Abstract
Artificial selection has greatly shaped crop agronomic traits1,2,3; however, the mechanistic basis of how immunity is selected remains unclear. Here we identify the Oryza sativa nucleotide-binding site and leucine-rich repeat (NLR) receptor XA48 and downstream transcription factors OsVOZ1 and OsVOZ2 (OsVOZ1/2), which confer resistance to bacterial blight. XA48 perceives the ancient pathogen effector XopG, activating effector-triggered immunity by degrading the negative regulator OsVOZ1/2. The XA48–OsVOZ1 module has undergone subspecies-specific selection: Xa48 is retained only in Oryza sativa indica and was lost in Oryza sativa japonica. By contrast, OsVOZ1 has diverged into two haplotypes—O. s. indica retains both OsVOZ1A/S alleles compatible with Xa48, whereas O. s. japonica has only OsVOZ1A. Reintroducing Xa48 into O. s. japonica severely compromises yield owing to the XA48–OsVOZ1A-mediated immune incompatibility. Stacking XA48-mediated effector-triggered immunity with XA21-mediated pattern-triggered immunity reconstitutes the broad-spectrum resistance from wild rice. Our study therefore reveals how asymmetric selection of an NLR–transcription factor module shapes disease resistance and reproductive development, providing a strategy for breeding crops by harnessing the relative immunity of wild rice.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 52 print issues and online access
$199.00 per year
only $3.83 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are available within this Article and its Supplementary Information. Original gel blots are shown in Supplementary Fig. 1. The full genomic sequence of Xa48 can be found in GenBank under accession numbers OR712925 and OR712926. The RNA-seq data generated in this study have been deposited in the SRA database under accession number PRJNA1028151. Gene sequences of the 3K rice germplasm were retrieved from the Rice Functional Genomics and Breeding (RFGB) database (https://www.rmbreeding.cn/public/searchbak), and the accession codes (BioProject) of the rice accessions used for allele frequency analysis are provided in Supplementary Table 4. Source data are provided with this paper.
References
Doebley, J. F., Gaut, B. S. & Smith, B. D. The molecular genetics of crop domestication. Cell 127, 1309–1321 (2006).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Barabaschi, D., Tondelli, A., Vale, G. & Cattivelli, L. Fitness cost shapes differential evolutionary dynamics of disease resistance genes in cultivated and wild plants. Mol. Plant 13, 1352–1354 (2020).
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Couto, D. & Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol. 16, 537–552 (2016).
Xin, X. F., Kvitko, B. & He, S. Y. Pseudomonas syringae: what it takes to be a pathogen. Nat. Rev. Microbiol. 16, 316–328 (2018).
Delaux, P. M. & Schornack, S. Plant evolution driven by interactions with symbiotic and pathogenic microbes. Science 371, eaba6605 (2021).
Han, X. & Tsuda, K. Evolutionary footprint of plant immunity. Curr. Opin. Plant Biol. 67, 102209 (2022).
Nino-Liu, D. O., Ronald, P. C. & Bogdanove, A. J. Xanthomonas oryzae pathovars: model pathogens of a model crop. Mol. Plant Pathol. 7, 303–324 (2006).
Lin, H. et al. An MKP-MAPK protein phosphorylation cascade controls vascular immunity in plants. Sci. Adv. 8, eabg8723 (2022).
Song, W. Y. et al. A receptor kinase-like protein encoded by the rice disease resistance gene Xa21. Science 270, 1804–1806 (1995).
Pruitt, R. N. et al. The rice immune receptor XA21 recognizes a tyrosine-sulfated protein from a Gram-negative bacterium. Sci. Adv. 1, e1500245 (2015).
Chen, F., Yan, B., Gong, X., Li, H. & He, Z. Genome sequencing of the bacterial blight pathogen DY89031 reveals its diverse virulence and origins of Xanthomonas oryzae pv. oryzae strains. Sci. China Life Sci. 64, 2175–2185 (2021).
Ercoli, M. F. et al. Plant immunity: rice XA21-mediated resistance to bacterial infection. Proc. Natl Acad. Sci. USA 119, e2121568119 (2022).
Yang, Y. et al. Research progress on cloning and function of Xa genes against rice bacterial blight. Front. Plant Sci. 13, 847199 (2022).
Vergish, S. et al. Rhomboid-mediated cleavage of the immune receptor XA21 protects grain set and male fertility in rice. Proc. Natl Acad. Sci. USA 122, e2502025122 (2025).
Chen, E., Huang, X., Tian, Z., Wing, R. A. & Han, B. The genomics of Oryza species provides insights into rice domestication and heterosis. Annu. Rev. Plant Biol. 70, 639–665 (2019).
Iyer, A. S. & McCouch, S. R. The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol. Plant Microbe Interact. 17, 1348–1354 (2004).
Lu, Y. et al. A new NLR disease resistance gene Xa47 confers durable and broad-spectrum resistance to bacterial blight in rice. Front. Plant Sci. 13, 1037901 (2022).
Yoshimura, S. et al. Expression of Xa1, a bacterial blight-resistance gene in rice, is induced by bacterial inoculation. Proc. Natl Acad. Sci. USA 95, 1663–1668 (1998).
Wu, Q. et al. Salicylic acid acts upstream of auxin and nitric oxide (NO) in cell wall phosphorus remobilization in phosphorus deficient rice. Rice 15, 42 (2022).
Duan, L., Liu, H., Li, X., Xiao, J. & Wang, S. Multiple phytohormones and phytoalexins are involved in disease resistance to Magnaporthe oryzae invaded from roots in rice. Physiol. Plant 152, 486–500 (2014).
He, J. et al. An R2R3 MYB transcription factor confers brown planthopper resistance by regulating the phenylalanine ammonia-lyase pathway in rice. Proc. Natl Acad. Sci. USA 117, 271–277 (2020).
Sun, L. et al. Functions of rice NAC transcriptional factors, ONAC122 and ONAC131, in defense responses against Magnaporthe grisea. Plant Mol. Biol. 81, 41–56 (2013).
Swaminathan, S. et al. CYP76M7 is an ent-cassadiene C11α-hydroxylase defining a second multifunctional diterpenoid biosynthetic gene cluster in rice. Plant Cell 21, 3315–3325 (2009).
Erb, M. & Reymond, P. Molecular interactions between plants and insect herbivores. Annu. Rev. Plant Biol. 70, 527–557 (2019).
Yang, D. L., Yang, Y. & He, Z. Roles of plant hormones and their interplay in rice immunity. Mol. Plant 6, 675–685 (2013).
Bi, G. et al. The ZAR1 resistosome is a calcium-permeable channel triggering plant immune signaling. Cell 184, 3528–3541.e12 (2021).
Förderer, A. et al. A wheat resistosome defines common principles of immune receptor channels. Nature 610, 532–539 (2022).
Roberts, M., Tang, S., Stallmann, A., Dangl, J. L. & Bonardi, V. Genetic requirements for signaling from an autoactive plant NB-LRR intracellular innate immune receptor. PLoS Genet. 9, e1003465 (2013).
Gao, Q. et al. A receptor-channel trio conducts Ca2+ signalling for pollen tube reception. Nature 607, 534–539 (2022).
DeFalco, T. A. et al. Using GCaMP3 to study Ca2+ signaling in Nicotiana species. Plant Cell Physiol. 58, 1173–1184 (2017).
Baruch, K. et al. Metalloprotease type III effectors that specifically cleave JNK and NF-κB. EMBO J. 30, 221–231 (2011).
Zhai, K. et al. RRM transcription factors interact with NLRs and regulate broad-spectrum blast resistance in rice. Mol. Cell 74, 996–1009 (2019).
Cheong, H. et al. Xanthomonas oryzae pv. oryzae type III effector XopN targets OsVOZ2 and a putative thiamine synthase as a virulence factor in rice. PLoS ONE 8, e73346 (2013).
Wang, J. et al. Two VOZ transcription factors link an E3 ligase and an NLR immune receptor to modulate immunity in rice. Mol. Plant 14, 253–266 (2021).
Wen, Y. et al. VOZ1 and VOZ2 transcription factors regulate arsenic tolerance and distribution in rice and Arabidopsis. Front Plant Sci. 14, 1209860 (2023).
Mitsuda, N. et al. VOZ; isolation and characterization of novel vascular plant transcription factors with a one-zinc finger from Arabidopsis thaliana. Plant Cell Physiol. 45, 845–854 (2004).
Yang, D. L. et al. Plant hormone jasmonate prioritizes defense overgrowth by interfering with gibberellin signaling cascade. Proc. Natl Acad. Sci. USA 109, E1192–E1200 (2012).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Xie, W. et al. Breeding signatures of rice improvement revealed by a genomic variation map from a large germplasm collection. Proc. Natl Acad. Sci. USA 112, E5411–5419 (2015).
Li, X. et al. Analysis of genetic architecture and favorable allele usage of agronomic traits in a large collection of Chinese rice accessions. Sci. China Life Sci. 63, 1688–1702 (2020).
Wei, X. et al. A quantitative genomics map of rice provides genetic insights and guides breeding. Nat. Genet. 53, 243–253 (2021).
Deng, Y. et al. Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science 355, 962–965 (2017).
He, Z., Webster, S. & He, S. Y. Growth-defense trade-offs in plants. Curr. Biol. 32, R634–R639 (2022).
Ma, Y. et al. COLD1 confers chilling tolerance in rice. Cell 160, 1209–1221 (2015).
Xiao, N. et al. Identification of genes related to cold tolerance and a functional allele that confers cold tolerance. Plant Physiol. 177, 1108–1123 (2018).
Liu, C. et al. Early selection of bZIP73 facilitated adaptation of japonica rice to cold climates. Nat. Commun. 9, 3302 (2018).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Ngou, B. P. M., Ahn, H. K., Ding, P. & Jones, J. D. G. Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 592, 110–115 (2021).
Pruitt, R. N. et al. The EDS1-PAD4-ADR1 node mediates Arabidopsis pattern-triggered immunity. Nature 598, 495–499 (2021).
Tian, H. et al. Activation of TIR signalling boosts pattern-triggered immunity. Nature 598, 500–503 (2021).
Yuan, M. et al. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 592, 105–109 (2021).
Wang, C. et al. XA23 is an executor R protein and confers broad-spectrum disease resistance in rice. Mol. Plant 8, 290–302 (2015).
Xu, Z. et al. A varied AvrXa23-like TALE enables the bacterial blight pathogen to avoid being trapped by Xa23 resistance gene in rice. J. Adv. Res. 42, 263–272 (2022).
Bomblies, K. & Weigel, D. Hybrid necrosis: autoimmunity as a potential gene-flow barrier in plant species. Nat. Rev. Genet. 8, 382–393 (2007).
Calvo-Baltanás, V., Wang, J. & Chae, E. Hybrid incompatibility of the plant immune system: an opposite force to heterosis equilibrating hybrid performances. Front. Plant Sci. 11, 576796 (2021).
Chen, C. et al. A two-locus interaction causes interspecific hybrid weakness in rice. Nat. Commun. 5, 3357 (2014).
Li, L. & Weigel, D. One hundred years of hybrid necrosis: hybrid autoimmunity as a window into the mechanisms and evolution of plant-pathogen interactions. Annu. Rev. Phytopathol. 59, 213–237 (2021).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824 (2012).
Li, M. X., Yeung, J. M., Cherny, S. S. & Sham, P. C. Evaluating the effective numbers of independent tests and significant p-value thresholds in commercial genotyping arrays and public imputation reference datasets. Hum. Genet. 131, 747–756 (2012).
Yin, L. et al. rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study. Genom. Proteom. Bioinform. 19, 619–628 (2021).
McKenna, A. et al. The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Wang, H. Z. et al. Experimental assessment of the yield gap associated with maize production in the North China Plain. Field Crops Res. 295, 108897 (2023).
Acknowledgements
We thank Q. Zhang for discussion, P. Ronald for providing the Xa21 transgenic seeds, X. Wang for rice transformation, Z. Lei for field inoculation, Q. Li and Y. Gao for hormone analysis, G. Chen for EMSA assay, J. Li for tissue sectioning, D. Zhu and W. Cai for microscopy observation, Y. Xu for statistical analysis and J. Shou for helping with X. o. pv. oryzae nursery. This work was supported by Biological Breeding-National Science and Technology Major Projects (2023ZD04070 to Y.D.), the National Natural Science Foundation of China (32088102 to Z.H.; 32361143515, 31830072 to G. Chen; 32402392 to H.L.), the Chinese Academy of Sciences (DXB1490000 to H.L.), the National Key Research and Development Program of China (2024YFD1200600 to H.L.), Shanghai Science and Technology Development Funds (24YF2751900 to H.L.) and Shanghai Agricultural Science and Technology Innovation Program (K2025005 to Z.H.; K2024-02-08-00-12-F00050 to H.L.).
Author information
Authors and Affiliations
Contributions
H.L., F.C., Y.D., G. Chen and Z.H. conceived and designed the experiments. H.L., F.C., G. Cheng, B.Y., M.Y. and Y.C. performed experiments. Y.W. and G. Cheng performed Tn5 mutagenesis. J.Q., X.H. and B.H. performed population genomics and artificial selection analysis. Y.L., Y.C., K.C., X.G., S.L., J.L. and J.-S.J. assisted with pathogen inoculation and field trials. B.M. assisted in rice transformation experiments. J.W., R.L. and J.X. grew and provided wild rice and rice germplasm. D.-L.Y. assisted with hormone analysis. H.X. and M.S. assisted with statistical analysis. Q.G. and B.L. assisted with cation-permeable channel assays. Z.H., Y.D. and G. Chen supervised the project. Z.H. and Y.D. provided theoretical contributions to the project. H.L., F.C., Y.D., G. Chen and Z.H. analysed the data and wrote the paper. All of the authors discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Yoji Kawano and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Map-based cloning identifies Xa48.
a, SKZ displayed strong resistance to J18 but high susceptible to PXO99A. Disease symptoms of SKZ, NIPB, TP309, and 106 at 14 dpi with Xoo strains J18 and PXO99A (n = 36). Line 106 is transgenic TP309 expressing Xa21. b, Xa48 was initially mapped to chromosome 3 between two InDel markers InDel26 and InDel47. c, d, Independent CRISPR/Cas9 knockout lines (CR-C1, CR-C2, CR-C3, CR-C4) of SKZ were generated (c), and representative lines’ lesion length of knockout mutants were measured (d). Note that only CR-C1 became susceptible to J18. e, Premature termination of XA48 translation in susceptible varieties caused by a single-base deletion and SNP. XA48 encodes a typical CNL receptor consisting of 1,094 amino acids, with N-terminal coiled-coil (CC) domain (1-195), NB-ARC domain (215-440), and C-terminal leucine-rich repeat (LRR) domain (555-1012). f, Transcriptional level of XA48 was detected in TP309-Xa48 and NIPB-Xa48 complement plants (two-tailed t test, n = 3). g, h, NIPB-Xa48 transgenic plants are highly resistant to J18 as SKZ. The lesion length (g) and bacterial populations (h) were measured. Data are shown as boxplots displaying the distribution of individual values (dots). Error bars represent the estimated marginal means (EMM) ± SE from linear mixed-effects models (a, d, g) or arithmetic means ± SD (f, h). For a, d, g, EMMs derived from linear mixed-effects models fitted to data from two or three independent experiments are compared between each genotype and the wild-type control using two-tailed planned contrasts. Scale bars, 5 cm.
Extended Data Fig. 2 Xa48 confers lifetime Xoo resistance.
a-d, Xa48 confers Xoo resistance in SKZ (indica) (a, b) and NIPB-Xa48 (japonica) (c, d) seedlings. Lesion lengths and bacterial populations were measured in seedlings inoculated with J18. e, GUS staining of pXa48::GUS reporter transgenic plants revealed Xa48 expression in leaf, stem, root, seed, and young panicle. The boxed sections were magnified for close views of GUS staining in the spikelet and anther. Scale bars, 1 cm. f, Expression of Xa48 in different tissues was detected by qRT-PCR and normalized to expression in the root 1. Root1 was from 5-day-old seedling, leaf1, stem1 were from 25-day-old seedling, and root2, leaf2, stem2 were from 80-day-old plants. g, Xa48 was not induced by Xoo as revealed by qRT-PCR. Four-week-old seedling SKZ were inoculated with J18 and PXO99A, and leaf samples were collected during a time course of 0-72 hpi for RNA preparation. Mock wase used as a negative control (two-tailed t test, n = 3). h, i, Hierarchical clustering (h) and Gene Ontology (GO) (i) analysis of differentially expressed genes in TP309 and TP309-Xa48 leaves inoculated with J18. Note that Xa48 enhanced gene induction in salicylic acid, ethylene, and diterpene phytoalexin catabolic processes, which play important roles in rice immunity. j, qRT-PCR revealed that genes related to the salicylic acid catabolic process (OsPAL3, OsPAL4, OsPAL6), ethylene response (ONAC122, ONAC131), and diterpene phytoalexin biosynthesis (CYP76M7) were up-regulated in TP309-Xa48 inoculated with J18 (two-tailed t test, n = 3). k-m, No significant differences in salicylic acid (SA) (k), jasmonic acid (JA) (l), and ethylene (ET) (l) content were observed between TP309 and TP309-Xa48 plants inoculated with J18 (two-tailed t test, n = 3). Error bars represent the estimated marginal means (EMM) ± SE from linear mixed-effects models (a, c) or arithmetic means ± SD (b, d, g, j-m). For a, c, EMMs derived from linear mixed-effects models fitted to data from two or three independent experiments are compared between each genotype and the wild-type control using two-tailed planned contrasts.
Extended Data Fig. 3 Identification and functional analysis of XopG through Tn5 mutagenesis.
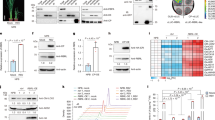
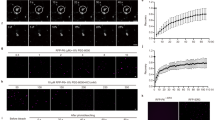
a, b, Generation of avrXa48/xopG insertion mutants. Tn5 transposon-insertion shows two independent insert mutants (Tn5#1 and Tn5#2) in the same site (a), which cause loss-of-function of XopG. Southern blot confirms single insertion of Tn5 in the two LN2 mutants (b). The non-TAL promoter of XopG is indicated with the featured sequence. c, Tn5 transposon-inserted mutants of strain LN2 lost avirulence to NIPB-Xa48 (n = 66). The lesion length was measured at 14 dpi. d, The introduction of XopGLN2 changed PXO99A into avirulence to NIPB-Xa48 (n = 75). e, XopG directly interacts with the XA48 CC domain by SLC in N. benthamiana. XA48-NBS and XA48-LRR domains, which do not interact with XopG, and XopV were used as negative controls. f, XopG associates with the XA48-CC mainly in the cell periphery and nucleus as revealed by BiFC in rice protoplasts. XA48-CC, XA48-NBS, XA48-LRR and XopG were fused to the N-terminal fragment of YFP (YN) and the C-terminal fragment of YFP (YC). YN and YC protein were detected by immunoblotting with anti-Myc and anti-HA antibodies, respectively. g, Representative localization images of XopG-GFP transgenic rice, showing main localization to the cell periphery and nucleus. Co-localization of XA48 with XopG was detected transiently expressed in rice protoplasts. FM4-64 and DAPI staining indicate the PM and nucleus. h-i, Reduction of relative luciferase activity upon co-expression of XA48-XopG in rice protoplasts, as readout for cell death. Relative luciferase activity was measured at 16 h post transfection. D519V served as a positive control (mean ± SD, two-tailed t test and Bonferroni correction). j-l, XA48-XopG is a non-selective cation channel. Typical whole-cell recordings and average current-voltage curves for K+ (j), Mg2+ (k), Na+ (l) conductance by XA48-XopG in HEK293T cells (two-tailed t test and Bonferroni correction, n = 8 cells). m, Schematic of the domain structure of XopG is indicated. Peptidase_M91 contains an HEXXH motif, characteristic of zinc metallopeptidases. n, o, XopG exhibits peptidase activity of not only self-cleavage but also cleaving β-casein, a commonly used substrate for assessing general protease activity (n). However, we did not observe that XopG cleavage XA48 (o). MBP acts as a control. XA48-FLAG with XopG-GFP detected by immunoblotting with anti-FLAG/GFP antibodies. Error bars represent the estimated marginal means (EMM) ± SE from linear mixed-effects models (c, d). For c, d, EMMs derived from linear mixed-effects models fitted to data from two or three independent experiments are compared between each genotype and the wild-type control using two-tailed planned contrasts. Scale bars, 5 cm. Experiments were independently repeated two times with similar results (e, f, n, o).
Extended Data Fig. 4 Functionality and specificity analysis suggest XopG is a conserved effector in bacterial pathogens.
a, A phylogenetic tree and protein alignment of XopG and homologous proteins from different plant and animal bacterial pathogens were constructed using the Neighbour-Joining method, with 1,000 bootstrap replicates for statistical support, in MEGA11 software. Bootstrap values greater than 70% are indicated on the tree. XopG-like proteins are grouped into three clades. Note that the functional XopG LN2 belongs to Class I, labelled with green (left). The conserved HEXXH motif is indicated with red box. Amino acids in red (XopGS155 and XopGA157) are conserved in Clade I and Clade II, but changed in Clade III (right). b, Disease resistance phenotypes of NIPB-Xa48 inoculated with PXO99A/XopGLN2, PXO99A/XopGPst DC3000, PXO99A/XopGPsa M228, PXO99A/XopGXtt KM9, PXO99A/XopGSea Serova, PXO99A/XopGE. coli O157, which contain three clades of XopG-like proteins. LN2 and PXO99A were used as avirulent and virulent control strain, respectively. c, SLC detected interactions between XA48-CC and XopG, XopG-like proteins from Pst DC3000, Psa M228, Xtt KM9, Sea Serova, E. coli O157. Notably, only XopG and XopG-like proteins from Clade I and Clade II, but not Clade III, interact with XA48-CC, consistent with their avirulence to NIPB-Xa48. Protein expression was detected by Western blot. d, e, Mutants of XopGS155A and XopGA157D lost interaction with XA48-CC, as detected by Co-IP (d) And SLC (e). Protein expression in N. benthamiana was detected by Western blot. f, Disease resistance phenotypes of NIPB-Xa48 inoculated with PXO99A expressing XopGS155A and XopGA157D (n = 60). Note that the S155 and A157 of XopG from Clade I and II were critical to its functional in XA48-mediated resistance. Error bars represent the estimated marginal means (EMM) ± SE from linear mixed-effects models (b, f). For b, f, EMMs derived from linear mixed-effects models fitted to data from two or three independent experiments are compared between each genotype and the wild-type control using two-tailed planned contrasts. Scale bars, 5 cm. Experiments were independently repeated twice with similar results (c-e).
Extended Data Fig. 5 HEXXH domain of XopG contributes to trigger XA48-mediated resistance in rice.
a, Decreased basal Xoo resistance of xopG-OE in NIPB. Leaves were inoculated with PXO99A. The lesion length was measured at 14 dpi. b, Disease resistance phenotype and lesion length of NIPB-Xa48 inoculated with PXO99A/XopGLN2H142R, PXO99A/XopGLN2E143R, and PXO99A/XopGLN2H146R. LN2 and PXO99A-XopGLN2 were used as avirulent controls. c, d, XopG mutants (H142R, E143R, H146R) interact with XA48-CC detected by Co-IP (c) and SLC (d). Western blotting was performed to detect protein levels. e, XopG mutants (H142R, E143R, H146R) could not trigger XA48-mediaed cell death in N. benthamiana leaves. Protein expression was detected by Western blot. Numbers in parentheses indicate leaves exhibited cell-death symptoms. f, The XopG mutants (H142R, E143R, H146R) promoted degradation in a cell-free assay. Cell extracts from transgenic plant expressing XopG-GFP, XopGH142R-GFP, XopGE143R-GFP, and XopGH146R-GFP were incubated and collected over a time course (0-2 h). Protein levels were analysed by Western blot using an anti-GFP antibody. g, h, Avirulence and virulence detection of Type I and Type V of XopG variant strains on TP309-Xa48 (g) and NIPB-Xa48 (h). Error bars represent the estimated marginal means (EMM) ± SE from linear mixed-effects models (a, b, g, h). For a, b, g, h, EMMs derived from linear mixed-effects models fitted to data from two or three independent experiments are compared between each genotype and the wild-type control using two-tailed planned contrasts. Scale bars, 5 cm. Experiments were independently repeated twice with similar results (c-f).
Extended Data Fig. 6 Identification of XA48-OsVOZ1/2 interaction.
a, The candidate XA48-interacting proteins revealed by Y2H screen. Note that OsVOZ1 is the most hit protein. b, Western blots detected expression of XA48-CC-nLuc, cLuc-OsVOZ1 and cLuc-OsVOZ2 in N. benthamiana for protein interaction. c, Representative images of XA48 subcellular localization. XA48-YFP was transiently expressed in rice protoplasts, which showed main localization to the cell periphery and nucleus. OsRAC1-mCherry and NLS-mCherry served as a plasma membrane (PM) and nuclear marker, respectively. Scale bars, 10 µm. d, Representative images showing the subcellular localization of OsVOZ1 and OsVOZ2 at the cell periphery and nucleus. OsVOZ1-YFP and OsVOZ2-YFP were transiently expressed in rice protoplasts. Scale bars, 10 µm. e, f, Co-localization of XA48 with OsVOZ1 and OsVOZ2 was detected in cell periphery and nucleus (e), and the presence of XopG did not alter the co-localization pattern of XA48 with OsVOZ1/OsVOZ2 (f). OsVOZ1-mCherry, OsVOZ2-mCherry, Xa48-CFP and XopG-GFP were transiently expressed in rice protoplasts. Scale bars, 10 µm. g-h, qRT-PCR and immunodetection analyses were performed on OsVOZ1-GFP and OsVOZ2-GFP transgenic overexpression (OE) lines, using SKZ and NIPB-Xa48 as wild type controls. Actin was detected as a loading control (mean ± SD, two-tailed t test; n = 3). i, Schematic of OsVOZ1/2 knockout (KO) lines in NIPB background. j, The degradation of OsVOZ1 and OsVOZ2 by XopG and XA48-CC in vitro was inhibited by the zinc metalloprotease-specific inhibitor phenanthroline. Purified OsVOZ1-MBP, OsVOZ2-MBP, XopG-MBP and XA48-CC-His were incubated in reaction mixture with or without 5 mM phenanthroline. Protein levels were analysed by Coomassie blue staining and Western blotting. k, XA48-CC promoted the interaction between XopG and OsVOZs. Notably, in vitro pull-down assays showed that OsVOZ1-MBP and OsVOZ2-MBP pulled down XopG-His in the presence of XA48-CC-GST. l, Resin sections of leaves from NIPB, CR-Osvoz1, and CR-Osvoz2 mutants. Microscopic observation revealed that the CR-Osvoz1 and CR-Osvoz2 mutants developed abnormal vascular bundles, including reduced xylem and phloem size, impaired xylem cell differentiation, and irregular vessel morphology. Xylem (xy), phloem (ph), and vessels (ve) are highlighted with in red, yellow, and black lines, respectively. m, The areas of xylem and phloem were measured, revealing that their sizes in the CR-Osvoz1 and CR-Osvoz2 mutants were smaller than those in NIPB (mean ± SD, two-tailed t test; n = 8). Experiments were independently repeated twice with similar results (b-f, j, k).
Extended Data Fig. 7 OsVOZ1 and OsVOZ2 specifically bind to the promoter of OsJAZ family, which contain the conserved CCCAC motif.
a, Gene Ontology (GO) analysis of differentially expressed genes in CR-Osvoz1 and NIPB. Note that loss of OsVOZ1 function activated genes involved in JA-related defence. b, EMSA was performed to investigate binding affinity of OsVOZ1 and OsVOZ2 to the cis-elements in OsJAZs promoter. Biotin-labelled probes were incubated with MBP or OsVOZs-MBP. Unlabelled competitor fragments were added to evaluate binding specificity. Experiments were independently repeated twice with similar results.
Extended Data Fig. 8 Xa48 decreases grain yield in japonica but not in indica.
a, Lesion length cause by J18 on Xa48 complement lines in indica Kasalath that contains a truncated Xa48 mutant (shown above) and harbours the OsVOZ1A allele (n = 62). Note that wild type Kasalath is also resistant to J18, mediated by an additional unrecognized Xa genes. Scale bars, 5 cm. b-d, Mature plants (b), plant height (c) and panicles (d) of NIPB-Xa48, TP309-Xa48, Kasalath-Xa48, and CR-Xa48/SKZ. Note that panicle size was not affected in these plants with or without Xa48, whereas Xa48 led to a decrease in plant height in both japonica varieties NIPB and TP309, but not in indica rice SKZ and Kasalath (mean ± SD, two-tailed t test, n = 16). Scale bars, 10 cm (b) and 5 cm (d). e, f, The introduction of Xa48 significantly decreased grain yield in japonica rice NIPB by reducing seed setting (e), and knockout of Xa48 (CR-Xa48) did not affect grain productivity in indica rice SKZ (f) (n = 540), with individual data points overlaid. Trials are arranged by location (Shanghai, Hainan) and year (2021-2023). g-i, KEGG analysis of differentially expressed genes in TP309-Xa48 vs TP309 young panicle. Note that the Xa48 introduction induced differential expression of many genes including those involved in sugar and amino acid metabolism, which may contribute to the seed development penalty in japonica. Xa48 does not cause much difference of gene expression in indica CR-Xa48/SKZ vs SKZ. (mean ± SD, two-tailed t test, n = 3). Error bars represent estimated marginal means (EMM) ± SE from linear mixed-effects models (a, e, f). For a, e, f, EMMs derived from linear mixed-effects models fitted to data from two or three independent experiments are compared between each genotype and the wild-type control using two-tailed planned contrasts.
Extended Data Fig. 9 XA48-OsVOZ1A/S immune modules shape different seed setting between japonica and indica.
a, Frequency changes of the Xa48-OsVOZ1 alleles combination during indica and japonica domestication and improvement. Note that the proportion of materials harbouring the Xa48-OsVOZ1S allele combination rises from 5.1% to 20.8%. b, c, OsVOZ1S interacts with XA48, determined by SLC (b) and Y2H (c) assays. d, The allelic variation of Xa48 and OsVOZ1 in different varieties was shown in the indica (i) and japonica (j) varieties sued in the study. e, Development of indica inbreed lines with four combinations of OsVOZ1A/S and Xa48, OsVOZ1S-Xa48−, OsVOZ1S-Xa48+, OsVOZ1A-Xa48− and OsVOZ1A-Xa48+, derived from SKZ (OsVOZ1S) crossing to Kasalath (OsVOZ1A) (F5), which showed no difference in seed setting rate (n = 90). f, Premature mutation of Xa48 in indica TN1 and 9311 at codon position 226. SKZ crossing to 9311 and TN1, which brings the same OsVOZ1S allele, to generate inbreed lines OsVOZ1S-Xa48− and OsVOZ1S-Xa48+ (F5) respectively (n = 90), which showed the same seed setting rate. g-j, Lesion lengths cause by J18 and seed setting rates of complement line TN1-Xa48 (n = 65, 90) (g, h) and ZS97-Xa48 (n = 66, 90) (i, j) (indica), harbouring OsVOZ1S and OsVOZ1A, respectively. The results indicated that the complement indica lines exhibited J18-resisatnce and no difference on seed setting rates. Scale bars, 5 cm. k, Performance of three-leaf stage seedlings of NIPB, CR-Osvoz1, CR-Osvoz2, OsVOZ1S-OE/NIPB, and OsVOZ1A-OE/NIPB was assessed after cold treatment at 4°C for 4 days, followed by a 5-day recovery period. The survival rates (the percentage of recovered seedlings) were then determined. Note that OsVOZ1A-OE/NIPB exhibited greater chilling tolerance than OsVOZ1S-OE/NIPB (mean ± SD, two-tailed t test, n = 8). l, BN-PAGE assay revealed that the oligomerization of OsVOZ1A was stronger than that of OsVOZ1S, which may contribute to chilling tolerance phenotype in japonica. Recombinant OsVOZ1A-His, OsVOZ1S-His and OsVOZ2-His was analysed by immunoblotting using anti-His antibody via SDS-PAGE. Error bars represent estimated marginal means (EMM) ± SE from linear mixed-effects models (e-j). For e-j, EMMs derived from linear mixed-effects models fitted to data from two or three independent experiments are compared between each genotype and the wild-type control using two-tailed planned contrasts.
Extended Data Fig. 10 Xa48-Xa21 stacking rice for broad-spectrum Xoo resistance was developed.
a, Development of transgenic TP309 (japonica) with integrating Xa21Xa48 through crossing and selfing selection (F5 generation). b, Broad-spectrum disease resistance of Xa21Xa48 plants against northeast Asian Xoo strain J18, LN1, LN2, LN3 and JL1. Scale bars, 5 cm. c, The defence genes PR4, PR5, and PR10 expression were significantly enhanced in Xa21Xa48 plants during Xoo infection (mean ± SD, two-tailed t test; n = 3). d, Development of indica rice combining endogenous Xa48 and Xa21 through crossing SKZ and BG139 (Xa21) and selfing (F4 generation). e, De novo development of japonica regaining Xa48 and OsVOZ1S by crossing CR-Osvoz1/NIPB and NIPB-Xa48. f, Mature plant and panicle of CR-Osvoz1, which showed defective growth and development phenotypes. Scale bars, 10 cm.
Supplementary information
Supplementary Fig. 1 (download PDF )
Uncropped blots and gel images.
Supplementary Table 1 (download XLSX )
Amino acid sequence of XA48.
Supplementary Table 2 (download XLSX )
XopG homologues: Xanthomonas stains.
Supplementary Table 3 (download XLSX )
Xa48 loci in 3K rice germplasm.
Supplementary Table 4 (download XLSX )
Rice accessions for allele frequency analysis.
Supplementary Table 5 (download XLSX )
Xa48 moderate high variant AltAlleleFreq.
Supplementary Table 6 (download XLSX )
OsVOZ1 moderate high variant AltAlleleFreq and paired XA48–OsVOZ1 ratio.
Supplementary Table 7 (download XLSX )
The collected strains.
Supplementary Table 8 (download XLSX )
Primer sequences.
Supplementary Table 9 (download XLSX )
Estimate, s.e. and P values.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, H., Chen, F., Cheng, G. et al. Asymmetric selection of a rice immune module and rebuild of disease resistance. Nature 653, 840–849 (2026). https://doi.org/10.1038/s41586-026-10361-6
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-026-10361-6