Abstract
Cell-tagging strategies with DNA barcodes have enabled the analysis of clone size dynamics and clone-restricted transcriptomic landscapes in heterogeneous populations. However, isolating a target clone that displays a specific phenotype from a complex population remains challenging. Here we present a multi-kingdom genetic barcoding system, CloneSelect, which enables a target cell clone to be triggered to express a reporter gene for isolation through barcode-specific CRISPR base editing. In CloneSelect, cells are first stably tagged with DNA barcodes and propagated so that their subpopulation can be subjected to a given experiment. A clone that shows a phenotype or genotype of interest at a given time can then be isolated from the initial or subsequent cell pools stored during the experiment using CRISPR base editing. CloneSelect is scalable and compatible with single-cell RNA sequencing. We demonstrate the versatility of CloneSelect in human embryonic kidney 293T cells, mouse embryonic stem cells, human pluripotent stem cells, yeast cells and bacterial cells.
Similar content being viewed by others
Main
Cells are not homogeneous in any system. Although they proliferate and replicate their genome, which encodes molecular regulatory programmes in their progeny, they also change their states in response to dynamic gene expression patterns and environmental signals. As typically shown in multicellular organisms, cells self-organize through mutual molecular and mechanical communications, dynamically creating complex structures. During these processes, spontaneous mutations in the genome may impair the cellular programme, leading to cellular malfunction. Other mutations may confer growth advantages to the cells, which can be either beneficial or catastrophic to the system.
For example, during cancer chemotherapy, resistant clones can arise and expand, contributing to cancer recurrence and metastasis1,2,3,4. In laboratory microbial evolution experiments, different cells within the initial population dynamically expand and shrink their clone sizes by acquiring new mutations over multiple generations5,6,7,8,9. There are also other examples in which the contribution of a genetic factor is unclear. In hematopoiesis, stem cells presenting an analytically indistinguishable set of cell surface markers show fate-restricted differentiation patterns, in which some cells seem to be primed for specific lineages by factors10,11,12,13. Similarly, in vitro stem cell differentiation and direct reprogramming experiments have demonstrated that ‘elite’ clones reproducibly transform into target cell states14,15,16.
These views on clonal heterogeneity and cell lineage bias have been rapidly shaped by cell clone tracing, whereby a library of short DNA sequences is introduced into a cell population to uniquely tag individual cells by a stable integration approach, such as lentiviral transduction. The change in abundance of the barcoded clones can be traced by subsampling the cell population over time and quantifying the DNA barcodes by PCR and deep sequencing. Furthermore, a transcribing barcode system with single-cell RNA sequencing (scRNA-seq) allows clonal lineages to be analyzed alongside cell states, revealing cell lineage-restricted state trajectories14,17,18,19,20,21,22,23,24. However, these methods are limited in their ability to analyze diverse molecular and environmental factors that derive specific fate outcomes. Flow cytometry cell sorting with immunostaining and emerging image cytometry cell sorting technologies enable the dissociation of heterogeneous cell populations into single cells with their observed phenotypes25,26,27,28 but cannot do the same for a population of clones before they exhibit a phenotype of interest.
Whether the chemotherapy-resistant clones existed in the initial cell population with the genetic mutations, whether molecular factors underlie the observed stem cell differentiation fate and whether the progression of the specific clone is conditional on the existence of any other clones are unanswered questions. To tackle these questions, the concept of ‘retrospective clone isolation’ has recently emerged29,30,31,32, in which a barcoded cell population is first propagated and its subpopulation is subjected to a given assay (Fig. 1a). After identifying a barcoded clone of interest, the same clone (or its close relative) is isolated in a barcode-specific manner from the initial or any other subpopulation stored during the experiment. The isolated live clone can then be subjected to various biological experiments, including omics measurements and the reconstitution of a synthetic cell population with the isolates.
a, Conceptual diagram of retrospective clone isolation. b, Different barcode-specific gRNA-dependent reporter activation circuits. CloneSelect C→T, CaTCH and ClonMapper. c, Barcode-dependent reporter activation of six barcoded cell lines by CloneSelect C→T. BC, barcode. Scale bar, 50 µm. d, Performance comparison of CloneSelect C→T, CaTCH and ClonMapper across the same set of barcode–gRNA pairs. Receiver operating characteristic curves were obtained by varying EGFP intensity thresholds for each target. The dashed line indicates the expected random classification. e, Percent positive cells with a uniform EGFP intensity gate applied to all of the tested systems (n = 3). Two-tailed Welch’s t-test was used for statistical analysis. f, Fold change between percent EGFP+ cells of OT (on-target) and NT (non-target) gRNA-barcode pairs for each barcoded cell line. Two-tailed Mann–Whitney U-test was used for statistical analysis. g, CloneSelect A→G. h, Barcode-dependent reporter activation of three barcoded cell lines by CloneSelect A→G. Scale bar, 50 µm. i, Performance comparison of CloneSelect A→G, CaTCH and ClonMapper across the same set of barcode–gRNA pairs. j, Percent positive cells with a uniform EGFP intensity gate applied to all of the tested systems (n = 3). Two-tailed Welch’s t-test was used for statistical analysis. k, Comparison of CloneSelect C→T and CloneSelect A→G. Activated cell frequencies of each barcoded cell sample by OT and NT gRNA queries were normalized by that of the cell sample with the same OT barcode–gRNA pair for CaTCH, ClonMapper or low-copy CRISPRa. The sample sizes for C→T barcodes and A→G barcodes were 18 and 9, respectively. Two-tailed Mann–Whitney U-test was used for statistical analysis; n.s., not significant; *P < 0.05; **P < 0.01; ***P < 0.001.
Most retrospective clone isolation methods have been implemented using CRISPR–Cas9 system. In the CRISPR activation (CRISPRa)-based approach29,30,31,32, cells are tagged with DNA barcodes upstream of a fluorescent reporter gene with an insufficient minimum promoter. Once a barcoded clone is identified for isolation, a CRISPR guide RNA (gRNA) targeting its barcode is introduced to the cell population with catalytically dead Cas9 (dCas9) fused to a transcriptional activator(s)33. As the fluorescent reporter is expressed in a barcode-specific manner, the cells with the same barcode can be isolated by flow cytometry cell sorting. Alternative approaches with the inversed configuration have been developed in which gRNAs with different sequences are used as barcodes, and a CRISPRa reporter containing a target sequence is introduced for barcode-specific fluorescent reporter activation30,31. These approaches, however, suffer from the leaky expression of a reporter without the targeting gRNA. Furthermore, CRISPRa-based retrospective clone isolation has only been demonstrated in mammalian cell systems.
Genetic circuits based on DNA code alteration generally show highly specific input responses. Although any inducible gene expression system may exhibit leakage in the absence of a gRNA, a circuit using genetic code alteration through genome editing cannot easily leak output signals without the intended genome editing. The wild-type Cas9 has also been used to establish a retrospective clone isolation method32. In this approach, a barcode is placed upstream of a reporter gene with a stop codon and an out-of-frame start codon. The reporter translation can be stochastically activated by Cas9-induced double-stranded DNA break and deletion through non-homologous end-joining DNA repair to remove the stop codon and bring the start codon into the coding frame. Although this does not usually show unexpected reporter activation for non-target clones, the system’s sensitivity relies on stochastic events, which creates a bottleneck in efficiency. Cas9-induced double-stranded DNA break has also been known to be cytotoxic and can potentially damage the target clone during the reporter activation procedure34,35.
Another approach has been proposed using RNA fluorescence in situ hybridization, in which cells expressing RNA barcodes are first fixed. A target clone is labelled with a fluorophore probe targeting its barcode RNA transcripts and isolated by cell sorting36. Although RNA fluorescence in situ hybridization is specific and sensitive, the isolated cells are fixed and cannot be used for further analyses that require live cells.
Here, we report a new CRISPR base editing approach, CloneSelect, that overcomes the technical limitations mentioned above. CloneSelect is based on restoring reporter protein translation by base editing of an impaired start codon or removing an upstream stop codon in a barcode-specific manner. The new method is highly scalable, programmable and compatible with scRNA-seq. Its specificity surpasses other CRISPR-based systems. We present the versatility of the method in human embryonic kidney (HEK) 293T cells, mouse embryonic stem (ES) cells, human pluripotent stem (PS) cells, yeast cells and bacterial cells.
Results
Mammalian CloneSelect
CRISPR base editing has been widely used to induce a single nucleotide substitution at a target genomic site without a double-stranded DNA break37. We first reasoned that a C→T base editor-based circuit38 would enable highly sensitive, precise barcode-specific clone isolation with better performance than the previous CRISPR-based methods. In CloneSelect C→T, a barcode is encoded immediately upstream of a reporter gene whose start codon is mutated to GTG (Fig. 1b, left). The reporter-encoding region is transcribed by a constitutively active promoter, but the impaired start codon renders its translation inactive. The C→T base editing on the antisense strand of the target barcode enables the first guanine of GTG (cytosine on its antisense strand) to be substituted into adenine (thymine on its antisense strand), restoring ATG and reporter translation. To achieve this outcome, we used Target-AID, which we previously developed as one of the first-generation C→T base editors that is highly active in mammalian cells and has a narrow C→T editing window in the gRNA target site35,37.
We first compared the performance of CloneSelect C→T, using enhanced green fluorescent protein (EGFP) as a reporter, with three other CRISPRa-based systems reported previously (CaTCH, ClonMapper and CaTCH alternative)29,31 and two other setups (low-copy CRISPRa and high-copy CRISPRa) that we prepared for this study. CaTCH (Fig. 1b, middle) and low-copy CRISPRa (Supplementary Fig. 1a, left) use the single-copy integration of barcode reporters for cell barcoding and the transfection of a gRNA for the target cell’s reporter activation. ClonMapper (Fig. 1b, right), CaTCH alternative (Supplementary Fig. 1a, middle) and high-copy CRISPRa (Supplementary Fig. 1a, right) take the inversed configuration, whereby a gRNA library is used to barcode cells in a population and a target reporter is introduced to the cell population by transfection for fluorescent activation. In this comparative analysis, we replaced a fluorescent marker downstream of dCas9-VPR in CaTCH with hygromycin to enable drug selection similar to the other systems for establishing cell lines.
The different retrospective clone isolation systems were tested using a common set of six orthogonal barcode–gRNA pairs that were randomly selected. The barcoding reagents were first individually introduced to HEK293T cells by lentivirus transduction with an infection rate of <0.1, ensuring the multiplicity of infection to be one (a single barcode per cell), and each barcoded cell sample was transfected with different fluorescent activation reagents (Fig. 1c and Supplementary Fig. 1b). In each reporter activation experiment, we analyzed EGFP intensities of single cells using flow cytometry (Supplementary Fig. 2a), obtained a true positive rate among expected positives and a false positive rate among expected negatives at each EGFP intensity threshold and generated a receiver operating characteristic curve (Fig. 1d and Supplementary Fig. 2b). When the false positive rates were all set to 0.5%, the true positive rates were 10.05–24.88%, 6.84–12.50%, 2.71–22.37%, 0.00–5.46%, 0.00% and 0.00% for CloneSelect C→T, CaTCH, low-copy CRISPRa, ClonMapper, CaTCH alternative and high-copy CRISPRa, respectively (Fig. 1d and Supplementary Fig. 3a). Given that it is practically impossible to find the best-performing EGFP intensity threshold for a given target barcode, we also arbitrarily selected a universal threshold for the different systems based on the signal intensity from the EGFP channel in negative control cells. With this threshold, the true positive rates and false positive rates were 2.39–20.74% and 0.00–0.62%, respectively, for CloneSelect C→T, 12.21–28.17% and 0.97–13.95%, respectively, for CaTCH, 10.27–21.48% and 0.09–5.54%, respectively, for low-copy CRISPRa, 10.94–23.23% and 2.48–14.08%, respectively, for ClonMapper, 60.66–80.00% and 42.45–73.09%, respectively, for CaTCH alternative and 59.31–88.45% and 49.30–88.27%, respectively, for high-copy CRISPRa (Fig. 1e and Supplementary Fig. 3b). The EGFP intensities of EGFP+ cells across different systems did not show marked differences (Supplementary Fig. 3c). Overall, we found that CloneSelect C→T performed the best in activating the expected barcode–gRNA pairs while minimizing the false positives in both metrics. Among the CRISPRa-based systems, the reporter-based barcoding systems were overall better than the gRNA-based barcoding systems in the receiver operating characteristic curve analysis, but ClonMapper showed comparable performance to the reporter-based barcoding systems when using the universal threshold (Fig. 1f).
Although it exhibits high efficiency and specificity in human cells, CloneSelect C→T is not applicable for clone isolation of several other eukaryotic and prokaryotic species, as GTG can be used as a non-canonical start codon39. Therefore, we designed another system, CloneSelect A→G, using an adenine base editor, ABE-7.10, that induces A→G base substitution at the gRNA target sequence40. In CloneSelect A→G, following a constitutively active promoter and a start codon, a barcode encoding a TAA stop codon prevents downstream reporter translation (Fig. 1g). The stop codon can be altered in a gRNA-dependent manner by mutating the antisense strand from thymine (adenine) to guanine, converting the stop codon into CAA (proline). Using another common set of three barcode–gRNA pairs, we compared the performance of CloneSelect A→G and the other five CRISPRa-based systems (Fig. 1h and Supplementary Fig. 4a–c). Similarly to CloneSelect C→T, CloneSelect A→G enabled tight activations of target barcoded cells with a minimal false positive level. When the false positive rate was set to 0.5%, the true positive rates were 14.12–35.19%, 7.50–17.42%, 1.92–6.60%, 0.00%, 0.00% and 0.00% for CloneSelect A→G, CaTCH, low-copy CRISPRa, ClonMapper, CaTCH alternative and high-copy CRISPRa, respectively (Fig. 1i and Supplementary Fig. 5a). When an arbitrarily selected threshold for sorting EGFP+ cells was applied to the different systems, the true positive rates and false positive rates were 12.27–31.47% and 0.00–0.30%, respectively, for CloneSelect A→G, 16.15–34.55% and 0.95–19.06%, respectively, for CaTCH, 5.88–23.80% and 3.08–16.07%, respectively, for low-copy CRISPRa, 12.70–37.21% and 2.92–16.92%, respectively, for ClonMapper, 74.46–83.73% and 59.78–69.86%, respectively, for CaTCH alternative and 48.33–76.30% and 35.14–77.21%, respectively, for high-copy CRISPRa (Fig. 1j and Supplementary Fig. 5b). The EGFP intensities of EGFP+ cells across different systems did not show marked differences (Supplementary Fig. 5c).
We also tested two wild-type Cas9-based systems: CloneSifter (reported elsewhere)32 and another approach we developed. Although we observed slight enrichments of true positive cells for low false positive rates, their overall performances were largely outperformed by the base editing and CRISPRa-based systems (Supplementary Fig. 6). Accordingly, CloneSelect C→T and CloneSelect A→G performed the best compared to the currently reported and our proposed CRISPRa-based methods in terms of orthogonality in barcode-specific, gRNA-dependent reporter activation. We also advanced the CloneSelect C→T and CloneSelect A→G circuits to test single-copy EGFP reporters for a three-gRNA-input OR gate and a three-gRNA-input AND gate in HEK293T cells. They exhibited the expected input-dependent output patterns, albeit with low efficiencies (Supplementary Fig. 7).
We were not able to directly compare CloneSelect C→T and CloneSelect A→G because they need to encode GTG and TAA in the antisense strand of the gRNA target sequences, respectively, and the efficacy of the gRNA in recruiting the effector Cas9, in general, has been known to depend on its targeting sequence41,42. Therefore, we normalized the true positive rate and false positive rate of each barcode in the CloneSelect systems by those obtained for the same barcode in CaTCH or low-copy CRISPRa. We did not observe a large discrepancy in efficiency between the two systems (Fig. 1k). Therefore, we used CloneSelect C→T in the following demonstrations with mammalian cells. We also showed that the efficiency of CloneSelect C→T could be optimized by the amount of the target gRNA-encoding DNA without changing the false positive rate (Supplementary Fig. 8).
Benchmarking different methods using complex populations
To examine whether CloneSelect C→T can isolate target barcoded cells from a complex population, we next generated a barcoded lentiviral library (Extended Data Fig. 1a–d). In this library, the barcode region was designed to be a semi-random sequence of WSNS repeats (W = A/T; S = G/C) to avoid additional start codons from appearing. We then isolated barcoded plasmid clones into a 96-well plate and pooled 93 that were confirmed to have single barcodes by Sanger sequencing (Extended Data Fig. 1e). The plasmid mini-pool was used to transduce HEK293T cells with an infection rate of <0.1. We amplified the barcode region from the plasmid mini-pool and the transduced cells by PCR and then analyzed them by high-throughput sequencing. We identified 115 barcodes (Fig. 2a,b) and found that the variation in barcode abundance was largely inherited from that in the plasmid pool (Extended Data Fig. 1f,g), showing no substantial barcode-dependent bias in lentiviral packaging and transduction and cell growth.
a, Nucleotide compositions of barcodes prepared for Mammalian CloneSelect Pool-100. Five barcodes with unexpected lengths were excluded from this visualization. The full barcode sequence list can be found in Supplementary Table 1. b, Barcode abundance distribution in the Pool-100 cell population. The pie chart represents the success rate of enriching target barcodes to more than 25% in each sorted cell population. The horizontal blue line shows the least abundant barcode successfully enriched after sorting. c, gRNA-dependent labeling of target barcoded cells in Pool-100 (n = 1). Scale bar, 40 µm. d, Conceptual diagram of the benchmarking experiment using Pool-100. e, Barcode enrichment analysis after the cell sorting of EGFP+ cells. Each row shows the barcode abundance profile for the predetermined barcodes in the pool corresponding to each target isolation assay. f, Conceptual diagram of the benchmarking experiment using Pool-10000. Query gRNAs or reporters were individually transfected. The cell samples were pooled later for combined cell sorting and analyzed by high-throughput sequencing. g, Barcoded cell frequencies in pre-sort and post-sort populations (replicate one of n = 2). h, Conceptual diagram representing the separation store of target barcoded cells from a background cell population of similar barcode abundances. i, Separation scores of different isolation attempts from Pool-10000 prepared for different retrospective clone isolation systems. The target barcode abundances were adjusted by the dilution factor introduced by the pooling of different experimental samples. Box plots display the median along with the 25th and 75th percentiles, with whiskers extending 1.5 times the interquartile range. Two-tailed Mann–Whitney U-test was used for statistical analysis; **P < 0.01; ***P < 0.001.
We then tested whether we could enrich cells with 16 arbitrarily selected barcodes of different abundances in the cell population. For each target barcode, the cell population was co-transfected with the targeting gRNA and Target-AID plasmids. After 4 days, we observed EGFP+ cells in each assay (Fig. 2c) and isolated them by flow cytometry cell sorting (Fig. 2d and Supplementary Fig. 9a). The enrichment of the target barcode was then analyzed by PCR and high-throughput sequencing. In sum, we successfully enriched the target barcoded cells in 14 out of 16 experiments (success rate of 87.5%) with an enrichment threshold of 25% in the sorted population (Fig. 2b,e and Extended Data Fig. 1h). We believed that setting the enrichment threshold at 25% was reasonable to define success because any features for a clone whose abundance is 25% can be observed as long as its effect size is over four-fold compared to the variance in the background population. Additionally, the target clone can be easily isolated as a single cell during the sorting of reporter-positive cells with odds of 1:3. For the 14 successful targets, the mutated start codon was restored to ATG with an efficiency of 91.63–99.85% (Extended Data Fig. 1i). A fraction of EGFP+ cells with the expected barcodes did not demonstrate the GTG→ATG mutation, suggesting that the cytidine deamination by Target-AID on the antisense strand might be sufficient to express the codon-repaired reporter transcripts, as suggested previously38.
Next, we assessed the performance of CloneSelect C→T to isolate a target clone from a highly complex population and compared it with the two best-performing reported systems, CaTCH and ClonMapper. To test each system, we established a complexity-bottlenecked lentivirus library of ~10,000 barcodes and transduced HEK293T cells with an infection rate of <0.1 to obtain around one million transduced cells with single barcodes, ensuring the controlled barcode complexity in the barcoded cell population. After establishing the barcoded cell populations (hereafter referred to as ‘Pool-10000’ populations), the genomic DNA was extracted and subjected to PCR and high-throughput sequencing to quantify the barcodes. We performed the same barcode quantification for the lentiviral plasmid pool with two library preparation replicates and confirmed that the barcoded cell populations representing the complexities of the original plasmid DNA pools were successfully established (Extended Data Fig. 2a–c). We then obtained EGFP− populations by flow cytometry cell sorting for CloneSelect C→T and CaTCH to ensure a high sample quality for the subsequent reporter activation experiments.
For each of the CloneSelect C→T, CaTCH and ClonMapper Pool-10000 populations, we arbitrarily selected 16 target barcodes of diverse clone abundances for isolation. We also confirmed their sequence distances from the other members in the corresponding pools, and there was minimal risk of isolating non-target barcoded cells because of CRISPR gRNA off-targeting (Extended Data Fig. 2d–f). The sorting of a cell sample of this scale requires a long sorting time, and the additional variations possibly introduced while waiting for the sorting of other cell samples were a concern. For this reason, after separately performing the transfection of the pool with 16 target reporter activation reagents, we combined the 16 large cell samples of each clone isolation system and sorted EGFP+ populations from ~6.0 × 107 to ~1.0 × 108 cells per system (Fig. 2f and Supplementary Fig. 9b). After cell sorting, the barcode abundances were quantified by PCR and high-throughput sequencing.
As we multiplexed 16 assays, the enrichment of each expected positive barcode was diluted 16-fold on average. Therefore, we adjusted their relative barcode frequencies to be 16-fold to allow for an intuitive interpretation of the data (Fig. 2g and Extended Data Fig. 3a) (note that this adjustment underrepresents false positives and overrepresents false negatives, but we also interpret the data with this assumption). In the CloneSelect C→T Pool-10000 population experiment, the enrichment of some barcodes, especially those that were extremely rare in the initial barcoded population, did not accompany the observation of GTG→ATG conversion. Although the ATG sequence on the sense strand might not be required for the transcription of a functional EGFP from the template antisense strand, this result is probably compounded by the overrepresentation of false negatives because of the abundance adjustment. To quantitatively compare the three systems independent of the tested barcode abundances in the initial barcoded populations, we calculated a score for separating each target barcode from other barcodes of the same abundance level in the initial population (Fig. 2h). CloneSelect C→T also performed best in this separation score metric, with ClonMapper next. ClonMapper showed high background false positive activations, presumably because of the leaky transcription of the high-copy EGFP reporter (Fig. 2i). We performed the same separation score analysis without the abundance adjustment of 16-fold for expected positives and found that the relative performance of CloneSelect C→T was still higher than the other two (Extended Data Fig. 3b).
Isolation of clones identified in a scRNA-seq platform
To extend the use of CloneSelect for the isolation of living clones identified according to their single-cell transcriptome profiles, we established single-cell CloneSelect (scCloneSelect) C→T, which is compatible with 3′ capture scRNA-seq platforms. In scCloneSelect, the barcode located upstream of the reporter with the mutated start codon (hereafter referred to as ‘uptag’) is paired with another barcode downstream of the reporter (‘dntag’), followed by a hard-coded 30 nt poly(A) sequence (Fig. 3a). The dntag is captured by the standard scRNA-seq 3′-end sequencing strategy43,44 and used to refer to its corresponding uptag for the reporter start codon restoration. This change in the circuit design did not affect the reporter activation performance of CloneSelect C→T in HEK293T cells (Fig. 3b,c and Extended Data Fig. 4a,b) or the high orthogonality between barcodes and gRNAs (Extended Data Fig. 4c,d). We also confirmed that dntag barcodes were transcribed and efficiently captured by a scRNA-seq platform (Extended Data Fig. 4e,f).
a, scCloneSelect. LTR, long terminal repeat. b,c, Micrographs (b) and %reporter+ cells (c) for barcode-specific gRNA-dependent reporter activation by CloneSelect C→T and scCloneSelect in HEK293T cells (n = 3). Scale bar, 50 µm. d,e, Micrographs (d) and %reporter+ cells (e) for barcode-specific gRNA-dependent reporter activation of three barcoded mouse ES cell lines by scCloneSelect (n = 2). Target-AID was stably integrated before the barcoding. gRNAs were delivered by lentiviral transduction. Scale bar, 100 µm. Two-tailed Welch’s t-test was used for statistical analysis. f, Schematic diagram of the scCloneSelect workflow. g, mouse ES cell assays and clone isolation performed in this work. h, scRNA-seq of a mouse ES cell population treated with LIF and 2i and that without LIF or 2i. Uniform manifold approximation and projection (UMAP) was used for the two-dimensional embedding of the high-dimensional gene expression space into a two-dimensional space. i, Distribution of cells for arbitrarily selected clones in the same UMAP. j, Abundance distribution of cell clones in the barcoded mouse ES cell population. The pie chart represents the success rate of enriching target clones to more than 25% in each sorted cell population. The horizontal blue line shows the least abundant clone successfully enriched after sorting. k, gRNA-specific activation of target barcoded clones in the mouse ES cell population. Scale bar, 50 µm. l, Barcode enrichment analysis after the cell sorting of EGFP+ cells. Each row shows the barcode abundance profile for the predetermined barcodes in the pool corresponding to each target isolation assay. The left heatmap was expanded from the dashed box area of the right heatmap. *P < 0.05.
One intriguing application of scCloneSelect is to study the fate-determining factors of stem cell differentiation and reprogramming. scCloneSelect can be used to retrospectively isolate, from the initial population, cell clones whose states have been identified using scRNA-seq after differentiation or reprogramming. As stem cells commonly suffer from low transfection efficiency, we established mouse ES cells and human PS cells that constitutively express Target-AID using piggyBac transposon and introduced single scCloneSelect barcodes separately to them (Extended Data Fig. 4g). In mouse ES cells, we found that the reporter activation was more efficient when delivering the target gRNAs by lentiviral transduction than by transfection (Fig. 3d,e and Extended Data Fig. 4h). In human PS cells, although the transfection of the target gRNA led to successful reporter activation (Extended Data Fig. 4i), we also established an approach that required a minimal number of steps and genomic transgene integrations, whereby human PS cells were first lentivirally barcoded and then the reporter was activated by electroporating both the target gRNA and Target-AID plasmids together (Extended Data Fig. 4j,k).
To examine whether target barcoded clones identified in a scRNA-seq platform can be isolated from a barcoded cell pool that was sub-populated in parallel with the one used in scRNA-seq, we set up the following pipeline using mouse ES cells (Fig. 3f and Supplementary Fig. 10a). The EGFP fragment is first amplified with forward primers encoding semi-random uptags of WSNS repeats followed by a mutated start codon and reverse primers encoding random dntags. They are ligated into a common lentivirus backbone plasmid (Supplementary Fig. 10b,c). The constructed plasmid pool is used to barcode cells by transduction. The barcoded cell pool is then cultured to propagate the clones (step 1) and separated into three subpools (step 2). The first subpool is stored for later clone isolation (step 3). The second group is used to construct the reference database of uptag–dntag combinations by PCR amplification and high-throughput sequencing (step 4). The last group is subjected to a defined assay, during which intermediate subpopulations can be stored at any point (step 5). Cell clones demonstrating gene expression profiles of interest can be identified with their dntags by scRNA-seq, and their corresponding uptags can be retrieved from the uptag–dntag reference database (step 7). Finally, the target clones can be isolated from the subpopulations stored either at the beginning or during the assay (step 8). Given that lentivirus transduction, in general, is prone to recombining the payload sequences during the genomic integration45,46, the uptag–dntag database needs to be determined every time after the barcoding process. To this end, we also optimized the sequencing library preparation to minimize artefact chimeric PCR products (Supplementary Fig. 10d,e).
Using a lentivirus plasmid pool of ~150,000 barcodes, we transduced mouse ES cells with Target-AID at an infection rate of <0.1. Following the creation of a clone variation bottleneck by sparse sampling and 10 days of expansion, we constructed a small pool of barcoded clones, from which we identified 216 unique barcode pairs. After preserving a subpopulation for clone isolation, the remaining cells were cultured with cell differentiation inhibitors leukemia inhibitory factor (LIF) and 2i or without LIF or 2i, to maintain or lose pluripotency, respectively (Fig. 3g). Then, four days later, scRNA-seq was performed independently for the two conditions. The RNA capture rates per cell of the two datasets were similar (Extended Data Fig. 5a); however, the gene expression profiles of single cells were clustered into two distinct groups based on the culture conditions (Fig. 3h and Extended Data Fig. 5b). Although the barcoded clones did not show a significantly biased distribution between the two conditions, we attempted to isolate the top ten abundant clones in the scRNA-seq datasets (Fig. 3i and Extended Data Fig. 5c,d) from the initial barcoded population. The abundances of these clones varied from 0.0133% to 9.21% in the initial population according to the analysis determined by the uptag–dntag database (Fig. 3j). Except for one experiment targeting clone 153, we obtained a sufficient number of EGFP+ cells after introducing the target gRNA by lentivirus transduction (Fig. 3k, Extended Data Fig. 5e and Supplementary Fig. 11). Unfortunately, we could not determine the reason for the failure to recover clone 153 among several potential factors, including low abundance of the clone, the low efficiency of the barcode sequence for base editing and the poor quality of the lentivirus packaging used to deliver base editing reagents. For each of the remaining nine clone isolation attempts, eight showed target clone enrichment above an enrichment threshold of 25%, whereas one (clone 028) showed an enrichment frequency of 18.9% (Fig. 3l). We also isolated and expanded clone 006 and clone 012 and confirmed their clonal purities (Extended Data Fig. 5f,g).
Elite human stem cells with a high naïve propensity
Human PS cells have multidirectional differentiation potential and self-renewal capacity. Clone-to-clone heterogeneity and line-to-line differences in the propensities of human PS cells toward various cell differentiation directions have been reported in diverse in vitro cell differentiation and organoid generation protocols47,48,49. Although human PS cells resemble epiblast cells of the post-implantation embryo, they cannot differentiate into the trophoblast lineage. Protocols have been developed to chemically induce naïve human PS cells resembling pre-implantation embryonic epiblast cells from primed human PS cells50,51,52,53. However, a naïve induction protocol cannot perfectly induce naïve human PS cells, leaving some cells partially primed.
Understanding the underlying molecular mechanism and the fate navigation of naïve human PS cells is one of the central interests of developmental biology and regenerative medicine; therefore, we aimed to isolate elite primed human PS cell clones that have a high propensity to be induced into naïve human PS cells. We prepared a barcoded human PS cell population using a scCloneSelect library by lentiviral transduction with an infection rate of <0.1 (Fig. 4a). After preserving its subpopulations, another subpopulation of the barcoded cells (referred to as tier 1 primed) was used immediately to induce naïve cells. Another subpopulation was passaged five times (tier 2 primed) and subjected to naïve induction. At 21–23 days of each induction experiment, we sorted CD320+ cells as naïve cells (tier 1 naïve and tier 2 naïve).
a, Overview of the experiment to identify elite human PS cells having high naïve induction potential. b, Clonal abundance distributions of barcoded clones in tier 1 (T1) and tier 2 (T2) primed and native stem cells before and after naïve induction. An average value of two replicates (n = 2) is reported for each barcoded clone. n.d., not determined. c, Correlation in barcoded clone abundance between T1 and T2 naïve cell samples. d, Isolation of elite human PS cell clone candidates having high naïve potential from the parental population. The CloneSelect C→T reporter was activated by electroporating Target-AID and gRNA plasmids. The sorted clones were expanded and subjected to naïve induction followed by trophoblast differentiation. e, Isolated elite human PS cell clones (n = 1). Scale bar, 100 µm. f, Microscopic images and flow cytometric profiles (n = 2) of the isolated elite human PS cell clones. Scale bar, 100 µm. CD24 and SUSD2 boundaries to call primed and naïve cells are represented by the red lines, and median marker intensities are shown by bold black lines. g, Microscopic images and flow cytometric profiles (n = 2) of the isolated clones after naïve induction. %+ve, percent positive. h, Differentially expressed genes in the primed state between the parental cells and isolated clones (n = 2). i, Transcriptome and DNA methylation track at a chromosome X locus for the parental cells and isolated elite clones in the primed state (n = 2). j, Differentially expressed genes in the naïve state between the parental cells and isolated clone cells. k, Gene set enrichment analysis for the elite clone-specific downregulated genes after naïve induction. Gene Ontology terms were clustered by spring-embedding a network representation of the Gene Ontology term hierarchy relationships using Cytoscape. FDR, false discovery rate.
Analyzing clonal barcode abundances of the primed and naïve human PS cell samples (Fig. 4b), we observed 693 clones in the union of tier 1 and tier 2 primed samples and a significant, high correlation in barcode abundance between tier 1 and tier 2 naïve samples (Fig. 4c) and the recurrent domination of the similar sets of clones after naïve induction (Fig. 4b). On the other hand, the correlations between tier 1 and tier 2 primed samples and between primed and naïve samples were also significant but markedly lower (Extended Data Fig. 6a). These data collectively demonstrated the presence of human PS cell clones with a high naïve induction propensity, and their fates were not attributed to a stochastic factor in the cell but rather were maintained for at least the mid-term during the five passages.
We arbitrarily chose six barcoded elite clones (Fig. 4b) and successfully isolated five (Fig. 4d,e, Extended Data Fig. 6b–d and Supplementary Fig. 12) from the initial primed human PS cell population. Clone 185, which we rejected for the following analyses, was accompanied by the enrichment of another barcode, but this clone might just be doubly barcoded given that the frequencies of the expected and unexpected barcodes were both nearly 50% (Extended Data Fig. 6b). We compared the isolated clones with the parental bulk human PS cells and the barcoded human PS cells, which all showed a typical flat, primed human PS cell morphology under the microscope (Fig. 4f). Note that the isolated cells underwent the stable genetic code restoration of the active EGFP expression (Extended Data Fig. 6d). When analyzed by flow cytometry, all the cell samples showed the CD24 primed cell marker expression, but the CD24 levels of the isolated clones were slightly lower than the parental cells (Fig. 4f), which was inconclusive but implied a difference in the molecular profile of the cell.
We tested whether the isolated clones retained and exerted the elite naïve transition propensity. By inducing naïve stem cells through chemical resetting, all of the samples presented a dome-shaped naïve stem cell colony morphology under the microscope (Fig. 4g). When we quantified the naïve transition efficiencies, the expected elite clones showed a nearly perfect transition, whereas this was not the case for the parental cells (Fig. 4g and Extended Data Fig. 6f–h). With arbitrary thresholds for CD24 (a primed marker) and SUSD2 (a naïve marker), 96.63–98.87% of the cells were found in the CD24−/SUSD2+ naïve cell fraction for the isolated clones and 56.85–85.68% for the parental cells.
To explore molecular factors underpinning the naïve transition potential, we performed an RNA-seq analysis of the elite primed cell clones and their parental cell population before and after barcoding, as well as those induced to naïve stem cells and purified using naïve stem cell markers in duplicates (Extended Data Fig. 7a). Although the global transcriptome profiles highlighted differences between the primed and naïve stem cell samples, the primed cell sample group and the naïve cell sample group each showed largely similar gene expression patterns. When differentially expressed genes in the primed state were explored between each of the five elite clones and parental populations, only five genes were detected as commonly downregulated genes in clone 006, clone 034, clone 116 and clone 216 (Fig. 4h and Extended Data Fig. 7b). By contrast, the same set of genes was upregulated in clone 332. Interestingly, four of them—CSAG1, MAGEA12, MAGEA6 and MAGEA3—were encoded in a proximal genomic locus on chromosome X (Fig. 4i). To explore their genome-wide methylation profiles, we selected clone 006, clone 216 and the outlier clone 332 in the primed state and analyzed them by enzymatic methyl-seq (EM-seq)54, along with their paired parental barcoded cell samples obtained following the introduction of corresponding base editing reagents before clone isolation (Extended Data Fig. 6f). Although no global difference in hypermethylation and hypomethylation patterns across the genome was observed (Extended Data Fig. 7c), we found elite clone-specific hypermethylation on the CSAG1 promoter region for clone 006 and clone 216 (Fig. 4i). Notably, the hypermethylation site overlapped with an ENCODE candidate promoter element and within a topologically associated domain harbouring the four genes55 (Extended Data Fig. 7d). By contrast, hypomethylation was observed in the same locus for clone 332 in agreement with its gene expression pattern.
When differentially expressed genes after naïve induction were explored between each of the five elite clones and parental populations, we observed 131 upregulated and 218 downregulated genes for the elite clone-derived samples compared to their parental cell-derived samples (Fig. 4j and Extended Data Fig. 7e). Interestingly, performing the gene set enrichment analysis (GSEA), we found that these genes were enriched for gene expression, RNA splicing and post-translational modifications, which somewhat aligned with the features of totipotent-like stem cells or 2C-like cells56 (Fig. 4k and Extended Data Fig. 7f). In concordance with the unique gene expression patterns in the naïve cells obtained from the elite cell clones, although they contributed to the trophoblast lineage, the overall efficiency was slightly lower than those obtained from the parental primed human PS cells, suggesting that intrinsic molecular fate determinants modulate their transitions into different cell states (Supplementary Fig. 13). Altogether, we observed an epigenetic signal that may explain the retention of a cell fate through multiple cell division generations, at least for some elite clones, highlighting the need for further investigation into the heterogeneity of epigenetic profiles and stem cell fates.
Yeast and Bacterial CloneSelect
Clonal barcoding approaches have also been used in microorganisms, such as yeast and Escherichia coli, to study their laboratory evolution and the genomic mutations accompanying clonal expansions of cells7,9. However, current analysis methods have been limited to time-course tracing of clone size dynamics. No retrospective clone isolation technology has been developed for yeast, and a recent clone isolation method developed for E. coli has been demonstrated on a limited scale57. Therefore, we extended CloneSelect C→T to yeast Saccharomyces cerevisiae (Yeast CloneSelect) and CloneSelect A→G to E. coli (Bacterial CloneSelect).
We used mCherry as a fluorescent reporter in Yeast CloneSelect (Fig. 5a). We first realized that mCherry translation could also be initiated from the second methionine coding codon in both mammalian and yeast cells and used an amino-terminus-truncated mCherry (Extended Data Fig. 8 and Supplementary Fig. 14). CRISPR base editors, including Target-AID, developed for mammalian species are fused with a uracil glycosylase inhibitor (UGI) to inhibit the base excision repair pathway, enhancing both the efficacy and purity of C→T substitution at the target site37. However, Target-AID was originally tested in yeast only without a UGI and was demonstrated to confer C→D (non-C) substitution at the target sequence at a high rate35. Therefore, we constructed a yeast Target-AID with a UGI and found that it did not largely impair the base editing activity (Extended Data Fig. 8b–d) but, as expected, greatly enhanced the frequency of C→T purity at the target site (Extended Data Fig. 8e). Efficient reporter activation was only possible with the UGI fusion (Extended Data Fig. 8f). Similar to the mammalian CloneSelect systems, Yeast CloneSelect was also demonstrated to activate the reporters in a highly target-specific manner (Fig. 4b–d and Extended Data Fig. 8g). Furthermore, unlike mammalian cells, the labelled clones could be isolated by picking fluorescent colonies formed on a solid agar plate (Fig. 5e).
a, Yeast CloneSelect. b,c, Micrographs (b) and reporter mCherry intensities (c) for barcode-specific gRNA-dependent mCherry activation. Scale bar, 25 µm. mCherry intensities measured by a plate reader were normalized by OD595 (n = 3). d, GTG→ATG editing frequencies observed by high-throughput sequencing. Two-tailed Welch’s t-test was used for the statistical test. e, Yeast colonies formed on a 10-cm agar plate after introducing a trigger plasmid encoding a target gRNA to the Pool-100 yeast population. f,g, Barcode abundance distributions in Pool-100 (f) and Pool-1580 (g). The pie charts represent the success rates of isolating target barcoded cells whose abundances were above the minimal abundance that was successfully isolated. The horizontal blue lines show the least abundant barcode successfully isolated. h,i, Colonies of barcoded cells isolated from Pool-100 (h) and Pool-1580 (i). j, Summary of CloneSelect yeast assays. k, Bacterial CloneSelect using a Zeocin resistance reporter gene. l, Barcode-specific gRNA-dependent reporter activation. m, Schematic diagram of the Bacterial CloneSelect workflow. n, Colonies formed on Zeocin selective and non-selective solid agar plates after introducing a trigger plasmid encoding a target gRNA and ABE to the Pool-100 E. coli population. o, Barcode abundance distributions in Pool-100 and Pool-1550. The pie charts represent the success rates of isolating target barcoded cells from the respective pools. The horizontal blue lines show the least abundant barcode successfully isolated. p, Analysis of colonies isolated from Zeocin selective and non-selective plates obtained from Pool-100 and Pool-1550. CFUs, colony-forming units; n.d., not determined. **P < 0.01; ***P < 0.001.
To test the sensitivity of Yeast CloneSelect, we generated a barcode plasmid pool by pooled ligation of the SWSN repeat barcode fragments to a backbone vector (Extended Data Fig. 9a–c). We then bottlenecked the barcode plasmid complexity, obtaining plasmid pools of 100 and ~1,580 colonies, and established yeast cell populations (referred to as ‘Pool-100’ and ‘Pool-1580’, respectively). From Pool-100 and Pool-1580, we attempted to isolate cells for 26 and 31 barcodes, respectively (Fig. 5f,g and Extended Data Fig. 9d). For each isolation, a target gRNA plasmid and Target-AID plasmid were co-transformed into the yeast cell pool. Fluorescent colonies were isolated, if any, together with four non-fluorescent colonies. The colony isolates were then cultured in liquid selective media to measure the fluorescence intensities, and barcode sequences were examined by PCR followed by Sanger sequencing (Fig. 5h,i and Extended Data Fig. 9e,f). For Pool-100, 16 out of the 26 attempts (61.58%) resulted in positive colonies, all of which had the expected barcodes with a GTG→ATG conversion rate of 48.92–97.41%, except for one of the three positive colonies obtained for barcode 030 (Fig. 5j). For Pool-1580, six out of the 31 attempts (19.35%) resulted in positive colonies, all of which had the expected barcodes with a GTG→ATG conversion rate of 81.51–97.20% (Fig. 5j). The least barcode abundances successfully isolated from Pool-100 and Pool-1580 were 0.81% and 0.12%, respectively.
To establish Bacterial CloneSelect, we first tested the CloneSelect A→G EGFP reporter expressed under the arabinose (Ara)-inducible promoter. E. coli cells were transformed using a reporter plasmid with a trigger plasmid encoding a target or non-target gRNA and ABE, each expressed under an isopropyl β-d-1-thiogalactopyranoside (IPTG)-inducible promoter consisting of a T7 promoter and lac operator (Extended Data Fig. 10a). In the IPTG-inducible promoter system, IPTG serves as a molecular mimic of allolactose and binds to the lac repressor, causing it to release from the lac operator sequence, thereby allowing gene expression. We found that the EGFP expression level by the target gRNA was only slightly higher than that by a non-target gRNA regardless of IPTG induction (Extended Data Fig. 10b–d). At the same time, the expected A→G substitution by a target gRNA was conferred without IPTG, suggesting that minimal gene expression of base editing reagents satisfies the edit. We also observed that the IPTG induction of base editing machinery instead led to the silencing of EGFP, probably because of silencing or bystander editing by ABE (Extended Data Fig. 10c). Therefore, we switched to a drug-selectable system in which EGFP was replaced with a Zeocin resistance gene (Sh ble). We realized that a tight gRNA-dependent reporter activation was only possible without Ara, as the addition of Ara led to false positive cells under Zeocin, showing that the reporter expression also needs to be minimized (Extended Data Fig. 10e). We also found that under the no-IPTG condition, removing the lac operator only from the target gRNA expression unit substantially dropped the number of colony-forming units (Extended Data Fig. 10f), probably because nickase Cas9 is toxic to bacterial cells58,59. Finally, we optimized the barcode reporter plasmid to use Sh ble under the Ara promoter and the trigger plasmid to encode a gRNA and ABE, both under the IPTG-inducible promoters, and use them without Ara or IPTG (Fig. 5k,l and Extended Data Fig. 10g). We also showed that the same setup can be used to construct a blasticidin S-resistance gene (bsr)-based reporter (Extended Data Fig. 10h,i).
Bacterial CloneSelect with the Sh ble reporter enabled isolating target barcoded E. coli clones with high sensitivity and high specificity. To demonstrate barcoded cell isolation from a complex population, we constructed a pooled plasmid library with semi-random barcodes of VNN repeats (V = non-T), preventing the appearance of stop codons (Fig. 5m and Supplementary Fig. 15a), and prepared cell pools by combining 100 and ~1,550 colonies, respectively (hereafter referred to as ‘Pool-100’ and ‘Pool-1550’). From Pool-100 and Pool-1550, we attempted to isolate ten and four barcoded clones, whose abundances ranged from 0.047–2.33% and 0.00089–0.211%, respectively (Fig. 5o and Supplementary Fig. 15b). For each target, the cell pool was transformed with a trigger plasmid encoding the target gRNA and ABE and selected under Zeocin (Fig. 5m). In every isolation experiment, the Zeocin selective conditions showed a substantially lower number of colonies than the non-selective conditions (Fig. 5n). For each of the successful target barcodes, except for barcode 036 of Pool-100 in which we obtained only two colonies in the selective condition, four and four colonies were isolated from the selective and non-selective conditions, respectively, and their barcodes and base editing patterns were analyzed by Sanger sequencing. All isolates from the selective conditions had the expected barcodes (Fig. 5p). By contrast, all of the isolates from the non-selective conditions had non-targeted barcodes. The least barcode abundances successfully isolated from Pool-100 and Pool-1550 were 0.047% and 0.0089%, respectively.
Accordingly, we demonstrated that Yeast CloneSelect and Bacterial CloneSelect are capable of isolating rare barcoded clones from a complex cell population with high sensitivity and near-perfect specificity.
Discussion
CloneSelect enables the isolation of target barcoded cells from a complex population using CRISPR base editing. Compared to the other retrospective clone isolation methods tested in this study, CloneSelect demonstrated an ability to isolate cells with an overall higher performance. Despite the gene circuit configurations optimized in some of the CRISPRa-based systems, they generally showed limited performance, probably because of the background reporter expression without the target gRNA. The wild-type Cas9-based systems share the same principle with CloneSelect and use the barcode-specific genetic code alternation for the reporter expression. However, the wild-type Cas9-based systems suffered from efficiency, probably because of the stochasticity in editing outcomes and the reported cytotoxicity.
CloneSelect benefits from the precision of base editing and the simplicity of altering the genetic code. The engineering of the evolutionarily conserved genetic code also enabled the implementation of the same concept across multi-kingdom species. We demonstrated the retrospective isolation of barcoded cells from complex yeast populations. Although the isolation of barcoded E. coli cells has recently been demonstrated using barcode-specific CRISPR interference of a counter-selection marker57, we showed that Bacterial CloneSelect isolated low-abundant target barcoded cells. Overall, we demonstrated that CloneSelect can enrich a target clone representation from more than one out of 10,000 for mammalian cells, from more than one out of 1,000 for yeast cells and from more than one out of 100,000 for bacterial cells. The success rates for isolating target clones at those abundance levels are estimated to be around 50% in mammalian cells and 100% in E. coli cells, respectively. The relatively low success rate of 19.3% in obtaining a target clone from the Pool-1000 yeast pool was probably a result of the limited sampling sensitivity in the fluorescence colony isolation approach, in which cells should be sparsely spread on the selectable plates. We expect the performance of Yeast CloneSelect to improve significantly with a flow cytometry cell sorting-based or growth-selective reporter approach, as in Mammalian and Bacterial CloneSelect.
The limited success rate per isolation attempt could be explained by the general gRNA-dependent genome editing efficacy41, as the isolation success did not correlate well with the abundance of the target in a population, and the relative performances across the same set of different barcodes were well correlated between different retrospective clone isolation systems. Therefore, given the current success rate, we suggest that CloneSelect is sufficient for most prospective assays. Given that a biological assay can be expected to show multiple barcoded clones exhibiting a target phenotype of interest, we optimistically suggest that one can plan multiple isolation attempts using different gRNAs.
In Mammalian CloneSelect systems, the purity of target barcoded cells after sorting reporter-positive cells was sometimes limited. The target barcoded cell enrichments from the HEK293T populations were generally not high, while the purities of isolating target barcoded clones from the mouse ES cell and human PS cell populations were high. Although we could not fully investigate this outcome, the level of clone enrichment appeared to depend largely on the cell sorting machines and sorting parameters. Nevertheless, if the purity of the target clone is not high after sorting the reporter-activated cells, we recommend performing single-cell isolation during cell sorting or from the enriched target population.
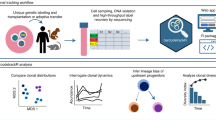
We propose CloneSelect to enable wide-ranging experiments in various fields of life sciences research. Existing time-course scRNA-seq measurement strategies already enable the interrogation of different clonal lineages in a barcoded population alongside the dynamic changes in their gene expression patterns, provided the clone population sizes are not too small14. By contrast, CloneSelect would allow clones isolated from different time points within a progressing population to be analyzed by diverse approaches (Fig. 6a). Such non-transcriptomic analyses could include morphological analyses under a microscope, molecular analyses available for small amounts of input cells and any currently available methods, as long as the given hypotheses permit the propagation of the isolated clones.
a, Clonal analysis of molecular profiles in a complex population. b, Replication of cell state trajectories. c, Reconstitution of synthetic cell populations. d, Transplantation of a fate-restricted elite clone. e, Isolation of a target product in a low efficient DNA assembly.
Cells isolated by CloneSelect are alive. The clones isolated from the initial population of a once-performed assay can be tested to determine whether they follow the same behavioural trajectories (Fig. 6b) or used to reconstitute a synthetic population with another cell population or other isolated clones (Fig. 6c). For example, a variety of human PS cell lines have been reported to be favourable for various cell differentiation and organoid models60,61,62,63,64, suggesting that there could also be fate priming of stem cell clones owing to undiscovered intrinsic factors. As exemplified in the naïve stem cell induction experiment, CloneSelect enables the mapping of cell states for stem cell clones after induction or differentiation together with their isolation from the initial population. The fate-mapped elite stem cell clones could be used to engineer new stem cell-based models or high-quality stem cell therapeutics. In general, CloneSelect could be used to obtain high-quality cells for cell-based therapies. Furthermore, cell clones isolated from diverse systems can also be transplanted into animal models (Fig. 6d). Examples include xenotransplantation of a cancer stem cell clone and aggregation of a fate-mapped stem cell clone with an early embryo. When a fluorescent reporter gene is used, the spatial distribution of the target clone and its interactions with others can be traced.
Lastly, in alignment with using DNA sequencing as a readout, CloneSelect would also promote the engineering of cells and DNA65,66,67,68,69. In the genetic engineering of cells, only a fraction of cells in the product pool typically encodes the target genetic product. Thus, obtaining successful cells becomes difficult when the efficiency of the genetic manipulation is low. In such situations, CloneSelect can enrich the target cells through barcoding and sequencing of the product cells. Similarly, we envision CloneSelect to improve DNA assembly (Fig. 6e). Currently, it is common practice to transform a DNA assembly reaction sample into E. coli cells to isolate assembly product clones, followed by colony isolation and screening of the correctly assembled products by sequencing. We propose using a pool of CloneSelect barcodes in the DNA assembly reaction to molecularly tag assembly products. The assembly products are used to transform E. coli, followed by pooling of transformants and extraction and long-read sequencing of the pooled plasmid products with their barcodes70. Finally, the barcoded clone harbouring the target product can be isolated by Bacterial CloneSelect. This strategy would enable the isolation of a target product from an inefficient DNA assembly reaction that would have previously been excluded from consideration.
Accordingly, CloneSelect is a method to precisely isolate a cell clone from a complex population. Its performance across multi-kingdom species opens a wide array of possibilities for addressing unresolved questions and tackling challenging engineering tasks in diverse areas of biology.
Methods
Plasmids
Oligonucleotides were chemically synthesized by FASMAC, Integrated DNA Technologies or Eurofins Genomics. All oligonucleotides and cloning procedures used to construct the plasmids in this study are listed in Supplementary Table 2. We used QUEEN (v.1.2.0) (https://github.com/yachielab/QUEEN) to design each plasmid construction and generate annotated plasmid files in QUEEN’s GenBank (gbk) file format, embedding the full construction procedure (see Supplementary Table 2). A QUEEN gbk file acts as a quine code that enables retrieving the plasmid construction process that generates the same plasmid map in the gbk format71. We believe that providing these QUEEN gbk files fulfils the requirement for reporting reproducible plasmid construction protocols. We also provided natural language descriptions for all the plasmid construction protocols in the QUEEN gbk files. Users can retrieve the protocols by executing ‘QUEEN --protocol_description --input [gbk file]’ in a QUEEN-installed environment. A custom QUEEN wrapper that generated all QUEEN-generated gbk files is also available at https://github.com/yachielab/CloneSelect_v1/tree/main/QUEEN. Accordingly, we do not include plasmid construction protocols in this paper. All plasmid DNA sequences were confirmed by Sanger sequencing. The representative plasmids are available from Addgene along with their QUEEN gbk files, as agreed upon with Addgene.
Common methods
Lentivirus preparation
Packaging
HEK293T cells were plated either in a 10 cm cell culture dish at a density of ~2 × 106 cells in 10 ml of culture medium or in six-well cell culture plate wells at a density of ~2 × 105 cells per well in 2 ml of culture medium, 1 day before plasmid transfection.
For virus packaging in a 10 cm dish, 3.0 µg of the transgene vector, 2.25 µg of psPAX2 (Addgene, no. 12260), 0.75 µg of pMD2.G (Addgene, no. 12259) and 18 µl of 1 mg ml−1 PEI MAX (Polysciences, no. 24765-100) were dissolved in 1,000 µl of 1× PBS and added to the cell culture. For packaging in a six-well plate, 489 ng of the transgene plasmid, 366.7 ng of psPAX2, 122.3 ng of pMD2.G and 2.93 µl of 1 mg ml−1 PEI MAX were dissolved in 300 µl of 1× PBS and added to the culture.
The culture medium was replaced with fresh medium 1 day after transfection. Transfected cells were then incubated for an additional 48–72 h. The recombinant lentivirus supernatant was collected and filtered through 0.22 µm sterile syringe filters. The lentivirus samples were aliquoted in 500–1,000 µl volumes into 1.5 ml tubes and stored at −80 °C.
Virus concentration
To increase the viral infection titre, collected virus samples were concentrated using a polyethylene glycol (PEG)-based method72 with PEG 6000 (Wako, no. 169-09125) or with Lenti-X Concentrator (Takara, no. 631231).
For concentration with PEG 6000, approximately 10 ml of the recombinant virus sample was combined with 2.55 ml of 50% w/v PEG 6000, 1.085 ml of 4 M NaCl and 1.365 ml of 1× PBS in a 50 ml tube. The mixture was rotated continuously at 4 °C for 90 min, then centrifuged at 4,000g and 4 °C for 20 min. The supernatant was discarded, and the virus pellet was resuspended in 1.1 ml of Opti-MEM (Gibco, no. 31985062) by pipetting and vortexing until fully dissolved, achieving a tenfold concentration of the virus sample.
Virus concentration using Lenti-X Concentrator followed the manufacturer’s protocol, with the virus dissolved in Opti-MEM (Gibco, no. 31985062) for a tenfold or 15-fold concentration. The concentrated virus samples were stored at −80 °C.
Preparing microscope imaging samples
All live-cell imaging was conducted using a BZ-X710 (Keyence), InCellAnalyzer 6000 (GE Healthcare) or IX83 (Olympus) with a ×4, ×10 or ×20 objective lens. The contrast and brightness of images obtained in a single experimental batch were uniformly adjusted using ImageMagick (v.7.1.0-20) or Fiji (v.1.0).
HEK293T cells and mouse ES cells were analyzed with Hoechst staining. For HEK293T cells, 25 µl of 0.1 mg ml−1 Hoechst 33342 (Invitrogen, no. H3570) dissolved in DMEM was directly added to each well of 24-well cell culture plates 3 days after transfection for nuclear counterstaining. The specimens were incubated at room temperature (18–25 °C) for 10 min, after which the culture medium was removed. Cells were gently washed with 500 µl of fresh DMEM and filled with 500 µl of fresh DMEM before imaging. For mouse ES cells, 5.0 µg ml−1 Hoechst 33342 dissolved in cell culture medium was directly added to each well and incubated at room temperature for 10 min before imaging.
Flow cytometry analysis
Cells were detached with 0.25% w/v trypsin-EDTA (Wako, no. 201-18841), incubated at 37 °C for 5 min, collected into a 1.5 ml tube or a 96-well round-bottom plate and centrifuged at 100g at room temperature for 5 min. After aspirating the supernatant, cell pellets were gently resuspended in 150–500 µl of ice-cold FACS buffer (2% FBS in 1× PBS). Samples were immediately placed on ice until flow cytometry analysis.
Flow cytometry analysis was performed using a BD FACSVerse cell analyzer (BD Biosciences) or CytoFLEX flow cytometer (Beckman Coulter). Samples were gently mixed by pipetting or vortexing immediately before analysis, and approximately 10,000–20,000 raw events were acquired per sample. Data analysis was conducted with custom R scripts using flowWorkspace (v.0.5.40) (https://github.com/RGLab/flowWorkspace), flowCore (v.1.11.20) (https://github.com/RGLab/flowCore) and CytoExploreR (v.1.1.00) (https://github.com/DillonHammill/CytoExploreR) or with the Python package FlowCytometryTools (v.0.5.0) (https://github.com/eyurtsev/FlowCytometryTools). The codes are available at https://github.com/yachielab/CloneSelect_v1/tree/main/FACS.
High-throughput sequencing
All amplicon sequencing libraries were combined with a 20–30% PhiX spike-in DNA control (Illumina, no. FC-110-3001) to enhance cluster generation on the flow cell. Libraries were sequenced using Illumina MiSeq (MiSeq v.3 150-cycle kit no. MS-102-3001 or 300-cycle kit no. MS-102-3003) or HiSeq 2500 (TruSeq rapid SBS kit v.2 no. FC-402-4022). Base calling was performed with bcl2fastq2 (v.2.20.0) to generate FASTQ files. Detailed sequencing conditions for each library and NCBI Sequence Read Archive IDs for each raw FASTQ file are provided in Supplementary Table 3.
Barcode identification and analysis
In barcode identification of each different barcoding system, sequencing reads were aligned to the constant sequences of the library structure using NCBI BLAST+ (v.2.6.0)73 with the blastn-short option to identify sample indices for demultiplexing and barcode sequences. For the clone isolation experiments, a barcode allowlist was generated by identifying barcode sequences present in both the plasmid DNA library and the genomic DNA library. Sequencing errors were corrected using Starcode (v.1.4) (https://github.com/gui11aume/starcode) with a maximum Levenshtein distance threshold of four, merging minor barcodes into major ones.
Barcode counts in each sample were normalized by the total barcode count. Barcode frequencies for each cell or DNA pool sample were estimated by averaging frequencies across replicates, where applicable. The barcode sequence and frequency data generated in this study are provided in Supplementary Table 1.
Statistical analysis
Statistical tests were conducted using R (v.4.2.0 and v.4.3.1). Specific details for each test are provided in the corresponding figure legends. Additionally, the statistical methods and associated P values used in this study are listed in Supplementary Table 4.
Experiments using HEK293T cells
Cell culture
HEK293Ta and HEK293T Lenti-X cells were purchased from GeneCopoeia (no. LT008) and Takara (no. 632180), respectively. Cells were cultured in DMEM (Sigma-Aldrich, no. 11965084) supplemented with 10% FBS (Gibco, no. 16000044) and 1% penicillin–streptomycin (Wako, no. 168-23191) at 37 °C with 5% CO2 in a cell culture incubator. Cells were detached and passaged using 0.25 w/v% trypsin-EDTA (Wako, no. 203-20251) once they reached 70–90% confluency. For microscopic imaging of HEK293T cells with Hoechst 33342 (Invitrogen, no. H3570) counterstain, 100–200 µl of Collagen-I (Nippi, no. PSC-1-100-100) diluted in 5 mM acetic acid was added to each cell culture plate well and incubated for 30 min at 37 °C. The collagen-coated plate wells were washed with 100–200 µl of 1× PBS before use. Cells were regularly tested for mycoplasma contamination.
Barcode plasmid pool preparation
CloneSelect C→T barcode library
To generate the CloneSelect C→T barcode library, a semi-random oligonucleotide pool, SI#679, encoding 5′-CCGWSNSWSNSWSNSWSNSNGTG-3′, was first chemically synthesized (Supplementary Table 2). This sequence includes the antisense strand of the 5′-CGG-3′ PAM sequence, followed by a quadruple repeat of WSNS (where W = A or T; S = G or C) and a mutated start codon (GTG). The WSNS repeat prevents the formation of additional start and stop codons upstream of the reporter. An EGFP coding sequence was then amplified from pLV-eGFP (Addgene, no. 36083) in 25 separate 50 µl PCR reactions, each containing 1 ng µl−1 of pLV-eGFP template plasmid, 1.25 µl of 20 µM SI#679 oligonucleotide pool as the forward primer, 1.25 µl of 20 µM SI#680 as the common reverse primer, 0.5 µl of Phusion High-fidelity DNA Polymerase (NEB, no. M0530), 10 µl of 5× Phusion HF Buffer (NEB, no. B0518S) and 5 µl of 2.5 mM deoxynucleotide triphosphates (dNTPs; Takara, no.4025). The thermal cycling conditions were as follows: 98 °C for 30 s; 30 cycles of 98 °C for 10 s, 72 °C for 10 s and 72 °C for 60 s; with a final extension at 72 °C for 5 min.
The amplified fragment was digested with DpnI (NEB, no. R0176) for 1 h at 37 °C, pooled into a single 1.5 ml tube and purified using the FastGene PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302). The purified fragment was then subjected to overnight digestion with EcoRI-HF (NEB, no. R3101S) and XbaI (NEB, no. R0145S) at 37 °C, followed by another purification with the FastGene PCR/Gel Extraction Kit. To obtain a highly complex lentiviral plasmid pool, we performed five ligation reactions using PCR strip tubes, each containing ~30 fmol of EcoRI-XbaI-digested pLVSIN-CMV-Pur backbone plasmid (Takara, no. 6183), ~300 fmol of the insert fragment, 2.5 µl of T4 DNA Ligase (NEB, no. M0202) and 5 µl of 10× T4 DNA Ligase Reaction Buffer (NEB, no. B0202) in a total volume of 50 µl. Reaction samples were incubated at room temperature for 2 h and then purified using the FastGene PCR/Gel Extraction Kit.
The ligation samples were used to transform NEB Stable Competent E. coli cells (NEB, no. C3040I) in five separate reactions, each with 1,250 ng of the ligation sample in 200 µl of competent cells, following the manufacturer’s high-efficiency transformation protocol. After a 1 h outgrowth in SOC medium (NEB, no. B9020) at 37 °C, cells were spun down and plated across 25 LB agar plates containing 100 µg ml−1 ampicillin (Wako, no. 014-23302). Colonies that formed on each plate after overnight incubation at 37 °C were scraped with 1–2 ml of double-distilled water (ddH2O). The cells collected from the plates were pooled into a flask and incubated in 200–300 ml of LB liquid medium with 100 µg ml−1 ampicillin (Wako, no. 014-23302) overnight at 37 °C. The transformation sample was plated with a 500-fold dilution in triplicate, and the library’s complexity was estimated to be ~6.8 × 105. The plasmid library was then purified using the NucleoBond Midi-prep Kit (Macherey-Nagel, no. 740410) and stored at −20 °C.
We isolated 16 random clones and verified the presence of the expected barcode inserts through triple restriction enzyme digestion with BsrGI-HF (NEB, no. R3575S), ClaI (NEB, no. R0197S) and PvuI-HF (NEB, no. R3150S), confirming that 16 out of 16 clones contained the desired inserts. To generate the mini-pool library for proof-of-concept assays in HEK293T cells, we sequenced barcode regions from 96 isolated clones by Sanger sequencing with primer SI#471. After excluding three clones with mixed sequencing spectra in the barcode region, the remaining barcoded plasmids were pooled in equimolar ratios and subjected to high-throughput sequencing and lentiviral packaging.
CloneSelect C→T Pool-10000 barcode library
To generate the CloneSelect C→T Pool-10000 barcode library, 100 ng of the original 700K library plasmid pool was re-transformed into 10 µl of NEB Stable Competent E. coli cells (NEB, no. C3040I). This transformation was controlled to confer approximately 10,000 colonies. The collected colonies were pooled and cultured overnight in 5 ml LB liquid medium containing 100 µg ml−1 ampicillin (Wako, no. 014-23302) at 30 °C. Plasmid DNA was extracted using the GeneJET Plasmid Miniprep Kit (Thermo Fisher Scientific, no. K0502) and stored at −20 °C until use.
CaTCH and ClonMapper Pool-10000 libraries
The CaTCH and ClonMapper Pool-10000 libraries were constructed using Golden Gate Assembly74 with the same protocol. To prepare an insert fragment pool, two single-stranded DNA oligonucleotide pools containing a random 19-mer nucleotide sequence were synthesized by Integrated DNA Technologies and annealed to generate sticky-end overhangs (underlined): 5′-CACCCNNNNNNNNNNNNNNNNNNNG-3′ and 5′-AAACCNNNNNNNNNNNNNNNNNNNG-3′ for CaTCH; 5′-CACCGNNNNNNNNNNNNNNNNNNG-3′ and 5′-AAACCNNNNNNNNNNNNNNNNNNC-3′ for ClonMapper (Supplementary Table 2). Equal volumes of top and bottom strand oligonucleotide pools were combined for phosphorylation and annealing in a 30 µl reaction volume in an eight-strip PCR tube. The reaction mixture included 3 µl of 10× T4 PNK Buffer (Takara, no. 2021A), 1.5 µl of T4 PNK (Takara, no. 2021A) and 3 µl each of 100 µM top and bottom strand oligonucleotide pools. The mixture was incubated with the following thermal cycling conditions: 37 °C for 30 min, 95 °C for 5 min, 70 cycles of 12 s at 95 °C with a ramp down of 1 °C per cycle, and then maintained at 25 °C. The annealed oligonucleotide pool was diluted to 1/10 with ddH2O and used for Golden Gate Assembly with the appropriate lentiviral cloning backbone (pLV-CS-307 and lentiTRACE-hU6-Puro for CaTCH and ClonMapper, respectively). The Golden Gate Assembly reaction mix was prepared in a 12.5 µl volume in an eight-strip PCR tube, consisting of 1 µl of insert, 1.25 µl of 10× T4 DNA Ligase Reaction Buffer (NEB, no. B0202S), 0.625 µl of 2 mg ml−1 BSA (NEB, no. B9000S), 0.5 µl of T4 DNA Ligase (Nippon Gene, no. 317-00406), 0.5 µl of BsmBI (NEB, no. R0580), 1.25 µl of 25 mM ATP (NEB, no. P0756S) and 12.5 ng of the backbone plasmid. The assembly reaction underwent the following thermal cycling conditions: 15 cycles of 37 °C for 5 min and 20 °C for 5 min, followed by 55 °C for 30 min, then held at 4 °C.
Following assembly, 3 µl of the product was transformed into NEB Stable Competent E. coli cells (NEB, no. C3040I) using the high-efficiency transformation protocol. After 1 h of outgrowth in SOC medium (NEB, no. B9020) at 30 °C, cells were spun down and plated on LB agar plates containing 100 µg ml−1 ampicillin (Gibco, no. 11593027). This transformation was controlled to confer approximately 10,000 colonies. After overnight incubation at 30 °C, colonies on each plate were scraped into 1–2 ml of LB medium containing 100 µg ml−1 ampicillin and pooled in a 5 ml tube. The collected cell samples were further incubated overnight with 4–6 culture tubes, each with 5 ml of LB liquid medium with 100 µg ml−1 ampicillin at 30 °C. Plasmid DNA was extracted using the GeneJET Plasmid Miniprep Kit (Thermo Fisher Scientific, no. K0502) and stored at −20 °C.
To confirm library quality, a random subset of clones was isolated and subjected to genotyping PCR with primer pairs SI#157–SI#766 for the ClonMapper library or SI#2040–SI#330 for the CaTCH library. Barcode sequences of the clones were further verified by Sanger sequencing.
Barcode sequencing library preparation
CloneSelect C→T mini-pool library
To identify barcodes in the CloneSelect C→T mini-pool library by high-throughput sequencing, ~10 ng of plasmid DNA (approximately 1.0 × 109 molecules) was used as a PCR template. For identifying barcodes in the initial barcoded HEK293Ta cell population, genomic DNA was purified using NucleoSpin Tissue (Macherey-Nagel, no. 740952) according to the manufacturer’s protocol, and 119 ng of extracted genomic DNA (4 × 104 molecules, 400-fold of the estimated barcode complexity) was used as a PCR template.
The sequencing libraries were prepared using a two-step PCR method. The first-round PCR was performed in a 20 µl reaction containing template DNA, 0.5 µl each of 20 µM forward (SI#682) and reverse (SI#683) primers, 0.2 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530), 4 µl of Phusion HF Buffer (NEB, no. B0518S), 2 µl of 2 mM dNTPs (Takara, no. 4025) and 0.6 µl of 100% DMSO (NEB, no. 12611P). The thermal cycling conditions were as follows: 98 °C for 10 s; 30 cycles of 98 °C for 10 s, 61 °C for 10 s and 72 °C for 30 s; followed by a final extension at 72 °C for 5 min. Each PCR product was size-selected using 2% agarose gel and purified with the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
To add Illumina sequencing adaptors and custom indices, the second-round PCR was performed on each first-round PCR product in a 20 µl reaction containing 2.5 ng of the first PCR product, 1 µl each of 10 µM P5 and P7 custom index primers, 0.2 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530), 4 µl of Phusion HF Buffer (NEB no. B0518S), 2 µl of 2 mM dNTPs (Takara, no. 4025) and 0.6 µl of 100% DMSO (NEB, no. 12611P). The thermal cycling conditions were as follows: 98 °C for 10 s; 20 cycles of 98 °C for 10 s, 61 °C for 10 s and 72 °C for 30 s; followed by a final extension at 72 °C for 5 min. Custom indices for the second-round PCR products are listed in Supplementary Table 3. Each second-round PCR product was size-selected and purified using the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302). Sequencing samples were pooled, quantified by qPCR using the Kapa Library Quantification Kit Illumina (Kapa Biosystems, no. KK4824) and analyzed by paired-end sequencing using Illumina MiSeq.
CloneSelect C→T, CaTCH and ClonMapper Pool-10000 libraries
To identify barcodes in each Pool-10000 plasmid library by high-throughput sequencing, ~1 pg of plasmid DNA was used as a PCR template. To identify barcodes in each barcoded HEK293Ta cell population, genomic DNA was purified using the NucleoSpin Tissue Kit (Macherey-Nagel, no. 740952) according to the manufacturer’s protocol, and a total of ~2 µg of genomic DNA was used as a PCR template. Sequencing libraries were prepared using a two-step PCR method.
The first-round PCR reaction mixture was split into 20 subreactions and distributed into 20 wells of a 96-well plate for each of the two replicates. Each 25 µl subreaction contained template DNA, 1.0 µl each of 10 µM forward and reverse primers, 0.5 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530), 5 µl of 5× Phusion HF Buffer (NEB, no. B0518S) and 0.5 µl of 10 mM dNTPs (NEB, no. N0447S). For CloneSelect C→T, the primer pair and the thermal cycle conditions were the same as described above. For CaTCH, the primer pair was CS-310-PS1.0-FW4 and CS-310-PS2.0-RV1, and the thermal cycling conditions were as follows: 98 °C for 10 s; 10 cycles of 98 °C for 10 s, 67 °C for 15 s and 72 °C for 30 s; followed by 20 cycles of 98 °C for 10 s and 72 °C for 1 min; with a final extension at 72 °C for 7 min. For ClonMapper, the primer pair was PS1.0-hU6-FW5 and Scaffold-PS2.0-RV5, and the thermal cycling conditions were as follows: 98 °C for 10 s; 30 cycles of 98 °C for 10 s, 67 °C for 15 s and 72 °C for 30 s; with a final extension at 72 °C for 7 min. PCR products were pooled and purified with a 1.8× volume of Agencourt AMPure XP magnetic beads (Beckman Coulter, no. A63881) following the manufacturer’s protocol.
To add Illumina sequencing adaptors and custom indices, the second-round PCR was performed on each first-round PCR product in a 25 µl reaction containing 10 ng of the first PCR product, 0.75 µl each of 10 µM P5 and P7 custom index primers, 0.5 µl of Kapa HiFi DNA Polymerase (Kapa Biosystems, no. KK2101), 5 µl of 5× Kapa HiFi Fidelity Buffer (Kapa Biosystems, no. KK2101) and 0.75 µl of 10 mM dNTPs (NEB, no. N0447S). The thermal cycling conditions were as follows: 95 °C for 5 min; 10 cycles of 98 °C for 20 s, 67 °C for 15 s and 72 °C for 1 min; followed by 10 cycles of 98 °C for 20 s and 72 °C for 1 min; with a final extension at 72 °C for 1 min. Custom indices for the second-round PCR products are listed in Supplementary Table 3. The second-round PCR products were purified using a 1.8× volume of Agencourt AMPure XP magnetic beads (Beckman Coulter, no. A63881). Sequencing samples were pooled, quantified by qPCR using the Kapa Library Quantification Kit Illumina (Kapa Biosystems, no. KK4824) and analyzed by paired-end sequencing using Illumina MiSeq.
Sorted cells
For amplicon sequencing-based barcode identification for low-volume cells obtained by barcode-specific clone isolation in the CloneSelect C→T mini-pool assays, a cell lysate was prepared for each sample as a PCR template. Cells in eight-strip PCR tubes were first incubated with 2.0 µl of lysis buffer containing 600 mM KOH, 10 mM EDTA and 100 mM dithiothreitol. The samples were then neutralized with 2.0 µl of neutralization buffer composed of 0.4 µl of 1 M Tris-HCl and 1.6 µl of 3 M HCl. For the first-round PCR, 2.0 µl of the cell lysate was used as the template. Although no visible bands were observed on gel electrophoresis for the first-round PCR products, the PCR product of the expected size was isolated using 2% agarose gel, purified and eluted in 15 µl of ddH2O with the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
The PCR products were quantified using the Quant-iT PicoGreen dsDNA Assay Kit (Thermo Fisher Scientific, no. P7589) and the Infinite 200 PRO plate reader (TECAN) with Tecan i-control software (v.1.10.4.0). For the second-round PCR, 2.0 ng of the first-round PCR product was used as the template. Custom indices assigned to the second-round PCR products are provided in Supplementary Table 3.
The second-round PCR products were size-selected and purified with the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302). The samples were pooled into a DNA LoBind 1.5 ml tube (Eppendorf, no. 13-698-791), quantified by qPCR using the Kapa Library Quantification Kit Illumina (Kapa Biosystems, no. KK4824) and analyzed by paired-end sequencing on an Illumina MiSeq.
For the Pool-10000 assays, following cell sorting, genomic DNA was extracted using the NucleoSpin Tissue Kit (Macherey-Nagel, no. 740952). Sequencing libraries were constructed using the protocols described for the barcoded Pool-10000 cell populations.
Cell barcoding
Cells were seeded in six-well cell culture plates at a density of ~2 × 105 cells per well in 2 ml of culture medium for barcoding of cells with single barcodes, a 10 cm dish at a density of ~2 × 106 cells per dish in 10 ml of culture medium for establishing the CloneSelect C→T mini-pool cell population and a 15 cm dish at a density of ~1 × 107 cells per dish for establishing the Pool-10000 pool cell populations. The next day, a total of 500–1,000 µl transduction mix containing 2 µg ml−1 Polybrene (Sigma-Aldrich, no. TR-1003), recombinant lentivirus and cell culture medium was applied to each well alongside non-virus controls. The following day, the culture medium was replaced with fresh medium containing 2.0 µg ml−1 puromycin (Gibco, no. A1113803) or 5.0 µg ml−1 blasticidin S (Wako, no. 029-18701) to select infected cells over 2–5 days. Barcoded cell populations were maintained with 500–1,000 coverages while expanding and passaging.
After drug selection, cell viability was measured using CellTiter-Glo (Promega, no. G7570) according to the manufacturer’s protocol, and luminescence was quantified with the Infinite 200 PRO plate reader (TECAN). Background luminescence from wells without cells was subtracted from all readings.
For each condition, the infection rate was calculated as the fraction of surviving cells compared to non-selective controls. Samples with an infection rate close to but not exceeding 0.1 were used in subsequent analyses, in which most selected cells were expected to contain a single viral integration based on Poisson statistics.
Preparing Pool-10000 cell pools
After barcoding cells with the CloneSelect C→T and CaTCH Pool-10000 libraries, background EGFP reporter expression was observed in some cells. To eliminate possible false positives before the experiment, EGFP− cells were first collected by flow cytometry cell sorting while maintaining the original barcode complexity. Approximately ~8 × 106 cells, representing an average of 800 clones per barcode, were sorted for both libraries using the MoFlo Astrios (Beckman Coulter). Following sorting and expansion to ~90% confluency, genomic DNA was purified for the barcode sequencing analysis using the NucleoSpin Tissue Kit (Macherey-Nagel, no. 740952) according to the manufacturer’s protocol.
Selecting target clones for isolation from the Pool-10000 cell populations
For each CloneSelect C→T, CaTCH and ClonMapper Pool-1000 cell population, 16 clones of a diverse range in abundance were arbitrarily selected for isolation. In each population, the barcode abundance rates were grouped into four bins: (0.01, 0.02], (0.025, 0.05], (0.05, 0.1] and (0.1, 1.0]. From each bin, four target clones were randomly chosen. If a bin contained fewer than four clones, additional clones were randomly selected from the next higher bin to reach a total of 16 target barcodes for testing. The targeted clones in each assay are listed in Supplementary Table 1.
Reporter activation
For all experiments delivering a reporter activation reagent to a cell sample of a single barcode, cells were seeded in 24-well cell culture plates at a density of ~5 × 104 cells per well in 500 µl of culture 1 day before transfection. A total of 400 ng of plasmids with a 3:1 mass ratio of a Cas9 effector or decoy plasmid to gRNA plasmid, 1.2 µl of 1 mg ml−1 PEI MAX (Polysciences, no. 24765) and 100 µl of 1× PBS were combined, incubated for 5–10 min at room temperature and applied to each well (for CaTCH, 300 ng of a decoy plasmid PLVSIN-CMV-Pur and 100 ng of a gRNA plasmid were used). For the dose-dependent reporter activation assay with different Target-AID expression plasmids, transfections were performed in 24-well plates with plasmid amounts per well ranging from 50 to 800 ng. The PEI MAX volume was adjusted to 3 µl per 1 µg of plasmid.
For isolating a target clone from the CloneSelect C→T mini-pool, cells were seeded in six-well plates at a density of ~2 × 105 cells per well in 2,000 µl of culture medium 1 day before transfection. A total of 800 ng of a plasmid encoding both Target-AID and a gRNA were combined with 2.5 µl of 1 mg ml−1 PEI MAX (Polysciences, no. 24765) and 200 µl of 1× PBS, then applied to each well after a 5–10 min incubation at room temperature.
For isolating a target clone from each Pool-10000 cell population, cells were cultured in 15 cm dishes with a seeding density of approximately 2–4 × 106 cells. Then, 1 day before transfection, CloneSelect C→T and CaTCH Pool-10000 cells were seeded in 10 cm dishes at a density of ~2 × 106 cells per dish in 10 ml of culture medium. ClonMapper Pool-10000 cells were seeded in six-well plates at a density of ~2 × 105 cells per well in 2 ml of culture medium.
The following day, CloneSelect C→T Pool-10000 cells were co-transfected with 5,250 ng of the Target-AID expression plasmid (pRS0035) and 1,750 ng of the barcode-targeting gRNA plasmid using 22.5 µl of 1 mg ml−1 PEI MAX (Polysciences, no. 24765) and 300 µl of 1× PBS. CaTCH Pool-10000 cells were co-transfected with 5,250 ng of a decoy plasmid (pcDNA3.1 V5-HisA) and 1,750 ng of the barcode-targeting gRNA plasmid using 22.5 µl of 1 mg ml−1 PEI MAX and 300 µl of 1× PBS. ClonMapper Pool-10000 cells were co-transfected with 550 ng of the dCas9-VPR expression plasmid (pLV-CS-282 v2) and 450 ng of the barcode-targeting reporter plasmid using 3 µl of 1 mg ml−1 PEI MAX and 100 µl of 1× PBS. The transfection mix was incubated for ~5 min at room temperature and then applied to each sample.
Flow cytometry cell sorting
In the isolation of a target clone from the CloneSelect C→T mini-pool, cells were detached using 0.25% w/v trypsin-EDTA (Wako, no. 201-18841) 4 days after transfection of the reporter activation reagents, incubated at 37 °C for 5 min, collected into a 1.5 ml tube and centrifuged at 100g at room temperature for 5 min. Cells were then resuspended in a 5 ml polystyrene round-bottom tube (FALCON) containing 150–500 µl of 1% FBS in 1× PBS and immediately placed on ice until sorting. Sorting was conducted using the BD FACSJazz (BD Biosciences) in 1.0 drop single sort mode. Cells were initially gated using FSC-A and SSC-A, with the gate for EGFP+ cells defined by selecting those with high FITC-A intensities, which were absent in a control sample transfected with Target-AID and non-target gRNA plasmids. EGFP+ cells were sorted into eight-strip PCR tubes (Nippon Genetics, no. FG-018WF), each containing 2.5 µl of 1× PBS. For optimal recovery, the collection tube’s cell destination position was manually adjusted for each sample. Sorted cells were immediately placed on an ice-cold 96-well aluminum block. Although the rate of EGFP+ cells varied across samples, approximately 50–600 EGFP+ cells were recovered per experiment.
In the Pool-10000 experiments, cell samples of different activation reagents were each detached using 1× PBS, detached with 0.25% trypsin-EDTA, phenol red (Gibco no. 25200072) 3 days after transfection and combined into 50 ml tubes for each replicate group. The pooled cell samples were resuspended in FACS buffer (2% FBS in 1× PBS) and kept on ice before sorting.
Cell sorting was performed on a MoFlo Astrios (Beckman Coulter). Owing to a low frequency (~0.01%) of EGFP+ cells for CloneSelect C→T and CaTCH, an initial enrichment sort was performed for ~1.4 × 108 cells to increase EGFP+ cells to 20–30%. The EGFP-enriched cells were then sorted again using the purity sort mode to obtain ~5 × 103 EGFP+ cells per sample. For ClonMapper, cells were sorted directly using a purity sort mode to obtain ~3 × 105 EGFP+ cells per sample. The EGFP+ gate was defined using a non-transfected cell sample for each sample.
The raw data for cell sorting is available at https://github.com/yachielab/CloneSelect_v1/tree/main/FACS/Raw_flow_data.
Experiments using mouse ES cells
Cell culture
Under approval from the Institutional Animal Care Committee of the University of Tokyo (RAC180003), mouse ES cells were derived from embryos of a 129(+Ter)/SvJcl (female mouse) × C57BL/6NJcl (male mouse) cross and maintained in DMEM low glucose (Sigma-Aldrich, no. D6046-500ML) supplemented with 1% penicillin–streptomycin (Gibco, no. 15140122), 1% MEM non-essential amino acids (Wako, no. 139-15651), 1% GlutaMAX supplement (Gibco, no. 35050061), 1% sodium pyruvate (Gibco, no. 11360070), 15% FBS (Gibco, no. 16000044), 100 µM 2-mercaptoethanol (Wako, no. 131-14572), 1,000 units per ml ESGRO Recombinant Mouse LIF Protein (Millipore, no. ESG1107), 3.0 µM CHIR99021 (GSK-3 inhibitor) (Wako, no. 038-23101) and 1.0 µM PD0325901 (MEK inhibitor) (Tocris, no. 4423). Before seeding cells, 0.1% gelatin (Sigma-Aldrich, no. G9391) in 1× PBS (Takara, no. T9181 or Gibco, no. 70011044) was added to each well, covering the entire surface, and then aspirated after 1 h at 37 °C. Cells were cultured at 37 °C with 5% CO2 in a cell culture incubator, and the cell culture medium was replaced at least every 2 days. Cells were regularly tested for mycoplasma contamination.
Cells with stably integrated Target-AID
The mouse ES cell line with stably integrated Target-AID was established by electroporation using the NEPA21 Super Electroporator (Nepa Gene). After detaching cells from culture plate wells, ~2 × 106 cells were mixed with 100 µl of Opti-MEM (Gibco no. 31985062), 2.0 µg of pNM1325 and 0.7 µg of a Super piggyBac transposase vector (SBI, no. PB210PA-1), then transferred to an electroporation cuvette (Nepa Gene, no. EC-002S). The electroporation was performed with two poring pulses of positive polarity at 115 V for 5 ms, with 50 ms intervals and a 10% decay rate. Five transfer pulses were then applied for both positive and negative polarities at 20 V for 50 ms, with 50 ms intervals and a 40% decay rate. After electroporation, cells were transferred to a 10 cm culture dish with fresh medium, which was replaced with fresh medium again 1 day post electroporation. Then, 2 days after electroporation, the medium was replaced with medium containing 5 µg ml−1 of blasticidin S (Wako, no. 029-18701) to select cells with stable integration. Cells were incubated for about 2 weeks in the selection medium.
Transfection
Cells were seeded in 48-well cell culture plates at a density of ~6 × 104 cells in 200 µl of culture medium. For each transfection reaction, 200 ng of a gRNA plasmid was diluted in 20 µl of Opti-MEM (Gibco, no. 31985062). Separately, 0.6 µl of Lipofectamine 2000 (Invitrogen no. 11668019) was combined with 19.4 µl of Opti-MEM to form the transfection mix. The plasmid solution and transfection mix were then combined and applied to each well after a 5 min incubation at room temperature.
Barcode plasmid pool preparation
The scCloneSelect barcode library was prepared similarly to the CloneSelect C→T barcode library. An EGFP coding sequence was first amplified from pLV-CS-112 (Addgene, no. 131127) by PCR using the semi-random oligonucleotide pool SI#679 as the forward primer and RS#244 as the reverse primer. The PCR was performed in 25 separate 40 µl reactions, each containing 0.12 µl of 10 ng µl−1 pLV-CS-112 template plasmid, 2 µl each of forward and reverse primers, 0.6 µl of Phusion High-fidelity DNA Polymerase (NEB, no. M0530), 8 µl of 5× Phusion HF Buffer (NEB, no. B0518S) and 3.2 µl of 2.5 mM dNTPs (NEB, no. N0447). The thermal cycling conditions were as follows: 98 °C for 30 s; followed by 30 cycles of 98 °C for 10 s, 65 °C for 10 s and 72 °C for 60 s; with a final extension at 72 °C for 5 min.
The amplified barcode-EGFP fragment was pooled into a single 1.5 ml tube, digested with 12.5 µl of DpnI (NEB, no. R0176) at 37 °C for 1 h and size-selected using the FastGene PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302). The purified product was then subjected to overnight digestion with EcoRI-HF (NEB, no. R3101S) and XbaI (NEB, no. R0145) at 37 °C and purified again using the FastGene PCR/Gel Extraction Kit. For backbone preparation, 25 µg of the pRS193 lentiviral cloning backbone plasmid was digested with EcoRI-HF (NEB, no. R3101S) and XbaI (NEB, no. R0145) overnight at 37 °C and then size-selected with the FastGene PCR/Gel Extraction Kit.
The ligation reaction was prepared by mixing 1.25 µg of the digested backbone, 320 ng of the purified insert, 25 µl of T4 DNA Ligase (Nippon Gene, no. 317-00406) and 25 µl of 10× T4 DNA Ligase Reaction Buffer (NEB, no. B0202) in a total volume of 250 µl, followed by overnight incubation at 16 °C. The ligation mixture was then transformed into NEB Stable Competent E. coli cells (NEB, no. C3040I) across 17 reactions, each containing 4 µl of the ligation sample and 50 µl of competent cells, following the manufacturer’s high-efficiency transformation protocol. After 1 h of outgrowth in SOC medium (NEB, no. B9020) at 37 °C, cells were centrifuged and plated across 15 LB agar plates containing 100 µg ml−1 ampicillin (Wako, no. 014-23302). Colonies that formed on each plate after overnight incubation at 37 °C were scraped with 1–2 ml ddH2O. The collected cell samples were pooled and further incubated in 200–300 ml of LB liquid medium with 100 µg ml−1 ampicillin (Wako, no. 014-23302) overnight at 37 °C. A 300-fold diluted transformation sample was plated in duplicate on agar, estimating the barcode complexity at ~1.5 × 105. The plasmid library was purified using the NucleoBond Midi-prep Kit (Macherey-Nagel, no. 740410) and stored at −20 °C.
We isolated 20 random clones and verified fragment insertion by genotyping PCR with primer pair RS#147 and SI#514, confirming the expected insertion in 17 out of 20 clones. From these, we selected six clones (including three without expected genotyping bands) for double digestion with EcoRI-HF (NEB, no. R3101S) and BamHI-HF (NEB, no. R3136S), followed by Sanger sequencing using primers SI#514 and RS#147 for the uptag and dntag, respectively. All tested clones contained the expected uptag and dntag inserts.
Barcode sequencing library preparation
Uptag–dntag combination reference database
To establish the uptag–dntag combination reference database for the barcoded mouse ES cell population, genomic DNA was first extracted from ~1 × 105 cells using NucleoSpin Tissue (Macherey-Nagel no. 740952) following the manufacturer’s protocol. Sequencing libraries were prepared using a two-step PCR method, with 50 ng of genomic DNA per PCR reaction.
The first-round PCR was performed in a 20 µl reaction containing template DNA, 0.7 µl each of 10 µM forward (SI#682) and reverse (RS#250) primers, 0.2 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530), 4.5 µl of Phusion HF Buffer (NEB, no. B0518S) and 1.6 µl of 2.5 mM dNTPs (NEB, no. N0447). The thermal cycling conditions were as follows: 98 °C for 10 s; 15 cycles of 98 °C for 10 s, 60 °C for 10 s and 72 °C for 2 min; followed by a final extension at 72 °C for 5 min. Each PCR product was size-selected using 2% agarose gel, purified and eluted in 20 µl of ddH2O with the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
To add Illumina sequencing adaptors and custom indices, the second-round PCR was performed in a 20 µl reaction using a 20-fold dilution of the first-round PCR product, 0.7 µl each of 10 µM P5 and P7 custom index primers, 0.2 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530), 4.5 µl of Phusion HF Buffer (NEB, no. B0518S) and 1.6 µl of 2.5 mM dNTPs (NEB, no. N0447). The thermal cycling conditions were as follows: 98 °C for 10 s; 20 cycles of 98 °C for 10 s, 60 °C for 10 s and 72 °C for 30 s; followed by a final extension at 72 °C for 5 min. Custom indices for the second-round PCR products are listed in Supplementary Table 3. The second-round PCR products were size-selected and purified using the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302). Sequencing samples were pooled, quantified by qPCR with the Kapa Library Quantification Kit Illumina (Kapa Biosystems, no. KK4824) and analyzed by paired-end sequencing using Illumina MiSeq.
Sorted cells
For cells sorted after gRNA-dependent barcode-specific clone isolation, a cell lysate was prepared for each sample as a PCR template. The sequencing library of each sample was generated by modifying the two-step PCR method for the uptag–dntag combination reference database.
Cell samples were first expanded in 96-well culture plate wells until confluent. After aspirating the culture medium, 20 µl of 50 mM NaOH was added to each well, and the contents were transferred to a 96-well PCR plate for direct cell lysis. The samples were then heated at 95 °C for 15 min and cooled on ice, followed by neutralization with 2.0 µl of 1 M Tris-HCl (pH 8.0).
The first-round PCR was performed in a 40 µl reaction, with 3.5 µl of cell lysate as the template. The second-round PCR was performed in a 20 µl reaction, using a tenfold dilution of the first-round PCR product as the template. Custom indices assigned to the second-round PCR products are provided in Supplementary Table 3. The second-round PCR products were size-selected and purified using the GeneJET Gel Extraction Kit (Thermo Fisher Scientific, no. K0691). Sequencing samples were pooled into a DNA LoBind 1.5 ml tube (Eppendorf, no. 0030108051), quantified by qPCR with the Kapa Library Quantification Kit Illumina (Kapa Biosystems, no. KK4824) and analyzed by paired-end sequencing using Illumina HiSeq 2500.
Reamplification of dntags from Drop-seq library
To increase the sensitivity of identifying dntags associated with single-cell transcriptome profiles, the DNA region encoding dntags and cell IDs were selectively reamplified from the intermediate Tn5 transposon-fragmented sample of the Drop-seq process and sequenced separately.
The reamplification PCR was performed in a 20 µl reaction containing 1 ng of template DNA (quantified using TapeStation with High Sensitivity D5000 ScreenTape; Agilent, nos. 5067-559 and 5067-5593), 0.7 µl each of 20 µM forward primer P5-TSO_Hybrid43 and reverse primer SI#682, 0.2 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530), 4.5 µl of 5× Phusion HF Buffer (NEB, no. B0518) and 1.6 µl of 2.5 mM dNTPs (NEB no. N0447). The thermal cycling conditions were as follows: 95 °C for 30 s; 30 cycles of 98 °C for 30 s, 60 °C for 10 s and 72 °C for 2 min; followed by a final extension at 72 °C for 5 min. The first-round PCR product was purified and eluted in 20 µl of ddH2O using the GeneJET Gel Extraction Kit (Thermo Fisher Scientific, no. K0691).
The second-round PCR was performed in a 20 µl reaction using a tenfold dilution of the first-round PCR product, 0.7 µl each of 10 µM P5 and P7 custom dual index primers, 0.2 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530), 4.5 µl of 5× Phusion HF Buffer (NEB, no. B0518) and 1.6 µl of 2.5 mM dNTPs (NEB, no. N0447). The thermal cycling conditions were as follows: 95 °C for 30 s; 15 cycles of 98 °C for 10 s, 65 °C for 10 s and 72 °C for 2 min; followed by a final extension at 72 °C for 5 min. Custom indices for the second-round PCR products are listed in Supplementary Table 3. The second-round PCR products were size-selected using 2% agarose gel, purified with the GeneJET Gel Extraction Kit (Thermo Fisher Scientific, no. K0691), pooled, quantified by qPCR using the Kapa Library Quantification Kit Illumina (Kapa Biosystems, no. KK4824) and analyzed by paired-end sequencing using Illumina HiSeq 2500.
Cell barcoding
To introduce a single barcode, cells with or without stably integrated Target-AID were seeded in six-well cell culture plates at a density of ~2 × 105 cells per well in 2 ml of culture medium 1 day before transduction. A recombinant virus sample with a 10–100 µl volume was thawed on ice, mixed with 1.5 µl of 8 µg ml−1 Polybrene (Sigma-Aldrich, no. TR-1003) and 1.5 ml of fresh culture medium and then applied to the cells. To select transduced cells, the culture medium was replaced with a fresh medium containing 1.0 µg ml−1 puromycin (Gibco no. A1113803) 2 days after infection, followed by an additional 3 days of incubation. Surviving cells were detached, and cell counts were measured using an automated cell counter (BioRad TC20). The infection rate was calculated as the fraction of surviving cells compared to the non-selective control condition. Samples with an infection rate close to but not exceeding 0.1 were used in subsequent analyses.
For the barcoding of a cell population, cells with stably integrated Target-AID were seeded in six-well cell culture plate wells at a density of ~2 × 105 cells per well with 2 ml of culture medium 1 day before transduction. The following day, cells were transduced with 500 µl of a 15-fold concentrated barcoding lentivirus pool using the same transduction protocol and selected 2 days after infection. For the downstream proof-of-principle differentiation and clone isolation assays, a clonal population bottleneck was created by seeding ~1,000 cells in a single six-well plate and culturing them for 10 days.
Mouse ES cell differentiation assay
The barcoded cell population with the clonal complexity bottleneck was then divided as follows: ~1 × 104 cells were seeded into culture medium with LIF and 2i (1.0 µM PD0325901; Tocris, no. 4423 and 3.0 µM CHIR99021; Wako, no. 038-23101) (LIF+2i+), ~1 × 104 cells were seeded into culture medium without LIF or 2i (LIF−2i−), two samples of ~1 × 105 cells each were set aside to establish the uptag–dntag combination reference database and five replicates of ~1 × 105 cells were stored at −80 °C in CELLBANKER 1 freeze medium (ZENOAQ, no. 11910). Then, 4 days later, cells in both the LIF+2i+ and LIF−2i− conditions were subjected to scRNA-seq.
Drop-seq
scRNA-seq was performed by Drop-seq with devices manufactured by Dolomite Bio according to the manufacturer’s protocol. Microfluidic devices were fabricated by YODAKA. Cell samples were prepared at a concentration of ~2 × 105 cells per ml for analysis.
Sequencing libraries were prepared following the original Drop-seq protocol43. In brief, after emulsion breakage and reverse transcription, ‘single-cell transcriptomes attached to microparticles’ (STAMPs) were washed and treated with Exonuclease I (NEB, no. M0293L). Approximately 2,000 STAMPs were used for the whole cDNA amplification of each sample. Following second-strand synthesis, library DNA was purified with AMPure XP beads (Beckman Coulter, no. A63881), quantified using a TapeStation with High Sensitivity D5000 ScreenTape (Agilent, nos. 5067-5592 and 5067-5593) and fragmented with Tn5 transposon using the Nextera XT DNA Library Preparation Kit (Illumina, no. FC-131-1024) as per the manufacturer’s protocol. The fragmented sequencing library was purified with AMPure XP beads (Beckman Coulter, no. A63881) and quantified again using the TapeStation with High Sensitivity D5000 ScreenTape (Agilent, nos. 5067-5592 and 5067-5593). Each library’s average size was confirmed to be ~500 bp. Multiple scRNA-seq libraries were pooled and subjected to high-throughput sequencing using Illumina MiSeq or HiSeq 2500. The sequencing library index information is provided in Supplementary Table 3.
RT–PCR
The transcription of polyadenylated scCloneSelect barcode products was assessed by PCR with reverse transcription (RT–PCR) and gel electrophoresis. Total RNA was extracted using the ISOSPIN Cell & Tissue RNA Kit (Nippon Gene, no. 314-08211) according to the manufacturer’s instructions. The RNA was then treated with DNase I (Takara, no. 2270B) to eliminate residual DNA and purified again using the ISOSPIN Cell & Tissue RNA Kit (Nippon Gene, no. 314-08211).
First-strand cDNA was synthesized using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, no. 4368814) in a 10 µl reaction volume containing 5 µl of DNase I-treated RNA (~1 µg), 0.5 µl of 100 µM oligonucleotide dT primer SI#4, 0.5 µl of MultiScribe Reverse Transcriptase, 1 µl of 10× RT buffer, 0.4 µl of 100 mM dNTPs and 0.5 µl of RNase Inhibitor (Applied Biosystems, no. N8080119). The thermal cycling conditions were as follows: 25 °C for 10 min, 37 °C for 12 min and 85 °C for 5 min.
The transcription of the target barcode was then analyzed by PCR alongside a GAPDH control. Each PCR reaction was conducted in a 20 µl volume, containing 2 µl of 50-fold diluted first-strand cDNA, 2.8 µl total of either the primer pair SI#116–SI#7 to amplify the dntag or the primer pair RS#507–RS#508 to amplify GAPDH, 0.2 µl of Phusion DNA Polymerase (NEB, no. M0530S), 4 µl of 5× Phusion HF Buffer (NEB, no. B0518) and 1.6 µl of 2.5 mM dNTPs (NEB, no. N0447). The thermal cycling conditions were as follows: 98 °C for 30 s; 30 cycles of 98 °C for 10 s, 60 °C for 10 s and 72 °C for 30 s; followed by a final extension at 72 °C for 5 min. The PCR products were analyzed on a 2% agarose gel.
Flow cytometry cell sorting
Each cell sample was expanded in a 10 cm cell culture dish 3 days after transduction with a query gRNA. Cells were detached with 0.25% w/v trypsin-EDTA (Gibco, no. 25200072), incubated at 37 °C for 5 min, collected into a 1.5 ml tube and centrifuged at 100g at room temperature for 5 min. The cells were then resuspended to approximately 1 × 106 cells in PBS containing 2% FBS and transferred to a 5 ml polystyrene round-bottom tube (Falcon, no. 352054). The cell suspension was immediately placed on ice until sorting.
Sorting was conducted using MoFlo Astrios EQ Cell Sorter (Beckman Coulter). Cells were initially gated using FSC-A and SSC-A, with the gate for EGFP+ cells set to include those with high FITC-A intensities, which were absent in a non-transduced control sample. EGFP+ cells were single-cell sorted into 96-well plate wells, while the remaining cells were sorted in bulk into a single well of a 96-well plate, each containing 100 µl of mouse ES cell culture medium. Approximately 100–1,000 EGFP+ cells were recovered per experiment, except for clone 153, for which no EGFP+ cells above the gating threshold were observed.
The raw data for cell sorting is available at https://github.com/yachielab/CloneSelect_v1/tree/main/FACS/Raw_flow_data.
Barcode analysis
To identify uptag and dntag barcodes in a cell population, sequencing reads were first demultiplexed, and cutadapt (v.4.1) (https://github.com/marcelm/cutadapt) was used to extract uptag and dntag sequences located between their 20 bp upstream and downstream constant sequences. Extracted uptags and dntags were filtered with a Q-score threshold of 30, then clustered and further filtered by length (17 bp for uptags and 30 bp for dntags) using bartender-1.1 (https://github.com/LaoZZZZZ/bartender-1.1)75.
In constructing the uptag–dntag combination reference database, redundant uptag–dntag pairs with either uptag or dntag found in more abundant pairs were discarded. For mapping dntags to the uptag–dntag database, symspellpy (v.6.7) (https://github.com/mammothb/symspellpy) was used to find the match with the shortest edit distance. If multiple dntags with the same edit distance were found, the dntag with the highest frequency in the database was selected.
To analyze uptag frequencies in cell populations following gRNA-dependent barcode-specific EGFP reporter activation and flow cytometry sorting, sequencing reads were mapped to the uptag–dntag database, and uptag read counts were obtained using bartender-1.1 and symspellpy (v.6.7).
The codes used for the barcode identification are available at https://github.com/yachielab/CloneSelect_v1/tree/main/Barcode_identification/scCloneSelect.
Drop-seq data analysis
After sample demultiplexing of Illumina sequencing reads, FASTQ files were processed with Drop-seq Tools (v.2.5.1) (https://github.com/broadinstitute/Drop-seq) for base quality filtering, adaptor trimming and extraction of cell ID and unique molecular identifier sequences.
Picard (v.2.18.14) (https://github.com/broadinstitute/picard) was used to convert BAM files back to FASTQ files for subsequent steps. Filtered reads were aligned using STAR (v.2.7) (https://github.com/alexdobin/STAR)76 with the mm10 reference genome.
Differential gene expression and clustering analyses were conducted using Seurat (v.3) (https://github.com/satijalab/seurat)77. Cells were filtered based on thresholds of Feature_RNA > 200, nFeature_RNA < 2500 and percent.mt < 5, and gene expression profiles were normalized using the Seurat::sctransform function before clustering.
To identify dntags for analysis, we performed an initial Drop-seq run and determined dntags based on the cumulative read count distribution of cell IDs, with a threshold set at the knee point using the Python package kneed (v.0.8.1) (https://github.com/arvkevi/kneed). For higher sensitivity in mapping dntag distributions to single-cell transcriptome data, we also sequenced the reamplified dntag library and used the dntag–uptag combination reference database to identify cell ID and dntag associations, as described for the scCloneSelect library preparation. When multiple dntags were associated with a single cell ID, the dntag with the highest unique molecular identifier count was selected.
The codes are available at https://github.com/yachielab/CloneSelect_v1/tree/main/Drop-seq.
Experiments using human PS cells
Cell culture
The CA1 human PS cell line was used with approval from the Canadian Institutes of Health Research Stem Cell Oversight Committee. CA1 human PS cells were cultured in mTeSR Plus medium (STEMCELL Technologies, no. 100-0276) in a humidified incubator at 37 °C with 21% O2 and 5% CO2. Culture plates were coated with Geltrex LDEV-Free Reduced Growth Factor Basement Membrane Matrix (Gibco, no. A1413201). To prepare the Geltrex working solution, DMEM/Nutrient Mixture F-12 (DMEM/F-12) (Gibco, no. 11320033) was diluted 1:100 with Geltrex. A sufficient volume of this solution was added to each well, covering the surface, and was aspirated after 1 h of incubation at 37 °C before plating cells. The cell culture medium was replaced every other day after cell seeding. Cells were routinely passaged as medium-sized clumps. After aspirating the medium, ReLeSR (STEMCELL Technologies, no. 05872) was added, and cells were incubated at room temperature for about 1 min before a second aspiration. Cells were then placed in the incubator for 4–5 min, fresh medium was added and cells were dissociated by gentle pipetting. The cells were then plated and returned to the incubator.
For single-cell passaging, TrypLE Express (Gibco, no. 12604021) was used. The cells were incubated for 4 min before adding fresh medium to stop the action of TrypLE Express. Cells were collected in centrifuge tubes, dissociated by pipetting and filtered through a 40 µm cell strainer (Sarstedt, no. 83.3945.040) to remove clumps. Tubes were centrifuged at 300–400g for 5 min, and the supernatant was aspirated. Pellets were resuspended in fresh medium supplemented with 10 µM ROCK inhibitor Y-27632 (Tocris Bioscience, no. 1254) for 24 h to support single-cell survival.
For culturing H1 human PS cells, we used StemFit AK02N medium (REPROCELL AHS, no. RCAK02N), with Y-27632 (Cayman, no. 10005583) added for 1–2 days after plating. Culture plates were coated with recombinant Laminin-511 E8 fragment using iMatrix-511 Silk (MAX, no. 892021).
The Center for iPS Cell Research and Application (CiRA) Ethics Committee, an internal committee at Kyoto University’s CiRA, approved our research plan for human ES cell research (CiRA21-03) and recombinant DNA experiments (240283). The WiCell line H1 (WA01) was used under agreements 10-WO-0098, 23-W0713 and 24-W0434.
Cells were regularly tested for mycoplasma contamination.
Cells with stably integrated Target-AID
To establish a human PS cell line with stably integrated Target-AID, CA1 cells were seeded in 24-well cell culture plates at a density of ~5 × 104 cells per well in 1 ml of culture medium 1 day before transfection. The transfection mix was prepared by combining 450 ng of pNM1325 (CAGp-Target-AID-2A-Blast), 50 ng of a hyperactive piggyBac transposase plasmid, 1 µl of Lipofectamine Stem Transfection Reagent (Invitrogen, no. STEM00001) and 49 µl of Opti-MEM (Gibco, no. 31985062) and was applied to the wells after 10 min of incubation. The following day, the culture medium was replaced with fresh medium to remove residual transfection reagent. Then, 3 days post transfection, the medium was replaced with fresh medium containing 5 µg ml−1 of blasticidin S to initiate selection for 24 h. An additional two-day selection was performed until cells reached confluency, at which point they were passaged into a new culture plate. A final selection round was conducted to ensure the selection of the cells.
Cell barcoding
For the introduction of a single barcode, cells with or without stably integrated Target-AID, cells were seeded in six-well cell culture plates at a density of ~1 × 105 cells per well in 2 ml of culture medium 1 day before transduction. For transduction, recombinant virus samples with a volume of 10–100 µl were thawed on ice, mixed with 1.5 µl of 8 µg ml−1 Polybrene (Sigma-Aldrich, no. TR-1003) and 1.5 ml of fresh culture medium and then applied to the cells. After 48 h of infection, the culture medium was replaced with fresh medium containing 1.0 µg ml−1 puromycin (Gibco, no. A1113803) for 3 days. The reporter-integrated cells were then dissociated into single cells and subjected to flow cytometry sorting to enrich EGFP− cells. The sorted cells were maintained in StemFit AK02N culture medium (REPROCELL, no. RCAK02N).
For the barcoding of the H1 cell population, cells were seeded at a density of ~2.1 × 104 cells per cm2 1 day before transduction. The following day, freshly prepared medium containing 50 µl of the barcoded virus library and 2 µl of 8 mg ml−1 Polybrene (Nacalai Tesque, no. 12996-81) was added to each well. After 48 h, the culture medium was replaced with fresh medium containing 1.0 µg ml puromycin (Life Technologies, no. A1113802) for 3 days to select for transduced cells. Following puromycin selection, EGFP− cells were enriched by flow cytometry cell sorting using the BD FACS Aria (BD Biosciences).
Naive induction
Barcoded cells were seeded at a density of ~1.6 × 104 cells per cm2 with iMatrix-511 silk (MATRIXOME, no. 387-10131) in StemFit AK02N (Ajinomoto, no. RCAK02N). After 48 h, naïve induction was initiated with cRM-1 + Y culture medium (designated as day 0), consisting of NDiff 227 (Takara Bio, no. Y40002) supplemented with 1 µM PD0325901 (Tocris, no. 4192), 10 ng ml−1 Recombinant Human LIF (Peprotech, no. 300-05), 1 mM valproic acid (Sigma-Aldrich, no. P4543) and 10 µM Y-27632 (Cayman, no. CAY-10005583-50). Then, 2 days later, the culture medium was switched to PXGL + Y medium, composed of NDiff 227 (Takara Bio, no. Y40002) with added 1 µM PD0325901 (Tocris, no. 4192), 10 ng ml−1 Recombinant Human LIF (Peprotech, no. 300-05), 2 µM Go 6983 (Tocris, no. 2285), 2 µM XAV-939 (Selleck, no. S1180) and 10 µM Y-27632 (Cayman, no. CAY-10005583-50). Cells were passaged using TrypLE Express Enzyme (Invitrogen, no. 12604021) and Enzyme Free Cell Dissociation Solution (Sigma-Aldrich, no. S-014-B) and cultured in the same medium for 23–25 days.
Reporter activation
Reporter activation assays using CA1 human PS cells
To activate the reporter of a barcoded CA1 human PS cell sample with stably integrated Target-AID, we used the Neon Transfection System (Invitrogen, no. MPK5000) to deliver the gRNA plasmid by electroporation. Cells were detached from culture plate wells, and ~1 × 105 cells were mixed with 100 µl of Neon Resuspension Buffer (Invitrogen, no. MPK10096) and 2.0 µg of gRNA plasmid. Electroporation was performed with the following settings: 1,200 V, 30 ms, single pulse.
Reporter activation assays using H1 human PS cells
To activate the reporter of a barcoded H1 human PS cell sample, Target-AID and gRNA expression plasmids were co-delivered by electroporation. Cells were detached from culture plate wells, and ~1 × 105 cells were mixed with 100 µl of Neon Resuspension Buffer (Invitrogen, no. MPK10096), 3.0 µg of Target-AID plasmid and 3.0 µg of gRNA plasmid. Electroporation was performed with the following settings: 1,200 V, 20 ms, two pulses.
Elite clone isolation from the barcoded H1 human PS cell clone population
To isolate a target clone from the barcoded H1 human PS cell population, Target-AID and gRNA expression plasmids were also co-delivered by electroporation using the Neon Transfection System (Invitrogen, no. MPK5000). Cells were detached with Accutase (Sigma-Aldrich, no. A6964-500ML), and ~2.0 × 106 cells were transferred to a 1.5 ml tube. The cells were washed once with 1× D-PBS (-) (Nacalai Tesque, no. 14249-24) and resuspended in 100 µl of Neon Resuspension Buffer (Invitrogen, no. MPK10096) containing 9 µg of the Target-AID plasmid and 6 µg of the gRNA plasmid. Electroporation was performed with the following settings: 1,200 V, 20 ms, two pulses.
Flow cytometry cell sorting
Cell samples were washed with 1× D-PBS (-) and detached using 2 ml of Accutase (Sigma-Aldrich, no. A6964-500ML) to create a single-cell suspension. Cells were resuspended in FACS buffer composed of 450 ml MilliQ water, 50 ml 10× Hanks’ Balanced Salt Solution (no calcium, no magnesium, no phenol red) (Invitrogen, no. 14185052) and 5 g BSA (Sigma-Aldrich, no. A2153-100G) and kept on ice for 30 min.
Immunostaining was conducted on ice with antibodies in FACS buffer for 30 min. Flow cytometry and cell sorting were performed using the BD LSR Fortessa or FACS Aria II systems (BD Bioscience). The following antibodies were used: anti-human SUSD2 antibody (PE) (Biolegend, no. 327406; 1:200 dilution), CD24 monoclonal antibody (APC) (Thermo Fisher Scientific, no. 17-0247-42; 1:200 dilution), TROP2 antibody, anti-human, REAfinity (Biotin) (Miltenyi Biotec, no. 130-115-054; 1:200 dilution), mouse anti-human CD249 (BV421) (BD Bioscience, no. 744872; 1:200 dilution) and APC streptavidin (Biolegend, no. 405207; 1:1,000 dilution). Data analysis was conducted with FlowJo (v.10.7.2).
The raw data for cell sorting is available at https://github.com/yachielab/CloneSelect_v1/tree/main/FACS/Raw_flow_data.
Trophoblast differentiation
The protocol for trophoblast differentiation was previously established and described78. In brief, H1 naïve stem cells were seeded at a density of ~2.0 × 104 cells per cm2 onto iMatrix-511 silk in NDiff 227 medium supplemented with 2 µM A 83-01 (Tocris, no. 2939), 2 µM PD0325901 and 10 ng ml−1 BMP-4 (R&D, no. 314-BP-500). The following day, the medium was replaced with NDiff 227 supplemented with 2 µM A 83-01, 2 µM PD0325901 and 1 µM JAK Inhibitor I (Calbiochem, no. 420099). On day 3, cells were detached using Accutase (Sigma-Aldrich, no. A6964-500ML), immunostained with anti-human TROP2 (Miltenyi Biotec, no. 130-115-054) and anti-human CD249 (BD Bioscience, no. 744872) and then sorted using the BD LSR Fortessa or FACS Aria II systems (BD Bioscience). Trophoblast marker genes used in this study were curated from a previous report78.
qPCR
HAVCR1+/ENPEP+ cells were subjected to total RNA extraction using the Quick-RNA Kit Micro-Prep (ZYMO, no. R1051). Total RNA (0.5 µg) was reverse-transcribed into cDNA with an oligonucleotide dT primer using SuperScript IV (Invitrogen, no. 18090050). qPCR was conducted using PowerUP SYBR Green Master Mix (Applied Biosystems no. A25743), following the manufacturer’s instructions. Results were analyzed with QuantStudio Design & Analysis Software (Thermo Fisher Scientific, v.1.4.1). Cycle threshold values were normalized to GAPDH to calculate the relative expression of trophoblast marker genes. Primer pairs used for qPCR are listed in Supplementary Table 2.
RNA-seq
Sequencing library preparation
RNA-seq libraries were prepared from 1 ng of total RNA using the SMART-Seq HT Kit (Takara, no. Z4436N) following the manufacturer’s instructions. Sequencing libraries were pooled with PhiX Control (v.3) (Illumina, no. FC-110-3001) and sequenced using Illumina NovaSeq 6000 with paired-end sequencing.
Data processing
RNA-seq reads were trimmed to remove adaptor sequences and low-quality bases using cutadapt (v.4.1) (https://github.com/marcelm/cutadapt). The trimmed reads were then aligned to the human reference genome (hg38) with STAR (v.2.7.10a)76. Read counts for each gene were obtained from the resulting BAM files using HTSeq (v.2.0.2).
Differential gene expression analysis was performed with DESeq2 (v.1.34.0)79 in R (v.4.1.1). The differentially expressed genes were identified from the DESeq2 output with an adjusted P value threshold of 0.05. To obtain normalized gene expression data for z-score standardization, a regularized log transformation was applied using the rlog function.
For visualizing read mapping in Integrated Genomics Viewer (IGV; v.2.16.2)80, BAM files were converted to BigWig format using the bamCoverage command in deepTools (v.3.5.4)81. Gene expression matrices were further processed for hierarchical clustering and visualization with the pheatmap package (v.1.0.12) (https://github.com/raivokolde/pheatmap) in R (v.4.3.1).
GSEA
To identify robust gene expression signatures in the isolated clones, clone 006, clone 034, clone 116, clone 216 and clone 332 were grouped as case samples, while the wild-type and barcoded wild-type samples were grouped as control samples. GSEA was performed on the log-transformed gene expression data using GSEApy (v.1.1.1)82. GSEA was conducted against the ‘GO_Biological_Process_2023’ gene set using the gseapy.gse function, and the enriched Gene Ontology terms were filtered with a false discovery rate threshold of 0.1. The Gene Ontology term database was obtained from the Enrichr website83.
The resulting GSEA data was converted to a graph structure using the Gene Ontology database go-basic.obo (release date 2024-01-17), obonet (v.1.0.0) (https://github.com/dhimmel/obonet) and networkx (v.3.2.1) (https://github.com/networkx/networkx) on Python (v.3.10.0). Cytoscape (v.3.10.1)84 was used for visualization.
EM-seq
Sequencing library preparation
EM-seq libraries were constructed according to the original protocol54. Genomic DNA was purified from ~1.0 × 106 input cells using the Wizard Genomic DNA Purification Kit (Promega, no. A1120). DNA concentration was quantified with the Qubit 1× dsDNA High Sensitivity Assay Kit (Thermo Fisher Scientific, no. Q33231), and 200 ng of genomic DNA was mixed with 20 pg of unmethylated lambda DNA and 1 pg of CpG-methylated pUC19 as internal controls.
The mixed DNA was fragmented using a Covaris E220 focused-ultrasonicator with the following settings: peak incident power at 175 W, duty factor at 10%, cycles per burst at 140, treatment time of 90 s and temperature range between 0 and 40 °C. DNA fragment size was verified with the High Sensitivity D5000 ScreenTape Assay (Agilent, no. 5067-5588) on the 4200 TapeStation System (Agilent), confirming predominant fragment sizes between 150 and 600 bp.
Library preparation followed the standard NEBNext Enzymatic Methyl-seq Kit protocol (no. E7120). After end-repair, A-tailing and EM-seq adaptor ligation, 5-methylcytosines and 5-hydroxymethylcytosines were oxidized with TET2 and deaminated with APOBEC. The library was then PCR-amplified and purified. Quantification was conducted using the High Sensitivity D5000 ScreenTape Assay Kit (Agilent, no. 5067-5588) on the 4200 TapeStation System (Agilent) and Qubit 1× dsDNA High Sensitivity Assay Kit (Thermo Fisher Scientific, no. Q33231). All EM-seq libraries were pooled with PhiX Control (v.3) (Illumina, no. FC-110-3001) and sequenced using Illumina NovaSeq 6000.
Data processing
EM-seq adaptor sequences were trimmed, and low-quality reads were discarded using Trim Galore (v.0.6.10) (https://github.com/FelixKrueger/TrimGalore). The processed reads were aligned to the human reference genome (hg38) with Bismark (v.0.24.1)85. The aligned reads were deduplicated using the deduplicate_bismark command, and methylated bases were called with the bismark_methylation_extractor command, applying the options --ignore 2, --ignore_r2 2 and --ignore_3prime_r2 3 to minimize methylation biases near the read ends.
BedGraph files for methylated bases were generated using the bismark2bedGraph command with the options --CX and --cutoff 3. Methylation reports for each nucleotide context were computed using the coverage2cytosine command with the --CX option. To visualize the methylation profile in IGV, we extracted cytosines in the CpG context and calculated the proportion of methylated cytosines in 500 bp bins with read counts of >20. The data were then converted into BigWig format using bedGraphToBigWig (v.2.10) (https://github.com/ENCODE-DCC/kentUtils). Unless otherwise specified, default settings were applied for all commands.
Methylation profiling was conducted with methylKit (v.0.9.7)86. Cytosines in the CpG context with a minimum coverage of three reads were extracted, and the reference genome was divided into 1,000-bp windows. Bins with fewer than ten reads were discarded. The binned CpG profiles were subjected to differential methylation analysis between the sorted clones and their corresponding parental samples using the calculateDiffMeth function in methylKit. Differentially methylated bins were extracted with a SLIM-adjusted P value threshold of <0.01 and a >25% change in methylation level.
Relative methylation levels in each bin for the sorted clones and their parental samples clones were calculated using the pyBigWig library (v.0.3.22) (https://github.com/deeptools/pyBigWig) on Python (v.3.10.0), and the resulting BigWig files were visualized in IGV (v.2.16.2)80.
Experiments using yeast
Strains
S. cerevisiae BY4741 (MATa his3∆0 leu2∆0 met15∆0 ura3∆0) was used for the yeast CloneSelect experiments.
Barcode plasmid pool preparation
To generate the yeast CloneSelect barcode library, a semi-random oligonucleotide pool, KI#200, encoding 5′-CCGWSNSWSNSWSNSWSNSNGTG-3′, was chemically synthesized (Supplementary Table 2) and amplified by PCR in a 40 µl reaction containing 2 µl of 0.01 µM template, 2 µl each of 10 µM forward primer SI#368 and 10 µM reverse primer SI#369, 0.8 µl of Phusion High-fidelity DNA Polymerase (NEB, no. M0530S), 8 µl of 5× Phusion HF Buffer (NEB, no. B0518S) and 2 µl of 2 mM dNTPs. The thermal cycling conditions were as follows: 98 °C for 30 s; 35 cycles of 98 °C for 10 s, 68 °C for 20 s and 72 °C for 5 s; followed by a final extension at 72 °C for 5 min. The PCR product was analyzed on a 2% agarose gel, size-selected and purified using the FastGene PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
The purified barcode fragment was then assembled into the cloning backbone plasmid pKI110 by Golden Gate Assembly using BsmBI (NEB, no. R0580S). Two assembly reactions were performed, each in a 25 µl volume containing 500 fmol barcode fragments, 50 fmol backbone plasmid, 0.5 µl of BsmBI (NEB, no. R0580S), 0.5 µl of T4 DNA Ligase (Nippon Gene, no. 317-00406), 2.5 µl of 10× T4 DNA Ligation Reaction Buffer (NEB, no. B0202S) and 0.125 µl of 60 mg ml−1 BSA (NEB, no. B9001S). The thermal cycling conditions were as follows: 15 cycles of 37 °C for 5 min and 20 °C for 5 min, followed by 55 °C for 30 min for complete backbone digestion.
For bacterial transformation, 5 µl of the assembly product was used to transform 50 µl of DH5α chemically competent cells (NEB, no. C2987I) following the manufacturer’s high-efficiency transformation protocol. After a 1 h outgrowth in 1 ml of SOC medium (NEB, no. B9020S) at 37 °C, cells were plated on four LB agar plates containing 100 µg ml−1 ampicillin (Wako, no. 014-23302). Diluted samples were also plated to estimate clone complexity. Random clones were isolated and analyzed by genotyping PCR using primers KI#169 and KI#170 to validate the presence of the expected barcode insert.
To construct the Pool-100 plasmid pool, 100 colonies were isolated, dissolved in 80 µl of LB medium containing 100 µg ml−1 ampicillin, combined in 5 µl aliquots and cultured overnight at 37 °C. Plasmid DNA was extracted using the FastGene Plasmid Mini Kit (Nippon Genetics, no. FG-90502). The Pool-1580 was constructed by scraping colonies from a plate with ~1,000 colony-forming units into 1.5 ml LB medium with 100 µg ml−1 ampicillin. Cells were centrifuged at 15,000g for 2 min, and the supernatant was discarded. Plasmid DNA pools were then purified from the collected cells using the FastGene PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
Barcoding of cells and introduction of genome editing reagents
For barcoding cells and introducing genome editing reagents, we used the Frozen-EZ Yeast Transformation II kit (Zymo Research, no. T2001) with slight modifications. Cells were initially pre-cultured in 5 ml of YPDA or SC–Dropout medium (adjusted to meet auxotrophic requirements for plasmid maintenance) in a cell culture tube rotating overnight at 30 °C. The following day, cells were cultured in 5 ml of fresh YPDA medium with a starting optical density at 600 nm (OD600) of 0.3 and incubated until the OD600 reached 0.8–1.0. After preparing competent cells according to the manufacturer’s protocol, plasmid DNA and 50 µl of competent cells were added to a 1.5 ml tube, mixed thoroughly with 500 µl of EZ3 solution as per the manufacturer’s protocol and incubated at 30 °C for 1 h with rotation. The cell sample was then centrifuged at 15,000g for 2 min, and the supernatant was discarded. For recovery, 2.5 ml of YPDA medium was added, and cells were allowed a 2 h outgrowth at 30 °C with rotation. After recovery, cells were centrifuged, the medium was removed and the cells were washed twice with 1 ml of TE buffer. Finally, cells were spread on SC–Dropout agar plates and incubated for 2–4 days at 30 °C.
Barcoding of cells
When the background BY4741 cells were transformed with the barcode plasmid library containing the HIS3 marker, YPDA medium was used for pre-culturing, and SC–His+Ade plates were used for selecting transformants. For pooled cell barcoding, the reaction was scaled up to transform 250 µl of competent cells using 200 ng of plasmid DNA. Colonies that formed on selective plates were pooled and collected by scraping with 3–4 ml of SC–His+Ade medium. For barcoding cells with a single barcode plasmid clone, 200 ng of plasmid was used to transform 15 µl of competent cells.
Introduction of genome editing reagents
When cells containing the barcode plasmid with the HIS3 marker were subjected to clone isolation, they underwent two rounds of transformation: first with the constitutively active Target-AID plasmid pKI086 containing the LEU2 marker and then with the targeting gRNA expression plasmid containing the URA3 marker. For the first transformation, cells were pre-cultured in SC–His+Ade medium and selected on SC–His–Leu+Ade plates. For the second transformation, cells were pre-cultured in SC–His–Leu+Ade medium and selected on SC–His–Leu–Ura+Ade plates.
When transforming the background BY4741 cells with one of the galactose-inducible Cas9-based enzyme plasmids (Cas9, dCas9, dCas9-PmCDA1, dCas9-PmCDA1-UGI, nCas9, nCas9-PmCDA1 or nCas9-PmCDA1-UGI) containing the LEU2 marker along with a CAN1-targeting gRNA plasmid containing the URA3 marker, YPDA medium was used for pre-culturing, and transformants were selected on SC–Leu–Ura+Ade plates.
For barcode-specific reporter activation within a complex barcoded population, the reaction was scaled up to transform 250 µl of competent cells with 200 ng of the enzyme plasmid and 200 ng of the targeting gRNA plasmid. For smaller-scale transformations, 200 ng of the enzyme plasmid and 200 ng of the target gRNA plasmid were used to transform 15 µl of competent cells.
Barcode sequencing library preparation
The barcode sequencing libraries of the plasmid DNA pools were prepared using a two-step PCR method. The first-round PCR was performed in a 40 µl volume, containing 1.0 µg of template DNA, 1 µl each of 10 µM forward primer KI#169 and 10 µM reverse primer KI#289, 0.4 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530S), 8 µl of Phusion HF Buffer (NEB, no. B0518S) and 0.8 µl of 10 mM dNTPs (Takara, no. 4030). The thermal cycling conditions were as follows: 98 °C for 30 s; 20 cycles of 98 °C for 10 s, 61 °C for 20 s and 72 °C for 25 s; with a final extension at 72 °C for 5 min. Each PCR product was size-selected on a 2% agarose gel, purified and eluted into 50 µl of ddH2O using the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
To add Illumina sequencing adaptors and custom indices, a second-round PCR was performed in a 40 µl volume containing 2 µl of the first-round product, 1 µl each of 10 µM P5 and P7 custom index primers, 0.4 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530S), 8 µl of 5× Phusion HF Buffer (NEB, no. B0518S) and 0.8 µl of 10 mM dNTPs (Takara, no. 4030). The thermal cycling conditions were as follows: 98 °C for 30 s; 15 cycles of 98 °C for 10 s, 60 °C for 10 s and 72 °C for 1 min; followed by a final extension at 72 °C for 5 min. Custom indices for the second-round PCR products are listed in Supplementary Table 3. Each second-round PCR product was size-selected and purified using the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
The sequencing libraries were pooled, quantified by qPCR with the Kapa Library Quantification Kit for Illumina (Kapa Biosystems, no. KK4824), combined in equimolar ratios and analyzed by paired-end sequencing using Illumina HiSeq 2500.
To identify barcodes in the yeast CloneSelect plasmids introduced into cells, yeast cells were first centrifuged at 20,000g for 3 min, and the supernatant was discarded. The cell pellet was resuspended in 20 µl of Zymolyase Buffer containing 2.5 mg ml−1 Zymolyase (Zymo Research, no. E1005) and 500 µl of Solution I Buffer (supplied with Zymolyase, no. E1005) containing 0.1 M EDTA and 1 M sorbitol. The sample was incubated at 37 °C for 1 h, centrifuged at 20,000g for 1 min, and the supernatant was discarded. The cell lysate was then treated with 250 µl of Solution II Buffer (supplied with Zymolyase, no. E1005) containing 20 mM EDTA, 50 mM Tris-HCl and 1% SDS and then incubated at 65 °C for 30 min. After this, 100 µl of 5 M potassium acetate was added, and the sample was incubated on ice for 30 min, followed by centrifugation at 20,000g for 3 min. The supernatant was transferred to a 1.5 ml tube, and plasmid DNA was precipitated by adding 400 µl of isopropanol, followed by a cleanup with 400 µl of 70% ethanol. The DNA pellet was resuspended in 50 µl of ddH2O containing 10 µg ml−1 RNase and incubated at 65 °C for 10 min. The sequencing library for each sample was prepared using the same method described above for the plasmid DNA pools, with custom indices for the second-round PCR detailed in Supplementary Table 3.
Analysis of reporter activation efficiency
To evaluate the efficiency of gRNA-dependent, barcode-specific mCherry reporter activation, we treated three independent barcoded cell samples with their corresponding gRNAs in a 3 × 3 assay. Each sample was spread on SC–His–Leu–Ura+Ade agar plates, scraped, inoculated into a 1.5 ml tube containing 500 µl of SC–His–Leu–Ura+Ade medium and cultured for 2–4 days at 30 °C.
A 20 µl aliquot of each pre-cultured sample was mixed with 180 µl of SC–His–Leu–Ura+Ade medium and transferred to a flat-bottom transparent 96-well plate (Greiner Bio-One, no. 655090). mCherry fluorescence intensities, normalized by OD595 values, were measured using the Infinite 200 PRO plate reader (TECAN) with TECAN i-control software (v.1.10.4.0). For microscopic observations, 2.5 µl of each cell sample was placed on a glass slide, covered with a coverslip and observed under a BZ-X710 microscope (Keyence) with ×20 and ×40 objective lenses.
To directly measure the GTG→ATG conversion rate in each sample, cells were collected from selective plates and lysed with DNAZol (COSMO BIO, no. DN127) according to the manufacturer’s protocol. Sequencing libraries were prepared using a two-step PCR method. The first-round PCR was conducted in a 32 µl reaction containing 1.6 µl of cell lysate, 1.6 µl each of 10 µM forward primer KI#168 and 10 µM reverse primer KI#169, 0.64 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530S), 6.4 µl of Phusion HF Buffer (NEB, no. B0518S) and 0.64 µl of 10 mM dNTPs (Takara, no. 4030). Thermal cycling conditions were as follows: 98 °C for 30 s; 30 cycles of 98 °C for 10 s, 61 °C for 10 s and 72 °C for 1 min; with a final extension at 72 °C for 5 min. The remaining library preparation and sequencing followed the same protocols described for barcode sequencing, with custom indices for the second-round PCR detailed in Supplementary Table 3.
Isolation and analysis of barcoded colonies
After barcode-specific reporter activation in a complex population, cells from test and control conditions were spread on SC–His–Leu–Ura+Ade agar plates and imaged under a blue light illuminator (FAS-IV, Nippon Genetics) to isolate mCherry+ or mCherry− colonies. Colonies were then isolated into 96-well cell culture plate wells containing 98 µl of SC–His–Leu–Ura+Ade medium and cultured overnight at 30 °C.
For analysis, samples were measured for mCherry fluorescence intensities normalized by OD595 values using an Infinite 200 PRO plate reader (TECAN) with TECAN i-control software (v.1.10.4.0). The same isolated colonies were also subjected to Sanger sequencing to identify their barcode sequences and assess base editing outcomes. Barcode DNA fragments were obtained using the same protocols for cell lysis, first-round PCR and PCR cleanup as in the reporter activation efficiency analysis. Each PCR product was analyzed by Sanger sequencing with sequencing primer SI#658, and sequencing traces were processed using PySanger (https://github.com/ponnhide/PySanger).
Canavanine assay
Genome editing efficiencies of different Cas9-based genome editing enzymes (Cas9, dCas9, dCas9-PmCDA1, dCas9-PmCDA1-UGI, nCas9, nCas9-PmCDA1 and nCas9-PmCDA1-UGI) were estimated using a canavanine assay. In this assay, Cas9-based enzymes under a galactose-inducible GAL1/10 promoter were introduced to cells along with a gRNA targeting the arginine transporter gene CAN1, allowing assessment of knockout efficiency through cell survival in the presence of the toxic arginine analogue canavanine.
To induce genome editing, cells containing both enzyme and gRNA plasmids were first cultured in SC–Leu–Ura medium with 2% glucose at 30 °C until saturation. Cells were then resuspended in SC–Leu–Ura medium with 2% raffinose at a 16-fold dilution and cultured at 30 °C until saturation. Finally, cells were resuspended in SC–Leu–Ura medium containing 2% raffinose and 0.02% galactose at a 32-fold dilution and cultured at 30 °C for 2 days.
Each sample was spread on SC–Leu–Ura–Arg+Ade plates and SC–Leu–Ura–Arg+Ade plates containing 60 mg ml−1 canavanine. Plates were incubated at 30 °C for 2–4 days to estimate colony-forming units and for spot assays. After examining colony-forming units, colonies were scraped from the SC–Leu–Ura–Arg+Ade control plates for genomic DNA extraction to assess mutation spectra by high-throughput sequencing.
For DNA extraction, 20 µl of cells at OD595 of 1.0 were lysed in 100 µl of DNAzol (COSMO BIO, no. DN127) and incubated at room temperature for 15 min. The lysate was mixed with 30 µl of 1 M NaCl and 50 µl of 100% ethanol, then centrifuged at 15,000g for 10 min. The supernatant was discarded, and the pellet was washed with 550 µl of 70% ethanol. After air-drying, the sample was resuspended in 50 µl of ddH2O.
Amplicon sequencing libraries were prepared for each sample using a two-step PCR method in triplicate. The first-round PCR was conducted in a 40 µl volume containing 2 µl of template DNA, 2 µl each of 10 µM forward primer no. KN85F3 and 10 µM reverse primer no. KN85R2, 0.8 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530S), 8 µl of Phusion HF Buffer (NEB, no. B0518S) and 0.8 µl of 10 mM dNTPs (Takara, no. 4030). The thermal cycling conditions were as follows: 98 °C for 30 s; 30 cycles of 98 °C for 10 s, 60 °C for 10 s and 72 °C for 1 min; with a final extension at 72 °C for 5 min. Control samples were prepared using primer pair HO2F2–HO2R2. The remaining library preparation and sequencing followed the same protocols described for barcode sequencing, with custom indices for the second-round PCR detailed in Supplementary Table 3.
Mutational spectra analysis
Amplicon sequencing reads obtained to assess mutational patterns at the CAN1 target site, induced by each Cas9-based genome editing enzyme, were processed using a previously established pipeline38. The codes specific to this analysis are available at https://github.com/yachielab/CloneSelect_v1/tree/main/Mutational_Spectra_Analysis.
Experiments using E. coli
Preparation of cells for various Bacterial CloneSelect systems
Cell samples containing single barcode plasmids were prepared for different Bacterial CloneSelect systems (Supplementary Table 2). For the EGFP reporter-based system, the plasmid was introduced into BL21(DE3) E. coli cells (NEB, no. C2527I). For the blasticidin and Zeocin resistance marker-based systems, plasmids were introduced into T7 Express chemically competent E. coli cells (NEB, no. C2566I), following the manufacturer’s high-efficiency transformation protocols. Transformants were selected on LB agar plates containing 100 µg ml−1 ampicillin (Wako, no. 014-23302) and/or 50 µg ml−1 kanamycin (Wako, no. 111-00344).
Barcode plasmid pool preparation
To generate the bacterial CloneSelect barcode library for the Zeocin resistance marker, a semi-random oligonucleotide pool KI#405 encoding 5′-ATGCCGVNNVNNVNNVNNVNNTAA-3′ was chemically synthesized (Supplementary Table 2). This sequence includes a start codon (ATG), the antisense strand of the 5′-CGG-3′ PAM, a quintuple repeat of VNN (V = non-T) and a stop codon (TAA). The VNN repeat restricts the appearance of in-frame stop codons upstream of the reporter.
The oligonucleotide pool was amplified by PCR in a 20 µl reaction containing 1 µl of 1 µM template, 1 µl each of 10 µM forward primer SI#368 and 10 µM reverse primer SI#369, 0.4 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530L), 4 µl of 5× Phusion HF Buffer (NEB, no. B0518S) and 0.4 µl of 10 mM dNTPs. The thermal cycling conditions were as follows: 98 °C for 30 s; 20 cycles of 98 °C for 10 s, 68 °C for 20 s and 72 °C for 20 s; followed by a final extension at 72 °C for 5 min. The PCR product was analyzed on a 2% agarose gel, size-selected and purified using the FastGene PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
The purified barcode fragment was assembled into the cloning backbone plasmid pKI243 by Golden Gate Assembly using BsmBI. The assembly reaction was performed in a 12.5 µl volume containing 2.91 fmol barcode fragments, 14.9 fmol backbone plasmid, 0.25 µl of BsmBI (NEB, no. R0580L), 0.5 µl of T4 DNA Ligase (Nippon Gene, no. 317-00406), 1.25 µl of 10× T4 DNA Ligation Reaction Buffer (NEB, no. B0202S) and 0.62 µl of 2 mg ml−1 BSA (NEB, no. B9001S). Thermal cycling conditions were 15 cycles of 37 °C for 5 min and 20 °C for 5 min, followed by 55 °C for 30 min for complete backbone digestion.
For transformation, 3 µl of the assembly product was used to transform 65 µl of T7 Express chemically competent cells (NEB, no. C2566I) following the high-efficiency transformation protocol. After a 1 h outgrowth in 500 µl of SOC medium (NEB, no. B9020S) at 37 °C, the cell sample was plated in 250 µl portions on three LB agar plates containing 100 µg ml−1 ampicillin (Wako, no. 014-23302). Diluted samples were also plated on selective plates to estimate clone complexity. Assembly quality and efficiency were checked by isolating 12 random clones and validating the barcode inserts by Sanger sequencing, with 11 out of 12 clones showing the expected barcode insert.
To construct the Pool-100 plasmid pool, 100 colonies were isolated, each resuspended in 80 µl of LB medium with 100 µg ml−1 ampicillin, combined in 5 µl aliquots and cultured overnight at 37 °C. Plasmid DNA was extracted using the FastGene Plasmid Mini Kit (Nippon Genetics, no. FG-90502). The Pool-1550 was constructed by scraping colonies from a plate with ~1,000 colony-forming units into 1.5 ml LB medium with 100 µg ml−1 ampicillin. The barcode plasmid libraries were used to transform T7 Express chemically competent cells (NEB, no. C2566I) to establish barcoded E. coli cell populations.
Barcode sequencing library preparation
For the Pool-100 and Pool-1550 barcode plasmid libraries, barcode sequencing libraries were prepared in triplicate using a two-step PCR method. The first-round PCR was performed in five separate 40 µl reactions, each containing 2.0 ng of plasmid template DNA, 1 µl each of 10 µM forward primer KI#403 and 10 µM reverse primer KI#404, 0.4 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530L), 8 µl of Phusion HF Buffer (NEB, no. B0518S) and 0.8 µl of 10 mM dNTPs (Takara, no. 4030). Thermal cycling conditions were as follows: 98 °C for 30 s; 20 cycles of 98 °C for 10 s, 54 °C for 20 s and 72 °C for 25 s; followed by a final extension at 72 °C for 5 min. For each replicate, the five PCR products were pooled, size-selected on a 2% agarose gel, purified and eluted in 30 µl of ddH2O using the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302).
In the second-round PCR, Illumina sequencing adaptors and custom indices were added to each first-round PCR product. Each 40 µl reaction contained 2 µl of the first-round PCR product, 1 µl each of 10 µM P5 and P7 custom index primers, 0.4 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530L), 8 µl of 5× Phusion HF Buffer (NEB, no. B0518S) and 0.8 µl of 10 mM dNTPs (Takara, no. 4030). The thermal cycling conditions were as follows: 98 °C for 30 s; 15 cycles of 98 °C for 10 s, 60 °C for 10 s and 72 °C for 60 s; followed by a final extension at 72 °C for 5 min. Custom indices for the second-round PCR products are listed in Supplementary Table 3.
The second-round PCR products were size-selected and purified using the PCR/Gel Extraction Kit (Nippon Genetics, no. FG-91302). The sequencing libraries were pooled, quantified by qPCR using the Kapa Library Quantification Kit for Illumina (Kapa Biosystems, no. KK4824), combined in equimolar ratios and analyzed by paired-end sequencing using Illumina MiSeq.
Introduction of genome editing reagents
To introduce a plasmid containing ABE-7.10 and a gRNA to barcoded cells, we used the Mix&Go! E. coli Transformation Kit (Zymo Research, no. T3001) following the manufacturer’s protocol. Transformants were selected by plating the transformation reaction on LB agar plates containing 100 µg ml−1 ampicillin (Wako, no. 014-23302) and 50 µg ml−1 kanamycin (Wako, no. 111-00344) and incubating overnight at 37 °C.
For experiments involving induction with Ara (Sigma-Aldrich, no. A3256-10MG) and IPTG (ThermoFisher Scientific, no. 15529019), cells were cultured overnight at 37 °C in medium containing 100 mM Ara and 0.1 mM IPTG before analysis. For barcoded cell isolation using the Zeocin resistance marker-based system, a low-salt LB medium adjusted to pH 7.5 with 1 M NaOH (Nakalai, no. 37421-05) was used to optimize Zeocin activity.
For genome editing and selection of reporter-activated cells without inducers, we used 100 µg ml−1 Zeocin (Invitrogen, no. R25001) or 100 µg ml−1 blasticidin S (Wako, no. 029-18701), as the leaky expression from inducible promoters in the absence of inducers was sufficient for gene editing while maintaining high cell viability. Details of the genome editing plasmids used in this study are provided in Supplementary Table 2.
Analysis of reporter activation efficiency
To evaluate the efficiency of gRNA-dependent, barcode-specific EGFP reporter activation, 200 µl of cell samples were transferred into a flat-bottom transparent 96-well plate (Greiner Bio-One, no. 655090) and analyzed using the Infinite 200 PRO plate reader (TECAN) with TECAN i-control software (v.1.10.4.0) to measure EGFP fluorescence intensities normalized to OD595 values.
For microscopic observation, 2.5 µl of each cell sample was placed on a glass slide (MATSUNAMI, no. S2441), gently covered with a glass coverslip and examined under a BZ-X710 microscope (Keyence) using ×20 and ×40 objective lenses.
Isolation and analysis of barcoded colonies
After barcode-specific activation of the Zeocin resistance marker in the barcoded cell population, barcodes from colonies under test and control conditions were analyzed by Sanger sequencing. For each colony, the barcode region was amplified by PCR in a 20 µl reaction containing 1 µl of cell suspension, 0.5 µl each of 10 µM forward primer KI#403 and 10 µM reverse primer KI#404, 0.2 µl of Phusion High-Fidelity DNA Polymerase (NEB, no. M0530L), 4 µl of Phusion HF Buffer (NEB, no. B0518S) and 0.4 µl of 10 mM dNTPs (Takara, no. 4030). The thermal cycling conditions were as follows: 98 °C for 30 s; 30 cycles of 98 °C for 10 s, 54 °C for 20 s and 72 °C for 30 s; followed by a final extension at 72 °C for 5 min.
The PCR products were analyzed on a 2% agarose gel and transferred to wells of a 96-well PCR plate for cleanup using 20 µl of AMPure XP beads (Beckman Coulter, no. A63881) according to the manufacturer’s protocol. Sanger sequencing was conducted using primer KI#403, and sequencing traces were analyzed with PySanger (https://github.com/ponnhide/PySanger).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
High-throughput sequencing data generated in this study are available at the NCBI BioProject (PRJNA901977). The list of plasmids used in this study can be found in Supplementary Table 1. The new plasmids necessary to reproduce the work have been deposited at Addgene (https://www.addgene.org/browse/article/28233756). Source data are provided with this paper.
Code availability
All the custom codes used in this study are available at https://github.com/yachielab/CloneSelect_v1 (ref. 87).
Change history
06 October 2025
In the version of Supplementary Table 1 initially published, in the “mESC_BC” tab, the labels for columns B and C were switched and have now been amended so that column B is “uptag sequence” and column C is “dntag sequence”. The corrected Supplementary Table 1 is now available alongside the online version of the article.
References
Bhang, H. E. et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nat. Med. 21, 440–448 (2015).
Hata, A. N. et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat. Med. 22, 262–269 (2016).
Ben-David, U. et al. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560, 325–330 (2018).
Nguyen, L. V. et al. Barcoding reveals complex clonal dynamics of de novo transformed human mammary cells. Nature 528, 267–271 (2015).
Venkataram, S. et al. Development of a comprehensive genotype-to-fitness map of adaptation-driving mutations in yeast. Cell 166, 1585–1596.e22 (2016).
Barrick, J. E. et al. Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature 461, 1243–1247 (2009).
Levy, S. F. et al. Quantitative evolutionary dynamics using high-resolution lineage tracking. Nature 519, 181–186 (2015).
Blundell, J. R. et al. The dynamics of adaptive genetic diversity during the early stages of clonal evolution. Nat. Ecol. Evol. 3, 293–301 (2019).
Nguyen Ba, A. N. et al. High-resolution lineage tracking reveals travelling wave of adaptation in laboratory yeast. Nature 575, 494–499 (2019).
Pei, W. et al. Polylox barcoding reveals haematopoietic stem cell fates realized in vivo. Nature 548, 456–460 (2017).
Lu, R., Neff, N. F., Quake, S. R. & Weissman, I. L. Tracking single hematopoietic stem cells in vivo using high-throughput sequencing in conjunction with viral genetic barcoding. Nat. Biotechnol. 29, 928–933 (2011).
Naik, S. H. et al. Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature 496, 229–232 (2013).
Gerrits, A. et al. Cellular barcoding tool for clonal analysis in the hematopoietic system. Blood 115, 2610–2618 (2010).
Biddy, B. A. et al. Single-cell mapping of lineage and identity in direct reprogramming. Nature 564, 219–224 (2018).
Hollmann, J. et al. Genetic barcoding reveals clonal dominance in iPSC-derived mesenchymal stromal cells. Stem Cell Res. Ther. 11, 105 (2020).
Tonge, P. D. et al. Divergent reprogramming routes lead to alternative stem-cell states. Nature 516, 192–197 (2014).
Rodriguez-Fraticelli, A. E. et al. Single-cell lineage tracing unveils a role for TCF15 in haematopoiesis. Nature 583, 585–589 (2020).
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020).
Wagner, D. E. et al. Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 360, 981–987 (2018).
Rodriguez-Fraticelli, A. E. et al. Clonal analysis of lineage fate in native haematopoiesis. Nature 553, 212–216 (2018).
He, Z. et al. Lineage recording in human cerebral organoids. Nat. Methods 19, 90–99 (2022).
Chan, M. M. et al. Molecular recording of mammalian embryogenesis. Nature 570, 77–82 (2019).
Raj, B. et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat. Biotechnol. 36, 442–450 (2018).
Ishiguro, S., Mori, H. & Yachie, N. DNA event recorders send past information of cells to the time of observation. Curr. Opin. Chem. Biol. 52, 54–62 (2019).
Ota, S. et al. Ghost cytometry. Science 360, 1246–1251 (2018).
Schraivogel, D. et al. High-speed fluorescence image-enabled cell sorting. Science 375, 315–320 (2022).
Nitta, N. et al. Intelligent image-activated cell sorting. Cell 175, 266–276.e13 (2018).
Hasle, N. et al. High-throughput, microscope-based sorting to dissect cellular heterogeneity. Mol. Syst. Biol. 16, e9442 (2020).
Umkehrer, C. et al. Isolating live cell clones from barcoded populations using CRISPRa-inducible reporters. Nat. Biotechnol. 39, 174–178 (2021).
Al’Khafaji, A. M., Deatherage, D. & Brock, A. Control of lineage-specific gene expression by functionalized gRNA barcodes. ACS Synth. Biol. 7, 2468–2474 (2018).
Gutierrez, C. et al. Multifunctional barcoding with ClonMapper enables high-resolution study of clonal dynamics during tumor evolution and treatment. Nat. Cancer 2, 758–772 (2021).
Feldman, D. et al. CloneSifter: enrichment of rare clones from heterogeneous cell populations. BMC Biol. 18, 177 (2020).
Pickar-Oliver, A. & Gersbach, C. A. The next generation of CRISPR–Cas technologies and applications. Nat. Rev. Mol. Cell Biol. 20, 490–507 (2019).
Kuscu, C. et al. CRISPR–STOP: gene silencing through base-editing-induced nonsense mutations. Nat. Methods 14, 710–712 (2017).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Emert, B. L. et al. Variability within rare cell states enables multiple paths toward drug resistance. Nat. Biotechnol. 39, 865–876 (2021).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Sakata, R. C. et al. Base editors for simultaneous introduction of C-to-T and A-to-G mutations. Nat. Biotechnol. 38, 865–869 (2020).
Hecht, A. et al. Measurements of translation initiation from all 64 codons in E. coli. Nucleic Acids Res. 45, 3615–3626 (2017).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Kim, N. et al. Prediction of the sequence-specific cleavage activity of Cas9 variants. Nat. Biotechnol. 38, 1328–1336 (2020).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Hegde, M., Strand, C., Hanna, R. E. & Doench, J. G. Uncoupling of sgRNAs from their associated barcodes during PCR amplification of combinatorial CRISPR screens. PLoS ONE 13, e0197547 (2018).
Cahan, P. & Daley, G. Q. Origins and implications of pluripotent stem cell variability and heterogeneity. Nat. Rev. Mol. Cell Biol. 14, 357–368 (2013).
Nishizawa, M. et al. Epigenetic variation between human induced pluripotent stem cell lines is an indicator of differentiation capacity. Cell Stem Cell 19, 341–354 (2016).
Nazareth, E. J. et al. High-throughput fingerprinting of human pluripotent stem cell fate responses and lineage bias. Nat. Methods 10, 1225–1231 (2013).
Theunissen, T. W. et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell 15, 471–487 (2014).
Ware, C. B. et al. Derivation of naive human embryonic stem cells. Proc. Natl Acad. Sci. USA 111, 4484–4489 (2014).
Chan, Y. S. et al. Induction of a human pluripotent state with distinct regulatory circuitry that resembles preimplantation epiblast. Cell Stem Cell 13, 663–675 (2013).
Gafni, O. et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 504, 282–286 (2013).
Vaisvila, R. et al. Enzymatic methyl sequencing detects DNA methylation at single-base resolution from picograms of DNA. Genome Res. 31, 1280–1289 (2021).
Krietenstein, N. et al. Ultrastructural details of mammalian chromosome architecture. Mol. Cell 78, 554–565.e7 (2020).
Li, S. et al. Capturing totipotency in human cells through spliceosomal repression. Cell 187, 3284–3302.e23 (2024).
Burian, J. et al. High-throughput retrieval of target sequences from complex clone libraries using CRISPRi. Nat. Biotechnol. 41, 626–630 (2023).
Banno, S., Nishida, K., Arazoe, T., Mitsunobu, H. & Kondo, A. Deaminase-mediated multiplex genome editing in Escherichia coli. Nat. Microbiol. 3, 423–429 (2018).
Shelake, R. M., Pramanik, D. & Kim, J. Y. In vivo rapid investigation of CRISPR-based base editing components in Escherichia coli (IRI-CCE): a platform for evaluating base editing tools and their components. Int. J. Mol. Sci. 23, 1145 (2022).
Kim, K. et al. Epigenetic memory in induced pluripotent stem cells. Nature 467, 285–290 (2010).
Bar-Nur, O., Russ, H. A., Efrat, S. & Benvenisty, N. Epigenetic memory and preferential lineage-specific differentiation in induced pluripotent stem cells derived from human pancreatic islet beta cells. Cell Stem Cell 9, 17–23 (2011).
Kim, K. et al. Donor cell type can influence the epigenome and differentiation potential of human induced pluripotent stem cells. Nat. Biotechnol. 29, 1117–1119 (2011).
Osafune, K. et al. Marked differences in differentiation propensity among human embryonic stem cell lines. Nat. Biotechnol. 26, 313–315 (2008).
Boulting, G. L. et al. A functionally characterized test set of human induced pluripotent stem cells. Nat. Biotechnol. 29, 279–286 (2011).
Casini, A., Storch, M., Baldwin, G. S. & Ellis, T. Bricks and blueprints: methods and standards for DNA assembly. Nat. Rev. Mol. Cell Biol. 16, 568–576 (2015).
Hutchison, C. A. III et al. Design and synthesis of a minimal bacterial genome. Science 351, aad6253 (2016).
Shao, Y. et al. Creating a functional single-chromosome yeast. Nature 560, 331–335 (2018).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Gibson, D. G. et al. Chemical synthesis of the mouse mitochondrial genome. Nat. Methods 7, 901–903 (2010).
Kijima, Y., Evans-Yamamoto, D., Toyoshima, H. & Yachie, N. A universal sequencing read interpreter. Sci. Adv. 9, eadd2793 (2023).
Mori, H. & Yachie, N. A framework to efficiently describe and share reproducible DNA materials and construction protocols. Nat. Commun. 13, 2894 (2022).
Kutner, R. H., Zhang, X. Y. & Reiser, J. Production, concentration and titration of pseudotyped HIV-1-based lentiviral vectors. Nat. Protoc. 4, 495–505 (2009).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Engler, C., Kandzia, R. & Marillonnet, S. A one pot, one step, precision cloning method with high throughput capability. PLoS ONE 3, e3647 (2008).
Zhao, L., Liu, Z., Levy, S. F. & Wu, S. Bartender: a fast and accurate clustering algorithm to count barcode reads. Bioinformatics 34, 739–747 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Okubo, T. et al. Hypoblast from human pluripotent stem cells regulates epiblast development. Nature 626, 357–366 (2024).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Fang, Z., Liu, X. & Peltz, G. GSEApy: a comprehensive package for performing gene set enrichment analysis in Python. Bioinformatics 39, btac757 (2023).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Akalin, A. et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 13, R87 (2012).
Ishiguro, S. et al. Source codes for the CloneSelect Project 2025. Github https://github.com/yachielab/CloneSelect_v1 (2025).
Acknowledgements
We thank the members of the Yachie lab currently at the University of British Columbia and Osaka University, as well as those formerly at the University of Tokyo. We especially appreciate D. Sheykhkarimli, S. Chopra, S. Okawa, S. Romero, Y. Liu, Y. Dorri, M. Kagieva, H. Yao, C. Ye, P. Lam, S. Khalilitousi, A. Uchida and J. Plant for reviewing the paper and providing constructive feedback. We also thank K. Shiina, S. Fukuda and T. Stach for their technical support in high-throughput sequencing, along with A. Johnson and J. Wong for their assistance with flow cytometry cell sorting. Our gratitude extends to Mitinori Saitou Laboratory and the Single-cell Genome Information Analysis Core (SignAC) at WPI-ASHBi, Kyoto University, for their support. This study was funded by the World Premier International Research Center Initiative (WPI), MEXT, Japan, the Japan Society for the Promotion of Science (JSPS) KAKENHI grant number (24H00867), the Japan Science and Technology Agency (JST)’s PRESTO Single Cell Analysis Program (JPMJPR14FE), the CREST Cell Control (yuCell) Program (JPMJCR23B7), the JSPS (24H00867), the Takeda Science Foundation, the SECOM Science and Research Foundation, the Canada Foundation for Innovation (CFI) (39968), the Canadian Institute for Advanced Research MacMillan Multiscale Human program (FL-001525) (all to N.Y.), the New Energy and Industrial Technology Development Organization (NEDO) (17101603-0) (to N.Y. and K.N.), the Allen Distinguished Investigator Award (12964), the Canadian Institutes of Health Research (CIHR) (175622, 479568 and 202107DT1-472331-DMT-CAAA-36569) (to N.Y. and N.S.), the Japan Agency for Medical Research and Development (AMED) (JP20gm6110007, JP21gm1310011, JP23bm1323001, 21ek0109448h0002 and 24bk0104169s0201) (to N.Y., Y.T. and K.N.), the Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grant (RGPIN-2020-04198), the Michael Smith Health Research BC Scholar Award (SCH-2021-1673), the Stem Cell Network (ECR-C4R1-3), the SickKids-CIHR New Investigator Grant (KF-179017), the Breakthrough T1D Team Grant (5-SRA-2021-1150-S-B) (all to N.S.) and the GteX Program Japan (JPMJGX23B4) (to K.N.). We were also supported by diverse fellowships, including the Canada Research Chair program through CIHR (N.Y. and N.S.), the Ryoichi Sasakawa Young Leaders Fellowship (S.I.), the Banting Postdoctoral Fellowship (S.I. and A.-S.A.), the Taikichiro Mori Memorial Research Fund (S.I., H.M. and N.M.), the JSPS DC fellowship (S.I., H.M., Y.K. and N.M.), the JSPS Overseas Research Fellowships (S.I., Y.K. and R.Y.), the Japan Student Services Organization (R.C.S.), the Funai Overseas Scholarship (R.C.S.), the Takenaka Scholarship Foundation (R.T.), the Ministry of Science and Education (MEXT) Research Scholarship (A.A.), the NSERC Undergraduate Student Research Award (S. King), the UBC Four Year Doctoral Fellowship (J.H.O.), the Michael Smith Health Research BC Scholar Award (R.I.K.G.), the Michael Smith Health Research BC Trainee Award (A.-S.A.), and the Killam Doctoral Scholarship, University of British Columbia (O.B.). Part of the high-throughput sequencing data analysis was conducted using the SHIROKANE Supercomputer at the University of Tokyo Human Genome Center.
Author information
Authors and Affiliations
Contributions
S.I. and N.Y. conceived the study. S.I., R.C.S. and M.T. performed the major mammalian cell experiments. K.I. and M.T. performed the yeast and bacterial experiments. R.T. and R.Y. supported the lentiviral library screens. O.B. and N.S. performed a part of the experiments using mouse ES cells and human PS cells. M.I. and Y.T. performed the human PS cell naïve induction screen and the subsequent trophoblast differentiation assay. T.Y., S.T., T.T. and H.A. supported data acquisition using high-throughput sequencing. S.I., Y.K., H.M., H.T., R.C.S. and H.A. performed the high-throughput sequencing analyses. R.T., A.A. and N.M. performed preliminary assays and contributed to the design of the system. J.H.O., A.-S.A. and R.I.K.G. contributed to the optimization of the system. S. King supported plasmid construction. K.N., A.K. and S. Kuhara also provided substantial ideas in designing the proposed system. M.S. supported the flow cytometry cell sorting. H.A., T.Y., S.T. and T.T. supported the high-throughput sequencing. H.A. supported the flow cytometry analysis and the microscope imaging. S.I., R.C.S., K.I. and N.Y. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
K.I. is an employee of Spiber Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Supplementary data for the isolation of target barcoded cells from a population using CloneSelect C → T.
a, Schematic diagram for the barcode library construction. The ∆EGFP fragment (GTG mutated start codon) was amplified by PCR using pooled forward primers encoding the PAM followed by semi-random barcode sequences encoding the GTG mutated start codon and a common reverse primer. The PCR product was then enzymatically digested and ligated to the lentivirus plasmid backbone. b, The PCR product. c, Estimated complexities of the generated plasmid pools by colony forming units (n = 3). d, Library QC by single colony isolation followed by plasmid purification and restriction digestion using BsrGI, ClaI and PvuI. e, Sanger sequencing of the barcode region of the colony isolates. f and g, Barcode distribution in the lentiviral plasmid DNA pool and that in the cell population transduced using the same plasmid DNA pool. h, Barcode distributions of the EGFP-positive cell samples obtained by cell sorting after barcode-specific activation. The results of the 16 independent samples were combined after read count normalization applied to each sample. i, Frequency of the GTG → ATG mutation observed for each barcode after sorting the EGFP-positive cells. Each row represents the GTG → ATG mutation frequency profile in each target isolation assay.
Extended Data Fig. 2 CloneSelect C → T Pool-10000 libraries.
a–c, Barcode abundances measured by high-throughput sequencing for each of the lentivirus Pool-10000 libraries (n = 2). d–f, Levenshtein distance distributions between the target barcodes (white dots) used in the study and non-target barcodes.
Extended Data Fig. 3 Benchmarking of CloneSelect and CRISPRa-based systems using Pool-10000 libraries.
a, Barcoded cell frequencies in pre- and post-sort populations (Replicate 2 of n = 2). b, Separation scores of different isolation attempts from Pool-10000 prepared for different retrospective clone isolation systems. The target barcode abundances were not adjusted by the dilution factor introduced by the pooling of different experimental samples. The two-tailed Mann-Whitney U test was used for statistical analysis. *P < 0.05; **P < 0.01; ***P < 0.001.
Extended Data Fig. 4 Supplementary data for scCloneSelect.
a, EGFP expressions of the ATG positive controls for the original CloneSelect C → T and scCloneSelect in HEK293T cells with the same genome-editing conditions tested for the respective reporters (n = 3). Scale bar, 50 µm. b, Median EGFP intensities of base editing-activated EGFP positive cells (n = 3). c and d, Barcode-specific gRNA-dependent reporter activation of six barcoded cell lines by scCloneSelect. The two-tailed Welch’s t-test was used for statistical analysis. Scale bar, 50 µm. e, RT-PCR of the scCloneSelect dntags in HEK293T. f, The fraction of mESC single-cell transcriptome profiles (Drop-seq) that contained dntags and the fraction of dntags reported in the uptag-dntag reference database. g, Schematic representation of a scCloneSelect reporter activation assay. Target-AID was stably introduced to the cell population prior to barcoding and gRNA-dependent reporter activation. h and i, Barcode-specific gRNA-dependent reporter activation of barcoded mESCs and CA1 hPSCs by scCloneSelect (n = 2). Target-AID was stably integrated prior to the barcoding. Target gRNAs were delivered by transfection. The two-tailed Welch’s t-test was used for statistical analysis. Scale bar, 100 µm. j, Schematic representation of another scCloneSelect reporter activation assay. The target gRNA and Target-AID were electroporated together to the barcoded cell population. k, Barcode-specific gRNA-dependent reporter activation of barcoded H1 hPSCs by scCloneSelect (n = 2). Targeting gRNA and Target-AID were electroporated together. Scale bar, 100 µm. *P < 0.05; **P < 0.01; ***P < 0.001.
Extended Data Fig. 5 Supplementary data for the isolation of barcoded cells identified in scRNA-seq data.
a, QC of the scRNA-seq datasets obtained for the barcoded mESC population cultured with LIF and 2i and that cultured without LIF or 2i. b, Single-cell expression patterns of key genes. c, Distribution of cells in a two-dimensional UMAP space for all the clones targeted for isolation. d, Abundances of barcoded cell clones in the mESC population. The data was generated based on dntags identified in the scRNA-seq dataset with no reamplification of the dntag reads. e, Barcode-specific gRNA-dependent activation of the reporter for each target clone in the initial mESC population. f, Culturing of isolated single cells. Scale bar, 100 µm. g, Sanger sequencing of the barcodes for the isolated target clones after expansion.
Extended Data Fig. 6 Supplementary data for the isolation of elite hPSC clones having high naïve induction potential.
a, Correlation in barcoded clone abundance between two different samples. b, Barcode enrichment analysis after the cell sorting of EGFP-positive cells from the initial primed hPSC population. Each row represents the barcode abundance profile in each target isolation assay. c, EGFP intensities of cells before cell sorting in each isolation attempt. The dashed lines indicate a gating threshold of cell sorting. d, EGFP intensities of each primed hPSC clone after sorting. e, Sanger sequencing of the barcode region of each isolated clone. f, Isolation of each elite hPSC clone candidate having high naïve potential from the parental population. CloneSelect C → T reporter was activated by electroporating Target-AID and gRNA plasmids. The pre-sorting population after the introduction of base editing was also kept, analyzed, and subjected to the naïve induction assay. g, Correlation in barcode abundance between pre-sorting and post-sorting primed hPSC populations. Pink dots represent the target clones. h, Flow cytometric profiles of pre-sorting and post-sorting primed hPSC populations after naïve induction.
Extended Data Fig. 7 Transcriptome and DNA methylation analysis of isolated hPSC clones.
a, Clustering of cell samples in transcriptome profile (n = 2). b, Downregulated genes in each isolated clone, compared to its parental sample. c, Global CpG methylation profile similarities between three elite hPSC clones. d, High-order chromatin structure of the CSAG1 region in Chromosome X previously measured by Micro-C in Human ESC H155. e, Upregulated genes in each isolated clone, compared to its parental sample. f, Enrichment and depletion of gene ontology terms in downregulated genes in naïve cells induced from the elite hPSC clones, compared to their corresponding pre-sorting samples with FDR < 0.1.
Extended Data Fig. 8 Supplementary data for the development of yeast CloneSelect.
a, Different mCherry reporter variants tested to establish CloneSelect C → T. Different variants were tested with the first codon as GTG or ATG. Scale bar, 100 µm. b, Canavanine resistance assays for different CRISPR genome editing enzymes with a gRNA targeting CAN1 gene and a control NT gRNA. For each experiment, cell concentration was normalized to 1.0 OD595 nm and serially diluted with 10-fold increments for spotting. c, Estimated colony forming unit (CFU) counts for the same assay in b. d, Genome editing outcomes observed by amplicon sequencing. Frequencies of mutation patterns observed across the target sequence region are shown for the same assay in b. e, Genome editing frequencies at the target CAN1 locus estimated by amplicon sequencing for the different enzymes. f, Activation of the mCherry M1V (GTG) + ∆2-9 mutant reporter by OT and NT gRNAs. Scale bar, 200 µm. g, mCherry expressions of the ATG positive control. Yeast cells having the positive control reporters with three different barcodes (BC-C1, BC-C2, and BC-C3) were each treated with Target-AID and three different targeting gRNAs. Scale bar, 25 µm.
Extended Data Fig. 9 Supplementary data for Yeast CloneSelect.
a, Schematic diagram for the barcode library construction. The barcode fragment pool was prepared by PCR using a common primer pair amplifying a template DNA pool encoding the PAM, WSNS semi-random repeat, and the mutated start codon GTG. The PCR product was digested and ligated to a backbone plasmid. b, The PCR product. c, Library QC by colony isolation and PCR amplification of the barcode insert (Pool-100). d, Barcode abundance distribution (read per million) in the constructed barcode library pool. The sequencing library was prepared and analyzed in triplicate (n = 3). e and f, mCherry intensities for expected negative isolates obtained from Pool-100 and Pool-1580. The intensities were measured by a plate reader in triplicate (n = 3).
Extended Data Fig. 10 Supplementary data for Bacterial CloneSelect.
a, Bacterial CloneSelect with the EGFP reporter. band c, Activation of the EGFP and control reporters using on-target (OT) and non-target (NT) gRNAs (n = 3). ABE and gRNA expression were controlled by an IPTG-inducible promoter, and the EGFP reporter expression was controlled by an arabinose-inducible promoter. The two-tailed Welch’s t-test was used for statistical analysis. d, Base editing outcomes of the positive control reporters analyzed by Sanger sequencing. e, Testing of Zeocin resistances conferred by two promoters expressing a Zeocin resistance gene with and without the upstream stop codon. Each cell sample concentration was first adjusted to 0.1 OD595 nm and serially diluted with 10-fold increments for spotting 5 µL. f, Testing of cell viability under a non-selective condition for a constitutively active T7 promoter and the IPTG-inducible promoter to express the gRNA. OT and NT gRNAs were tested for the gRNA-dependent EGFP reporter and the positive control EGFP reporter. ABE was expressed under the IPTG-inducible promoter without IPTG. g, Barcode-specific gRNA-dependent Zeocin resistance reporter activation tested for the IPTG-inducible promoters with and without IPTG. h, Bacterial CloneSelect using a Blasticidin S resistance gene, bsr. i, Barcode-specific gRNA-dependent Blasticidin S-resistance reporter activation tested for different inducer conditions. Each cell sample concentration was adjusted to 0.1 OD595 nm for spotting 5 µL. *P < 0.05; **P < 0.01; ***P < 0.001.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–15.
Supplementary Table 1 (download XLSX )
Barcode count distributions observed in this study.
Supplementary Table 2 (download XLSX )
Oligonucleotides and plasmids used in this study.
Supplementary Table 3 (download XLSX )
List of high-throughput sequencing datasets.
Supplementary Table 4 (download XLSX )
P values and statistical test methods used in this study.
Supplementary Table 5 (download XLSX )
Values obtained from flow cytometry analyses.
Source data
Source Data Extended Data Figs. 1, 4 and 9 (download PDF )
Unprocessed gel images.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ishiguro, S., Ishida, K., Sakata, R.C. et al. A multi-kingdom genetic barcoding system for precise clone isolation. Nat Biotechnol (2025). https://doi.org/10.1038/s41587-025-02649-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-025-02649-1