Abstract
Neuronal axons have traditionally been considered to be the primary mediators of functional connectivity among brain regions. However, the role of astrocyte-mediated communication has been largely underappreciated. Astrocytes communicate with one another through gap junctions, but the extent and specificity of this communication remain poorly understood. Astrocyte gap junctions are necessary for memory formation1,2, synaptic plasticity3,4,5, coordination of neuronal signalling6, and closing the visual and motor critical periods7,8. These findings indicate that this form of communication is essential for proper central nervous system development and function. Despite the importance of astrocyte gap junctional networks, studying them has been challenging. Current methods such as slice electrophysiology disrupt network connectivity and introduce artefacts due to tissue damage. Here, we developed a vector-based approach that labels molecules as they are fluxed by astrocyte gap junctions in awake, behaving animals to overcome these limitations. We then used whole-brain tissue clearing9,10 to image these intact, three-dimensional astrocyte networks. We show that multiple astrocyte networks traverse the mouse brain. These networks selectively connect specific regions, rather than diffusing indiscriminately, and vary in size and organization. We observe local networks that are confined to single brain regions and long-range networks that robustly interconnect multiple regions across hemispheres, often exhibiting patterns distinct from known neuronal networks. We also demonstrate that astrocyte networks undergo structural reorganization in the adult brain after sensory deprivation. These findings reveal a mode of communication between distant brain regions that is mediated by plastic networks of gap junction-coupled astrocytes.
Similar content being viewed by others
Main
Astrocyte intercellular communication is critical to proper CNS function. This communication occurs through gap junctions—membrane channels that connect the cytoplasm of neighbouring cells, enabling them to redistribute resources and share biochemical signals. Studies using mice lacking astrocyte gap junctions have shown that these gap junctions are necessary for memory formation1,2, synaptic plasticity3,4,5, coordination of neuronal signalling6, and closing the visual and motor critical periods7,8. In disease, networks of gap junction-linked astrocytes redistribute metabolic resources across the CNS to protect degenerating neurons11,12. Despite these insights, our understanding of the spatial architecture and functional topology of astrocyte networks remains limited. Existing methods, such as dye diffusion in acute slices or reporter activation in injury models, are inherently constrained to local environments and often disrupt native connectivity. As a result, it remains unclear whether astrocytes form a continuous brain-wide syncytium or operate through discrete, region-specific subnetworks. It is also unclear whether their anatomical connectivity aligns with neuronal networks or establishes an independent framework for long-range, non-neuronal signalling.
Tracing astrocyte gap junction networks
To address this knowledge gap, we developed a vector-based approach to express a fusion protein comprising connexin 43 (Cx43, encoded by Gja1), the main gap junction protein used by astrocytes, and TurboID (TID), a rapid and promiscuous biotinylating enzyme13,14,15, under the shortened Gfap promoter16,17,18 (AAV5-GfaABC1D-Cx43:TID:HA; Fig. 1a). When this fusion protein incorporates into a connexon as any one of its six constituent connexins (Fig. 1b), molecules that flux through the infected astrocyte’s gap junctions are rapidly tagged with small, inert biotin (Fig. 1c). This enables us to detect the infected astrocyte population (through the haemagglutinin (HA) tag on the fusion protein), the in-network astrocytes (by staining biotinylated moieties with streptavidin) and cells that are not in-network with the infected population (gaps in astrocyte tiling; Fig. 1d). As the mouse CNS has minimal-to-no native biotin13,14,15, streptavidin staining provides little background. Moreover, each biotin can bind to only a single streptavidin, eliminating variability introduced through antibody multiplexing and making images more quantitative.
a, Diagram of the astrocyte network tracer construct. b, Cx43 connexons contain six connexins, only one of which needs to be the fusion protein for TID to reside on the gap junction. c, Infected astrocytes biotinylate molecules that flux through Cx43 gap junctions into adjacent, uninfected cells. d, This volume-fills in-network astrocytes with biotinylated molecules such that infected (HA+streptavidin+), in-network (HA−streptavidin+) and out-of-network (HA−streptavidin−) cells can be visualized. e, Infection of primary immunopanned rat astrocytes (GFAP, green) resulted in Cx43–TID–HA expression (cyan) in 10% of cells, forming a biotin-labelled network containing 80% of cells (streptavidin, magenta) (Extended Data Figs. 1 and 2). f, Primary rat astrocytes were expanded 8.7× to visualize individual Cx43 gap junctions (grey) and incorporated fusion protein (HA, cyan). g, Super-resolution microscopy analysis of expanded astrocytes20 enabled single-molecule imaging of the fusion protein incorporated into astrocyte gap junctions expressed in immunopanned primary rat astrocytes. Polyclonal Cx43 antibodies conjugated directly to fluorophores reveal gap junction structure (grey), and the HA tag (cyan) shows TID within the gap junction vestibule of a small gap junction plaque. h, Single-molecule localization microscopy (SMLM) analysis of HA (cyan) and Cx43 (magenta). i, Quantification of the intrafluorophoric distance from HA to the nearest Cx43. The median (20.73 Å) is indicated by a solid blue line; the dotted lines denote the 25th (18.09 Å) and 75th (25.74 Å) percentiles. j, In vivo experimental timeline. Three weeks after viral injection, mice received biotin-supplemented drinking water for 1 week, after which they were perfused. The brains were then delipidated, stained with streptavidin to reveal biotinylated molecules, cleared and imaged using light-sheet microscopy. k, Light-sheet micrograph of the HA-tag (cyan) shows gap junctions containing fusion protein in an infected cell within a cleared brain. l, Streptavidin (magenta) labelling of biotinylated molecules. m, Merged image showing a two-cell example of an infected (left) and an uninfected in-network (right) cell. Scale bars, 20 μm (k–m); 100 μm (e); 23 μm, expanded to 200 μm (f); 574 Å, expanded to 500 nm (g); and 114 Å, expanded to 100 nm (h).
To validate the construct, we first infected cultures of primary immunopanned serum-free rat astrocytes (Fig. 1e; purity confirmation is shown in Extended Data Fig. 1a, and full blots are provided in Supplementary Fig. 1). When around 10% of cells expressed Cx43–TID, over 80% of cells in the plate were identifiably in-network (Fig. 1e; quantification and controls are shown in Extended Data Fig. 2). Streptavidin staining beyond the infected astrocyte population was confirmed to be gap junction-dependent in primary astrocyte cultures from mice with inducible knockout (KO) of astrocyte gap junctions (Extended Data Figs. 1 and 3).
We next confirmed that, when incorporated into a gap junction, the tertiary structure of the fusion protein was optimized. Fusion of TID to the C terminus of Cx43 through a linker that places the enzyme within the gap junction vestibule ensures that only molecules that can fit into the vestibule and are primed for gap junction flux are biotinylated, meaning that the system should be more precise than a cytoplasmically expressed TID. To determine that TID resides within the gap junction vestibule and was therefore proximal to molecules staged for gap junction flux19, we used a combination of expansion microscopy20 (Fig. 1f) and super-resolution imaging (Zeiss Elyra 7 SIM2). This approach enabled us to visualize the fusion protein incorporated within a gap junction in expanded primary cultured rat astrocytes21 (Fig. 1g). It also enabled us to measure the distance between the Cx43 vestibule and the HA tag on the N terminus of TID (Fig. 1h,i). The 20.73 Å median distance indicates that the HA tag lies immediately outside the approximately 15 Å gap junction vestibule19. While the 3D structure of TID has not been solved, the labelling radius of biotin ligases throughout the TID family of enzymes has been estimated at around 10–30 nm (100–300 Å)13,22,23, well within all measured intrafluorophoric distances.
After our in vitro validation, we moved to in vivo experiments (Fig. 1j). For each in vivo experiment, mice were unilaterally injected with AAV5-GfaABC1D-Cx43:TID:HA (hereafter, astrocyte network tracer) in a single defined brain region. Then, 3 weeks later, biotin was administered in the drinking water for 1 week. On day 28, the mice were perfused and the brains carefully dissected. The brains were then delipidated, stained, cleared and imaged using light-sheet microscopy. In cleared brains, infected cells expressed Cx43–TID–HA in a punctate pattern on the cell membrane, characteristic of gap junctions (Fig. 1k). Streptavidin staining revealed an adjacent in-network, uninfected cell (Fig. 1l); while this cell was negative for Cx43–TID–HA, it contained biotinylated molecules that had passed through the infected astrocyte’s gap junctions (Fig. 1m).
Astrocyte networks vary regionally
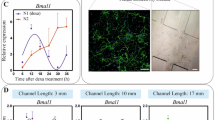
To visualize in vivo astrocyte networks, assess their reproducibility across mice and examine regional differences in connectivity, we injected male, 3-month-old C57BL/6 mice with astrocyte network tracer in one of three brain regions: motor cortex (Fig. 2 (top row) and Supplementary Video 1), hypothalamus (Fig. 2 (middle row) and Supplementary Video 2) or prefrontal cortex (Fig. 2 (bottom row) and Supplementary Video 3). Dorsal, sagittal and oblique views of the light-sheet datasets illustrate distinct network morphologies across regions and reveal the specificity of astrocytic connectivity. Infected cells displayed the canonical stellate morphology of astrocytes as well as the expected punctate HA-tag staining pattern (Extended Data Fig. 4). We found consistent patterns of regional connectivity among mice injected in the same area (Extended Data Figs. 5–7 and Supplementary Video 4), suggesting that astrocyte networks are conserved among mice.
Three-dimensional renderings of light-sheet-imaged brains infected in motor cortex (top row), hypothalamus (middle row) or prefrontal cortex (bottom row). Each image is presented in two views: the first shows the infected region (HA-tag, cyan) inside the autofluorescence capture (grey) for context; the second shows the same infected region within the streptavidin-stained astrocyte network originating from that brain region (magenta). The dorsal (left column), sagittal (middle column) and oblique (right column) views of the same light-sheet datasets show the network morphology at different angles. Full rotational views of each brain are provided in Supplementary Videos 1–3. Scale bars, 2 mm (dorsal and ventral) and 3 mm (sagittal).
To further probe the structure and consistency of astrocyte networks, we registered our samples to the Allen Reference Atlas and quantified the average streptavidin signal intensity per region (Fig. 3a). Networks in the hemispheres ipsilateral and contralateral to the injection site were then compared with one another to determine significantly enriched regions in each network and to begin to pull out similarities and differences among networks. Most of the motor cortex network regions were exclusively enriched in that network (Fig. 3b,c and Extended Data Fig. 8), meaning that the motor cortex network was largely distinct among the networks quantified. By contrast, most regions enriched in the frontal cortex and hypothalamic networks were enriched in both networks, indicating that these two networks may in fact represent bidirectional communication across the same sequence of brain regions. There are a few regions that exhibit robust bilateral signal in all networks, such as the superior colliculus these regions may serve as local hubs of astrocyte network connectivity. Multiple other regions contain signal only on either the ipsilateral or contralateral side of the brain (Fig. 3d and Supplementary Video 5). Thus, a contralateral network is not necessarily a smaller reflection of its corresponding ipsilateral network.
a, The summed intensity per region normalized to a region’s volume, for each brain unilaterally injected in motor cortex (left), prefrontal cortex (middle) or hypothalamus (right). Ipsilateral and contralateral hemispheres are grouped and shown sequentially. Regions (rows, right labels) are grouped by overarching functional or structural properties (rows, left labels). b,c, Euler diagrams demonstrating the number of significantly enriched regions (corresponding to heat map right row labels) in brains injected in the motor cortex (yellow), prefrontal cortex (blue) or hypothalamus (pink) for the ipsilateral (b) or contralateral (c) hemispheres. d, Heat map (colour defined as in a) of coronal sections showing the average values in the motor cortex (M), prefrontal cortex (P) and hypothalamus (H) networks. Top row, atlas outline with injected regions highlighted (Supplementary Video 5). Significance for b and c was determined using t-tests with Benjamini–Hochberg correction (α = 0.05). Significance had to be achieved for subregions accounting for 60% of the volume of the composite subregions for each region listed (at minimum).
Virtual coronal sections of the raw data mapped24 to the Allen Brain Atlas further illustrate the specificity and local complexity of astrocyte networks (Extended Data Fig. 9). Regions with robust connectivity are often directly adjacent to areas that lack any detectable signal. Within a single astrocyte network, different brain regions can display distinct patterns of network-linked cells (Extended Data Fig. 10). While some regions remain largely unilaterally connected (Extended Data Fig. 10a′′), others show robust bilateral connectivity (Extended Data Fig. 10a′). In rare instances, specific neuronal populations contain biotinylated molecules. When present, this labelling typically occurs at the terminus of an interconnected chain of astrocytes and is restricted to neurons within a localized area (Extended Data Fig. 10c). Most cells (and, in many networks, all cells) that contain biotinylated molecules exhibit the characteristic stellate morphology of astrocytes (Supplementary Video 6).
Astrocyte networks can link multiple astrocyte subtypes. In bilateral networks, chains of interconnected astrocytes originating in the grey matter before spanning white-matter tracts such as the corpus collosum are clearly visible (Extended Data Fig. 11). Rather than filling the entire corpus callosum, these chains of astrocytes appear to follow specific neuronal axons. Networks can also include both parenchymal astrocytes and glia limitans superficialis astrocytes25 located at the brain’s surface (Extended Data Fig. 12). These two astrocytic populations contact one another, although the glia limitans superficialis astrocytes are not necessarily directly coupled.
Gap junctions link distant astrocytes
To examine whether non-gap-junctional mechanisms could account for biotinylated molecules beyond the initially infected cells, we used tamoxifen-inducible, astrocyte-specific Gjb6 (encoding Cx30) and Gja1 double KO mice (Slc1a3:cre-ERT2+/ × Gja1fl/fl × Gjb6fl/fl, hereafter, conditional KO (cKO) mice)1. KO of Gja1 alone is insufficient, as astrocytes also express connexin 30 (Cx30), albeit at lower levels1. We designed our vector to biotinylate molecules fluxed by Cx43 because it is the most permissive connexin26. However, we knocked out both Gja1 and Gjb6 in this cKO to control for the possibility that a subset of small molecules might still pass through Cx30 in the absence of Cx43.
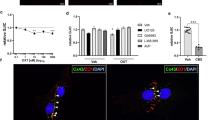
We first used immunohistochemistry (Fig. 4 and Extended Data Fig. 3) and quantitative PCR (qPCR) (Extended Data Fig. 1) to confirm reduction of connexin expression in cKO mouse astrocytes. One week after tamoxifen gavage, immunoreactivity for Cx43 (Fig. 4a,c,e) and Cx30 (Fig. 4b,d,f) was diminished in cKO mice relative to in the littermate controls. As expected for Cx43, a small degree of immunoreactivity was retained around vasculature in cKO mice (vasculature marked in Fig. 4c–f using Lycopersicon esculentum (tomato) lectin (TL), red). Pericytes and a subset of endothelial cells both express Cx4327,28 and are not Cre+ in this model. The persistence of connexins adjacent to vasculature, but not elsewhere, confirms that our gap junction KO was specific to astrocytes. We quantified this by determining the fluorescence intensity relative to the distance from the nearest blood vessel in two-dimensional sections. Immunoreactivity in cKO brains (n = 3 per group) was significantly reduced compared with the littermate controls at all locations except for around 3 µm from vessel walls, as expected (Fig. 4i).
a,b, Sagittal sections of a Gja1fl/fl × Gjb6 fl/fl mouse brain (a; littermate) and a Slc1a3-cre-ERT2 × Gja1fl/fl × Gjb6fl/fl (cKO) mouse brain (b). Both mice were given tamoxifen, inducing robust connexin ablation in cKO mice; the sections were stained for Cx43. c–f, Higher-resolution images of hippocampi stained for GFAP (green, astrocytes), TL (red, vasculature), and either Cx43 (c,e) or Cx30 (d,f) (white) in control (c,d) or cKO (e,f) mice. g, Control mice injected in the hippocampus express the astrocyte network tracer in CA2 (HA tag, cyan) and exhibit bilateral streptavidin labelling across the hippocampus and hypothalamic regions. h, In cKO mice, streptavidin labelling is restricted to infected astrocytes where Cx43 is reintroduced through the astrocyte network tracer. The white asterisks in g and h mark ventricular autofluorescence. i, Quantification of Cx43 expression across vasculature and parenchyma. In cKO mice, Cx43 immunoreactivity is significantly diminished in all locations except at the periphery of blood vessels, where pericytes reside. Statistical analysis was performed using paired two-sided t-tests across 180 locations relative to vasculature with Bonferroni correction; *P < 0.001 in all significant locations. n = 3 mice per condition; 3 micrographs were averaged to generate values for each mouse. j, Quantification of the astrocyte network volume normalized to the volume of infected astrocytes. When astrocytes are infected in either hippocampus (HPC), prefrontal cortex (PFC) or barrel cortex (BC), cKO mouse networks are significantly reduced from those in control mice. Statistical analysis was performed using unpaired two-sided t-tests; *P < 0.001 in all regions. n = 4 mice per region per condition. Data are mean ± s.d. k, cKO mice induced through tamoxifen gavage were injected with astrocyte network tracer according to the experimental timeline in Fig. 1j. TL was injected during perfusion to stain the vasculature. The micrographs show the streptavidin-labelled astrocyte network (magenta) and vasculature (red). The network does not appear to follow or expand along blood vessels after astrocyte network loss. Scale bars, 1 mm (a, g (top) and h (top)), 200 μm (g (bottom) and h (bottom)) and 50 μm (c–f).
We infected four cKO and four littermate control mice per condition in the hippocampus, prefrontal cortex or barrel cortex. Astrocyte networks originating in each region were significantly smaller in cKO mice than in the control mice (Fig. 4j). The network volume within each brain region was consistent among animals in the same condition, even among networks of different sizes, such as the relatively small barrel cortex astrocyte network. This internal consistency parallels that observed in C57BL/6 mice injected in the motor cortex, prefrontal cortex and hypothalamus (Extended Data Figs. 5–7 and Supplementary Video 4). After KO, the network volume in each brain region was restricted to the infected astrocytes, which had Cx43 reintroduced through the astrocyte network tracer. These cells could form a network with one another, but not with neighbouring uninfected astrocytes lacking Cx43 and Cx30.
We next examined whether endothelial- or pericyte-expressed Cx43, or other vasculature-related mechanisms, could compensate for the loss of astrocyte gap junctions and support the distribution of biotinylated molecules in this system. After infection with the astrocyte network tracer and 1 week of biotin-supplemented water, mice were perfused with TL. Brains were subsequently stained and cleared. We found no evidence that the streptavidin signal followed the vasculature in cKO brains (Fig. 4k), indicating that vasculature-related mechanisms did not compensate for the loss of astrocyte gap junctions.
Astrocyte networks are plastic
The barrel cortex is a pliable and accessible forebrain region that is commonly used to investigate neuroplasticity29,30,31. To test whether astrocyte networks, like neurons, exhibit structural plasticity, the whiskers of 4-week-old mice were unilaterally trimmed for 28 days (Fig. 5a). This manipulation is known to induce robust structural remodelling in neurons29,30,31,32,33. At day 28, a viral mixture containing the astrocyte network tracer and CaMKIIα-driven mCherry (to label excitatory neurons) was injected into the barrel cortex corresponding to the trimmed whiskers (Fig. 5b). Whiskers were trimmed for an additional 4 weeks, with biotin administered in the drinking water during the final week.
a, Experimental timeline. P28, postnatal day 28. b, Diagram of the constructs in the viral mixture. c, After whisker trimming, the number of streptavidin-positive cells relative to the number of HA-positive cells was significantly reduced in corresponding barrel cortex relative to naive animals. n = 4 (naive) and n = 3 (trimmed) mice. Statistical significance (P value indicated on graph) was determined using the one-sided Mann–Whitney U-test (α = 0.05). Data are mean ± s.d. d, Three-dimensional rendering of a naive mouse infected with both constructs in a single injection. Streptavidin (magenta) shows astrocyte connectivity originating from the same region in which infected neurons (yellow, mCherry) reside. e,f, Horizontal (e) and coronal (f) virtual sections provide a detailed view of connectivity among specific regions. The magenta arrows highlight regions in which astrocyte connectivity was altered after whisker trimming; the yellow arrows highlight reduced neuronal dendritic arbours after whisker trimming. g–i, Images corresponding to d–f, respectively, in a brain from a mouse with unilaterally trimmed whiskers. Scale bars, 3 mm (g (right)), 2 mm (d, e, g (left) and h) and 1 mm (f and i).
We quantified the network size in both trimmed and naive conditions by calculating the ratio of streptavidin-positive to HA-positive cells in a cohort with reduced injection volumes to improve measurement precision (Fig. 5c and Extended Data Fig. 13). Mice with unilaterally trimmed whiskers had significantly smaller astrocyte networks in the corresponding barrel cortex (P = 0.002; naive 3.54 ± 0.39 versus trim 2.16 ± 0.22), indicating that astrocyte networks are indeed structurally plastic.
Astrocyte and neuronal networks differ
We next increased the injection volume to assess commonalities and differences among long-range astrocytic networks and neuronal projections. While the astrocyte streptavidin signal occasionally overlapped with neuronal projections, this was not uniformly the case. Figure 5d shows a three-dimensional rendering of astrocyte networks and neuronal projections in naive barrel cortex. While local connections and some long-range projections were shared among astrocytes and neurons, many neuronal projections lacked corresponding astrocyte connectivity. This divergence became more pronounced after whisker trimming. As shown in Fig. 5g, the astrocyte network contracted markedly, particularly in prefrontal cortex. Virtual horizontal (Fig. 5e,h) and coronal (Fig. 5f,i) sections of these brains reveal the extent of this effect. In the naive brain, barrel cortex astrocyte networks project to prefrontal cortex and show limited contralateral signal, both of which are largely absent after whisker trimming. By contrast, midbrain connectivity remained relatively stable. These findings demonstrate that long-range astrocyte networks are structurally plastic, and that their architecture can diverge significantly from that of the neuronal connectome.
Conclusion
Here we demonstrate that multiple astrocyte networks traverse the mouse brain, each selectively connecting and omitting brain regions rather than diffusing indiscriminately (Fig. 2). These networks vary in size and organization, with some local networks limited to a single brain region and other long-range networks robustly interconnecting multiple regions across hemispheres. Interconnected astrocytes display region-specific patterns of connectivity within the same network (Fig. 3). In some regions, such as prefrontal cortex in the motor cortex-derived astrocyte network, nearly all astrocytes contain biotinylated molecules. In other regions, only subsets of astrocytes are linked, sometimes in linear chains and other times in honeycomb-like patterns that exclude scattered astrocytes. The morphology of networks in different regions may provide clues about their local functions. In brain regions that are fully interconnected, we hypothesize that the network buffers ions across as many astrocytes as possible. In regions with honeycomb-like patterns, we propose that multiple interleaved astrocyte networks may occupy distinct functional niches for the same neuronal population.
Astrocyte networks can directly link brain regions that are not connected by neurons, suggesting that previously unassociated brain regions communicate with one another through gap junction-coupled astrocytes. The physiological role that such an overlapping multicellular network might have in the brain is likely to be multifaceted and is beyond the scope of this study. However, it is tantalizing to hypothesize that, under physiological conditions, these networks facilitate the redistribution of metabolic and antioxidant support from regions of low neuronal activity to those with high demand. Glutathione and phosphocreatine could mediate such support by serving antioxidant and energetic buffering roles, respectively, and both are small enough to diffuse through Cx43 gap junctions26,34,35,36. By contrast, during pathology, astrocyte networks may help to dissipate locally accumulated metabolic byproducts or pathogenic molecules by redistributing them across the network for degradation or clearance from the CNS37,38,39.
Some networks also exhibit streptavidin labelling in small, specific populations of neurons. Connexin 36, the canonical neuronal connexin40, is a delta-type connexin and is therefore incompatible for heterodimerization with the alpha-type Cx43 or beta-type Cx30 expressed by astrocytes—explaining why neuronal streptavidin staining is so rare. A notable exception is motor neurons, which have been reported to express Cx43 during development or in response to injury41,42,43. Although we did not induce a substantial injury and studied only adult mice, we observed streptavidin-positive motor neurons projecting through the spinal cord one month after astrocyte network tracer injection into the motor cortex (Extended Data Fig. 10). While this signal could have originated from astrocyte–oligodendrocyte–neuron communication, the continuous linear signal and lack of distinguishable cell bodies in the white matter imply that is not the case, at least at the moment of sample collection. Rather, this observation suggests that connexin-mediated astrocyte–neuron communication in motor neurons may be more prevalent than previously appreciated.
The reason for astrocyte-to-neuron molecular transfer is unclear, but there are several potential explanations. Neurotransmitters have been shown to flux gap junctions44, raising the possibility that astrocyte networks resupply neurons with neurotransmitters, allowing them to conserve energy. This may be particularly important for neurons with long, energetically intensive projections. Moreover, highly active neuronal populations may also receive antioxidants such as glutathione (~307 Da) from astrocytes to mitigate oxidative stress without incurring the metabolic cost of de novo antioxidant production. Future work should use mass spectrometry to identify the specific biotinylated molecules that are transferred through astrocyte networks, which will help us to understand the functional roles of astrocyte networks across conditions.
Crucially for future experimentation, astrocyte networks link both hemispheres, typically including at least the contralateral counterpart of the infected region. This raises critical considerations: internally controlled experiments must be interpreted cautiously, as astrocyte connectivity may undermine the assumption of regional independence. Moreover, gap junction expression and coupling are known to change through injury—even in localized injury models such as traumatic brain injury45,46, ischaemia47 and focal seizure48. We may therefore need to modify our interpretations of previous work that used an internal control, particularly if it was the contralateral counterpart of an injured region. Regions that are functionally linked to an experimental region by astrocyte networks are unlikely to remain biologically naive.
This bilateral connectivity may also explain other contralateral astrocytic responses to unilateral stressors. For example, ischaemic stroke in somatosensory cortex causes an array of astrocytic responses in the contralateral hemisphere after ipsilateral limb stimulation49. Similarly, a unilateral model of glaucoma elicits astrocytic responses in both the contralateral retina and the contralateral hemisphere of the brain11,50. Importantly, these contralateral responses are not restricted to a single sensory system, suggesting that astrocyte-mediated interhemispheric communication may represent a broader feature of neurodegenerative progression.
We also find that astrocyte networks are plastic. Barrel cortex astrocyte networks shrink in response to whisker trimming (Fig. 5). To mirror the canonical timeline of neuronal remodelling, we trimmed the whiskers for a month before astrocyte infection30. However, both astrocyte gap junctions and the fine processes that house them can remodel on much shorter timescales. The half-life of Cx43 is estimated to be between 1.5 and 5 h (refs. 51,52), and astrocyte process remodelling occurs over similarly rapid intervals53. Our whisker barrel experiments indicate that astrocyte networks can reconfigure in a stable manner, but it is also likely that rapid remodelling occurs on a smaller scale during more acute perturbations. Future studies should explore whether acute energetic stress triggers dynamic reorganization of astrocyte networks to meet varied metabolic demands across the CNS.
A strength of this method is the ability to implement the astrocyte network tracer in numerous models, but it is true that the five allele modifications on the cKO mice used herein as controls are not feasible to add to other genetic models. To strengthen reliability in future experiments where cKO mice may be unfeasible, there are two control experiments that should be considered. First, fusion of TID to the amino terminus of Cx43 would generate a fusion protein expected to block the channel function of endogenously expressed Cx43 when incorporated into gap junctions. This would be a complimentary and more definitive test of whether the biotin transfer requires open Cx43 gap junction channels, rather than simply the presence of Cx43 protein. Second, cytoplasmic (unfused) TID expression could be a useful control for specificity of biotinylation in the original construct. This experiment should generate lower, but still detectable, biotin across a network but would be more likely to also mark other methods of cell–cell communication—a potentially useful tool for looking at combinations of mechanisms of cell–cell transfer for specific molecules.
While our approach visualizes astrocyte networks through a binary lens (connected or not through viral labelling), it is important to recognize that a diverse array of molecules may flux through these networks, and their distribution is unlikely to be uniform. Local and long-distance astrocyte networks may have distinct physiological roles, even within the same interconnected chain of cells. Occasionally, biotin-containing astrocytes that appear to be unconnected to the rest of a network are visible. We hypothesize that this may represent cells previously connected to the network, perhaps earlier in the day, that are no longer linked but still contain biotinylated molecules that have not yet been degraded. Identifying the specific dynamics of different fluxed molecules and how these dynamics are shaped by the molecular identity and subtypes of each astrocyte will be a complex task. However, such studies may ultimately reveal a rich array of functional niches that astrocyte networks occupy across the brain.
Here we define the specificity and spatial extent of astrocyte networks across brain regions, demonstrate how they differ from neuronal networks and reveal their ability to remodel. These findings establish a foundation for future exploration of how astrocyte network structure and function are shaped by injury, disease, development, ageing and experience-dependent processes such as learning and memory.
Methods
Mice
Animal procedures were performed in accordance with National Institutes of Health guidelines with the approval of NYU Grossman School of Medicine’s Institutional Animal Care and Use Committee (IACUC). All animals were housed at 22–25 °C with 50–60% humidity. Animals had access to food and water ad libitum and were housed under a 12 h–12 h light–dark cycle. Experiments in Figs. 2 and 3 were on male 12-week-old C57BL/6J mice. For experiments in Fig. 4, the Saab laboratory crossbred mice1 expressing the tamoxifen-sensitive Cre recombinase cre-ERT2 under the control of the mouse Slc1a3 (encoding GLAST) promoter54 with mice carrying floxed Gjb6 (Gjb6fl/fl)55 and floxed Gja1 (Gja1fl/fl)56. Mice hemizygous for Slc1a3:cre-ERT2 were bred to noncarriers to generate Slc1a3:cre-ERT2+/ × Gja1fl/fl × Gjb6fl/fl experimental mice and Gja1fl/fl × Gjb6fl/fl littermate controls for in vivo experiments; in vivo experiments on this genotype were balanced for sex. When primary mouse astrocytes were isolated, Slc1a3:cre-ERT2 was kept homozygous to obtain a culture in which all astrocytes could be induced through 4-hydroxytamoxifen. All mice were on a C57BL/6 background. The primer sequences used for genotyping were as follows: for Gjb6 flox: 5′-TTCCCTATGCTGGTAGAGTGCTTGT-3′ and 5′-GCAGTAACTTATTGAAACCCTTCACCT-3′; for Gja1 flox: 5′-GGGATACAGACCCTTGGACTCC-3′ and 5′-TCACCCCAAGCTGACTCAACCG-3′; and for Slc1a3-creERT2: 5′-GAGGCACTTGGCTAGGCTCTGAGGA-3′, 5′-GAGGAGATCCTGACCGATCAGTTGG-3′ and 5′-GGTGTACGGTCAGTAAATTGGACAT-3′. The Saab laboratory previously analysed Cre reporter function1 by crossing with ROSA26-floxed-STOP-GCaMP6s (Ai96, JAX, 024106). After shipment, mice were rederived by NYU Grossman School of Medicine’s Rodent Genetic Engineering Laboratory.
Tamoxifen treatment
Tamoxifen treatment was performed as previously described11. For 3 consecutive days, both Slc1a3:cre-ERT2+/ × Gja1fl/fl × Gjb6fl/fl experimental and littermate control mice received daily (between 15:00 and 17:00) gavage of a tamoxifen solution (20 mg ml−1 dissolved in corn oil). The solution was prepared 2 h before administration, during which it was shaken at 37 °C in the dark. Any further experimental manipulations were performed at least 7 days after the last dose of tamoxifen.
Primary rat astrocyte cultures
Isolation of primary rat astrocytes by immunopanning was performed as previously described21. In brief, cortices of postnatal day 6/7 Sprague–Dawley rats (Charles River) were blunt dissected, the meninges were removed and the cortices were enzymatically dissociated at 37 °C and 10% CO2 using papain (Worthington Biochemical, LS003126). Tissue was then triturated using a 5 ml serological pipette to generate a single cell suspension, resuspended in Dulbecco’s PBS (VWR, SH30264.FS) with BSA (BSA, Sigma-Aldrich, A4161) and DNase (Worthington Biochemical, LS002007), and filtered using a 20 µm nitex filter. The suspension was negatively panned for non-specific secondary antibody binding, endothelial cells (BSL-1, Vector Labs L-1100), microglia (CD45, BD Pharmingen, 553076) and oligodendrocyte lineage cells (O4 hybridoma, generated in house), and positively panned for astrocytes (ITGB5, Thermo Fisher Scientific, 14-0497-82). Astrocytes were removed from the positive panning plate using TrypLE (Thermo Fisher Scientific, 12-605-010) and plated at 10,000 cells per well in an eight-well chamber slide precoated with poly-l-lysine (Ibidi 80804). Astrocytes were cultured in serum-free medium containing 50% neurobasal, 50% DMEM, 100 U ml−1 penicillin, 100 μg ml−1 streptomycin, 1 mM sodium pyruvate, 292 μg ml−1 l-glutamine, 1× SATO, 5 μg ml−1 N-acetylcysteine and 5 ng ml−1 HBEGF (Sigma-Aldrich, E4643-50UG). After 7 days, AAV was added to the medium change, resulting in an effective titre of 105. Cells were incubated for a further 7 days, with biotin (effective concentration, 250 µM) added to the medium 2 days before fixation.
Expansion microscopy
Expansion microscopy was performed based on published protocols20. Primary astrocytes were quickly washed in PBS, then fixed in 4% PFA in PBS at room temperature for 15 min. After three 5-min washes in PBS, the cells were permeabilized for 15 min in PBS with 0.5% Triton X-100, then blocked for 1 h in 5% BSA and 0.2% Tween-20 in PBS with shaking at room temperature. They were then incubated in primary antibody solution overnight at 4 °C (3% BSA, 0.2% Tween-20 in PBS containing mouse monoclonal anti HA-tag (Cell Signaling, 2367) directly conjugated to ATTO488-NHS (ATTO-TEC AD 488-31), rabbit polyclonal anti-Cx43 (Cell Signaling, 3512) directly conjugated to ATTO565-NHS (ATTO-TEC AD 565-31) and ATTO643-streptavidin (ATTO-TEC, AD 643-61)). Dye conjugation was performed according to the manufacturer’s protocol. Antibodies were added for an effective concentration of 2 µM for anti-Cx43 and 10 µM for anti-HA, and streptavidin (stock concentration 2 μg ml−1) was added at 1:1,000. Cells were then washed three times for 5 min in PBS and incubated for 10 min in 300 nM DAPI in PBS, then imaged on a spinning-disk confocal microscope (CrestOptics X-LIGHT V3 Confocal on Nikon Ti2) with a ×60/1.4 NA oil lens to later establish an expansion coefficient. Then, 250 µl Acryloyl-X, SE (Invitrogen, A20770) was added for overnight incubation at room temperature.
Cells were washed twice for 15 min in PBS, then incubated in 300 µl Gelation Solution per well (for 2 ml: 542 µl 4 M Na acrylate (VWR, S03880), 1 ml PROTOGEL (Thermo Fisher Scientific, 50-899-90119), 200 µl 10× PBS, 198 µl H2O (MQ), 30 µl 10% ammonium persulfate (Thermo Fisher Scientific, 17874), 30 µl tetramethylethylenediamine (TEMED, Thermo Fisher Scientific, 17919)) for 1 h at 37 °C. Then, 300 μl digestion buffer was added to each well and the gels were carefully outlined with a needle to facilitate detachment; once the gels detached, they were each transferred to one well of a 12-well plate for the remainder of the 4 h incubation at 37 °C (volume per gel/well of 12-well plate: 1,550 µl Tris-acetate-EDTA buffer, 100 µl 10% Triton X-100 in PBS, 320 µl 5 M NaCl with 26 µl proteinase K (Thermo Fisher Scientific, EO0491) added immediately before use). The gels were then each transferred to 15 cm Petri dishes filled with room-temperature H2O (MQ) and incubated for 30 min at room temperature. H2O (MQ) was then replaced and the gels were left overnight at room temperature to expand. The gels were then stored at 4 °C until imaging.
Single-molecule imaging and localization
Gels were mounted into 35 mm dishes with #1.5 coverslip bottoms (Ibidi, 81158) with SlowFade Glass soft-set antifade mountant (RI 1.52; Invitrogen, S36917) immediately before imaging. For lattice-SIM imaging, we used the ZEISS Elyra 7 system with an Alpha Plan Apochromat ×63/1.46 NA oil total internal reflection fluorescence microscopy objective. Dual PCO.Edge 4.2 sCMOS cameras collected data (1,280 × 1,280 pixels) with a Dual Camera Beam Splitter (SBS LP 560) used for channel separation on Zen Black 3.0 SR software corresponding to 15 phases of a 27.5 µm grating period for both 488 and 561 channels. SIM2 (structured illumination microscopy) 3D Leap processing was run using the following parameters: input SNR, low; Iterations, 25; regularization weight, 0.01; processing sampling and output sampling, 2. At least 10,000 frames with a 30 ms acquisition time were collected from each sample for each wavelength channel and processed using the Localization Microscopy processing function in 3D. All single- and multi-emitter events were fitted as single emitter events.
Primary mouse astrocyte cultures
Mouse astrocytes were obtained from Slc1a3:cre-ERT2+/+ × Gja1fl/fl × Gjb6fl/fl (note the homozygous Cre, opposed to hemizygous for in vivo experiments where mice could be genotyped). Cortices of postnatal day 3 mice were blunt dissected, meninges were removed, and the cortices were enzymatically dissociated at 37 °C and 10% CO2 by papain (Worthington Biochemical, LS003126). Tissue was then triturated using a 5 ml serological pipette to generate a single-cell suspension, resuspended in Dulbecco’s PBS (VWR, SH30264.FS) with BSA (Sigma-Aldrich, A4161) and DNase (Worthington Biochemical, LS002007), and filtered using a 20 µm nitex filter. The suspension was negatively panned for non-specific secondary antibody binding, microglia (CD45, BD Pharmingen, 553076), oligodendrocyte lineage cells (O4 hybridoma57, generated in house) and L1 (Millipore, rat anti-L1, MAB5272), and positively panned for astrocytes (HepaCAM58; human HepaCAM antibody, MAB4108, R&D Systems). Astrocytes were removed from the positive panning plate using TrypLE (Thermo Fisher Scientific, 12-605-010) and plated at 10,000 cells per well in an eight-well chamber slide pre-coated with poly-l-lysine (Ibidi 80804). Astrocytes were cultured in serum-free medium containing 50% neurobasal, 50% DMEM, 100 U ml−1 penicillin, 100 μg ml−1 streptomycin, 1 mM sodium pyruvate, 292 μg ml−1 l-glutamine, 1× SATO, 5 μg ml−1 N-acetylcysteine and 5 ng ml−1 HBEGF (Sigma-Aldrich, E4643-50UG). On day 6, 4-hydroxytamoxifen (Sigma-Aldrich, SML1666) was added at 1:5,000 (or concentration appropriate for gradient experiments); ethanol:isopropanol (95:5) was added to control wells at the same concentration. On day 7, AAV was added to the medium change, resulting in an effective titre of 1.5 × 104. Cells were incubated for a further 7 days, with biotin (effective concentration 250 µM) added to the medium 2 days before fixation. Cells were washed twice in PBS, then fixed in 4% PFA in PBS for 15 min before immunocytochemistry.
Immunocytochemistry
Fixed cells were washed twice in PBS/azide, then blocked for 2 h in 5% normal donkey serum (NDS) with 0.1% Tween-20 in PBS/azide. They were then incubated overnight at 4 °C with shaking in 3% NDS, 0.1% Triton X-100 in PBS/azide with corresponding primary antibodies (rabbit polyclonal Anti-HA: Cell Signaling, 3724, 1:500; goat polyclonal anti-GFAP, Abcam, ab53554, 1:500). Cells were washed three times for 5 min in PBS/azide, then incubated for 2 h at room temperature with shaking in 1% NDS, 0.1% Triton X-100 in PBS/azide with the corresponding secondary antibodies (donkey anti-goat conjugated to Alexa Fluor 488, Invitrogen, A-110055, 1:500; donkey anti-rabbit conjugated to Alexa Fluor 555, Invitrogen, A-31572, 1:500) and streptavidin conjugated to Alexa Fluor 647 (Thermo Fisher Scientific, S32357, 1:500). Cells were washed three times for 5 min in PBS/azide, then imaged (2,048 × 2,048) on the Zeiss 880 laser-scanning confocal microscope using a ×20/0.8 NA plan apochromat air objective.
Western blotting
Cells were lysed in ice-cold 1× RIPA buffer (diluted in double-distilled H2O from 10×, Cell Signaling, 9806) with 1 mM PMSF (Cell Signaling, 8553) and Halt Protease & Phosphatase Single-Use Inhibitor Cocktail (Thermo Fisher Scientific, 78440). The samples were sonicated and the lysates were separated from insoluble material by centrifugation at 20,000g. Then, 1.5 mm 10% acrylamide gels (Running gel: 4 ml PROTOGEL (Thermo Fisher Scientific, 50-899-90119), 3 ml 1.5 M Tris pH 8.8 + SDS, 5 ml double-distilled H2O, 120 µl 10% ammonium persulfate (Thermo Fisher Scientific, 17874) and 12 µl TEMED (Thermo Fisher Scientific, 17919); stacking gel: 780 µl PROTOGEL (Thermo Fisher Scientific, 50-899-90119), 1.5 ml 0.5 M Tris pH 6.8 + SDS, 3.75 ml double-distilled H2O, 60 µl 10% ammonium persulfate (Thermo Fisher Scientific, 17874) and 18 µl TEMED (Thermo Fisher Scientific, 17919)). The gels were loaded with 10 µg protein per well and run at 120 V; transfers onto PVDF membranes (Immoblion-FL, IPFL00010; pore size 0.45 µm) occurred overnight at 4 °C at 20 V with stirring. After a 1 h block (LI-COR Intercept PBS blocking buffer (LI-COR 927-70001)), the blots were incubated with rabbit anti-Cx43 (1:1,000; Cell Signaling, 3512) and mouse anti-actin (1:1,000; Abcam, ab8226) antibodies in LI-COR Intercept PBS blocking buffer (LI-COR 927-70001) with 0.1% Tween-20 (Sigma-Aldrich, P9416) for 4 h, then washed four times for 5 min with PBST. They were then incubated for 1 h in LI-COR Intercept PBS blocking buffer (LI-COR 927-70001) containing 0.1% Tween-20 and 0.1% SDS with donkey anti-mouse IRDye 680LT (1:20,000, LI-COR, 926-68072) and donkey anti-rabbit IRDye 800CW (1:20,000, LI-COR, 926-32213). After three 5 min PBST washes and one 5 min PBS wash, the blots were imaged on the ChemiDoc MP (Bio-Rad) system using Image Lab v.2.4 (Bio-Rad). The band peaks were quantified in FIJI (ImageJ) using the Gels function.
qPCR
RNA was obtained from cultures using the Qiagen RNeasy Micro Kit (74004) and converted to cDNA using the Qiagen QuantiTect Reverse Transcription Kit (205311) according to the manufacturer’s instructions. qPCR was performed using SYBR Green Universal Master Mix (Thermo Fisher Scientific, 4309155) on the StepOne Plus Real-Time PCR System (Applied Biosystems) using the following primers: Gja1 (F: TCATTGGGGGAAAGGCGTGA, R: CATGTCTGGGCACCTCTCTTTCA), Aldh1l1 (F: TCCCGTCTTTGACCTTGGGT, R: CGCCACCGAGGGAACTTAAA), Slc1a3 (F: CCCCTTACAAAATCAGAAAAGTTGT, R: CCCATCTTGGGCTCTTCTCC), Sox10: (F: GAAGAAGGCTCCCCCATGTC, R: TTGGGTGGCAGGTATTGGTC), Mog (F: GCAGGTCTCTGTAGGCCTTG, R: CCCTCAGGAAGTGAGGATCAAA), Aif1 (F: CTGGGCAAGAGATCTGCCAT, R: ACCAGTTGGCCTCTTGTGTT), Cd14 (F: ACTGAAGCCTTTCTCGGAGC, R: TGAAAGCGCTGGACCAATCT), Tubb3 (F: ACCATGGACAGTGTTCGGTC, R: ACACTCTTTCCGCACGACAT) and Syp (F: CGGCTACCAGCCTGACTATG, R: CTGGGCTTCACTGACCAGAC).
Perfusion and tissue preparation
Mice were always perfused between 14:00 and 17:00 to control for circadian effects. Mice were heavily anaesthetized with an overdose of pentobarbital (Euthasol: 390 mg pentobarbital, 50 mg phenytoin per ml at 2 μl g−1) and transcardially perfused with PBS containing 10 mg l−1 heparin (Sigma-Aldrich, heparin sodium salt from porcine intestinal mucosa, H3393) until the liver and right ventricle were clear of blood. Perfusion solution was then switched to 4% PFA in PBS; all solutions were administered at room temperature at 70% cardiac perfusion pressure. Brains were immediately dissected, taking care to leave the surface fully intact, and then post-fixed in 4% PFA in PBS overnight to stabilize protein tertiary structures. The fixed samples were washed in PBS with 0.01% sodium azide (NaN3) three times for 1 h, then stored in PBS/azide at 4 °C.
When L. esculentum (tomato) lectin was used to image vasculature, 100 µg of TL conjugated to DyLight 649 diluted in 100 µl PBS was injected directly into the left ventricle of the heart before perfusion. After circulating through the beating heart for 1 min, transcardial perfusion resumed as above.
Slice immunohistochemistry
Hemibrains were delipidated (see the ‘Clearing’ section), rehydrated through the same methanol gradient to B1n, then washed three times in PBS/azide. The samples were then sectioned sagittally at 200 µm on a vibrating microtome (Leica, VT1000 S). Floating sections were blocked for 2 h in 5% NDS with 0.1% Tween-20 in PBS/azide, then incubated overnight at 4 °C with shaking in 3% NDS, 0.1% Triton X-100 in PBS/azide with corresponding primary antibodies (rabbit polyclonal anti-Cx43, Cell Signaling, 3512, 1:500; or rabbit polyclonal anti-Cx30, Invitrogen, 71-2200, 1:250; goat polyclonal anti-GFAP, Abcam, ab53554, 1:500). The sections were washed five times for 10 min in PBS/azide, then incubated overnight at 4 °C with shaking in 1% NDS, 0.1% Triton X-100 in PBS/azide with the corresponding secondary antibodies (donkey anti-goat conjugated to Alexa Fluor 488, Invitrogen, A-110055, 1:500; donkey anti-rabbit conjugated to Alexa Fluor 555, Invitrogen, A-31572, 1:500). The sections were washed five times for 10 min in PBS/azide and mounted in ProLong Diamond Antifade Mountant with DAPI (Invitrogen, P36962) to obtain a close refractive index match to the delipidated sections.
Confocal imaging
Sections were imaged on the Zeiss 800 confocal microscope; montages were obtained using a ×20 plan apochromat NA 0.75 air objective at 512 × 512 per image and high-resolution inset images were obtained using a ×63 plan apochromat NA 1.4 oil objective at 2,048 × 2,048 resolution. z stacks were obtained at a step size of 8.2 µm (×20) or 4.4 µm (×63) using Zen Blue 3.2 (Zeiss). All directly compared sections were obtained at the same magnification and were imaged using the same settings. Maximum z projections were rendered using FIJI (ImageJ v.1.54f).
Connexin and vasculature relative localization analysis
Confocal images were analysed in FIJI/ImageJ (v.1.54m). Three confocal micrographs for each of three mice per condition were analysed using ImageJ’s Plot Profile function for three equally spaced line segments perpendicular to imaged vasculature. Line segments were placed so that they began on the edge of a vessel, completely transected it and then passed through the remainder of the micrograph. Profile values for each line were averaged to represent one micrograph. The graph is presented as the distance from the far end of each blood vessel with the average in-vessel portion of all images demarcated.
Intracerebral viral injections
Surgeries were performed under aseptic conditions in accordance with NYU Grossman School of Medicine’s institutional biosafety guidelines. A glass micropipette was pulled to a tip diameter of approximately 20 μm, filled with mineral oil and attached to a Nanoject III microinjector (Drummond Scientific, 3-000-207). The micropipette was then backfilled with 1 μl of adeno-associated Virus (AAV) at a titre of 1011 genomic units per μl. Mice were anaesthetized with 0.7–2.5% isoflurane (adjusted on the basis of scored reflexes and breathing rate during surgery) and placed into a stereotaxic apparatus (Kopf), and a single craniotomy was performed over the experimental brain region (coordinates are provided below); the micropipette was inserted and 200 nl of virus was injected (40 cycles, 5 s delay between cycles, 5 nl injected per cycle and 5 nl s−1). Then, 10 min after injection, the micropipette was withdrawn and the wound was sutured closed. Buprenorphine (0.1 mg per kg) was subcutaneously administered immediately after surgery and twice daily for the next 72 h. We used the following stereotaxic coordinates: motor cortex (anteroposterior (AP), −2.6 mm; mediolateral (ML), 1.3 mm; dorsoventral (DV), −0.8 mm), hippocampus (AP, −2.0 mm; ML, 1.5 mm; DV, −1.5 mm), paraventricular nucleus (AP, 0.12 mm; ML, −0.71 mm; DV, −4.8 mm), prefrontal cortex (AP, 0.35 mm; ML, 1.9 mm; DV, −2.5 mm), barrel cortex (AP, −1.5 mm; ML, 3.5 mm; DV, −0.7 mm). The astrocyte network tracer (AAV5-GfaABC1D-Cx43:TID:HA, Addgene, 252501) was constructed and packaged by VectorBuilder. The AAV9-CaMKIIa-mCherry was obtained from Addgene as a gift from K. Deisseroth (Addgene, 114469-AAV9). All vectors were injected at a titre of 1 × 211.
Biotin administration
Biotin (Sigma-Aldrich,B4639) stocks were diluted to 100 mM in DMSO, aliquoted for single-use and stored frozen at −20 °C. Then, 21 days after intracerebral injections, biotin stock was diluted in distilled water to 160 µM (400 µl of 100 mM stock per 250 ml of distilled water) and provided ad libitum for 7 days, after which the mice were perfused (always between 14:00 and 17:00 to control for circadian effects).
Whisker trimming
Every other day from P28 to P84, mice were lightly anaesthetized with 0.7–1.5% isoflurane (adjusted on the basis of scored reflexes and breathing rate). All whiskers one side of the face were trimmed with scissors. On P56, whisker trimming coincided with intracerebral viral injections (see above). For P77–P84, mice received biotin-supplemented drinking water (see above); perfusion occurred on P84 (see above).
Clearing
The clearing protocol uses similar principles to the iDISCO10 or AdipoClear9 protocol with several adjustments specific to brain tissue and astrocyte morphological maintenance. To delipidate, fixed samples were washed in B1N buffer (H2O, 0.1% Triton X-100, 0.3 M glycine, 0.01% sodium azide, pH 7) for 1 h, followed by 1 h washes each in 20%, 40%, 60% and 80% methanol in B1N. The samples were fully dehydrated in 100% methanol for three 1 h washes, then delipidated in 2:1 dichloromethane (DCM; Sigma-Aldrich, SHBR8133) overnight. Delipidation resumed the next day in three 1 h washes in 100% DCM. The samples were then washed twice in 100% methanol for 1 h. Residual fluorescence was quenched through an overnight incubation in 80% methanol, 5% H2O2 and 15% H2O at 4 °C. The brains were then rehydrated by 1 h incubations in each 60%, 40% and 20% methanol in B1N. The samples were then washed in B1N overnight.
The samples were permeabilized in PTxwH buffer (PBS, 0.1% Triton X-100, 0.05% Tween-20, 2 µg ml−1 heparin, 0.02% sodium azide) with added 45 g l−1 glycine and 50 ml l−1 dimethyl sulfoxide (DMSO), two times for 1 h each. The samples were then washed three times in PTxwH for 1 h.
The samples were incubated in primary antibody (HA-tag, Cell Signaling, 3724, 1:2,000) and conjugated streptavidin (Alexa Fluor 647 streptavidin, Thermo Fisher Scientific, S32357, 1:800) for 1 week at 37 °C with rotation. After primary incubation, the samples were washed in PTxwH with eight solution changes over a period of 3 days. The samples were then incubated in secondary antibody (donkey anti-rabbit conjugated to Alexa Fluor Plus 555, Thermo Fisher Scientific, A32794) and conjugated streptavidin (1:800) for 1 week at 37 °C with rotation. As after secondary incubation, the samples were washed in PTxwH with 8 solution changes over a period of 3 days.
For tissue clearing, the samples were dehydrated in a 20%, 40%, 60%, 80%, 100%, 100%, 100% methanol/H2O series for 45 min each. After dehydration, samples were washed in 2:1 DCM:methanol overnight followed by three 1 h washes in 100% DCM. The samples were cleared overnight in dibenzyl ether (Sigma-Aldrich) and stored at room temperature in the dark in amber glass vials until imaging.
Light-sheet imaging
Whole, cleared brains were imaged on a light sheet microscope (Zeiss Z1) equipped with a ×5 EC plan neofluar NA 0.16 objective lens and ×20 Clr plan neofluar NA 1.0 objective lens. The images were collected with two 1,920 × 1,920 pixel sCMOS cameras and acquired using Zen Blue 3.1 LS (Zeiss). The samples were staged in a custom imaging chamber (Translucence Biosystems) filled with dibenzyl ether and illuminated from both sides by the laser light sheet (5× light sheet thickness, 13.08 µm; step size, 4.5 µm; ×20 light sheet thickness, 4.35 µm; step size, 0.61 µm).
Light-sheet image analysis
For cKO experiments, Imaris (v.9.9) was used to generate surfaces around the positive signal (calculated by thresholding) in the HA and streptavidin channels. The total volume of streptavidin signal divided by the total volume of HA signal was used to generate a relative network volume representing each brain. For whisker-trimming experiments, individual astrocytes were counted in FIJI (ImageJ v.1.54b) after background thresholding. The number of streptavidin-positive astrocytes was divided by the number of HA positive astrocytes to generate a network size value representing each brain.
For the analyses in Fig. 3, image tiles were stitched into full-volume 3D TIFF stacks using Stitchy v.1.15.0 (Translucence Biosystems)—proprietary software that performs automated tile registration by aligning overlapping regions to generate seamless volumetric reconstructions. Stitching was performed on Windows workstations with a sigmoidal blending algorithm applied across overlapping tiles to minimize boundary artifacts. Each reconstructed brain volume averaged approximately 280 GB in size. In some instances, the software reoriented spatial axes as needed to maintain a consistent anatomical orientation across samples.
Segmentation and quantification were performed using Voxels v.1.10.0 (client) and v.1.9.1 (server) (Translucence Biosystems), a cloud-ready image-analysis platform for large-scale processing. Raw fluorescence volumes were converted into 3D voxel-wise probability maps using a random-forest pixel classifier trained on manually annotated crops from seven representative brain volumes to classify voxels as signal of interest or background. Probability maps were refined with threshold- and morphology-based filters to generate binary total-signal masks at full imaging resolution, which were then used to exclude background voxels during downstream regional quantification and atlas registration.
Each sample was registered to the Allen Reference Atlas 2, based on the Allen Mouse Common Coordinate Framework (CCFv3), using the 647 nm fluorescence channel as the reference. The resulting full-resolution total-signal masks were resampled to the atlas voxel resolution for non-linear registration, preserving segmentation fidelity and quantitative accuracy. After alignment, masked signal volumes were partitioned by the midline into left and right hemispheres to assess potential lateralization.
A hierarchical atlas annotation encompassing >500 regions enabled multiscale quantification, allowing integrated intensity values to be aggregated from fine substructures (for example, dentate gyrus) to parent structures (for example, hippocampal formation) and to whole-hemisphere levels. All samples met quality-control criteria for registration accuracy, confirmed by visual inspection of anatomical landmarks relative to the reference atlas and annotation boundaries.
Statistical analysis and reproducibility
Data are presented as mean ± s.d. Mice were assigned at random to groups. Experiments were not performed in a blinded manner. Sample sizes were determined through reference to previous literature. Statistical significance for each experiment was determined as described in the respective figure legend. In the figure legends, n values represent the number of biologically independent animals per group. For data in all figures other than Fig. 3 and Extended Data Fig. 8, statistical analyses were performed in Prism 10 software (GraphPad). For Fig. 3 and Extended Data Fig. 8, regions smaller than 10,000 voxels were excluded owing to potential variance in atlas registration. Student’s t-tests assess regional differences and the Benjamini–Hochberg procedure was used to control the false discovery rate. No datapoints were excluded.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Owing to size constraints and hosting difficulties, light-sheet data are available from the corresponding authors. Source data are provided with this paper.
References
Hosli, L. et al. Decoupling astrocytes in adult mice impairs synaptic plasticity and spatial learning. Cell Rep. 38, 110484 (2022).
Frisch, C. et al. Mice with astrocyte-directed inactivation of connexin43 exhibit increased exploratory behaviour, impaired motor capacities, and changes in brain acetylcholine levels. Eur. J. Neurosci. 18, 2313–2318 (2003).
Murphy-Royal, C. et al. Stress gates an astrocytic energy reservoir to impair synaptic plasticity. Nat. Commun. 11, 2014 (2020).
Pannasch, U. et al. Astroglial networks scale synaptic activity and plasticity. Proc. Natl Acad. Sci. USA 108, 8467–8472 (2011).
Pannasch, U. et al. Connexin 30 sets synaptic strength by controlling astroglial synapse invasion. Nat. Neurosci. 17, 549–558 (2014).
Chever, O., Dossi, E., Pannasch, U., Derangeon, M. & Rouach, N. Astroglial networks promote neuronal coordination. Sci. Signal. 9, ra6 (2016).
Ribot, J. et al. Astrocytes close the mouse critical period for visual plasticity. Science 373, 77–81 (2021).
Ackerman, S. D., Perez-Catalan, N. A., Freeman, M. R. & Doe, C. Q. Astrocytes close a motor circuit critical period. Nature 592, 414–420 (2021).
Chi, J., Crane, A., Wu, Z. & Cohen, P. Adipo-Clear: a tissue clearing method for three-dimensional imaging of adipose tissue. J. Vis. Exp. https://doi.org/10.3791/58271 (2018).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Cooper, M. L. et al. Redistribution of metabolic resources through astrocyte networks mitigates neurodegenerative stress. Proc. Natl Acad. Sci. USA 117, 18810–18821 (2020).
Rouach, N., Koulakoff, A., Abudara, V., Willecke, K. & Giaume, C. Astroglial metabolic networks sustain hippocampal synaptic transmission. Science 322, 1551–1555 (2008).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
Rayaprolu, S. et al. Cell type-specific biotin labeling in vivo resolves regional neuronal and astrocyte proteomic differences in mouse brain. Nat. Commun. 13, 2927 (2022).
Takano, T. et al. Chemico-genetic discovery of astrocytic control of inhibition in vivo. Nature 588, 296–302 (2020).
Jiang, R., Haustein, M. D., Sofroniew, M. V. & Khakh, B. S. Imaging intracellular Ca2+ signals in striatal astrocytes from adult mice using genetically-encoded calcium indicators. J. Vis. Exp. https://doi.org/10.3791/51972 (2014).
Nagai, J. et al. Specific and behaviorally consequential astrocyte Gq GPCR signaling attenuation in vivo with ibetaARK. Neuron 109, 2256–2274 (2021).
Shigetomi, E. et al. Imaging calcium microdomains within entire astrocyte territories and endfeet with GCaMPs expressed using adeno-associated viruses. J. Gen. Physiol. 141, 633–647 (2013).
Lee, H. J. et al. Conformational changes in the human Cx43/GJA1 gap junction channel visualized using cryo-EM. Nat. Commun. 14, 931 (2023).
Damstra, H. G. J. et al. Ten-fold robust expansion microscopy. Bio Protoc. 13, e4698 (2023).
Foo, L. C. Purification of rat and mouse astrocytes by immunopanning. Cold Spring Harb. Protoc. 2013, 421–432 (2013).
Mair, A. & Bergmann, D. C. Advances in enzyme-mediated proximity labeling and its potential for plant research. Plant Physiol. 188, 756–768 (2022).
Cho, K. F. et al. Proximity labeling in mammalian cells with TurboID and split-TurboID. Nat. Protoc. 15, 3971–3999 (2020).
Chiaruttini, N. et al. ABBA+BraiAn, an integrated suite for whole-brain mapping, reveals brain-wide differences in immediate-early genes induction upon learning. Cell Rep. 44, 115876 (2025).
Hasel, P. et al. Defining the molecular identity and morphology of glia limitans superficialis astrocytes in vertebrates. Cell Rep. 44, 115344 (2025).
Weber, P. A., Chang, H. C., Spaeth, K. E., Nitsche, J. M. & Nicholson, B. J. The permeability of gap junction channels to probes of different size is dependent on connexin composition and permeant-pore affinities. Biophys. J. 87, 958–973 (2004).
Alarcon-Martinez, L. et al. Interpericyte tunnelling nanotubes regulate neurovascular coupling. Nature 585, 91–95 (2020).
Krolak, T. et al. Brain endothelial gap junction coupling enables rapid vasodilation propagation during neurovascular coupling. Cell 188, 5003–5019 (2025).
Renier, N. et al. A mutant with bilateral whisker to barrel inputs unveils somatosensory mapping rules in the cerebral cortex. eLife 6, e23494 (2017).
Margolis, D. J. et al. Reorganization of cortical population activity imaged throughout long-term sensory deprivation. Nat. Neurosci. 15, 1539–1546 (2012).
Kelly, M. K., Carvell, G. E., Kodger, J. M. & Simons, D. J. Sensory loss by selected whisker removal produces immediate disinhibition in the somatosensory cortex of behaving rats. J. Neurosci. 19, 9117–9125 (1999).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Sellien, H. & Ebner, F. F. Rapid plasticity follows whisker pairing in barrel cortex of the awake rat. Exp. Brain Res. 177, 1–14 (2007).
Saccheri, F. et al. Bacteria-induced gap junctions in tumors favor antigen cross-presentation and antitumor immunity. Sci. Transl. Med. 2, 44ra57 (2010).
Harris, A. L. Emerging issues of connexin channels: biophysics fills the gap. Q. Rev. Biophys. 34, 325–472 (2001).
Valiunas, V., Cohen, I. S. & Brink, P. R. Defining the factors that affect solute permeation of gap junction channels. Biochim. Biophys. Acta Biomembr. 1860, 96–101 (2018).
Le, H. T. et al. Gap junction intercellular communication mediated by connexin43 in astrocytes is essential for their resistance to oxidative stress. J. Biol. Chem. 289, 1345–1354 (2014).
Spray, D. C. et al. Gap junctions and bystander effects: good samaritans and executioners. Wiley Interdiscip. Rev. Membr. Transp. Signal. 2, 1–15 (2013).
Xie, L. et al. Sleep drives metabolite clearance from the adult brain. Science 342, 373–377 (2013).
Lee, S. N. et al. Cryo-EM structures of human Cx36/GJD2 neuronal gap junction channel. Nat. Commun. 14, 1347 (2023).
Wang, M. et al. Connexin43 in neonatal excitatory neurons is important for short-term motor learning. Brain Res. 1720, 146287 (2019).
Chang, Q. & Balice-Gordon, R. J. Gap junctional communication among developing and injured motor neurons. Brain Res. Brain Res. Rev. 32, 242–249 (2000).
Chang, Q., Pereda, A., Pinter, M. J. & Balice-Gordon, R. J. Nerve injury induces gap junctional coupling among axotomized adult motor neurons. J. Neurosci. 20, 674–684 (2000).
Cheung, G. et al. Physiological synaptic activity and recognition memory require astroglial glutamine. Nat. Commun. 13, 753 (2022).
Shandra, O. et al. Repetitive diffuse mild traumatic brain injury causes an atypical astrocyte response and spontaneous recurrent seizures. J. Neurosci. 39, 1944–1963 (2019).
Munoz-Ballester, C. et al. Astrocytic connexin43 phosphorylation contributes to seizure susceptibility after mild traumatic brain injury. Preprint at bioRxiv https://doi.org/10.1101/2024.11.12.623104 (2025).
Cotrina, M. L. et al. Astrocytic gap junctions remain open during ischemic conditions. J. Neurosci. 18, 2520–2537 (1998).
Onodera, M. et al. Exacerbation of epilepsy by astrocyte alkalization and gap junction uncoupling. J. Neurosci. 41, 2106–2118 (2021).
Takatsuru, Y. et al. Critical role of the astrocyte for functional remodeling in contralateral hemisphere of somatosensory cortex after stroke. J. Neurosci. 33, 4683–4692 (2013).
Cooper, M. L. et al. Astrocytes in the mouse brain respond bilaterally to unilateral retinal neurodegeneration. Proc. Natl Acad. Sci. USA 122, e2418249122 (2025).
Beardslee, M. A., Laing, J. G., Beyer, E. C. & Saffitz, J. E. Rapid turnover of connexin43 in the adult rat heart. Circ. Res. 83, 629–635 (1998).
Laing, J. G. & Beyer, E. C. The gap junction protein connexin43 is degraded via the ubiquitin proteasome pathway. J. Biol. Chem. 270, 26399–26403 (1995).
Lawal, O., Ulloa Severino, F. P. & Eroglu, C. The role of astrocyte structural plasticity in regulating neural circuit function and behavior. Glia 70, 1467–1483 (2022).
Mori, T. et al. Inducible gene deletion in astroglia and radial glia—a valuable tool for functional and lineage analysis. Glia 54, 21–34 (2006).
Boulay, A. C. et al. Hearing is normal without connexin30. J. Neurosci. 33, 430–434 (2013).
Theis, M. et al. Accelerated hippocampal spreading depression and enhanced locomotory activity in mice with astrocyte-directed inactivation of connexin43. J. Neurosci. 23, 766–776 (2003).
Bansal, R. & Pfeiffer, S. E. Reversible inhibition of oligodendrocyte progenitor differentiation by a monoclonal antibody against surface galactolipids. Proc. Natl Acad. Sci. USA 86, 6181–6185 (1989).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Acknowledgements
We thank the members of the Microscopy Laboratory at NYU Grossman School of Medicine for experimental and technical support; G. Bertoli for her advice on expansion microscopy and the course directors of the Kavli iDISCO course at Rockefeller University for their expertise in tissue clearing; the staff at the NYU Grossman School of Medicine Genotyping Core Laboratory and the Advanced Rodent Transgenic Laboratory for their assistance with colony rederivation and genetic maintenance; E. Katafygiotou, B. S. Balza, J. M. Santos and A. Amponsah for their assistance with tissue clearing; J. Minder for her cloning and surgical expertise; S. Peron for his advice on working with whisker barrel cortex; P. Glimcher, S. Davis Jr and F. X. Castellanos for proofreading and comments on the manuscript. We acknowledge the Leon Levy Fellowship in Neuroscience (M.L.C.), the CureAlzheimer’s Fund (S.A.L.), the PEW Charitable Trusts Postdoctoral Fellowship (M.C.S.), the Simons Foundation SURFiN Fellowship (K.E.C., M.L.C. and M.V.C.), the Brain & Behavior Research Foundation Young Investigator Award (M.L.C.), the Belfer Neurodegeneration Consortium (S.A.L.), the Carol and Gene Ludwig Family Foundation (S.A.L.), the Swiss National Science Foundation (320030-232028, A.S.S.), and the NIH (K99 NS139313 (M.L.C.), K00 AG068343 (H.K.G.), U19 NS107616 (M.V.C.), T32 MH019524 (M.L.C.), P30 AG066512 (M.L.C.) and R01 EY033353 (S.A.L.)). NYU shared resources are partially supported by the Cancer Center Support Grant P30CA016087 at the Laura and Isaac Perlmutter Cancer Center.
Author information
Authors and Affiliations
Contributions
M.L.C. designed research. S.A.L. and M.V.C. supervised the work. M.L.C., M.C.S., M.C., H.K.G., J.S. and K.E.C. performed research. M.L.C. and A.S.S. contributed new reagents/analytic tools. M.L.C., C.R., P.C. and D.G.W. analysed the data. M.L.C. visualized the data. M.L.C. wrote the paper. M.L.C., M.C.S., M.C., C.R., H.K.G., K.E.C., A.S.S., S.A.L. and M.V.C. edited the paper. M.L.C., M.C.S., H.K.G., S.A.L. and M.V.C. acquired funding.
Corresponding authors
Ethics declarations
Competing interests
S.A.L. maintains a financial interest in AstronauTx and Synapticure. S.A.L. is on the scientific advisory board of the Global BioAccess Fund. C.R., P.C. and D.G.W. are employed by Translucence Biosystems.
Peer review
Peer review information
Nature thanks David Lyons and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Primary immunopanned astrocyte purity and knockout induction.
a. qPCR of marker gene expression for astrocytes (Gja1, Aldh1l1, Slc1a3), oligodendrocytes (Sox10, Mog), microglia (Aif1, Cd14), and neurons (Tubb3, Syp) in primary immunopanned astrocytes from rat or cKO mouse normalized to the ribosomal gene RPL19 and relative to naïve cortex from simultaneously dissected littermate controls. b. Primary cKO mouse astrocyte viability determined as the ratio of Hoechst positive, Propidium Iodine (PI) negative cells to the total number of Hoechst positive cells. Viability was significantly decreased at 4-Hydroxytamoxifen (4-OHT) concentrations of 1:2000 and above (p < 0.001). c. qPCR of Gja1(Cx43) and Gjb6(Cx30) in primary immunopanned cKO astrocytes at a 4-OHT concentrations of 1:5000, 1:4000, and 1:3000 relative to naïve cells. All 4-OHT concentrations decreased expression of both RNAs, with the largest decrease at 1:3000. d. Quantification of western blots from immunopanned cKO astrocytes in the following conditions: Naïve, Naïve + Vehicle (EtOH, 1:3000), 4-OHT 1:6000, 4-OHT 1:5000, 4-OHT 1:4000, and 4-OHT 1:3000. Blots were costained for Cx43 and Actin (example on right; all quantified blots in Extended Data Fig. 1). Cx43/Actin normalized to Naïve significantly decreased at all 4-OHT concentrations, with the largest decrease at 1:3000. n = 3 independent isolations for all panels. Statistical significance was determined by repeated measures one-way ANOVAs (p values indicated on graphs) followed by Dunnett’s multiple comparisons test (α = 0.05). Values are mean ± standard deviation.
Extended Data Fig. 2 Both AAV5-GfaABC1D-Cx43:TID:HA and biotin are required for astrocyte network detection.
a. (also in Fig. 1e) Primary rat astrocytes (GFAP, green) infected with AAV5-GfaABC1D-Cx43:TID:HA and given biotin have punctate staining patterns for HA-tag in about 20% of cells; ~80% of cells are positive for streptavidin. b. Without supplemented biotin, HA-tag is detected but not streptavidin. Without AAV (c) or with neither AAV nor biotin (d), neither HA-tag nor streptavidin are detected. e. Primary rat astrocytes were incubated with a range of MOIs (multiplicity of infection) to estimate the viral load necessary for ~10% of astrocytes to express Cx43-TID. Across each MOI tested, about 80~ of astrocytes were positive for biotinylated fluxed molecules. n = 2 isolations f. 87.7 ± 3.0% of cells from the AAV + /Biotin+ condition (MOI = 1 ×104) were positive for streptavidin; the other 3 conditions contained no detectable streptavidin signal. g. 13.3 ± 0.8% of cells were positive for HA-tag in both AAV+ conditions; neither AAV- condition contained detectable HA-tag immunopositivity. n = 3 isolations. Statistical significance (p values for all comparisons < 0.0001) determined via one-way ANOVA followed by Tukey’s multiple comparisons test (α = 0.05).
Extended Data Fig. 3 Genetically-expressed Cx43 is necessary for biotinylated molecules to distribute beyond infected cells.
a. Primary astrocytes isolated from Slc1a3:Cre-ERT2+/+ x Gja1fl/fl x Gjb6fl/fl mice (GFAP, green) demonstrate distributed streptavidin staining and punctate HA-tag staining when infected with AAV5-GfaABC1D-Cx43:TID:HA and given biotin. b. Only HA-tag is visible when AAV is given without biotin. Without AAV (c) or with neither AAV nor biotin (d), neither HA-tag nor streptavidin are detected. e. Astrocytes induced for Cx43 and Cx30 knockout via 4-OHT are only positive for streptavidin in HA-tag positive cells in the AAV and biotin condition. As in b–d, only HA-tag is visible when AAV is given without biotin (f). Without AAV (g) or with neither AAV nor biotin (h), neither HA-tag nor streptavidin are detected. i. Primary mouse astrocytes isolated from uninduced cKO mice were incubated with a range of MOIs to estimate the viral load necessary for ~10% of astrocytes to express Cx43-TID. Across each MOI tested other than 1 ×103, about 80% of astrocytes were positive for biotinylated fluxed molecules; for 1 ×103, ~60% of astrocytes were streptavidin positive. j. 85.2 ± 1.6% of cells from the control AAV+ and Biotin+ condition (MOI = 1 ×104) were positive for streptavidin; the other 3 control conditions contained no detectable streptavidin signal. For cKO astrocytes induced with 4-OHT, 15.7 ± 2.6% of cells from the AAV+ and Biotin+ condition were positive for streptavidin; the other 3 cKO condition contained no detectable streptavidin signal. k. ~ 10% of cells were positive for HA-tag in all AAV+ conditions (control ++ 10.3 ± 1.2%; control +- 10.4 ± 1.1%; cKO ++ 10.3 ± 1.7%; cKO +- 10.4 ± 0.8%); no AAV- condition contained detectable HA-tag immunopositivity. n = 3 isolations for both j and k. Statistical significance (p values for all comparisons < 0.0001) determined via one-way ANOVA followed by Tukey’s multiple comparisons test (α = 0.05). Values are mean ± standard deviation.
Extended Data Fig. 4 Infected cells exhibit astrocytic morphology and punctate HA-tag staining.
a) Three-dimensional rendering of infected cells in motor cortex stained for streptavidin (magenta) and HA-tag (cyan). b) Single slice shows neighbouring infected and uninfected cells. All cells demonstrate astrocytic morphology when filled with biotinylated molecules (streptavidin, magenta); infected cells exhibit a punctate pattern of HA-tag staining (cyan) on the surface of each cell indicative of gap junctional localization.
Extended Data Fig. 5 Motor Cortex Network Reproducibility.
Light sheet imaged brains from 3 independent mice (columns) shown in dorsal (top row), sagittal (middle row), and ventral/oblique views (bottom row). The infection site is stained for HA-tag (cyan), the astrocyte network originating in the infection site is stained with streptavidin (magenta), and an empty channel’s autofluorescence (grey) shows the outline of the brain. See also Supplemental Video 4.
Extended Data Fig. 6 Hypothalamic Network Reproducibility.
Light sheet imaged brains from 3 independent mice (columns) shown in dorsal (top row), sagittal (middle row), and ventral/oblique views (bottom row). The infection site is stained for HA-tag (cyan), the astrocyte network originating in the infection site is stained with streptavidin (magenta), and an empty channel’s autofluorescence (grey) shows the outline of the brain.
Extended Data Fig. 7 PFC Network Reproducibility.
Light sheet imaged brains from 3 independent mice (columns) shown in dorsal (top row), sagittal (middle row), and ventral/oblique views (bottom row). The infection site is stained for HA-tag (cyan), the astrocyte network originating in the infection site is stained with streptavidin (magenta), and an empty channel’s autofluorescence (grey) shows the outline of the brain.
Extended Data Fig. 8 Astrocyte network specificity (expanded).
Euler diagrams showing the number of significantly enriched regions (corresponding boxes) in brains injected in either motor cortex (yellow), prefrontal cortex (blue), or hypothalamus (pink) for ipsilateral (top) or contralateral (bottom) hemispheres.
Extended Data Fig. 9 Astrocyte networks link specific brain regions.
Virtual coronal sections of light sheet imaged brains infected in motor cortex (left column), hypothalamus (middle column), or prefrontal cortex (right column). Sections from each brain are arranged from rostral to caudal (top to bottom images) and were selected as representative examples of robust astrocyte network connections. Brain regions are outlined using an overlay of the Allen Brain Atlas (fitted via the FIJI ABBA plugin24; 3D mouse Allen Brain Atlas, mouse.brain-map.org and atlas.brain-map.org).
Extended Data Fig. 10 Diversity of cell types within a motor cortex astrocyte network.
a, b, c. Individual slices at varied depths in the horizontal plane through a light sheet imaged brain infected in motor cortex (yellow lines in reference 3D image, top right; HA, cyan; Streptavidin, magenta). a’. Biotinylated compounds fill many, but not all, adjacent cells. a”. While many brain regions show signal bilaterally, some regions, such as prefrontal cortex in this brain, do not. b’. Following motor cortex infection, sparse fluorescence in some neurons is visible. b”. Chains of astrocytes contact and diverge from neuronal axons, which qualitatively become brighter following astrocyte network contact points. b”’. Most astrocyte network termini, such as this terminus in prefrontal cortex, do not show detectable streptavidin signal among adjacent neurons. c’. Following motor cortex infection, axons descending to the spinal cord show visible streptavidin signal.
Extended Data Fig. 11 Chains of fibrous astrocytes interconnect bilateral astrocyte networks.
Three-dimensional rendering (horizontal and oblique) of streptavidin-positive astrocytes in corpus callosum following infection in visual cortex.
Extended Data Fig. 12 Cortical astrocyte networks can contain both parenchymal and glia limitans superficialis astrocytes on the brain’s surface.
a. Three-dimensional rendering of streptavidin positive astrocytes on the surface of the brain (top) connected to astrocytes in the parenchyma. b. Slice (marked in a) showing glia limitans superficialis astrocytes on the brain’s surface. c. Slice (marked in a) showing the parenchymal astrocyte in contact with the glia limitans superficialis astrocytes.
Extended Data Fig. 13 Whisker trim reduces astrocyte network size.
Z-projections of all fluorescent cells in animals infected to express Cx43-TID-HA (cyan) in barrel cortex, resulting in biotinylation of fluxed molecules in an astrocyte network (magenta). Small-volume (20 nl) injections into barrel cortex corresponding to naïve (a) or trimmed whiskers (b) resulted in cells labelled at a sparse enough density to be counted and quantified (see Fig. 5c).
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1: western blot data.
Supplementary Video 1 (download MP4 )
Motor cortex astrocyte network. Three-dimensional video rendering of light sheet-imaged brain infected in motor cortex. The video contains two subsequent renderings. The first shows the infected region (HA tag, cyan) inside of the autofluorescence capture (grey) for context; this is first shown completely and then slices are removed dorsal to ventral. The second shows the same infected region within the streptavidin-stained astrocyte network originating from that brain region (magenta); this was first built in slices from ventral to dorsal and then rotated.
Supplementary Video 2 (download MP4 )
Hypothalamic astrocyte network. Three-dimensional video rendering of light sheet-imaged brain infected in hypothalamus. The video contains two subsequent renderings. The first shows the infected region (HA tag, cyan) inside of the autofluorescence capture (grey) for context; this is first shown completely and then slices are removed dorsal to ventral. The second shows the same infected region within the streptavidin-stained astrocyte network originating from that brain region (magenta); this first built in slices from ventral to dorsal, and then rotated.
Supplementary Video 3 (download MP4 )
Prefrontal cortex astrocyte network. Three-dimensional video rendering of light sheet-imaged brain infected in prefrontal cortex. The video contains two subsequent renderings. The first shows the infected region (HA-tag, cyan) inside of the autofluorescence capture (grey) for context; this is first shown completely and then slices are removed dorsal to ventral. The second shows the same infected region within the streptavidin-stained astrocyte network originating from that brain region (magenta); this is first built in slices from ventral to dorsal, and then rotated.
Supplemental Video 4 (download MP4 )
Astrocyte networks are consistent among mice. Sequential horizontal and coronal virtual sections from 3 independent mice, each injected in motor cortex (samples also shown in Extended Data Fig. 5). The autofluorescence channel (sample 1) and its registration are included for reference.
Supplemental Video 5 (download MP4 )
Astrocyte networks differ. Coronal-atlas-mapped heat maps demonstrating the streptavidin signal in each brain region as the average of all mice injected in either motor cortex (left), prefrontal cortex (middle) or hypothalamus (right).
Supplemental Video 6 (download MP4 )
Interconnected astrocytes in motor cortex contralateral to injection site. z stack of streptavidin-containing (magenta) astrocytes in motor cortex, contralateral to the infection site. Chains of astrocytes contacting one another are visible as the z stack moves dorsal to ventral.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cooper, M.L., Selles, M.C., Cammer, M. et al. Astrocytes connect specific brain regions through plastic networks. Nature (2026). https://doi.org/10.1038/s41586-026-10426-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10426-6