Abstract
The vertebrate central nervous system is protected by the blood–brain barrier and meningeal membranes, which ensure immune privilege1. In the mammalian brain, microglia and barrier-associated or border-associated macrophages (BAMs) provide immune surveillance and scavenge wastes2, yet how evolution shaped immune-cell diversity and function is not understood. In zebrafish, a vascular-derived mural lymphatic endothelial cell (muLEC) lineage fulfils scavenger cell functions at central nervous system borders3,4,5. Here we identify the transcription factor odd-skipped related 2 (osr2) as a specific marker and regulator of muLEC differentiation and maintenance. osr2 controls the transition of muLECs from interconnected endothelial cells to individual scavenger cells in part by means of control of cadherin-6. muLECs are more transcriptionally similar to BAMs than to other mammalian meningeal cells and share several functions in tissue homeostasis. However, BAMs are absent from zebrafish and muLECs from mice and humans. Analysis of osr2, lymphatic endothelial cell (LEC) and BAM markers in diverse vertebrate species reveals muLECs as an ancient lineage and BAMs a recent mammalian specialization. muLECs and BAMs share functional analogies but are not homologous, providing an example of convergent evolution. This highlights the physiological importance of meningeal scavenger cells and the developmental plasticity of LECs in generating specialized cell types throughout evolution.
Similar content being viewed by others
Main
The vertebrate blood–brain barrier and meningeal membranes serve as barriers that together control influx and efflux of cells, nutrients and metabolites at the central nervous system (CNS)1. As such, the CNS is immune privileged and patrolled by highly specialized immune cells2. Key innate immune cells in mammals are the parenchymal microglia and the border- (also known as barrier-) associated macrophages (BAMs)6,7. BAMs develop in the mammalian embryo from yolk sac-derived monocytic progenitors, are replenished throughout life by low level self-repopulation, and in disease settings by influx of circulating monocytes7. BAMs express a unique repertoire of scavenger receptors such as MRC1 (CD206), LYVE1, STAB1 and others that are involved in the phagocytosis of diverse ligands and tissue wastes7. They are localized to perivascular spaces at brain borders, the choroid plexus and the meninges, where they are thought to both clear wastes and regulate immune responses. BAMs have been shown to influence progression of Alzheimer’s and Parkinson’s disease phenotypes and are implicated in stroke and brain cancer8,9,10,11. Most of our understanding of immune-cell types in the CNS comes from studies in mammalian systems. Consequently, we know very little about the evolutionary diversification of CNS immunity at a cellular and functional level, in other parts of the phylogeny where immune privilege is intact.
The teleost fish Danio rerio (zebrafish) has a conserved blood–brain barrier and cell lineages of the neurovascular unit as well as a conserved innate immune system12,13. Whereas zebrafish macrophages, neutrophils and microglia have homologous roles to their mammalian counterparts13, BAMs have not been described. However, a new population of isolated muLECs (also known as brain LECs or fluorescent granular perithelial cells) was previously discovered within the CNS adjacent to the pial meningeal layer in an equivalent location to BAMs in mammals3,4,5. muLECs develop from the choroidal vascular plexus to form loop-like vascular structures on the dorsal surface of the zebrafish brain by 5–7 days postfertilization (dpf). Cells within these loop-like structures then disperse into individual cells as a sparse mural population covering the meningeal blood vascular network in juvenile and adult fish3. This developmental process is dependent on canonical drivers of lymphangiogenesis (vegfc, flt4, ccbe1, prox1a)3,4,5,14, glial cell function15 and independent of haematopoietic programs5. muLECs are a specialized lymphatic cell type with a distinctive mesenchymal morphology and established functions in scavenging macromolecular wastes into endocytic vesicles in a manner similar to scavenger endothelial cells (ECs)16,17. This scavenging function is dependent on Mrc1 (also known as CD206), Stab1, Stab2 and the endocytic adaptor protein Dab2 (refs. 16,18,19,20). So far, a lineage-specific marker and developmental program controlling muLEC differentiation has not been identified.
We examined the developmental transition of muLECs as they populate the zebrafish midbrain surface. At a gross morphological level, muLECs adjacent to the mesencephalic veins progressively disconnect from each other and lose their elongated EC appearance to become separated from one another and more rounded in morphology between 7 dpf and 30 dpf (Fig. 1a,b and Extended Data Fig. 1a). To identify the earliest transcriptional drivers of this change, we collated a single-cell RNA sequencing (scRNA-seq) time course for ECs, integrating datasets spanning key stages of muLEC development in the brain. This included 40 h postfertilization (hpf), 3 dpf, 4 dpf, 5 dpf and 7 dpf, encompassing sprouting of LECs from embryonic veins (40 hpf), emergence of muLECs at midbrain meninges (3–4 dpf) and the formation of muLEC loops (5–7 dpf) (Extended Data Fig. 1b–e). We annotated cell clusters on the basis of known marker genes (Extended Data Fig. 1f and Supplementary Data Table 1) and ECs (n = 15,898) were reclustered (Fig. 1c,d and Extended Data Fig. 1g–j). A putative muLEC cluster was identified as transcriptionally distinct from other EC clusters, including LECs. To investigate transcriptional differences between muLECs and LECs, we performed differentially expressed genes (DEG) analysis at 3 dpf and 7 dpf. Several genes marking LECs were strongly downregulated or absent in the muLEC cluster (for example, cadherin-6 (cdh6), tbx1)14, whereas others were reduced in expression (for example, prox1a). Conversely, genes associated with scavenger activity (for example, lyve1b, mrc1a) were upregulated (Fig. 1e–g and Supplementary Table 2). Notably, the transcription factor odd-skipped-related 2 (osr2), previously described for its role in the development of palate, synovial joint, kidney and bone formation in mammals21,22,23 and fin chondrogenesis in zebrafish24, was the most upregulated gene in the muLEC cluster at 7 dpf (Fig. 1e–g). osr2 expression initiated in ECs between 40 hpf and 3 dpf and was specifically restricted to the muLEC cluster at all timepoints, being the most cell type specific marker across the whole transcriptome (Fig. 1f,g and Extended Data Fig. 2a–c’). cdh6 and tbx1 were LEC-specific markers not expressed in muLECs (Fig. 1g, Supplementary Table 2 and Extended Data Fig. 2b,c”).
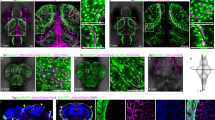
a, Schematic of anatomical areas imaged. a′, EC nuclei (Tg(fli1a:nGFP)y7, blue) and muLECs (Tg(−5.2lyve1b:DsRed)nz101, orange) in 3-dpf, 7-dpf, 14-dpf and 30-dpf zebrafish right midbrains. b, muLEC length at mesencephalic vein scored as longest distance along any cellular axis from a′ (n = 12, n = 106, n = 10, n = 274, n = 123, n = 191 cells at stages indicated). c,d, UMAP showing EC scRNA-seq at different timepoints (n = 15,898 cells). Coloured by phenotype (c) and stage (d). e, Volcano plot of DEGs between muLECs and LECs at 3 dpf and 7 dpf (n = 220 (102 up, 118 down), and n = 211 (75 up, 136 down), respectively). Pink indicates DEGs (adjusted P < 0.05 and absolute log2[FC] > 1) and purple genes of interest. Two-sided Wilcox test, Bonferroni correction. f, UMAPs showing ECs by timepoint (40 hpf, 3d pf and 7 dpf), coloured by phenotype (upper) and osr2, mrc1a, lyve1b and prox1a expression (lower). Colour scale represents log-normalized expression. g, Dot plot showing marker gene expression in muLEC, VEC and LECs. Gene names and stages are indicated. Size of dots represents proportion of expressing cells, colour scale log-normalized expression. h–h′′, osr2 expression (Tg(osr2-gal4, UAS:mTagBFP2);Tg(UAS:RFP) knock-in, magenta) and lymphatics (Tg(−5.2lyve1b:DsRed)nz101, cyan) at 7 dpf (h), and higher magnification of facial (h′) and trunk lymphatics (h′′). i–i′′, osr2 expression (magenta) and LECs and VECs (cyan) in a dorsal view of the 7-dpf brain (i) and magnification of muLEC loop adjacent to the mesencephalic vein (individual channels, i′ and i′′). Co-expression confirmed in more than ten animals. AEC, arterial EC; imVEC immature VEC; MV, mesencephalic vein; sEC, sparc+ EC. Images are confocal projections. Error bars represent s.e.m. Scale bars, 100 μm (a′,h,h′,h′′,i), 25 μm (i′′). Illustration in a created in BioRender. Usseglio gaudi, A. (2025) https://BioRender.com/w7lx2hd.
To validate osr2 as a definitive muLEC lineage marker, we generated a knock-in line. This identified expression in embryonic tissues previously reported to express osr2 (ref. 24) and expression colocalized with lyve1b in muLECs but not LECs at 7 dpf (Fig. 1h,i″ and Extended Data Fig. 2d). To further validate this, we took advantage of a published single-nuclei assay for transposase accessible chromatin sequencing (snATAC-seq) dataset14, and analysed differentially accessible chromatin peaks at the osr2 locus at 4 dpf. In muLECs, we discovered a putative enhancer in a distal region 45 kilobases (kb) (+45 osr2) downstream of the osr2 promoter. This region was cloned into a zebrafish enhancer detection vector and was sufficient to drive muLEC expression in a stable transgenic line (Extended Data Fig. 2e–g’). Taken together, these data confirm osr2 as a EC lineage-specific marker of muLECs.
To understand osr2 function in muLEC development, we generated a predicted loss-of-function mutant using CRISPR genome editing (Extended Data Fig. 3a). osr2 mutants and their wild-type siblings showed indistinguishable growth and morphology at all timepoints (Extended Data Fig. 3b). Mature muLECs are typically isolated, individual cells with limited cell-to-cell contact, unlike ECs in vessels. In mutants, muLEC number was unchanged (Extended Data Fig. 3c–f), but the characteristic transition to individual scavenger cells was abnormal (7 dpf, 14 dpf, 21 dpf and 30 dpf; Fig. 2a,a′ and Extended Data Fig. 3g–i’). muLEC length was increased in osr2 mutants compared with sibling controls at all 14 dpf, 21 dpf and 30 dpf, but unchanged at 7 dpf (Fig. 2b, Extended Data Fig. 3j). In addition, the percentage of muLECs engaged in cell-to-cell contact with other muLECs was reduced in osr2 mutants from 14 dpf up to 30 dpf (Fig. 2c). Together, these data show a role for osr2 in the morphological transition of muLECs from ECs to individual cells. In addition, we raised mutant and sibling animals to 1 year of age and assessed muLECs. Although mutant brains were morphologically normal, we observed a marked loss of muLECs (Fig. 2d,d′), identifying a requirement for osr2 in the maintenance of muLECs.
a,a′, muLECs (Tg(−5.2lyve1b:DsRed)nz101) of sibling (upper) and osr2 mutant (lower) at 30-dpf right midbrain (a) and mesencephalic vein (a′). b,c, Average muLEC length (b) and percentage of non-contacting muLECs (c) at mesencephalic vein in 7 dpf (n = 40, n = 23); 14 dpf (n = 14, n = 8); 21 dpf (n = 11, n = 5); 30 dpf (n = 10, n = 10) siblings (green) and mutants (blue), respectively. b, **P = 0.0023, ****P < 0.0001. c, ****P < 0.0001. d, muLECs in 1-year-old (yo) sibling (upper) and osr2 mutant (lower). d′, Brain width (upper) and total muLECs (lower) for d (n = 5 siblings, n = 6 mutants), ****P < 0.0001. e,e′, UMAP of scRNA-seq at 30 dpf (level 02, RNA_snn_res.0.2, n = 6,942) coloured by phenotype (e) and genotype (e′). f, Stacked bar plot showing genotype composition per cell type. g, UMAP coloured by mean mixing metric score. h,h′, UMAPs of sibling (h) and mutant osr2−/− (h′) ECs, coloured by phenotype (sibling n = 3,473 cells, mutant n = 3,543 cells). Arrows indicate RNA velocity. i,j, Stacked bar plot for DEGs (ECs, adjusted P < 0.05 and absolute log2[FC] > 0.5, i) and volcano plot for muLEC DEGs (n = 82 (55 up, 27 down, j). Pink, DEGs (adjusted P < 0.05 and absolute log2[FC] > 0.5). Purple, genes of interest. Two-sided Wilcox tests with Bonferroni correction. k, cdh6 expression (Tg(−48cdh6:E1b:EGFP)uom134, magenta) in muLECs (green) in siblings (top) and osr2 mutants (bottom). k′, Higher magnification. k′′, cdh6 mean intensity from k (n = 54 siblings, n = 30 mutants), ****P < 0.0001. l, muLECs at mesencephalic vein (Tg(−5.2lyve1b:DsRed)nz101, magenta) and blood vessels (Tg(kdrl:EGFP)s843, green) in sibling (top) and cdh6 mutants (bottom). m,n, Average muLEC length (m) and percentage non-contacting muLECs (n) from l. 5 dpf n = 30, n = 13; 7 dpf n = 32, n = 11; 10 dpf n = 24, n = 13, for siblings and mutants, respectively. ***P < 0.0001. One-way ANOVA in b,c,m,n. Unpaired two-tailed t-test for d′, k′′. All images are confocal projections. Error bars represent s.e.m. Scale bars, 100 μm (a,d,k,l), 25 μm (a′,k′).
To further understand the role of osr2 in muLEC development, we performed scRNA-seq of ECs from siblings and osr2 mutants at 30 dpf (Extended Data Fig. 3k–m). We annotated cell clusters on the basis of known marker genes (Extended Data Fig. 3n and Supplementary Data Table 1) and ECs (n = 6,942) were reclustered (Fig. 2e,f and Extended Data Fig. 3o,p). On clustering, we observed that although mutant and sibling muLECs were both present, the mixing of cells in the muLEC cluster was heterogeneous (Fig. 2g and Extended Data Fig. 3q). Separate clustering of sibling and mutant datasets coupled with velocyto trajectory analysis revealed a notable defect in the differentiation trajectory of osr2 mutant compared with sibling control muLECs. Mutant muLEC transcriptomes were closely associated with LEC transcriptomes with notable mixing of the lineages in velocyto trajectory analysis, whereas wild-type muLECs had transitioned to a distinctly separate cell type (Fig. 2h,h′ and Extended Data Fig. 4a–c). Furthermore, analysis of Slingshot pseudotime differentiation trajectories confirmed a reduction in differentiation of mutant muLECs (Extended Data Fig. 4e,f). A DEG analysis comparing mutant and sibling muLECs identified 82 DEGs (27 downregulated, 55 upregulated) (Fig. 2i and Supplementary Data Table 3). Changes in gene expression included the dysregulation of angpt2a and its receptor tie1 (ref. 25) (involved in EC adhesion and lymphatic development) and the increased expression of some cell–cell adhesion molecules in muLECs (Fig. 2j and Extended Data Fig. 4d). This included expression of both cldn5b and cdh6, normally high in wild-type LECs, but lost in differentiated wild-type muLECs. Both genes were highly expressed in osr2 mutant muLECs that had failed to fully differentiate (Extended Data Fig. 4g). As this data is consistent with a defect in differentiation and muLECs are active in scavenging meningeal wastes, we assessed muLEC scavenging in osr2 mutants. We found that mutants show reduced scavenging capacity as early as 7 dpf, which is consistent with abnormal differentiation (Extended Data Fig. 4h–j). Overall, this suggests that osr2 mutant muLECs still form, but do not differentiate as well as their wild-type counterparts, leading to morphological and functional deficits in the lineage and a failure in maintenance over time.
cdh6 encodes a cadherin predicted to control cell–cell adhesion and expressed specifically in LECs in zebrafish, whereas cdh5 is only found in blood vessels in this model system14. Notably, the developmental transition to individual muLECs is concomitant with the progressive downregulation of cdh6 (Extended Data Fig. 2b,c”). Using a cdh6 enhancer reporter transgenic line, we confirmed that cdh6 expression labels all LECs but is downregulated during muLEC development (Extended Data Fig. 4k–l’ and Extended Data Fig. 2b,c”). Crossing this line onto the osr2 mutant, we confirmed an increase in cdh6 expression in muLECs lacking osr2 (Fig. 2k–k′′). To determine whether loss of cdh6 function might contribute to the morphological transition from EC to muLEC, we generated a CRISPR mutant for cdh6 (Methods). We found that mutants lacking cdh6 showed muLECs with more rounded and isolated morphologies at 5 dpf, 7 dpf and 10 dpf, whereas there was no gross change to lymphatic vessel morphology in these animals (Fig. 2l–n and Extended Data Fig. 4m). This is consistent with an earlier morphological transition from EC towards individual scavenger cells occurring in cdh6 mutants than in wild-type animals. Thus, downregulation of cdh6 and cell–cell adhesion is probably an important aspect of the osr2-regulated morphological changes observed during development of the lineage.
The mammalian CNS is devoid of LECs, with the presence of lymphatic vessels only seen at the dura in close proximity to the skull26. Although rare cells expressing some LEC markers have been observed in human histological analyses27, a large population resembling zebrafish muLECs has never been reported. To hunt for a mammalian muLEC population in an unbiased manner, we isolated cells from mouse meninges for scRNA-seq on the basis of their ability to scavenge Alexa Fluor 488-labelled acetylated-LDL (488acLDL). Previous work has shown that 488acLDL is taken up by zebrafish muLECs3 and this moiety would be expected to be taken up broadly by cells with scavenging ability. We finely dissected the leptomeninges and some associated parenchyma from the mouse cortex, dissociated cells, cultured for 1 h in 488acLDL and fluorescence-activated cell-sorted all cells that had taken up the fluorophore. These cells were then sequenced to profile meningeal cells with scavenging capability (Fig. 3a and Extended Data Fig. 5a,b). We assigned clusters based on expression of known marker genes (Extended Data Fig. 5c and Supplementary Data Table 1). Large populations of ECs and immune cells were identified as well as smaller populations of glia and oligodendrocytes (Fig. 3a). We captured parenchymal microglia and two distinct macrophage populations, perivascular macrophages and BAMs on the basis of published profiles28. Despite isolating n = 16,262 cells capable of scavenging 488acLDL, we did not identify any cells expressing LEC markers or Osr2 that would be homologous to zebrafish muLECs. We further interrogated the publicly available Allen Brain Cancer Cell Atlas dataset29, which captured more than 4 million cells, including approximately 230,000 immune and vascular cells, and found no evidence of muLECs in the mouse brain (Extended Data Fig. 5d). To explore scavenging cell populations in human leptomeninges, we took advantage of two previously published human scRNA-seq datasets that included leptomeningeal cells30,31 (Fig. 3b and Extended Data Fig. 5e–i). We assigned clusters on the basis of expression of known marker genes and we investigated the expression of OSR2, LEC and muLEC marker genes across cell clusters (Extended Data Fig. 5j–l). We found that human meninges were also devoid of muLECs and, thus, muLECs appear to be absent from mammals.
a, UMAP of scRNA-seq of dissected mouse cortex scavenging cells (n = 16,262 cells) coloured by phenotype (right). b, UMAP of selection of cells from GSE2453111 (ref. 30) (23 weeks postconception human leptomeningeal cells). Level 02 (n = 10,986 cells) coloured by phenotype. c–e, Heatmap of spearman correlation coefficients (left) and Jaccard similarity (right) between indicated species: mouse versus zebrafish (c); human versus zebrafish (d) and human versus mouse (e). Sorted by Jaccard similarity (high to low). Colour represents positive correlation coefficient and Jaccard similarity, respectively. Those that did not pass significance testing are denoted with an x. f, Dot plot showing expression for lymphatic, scavenging, proteolytic and haematopoietic marker genes for zebrafish, mouse and human (data from refs. 30,31). Size of dots represents proportion of expressing cells, colour represents average log-normalized expression. g, Meningeal blood vessels (Tg(kdrl:EGFP)s843, green) in control (upper) and ccbe1 MO (lower) injected animals. muLECs (Tg(−5.2lyve1b:DsRed)nz101, magenta) shown in the inset. h, Variance of vascular density in control and ccbe1 MO. Two-sided Levene’s test ***P = 0.0005 (n = 7 control, n = 13 ccbe1 MO). i, Neutrophils (Tg(lyz:BFP)zf217Tg, green) in control (upper), flt4um203 mutant (middle) and ccbe1 MO (lower) 4-mpf brains. muLECs inset. j, Neutrophil numbers in control, flt4 mutant and ccbe1 MO (n = 22 control, n = 19 flt4 mutant, n = 9 ccbe1 MO). k, Microglia and/or macrophages (Tg(mpeg1.1:TagBFP)bcz53Tg, green) in control (upper), flt4um203 mutant (middle) and ccbe1 MO (lower) 4-mpf brains. muLECs in inset. l, Microglia and/or macrophage numbers in control, flt4 mutant and ccbe1 MO (n = 8 control, n = 6 flt4 mutant and n = 10 ccbe1 MO brains). Unpaired two-tailed t-test **P = 0.0033 for control versus flt4 mutants, P = 0.0017 for control versus ccbe1 MO in j. *P = 0.0251 for control versus flt4 mutants, **P = 0.0092 for control versus ccbe1 MO in l. NK cells, natural killer cells; MG, microglia; Prolif. BAM, proliferative BAM; PVM, perivascular macrophage; VSMC, vascular smooth EC; wpc, weeks postconception. Scale bars, 100 μm. All images confocal projections. Error bars represent s.e.m. Illustrations in a and b created in BioRender. Usseglio gaudi, A. (2025) https://BioRender.com/w7lx2hd.
To more deeply compare scavenging cells across species and to identify transcriptionally similar cell types in an unbiased manner, we compared our mouse and human datasets with the muLEC transcriptome from zebrafish. This approach used two independent methods (a correlation coefficient and Jaccard similarity score) correlating gene expression as well as the most specific markers of cell types across species. muLECs showed the highest correlation coefficient and Jaccard similarity score with the mouse and human BAM clusters (Fig. 3c–e, Extended Data Fig. 6a–e’ and Supplementary Data Table 4). Detailed analysis of shared gene expression between zebrafish muLECs and mouse BAMs revealed expression of scavenger receptors (for example, homologues of STAB1, MRC1, LYVE1) and factors involved in protein turnover (homologues of CTSD, CTSH, CTSZ, CTSLA, LGMN and LAMTOR4) (Fig. 3f). muLEC specific genes were lymphatic marker genes (for example, homologues of FLT4, PROX1) and BAM-specific genes were associated with the complement system (C1QA, C1QC), known BAM-specific markers (homologues of MS4A7) and well-known haematopoietic marker genes32 (Fig. 3f, Extended Data Fig. 6f,g and Supplementary Data Table 4). Expression of immunomodulatory genes by BAMs but not muLECs suggests specialized roles for BAMs. The development of muLECs at the zebrafish meninges has been demonstrated to be controlled by glial Vegfc production and meningeal fibroblast-produced Ccbe1 (ref. 15). We saw expression of these lymphangiogenic factors in the zebrafish but not mammalian meninges, potentially explaining the species specificity of the lineage (Extended Data Fig. 6h–k’). These data together indicate that muLECs and BAMs are distinct cell types that share transcriptional similarities attributable in large part to genes involved in scavenging of extracellular wastes.
In zebrafish, muLECs can clear tissue wastes and change morphology on injury33 reminiscent of macrophages, but muLECs are generated by lymphangiogenesis and do not develop through haematopoiesis5. To better understand the nature of muLECs and their activation on injury, we generated a scRNA-seq dataset that compared injured versus uninjured larval heads and brains at 24 h postinjury in 7 dpf larvae (Methods and Extended Data Fig. 7a–c). The injury assay led to local accumulation of macrophages, changes in muLEC morphology as previously reported34 and broadly increased lysosomal activity at the wound (Extended Data Fig. 7d,e). The dataset captured a large number of cell types and we correlated gene expression between muLECs and all other cell types (Extended Data Fig. 7f). Apart from high similarity to LECs (consistent with previous work3,5), the second highest correlation score was with a cluster of macrophages or microglia (Extended Data Fig. 7g). Analysis of individual marker genes revealed that muLECs and macrophages and/or microglia both expressed a selection of cathepsins (ctsd, ctsh, ctsz) involved in phagocytic functions35,36,37 (Extended Data Fig. 7h,i and Supplementary Data Table 5), but only muLECs expressed the scavenger receptors typical of mammalian BAMs (stab2, lyve1b, mrc1a)7. Zebrafish macrophages and/or microglia but not muLECs showed expression of myeloid lineage markers (mpeg1.1, csf1ra, spi1a, mfap4) (Extended Data Fig. 7j) and we found no evidence of BAMs present in the zebrafish brain in our own or published datasets38 (Extended Data Fig. 7k,l). Next, we examined the differences seen on wounding. We noticed that the muLEC cluster showed heterogeneous mixing of control and wounded cells and quantification of mixing metric suggested muLECs undergo a substantial transcriptional change on wounding (Extended Data Fig. 8a–d). DEG analysis revealed that muLECs had the highest number of DEGs in response to injury of any cell type (Extended Data Fig. 8e). Gene ontology (GO) terms associated with upregulated genes in muLECs following injury related to endocytosis, proteolysis and metabolic regulation (Extended Data Fig. 8f,g). Upregulated were scavenging cell markers (stab1, stab2, ctsla, ctsh, lgmn, lamtor4) and genes related to proteolysis, peptidase and lysosomal activity (Extended Data Fig. 8h). Taken altogether, these data indicated that muLECs should be considered ‘macrophage-like’ on the basis of anatomical, transcriptional and functional analogy. As a functional test, we delivered clodronate liposomes to 6 dpf zebrafish embryos by angiogram. Then 24 h postinjection, both macrophages and muLECs were significantly depleted (Extended Data Fig. 8i,j). In addition, as muLECs change morphology on wounding, we tested whether this change is dependent on normal differentiation of the lineage. We examined osr2 mutants following wounding and found that mutant muLECs did not undergo the expected morphological transition (Extended Data Fig. 8k–l). Altogether, muLECs differentiate into macrophage-like cells that are activated on injury and are transcriptionally reminiscent of mammalian BAMs, but the zebrafish brain lacks a true BAM population.
As well as scavenging wastes, other physiological functions attributed to BAMs include signalling to endothelium and modulating local immune responses (for a review, see refs. 7,28,39). muLECs can produce Vegfs and modulate meningeal angiogenesis early in development3, but their influence over mature vasculature and immune-cell populations has not been studied, whereas some immune-cell types (such as microglia) share related functions18,40. We found that injection of optimized doses of a morpholino oligomer (MO) depleting Ccbe1 (ref. 41) generated mature animals that lack muLECs but retain lymphatics, a scenario in which a temporary developmental block effectively deletes the lineage in older otherwise wild-type animals (Fig. 3g). Furthermore, we examined flt4hu4602 mutants that are hypomorphic, lacking muLECs either completely, or partially in an asymmetric manner (Extended Data Fig. 8m). In both models, we examined the number of mpeg1-expressing macrophages and/or microglia and lyz-expressing neutrophils, as well as the morphology of meningeal blood vasculature in adults. The brain showed no major morphological change but marked patterning defects were observed in meningeal blood vessels with increased regional variance in network density (Fig. 3g,h and Extended Data Fig. 8m–o). Furthermore, neutrophil and macrophage numbers were increased in both models at 4 months of age (Fig. 3i–l and Extended Data Fig. 8p,q), but unchanged at 7 dpf (Extended Data Fig. 8r,s). Direct interactions between muLECs and adaptive immune cells have been recently observed42. Although changes in innate immune cells and vasculature in flt4hu4602 mutants could be due to the absence of Flt4 in vasculature, the defects in ccbe1 MO-injected animals at 4 months of age (when the MO would be absent) rather suggests a specific function for muLECs interacting with cells in their microenvironment. Overall, muLECs are absent from the mammalian brain and BAMs from zebrafish brains, the two lineages develop from distinct origins and are controlled by independent mechanisms. Nevertheless, the two cell types localize to equivalent anatomical locations, show transcriptomic similarities and share functions in clearing leptomeningeal wastes, influencing the local immune landscape and the vasculature. muLECs and BAMs are therefore analogous but not homologous cell types.
To better understand the evolutionary relationship between these cell types, we investigated their presence in leptomeninges in each of the taxonomic classes; Elasmobranchii, Actinopterygii, Amphibia, Aves and Mammalia. We explored the ancestral state of meningeal scavenger cells by examining their marker gene expression in the meninges of the epaulette shark, the axolotl, the chicken, the fat-tailed dunnart (a marsupial), mice and zebrafish. We used hybridization chain reaction in situ hybridization (HCR ISH) and analysed OSR2 as a definitive lineage marker, as well as known markers of BAMs (MRC1 and CSF1R) and in select species LECs (FLT4 and PROX1). We performed conservation analysis to identify different orthologues in these species and used a polyA positive control probe to confirm intact RNA in our tissue sections (Methods). Staining of muLEC markers (OSR2, MRC1) revealed that consistent with zebrafish meningeal cellular composition, the epaulette shark, the axolotl and the chicken showed the presence of OSR2, MRC1 positive muLECs (Fig. 4a–d, white arrows). We observed co-expression of muLEC and LEC markers Osr2 and Flt4 in shark and LEC markers Flt4 and Prox1 in chicken, further confirming their identity as muLECs (Fig. 4g,h, see also Extended Data Fig. 9a–f). We did not detect Osr2-expressing cells at the meningeal borders in either mammalian species (Fig. 4e,f, white asterisks). Notably, our Osr2 probe stained the forelimb of E12.5 mouse embryos as previously described in ref. 43 and we detected staining in the dunnart intestine (Extended Data Fig. 9g,h). Staining with macrophage marker (CSF1R) revealed CSF1R-positive brain parenchymal cell populations in all species (Fig. 4a′–f′, yellow arrows). This staining was not present in cells concentrated at brain borders in the epaulette shark, zebrafish, axolotl or the chicken (Fig. 4a′–d′). However, Mcr1 Csf1r double positive cells were found at the meningeal borders of the mouse brain and were present (albeit less abundant) at dunnart meninges, consistent with BAMs that derived through haematopoiesis in distant mammalian species (Fig. 4e′,f′, white and yellow arrows). Taken together, in cartilaginous fishes, ray-finned fishes, amphibians and birds, OSR2-expressing muLECs represent an ancient meningeal scavenging cell lineage. A distinct lineage is present in mammals (BAMs) and appears to have replaced muLECs (Fig. 4i and Extended Data Fig. 9i).
a–f, HCR ISH staining of epaulette shark (Hemiscyllium ocellatum) (a), zebrafish (Danio rerio) (b), axolotl (Ambystoma mexicanum) (c), chicken (Gallus gallus) (d), fat-tailed dunnart (Sminthopsis crassicaudata) (e) and mouse (Mus musculus) brain sections (f), for expression of OSR2 (green), MRC1 (magenta) orthologues with cell nuclei (DAPI, blue). White arrowheads point to OSR2 MRC1-double positive cells at meningeal borders, asterisks their absence, white dotted line marks the leptomeningeal border. HCR co-expression in all species, but individual probes in axolotl. n = 3 sections from n = 3 animals were analysed for all species. a′–f′, HCR ISH staining for expression of CSF1R (red) orthologues with cell nuclei (DAPI, blue) in the same species as a–f. Yellow arrows point to CSF1R-positive cells in the brain parenchyma. g, HCR ISH staining of epaulette shark (H. ocellatum) brain sections, showing expression of osr2 (green), flt4 (magenta) with cell nuclei (DAPI, blue). n = 3 sections from n = 3 animals. h, HCR ISH staining of chicken (G. gallus) brain sections, showing expression of prox1 (green), flt4 (magenta) with cell nuclei (DAPI, blue). n = 3 sections from n = 3 animals. White arrowheads point to co-expressing cells. i, Depiction of the phylogeny of animals involved in this study. Classes of animals were ordered based on their appearance during evolution. Tree branches and animals are colour-coded based on the developmental origin of their meningeal scavenger cells as indicated. DAPI, 4,6-diamidino-2-phenylindole; lm, leptomeninges; sk, skull. Scale bars, 100 μm. All images confocal projections. Illustrations in a–f created in BioRender. Usseglio gaudi, A. (2025) https://BioRender.com/w7lx2hd.
Evolution commonly involves the progressive adaptation of homologous structures or cells that arise from common developmental origins. However, functionally analogous cell types or structures can also evolve independently from distinct origins in different species, a rarer process known as convergent evolution. BAMs are essential innate immune cells populating mammalian leptomeningeal spaces and are specialized to scavenge wastes. Whereas mammalian BAMs develop from yolk sac-derived haematopoietic progenitors28, vascular-derived muLECs are found at equivalent anatomical locations in zebrafish. muLECs express many of the same scavenger receptors as BAMs and are established to scavenge wastes similar to scavenger ECs or BAMs4. muLECs are more transcriptionally similar to BAMs than any other cells of the mammalian CNS and share several analogous functions. Across taxa, the most prevalent state in vertebrates is a muLEC patrolled meningeal barrier, as found in zebrafish, epaulette shark, axolotl and chicken. Mammalian meninges (human, mouse and dunnart) are invested with BAMs as a newer specialized immune-cell type. Thus, we propose that muLECs and BAMs have evolved by means of convergent evolution. It is likely that changes in haematopoietic evolution in mammals allowed for a yolk sac-derived BAM to evolve and ultimately replace the lymphatic lineage, perhaps due to beneficial functions in modulating tissue immunity. We note it is possible that species could exist that harbour both cell types, although we have not observed evidence of this in our analyses. That meningeal scavenging cells have evolved twice, highlights the importance of the cleaning of wastes in meningeal spaces across the vertebrate phylogeny.
It should not be surprising that LECs could be a source of such a novel cell type in evolution. First, having canonical lymphatic vessels develop within a vertebrate CNS would be detrimental and selected against, whereas the rostral veins in developing embryos provide a source of LEC progenitors on which evolution could act. Second, lymphatic vasculature already expresses scavenger receptors and has been reported to have a ‘macrophage-like’ gene expression repertoire in mice44. In fact, developing venous ECs (VECs) and LECs have phagocytic scavenging capabilities16 that could be taken advantage of by enacting minimal evolutionary change. Third, lymphatic vasculature is highly plastic, having recently been shown to produce macrophages on downregulation of Prox1 (ref. 44), and to differentiate into specialized blood carrying vessels in teleost fins45. Thus, LECs are perhaps an ideal source with which to diversify new, specialized cell types throughout evolution. We suggest these characteristics have been taken advantage of by Darwinian selective pressures, to generate a new scavenger cell population at non-mammalian vertebrate meninges. This involved evolution of a unique developmental mechanism, with Osr2 an important regulator of the most common vertebrate meningeal scavenger cell lineage.
Methods
Animals
All animal work was conducted in compliance with animal ethics committees at the Peter MacCallum Cancer Centre (PMCC), the University of Melbourne, Monash University, the University of Queensland and Institutional Animal Care and Use Committee at Weill Cornell Medical College.
Zebrafish
The following published transgenic lines were used at the stages indicated in the figures: Tg(kdrl:EGFP)s843 (ref. 46); Tg(fli1a:nEGFP)y7 (ref. 47); Tg(−5.2lyve1b:DsRed)nz101 (ref. 48); Tg(14xUAS:NfsB-mCherry)c264 (ref. 49); Tg(lyve1b:ERK-KTRClover)uom117; Tg(kdrl:Hsa.HRAS-mCherry)s916 (ref. 50); Tg(mpeg1.1:gal4FF)gl25 (ref. 51); TgBAC(lamp2-RFP)pd1044 (ref. 52); Tg(adamts3:GAL4FF)mu400 (ref. 53); Tg(ccbe1:mCitrine)hu6741 (ref. 53); Tg(vegfc:GAL4FF)mu402 (ref. 53); Tg(5xUAS:EGFP)zf82 (ref. 54); Tg(lyz:BFP)zf217Tg (ref. 55) and Tg(mpeg1.1:TagBFP)bcz53Tg (ref. 56). flt4um203 mutants were previously published in ref. 57 and were used at the stages indicated.
Shark, axolotl, chicken, fat-tailed dunnart and mouse
Tissue samples were collected from stage 38 axolotl (Ambystoma mexicanum) larvae and prehatching epaulette sharks (Hemiscyllium ocellatum) at Monash University. Samples from chicken embryos were collected at 10 dpf at the University of Melbourne. Mouse brains from 6-month-old mice were collected at the PMCC. Following euthanasia mice were perfused with PBS transcardially by means of injection through the right ventricle. Brains and abdominal organs were collected from 1-year-old fat-tailed dunnarts (Sminthopsis crassicaudata) during postmortem dissections at the University of Melbourne.
Transgenesis
The osr2 targeted knock-in line Ti(osr2int2-Gal4vp16/4xnrUAS-mTagBFP2) was generated using the CRISPR–Cas9 Insertional Mutagenesis Protocol and tool kit58. CRISPR RNA (crRNA)(tacatcacacttttaccccg GGG, PAM sequence in uppercase) targeting the second intron of the Osr2 gene was designed using the IDT online tool (https://sg.idtdna.com/site/order/designtool/index/CRISPR_CUSTOM). A solution containing the pSA1-T2A-Gal4vp16_synCoTC/4xnrUAS-mTagBFP2 plasmid (17 ng μl−1), precomplexed guide RNAs (gRNAs) (Alt-R) targeting the Osr2 intronic sequence (6 μM) and plasmid (3 μM) and Cas9 protein (Cas9 HiFi v.3, Integrated DNA Technologies) was injected into one-cell stage zebrafish eggs. Injected embryos were screened for mTagBFP2 fluorescence, raised to adulthood and outcrossed with wild-type zebrafish to identify founders. Cassette integration in the genome was confirmed by PCR and Sanger sequencing using the following primers:
mTagBFP2 forward: AGCTGGGACACAAGCTGAAT
osr2 reverse: TCTGAGGAACAGGCGAGAG
T2A reverse: GGTTCTCCTCCACATCTCCA
osr2 forward: AGTGGAGAGAGCTGAACACC
Tg(+45osr2:E1b:GFP)uom135 and Tg(−48cdh6:E1b:GFP)uom134 transgenic lines were generated for this study by cloning a 500-bp PCR fragment of the osr2 enhancer and a 501-bp PCR fragment of the cdh6 enhancer, respectively, into E1b zebrafish enhancer detection vectors59.
The specific genomic region upstream of osr2 and cdh6 was obtained by PCR amplification from genomic DNA (gDNA) using the primers:
+45osr2 forward: 5′–CAGATGGGCCCTCGAGAACACACACACACTTCA AAGTCCC–3′
+45osr2 reverse: 5′–CCGCAAGCTTGCTAGCTGTGTTGTGAAAACATCAGACAGATGTTTAGG–3′
−48cdh6 forward: 5′–CCGCAAGCTTGCTAGCCTGTAAATTCAATTGTAACAATAGTTTAGTCTG-TAAATACCCT–3′
−48cdh6 reverse: 5′–CAGATGGGCCCTCGAGCGGTTTCGGAGCACATAAACAG–3′
Cloning into the E1b vector plasmid was performed by digestion of the plasmid with restriction enzymes XhoI and NheI (NEB), before cloning of the PCR product using In-Fusion (Takara Bio). The E1b zebrafish enhancer detection vector was a kind gift from E. Wong.
The transgenic lines were made by performing microinjections into single-cell staged embryos as previously described in ref. 60. Next 50 ng μl−1 of transposase messenger RNA (mRNA) was injected with fully assembled Tol2 DNA constructs (40 ng μl−1)61. Zebrafish embryos mosaically expressing the Tol2 construct were then selected and raised to adulthood. Stable transgenic lines were established by assessing F1 larvae for germline transmission and raising positive animals to adulthood.
Genome editing and embryo manipulations
CRISPR–Cas9 genome editing for generating knockout zebrafish embryos was performed as previously described in refs. 62,63. gRNA was generated by annealing and elongation using T4 DNA polymerase (M0203S, New England Biolabs). A gene specific oligonucleotide sequence (Integrated DNA Technologies) was annealed with a constant oligonucleotide, elongation was performed by adding dNTP Mix (U1511, Promega) and T4 DNA polymerase.
The single-guide RNA (sgRNA) target site for osr2 was as follows:
osr2: 5′–GCAGATGAACCGGTGGACGG–3′
The sgRNA target site for cdh6 was as follows:
cdh6-g2: 5′–TGTCACTCAAGTGACCGCCC–3′
This template was purified using DNA Clean & Concentrator-5 Kit (D4014, Zymo Research). Template DNA was transcribed into sgRNA using MEGAscript T7 Transcription Kit (AM1334, ThermoFisher) and subsequently purified using RNA Clean & Concentrator-5 (R1013, Zymo Research). The sgRNA was then combined with Cas9 nuclease (10 mg ml−1) to form a ribonucleoprotein complex that guides targeted cleavage of gDNA. A final concentration of 160 ng ml−1 was used for osr2 crRNA. Ribonucleoprotein mixes were then microinjected into single-cell staged embryos as previously described in ref. 60. Uninjected embryos were used as controls.
To allow imaging of older animals in the absence of pigment (Figs. 1 and 2 and Extended Data Figs. 1–3), CRISPR–Cas9 genome editing was used to target slc45a2 gene (ENSDARG00000002593) for generating F0 melanocyte-deficient adult zebrafish as previously described in ref. 64.
Genotyping
For the use of osr2 and cdh6 F0 founders generated by CRISPR–Cas9 and gRNA injections, cutting of gDNA at the targeted sequence and putative biallelic loss was confirmed by PCR and indicated by running amplicons on a high-percentage agarose gel.
The following primers were used for PCR:
osr2 forward: 5′–ATCATTAACGGGGCCGTGC–3′
osr2 reverse: 5′–CCAGATTGGCGAAGTCGAAG–3′
cdh6-g2 forward: 5′–TTTCATTAGCCATGGGCAAGAGC–3′ cdh6-g2-reverse: 5′–GGATGGCCCTCTTTGATACTGT–3′
Targeting of osr2 resulted in a 7-bp deletion in exon 2, this was predicted to lead to a subsequent frameshift and generation of a premature stop codon at amino acid 107 of the osr2 protein sequence (pThr51AlafsTer57). The allele was transmitted to F1 animals and used to establish a stable mutant strain designated allele osr2uom133. The predicted effect of this allele is loss of osr2 three zinc-finger domains (Extended Data Fig. 3a). All experiments in osr2uom133 mutant fish were conducted in F3 animals or subsequent generations. Fish were imaged and analysed blindly with genotyping retrospectively.
Targeting of cdh6 resulted in a 5 bp deletion in exon 3, this predicted to lead to a subsequent frameshift and generation of a premature stop codon at amino acid 181 of the cdh6 protein sequence (pAla178GlyfsTer4). The allele was transmitted to F1 animals and used to establish a stable mutant strain designated allele cdh6uom132. The predicted effect of this allele is loss of four of the five cdh6 cadherin domains. All experiments in cdh6uom132 mutant fish were conducted in F2 animals. Fish were imaged and analysed blindly with genotyping retrospectively.
Clodronate injections
Zebrafish embryos at 24 hpf were incubated in system water supplemented with N-phenylthiourea (PTU) at a final concentration of 0.003% to inhibit pigmentation65. Clodronate injections were performed as previously described in ref. 66. Clodronate and control liposomes (CP-005-005, Liposoma) were used at a concentration of 5 mg ml−1 and supplemented with dextran, Cascade Blue, 10,000 molecular weight (D1976, Invitrogen) to a working concentration of 5 mg ml−1. A microinjection glass needle and a pneumatic pump were used to perform microinjections. Larval fish at 5 dpf were immobilized with tricaine (0.08 mg ml−1) and mounted laterally to ensure accessibility to the posterior cardinal vein for microinjection. The pressure of the pneumatic pump was adjusted to ensure consistent 2-nl volume of injection for each larval fish. Each injection was validated by seeing immediate distribution of the injected fluorescent dextran mix using a fluorescence microscope. All larval fish that show correct distribution of the fluorescent marker in the body vasculature were recovered in fresh system water supplemented with 0.003% PTU and incubated at 28.5 °C for 48 h. Fish were imaged 48 h postinjection and analysed for scavenging cell depletion.
Dye injections
Zebrafish osr2−/− mutants and siblings embryos at 24 hpf were incubated in system water supplemented with PTU at a final concentration of 0.003% to inhibit pigmentation. Dextran, Cascade Blue, 10,000 molecular weight (D1976, Invitrogen) was diluted to a working concentration of 5 mg ml−1. A microinjection glass needle and a pneumatic pump were used to perform microinjections. Larval fish at 7 dpf were immobilized with tricaine (0.08 mg ml−1) and mounted laterally to ensure accessibility to the posterior cardinal vein for microinjection. The pressure of the pneumatic pump was adjusted to ensure consistent 2-nl volume of injection for each larval fish. Each injection was validated by seeing immediate distribution of the injected fluorescent dextran mix using a fluorescence microscope. All larval fish that showed correct distribution of the fluorescent marker in the body vasculature were recovered in fresh system water supplemented with 0.003% PTU and incubated at 28.5 °C for 2 h. Fish were imaged and analysed blindly with genotyping retrospectively.
Morpholino injections
The ccbe1 start codon targeting morpholinos (MO) and control MO (Genetools) were injected at a concentration of 2.5 ng per embryo, as previously described in ref. 41.
The sequences of the ccbe1 start codon targeting MO and control MO are as follows:
ccbe1 ATG MO: 5′–CGGGTAGATCATTTCAGACACTCTG–3′
Control MO: 5′–CCTCTTACCTCAGTTACAATTTATA–3′
Larval brain wounding
Siblings and osr2−/− mutant Tg(fli1a:nEGFP)y7; Tg(−5.2lyve1b:DsRed)nz101 transgenic fish; Tg(−5.2lyve1b:DsRed)nz101; Tg(mpeg1.1:gal4FF)gl25; (5xUAS:EGFP)zf82 transgenic fish and Tg(lyve1b:ERK-KTRClover)uom117 and Tg(lamp2:RFP)pd1044 transgenic fish were raised at 28.5 °C to 6 dpf in system water supplemented with PTU at a final concentration of 0.003% to inhibit pigmentation. Fish were then immobilized with tricaine (0.08 mg ml−1), mounted ventrally in 0.5% low gelling agarose (BioRad) as previously described in ref. 67, and injured with a single 27 G needle insertion in the left hemisphere of the larval fish midbrain. Fish were then gently recovered in embryo water supplemented with 0.003% PTU and incubated at 28.5 °C for 24 h. Fish were imaged and analysed blindly with genotyping retrospectively.
HCR-RNA-FISH
Tissue preparation and fixation
Whole axolotl heads and spinal cords were fixed in 4% PFA (Sigma) in PBS for 2 days at 4 °C. Epaulette shark brains were prefixed for 10 min at room temperature by injecting 4% PFA in PBS into the chondrocranium before dissection. The shark brains were subsequently dissected free and fixed in 4% PFA in PBS for 3 days at 4 °C. Zebrafish brains were dissected free and fixed in 4% PFA in PBS overnight at 4 °C. Whole chicken heads were fixed in 4% PFA in PBS overnight at 4 °C, before removal of the beak and eyes. Fat-tailed dunnart and mouse organs were dissected free and fixed in 4% PFA in PBS overnight at 4 °C.
Cryogenic protection and embedding
Following fixation, axolotl and epaulette shark samples were washed 3 times with PBS for 15 min at room temperature. Samples were then infiltrated with a gelatin–sucrose embedding solution (8% fish gelatin (Sigma), 20% sucrose (Sigma), dissolved in PBS) for a minimum of 1 hour at room temperature. Samples were submerged in fresh embedding solution in easy-peel moulds (ProSciTech, catalogue no. RR88-T) and oriented for transverse section, then quickly frozen on dry ice. Following fixation, zebrafish, chicken, fat-tailed dunnart and mouse samples were washed 3 times with PBS for 15 min at room temperature. Samples were then submerged in a 30% sucrose (Sigma) solution in PBS and incubated overnight at 4 °C, until complete sinking of the tissues. Samples were then submerged in fresh optimal cutting temperature embedding solution in easy-peel moulds (ProSciTech) and oriented for transverse section, then quickly frozen in dry-ice cold isopropanol. Embedded samples were stored at −80 °C until sectioning.
Cryogenic sectioning
Axolotl and shark samples were sectioned to 20-μm thicknesses in a precooled Leica CM3050 S cryostat (chamber −30 °C and objective −28 °C); zebrafish, chicken, fat-tailed dunnart and mouse samples were sectioned to 4-μm thicknesses in a precooled ThermoScientific CryoStar NX50 cryostat (chamber −40 °C and objective −30 °C). All sections were collected on Superfrost Plus glass slides (Menzel Glaser) and allowed to completely dry at room temperature before being stored at −80 °C until further processing.
HCR-RNA-FISH staining
All stainings were performed following the HCR-RNA-fluorescence ISH (FISH) protocol for fixed frozen tissue sections (Molecular Instruments). Probe set designing and production was completed by Molecular Instruments, based on either gene National Center for Biotechnology Information accession numbers:
Mouse: Csf1r (NM_001037859.2); Lyve1 (NM_053247.4); Osr2 (NM_001368665.1); Flt4 (NM_008029.3); Prox1 (NM_008937.4)
Chicken: csf1r (NM_001321517.2); lyve1 (NM_001199587.2); osr2 (NM_001398381.1); flt4 (NM_001293093.2); prox1 (NM_001005616.1)
Zebrafish: csf1ra (NM_131672.1); lyve1b (NM_001327897.1); osr2 (NM_001017694.1), flt4 (NM_130945.2); prox1a (NM_131405.2)
or by blasting of zebrafish reference genes against non-model organism whole genome:
Axolotl: ambMex 6.0-DD
Epaulette shark: sHemOce1_pat_c1
Fat-tailed dunnart: in-house sequence data (unpublished)
Imaging acquisition and analysis
Image acquisition
Imaging was conducted at the Centre for Advanced Histology and Microscopy (PMCC) and Weill Cornell Medical College. Live embryos and adult fish at indicated stages were immobilized with tricaine (0.08 mg ml−1) and mounted either laterally or dorsally in 1% low gelling agarose (BioRad) as previously described in ref. 41. Embryos, larval and adult fish were imaged on either a Nikon Yokogawa CSU-W1 spinning disc confocal microscope, Zeiss LSM 900 confocal microscope or an Olympus Multiphoton FVMPE-RS microscope at ×4, ×10, ×20, ×25 or ×40 objectives. HCR-RNA-FISH sections were imaged on a Nikon Yokogawa CSU-W1 spinning disc confocal microscope at ×4, ×10 and ×20 objectives.
Quantification
Generally, quantification of muLECs was performed by identifying ECs using Tg(fli1a:nEGFP)y7 or Tg(kdrl:EGFP)s843 transgenic reporters and lymphatic cells using Tg(−5.2lyve1b:DsRed)nz101 expression. For Fig. 2k, Tg(lyve1b:ERK-KTRCloveruom117 was used. muLECs were defined as ECs found mural to blood vasculature and the mesencephalic vein specifically for quantification in Figs. 1 and 2 and Extended Data Figs. 3 and 4. The length of lyve1b-positive muLECs covering the mesencephalic vein in the wild-type midbrain of the zebrafish at different stages (3 dpf, 7 dpf, 14 dpf, 21 dpf, 28 dpf and 30 dpf) was quantified for each individual cell across z-stacks, using the point and segment tools from Imaris (v.10.0.0, Oxford Instruments).
Changes in muLEC phenotypes in response to osr2 and cdh6 mutants were analysed blindly with retrospective genotyping in osr2−/− mutants and control siblings at 7 dpf, 14 dpf, 21 dpf and 30 dpf and in cdh6−/− mutants at 5 dpf, 7 dpf and 10 dpf. The overall length of the fish was measured using a stereo microscope; the number of total muLECs covering the mesencephalic vein was counted manually examining each z-stack individually using the point tool from Imaris (v.10.0.0, Optical Instruments). The length of muLECs covering the mesencephalic vein was measured across different z-stacks using the segment tool from Imaris (v.10.0.0, Optical Instruments), whereas the number of contacting muLECs was defined as cells physically contacting along the mesencephalic vein across the different z-stacks.
The difference in dextran, Cascade Blue, 10,000 molecular weight (D1976, Invitrogen) uptake in response to osr2 knockout (Extended Data Fig. 4j) was analysed blindly with retrospective genotyping in osr2−/− mutants and control siblings at 7 dpf. Briefly, a mask of the lyve1b-positive cells was generated over a consistent region of the mesencephalic vein, using the surface tool from Imaris (v.10.0.0, Optical Instruments). The average intensity of the Cascade Blue fluorophore within the masked area was then recorded for each animal.
Quantification of scavenging cell depletion on clodronate liposomes injection (Extended Data Fig. 8j) was performed by individually counting the number of mpeg1-positive macrophages and lyve1b-positive muLECs, using the point tool from Imaris (v.10.0.0, Optical Instruments).
The difference in total number of microglia and/or macrophages and neutrophils in flt4 mutants and ccbe1 MO-injected animals (Fig. 3i–l and Extended Data Fig. 8s) was performed by individually counting the number of mpeg1-positive and lyz-positive cells, respectively, covering the midbrain of either flt4−/− mutants and ccbe1 MO-injected animals at 7 dpf and in adults (4 months postfertilization (mpf)), using ImageJ. As a method of normalization, the overall width of the brain was measured as the longest distance across the midbrain of 4-mpf control, flt4−/− and ccbe1 morphants (Extended Data Fig. 8p,q).
To assess overall brain vessel density, midbrains of 4 mpf flt4 mutants, control siblings and age-matched ccbe1 MO-injected animals were analysed using ImageJ. Vessels marked by fli1a:GFP were segmented using the threshold function, and the quantified vessel area was normalized to brain area to calculate vascular density as a percentage. Using the same set of images, we used an approach based on a previously published method to examine heterogeneous vascular network patterning phenotypes68. The density of five distinct regions per brain across the surface meningeal vasculature were quantified, with equivalent regions selected for each brain to avoid any sampling bias. This captured intra-sample spatial heterogeneity to measure vascular patterning defects in each animal. Then, variance values per animal were calculated and were compared across the groups; control, flt4 mutants and ccbe1 MO-injected animals as shown in Fig. 3h and Extended Data Fig. 8n.
All images were processed using ImageJ2 v.2.9.0/1.53t (National Institutes of Health) software.
Sample preparation for scRNA-seq and snATAC-seq
Wounding dataset
Tg(kdrl:Hsa.HRAS-mCherry)s916 transgenic zebrafish were raised at 28.5 °C to 6 dpf. Fish were then mounted dorsally in 1% low gelling agarose (BioRad) as previously described in ref. 67, and injured with a single 27 G needle insertion in 1 of the 2 hemispheres of the midbrain of the larval fish. Fish were then gently recovered in embryo water and incubated at 28.5 °C for 24 h. Injured fish and uninjured control siblings were then euthanized with 0.04% tricaine methanesulfonate in system water, on an ice-cold pad. Heads were dissected and collected in sterile Eppendorf tubes. Single cells were isolated from Tg(kdrl:Hsa.HRAS-mCherry)s916 wounded and control transgenic zebrafish larval heads as previously described in ref. 14. The BD FACSAria Fusion 5 Flow Cytometer (BD Biosciences) was used to sort a population of Zombie Violet−/DsRed+ cells. Cells were sorted directly into cold 100% FBS (Lonza). Sorted cells were taken immediately to the sequencing facility and used for scRNA-seq.
Mouse dataset
Meninges from two female C57 mice at 3 months of age were dissociated as previously described in refs. 69,70. In brief, mice were euthanized using an intraperitoneal injection of sodium pentobarbitone (1.6 mg g−1 body weight), before being transcardially perfused with 15 ml of PBS. Brains were immediately removed, and thin shavings of the cortical surface collected with scalpel. Collected tissue was diced with a scalpel and digested for 16 min at 37 °C in 0.1% papain (Worthington Biochemical) and 0.1% DNaseI (Roche Australia) in Hank’s buffered salt solution (ThermoScientific), with titration to dissociate the tissue. Cells were washed and resuspended in wash media (Hank’s buffered salt solution containing 10% fetal calf serum), then filtered into a single-cell suspension through a 100-µm sieve (Falcon, BD Biosciences). Dissociated cells were suspended in 500 μl of wash media and incubated with 6 μl of acetylated-LDL-Alexa488 (Invitrogen) for 1 h at 37 °C. Cells were washed twice with 5 ml of wash media and strained through a 70-μm cell strainer. Then 10 μl of Draq7 were added to the cell suspension, and 488acLDL+/Draq7− cells were fluorescence-activated cell sorted using a Cytopeia Influx Cell Sorter (BD Bioscience) into 100% FBS. Cells were subsequently quantified and loaded onto a 10X Chromium platform scRNA-seq at the Institute for Molecular Bioscience Genomics Facility (University of Queensland).
osr2 mutant dataset
Fifteen osr2−/− and 22 sibling animals were raised to 30 dpf, carefully genotyped and euthanized with 0.04% tricaine methanesulfonate in system water, on an ice-cold pad. As much as possible, tissues surrounding the brain were carefully removed by dissection to facilitate the dissociation of brain tissues without affecting the dorsal leptomeningeal layers. Tissues were collected in sterile Eppendorf tubes. Single cells were isolated from Tg(−5.2lyve1b:DsRed)nz101; Tg(fli1a:nGFP)y7 osr2−/− and sibling transgenic zebrafish 30-dpf brains as previously described in ref. 14. The BD FACSAria Fusion 5 Flow Cytometer (BD Biosciences) was used to sort a population of Zombie Violet−/GFP+/DsRed+ cells. Cells were sorted directly into cold 100% FBS (Lonza). Sorted cells were taken immediately to the sequencing facility and used for scRNA-seq.
scRNA-seq library preparation
Library preparation and sequencing was performed at the Molecular Genomics Core Facility (PMCC). Single-cell suspensions from fluorescence-activated cell sorting were spun down and a cell count was performed to determine postsort viability and cell concentration. The single-cell suspension was partitioned and barcoded using the 10X Genomics Chromium iX Controller (10X Genomics) and the Single Cell 3′ Library and Gel Bead Kit (Wounded dataset, V3.1; 10X Genomics; PN-1000121, mouse dataset V2). The cells were loaded onto the Chromium Single Cell Chip G (10X Genomics; PN-1000120) to target 10,000 cells. Gel bead in Emulsion generation and barcoding, complementary DNA amplification and library construction was performed according to the 10X Genomics Chromium User Guide. The resulting single-cell transcriptome libraries contained unique sample indices for each sample. The libraries were quantified on the Agilent TapeStation 4200 using High Sensitivity D1000 ScreenTape and reagents (Agilent, 5067-5585, 5067-5584). Single-cell transcriptome libraries were pooled in equimolar ratios before being sequenced (wounded dataset: Illumina NovaSeq 6000 using S4 300-cycle chemistry; mouse dataset: Illumina NextSeq 500using S4 300-cycle chemistry; osr2 mutant dataset: Illumina NextSeq 6000 using S4 300-cycle chemistry). Read1 supplies the cell barcode and UMI, i7 and i5 the sample indexes, and Read2 the 3′ sequence of the transcript. Sequencing read lengths were trimmed to 28 bp (read 1), 10 bp (i7 index), 10 bp (i5 index) and 91 bp (read 2), ensuring compatibility with the 10X Genomics analysis software, Cell Ranger. After sequencing, fastq files were generated using bcl2fastq2 (v.2.20.0.422).
scRNA-seq data processing
The following general workflow was applied to all datasets. Detailed parameter descriptions and version for software for preprocessing, genome versions, sample specific metrics and cluster annotation details are described in Supplementary Data Table 1. For details about R package versions, please see the renv.lock file at https://atlassian.petermac.org.au/bitbucket/projects/HL/repos/hogan_lab/browse/2024_Gaudi_et_al_muLEC. Sequencing quality was assessed using FastQC71 v.0.11.6 and MultiQC72 v.1.7 viewer for aggregated reports. Cell Ranger73 v.6.0.2 or v.5.0.0 count (and aggr where appropriate) was used to generate count files. Doublets were identified from the filtered aggregated count files using Scrublet74 in Python v.3.6 or scDblFinder v.1.12 in R statistical software v.4.2.2. Sequence reads corresponding to ribosomal and global genes were removed, cells filtered according to library size and mitochondrial content, normalized, followed by uniform manifold approximation and projection (UMAP) dimension reduction, clustering (louvain) and cell-cycle analysis using Seurat75 v.4.1.1. The quality of the dataset was evaluated before and after normalization using both custom plots and built-in functions in Seurat76 and scater77 v.1.26.1. Cluster solutions were evaluated, the most appropriate resolution selected using ClusTree78 v.0.5.0, and cluster identity was defined using key markers of phenotype.
The datasets in the following subsections had modifications to the general workflow.
Zebrafish developmental time-course dataset
In addition to the 7 dpf wild-type sample generated in this study, previously published14 fastq files for 40 hpf, 3 dpf, 4 dpf and 5 dpf were used. After normalization and filtering (as described above and in Supplementary Data Table 1), all samples were merged into one Seurat object using the merge function. The merged object was subsequently scaled using all features and splitting by sample name. FindVariableFeatures function was run with selection.method = ‘vst’ and nfeatures = 5,000 followed by principal components analysis dimension reduction (npcs = 20) and UMAP (dims = 1:20). Finally, we applied CSS simspec78 v.0.0.9 function cluster_sim_spectrum with label_tag = ‘Stage’ and repeated UMAP dimension reduction and clustering with reduction = ‘css’.
osr2 mutant dataset
Cellbender79 v.0.3.0 was applied to remove potential ambient RNA from these samples using the remove-background function with a learning rate of 106. Cellbender outputs were converted to a Seurat compatible format using pytables v.3.9.2.
GSE245311 human 23 weeks postconception dataset
Published H5Seurat files for the data presented in figure 3b from ref. 30 (GSE245311) were converted to Seurat objects using SeuratDisk80 v.0.0.0.9021. Following the original author’s code, cell type labels were adjusted and detailed immune-cell labels were obtained by matching barcodes from the subset of immune-cell dataset (figure 4d in the original publication). Finally, only cells from 23 weeks postconception from the leptomeninges were retained and reprocessed using a standard Seurat workflow without sc transformation81. The amended published cell type annotations were used for downstream analysis.
PRJNA826269 human dataset
In agreement with the original publication in ref. 31 (PRJNA826269), we identified a cluster expressing both barrier-associated macrophage (BAM) as well as microglia genes. For simplicity, this cluster was named ‘BAM’ in the main figure (Fig. 3b,f).
DanioCell dataset
The processed Seurat object and corresponding annotations published in ref. 38 were downloaded and subsequently subset into tissue specific objects as described by the original authors. The EC clusters in the ‘haematopoietic/vasculature’ object were then annotated in greater detail using expression of marker genes. Renaming of clusters is outlined in more detail in Supplementary Table 5.
ABC mouse atlas
The Allen Brain Cell (ABC) mouse atlas was accessed using the interactive web viewer using the ‘Mouse whole-brain transcriptomic cell type atlas’ dataset provided by the original authors of ref. 29. Plots presented in Extended Data Fig. 5d were acquired using the provided download function.
scRNA-seq data analysis
All plots and visualizations were performed using Seurat76 v.4.1.1, ggpubr v.0.4.0 or ggplot2 (ref. 82) v.3.4.1 as described in scripting.
Marker genes and differential gene expression
Marker genes for relevant datasets were calculated using the FindAllMarkers function with only.pos = TRUE, min.pct = 0.1, return.thresh = 1 and logfc.threshold = 0. For cross-species correlation analysis, marker genes were calculated using FindAllMarkers function with only.pos = FALSE, min.pct = 0.1, return.thresh = 1 and logfc.threshold = 0. Further filtering of this list was done as appropriate for each downstream analysis tool. For the larval brain wounding scRNA-seq dataset, we identify DEGs between wounded and unwounded cells for each cell type using the Seurat function FindMarkers with the default parameters except min.pct = 0.1, logfc.threshold = 0, return.threshold = 1 and only.pos = FALSE. For the lollipop plot in Extended Data Fig. 8e, genes with adjusted P < 0.05 and log2 fold change (log2[FC]) greater than 0.5 were considered DEG. For the osr2 mutant scRNA-seq dataset, we identified DEG between osr2 mutant and wild-type siblings for each cell type using the Seurat function FindMarkers with the default parameters except min.pct = 0.1, logfc.threshold = 0, return.threshold = 1 and only.pos = FALSE. For the stacked bar plot in Fig. 2i genes with adjusted P < 0.05 and absolute log2[FC] greater than 0.5 were considered DEG.
Gene set enrichment analysis
All gene set enrichment analyses were performed using enrichR83 v.3.2, setting EnrichR site84 to FishEnrichr and using GO_Biological_Process_2018, GO_Cellular_Component_2018 and GO_Molecular_Function_2018 databases. Genes with adjusted P < 0.05 and log2[FC] > 0.5 were considered significantly upregulated. For GO terms upregulated in wounded versus control muLEC, GO Biological Process terms with adjusted P < 0.05 were submitted to REVIGO85 v.1.8.2 (http://revigo.irb.hr/) using the httr86 v.1.4.4 package using the following parameters: cutoff = ‘0.7’, speciesTaxon = ‘7955’, measure = ‘STIMREL’, namespace = ‘1’ and output.type = ‘Xgmml’. Finally, the Xgmml files were imported to Cystoscape87 v.3.10.2 for network visualization. For GO terms upregulated in muLEC versus LEC in the wounded sample, all GO categories were considered and the top ten (number of genes overlap with gene set) terms are shown.
Correlation analysis
For the correlation analysis within the zebrafish larval brain wounding dataset, Spearman correlation coefficients between cell types were calculated using scaled normalized expression data for all genes and performed pairwise for each cell type using the R function cor.test in the stats v.3.6.2 package with method = ‘spearman’ and exact = F. For the cross-species analysis, only pairwise marker genes (adjusted P < 0.05, no log2[FC] threshold, upregulated and downregulated) present in both species were considered for correlation and gene names for the mouse and human datasets were translated to zfin or mgi symbols using biomaRt88 v.2.52.0 with host https://dec2021.archive.ensembl.org and uniqueRows = T. For mouse or human gene symbols that mapped to multiple zfin symbols, all orthologues were kept. Scatterplots were generated using the ggscatter function and genes with high shared or exclusive expression for muLEC, BAM or macrophages were coloured. Although all these genes were investigated in more detail, only selected genes have been labelled. Density plots were plotted using the geom_bin2d function and scores represent log-transformed counts. All genes and their cross-species translations can be found at https://atlassian.petermac.org.au/bitbucket/projects/HL/repos/hogan_lab/browse/2024_Gaudi_et_al_muLEC/data/cross_species.
Jaccard similarity analysis
Jaccard similarity was calculated between sets of marker genes between cell types using a custom function. Marker genes were defined as adjusted P < 0.05 and log2[FC] > 0.75 for all datasets, except the GSE245311 human dataset for which a log2[FC] threshold of 0.5 was used. For the cross-species analysis, gene names for the mouse and human dataset were translated to zfin and mgi symbols using biomaRt v.2.52.0 with host https://dec2021.archive.ensembl.org and uniqueRows = T.
Mixing metric
For relevant datasets, we calculated the mixing metric within each cell type separately using the Seurat function MixingMetric. If the number of cells exceeded 300, max.k was set to the number of cells, if not max.k was set to 300. Average mixing score per cell type was then added as a metadata column to the Seurat object for plotting. In addition, we performed statistical analysis using log-transformed metric scores to test for differences between cell types using the function compare_means in ggpubr v.0.4.0 with method = ‘t.test’ for pairwise, and method = ‘anova’ for global comparisons. Mixing metric scores for all cell types can be found at https://atlassian.petermac.org.au/bitbucket/projects/HL/repos/hogan_lab/browse/2024_Gaudi_et_al_muLEC/data/mixing_metric.
Velocyto
Estimates of RNA velocities were calculated for each sample using velocyto89 v.0.17.17, and combined for each dataset using loompy v.2.0.10. Finally, RNA velocities were transferred onto previously generated UMAPs using the velocyto.R89 v.0.6 package. Genes were filtered using the filter.genes.by.cluster.expression function with min.max.cluster.average = 0.1 (spliced) and 0.05 (unspliced). Velocity estimates were computed using the gene.relative.velocity.estimates function using the following parameters: deltaT = 3, kCells = 10, fit.quantile = 0.02. Velocity arrows were then plotted on embeddings using the show.velocity.on.embedding.cor function with the following parameters: n = 400 and scale = ‘sqrt’.
Trajectory analysis
In addition to velocyto, pseudotime analysis was performed using slingshot90 in the ExtendSeurat v.1.1.6 package with the RunSlingshot function using default parameters and group.by = ‘L2_Seurat_cluster_predicted_phenotype’ (for zebrafish developmental dataset) and ‘L2_predicted_phenotype’ (for osr2 mutant dataset). Smoothed gene expression trends were predicted by a generalized additive model following gene expression ~ s(Pseudotime) using the gam function from the mgcv91 v.1.8-42 package. Gene expression trend plots presented in Extended Data Fig. 2c–c” were created by adapting the GeneTrendCurve.Slingshot function in the ExtendSeurat package. In addition, we tested for statistically significant differences in pseudotime for each cell type between genotypes using the package ggpubr v.0.4.0 with method = ‘wilcox.test’ (default) for pairwise, and method = ‘kruskal.test’ for global comparisons (Extended Data Fig. 4f). Pseudotime predictions for all pseudotime trajectories can be found at https://atlassian.petermac.org.au/bitbucket/projects/HL/repos/hogan_lab/browse/2024_Gaudi_et_al_muLEC/data/pseudotime.
snATAC-seq data processing and analysis
The snATAC dataset was previously published in ref. 14. Using ArchR92 v.1.0.2, the ArchR object shared by the original authors was subset to only include cell types of interest (VEC, VEC_02_03, LEC, arterial EC, endocardium and muLEC) and pseudo replicates and peak calling using MACS2 (ref. 93) v.2.1.1 was repeated using groupBy = ‘Phenotype’. Local chromatin accessibility of osr2 (50 kb upstream and 12 kb downstream) was visualized using the plotBrowserTrack for muLEC, LEC and VEC.
Statistical analysis
Graphic representations and statistical analyses were performed using GraphPad Prism v.10. The Shapiro–Wilk test was applied to test normal distribution of the data. When comparing two groups with normal distribution a two-tailed Student’s t-test was used. For multiple group comparisons, a one-way analysis of variance (ANOVA) test was used for normally distributed data. The distribution of per-animal variance values in Fig. 3h and Extended Data Fig. 8n was tested using Levene’s test in Python (v.3.11).
All the scatterplots presented throughout the figures included a depiction of standard error of the mean (s.e.m.). Not significant (NS) P > 0.05, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. Further details on statistics can be found in each figure legend.
Illustration design
Illustrations were designed using BioRender (https://biorender.com; main-text figures, created in BioRender. Usseglio gaudi, A. (2025) https://BioRender.com/w7lx2hd; Extended Data figures, created in BioRender. Usseglio gaudi, A. (2025) https://BioRender.com/2l3p654). For brain sections depicted in Fig. 4 and Extended Data Fig. 9, we used either the imaged sections or published references to match the stage of development of the brains94.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw and processed files for all datasets published in this study can be found in Gene Expression Omnibus (GEO) under accession code GSE270486. Fastq files for previously published scRNA-seq datasets14 (GSE188342) were shared by the authors. The ArchR project for the previously published snATAC dataset14 (fastq files are available at GSE188342, sample GSM5677990) was shared by the authors for use in this paper. For the human meningioma dataset, fastq files for selected samples (DUNN_MEN10, DUNN_MEN09, DUNN_MEN08, DUNN_MEN13) were downloaded from National Center for Biotechnology Information (BioProject accession PRJNA826269, ref. 31). For the human developmental dataset, H5Seurat were downloaded from GEO (accession GSE245311, ref. 30). The DanioCell dataset was downloaded as a preprocessed Seurat object from https://daniocell.nichd.nih.gov/assets/obj-download. The ABC mouse atlas was accessed through the ABC Atlas portal at https://portal.brain-map.org/atlases-and-data/bkp/abc-atlas using the interactive web viewer at https://knowledge.brain-map.org/abcatlas#AQEBQVA4Sk5ONUxZQUJHVk1H S1kxQgACUTFOQ1dXUEc2RlowRE5JWEpCUQADAAQBAQKBrOWEgbxpYgOF4HIShP4xxwAFAAYBAQJGUzAwRFhWMFQ5UjFYOUZKNFFFAAN%2BAAAABAAACEc0STRHRkpYSkI5QVRaM1BUWDEACUxWREJKQVc4Qkk1WVNTMVFVQkcACgALAW5vbmUAAm5vbmUAAwEEAQACIzAwMDAwMAADyAEABQE BAiMwMDAwMDAAA8gBAAAAAgEA. All data generated to support this study are available within the paper and its Supplementary Information. Genomic datasets can be found in GEO under accession code GSE270486. Links to previously published datasets and code used to support this study can be found in Methods in the Data availability and Code availability sections. Source data are provided with this paper.
Code availability
All code and scripts associated with this analysis are publicly available under an open-source software licence through BitBucket at https://atlassian.petermac.org.au/bitbucket/projects/HL/repos/hogan_lab/browse/2024_Gaudi_et_al_muLEC.
References
Zhao, Z., Nelson, A. R., Betsholtz, C. & Zlokovic, B. V. Establishment and dysfunction of the blood–brain barrier. Cell 163, 1064–1078 (2015).
Herz, J., Filiano, A. J., Wiltbank, A. T., Yogev, N. & Kipnis, J. Myeloid cells in the central nervous system. Immunity 46, 943–956 (2017).
Bower, N. I. et al. Mural lymphatic endothelial cells regulate meningeal angiogenesis in the zebrafish. Nat. Neurosci. 20, 774–783 (2017).
van Lessen, M. et al. Intracellular uptake of macromolecules by brain lymphatic endothelial cells during zebrafish embryonic development. eLife https://doi.org/10.7554/eLife.25932 (2017).
Venero Galanternik, M. et al. A novel perivascular cell population in the zebrafish brain. eLife https://doi.org/10.7554/eLife.24369 (2017).
Kierdorf, K., Masuda, T., Jordao, M. J. C. & Prinz, M. Macrophages at CNS interfaces: ontogeny and function in health and disease. Nat. Rev. Neurosci. 20, 547–562 (2019).
Vara-Perez, M. & Movahedi, K. Border-associated macrophages as gatekeepers of brain homeostasis and immunity. Immunity 58, 1085–1100 (2025).
Schonhoff, A. M. et al. Border-associated macrophages mediate the neuroinflammatory response in an alpha-synuclein model of Parkinson disease. Nat. Commun. 14, 3754 (2023).
De Schepper, S. et al. Perivascular cells induce microglial phagocytic states and synaptic engulfment via SPP1 in mouse models of Alzheimer’s disease. Nat. Neurosci. 26, 406–415 (2023).
Gerganova, G., Riddell, A. & Miller, A. A. CNS border-associated macrophages in the homeostatic and ischaemic brain. Pharmacol. Ther. 240, 108220 (2022).
Ochocka, N. et al. Single-cell RNA sequencing reveals functional heterogeneity of glioma-associated brain macrophages. Nat. Commun. 12, 1151 (2021).
O’Brown, N. M., Pfau, S. J. & Gu, C. Bridging barriers: a comparative look at the blood-brain barrier across organisms. Genes Dev. 32, 466–478 (2018).
Renshaw, S. A. & Trede, N. S. A model 450 million years in the making: zebrafish and vertebrate immunity. Dis. Model Mech. 5, 38–47 (2012).
Grimm, L. et al. Single-cell analysis of lymphatic endothelial cell fate specification and differentiation during zebrafish development. EMBO J. 42, e112590 (2023).
Li, J. et al. Neural-activity-regulated and glia-mediated control of brain lymphatic development. Cell 188, 3274–3290.e3216 (2025).
Campbell, F. et al. Directing nanoparticle biodistribution through evasion and exploitation of Stab2-dependent nanoparticle uptake. ACS Nano 12, 2138–2150 (2018).
Sorensen, K. K. et al. The scavenger endothelial cell: a new player in homeostasis and immunity. Am. J. Physiol. Regul. Integr. Comp. Physiol. 303, R1217–R1230 (2012).
Huisman, Y. et al. Meningeal lymphatic endothelial cells fulfill scavenger endothelial cell function and cooperate with microglia in waste removal from the brain. Glia 70, 35–49 (2022).
Uphoff, K., Suarez, I., van Impel, A. & Schulte-Merker, S. dab2 is required for the scavenging function of lymphatic endothelial cells in the zebrafish meninges. Sci. Rep. 14, 27942 (2024).
Mishra, S. K. et al. Disabled-2 exhibits the properties of a cargo-selective endocytic clathrin adaptor. EMBO J. 21, 4915–4926 (2002).
Lan, Y. et al. Odd-skipped related 2 (Osr2) encodes a key intrinsic regulator of secondary palate growth and morphogenesis. Development 131, 3207–3216 (2004).
Kawai, S., Yamauchi, M., Wakisaka, S., Ooshima, T. & Amano, A. Zinc-finger transcription factor odd-skipped related 2 is one of the regulators in osteoblast proliferation and bone formation. J. Bone Miner. Res. 22, 1362–1372 (2007).
Gao, Y., Lan, Y., Liu, H. & Jiang, R. The zinc finger transcription factors Osr1 and Osr2 control synovial joint formation. Dev. Biol. 352, 83–91 (2011).
Lam, P. Y. et al. odd-skipped related 2 is required for fin chondrogenesis in zebrafish. Dev. Dyn. 242, 1284–1292 (2013).
Uphoff, K. et al. SVEP1 enables efficient binding of angiopoietin-2 to the TIE1 receptor, allowing receptor phosphorylation and downstream signaling. eLife 14, RP106338 (2025).
McDonald, D. M. et al. Cerebrospinal fluid draining lymphatics in health and disease: advances and controversies. Nat. Cardiovasc. Res. 4, 1047–1065 (2025).
Shibata-Germanos, S. et al. Structural and functional conservation of non-lumenized lymphatic endothelial cells in the mammalian leptomeninges. Acta Neuropathol. 139, 383–401 (2020).
Dermitzakis, I. et al. CNS border-associated macrophages: ontogeny and potential implication in disease. Curr. Issues Mol. Biol. 45, 4285–4300 (2023).
Yao, Z. et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317–332 (2023).
Sankowski, R. et al. Multiomic spatial landscape of innate immune cells at human central nervous system borders. Nat. Med. 30, 186–198 (2024).
Wang, A. Z. et al. Single-cell profiling of human dura and meningioma reveals cellular meningeal landscape and insights into meningioma immune response. Genome. Med. 14, 49 (2022).
Masuda, T., Amann, L. & Prinz, M. Novel insights into the origin and development of CNS macrophage subsets. Clin. Transl. Med. 12, e1096 (2022).
Chen, J. et al. Cerebrovascular injuries induce lymphatic invasion into brain parenchyma to guide vascular regeneration in zebrafish. Dev. Cell 49, 697–710.e695 (2019).
Chen, J. et al. Acute brain vascular regeneration occurs via lymphatic transdifferentiation. Dev. Cell 56, 3115–3127 (2021).
Wei, K. H. et al. Comparative single-cell profiling reveals distinct cardiac resident macrophages essential for zebrafish heart regeneration. eLife https://doi.org/10.7554/eLife.84679 (2023).
Jones, L. O. et al. Single-cell resolution of the adult zebrafish intestine under conventional conditions and in response to an acute Vibrio cholerae infection. Cell Rep. 42, 113407 (2023).
Rougeot, J. et al. RNAseq profiling of leukocyte populations in zebrafish larvae reveals a cxcl11 chemokine gene as a marker of macrophage polarization during mycobacterial infection. Front. Immunol. 10, 832 (2019).
Sur, A. et al. Single-cell analysis of shared signatures and transcriptional diversity during zebrafish development. Dev. Cell 58, 3028–3047 (2023).
Sun, R. & Jiang, H. Border-associated macrophages in the central nervous system. J. Neuroinflammation 21, 67 (2024).
Green, L. A., O’Dea, M. R., Hoover, C. A., DeSantis, D. F. & Smith, C. J. The embryonic zebrafish brain is seeded by a lymphatic-dependent population of mrc1(+) microglia precursors. Nat. Neurosci. 25, 849–864 (2022).
Hogan, B. M. et al. Ccbe1 is required for embryonic lymphangiogenesis and venous sprouting. Nat. Genet. 41, 396–398 (2009).
Venero Galanternik, M. et al. Anatomical and molecular characterization of the zebrafish meninges. Preprint at bioRxiv https://doi.org/10.1101/2025.04.09.646894 (2025).
Stricker, S., Brieske, N., Haupt, J. & Mundlos, S. Comparative expression pattern of Odd-skipped related genes Osr1 and Osr2 in chick embryonic development. Gene Expr. Patterns 6, 826–834 (2006).
Kazenwadel, J. et al. A Prox1 enhancer represses haematopoiesis in the lymphatic vasculature. Nature 614, 343–348 (2023).
Das, R. N. et al. Generation of specialized blood vessels via lymphatic transdifferentiation. Nature 606, 570–575 (2022).
Jin, S.-W., Beis, D., Mitchell, T., Chen, J.-N. & Stainier, D. Y. Cellular and molecular analyses of vascular tube and lumen formation in zebrafish. Development 132, 5199–5209 (2005).
Lawson, N. D. & Weinstein, B. M. In vivo imaging of embryonic vascular development using transgenic zebrafish. Dev. Biol. 248, 307–318 (2002).
Okuda, K. S. et al. lyve1 expression reveals novel lymphatic vessels and new mechanisms for lymphatic vessel development in zebrafish. Development 139, 2381–2391 (2012).
Davison, J. M. et al. Transactivation from Gal4-VP16 transgenic insertions for tissue-specific cell labeling and ablation in zebrafish. Dev. Biol. 304, 811–824 (2007).
Chi, N. C. et al. Foxn4 directly regulates tbx2b expression and atrioventricular canal formation. Genes Dev. 22, 734–739 (2008).
Palha, N. et al. Real-time whole-body visualization of Chikungunya virus infection and host interferon response in zebrafish. PLoS Pathog. 9, e1003619 (2013).
Park, J. et al. Lysosome-rich enterocytes mediate protein absorption in the vertebrate gut. Dev. Cell 51, 7–20 (2019).
Wang, G. et al. Specific fibroblast subpopulations and neuronal structures provide local sources of Vegfc-processing components during zebrafish lymphangiogenesis. Nat. Commun. 11, 2724 (2020).
Asakawa, K. & Kawakami, K. Targeted gene expression by the Gal4-UAS system in zebrafish. Dev. Growth Differ. 50, 391–399 (2008).
de Oliveira, S. et al. Metformin modulates innate immune-mediated inflammation and early progression of NAFLD-associated hepatocellular carcinoma in zebrafish. J. Hepatol. 70, 710–721 (2019).
Michael, C., Mendonca-Gomes, J. M., DePaolo, C. W., Di Cristofano, A. & de Oliveira, S. A zebrafish xenotransplant model of anaplastic thyroid cancer to study tumor microenvironment and innate immune cell interactions in vivo. Endocr. Relat. Cancer https://doi.org/10.1530/ERC-23-0195 (2024).
Kok, F. O. et al. Reverse genetic screening reveals poor correlation between morpholino-induced and mutant phenotypes in zebrafish. Dev. Cell 32, 97–108 (2015).
Miles, L. B. et al. CRIMP: a CRISPR/Cas9 insertional mutagenesis protocol and toolkit. Nat. Commun. 15, 5011 (2024).
Stolper, J. et al. The human FLT1 regulatory element directs vascular expression and modulates angiogenesis pathways in vitro and in vivo. Preprint at bioRxiv https://doi.org/10.1101/2021.03.03.433738 (2021).
Westerfield, M. A Guide for the Laboratory Use of Zebrafish (Danio rerio) (Univ. Oregon Press, 2000).
Kawakami, K. Tol2: a versatile gene transfer vector in vertebrates. Genome Biol. 8, S7 (2007).
Wu, R. S. et al. A rapid method for directed gene knockout for screening in G0 zebrafish. Dev. Cell 46, 112–125 (2018).
Gagnon, J. A. et al. Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS ONE 9, e98186 (2014).
Davis, A. E., Castranova, D. & Weinstein, B. M. Rapid generation of pigment free, immobile zebrafish embryos and larvae in any genetic background using CRISPR–Cas9 dgRNPs. Zebrafish 18, 235–242 (2021).
Karlsson, J., von Hofsten, J. & Olsson, P. E. Generating transparent zebrafish: a refined method to improve detection of gene expression during embryonic development. Mar. Biotechnol. 3, 522–527 (2001).
Yang, L., Rojas, A. M. & Shiau, C. E. Liposomal clodronate-mediated macrophage depletion in the zebrafish model. Bio Protoc. 11, e3951 (2021).
Okuda, K. S., Baek, S. & Hogan, B. M. Visualization and tools for analysis of zebrafish lymphatic development. Methods Mol. Biol. 1846, 55–70 (2018).
Kasper, D. M. et al. MicroRNAs establish uniform traits during the architecture of vertebrate embryos. Dev. Cell 40, 552–565 (2017).
Willis, E. F. et al. Repopulating microglia promote brain repair in an IL-6-dependent manner. Cell 180, 833–846 (2020).
Chauquet, S. et al. Exercise rejuvenates microglia and reverses T cell accumulation in the aged female mouse brain. Aging Cell https://doi.org/10.1111/acel.14172 (2024).
Andrews, S. FastQC: a quality control tool for high throughput sequencing data. bioinformatics.babraham.ac.uk http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010)
Ewels, P., Magnusson, M., Lundin, S. & Kaller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
McCarthy, D. J., Campbell, K. R., Lun, A. T. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Zappia, L. & Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. Gigascience https://doi.org/10.1093/gigascience/giy083 (2018).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat. Methods 20, 1323–1335 (2023).
Hoffman, P. SeuratDisk: interfaces for HDF5-based single cell file formats, version 0.0.0.9021 (2023).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis, version 3.4.1 (Springer, 2016).
Jawaid, W. enrichR: provides an R Interface to ‘Enrichr’, version 3.2 (2023).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinf. 14, 128 (2013).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Wickham, H. httr: tools for Working with URLs and HTTP, version 1.4.5 (2022).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Wood, S. N. Fast stable restricted maximum likelihood and marginal likelihood estimation of semiparametric generalized linear models. J. R. Stat. Soc. Ser. B: Stat. Methodol. 73, 3–36 (2010).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Gaspar, J. M. Improved peak-calling with MACS2. Preprint at bioRxiv https://doi.org/10.1101/496521 (2018).
Wei, X. et al. Single-cell Stereo-seq reveals induced progenitor cells involved in axolotl brain regeneration. Science 377, eabp9444 (2022).
Acknowledgements
We dedicate this paper to the memory of Dr Neil Bower, our colleague and friend, who co-led the early stages of the project. This project was supported by funding from the Bright Focus Foundation USA (grant no. A2018807S) to N.I.B. and B.M.H. and the Australian Research Council (ARC) (grant no. DP200101668). B.M.H. was supported by a National Health and Medical Research Council (NHMRC) Senior Research Fellowship (grant no. 1155221) and Investigator grant (no. 2033008). P.D.C. and F.J.T. were supported by the ARC (grant no. DP200103219) and P.D.C. by an NHMRC Investigator grant (no. 2016338). This work was made possible in part through B.M.H. and O.F.B.’s membership of, and financial support from, the Brain Cancer Centre. M.R.M.H. and S.C. were supported by the NINDS (grant no. R01NS126209) and S.C. by the Ehrenkranz Young Scientist Award. We thank T. Semple and the PMCC Molecular Genomics Core Facility (RRID:SCR_025695), R. Lupat and the PMCC Bioinformatics Core Facility (RRID:SCR_025901), Research Computing Facility, RLS core (RRID:SCR_025699), Research Flow Core Facility (RRID:SCR_025550), Centre for Advanced Histology and Microscopy (RRID:SCR_025432) and Genomics Core at IMB (UQ). We also thank E. Wong, M. Nakayama, C. Bladen, A. Cox, R. Brown, M. Ramialison, J. Mao, J. da Silva Viana, L. Tobar, V. Garside and R. de Iongh for academic discussions, technical support and providing biological samples.
Author information
Authors and Affiliations
Contributions
N.I.B. and B.M.H. initially acted as co-senior authors. A.U.G. performed and analysed most zebrafish experimentation, HCR ISH experiments on multiple species and functional assays presented throughout the paper (all figures), and cowrote the paper. B.M.H. conceptualized experiments, analysed data, resourced the project and cowrote the paper. N.I.B. conceptualized key aspects of the project, performed experiments profiling mouse meningeal scavenger cells (for Fig. 3), resourced the project and analysed data together with J.V. and A.K.L. J.V. performed experimental dissection and optimized profiling of mouse meninges for Fig. 3. M.M. provided most computational and bioinformatic analyses across all figures supported by experimentation and analysis from E.M. and H.Y. M.M. cowrote the paper. O.F.B. performed experimentation and analysis for Figs. 1 and 2l and Extended Data Figs. 7 and 8 with A.U.G. S.C. and M.R.M.H. conceptualized, and performed experimentation and analysis for Fig. 3g–l and Extended Data Fig. 8m–q. F.J.T., R.J.B.-R., L.B.M. and P.D.C. generated and provided the osr2 knock-in strain for Fig. 1h,i. S.D., S.P. and M.C.R.G. optimized zebrafish single-cell preparation, genomics and CRISPR genetics presented in Figs. 1 and 2 and Extended Data Figs. 1–4. S.-A.S., J.K. and P.D.C optimized experimental approaches, conceptualized experiments and provided materials for Fig. 4 and Extended Data Fig. 9 relating to shark and axolotl. A.J.P. provided isolated dunnart tissues and conceptualized experimentation in Fig. 4 and Extended Data Fig. 9. K.A.S. provided isolated chicken tissues and conceptualized experimentation in Figs. 1 and 4 and Extended Data Fig. 9.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Gou Young Koh and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Additional data related to key findings in Fig. 1.
a, Confocal projection of lymphatics in a lateral view of 30 dpf whole zebrafish (Tg(−5.2lyve1b:DsRed)nz101, black). This image demonstrates the absence of muLECs from any location other than the brain. AFL, anal fin lymphatics; CarL, cardinal lymphatics; CFL, caudal fin lymphatics; CL, caudal lymphatics; DFL, dorsal fin lymphatics; DL, dorsal lymphatics; DuL, dural lymphatics; FL, facial lymphatics; iLV, intersegmental lymphatic vessel; JL, jugular lymphatics; LL, lateral lymphatics; muLEC, mural lymphatics; OL, orbital lymphatics; SL, spinal lymphatics; TD, thoracic duct; VL, ventral lymphatics. b, Cartoon depicting the origin of the scRNA-seq and single-nuclei ATAC-sequencing (snATAC-seq) data used in the study. c, UMAP visualisation of all cells (Level 01, RNA_snn_res.0.2, n = 37,903) captured from zebrafish embryos at different timepoints (40 hpf, 3, 4, 5 and 7 dpf) for scRNA-seq coloured by seurat cluster. d, Stacked bar plot showing composition of samples for each cell phenotype (c) for Level 01 dataset. e, UMAP visualisation of all cells (Level 01, RNA_snn_res.0.2, n = 37,903) captured from zebrafish embryos at different timepoints (40 hpf, 3, 4, 5 and 7 dpf) for scRNA-seq coloured by developmental stage. f, Dot plot showing the gene expression for key marker genes used to annotate clusters at Level 01. Size of the dot plots represents the proportion of expressing cells and colour represents average log-normalised expression. g-i, UMAP visualisation of re-clustered endothelial cells (Level 02, RNA_snn_res.0.3, n = 15,898) from zebrafish embryos at different timepoints (40 hpf, 3, 4, 5 and 7 dpf) coloured by cell phenotype. AEC, arterial EC; VEC, venous EC; imVEC immature VEC; sEC, sparc + EC; LEC, lymphatic EC; muLEC, mural LEC (g). Stacked bar plot showing cellular (h) and sample (i) composition per cell phenotype annotated seurat cluster for Level 02 dataset. j, Dot plot showing the gene expression for key marker genes used to annotate clusters at Level 02. Size of the dot plots represents the proportion of expressing cells and colour represents average log-normalised expression. a, Scale bar 500 μm.
Extended Data Fig. 2 Supporting data and dataset analysis related to Fig. 1.
a, UMAP of Level 02 endothelial cells split by timepoint (40 hpf, 3, 4, 5 and 7 dpf) and coloured by cell phenotype (top); UMAP visualisation of genes of interest osr2, mrc1a, lyve1b, prox1a and cdh6 (top to bottom). Colour scale represents log-normalised expression. b, UMAP of pseudotime trajectory analysis of Level 02 endothelial cells. Colour scale represents predicted pseudotime. Cells coloured in grey were not assigned to this trajectory. c-c”, Smoothed gene expression trends for osr2 (pink) and cdh6 (grey) along pseudotime trajectory (c). Smoothed gene expression trend for osr2 (c’) and cdh6 (c”) coloured by cell type. d, Confocal projection of osr2 expression (Tg(osr2-gal4, UAS:mTagBFP2);Tg(UAS:RFP) knock in, magenta) in a dorsal view of zebrafish embryos brain at different timepoints (3, 4, 5, 6 and 7 dpf). Representative of >10 animals imaged. e, Single nuclei ATAC sequencing data genome accessibility track of osr2 locus in LEC, muLEC and VEC at 4 dpf. Green box represents peak region +45osr2. Data from Grimm et al., 2023. f-f”, Confocal projections of +45osr2 zebrafish enhancer detection (ZED) vector expression (Tg(+45osr2:E1b:GFP)uom135, magenta) and lymphatics (Tg(−5.2lyve1b:DsRed)nz101, cyan) in whole zebrafish embryos at 7 dpf (top). Magnification of the facial lymphatics (f’) and trunk lymphatics (f”). Representative of >10 animals imaged. g-g”, Confocal projection of +45osr2 expression (magenta) and lymphatics (cyan) in a dorsal view of 7 dpf zebrafish embryo brain (g) and magnification of the lyve1b+ loop-like structure adjacent to the MV (g’-g”). Representative of >10 animals imaged. d,f,f’,f”,g Scale bar 100 μm. g’ Scale bar 25 μm.
Extended Data Fig. 3 Additional data related to key findings in Fig. 2.
a, Schematic of the CRISPR/Cas9-mediated frameshift leading to a 7 bp deletion and premature stop codon in the osr2 sequence (pThr51AlafsTer57; osr2uom133), predicted to result in the loss of three zinc-finger domains and a null allele. b, Quantification of total body length of the siblings (green) and osr2 mutants (blue) at different timepoints from Fig. 2 (7 (n = 40 siblings and n = 23 mutants), 14 (n = 14 and n = 8), 21 (n = 11 and n = 5) and 30 dpf (n = 21 and n = 19)). c, Quantification of total number of muLECs covering the MV in siblings (green) and osr2 mutants (osr2−/−, blue) at 7 dpf (n = 40 siblings and n = 23 mutants) from g. d, Quantification of total number of muLECs covering the MV in siblings (green) and osr2 mutants (osr2−/−, blue) at 14 dpf (n = 14 siblings and n = 8 mutants) from h. e, Quantification of total number of muLECs covering the MV in siblings (green) and osr2 mutants (osr2−/−, blue) at 21 dpf (n = 11 siblings and n = 5 mutants) from i. f, Quantification of total number of muLECs covering the MV in siblings (green) and osr2 mutants (osr2−/−, blue) at 7 dpf (n = 10 siblings and n = 10 mutants) from Fig. 2a. (Right): schematic of brain imaged in this figure. g-i’, Confocal projection of muLECs (Tg(−5.2lyve1b:DsRed)nz101, orange) of sibling (top) and osr2 mutant (osr2−/−, bottom) zebrafish brains and (g’-i’) magnification of lyve1b+ muLECs covering the MV in siblings (top) and osr2 mutants (osr2−/−, bottom). Stages as indicated. j, Quantification of muLEC length at MV in siblings (green) and mutants (blue) (7 dpf (n = 481 cells/n = 40 siblings, n = 285 cells/n = 23 mutants); 14 dpf (n = 302 cells/n = 14 siblings, n = 169 cells/n = 8 mutants); 21 dpf (n = 318 cells/n = 11 siblings, n = 144 cells/n = 5 mutants); 30 dpf (n = 253 cells/n = 10 siblings, n = 221 cells/n = 10 mutants) from Fig. 2a. Dots=cells. **** <0.0001. k, UMAP visualisation of all cells (Level 01, SCT_snn_res.0.2, n = 7,748 cells) captured from siblings and osr2 mutants zebrafish at 30 dpf for scRNA-seq coloured by Seurat cluster. l, UMAP visualisation of all cells (Level 01, SCT_snn_res.0.2, n = 7,748 cells) captured for scRNA-seq coloured by genotype. m, Stacked bar plot showing genotype composition of samples for Seurat clusters in Level 01 dataset. n, Dot plot showing gene expression for key marker genes used to annotate clusters at Level 01. Size of dot plots represents proportion of expressing cells. Colour=average log-normalised expression. o, UMAP visualisation of re-clustered endothelial cells (Level 02, SCT_snn_res.0.2, n = 6,942 cells) from siblings and osr2 mutant zebrafish at 30 dpf coloured by Seurat clusters. p, Dot plot showing gene expression for key marker genes used to annotate clusters at Level 02. Size of dot plots represents proportion of expressing cells. Colour=average log-normalised expression. q, Boxplot showing log-transformed mixing metric score for each cell-phenotype for Level 02 dataset. Cell number analysed: n = 931 (muLEC), n = 4528 (LEC), n = 508 (EC), n = 510 (specialised BEC), n = 465 (sparc EC), total n = 6942 cells. Bounds of box = 25th − 75th quantile, centre = median, whiskers extend to +/−1.5 times interquartile range. ANOVA for multiple comparisons, two-sided T-test for comparison to muLEC, adjusted for multiple testing using holm. (ANOVA p value = 3.094896e-166), Comparison to muLEC adjusted p-values: 5.9e-46 (EC), 9.8e-48 (LEC), 2.4e-24 (sparc_EC), 8.2e-14 (specialised BEC)****p = <0.0001. g, h, i, Scale bar 100 μm. g’, h’, i’, Scale bar 25 μm. One-way Analysis of Variance (ANOVA) for multiple comparisons, T-test for pairwise comparisons. ****p = <0.0001.
Extended Data Fig. 4 Supporting data and dataset analysis related to Fig. 2.
a, UMAP visualisation of RNA velocity on Level 02 endothelial cells of 30 dpf osr2 mutants and siblings from Fig. 2e. b, UMAP visualisation of Level 02 endothelial cells (n = 3,473 cells) of 30 dpf sibling zebrafish coloured by cell phenotype. c, UMAP visualisation of Level 02 endothelial cells (n = 3,543 cells) of 30 dpf osr2 mutant zebrafish coloured by cell phenotype. d, Bar plot showing the different expression of selected genes (log2FC on y axis) for osr2 mutant compared to siblings in all endothelial cell clusters from Fig. 2e. Two-sided Wilcox test, adjusted for multiple correction using Bonferroni. Exact adjusted p values for significant hits are 3.196e-50 (cldn5b, muLEC), 3.195e-34 (angpt2a, muLEC), 5.505e-24 (tie1, muLEC), 4.508e-22 (lgmn, LEC), 2.992e-10 (ctsla, LEC), 6.335e-09 (lamp2, LEC), 1.4774e-03 (ctsla, muLEC), 2.070e-02 (tie1, LEC). For a complete list of exact p values, please refer to Supplementary Data Table 3. * adjusted p < 0.05. e, UMAP of pseudotime trajectory analysis of Level 02 endothelial cells for siblings (left) and osr2 mutants (right). Colour scale represents predicted pseudotime. Cells coloured in grey were not assigned to this trajectory. f, Boxplot showing difference in pseudotimes for sibilings and osr2 mutants in all endothelial cell clusters from e. Only muLECs in osr2 mutant cells show a significantly decreased pseudotime compared to siblings (adjusted p-value = 2.6e-43) Boxplot spans 25th- 75 h quantiles, with the central line representing the median, whiskers extending to +− 1.5x interquartile ranges and all data points shown. Cell number analysed: n = 560 mutant, 371 wildtype (muLEC), n = 2308 mutant, 2220 wildtype (LEC), n = 268 mutant, 240 wildtype (EC), n = 210 mutant, 300 wildtype (specialised BEC), n = 212 mutant, 253 wildtype (sparc BEC). Global comparison Kruskal-Wallis p = 0.0051, and pairwise comparisons were performed using a two-sided Wilcox test corrected for multiple testing using the holm method. **** = p value < 0.0001. g, UMAP visualisation of expression of selected genes in siblings and osr2 mutants as indicated, from b-c. h-i, Confocal projection of muLECs (Tg(−5.2lyve1b:DsRed)nz101, magenta), blood vessels (Tg(kdrl:EGFP)s843, green), and tracer dyes (10 kDa Dextran, cyan; 2000kDa Dextran, blue) in 7dpf siblings (top) and osr2 mutants (osr2−/−, bottom) zebrafish brains (h). Magnification of left lyve1b+ loop (muLEC, magenta; 10 kDa Dextran, cyan) (i) from h. j, Quantification of tracer uptake index in siblings (green) and osr2 mutants (blue) 7 dpf zebrafish embryos from i (n = 48 sibling embryos and n = 22 mutant embryos). **p = 0.0032 unpaired t-test. k-l’, Confocal projection of lymphatic and vascular endothelial cells (Tg(−5.2lyve1b:DsRed)nz101, magenta) and cdh6 reporter (Tg(−48cdh6:E1b:GFP)uom134, green) in 7 dpf whole zebrafish (k-l) and magnification of the facial lymphatics and cdh6 expression (k’-l’). m, Confocal projection of lymphatics (Tg(−5.2lyve1b:DsRed)nz101, magenta) and blood vessels (Tg(kdrl:EGFP)s843, green) in 10 dpf siblings (left) and cdh6 mutants (cdh6−/−, right) zebrafish trunks. Representative of 3 mutant and 4 control animals imaged. h, k, l, m, Scale bar 100 μm. i, k’, l’, Scale bar 25 μm. Unpaired two-tailed t-test **p = 0.0032 for sibling versus osr2 mutants in j.
Extended Data Fig. 5 Additional data related to key findings in Fig. 3.
a, UMAP visualization of meningeal scavenging cells (Level 01, SCT_snn_res.0.2, n = 16,262 cells) coloured by Seurat cluster. VSMC, vascular smooth muscle cell; BAM, barrier associated macrophage. b, Stacked bar plot showing sample composition for cell phenotypes for Level 01 dataset from (a). c, Dot plot showing the gene expression for key marker genes used to annotate clusters at Level 01. d, UMAP visualisation of immune and vascular cells from the Allen Brain Cell Atlas (n = ~230,000 cells) coloured by class as provided by the original authors (top left), and by selected lymphatic and/or scavenging markers. OPC, oligodendrocyte precursor cell; BAM, barrier associated macrophage; ABC, arachnoid barrier cell; VLMC, vascular leptomeningeal cell; SMC, smooth muscle cell; Peri., pericyte; Endo., endothelial cell. Images were captured using the export function on the Brain Knowledge Platform. e, Workflow of single-cell RNA-sequencing on human leptomeningeal layers from previously published study PRJNA82626931 (left). UMAP visualisation of re-clustered selection (endothelial cells, Mesenchymal cells, antigen presenting, monocytes, macrophages, DC and B cells) of cell types (Level 02, SCT_snn_res.0.4, n = 8,478 cells) coloured by cell phenotype. MG, microglia; BAM, barrier associated macrophage. f, UMAP visualization of cells captured (Level 01, SCT_snn_res.0.4, n = 23,782 cells) in Wang et al.31 coloured by cell phenotype. DC, dendritic cells; NK cells, natural killer cells. g, Dot plot showing gene expression for marker genes used to annotate clusters at Level 01 from f. h, Stacked bar plot showing sample composition for each cell phenotype for Level 02 from e. I, Dot plot showing the gene expression for key marker genes used to annotate clusters at Level 02 dataset from e. j, Dot plot showing the gene expression of lymphatic and/or hematopoietic markers in all cell clusters of Level 02 GSE245311 (human developmental dataset from Fig. 3b) dataset. k, Dot plot showing the gene expression of lymphatic and/or hematopoietic markers in all cell clusters of Level 01 PRJNA826269 (human meningioma dataset from f dataset. l, Dot plot showing the gene expression of lymphatic and/or hematopoietic markers in all cell clusters of Level 02 PRJNA826269 (human meningioma dataset from e dataset. For all dot-plots; size of dots represents proportion of expressing cells and colour represents average log-normalised expression.
Extended Data Fig. 6 Additional data related to key findings in Fig. 3.
a, Heatmap visualisation of spearman correlation coefficients of scaled gene expression (left) and Jaccard similarity of marker genes (right) for all pairwise comparisons of zebrafish and mouse cell phenotypes. Colour represents correlation coefficient and Jaccard similarity, respectively. Correlation coefficient that did not pass significance testing are denoted with a x. b, Heatmap visualisation of spearman correlation coefficients of scaled gene expression (left) and Jaccard similarity of marker genes (right) for all pairwise comparisons of human developmental dataset (23 wpc) (Fig. 3b) and mouse cell phenotypes. Colour represents correlation coefficient and Jaccard similarity, respectively. Correlation coefficient that did not pass significance testing are denoted with a x. c, Heatmap visualisation of spearman correlation coefficients of scaled gene expression (left) and Jaccard similarity of marker genes (right) for all pairwise comparisons of human developmental dataset (23 wpc) (Fig. 3b) and zebrafish cell phenotypes. Colour represents correlation coefficient and Jaccard similarity, respectively. Correlation coefficient that did not pass significance testing are denoted with a x. d-d’, Heatmap visualisation of spearman correlation coefficients of scaled gene expression of pairwise marker genes (left) and Jaccard similarity of marker genes (right) between zebrafish cells and all human leptomeningeal cells from the Level 02 meningioma dataset from (Extended Data Fig. 5e) (d). d’. Comparison of muLEC vs all other cell types. e-e’, Heatmap visualisation of spearman correlation coefficients of scaled gene expression of pairwise marker genes (left) and Jaccard similarity of marker genes (right) between mouse meningeal scavenging cells and all human leptomeningeal cells from the Level 02 meningioma dataset from (Extended Data Fig. 5b) (e). e’. Comparison of BAMs vs all other cell types. f, Scatterplot showing mean scaled gene expression for zebrafish muLEC and mouse BAM. Two-sided spearman correlation, the line represents the fitted regression line and spearman correlation is rho = 0.26, p < 2.2e-16. P value not adjusted. Genes are colour coded based on their expression in BAM, muLEC or shared, as shown. muLECs and BAMs share scavenger receptors but not complement factors (BAM specific) or lgmn and cstla (muLEC specific). p value = 2.537885e-25. g, Density plot showing the dot density of scatterplot in (f). Colour scale represents log(count) density and highlights regions of overplotting. h-h’, Confocal projection of lymphatics, veins and muLECs (Tg(−5.2lyve1b:DsRed)nz101, magenta) and adamts3 expression (Tg(adamts3:gal4FF)mu400;Tg(5xUAS:EGFP)zf82, green) in 7 dpf zebrafish larva and (h’) magnification of adamts3+ expression in a lateral view of the zebrafish larva head from h. Representative of >10 animals imaged. i-i’, Confocal projection of lymphatics, veins and muLECs (Tg(−5.2lyve1b:DsRed)nz101, magenta) and ccbe1 expression (Tg(ccbe1:mCitrine)Hu6741, green) in 7 dpf zebrafish larva (i) and (i’) magnification of the ccbe1+ expression in a lateral view of the zebrafish larva head from i. Representative of >10 animals imaged. j-j’, Confocal projection of lymphatics, veins and muLECs (Tg(−5.2lyve1b:DsRed)nz101, magenta) and vegfc expression (Tg(vegfc:gal4FF)mu402; Tg(5xUAS:EGFP)zf82, green) in 7 dpf zebrafish larva (j) and (j’) magnification of the vegfc+ expression in a lateral view of the zebrafish larva head from j. Representative of >10 animals imaged. k-k’, HCR in situ hybridization staining of adult mouse (Mus musculus) brain sections, showing the expression of ccbe1 (green), vegfc (magenta) and cell nuclei (DAPI, blue) (k) and the expression of adamts3 (green), vegfc (magenta) and cell nuclei (DAPI, blue) (k’). n = 3 sections from n = 3 animals. h-h’, i-i’, j-j’, k-k’, Scale bar 100 μm.
Extended Data Fig. 7 Additional data related to key findings in Fig. 3.
a, Depiction of larval wounding (left). UMAP visualization of all cells captured from control and wounded zebrafish heads (n = 8,879 cells) coloured by cell phenotype (right). AEC, arterial EC; LEC, lymphatic EC; muLEC, mural LEC; VEC, venous EC. b, UMAP visualization of all cells captured (Level 01, SCT_snn_res.0.8, n = 8,879 cells) for scRNA-seq coloured by seurat cluster. AEC, arterial endothelial cell; VEC, venous endothelial cell; LEC, lymphatic EC; muLEC, mural LEC. c, Dot plot showing the gene expression for key marker genes used to annotate clusters at Level 01. Size of the dot plots represents the proportion of expressing cells and colour represents average log-normalised expression. d, Confocal projections of muLECs (Tg(−5.2lyve1b:DsRed)nz101, green) and lysosome marker (TgBAC(lamp2-RFP)pd1044, magenta) in 7dpf zebrafish brains, 24 h post stab wound (white arrows) to the left midbrain hemisphere. Representative of >10 animals imaged. e, Confocal projections of muLECs (Tg(−5.2lyve1b:DsRed)nz101, green), MG/macrophages (Tg(mpeg1.1:gal4FF)gl25; Tg(5xUAS:EGFP)zf82, magenta) and 10 kDa Dextran (blue) in 7dpf zebrafish brains, 24 h post larval wounding to the left midbrain hemisphere. Representative of >10 animals imaged. f, Heatmap visualisation of spearman correlation coefficients of scaled gene expression of all genes for all pairwise comparisons of cell phenotypes present. Colour represents the correlation coefficients. Correlation coefficient that did not pass significance testing are denoted with an x. g, Heatmap of spearman correlation coefficients of scaled gene expression of all genes between muLEC and other phenotypes. Colour represents correlation coefficients. Coefficients that did not pass significance testing denoted with x. muLECs show transcriptional similarity to macrophages. h, Scatterplot showing mean scaled gene expression for muLECs and macrophages. Two-sided spearman correlation, the line represents the fitted regression line, and the spearman correlation is rho = 0.27, p value = 7.168918e-311. P value not adjusted. Genes are colour coded based on their expression in macrophages, muLECs or shared, as shown. i, Density plot showing the dot density of scatterplot in (h). Colour scale represents log(count) density and highlights regions of overplotting. j, Dot plot showing expression of haematopoietic or scavenging receptor genes in muLEC and macrophage clusters. k, UMAP visualisation of mural cells in DanioCell38 (hematopoietic and vasculature subset, n = 3,865 cells) coloured by cell phenotype with detailed endothelial cell annotation. l, Dot plot showing the gene expression for key marker genes used to annotate clusters in DanioCell dataset (k). Size of the dot plots represents the proportion of expressing cells and colour represents average log-normalised expression. No evidence of a population homologous to mammalian BAMs was found. d, e, Scale bar 100 μm. For all dot-plots; size of dots represents proportion of expressing cells and colour represents average log-normalised expression.
Extended Data Fig. 8 Additional data related to key findings in Fig. 3.
a, UMAP visualization of all cells coloured by wounded status from Extended Data Fig. 7a. Blow-up of the muLEC cluster boxed. b, UMAP visualization of all cells coloured by the mean mixing metric score per cell phenotype. c, Boxplot showing log-transformed mixing metric scores for each cell-phenotype. Cell number analysed: n = 1862 (AEC), n = 1088 (LEC), n = 644 (VEC), n = 935 (Glia), n = 1025 (muLEC), n = 598 (Neuron), n = 782 (Epithelial), n = 791 (Fibroblast), n = 322 (Macrophage), n = 224 (Neural_crest), n = 160 (Myeloid_progenitors), n = 228 (Photoreceptor), n = 84 (Thrombocyte), n = 55 (Adipocyte), n = 54 (Oligodendrocytes), n = 27 (Neutrophil), total n = 8879 cells. Bounds of box = 25th - 75th quantile, centre = median, whiskers extend to +/−1.5 times interquartile range. ANOVA for multiple comparisons, two-sided T-test for comparison to muLEC. corrected for multiple testing using holm method. (ANOVA p value = 0), Comparison to muLEC adjusted p-values: 1.1e-36 (Adipocytes), 8.6e-132 (AEC), 1.6e-114 (Epithelial), 2e-122 (Fibroblast), 1.7e-50 (Glia), 1.7e-117 (LEC), 4.8e-49 (Macrophages), 3.8e-86 (Myeloid_progenitors), 3.3e-129 (Neural_crest), 1.6e-62 (Neuron), 1.1e-70 (Oligodendrocyte), 1.3e-138 (Photoreceptor), 4.2e-25 (Thrombocyte), 3.4e-101 (VEC). ****adjusted p = <0.0001. d, Stacked bar plot showing composition of wounded status for each cell phenotype for Level 01 dataset. e, Lollipop plot showing the number of DEG (adjusted p-val<0.05 and |log2FC | > 0.1) between wounded and control cells for each cell phenotype (muLEC, n = 350). Two-sided Wilcox test, adjusted for multiple correction using Bonferroni method. f, Barplot of GO terms upregulated in muLEC vs LEC in wounded sample, coloured by GO category and sorted by number of genes overlapping with the gene set. Top 10 most overlapping terms are shown (full list in Supplementary Data Table 4). muLECs show the most change of any cell type due to increased endocytic and metabolic gene expression. One-sided Fisher’s exact test, adjusted for multiple correction using Benjamini-Hochberg method (employed through enrichR). padj <0.05 & log2FC > 0.5 (top10). g, Network visualisation of gene ontology (GO) terms (biological processes) upregulated in wounded compared to control muLEC. Each node represents a GO term, and edges reflect the pairwise similarity between nodes. Connected nodes annotated and coloured manually to highlight biological themes. h, Dotplot showing expression of select genes driving GO term enrichment, split by wounded status. I, Confocal projection of muLECs (Tg(lyve1b:ERK-KTRClover)uom117, green), macrophages (Tg(mpeg1:Gal4ff)gl25; Tg(14xUAS:Nfsb-mCherry)c264, red) and 10 kDa Dextran Alexa405 (blue) in empty-liposome (top) and clodronate-liposome injected (bottom) 7 dpf zebrafish heads (lateral view). j, Quantification of total number of macrophages and muLECs in empty control-liposome (green) and clodronate-liposome injected (orange) 7 dpf zebrafish (n = 13 control and n = 21 clodronate) from i. * 0.0235, **** <0.0001. k, Confocal projection of muLECs (Tg(−5.2lyve1b:DsRed)nz101, orange) in 7 dpf siblings (left) and osr2 mutants (right) zebrafish brains, 24 h post wounding (dotted white circle) to the left midbrain. l, Quantification of average length of muLECs of siblings (green) and osr2 mutants (blue) (n = 39 sibling and n = 21 mutant embryos) in both uninjured and injured hemispheres (as indicated) from k. **** <0.0001. m, Confocal projections of meningeal blood vessels (Tg(kdrl:EGFP)s843, green) in flt4−/− mutants. muLECs (Tg(−5.2lyve1b:DsRed)nz101, magenta) shown inset. n, Quantification of meningeal vessel density in control, flt4 mutants and ccbe1 morphants from Fig. 3g and m. (n = 7 control, n = 8 flt4 mutant and n = 12 ccbe1 MO brains). o, Variance of vascular density per animal (see methods) in control and flt4um203 mutants (flt4−/−). Two-sided Levene’s test *p = 0.015 for flt4−/− (n = 7 control, n = 8 flt4 mutant brains). p, Quantification of brain width in control, flt4 mutants and ccbe1 morphants from (Fig. 3i) (n = 22 control brains, n = 19 flt4 mutant brains and n = 9 ccbe1 morphants). q, Quantification of brain width in control, flt4 mutants and ccbe1 morphants from (Fig. 3k) (n = 8 control brains, n = 6 flt4 mutant brains and n = 9 ccbe1 morphants). r, Confocal projection of muLECs muLECs (Tg(−5.2lyve1b:DsRed)nz101, magenta) and neutrophils (Tg(lyz:BFP), green) in sibling (top left), flt4um203 mutants (flt4−/−, top right) and control morphants (control MO, bottom right), ccbe1 morphants (ccbe1 MO, bottom left) 7 dpf zebrafish brains. s, Quantification of total number of neutrophils populating the zebrafish leptomeninges in siblings, flt4 mutants and control morphants, ccbe1 morphants from (h) (n = 18 siblings brains, n = 25 flt4 mutant brains and n = 14 control morphants, n = 12 ccbe1 morphants). i, k, m, r, Scale bar 100 μm. For 8c: ANOVA for multiple comparisons, two-sided T-test for comparison to muLEC. corrected for multiple testing using holm method. (ANOVA p value = 0), ****p = <0.0001. One-way ANOVA *p = 0.0235 and ****p = <0.0001 for control versus clodronate-injected in j. One-way Analysis of Variance (ANOVA) ****p = <0.0001 for siblings versus osr2 mutants in l. For 8n, p, q: One way ANOVA; multiple comparison made between means of each condition (mean control vs mean flt4 vs mean ccbe1). For 8 s: unpaired two-tailed t-test (siblings vs flt4) and (control MO vs ccbe1 MO). For all dot-plots; size of dots represents proportion of expressing cells and colour represents average log-normalised expression.
Extended Data Fig. 9 Additional data related to key findings in Fig. 4.
a, HCR in situ hybridization staining of zebrafish (Danio rerio) brain section, showing expression of prox1a (green) and cell nuclei (DAPI, blue). n = 3 sections from n = 3 animals analysed. b, HCR in situ hybridization staining of axolotl (Ambystoma mexicanum) brain section, showing expression of flt4 (magenta) and cell nuclei (DAPI, blue). n = 3 sections from n = 3 animals analysed. c, HCR in situ hybridization staining of mouse (Mus musculus) brain section, showing expression of osr2 (green), flt4 (magenta) and cell nuclei (DAPI, blue). White dotted line marks leptomeningeal border. n = 3 sections from n = 3 animals analysed. d, HCR in situ hybridization staining of axolotl (Ambystoma mexicanum) whole spinal cord section, showing expression of csf1r (red) and cell nuclei (DAPI, blue). n = 3 sections from n = 3 animals analysed. e, HCR in situ hybridization staining of axolotl (Ambystoma mexicanum) whole brain section, showing expression of csf1r (magenta) and cell nuclei (DAPI, blue). n = 3 sections from n = 3 animals analysed. f, HCR in situ hybridization staining of mouse (Mus musculus) brain section, showing expression of prox1 (green), flt4 (magenta) and cell nuclei (DAPI, blue). n = 3 sections from n = 3 animals analysed. g, HCR in situ hybridization staining of adult fat-tailed dunnart (Sminthopsis crassicaudata) intestine with osr2 (green) and nuclei (DAPI, blue). This confirmed that the fat-tailed dunnart osr2 HCR probe works, despite not staining cells in the meninges. Representative of n = 3 biological replicates, and at least 3 technical replicates were imaged for each shown image. h, HCR in situ hybridization staining of E12.5 mouse embryo (Mus musculus) forelimb with osr2 (green) and nuclei (DAPI, blue). This confirmed that the mouse osr2 HCR probe works, despite not staining cells in the meninges Representative of n = 3 biological replicates, and at least 3 technical replicates were imaged for each shown image. i, Schematic depiction of the cellular composition of mammalian (left) and non-mammalian vertebrate (based on zebrafish, right) meninges. a-h, Scale bar 100 μm.
Supplementary information
Supplementary Table 1 (download XLSX )
Detailed information on how the datasets were preprocessed.
Supplementary Table 2 (download XLSX )
Detailed information on analysis performed on the zebrafish lymphatic time-course dataset.
Supplementary Table 3 (download XLSX )
Detailed information on analysis performed on the zebrafish osr2 dataset.
Supplementary Table 4 (download XLSX )
Detailed information on the cross-species analysis (all species).
Supplementary Table 5 (download XLSX )
Detailed information on analysis performed on the zebrafish wounded dataset.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gaudi, A.U., Meier, M., Baltaci, O.F. et al. Convergent evolution of scavenger cell development at brain borders. Nature 651, 743–751 (2026). https://doi.org/10.1038/s41586-025-10003-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-10003-3