EMD-70790 (composite map), EMD-70788 (consensus map) and EMD-70789 (focus map). CRBNIKZF1(ZF2)/DDB1ΔBPB bound to lenalidomide and SB-405483: EMD-70799 (composite map), EMD-70795 (consensus map) and EMD-70796 (focus map). CRBNCK1α/DDB1ΔBPB bound to DEG-47: EMD-70804 (composite map), EMD-70801 (consensus map), EMD-70802 (focus map on CRBN) and EMD-70803 (focus map on CK1α). CRBNCK1α/DDB1ΔBPB bound to DEG-47 and SB-405483: EMD-70862 (composite map), EMD-70827 (consensus map), EMD-70835 (focus map on CRBN) and EMD-70836 (focus map on CK1α), PDB 9OTY. CRBNIKZF1(ZF2)/DDB1ΔBPB bound to DEG-47 and SB-405483: EMD-70870 (composite map), EMD-70868 (consensus map) and EMD-70869 (focus map on CRBN and IKZF1), PDB 9OUL. CRBNIKZF1(ZF2)/DDB1ΔBPB bound to DEG-47: EMD-70867 (composite map), EMD-70865 (consensus map) and EMD-70866 (focus map on CRBN and IKZF1), PDB 9OUK. Source data are provided with this paper.
References
Ito, T. et al. Identification of a primary target of thalidomide teratogenicity. Science 327, 1345–1350 (2010).
Kronke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2014).
Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305–309 (2014).
Kronke, J. et al. Lenalidomide induces ubiquitination and degradation of CK1α in del(5q) MDS. Nature 523, 183–188 (2015).
Winter, G. E. et al. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348, 1376–1381 (2015).
Lu, J. et al. Hijacking the E3 ubiquitin ligase cereblon to efficiently target BRD4. Chem. Biol. 22, 755–763 (2015).
Nowak, R. P. et al. Plasticity in binding confers selectivity in ligand-induced protein degradation. Nat. Chem. Biol. 14, 706–714 (2018).
Donovan, K. A. et al. Thalidomide promotes degradation of SALL4, a transcription factor implicated in Duane Radial Ray syndrome. eLife 7, e38430 (2018).
Matyskiela, M. E. et al. SALL4 mediates teratogenicity as a thalidomide-dependent cereblon substrate. Nat. Chem. Biol. 14, 981–987 (2018).
Oleinikovas, V., Gainza, P., Ryckmans, T., Fasching, B. & Thoma, N. H. From thalidomide to rational molecular glue design for targeted protein degradation. Annu. Rev. Pharmacol. Toxicol. 64, 291–312 (2024).
Sasso, J. M. et al. Molecular glues: the adhesive connecting targeted protein degradation to the clinic. Biochemistry 62, 601–623 (2023).
Ichikawa, S. et al. The E3 ligase adapter cereblon targets the C-terminal cyclic imide degron. Nature 610, 775–782 (2022).
Beyett, T. S. et al. Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors. Nat. Commun. 13, 2530 (2022).
Schoepfer, J. et al. Discovery of asciminib (ABL001), an allosteric inhibitor of the tyrosine kinase activity of BCR-ABL1. J. Med. Chem. 61, 8120–8135 (2018).
Liu, J. & Nussinov, R. The role of allostery in the ubiquitin-proteasome system. Crit. Rev. Biochem. Mol. Biol. 48, 89–97 (2013).
Watson, E. R. et al. Molecular glue CELMoD compounds are regulators of cereblon conformation. Science 378, 549–553 (2022).
Orlicky, S. et al. An allosteric inhibitor of substrate recognition by the SCFCdc4 ubiquitin ligase. Nat. Biotechnol. 28, 733–737 (2010).
Petzold, G., Fischer, E. S. & Thoma, N. H. Structural basis of lenalidomide-induced CK1α degradation by the CRL4CRBN ubiquitin ligase. Nature 532, 127–130 (2016).
Hulme, E. C. & Trevethick, M. A. Ligand binding assays at equilibrium: validation and interpretation. Br. J. Pharmacol. 161, 1219–1237 (2010).
Christopoulos, A. & Kenakin, T. G protein-coupled receptor allosterism and complexing. Pharmacol. Rev. 54, 323–374 (2002).
Fischer, E. S. et al. Structure of the DDB1–CRBN E3 ubiquitin ligase in complex with thalidomide. Nature 512, 49–53 (2014).
Chamberlain, P. P. et al. Structure of the human cereblon–DDB1–lenalidomide complex reveals basis for responsiveness to thalidomide analogs. Nat. Struct. Mol. Biol. 21, 803–809 (2014).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Groisman, R. et al. The ubiquitin ligase activity in the DDB2 and CSA complexes is differentially regulated by the COP9 signalosome in response to DNA damage. Cell 113, 357–367 (2003).
Ohtake, F. et al. Dioxin receptor is a ligand-dependent E3 ubiquitin ligase. Nature 446, 562–566 (2007).
Riching, K. M. et al. Quantitative live-cell kinetic degradation and mechanistic profiling of PROTAC mode of action. ACS Chem. Biol. 13, 2758–2770 (2018).
Miyamoto, D. K. et al. Design and development of IKZF2 and CK1α dual degraders. J. Med. Chem. 66, 16953–16979 (2023).
Park, S. M. et al. Dual IKZF2 and CK1α degrader targets acute myeloid leukemia cells. Cancer Cell 41, 726–739 (2023).
Razumkov, H. et al. Discovery of CRBN-dependent WEE1 molecular glue degraders from a multicomponent combinatorial library. J. Am. Chem. Soc. 146, 31433–31443 (2024).
Ting, P. Y. et al. A molecular glue degrader of the WIZ transcription factor for fetal hemoglobin induction. Science 385, 91–99 (2024).
Russell, P. & Nurse, P. Negative regulation of mitosis by wee1+, a gene encoding a protein kinase homolog. Cell 49, 559–567 (1987).
Schwinn, M. K. et al. CRISPR-mediated tagging of endogenous proteins with a luminescent peptide. ACS Chem. Biol. 13, 467–474 (2018).
Sievers, Q. L. et al. Defining the human C2H2 zinc finger degrome targeted by thalidomide analogs through CRBN. Science 362, eaat0572 (2018).
Słabick, M. et al. Expanding the druggable zinc-finger proteome defines properties of drug-induced degradation. Mol. Cell 85, 3184–3201.e14 (2025).
Lindner, S. et al. Chemical inactivation of the E3 ubiquitin ligase cereblon by pomalidomide-based homo-PROTACs. J. Vis. Exp. 147, e59472 (2019).
Ichikawa, S. et al. The cyclimids: degron-inspired cereblon binders for targeted protein degradation. Cell Chem. Biol. 31, 1162–1175 (2024).
Donovan, K. A. et al. Mapping the degradable kinome provides a resource for expedited degrader development. Cell 183, 1714–1731 (2020).
Petzold, G. et al. Mining the CRBN target space redefines rules for molecular glue–induced neosubstrate recognition. Science 389, eadt6736 (2025).
Conn, P. J., Christopoulos, A. & Lindsley, C. W. Allosteric modulators of GPCRs: a novel approach for the treatment of CNS disorders. Nat. Rev. Drug Discov. 8, 41–54 (2009).
Wang, J., Gareri, C. & Rockman, H. A. G-protein-coupled receptors in heart disease. Circ. Res. 123, 716–735 (2018).
Adrian, F. J. et al. Allosteric inhibitors of Bcr-abl-dependent cell proliferation. Nat. Chem. Biol. 2, 95–102 (2006).
Wylie, A. A. et al. The allosteric inhibitor ABL001 enables dual targeting of BCR-ABL1. Nature 543, 733–737 (2017).
Acknowledgements
We thank members of the Woo laboratory for their helpful discussions, M. Leon Duque for insight on proteomics, D. Miyamoto for DEG library compounds, R. Mazitschek at Massachusetts General Hospital for discussions on TR-FRET experiments, S. Bedard for providing training in performing the dynamic light scattering assay, J. Kozole for insight and discussions on ASMS experiments, C. Martinez Fleites for support with data analysis, W. Lessin at the Scripps Research Electron Microscopy Facility for microscopy support, J. C. Ducom at Scripps Research High Performance Computing for computational support, the Harvard Mass Spectrometry and Proteomics Resource Laboratory for mass spectrometry support, the Harvard Bauer Flow Cytometry core for flow cytometry support, the Laukien–Purcell Instrumentation Center Magnetic Resonance Laboratory for their nuclear magnetic spectroscopy support and Viva Biotech for protein preparation and crystallographic support. We are grateful to L. Gerbert from I. MacRae’s laboratory for assistance with the CK1α protein. U937 Cas9 cells stably expressing SALL4(ZF1,2)-GFP were a generous gift from M. Słabicki’s laboratory at Massachusetts General Hospital Cancer Center and the Broad Institute. Support from the Blavatnik Biomedical Accelerator at Harvard (C.M.W.), Starr Cancer Consortium (C.M.W.), Merkin Family Foundation (C.M.W.), Promega Academic Access Program (C.M.W.), National Science Foundation Graduate Research Fellowship (V.N.D., X.B.T., F.K.) and National Institutes of Health (NIH) GM143805 and GM154216 (G.C.L.) is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
C.M.W. and A.B.B. conceived and oversaw the project. V.N.D. designed and performed the thermal shift assay, NanoBRET assays and cell viability assays. V.N.D. and I.F.A. designed and performed degradation assays. W.X. and V.N.D. designed and performed proteomics experiments, and analysed the proteomics data jointly with C.M.W. I.F.A. designed and performed HiBiT experiments. X.B.T. supported preparation of SB-405483. N.M.C. designed and synthesized DEG-47. F.K. and V.N.D. generated and validated MOLM-13 CRBN knockout cells. K.-H.C. generated and validated A2780 Wee1-HiBiT cells. A.E.C. designed and performed the TR-FRET assay and the dynamic light scattering assay. A.E.C. and A.J.J. analysed the high-throughput screen results. M.A.Q. conceived high-throughput screen experiments. A.E.C. and J.L.S. calculated the allosteric parametric data. C.-w.C., M.A.Q. and H.D. designed, analysed and interpreted the crystallography data, identified the allosteric site and proposed the mechanistic hypothesis for allosteric modulation. A.E.C. and C.M. designed and performed ASMS experiments, and analysed the data together with J.L.S. Z.R., W.C. and G.C.L. worked on the cryo-EM section of this project. Z.R. generated the constructs, purified the proteins and performed all cryo-EM structure determination, model building and refinement and mechanistic interpretation, with guidance from G.C.L. W.C. helped in data processing for the initial one dataset that helped identify the CRBN intermediate state. V.N.D., C.M.W., C.-w.C., A.E.C., J.L.S., M.A.Q., A.B.B., Z.R. and G.C.L. drafted the manuscript. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The Woo laboratory receives or has received support from Amgen, Ono Pharmaceuticals and Merck. A.E.C., C.-w.C., A.J.J., J.L.S., K.-H.C., M.A.Q., C.M., H.D. and A.B.B. are/were employees and/or shareholders of GSK. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Alessio Ciulli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 SB-405483 binds allosterically to CRBN in vitro.
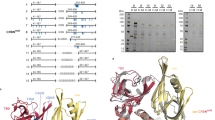
(a) Schematic of TR-FRET competition assay. (b) ASMS data demonstrating binding of both orthosteric and allosteric ligands. (c) ASMS signal for SB-405483 over increasing concentrations of SB-405483 and pomalidomide. (d) ASMS signal for pomalidomide over increasing concentrations of SB-405483 and pomalidomide. (e) TR-FRET IC50 values of orthosteric ligands in the presence or absence of 25 µM SB-405483. (f) X-ray maps around the ligand. OMIT difference map (Fo-Fc), after removal of the ligand from the final X-ray structure model, is shown. The map is contoured at +/- 3 sigma, in blue and green respectively. The modelling of the binding mode of ligands are unambiguous with all elements of the molecule clearly defined. (g) Side view of allosteric site. (h) Top view of allosteric binding site. (i) Bottom view of allosteric binding site. (j) 4CI2 overlaid with SB-405483. (k) 4TZ4 overlaid with SB-405483. (l) Crystal structure of human CRBN/DDB1 with SB-405483. (m) Front and rear site view of allosteric binding site from the crystal structure obtained in this study. Residues in grey are 100% conserved across species. Residues in dark red are conserved in 75-99% of species. Residues in pink are conserved in 50-74% of species. (n) AlphaFold structure of Mus musculus CRBN allosteric site, X-ray structure of Gallus gallus CRBN allosteric site (PDB: 4CI2), and AlphaFold structure of Danio rerio CRBN allosteric site with residues that differ from the Homo sapiens CRBN allosteric site in red. ASMS data are the mean values resulting from n = 2 biologically independent samples. TR-FRET data are mean ± s.d. from n = 4 biologically independent samples.
Extended Data Fig. 2 Functional engagement of CRBN by SB-405483 in cells.
(a) Western blot of CRBN levels after 1 h treatment of MM.1S cells with varying concentrations of SB-405483 in the presence or absence of lenalidomide and exposure to 57 °C for 5 min. (b) Quantification of n = 3 biologically independent replicates of a. (c) Western blot of CRBN levels after 1 h treatment of MM.1S cells with varying concentrations of lenalidomide in the presence or absence of SB-405483 and exposure to 57 °C for 5 min. (d) Quantification of n = 3 biologically independent replicates of c. (e) Western blot of CRBN levels after treatment of MM.1S cells with lenalidomide in the presence or absence of SB-405483 for 1 h and exposure to varying temperature for 5 min. (f) Quantification of n = 3 biologically independent replicates of e. (g) Western blot of ubiquitin and CRBN levels following affinity purification of Flag-CRBN from HEK-CRBN cells. Len, lenalidomide. (h) Quantification of n = 3 biologically independent replicates of ubiquitin blots of g. (i) Schematic of NanoBRET assay. (j) NanoBRET assay with lenalidomide and SB-405483. (k) NanoBRET assay with CC-90009 in indicated mutant CRBN-nLuc. (l) NanoBRET assay with dBET6 in indicated mutant CRBN-nLuc. Comparisons were performed using a one-way ANOVA with Šídák’s multiple comparisons test. P-values are shown. Data are mean ± s.d. All Western blot and NanoBRET data are representative of n = 3 biologically independent samples. For uncropped Western blot images, see Supplementary Fig. 2.
Extended Data Fig. 3 SB-405483 potentiates the degradation and antiproliferative effects of CK1α.
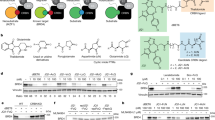
(a) Western blot of CK1α levels after treatment of KG-1α cells with lenalidomide in the presence or absence of SB-405483 for 24 h. (a) Quantification of n = 3 biologically independent replicates of b. (c) Western blot of CK1α levels after treatment of MOLM-13 cells with increasing concentration of DEG-47 in the presence or absence of SB-405483 for 24 h. (d) Protein expression levels of CK1α after treatment with indicated compounds in MOLM-13 cells for 24 h (n = 3 biologically independent replicates). (e) Flow cytometry analysis of GFP levels in HEK293T cells expressing GFP-CK1α after treatment with DEG-47 in the presence and absence of SB-405483 for 24 h. (f) Western blot of CK1α levels after MOLM-13 treatment with DEG-47 with and without SB-405483 at various time points. (g) Quantification of CK1α levels after MOLM-13 treatment with DEG-47 in the presence or absence of SB-405483 at various time points. (h) Cell viability (MTT) assay of the indicated compounds after treatment of MOLM-13 cells for 3 d. (i) Western blot analysis of CK1α levels after treatment of MOLM-13 CK1α WT and MOLM-13 CK1α G40N cells with 1 µM DEG-47 for 24 h. (j) Structure of DEG-35. (k) Western blot of CK1α levels after RAW 264.7 treatment with DEG-35 in the presence or absence of SB-405483 for 3 h. (l) Quantification of n = 3 biologically independent samples of k. Comparisons were performed using a one-way ANOVA with Šídák’s multiple comparisons test. P-values are shown. Data are represented as mean ± SD. Western blot, cell viability, and flow cytometry data are representative of n = 3 biologically independent samples. For uncropped Western blot images, see Supplementary Fig. 3.
Extended Data Fig. 4 Supplemental global proteomics volcano plots and substrate validation with DEG-47 in the presence and absence of 10 µM SB-405483.
(a–c) Volcano plots of MOLM-13 cells after 18 h incubation with (a) 10 nM DEG-47 and 10 µM SB-405483 (b) 10 µM SB-405483 and (c) 10 nM DEG-47. (d) Western blot analysis of AHRR, CESI, and TXNIP after treatment of MOLM-13 cells with SB-405483 for 18 h. (e–f) Volcano plots of MOLM-13 cells after 18 h incubation with 10 nM DEG-47 and 10 µM SB-405483 versus MOLM-13 cells treated with (e) 10 µM SB-405483 (f) 10 nM DEG-47 for 18 h. (g) Western blot analysis of Wee1 levels after treatment of MOLM-13 cells with 10 nM DEG-47 in the presence or absence of SB-405483 for 18 h. (h) Western blot analysis of WIZ levels with increasing concentration of DEG-47 with and without 10 µM SB-405483 for 18 h. All proteomics experiments were performed with n = 4 biologically independent samples. P-values by t-test (background) method. For uncropped Western blot images, see Supplementary Fig. 4.
Extended Data Fig. 5 Wee1 is a CRBN neosubstrate targeted by DEG-47.
(a) Western blot of Wee1, PLK1, and pCDK1 after treatment of MOLM-13 cells with increasing concentration of DEG-47 in the presence or absence of SB-405483 for 24 h. (b) Quantification of n = 3 biologically independent Western blots measuring Wee1 levels shown in a. (c) Western blot of Wee1 levels after treatment with DEG-47 over various time points. (d) Quantification of n = 3 biologically independent Western blots measuring pCDK1 levels shown in a. (e) Quantification of n = 3 biologically independent Western blots measuring PLK1 levels shown in a. (f) Western blot analysis of MOLM-13 cells pretreated with DMSO, MG132, or MLN4924 for 1 h, and then treated with DEG-47 in the presence or absence of SB-405483 for 24 h. (g) Western blot analysis of MOLM-13 CRBN KO or MOLM-13 WT cells treated with DEG-47 in the presence or absence of SB-405483 for 24 h. (h) Western blot analysis of MOLM-13 cells pretreated with pomalidomide or DMSO for 1 h prior to treatment with DEG-47 in the presence or absence of SB-405483 for 24 h. (i) Wee1-HiBiT luminescence signal in cells transiently expressing Wee1-HiBiT or Wee1-HiBiT G322N in HEK293T cells dosed with DEG-47 in the presence or absence of 10 µM SB-405483 for 24 h. (j) Protein expression levels of Wee1 after treatment with indicated compounds in MOLM-13 cells for 24 h. Data are represented as mean ± SD. Comparisons were performed using a one-way ANOVA with Šídák’s multiple comparisons test with * = p < 0.05, ** = p < 0.01, *** = p < 0.001, **** = p < 0.0001. All Western blot, HiBiT, and protein expression data are representative of n = 3 biologically independent samples. For uncropped Western blot images, see Supplementary Fig. 5.
Extended Data Fig. 6 SB-405483 differentially modulates degradation of CRBN neosubstrates in the presence of monovalent degraders.
(a) Structures and reporter cell line targets of select orthosteric ligands tested in reporter cell line screen. (b–d) Neosubstrate degradation with indicated orthosteric ligands in the presence and absence of 10 µM SB-405483 treated for 24 h: (b) CK1α-HiBiT (c) IKZF1/3(ZF2)-GFP (d) IKZF2(ZF2)-GFP (e) Levels of SALL4(ZF1,2)-GFP in HEK293T reporter cell line treated with SB-405483 and orthosteric ligands relative to orthosteric ligand alone. (f) Degradation of indicated neosubstrate after treatment with lenalidomide in the presence or absence of 10 µM SB-405483 for 24 h. (g) Degradation of indicated neosubstrate after treatment with avadomide in the presence or absence of 10 µM SB-405483 for 24 h. (h) Degradation of indicated neosubstrate after treatment with mezigdomide in the presence or absence of 10 µM SB-405483 24 h. Data are mean ± s.e and error propagation were performed using the delta method. All flow cytometry data is representative of n = 3 biologically independent samples.
Extended Data Fig. 7 SB-405483 universally potentiates the degradation of PROTAC targets.
(a) Structures and targets of select PROTACs. (b–c) Levels of the indicated protein with SB-405483 with orthosteric ligand relative to orthosteric ligand alone: (b) HEK293T BRD4(BD1)-GFP (c) BRD4(BD2)-GFP. (d) Western blot of CRBN levels after dosing HEK293T cells with OUN20985 with and without SB-405483 for 24 h. (e–f) Western blot of FKBP12 levels after dosing HEK293T cells with (e) dFKBP12-AcN (f) dFKBP12-AcQ with and without SB-405483 for 24 h. (g) Structure of WH-10417-099. (h–j) Volcano plots of HEK293T cells after 24 h incubation with (h) 100 nM WH-10417-099 and 10 µM SB-405483 versus DMSO. (i) 100 nM WH-10417-099 and 10 µM SB-405483 versus 100 nM WH-10417-099. (j) 100 nM WH-10417-099 and 10 µM SB-405483 versus 10 µM SB-405483. (k) Venn diagram analysis of significantly downregulated proteins across comparisons. (l) List of proteins significantly downregulated in global proteomics comparisons. Known targets of WH-10417-099 are in red. Additional kinases are in blue. Previously associated neosubstrates in green. Proteomics were performed with n = 4 biologically independent samples. P-values by t-test (background) method. Data are mean ± s.e and error propagation were performed using the delta method. Flow cytometry data and Western blots are representative of n = 3 biologically independent samples. For uncropped Western blot images, see Supplementary Fig. 6.
Extended Data Fig. 8 Cryo-EM reveals how SB-405483 modulates CRBN conformation and neosubstrate recruitment.
(a) The distribution of conformational populations observed for CRBN in the absence or presence of substrates upon introducing SB-405483 and different orthosteric ligands. (b) EM density map of the DDB1ΔBPB/CRBNclosed complex in the presence of lenalidomide, SB-405483, and CK1α. CK1α density is shown as low-pass filtered image and zoomed-in view below shows both compound densities. (c) EM density map of the DDB1ΔBPB/CRBNclosed complex in the presence of lenalidomide, SB-405483, and IKZF1(ZF2,3). Notably, IKZF1 density is not seen; the zoomed-in view below shows both ligand densities. (d) EM density map of DDB1ΔBPB/CRBNclosed complex in the presence of DEG-47 and IKZF1(ZF2,3) fit with atomic model. The zoomed-in view below shows density to model fit for DEG-47 and IKZF1(ZF2,3). (e). EM density map of DDB1ΔBPB/CRBNclosed complex in the presence of DEG-47, SB-405483 and IKZF1(ZF2,3). Notably, IKZF1 density is not seen at high threshold but present at very low threshold; the zoomed-in view below shows DEG-47 drug density and zoomed-in view on right side confirms density for SB-405483. (f) Mechanistic model of allosteric and orthosteric modulation of CRBN conformation that drives differential neosubstrate recruitment and protein degradation.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–12, materials and methods, synthetic procedures, additional references, NMR spectra and IR spectra.
Supplementary Table 1 (download XLS )
Sequence similarity matches conducted using PSI-Blast at NCBI (http://blast.ncbi.nlm.nih.gov/) on the non-redundant protein sequence database with an E-value threshold of 0.005. Sequences reached convergence after three iterations. Sequence alignment was preformed using Geneious Alignment (shown in Fig. 2h and Extended Data Fig. 1n,m).
Supplementary Table 2 (download XLS )
Proteins identified from MOLM-13 cells treated with DMSO, DEG-47, DEG-47 + SB-405483 and SB-405483 for 18 h and analysed via global proteomics (shown in Fig. 3i,j and Extended Data Fig. 4a–c,e,f). P values for the abundance ratios were calculated by one-way ANOVA with Tukey’s HSD post hoc test.
Supplementary Table 3 (download XLS )
FITC-A mean or luminescence raw values and subsequent log2(fold change AUC) resulting from flow cytometry analysis or luminescence readings in reporter cell lines stably expressing: CK1α-HiBiT, Wee1-HiBiT, GSPT1-GFP, IKZF1/3(ZF2)-GFP, IKZF2(ZF2)-GFP, SALL4(ZF1,2)-GFP, BRD4(BD1)-GFP or BRD4(BD2)-GFP (shown in Figs. 3l–n and 4b-f and Extended Data Figs. 6b–h and 7b,c).
Supplementary Table 4 (download XLS )
Proteins identified from MOLM-13 cells treated with DMSO, WH-10417-099, WH-10417-099 + SB-405483 and SB-405483 for 18 h and analysed via global proteomics (shown in Extended Data Fig. 7h–l). P values for the abundance ratios were calculated by one-way ANOVA with Tukey’s HSD post hoc test.
Supplementary Table 5 (download XLS )
Cryo-EM data collection and image analysis (shown in Fig. 5 and Extended Data Fig. 8b–e).
Supplementary Table 6 (download XLS )
Cryo-EM atomic modelling, refinement, and validation statistics (shown in Fig. 5 and Extended Data Fig. 8b–e).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dippon, V.N., Rizvi, Z., Choudhry, A.E. et al. Identification of an allosteric site on the E3 ligase adapter cereblon. Nature 651, 482–490 (2026). https://doi.org/10.1038/s41586-025-09994-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09994-w