Abstract
Small molecules can bind RNAs to regulate their fate and functions, providing promising opportunities for treating human diseases. However, current tools for predicting small molecule–RNA interactions (SRIs) require prior knowledge of RNA tertiary structures. Here we present SMRTnet, a deep learning method that uses multimodal data fusion to integrate two large language models with convolutional and graph attention networks to predict SRIs on the basis of RNA secondary structure. SMRTnet achieves high performance across multiple experimental benchmarks, substantially outperforming existing tools. SMRTnet predictions for ten disease-associated RNA targets identified 40 hits of RNA-targeting small molecules with nanomolar-to-micromolar dissociation constants. Focusing on the MYC internal ribosome entry site, SMRTnet-predicted small molecules showed binding scores correlated closely with observed validation rates. One predicted small molecule downregulated MYC expression, inhibited proliferation and promoted apoptosis in three cancer cell lines. Thus, by eliminating the need for RNA tertiary structures, SMRTnet expands the scope of feasible RNA targets and accelerates the discovery of RNA-targeting therapeutics.
Similar content being viewed by others
Main
RNA has recently emerged as an attractive target for drug discovery, as its complex structures can be selectively modulated by small molecules to influence diverse biological processes, including pre-mRNA splicing1,2, mRNA translation3,4, RNA–protein interaction5,6, noncoding RNA processing7,8 and the replication of RNA viruses9. The ability to target RNAs expands the spectrum of therapeutic targets for small-molecule drugs, that is, genes of disease-associated ‘undruggable’ proteins. A notable example is Evrysdi (risdiplam), the first RNA-targeting drug that promotes the inclusion of exon 7 in SMN2 during splicing and is used for treating spinal muscular atrophy10,11. However, despite this promise, RNA remains far less explored than proteins because of several key challenges—chief among them being the difficulty of determining RNA tertiary structures, which constrains the discovery of RNA-targeting small molecules. Recently, high-throughput experimental methods such as automated ligand identification system12 and small-molecule microarray13 have been developed to circumvent this challenge and facilitate large-scale screening of small molecule–RNA interactions (SRIs). Nevertheless, the enormous chemical space of potential RNA binders poses a formidable challenge to experimental methods14,15.
Beyond methods based on direct experimental measurements, computational methods have also been developed to predict SRIs16,17. For instance, molecular docking tools, such as AutoDock Vina18, RLDOCK19, NLDock20 and rDock21, have been adapted or developed to perform docking between small molecules and the tertiary structures of nucleic acids22. Deep learning approaches have recently been applied to predict SRIs23,24,25. Among them, RNAmigos2 (ref. 24) uses a variational autoencoder and graph neural network to predict likely binders to RNA, while RLaffinity25 introduces a three-dimensional (3D) convolutional neural network (CNN) to predict the binding affinity of small molecules to RNA targets. Despite these advances, these computational methods require known RNA tertiary structures, limiting their practicality because most disease-related RNAs lack defined tertiary structures and only a few have known active sites.
Here we developed SMRTnet (predicting small molecule and RNA target interactions using deep neural networks), a deep learning method that predicts SRIs without requiring prior knowledge of RNA tertiary structures. SMRTnet incorporates two large language models (LLMs), CNNs and graph attention networks (GATs) to capture both sequence and structural features from small molecules and RNAs and uses multimodal data fusion (MDF) to integrate these features for accurate SRI prediction. SMRTnet demonstrates high performance on multiple benchmarks of experimentally validated SRIs, substantially outperforming existing methods. We also used SMRTnet to identify potential binders for ten disease-associated RNA targets by screening a library comprising 7,350 compounds. Microscale thermophoresis (MST)26 assays validated 40 hits with dissociation constants (Kd) in the nanomolar-to-micromolar range. Focusing on the MYC internal ribosome entry site (IRES)27, we found that SMRTnet-predicted binding scores showed a strong correlation with MST validation rates. One predicted compound exhibited bioactivity in three cancer cell lines by reducing MYC expression, inhibiting proliferation and promoting apoptosis. In summary, SMRTnet represents an effective approach for SRI screening and showcases how artificial intelligence (AI) methods can be harnessed for the development of RNA-targeting small-molecule therapeutics.
Results
Overview of SMRTnet
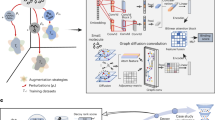
SMRTnet is a deep learning method that takes an RNA sequence with its secondary structure and a small-molecule simplified molecular input line entry system (SMILES) as input to predict a binding score (Fig. 1a). The architecture of SMRTnet consists of an RNA encoder, a small-molecule encoder, an MDF module and a decoder (Fig. 1b and Methods). The RNA encoder integrates an in-house RNA language model (RNASwan-seq) (Extended Data Fig. 1a) and a two-layer CNN with residual neural networks (ResNets) (Extended Data Fig. 1b) to extract both nucleotide and base-pairing information as the representation for the input RNA. The small-molecule encoder incorporates a published chemical language model (MoLFormer)28 and a three-layer GAT (Extended Data Fig. 1c) to capture atomic composition and chemical structure as the representation for the input small molecule. We also developed an MDF module to capture the complex interplay between RNA and small-molecule representations in defining SRIs by progressively integrating pairwise binding information through co-attention29 and self-attention neural networks30 (Extended Data Fig. 1d) and output an interaction representation, which is passed to a fully connected neural network decoder to predict the binding score.
a, Schematic overview of SMRTnet, a deep learning method designed to predict SRIs. The method accepts an RNA sequence along with its secondary structure and a small molecule represented by its SMILES notation and outputs the binding score and potential binding site. b, Architecture of SMRTnet. The training dataset was derived from the PDB. SMRTnet comprises three main components: an RNA encoder, a small-molecule encoder, and an MDF module. The RNA encoder processes RNA sequences using an RNA language model (RNASwan-seq) and RNA structure using CNNs with ResNets. The small-molecule encoder processes small-molecule SMILES using a chemical language model (MoLFormer) and the chemical structure of small molecules using GATs. The MDF module uses attention-based neural networks to progressively integrate pairwise binding information, which is decoded by the fully connected neural network to predict the binding score for the input small molecule and RNA pair. Finally, an ensemble scoring strategy is applied to generate the final binding score.
To train SMRTnet, we constructed the SMRTnet dataset by collecting 1,061 high-quality 3D structures from the Protein Data Bank (PDB) that contain at least one RNA and one small molecule (Extended Data Fig. 2a and Supplementary Table 1). As each interaction site usually involves multiple RNA fragments, we obtained the secondary structure of the fragments and generated a total of 8,672 interactions of RNA fragments and small molecules (Extended Data Fig. 2b and Methods). These interactions are used as positive samples for training and testing. We also randomly paired RNA fragments and small molecules to create noninteraction pairs as negative samples after filtering out those with known interactions. To evaluate the model’s robustness, we sampled negative samples at varying ratios (1:1, 1:2, 1:3, 1:4, 1:5 and 1:10) relative to positive samples, aligning with the range of ratios employed in prior studies31,32,33.
We divided the SMRTnet dataset into training, validation and test sets at an 8:1:1 ratio and applied a ligand-based data-splitting strategy to ensure that small molecules in test sets were absent from training and validation sets34,35,36,37,38,39,40. We also applied fivefold cross-validation (CV) to assess the model stability and ultimately used an ensemble scoring strategy41 to calculate the median binding score as the final binding score on the basis of the five models from fivefold CV to minimize random prediction errors in individual models (Extended Data Fig. 2c and Methods). Additionally, the F0.5 score, which assigns twice the weight to precision compared to recall, has been widely adopted in virtual screening42,43,44 and was used to determine the classification threshold of SMRTnet (Extended Data Fig. 2d and Methods).
Evaluating SMRTnet’s performance on SRI data from PDB
SMRTnet demonstrated robust performance with an average area under the receiver operating curve (auROC) of 0.830–0.844 across varying positive-to-negative sample ratios (1:1, 1:2, 1:3, 1:4, 1:5 and 1:10) on the SMRTnet dataset over fivefold CV (Fig. 2a and Extended Data Fig. 2e). This performance remained consistent under different random seeds used to generate negative samples, supporting that the randomization process in sampling had minimal impact on model performance (Extended Data Fig. 2g). For comparison, RNAmigos2—the only other tool that can be applied to the test set of SMRTnet—achieved lower auROC values of 0.567–0.596 (Fig. 2a and Extended Data Fig. 2e).
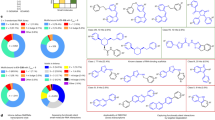
a, Performance comparison between SMRTnet and RNAmigos2 on the original test set of SMRTnet, the revised test set applying a maximum Tanimoto similarity threshold of 0.7 and the revised test set excluding identical multistrand binding sites. Bar plots with underlaid dot plots represent the auROC of SMRTnet and RNAmigos2 evaluated on the SMRTnet dataset with a 1:2 positive-to-negative ratio and its variants based on fivefold CV (n = 5). The bold solid line represents the s.d. of fivefold CV, presented as the mean ± s.d. b, Performance comparison between SMRTnet retrained on the RNAmigos2 subset versus RNAmigos2 on the full test set of RNAmigos2 using decoys from different libraries, including ChEMBL (n = 500), PDB (n = 264) and ChEMBL + PDB (n = 764). Data are presented as box and violin plots with underlaid dot plots (n = 37), where the middle line is the median, the lower and upper edges of the rectangle are the first and third quartiles, the lower and upper whiskers of the box plot represent the interquartile range (IQR) × 1.5 and mean values for each distribution are shown as numbers. c, Statistics of small molecule–RNA pairs across five experimental validated benchmarks. Bar plots represent the number of interacting (colored) and noninteracting (gray) small molecule–RNA pairs across five benchmarks, collectively referred to as the SMRTnet-benchmark dataset. d, ROC curves and auROC values of SMRTnet across the SMRTnet-benchmark dataset, including NALDB (n = 524), SMMRNA (n = 459), R-SIM (n = 423), R-BIND (n = 389) and NewPub (n = 216). e, Donut chart shows the percentage distribution of eight RNA types across the SMRTnet-benchmark dataset. f, Performance of SMRTnet across eight RNA types, including aptamer (n = 463), viral RNA (n = 294), miRNA (n = 202), riboswitch (n = 200), repeat expansion (n = 148), mRNA (n = 118), rRNA (n = 117) and others (n = 107). Right, bar chart representing the number of interacting (colored) and noninteracting (gray) small molecule–RNA pairs for each RNA type. Top: bar chart representing the overall performance across different RNA types (n = 8); the bold solid line represents the s.d. of the performance across eight RNA types. g, Overall performance of SMRTnet against current field-leading tools in the decoy evaluation task using the test set of SMRTnet (n = 684), including RNAmigos (n = 605), RNAmigos2 (n = 547), AutoDock Vina (n = 659), NLDock (n = 351), RLDOCK (n = 212) and rDock (n = 391). Data are presented as violin plots with underlaid box plots, where the middle line is the median, the lower and upper edges of the rectangle are the first and third quartiles and the lower and upper whiskers of the violin plot represent the IQR × 1.5. h, Comparison of the target molecule’s rank within the given decoy library across different computational tools. Overlapping arrows represent tied rankings. Arrows closer to the left indicate a higher rank for the target molecule, while those closer to the right indicate a lower rank. Each target molecule has 30 decoys. i, Performance of SMRTnet in identifying binding sites across different datasets. Violin plots represent the auROC of SMRTnet for binding-site identification on the test set of SMRTnet (n = 2,508), TE18 (n = 9), RB19 (n = 11) and interactions with experimentally determined binding sites in the SMRTnet-benchmark dataset (n = 5), where the middle line is the median, the lower and upper edges of the rectangle are the first and third quartiles and the lower and upper whiskers of the violin plot represent the IQR × 1.5. Numbers above each violin plot indicate the average auROC values for binding-site identification in each dataset. j–n, Binding-site predictions for the MYC IRES (j), HIV-1 TAR element (k), the r(CUG) repeat expansion in the HTT gene (l), the pre-miR-18a (m) and the guanine riboswitch (n). The two heat map tracks show the SMRTnet model’s response at each nucleotide position, with red color indicating HARs (top track, sequence response; bottom track, structure response). In the RNA structure diagram, red circles and shading indicate the known binding sites for the corresponding small molecules and labels around the binding sites denote their names. The binding scores are calculated between each small molecule and RNA pair and the Pearson correlation coefficients (r) and auROC values are calculated between the gradient signals and the proximity to experimentally determined binding sites.
To evaluate potential data leakage from small-molecule structural similarity, we calculated the Tanimoto similarity between molecules in the training and test sets and found that the average value consistently remained <0.75 across fivefold CV (Extended Data Fig. 2h). We further enforced a maximum Tanimoto similarity threshold of 0.7–1.0 and found that model performance remained stable with average auROC values of 0.844–0.855, indicating that no obvious data leakage occurred even without small-molecule similarity constraints (Fig. 2a, Extended Data Fig. 2i and Methods).
To examine data leakage from RNA-binding-site similarity, we excluded RNAs in the test set that shared identical multistrand binding sites with training set RNAs (Methods). This resulted in a performance decrease in the average auROC value, from 0.844 to 0.798 (Fig. 2a and Extended Data Fig. 2j). Furthermore, we retrained SMRTnet on the SMRTnet dataset using a structure-based data-splitting strategy24, which clusters structurally similar binding pockets (RMscore ≥ 0.75) into the same training or test set, resulting in an average auROC of 0.806 (Extended Data Fig. 2k). These results indicated that, while minor data leakage may occur on the RNA side, SMRTnet maintains robust predictive performance even when excluding identical or similar RNA-binding sites between training and test sets.
We further investigated SMRTnet’s robustness by benchmarking it against RNAmigos2 on RNAmigos2’s own dataset, either retraining and evaluating SMRTnet on the RNAmigos2 dataset or assessing the original SMRTnet on a filtered RNAmigos2 test set that excluded data with similar ligands and binding sites (Methods). In both tests, SMRTnet demonstrated highly competitive performance to RNAmigos2, despite being trained on a much smaller training dataset and/or evaluated on novel chemical and structural spaces (Fig. 2b and Extended Data Fig. 2g). These results underscore that SMRTnet is a robust and generalizable method for RNA-targeting drug discovery.
To investigate whether SMRTnet learns specific RNA–ligand interactions rather than relying on ligand-centric features. We created a test set of mismatched RNA–ligand pairs by swapping the ligand in each positive sample in the test set with a randomly selected ligand from another pair, while preserving the RNA and the ‘positive’ label (Methods). We found that SMRTnet exhibited a drastic performance decline (auROC = 0.572). We then modified the labels for these same mismatched pairs from ‘positive’ to ‘negative’. This single change fully restored the model’s high performance to an auROC of 0.830 (Extended Data Fig. 2l). These results provide evidence that SMRTnet has learned the underlying principles of SRIs.
Evaluating SMRTnet’s performance on SRI data from published studies
In addition to the SMRTnet dataset generated from PDB, we further curated a dataset of experimentally validated SRIs (together with noninteracting small molecule–RNA pairs) from four databases (R-BIND45,46, R-SIM47, SMMRNA48 and NALDB49) and 22 new publications (NewPub). After excluding the SRIs that already appeared in the SMRTnet dataset, we obtained 1,665 SRIs and 346 noninteracting small molecule–RNA pairs (that is, negative samples) (Fig. 2c, Supplementary Table 2 and Methods), collectively termed the SMRTnet-benchmark dataset.
We extracted RNA sequences and their secondary structures from the relevant publications to run SMRTnet and found that SMRTnet achieved an average auROC of 0.720 on the SMRTnet-benchmark dataset, with performance ranging from 0.684 on the SMMRNA subset to 0.765 on the NewPub subset (Fig. 2d). Furthermore, we categorized these data into eight categories on the basis of RNA types and observed that SMRTnet exhibited no clear preference for any specific RNA types, except for riboswitches (Fig. 2e,f). These results underscore SMRTnet’s broad applicability to diverse RNA types and its utility in predicting unseen SRIs.
SMRTnet outperforms other computational methods in the decoy evaluation task
We benchmarked SMRTnet against existing methods using decoy evaluation, a commonly used task for assessing the discriminative capacity of docking tools23,50. In this task, for each RNA target and its true binder, a library of decoys is designed to be physically similar to yet chemically distinct from the true binder51. The tools then rank the true binder among these decoys. Specifically, for each SRI in the test set of SMRTnet, we used DecoyFinder51 to retrieve up to 30 decoys23 from the ZINC15 database52 (Methods) and compared SMRTnet to four leading docking tools (AutoDock Vina, NLDOCK, RLDOCK and rDock) and two deep learning tools (RNAmigos and RNAmigos2).
SMRTnet achieved a mean ranking of 92.6% on the test set of SMRTnet, outperforming the four docking tools (ranging from 27.3% to 46.6%) and surpassing the two deep learning tools (ranging from 16.0% to 23.8%) (Fig. 2g). Specifically, SMRTnet consistently ranked the true binder within the top five across seven representative cases from the test set (Fig. 2h). As an example, for the theophylline-binding aptamer (PDB 1O15), SMRTnet ranked theophylline at the top, followed by decoys sharing the same functional group, while decoys lacking this functional group were ranked at the bottom (Extended Data Fig. 3a–h). These results demonstrate SMRTnet’s superior ability to identify true binders from structurally closely related small molecules.
We also evaluated SMRTnet’s computational efficiency by measuring inference times for predicting interactions across varying numbers of SRIs (Extended Data Fig. 2m and Methods). The results demonstrate that SMRTnet offers substantially greater computational efficiency than GPU-accelerated docking tool Vina-GPU 2.0 (ref. 53).
RNA encoders and experimentally derived RNA secondary-structure data are essential for accurate SRI prediction
We conducted ablation studies to assess the contributions of different SMRTnet components to its performance. Specifically, we defined seven test variants of SMRTnet: (1) RNA structure and small-molecule structure encoder; (2) RNA sequence and small-molecule SMILES encoder; (3) two small-molecule encoders with RNA structure encoder; (4) two small-molecule encoders with RNA sequence encoder; (5) two RNA encoders with small-molecule SMILES encoder; (6) two RNA encoders with small-molecule structure encoder; and (7) the full SMRTnet model without the MDF module (Methods).
The auROC of SMRTnet decreased from 0.844 to 0.812 for variant 7 and slightly further to 0.808 and 0.802 for variants 5 and 6, respectively. It then dropped substantially to 0.561, 0.552, 0.571 and 0.591, respectively, for variants 1–4 (Extended Data Fig. 2n). These findings suggest that both RNA sequence and structure information are the most critical, while the MDF module also contributes to achieving high predictive performance.
Next, we focused on RNA secondary structure and investigated its contributions to SMRTnet’s performance. We trained a sequence-only version of SMRTnet (termed SMRTnet-seq) by excluding RNA structural features. This reduced the average auROC from 0.844 to 0.760 on the SMRTnet dataset and from 0.720 to 0.578 on the SMRTnet-benchmark dataset (Extended Data Fig. 2o–p). We also replaced the retrieved structural data (which, for more than 80% cases, were experimentally derived) with predictions from RNAstructure54 on the SMRTnet-benchmark dataset. This substitution reduced the auROC value to 0.664 (Extended Data Fig. 2p). These findings underscore the essential role of RNA secondary structures—especially those determined experimentally—in attaining high predictive accuracy.
Additionally, we evaluated the impact of using different RNA LLMs on SMRTnet’s performance, comparing implementations based on RNASwan-seq to RNA-FM55 and RNAErine56. The results show that SMRTnet performed slightly better when using RNASwan-seq over the two alternatives (Extended Data Fig. 2q).
SMRTnet identifies small-molecule-binding sites on RNA
Previous studies have shown that model interpretability analysis can reveal binding sites for various molecular interactions57,58. Following this approach, we identified small-molecule-binding sites on RNA targets as regions of heightened importance, termed high-attention regions (HARs). Specifically, we applied the Grad-CAM algorithm59 to quantify the contribution of each nucleotide to the predicted binding scores of the SRIs (Methods).
We assessed the accuracy of this binding-site identification approach by comparing the gradient signals to experimentally determined binding sites from four datasets. First, on the SMRTnet dataset, where binding-site information is fully available, SMRTnet achieved an average auROC of 0.695 across fivefold CV. Second, on two additional benchmark datasets from a prior study presenting RNAsite, a specialized binding site prediction tool57, SMRTnet achieved auROCs of 0.741 and 0.770, after excluding invalid entries, matching the performance of RNAsite57 (Fig. 2i and Methods).
Lastly, focusing on the SMRTnet-benchmark dataset, we examined five representative SRIs with known binding-site information: MYC-RiboTAC binding to the MYC IRES at the internal loop (5′ UUCG/3′ ACCC)60, a thienopyridine derivative binding to the HIV-1 TAR RNA around the 5′ AUCUG/3′ U_C bulge61, lomofungin binding to the highly structured r(CUG) repeat expansion in the 3′ untranslated region (UTR) of the dystrophia myotonica protein kinase mRNA at the internal loop (5′ CUG/3′ GUC)62, a benzimidazole-derivative dimer binding to pre-miR-18a at the 5′ GAUAG_U/3′ C_AUCUA bulges63 and 6-N-hydroxylaminopurine binding to the guanine riboswitch at the three-way junction (5′ AUAAGUU/UUCGUUUCUACC/3′ UCG)64. SMRTnet attained an average auROC of 0.793 on these interactions (Fig. 2i). We further examined how precisely predicted binding sites matched experimentally determined ones by calculating the Pearson correlation coefficient (r) between the gradient signals and the proximity to experimentally determined binding sites (Methods). We found that SMRTnet’s predicted HARs closely matched all these sites (Fig. 2j–n; r = 0.550, 0.783, 0.214, 0.322 and 0.527). Together, these findings highlight SMRTnet’s precision in identifying small-molecule-binding sites across diverse RNA structures.
SMRTnet predicts compounds that bind to disease-associated RNA targets
We applied SMRTnet to screen for compounds capable of binding to ten disease-associated RNA targets using a curated library consisting of 7,350 compounds of natural products and metabolites (Supplementary Table 3 and Methods). In addition to the MYC IRES, these RNA targets included pre-miR-155, which is overexpressed in various cancers60, HOTAIR helix 7, which represses tumor and metastasis suppressors65, HIV-1 Rev response element (RRE) IIB element, a critical determinant of viral replication66, and CAG repeat expansion of HTT, which causes Huntington disease67. Additionally, we included five other RNA structural elements from the 5′ UTR of SARS-CoV-2 (ref. 68): stem loop 1 (SL1), SL2/3 involved in viral replication69,70,71,72, SL4 associated with subgenomic RNA synthesis72,73 and SL5a and SL5b implicated in viral packaging74,75. We obtained both the sequence and secondary structures of these RNA targets as the input for SMRTnet. These RNAs exhibit distinct secondary structures, allowing us to probe the performance of SMRTnet across a broad range of RNA molecules and structural features (Fig. 3a and Supplementary Table 4).
a, Top: the secondary structures of ten disease-associated RNA targets with predicted minimum free energy scores computed by RNAstructure. Bottom: the distribution of binding scores for these RNA targets as predicted by SMRTnet on the natural product library. Data are presented as box plots with underlaid dot plots, where the middle line is the median, the lower and upper edges of the rectangle are the first and third quartiles, the lower and upper whiskers of the box plot represent the IQR × 1.5 and the green triangle is the average. Each point represents the predicted binding score of a compound to a target (n = 7,350) and the gray dashed line marks the classification threshold (0.704) of SMRTnet. b, Validation rate of SMRTnet predictions selected for MST, where the colored areas represent compounds validated as binding and the gray areas represent compounds validated as nonbinding. The numerator indicates the number of compounds experimentally validated as binders, while the denominator shows the total number of top-ranked compounds selected for experimental validation. The percentage denotes the validation rate for each RNA target. c, Structural similarity (measured by the Tanimoto similarity) of the 40 compounds that interact with ten disease-associated RNA targets. Compounds were grouped by their respective RNA targets. d, Bar plots represent the Kd for 40 experimentally validated compound–RNA interactions across ten disease-associated RNA targets. Compounds were grouped by their respective RNA targets and labeled by CAS number. Kd values (μM) are plotted on a logarithmic scale. The gray dashed line indicates Kd > 100 μM, serving as a threshold for weaker interactions.
For each RNA target, we predicted the binding scores for all 7,350 compounds using SMRTnet. The top 20 compounds for each target were then selected on the basis of their binding scores (also requiring the scores to be higher than the classification threshold of 0.704), resulting in 190 predicted SRIs for experimental validation (Fig. 3a). We used the binding check mode of MST as the main assay and incorporated the SDS denaturation test (SD-test) to validate intrinsically fluorescent compounds that may interfere with MST measurements (Extended Data Fig. 4 and Methods). Of the 190 predicted SRIs, 40 were experimentally validated, achieving an average validation rate of 21.1% (Fig. 3b). Each target also exhibited a unique profile of validated binders (Fig. 3c), underscoring SMRTnet’s ability to discern subtle differences in RNA structure and ligand specificity.
We subsequently determined the Kd of the 40 validated SRIs using the binding affinity mode of MST and found that most SRIs exhibited micromolar-level affinities, a range commonly observed for initial hits in RNA-targeting drug discovery. However, six SRIs exhibited nanomolar-level affinities (Fig. 3d and Extended Data Fig. 5). These findings highlight SMRTnet’s robust ability to predict SRIs across diverse disease-related RNA targets with the potential for the identification of high-affinity binders.
SMRTnet-predicted binding scores correlate with experimental validation for the MYC IRES
The oncogenic transcription factor MYC is a hallmark of many human cancers, yet it is generally considered ‘undruggable’. However, recent studies suggested that targeting the MYC IRES, located in the 5′ UTR of MYC mRNA and responsible for cap-independent translation of MYC, may represent a viable strategy for controlling this oncogene60,76. To explore this further, we performed a large-scale experimental validation by randomly sampling a subset of 376 compounds from the aforementioned compound library with varying predicted binding scores and high chemical structure diversity (Fig. 4a–c, Supplementary Table 5 and Methods).
a, Distribution of predicted binding scores predicted by SMRTnet for the natural product library (left) and a downsampled subset (right) targeting MYC IRES. Data are presented as box plots with underlaid dot plots, where the middle line is the median, the lower and upper edges of the rectangle are the first and third quartiles, the lower and upper whiskers of the box plot represent the IQR × 1.5, and the green triangle is the average. Each point represents the predicted binding score of a compound to MYC IRES. b, Donut chart shows the number and percentage of compounds sampled across ten prediction intervals. c, A t-distributed stochastic neighbor embedding (TSNE) analysis of Morgan fingerprint for different compounds datasets, including the natural product library (n = 7,350), the downsampled subset (n = 376), and experimentally validated binders determined by MST from the downsampled subset (n = 15). d, Distribution of selected compounds from the downsampled subset (right) and their validation rate (left) across ten prediction intervals. The numerator indicates the number of compounds experimentally validated as binders, while the denominator shows the total number of compounds randomly selected for experimental validation. The percentage denotes the validation rate within each interval. e, Structural similarity (measured by the Tanimoto similarity) of the 15 experimentally validated compounds that interact with MYC IRES. f, Bar plots represent the predicted binding scores and Kd values (μM) for 15 experimentally validated compounds targeting MYC IRES. Compounds were grouped by their respective prediction intervals and labeled by CAS number. Top: the gray dashed line represents the classification threshold (0.704) of SMRTnet. Bottom: the gray dashed line indicates Kd > 100 μM, serving as a threshold for weaker interactions.
A total of 15 compounds from the subset library were validated as binders of MYC IRES through MST, revealing a clear positive correlation between the predicted binding scores and the validation rate. Specifically, two of seven predictions (28.6%) with binding scores in the range of 0.9–1.0 were validated as positive and the validation rate decreased steadily with lower binding scores, dropping to zero of 93 (0%) in the range of 0.0–0.1 (Fig. 4d, Extended Data Fig. 6a and Methods). These results demonstrate that SMRTnet effectively prioritizes true binders with high prediction scores. Additionally, the Tanimoto similarity analysis of these 15 compounds revealed generally low structural similarity (Fig. 4e). We further quantified the Kd of these 15 hits and found that they all exhibited micromolar-level affinities with no correlation between binding affinities and binding scores (Fig. 4f and Extended Data Fig. 6b).
SMRTnet identifies the binding site of irinotecan hydrochloride trihydrate (IHT) on the MYC IRES
Among the 15 compounds identified as binders of the MYC IRES, only IHT (CAS: 136572-09-3) meets the criteria for drug-development potential, including physicochemical properties and medicinal chemistry characteristics, as evaluated by ADMETlab 3.0 (ref. 77) (Extended Data Fig. 7a–j). To further elucidate the molecular basis of its interaction with the MYC IRES, we used SMRTnet to predict the binding site of IHT and HAR analysis pinpointed the interaction to the internal loop (5′ UUCG/3′ ACCC) of the MYC IRES (Fig. 5a and Methods). This prediction was further corroborated by molecular docking, which localized IHT precisely within the same internal loop (Fig. 5b).
a, Potential binding site of IHT on the MYC IRES. The RNA secondary structure diagram highlights the HARs in red and the two heat map tracks show the model’s response at each nucleotide position—with the top track representing sequence response and the bottom track representing structure response (red indicates high attention). b, The tertiary structure of MYC IRES precited using SimRNA87, with the cyan-colored region indicating the predicted binding site by SMRTnet; molecular docking results for IHT at this predicted site are visualized in ChimeraX88. c, The secondary structure of 20 MYC IRES mutants, predicted using RNAstructure. Different background colors represent different binding-site types (labeled above) and colored nucleotides mark the positions and types of mutations. The lavender background denotes a fully base-paired structure, the linen background denotes an upper 1 × 1 internal loop, the misty-rose background denotes a lower 1 × 1 internal loop, the papaya-whip background denotes a 2 × 2 internal loop and the light-gray background denotes a 3 × 3 internal loop. WT, wild type. d, Distribution of predicted binding score for different mutant MYC IRES against IHT. Bottom, the bar plots indicate experimental validation results using MST. The red dot represents the average binding scores across the same binding-site types and the red dashed line represents the trend in binding scores as the binding-site type changes. The gray dashed line marks the classification threshold (0.704) of SMRTnet. e, Radar plot shows the agreement between computational predictions and experimental validations for the 20 MYC IRES mutants with respect to IHT. f, The chemical structure and full name of the IHT.
To validate this predicted binding site, we designed 20 mutant variants of the MYC IRES, which we grouped into five categories: (1) mutants preserving the 2 × 2 internal loop but with altered sequence; (2–4) mutants converting the internal loop into 3 × 3 or 1 × 1 configurations with concomitant sequence changes; and (5) mutants in which the loop was entirely removed, yielding a fully base-paired structure (Fig. 5c and Supplementary Table 6). Analysis revealed a progressive decline in predicted binding scores from the 2 × 2 configuration to the 1 × 1 forms, with the lowest scores observed in the fully base-paired mutants. Interestingly, predicted binding scores increased when altering from the 2 × 2 configuration to the 3 × 3 forms (Fig. 5d and Extended Data Fig. 8a–e). These trends in predicted binding scores were highly consistent with experimental validation rates, attesting to the binding sites of IHT on the MYC IRES and underscoring the reliability of SMRTnet’s binding-site predictions (Fig. 5e,f).
An MYC IRES-targeting compound represses MYC expression and cell proliferation
We examined the biological consequences of IHT binding to the MYC IRES in various cancer cells. In HeLa cells, IHT treatment resulted in a reduction in MYC mRNA levels by ~56.9% and MYC protein levels by ~71.6%, surpassing the effects observed with MYC-RiboTAC (Fig. 6a,b and Supplementary Tables 7 and 8), a chimeric molecule that binds the MYC IRES and induces MYC mRNA degradation by recruiting RNaseL60. Additionally, in three cancer cell lines (HeLa, Jurkat and Raji) that are known to require MYC for optimal proliferation60,78,79, IHT decreased cell proliferation by 19.6–48.4% and increased cell apoptosis by 56.6–124.2% in these three cancer cell lines that depend on MYC for optimal growth. It is again noteworthy that IHT had a higher impact on cell proliferation and viability than MYC-RiboTAC, consistent with IHT’s higher impact in reducing MYC mRNA and protein levels than MYC-RiboTAC (Fig. 6c–h and Supplementary Tables 9 and 10).
a,b, The effect of MYC-RiboTAC and IHT on MYC mRNA levels (a) and MYC protein levels (b) in HeLa cells was determined using qPCR and western blotting at a 10 μM dose after 48 h of treatment. Data are presented as the mean ± s.d. from n = 3 biological replicates. The P values were determined using a two-sided unpaired Student’s t-test. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 and ****P ≤ 0.0001; NS, not significant (P > 0.05). c–e, The effect of MYC-RiboTAC and IHT on the proliferation of HeLa (c), Jurkat (d) and Raji (e) cells was measured at a 10 μM dose after 48 h of treatment. Data are presented as the mean ± s.d. from n = 6 biological replicates. P values were determined using a two-sided unpaired Student’s t-test. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 and ****P ≤ 0.0001; NS (P > 0.05). f–h, The effect of MYC-RiboTAC and IHT on the apoptosis of HeLa (f), Jurkat (g) and Raji (h) cells was measured at a 10 μM dose after 48 h of treatment. Data are presented as the mean ± s.d. from n = 6 biological replicates. P values were determined using a two-sided unpaired Student’s t-test. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 and ****P ≤ 0.0001; NS (P > 0.05). i, Schematic diagram of the luciferase reporter system used to measure the translational activity of MYC IRES. j,k, The effect of MYC-RiboTAC and IHT on MYC IRES (j) and fully base-paired MYC IRES reporter (k) in HEK293T cells was measured at a 10 μM dose after 48 h of treatment. Renilla luciferase expression was normalized to firefly luciferase expression, showing the translational activity of MYC IRES. Data are presented as the mean ± s.d. from n = 6 biological replicates. P values were determined using a two-sided unpaired Student’s t-test. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001 and ****P ≤ 0.0001; NS (P > 0.05).
We further studied the impact of IHT on an MYC IRES luciferase reporter in HEK293T cells, using the fully base-paired MYC IRES as the control luciferase reporter (Fig. 6i and Methods). Similar to the effect of MYC-RiboTAC, IHT reduced luciferase levels for the MYC IRES reporter (5′ UUCG/3′ ACCC) by ~14.2% but not for the control luciferase reporter (5′ UAUG/3′ AUGC) (Fig. 6j,k and Supplementary Table 11). These results indicate that SMRTnet could assist in identifying small molecules that bind to disease-associated RNA targets with bioactivity and therapeutic potential.
Discussion
Here, we present SMRTnet, a deep learning method to predict SRIs using RNA sequence and secondary structure alongside small-molecule SMILES. SMRTnet distinguishes itself from current learning-based tools in three key aspects23,24. First, by incorporating secondary-structure information, which is readily obtained from experimental approaches such as icSHAPE80 and SHAPE-MaP81, SMRTnet captures accurate SRI determinants that sequence-only methods miss, thus improving prediction accuracy. This secondary-structure-based design also extends applicability to disease-associated RNA targets that lack well-defined tertiary structures. Second, SMRTnet integrates two LLMs trained on billions of RNA or small-molecule data, improving the representation capacity for both modalities and facilitating accurate predictions. Third, the MDF module integrates the features of RNA and small-molecule encoders to generate an interaction-aware representation that robustly characterizes the probability of an SRI. We also demonstrate the high prediction performance of SMRTnet across various experimental benchmarks and large-scale experiments. These results showcase SMRTnet’s potential for accelerating the discovery of RNA-targeting therapeutics.
The current knowledge of experimentally validated SRIs remains limited in terms of RNA types and small-molecule diversity82,83. Therefore, experimental efforts must prioritize high-throughput screening platforms capable of efficiently profiling large-scale interactions in a multiplexed format (for example, small-molecule libraries versus diverse RNA targets). Such datasets would serve as critical training resources for AI-driven drug discovery, mirroring how the vast number of small molecule–protein interactions has transformed our ability to predict small molecule–protein recognition and binding84,85,86.
It bears emphasis that binding alone does not inherently imply that a small molecule will regulate RNA expression or exert biological activity. As multi-omics data continue to accumulate (for example, chemical-RNA interactomes, transcriptomics and functional screens), future research can focus on integrating these multimodal layers—particularly gene expression profiles and phenotypic readouts—to develop AI methods capable of simultaneously predicting binding interactions and downstream biological effects, bridging the gap between target engagement and functional impact and advancing RNA-targeting therapeutics.
Methods
SMRTnet dataset collection and processing
To train SMRTnet, we first filtered 2,477 structures (1.27%) from 195,340 structures in the PDB (as of January 2024) that contained at least one RNA and one small molecule using atomium89. We then removed nontherapeutically relevant small molecules and structures with fewer than 31 RNA residues, as well as binding sites where more than 50% of the residues within 10 Å were proteins. This filtering process yielded 1,061 high-quality SRI structures. Next, we converted the RNA tertiary structures from these 1,061 SRIs into secondary structures using DSSR90 and identified the binding positions of RNA residues within 10 Å of the small molecule using atomium23,89. These binding positions were subsequently extended by 15 nt in both the 5′ and the 3′ directions to generate 31-nt RNA fragments. Finally, we converted the chemical structures of small molecules to canonical SMILES using the RDKit91 and each SMILES was paired with its corresponding RNA fragment to form a positive sample. Additionally, we generated negative samples by randomly pairing RNA fragments with small molecules after removing known interaction pairs and maintained specific positive-to-negative sample ratios (1:1, 1:2, 1:3, 1:4, 1:5 and 1:10) using different random seeds (1, 2, 3, 4 and 42). This process resulted in the SMRTnet dataset. The SMRTnet dataset was divided into training (80%), validation (10%) and test (10%) sets using a ligand-based data-splitting strategy, ensuring that no SMILES in the test set appeared in the training or validation sets. We also used fivefold CV to evaluate model stability and ensured that no SMILES were shared between the validation or test sets across different folds. Details of the SMRTnet dataset are provided in Supplementary Table 1.
To evaluate potential data leakage in SMRTnet, we constructed several revised versions of the test set of SMRTnet. On the ligand side, we tested a range of maximum similarity threshold from 0.7 to 1.0, with intervals of 0.1 (with 1.0 corresponding to our ligand-based data-splitting strategy). For each threshold, we excluded small molecules (along with their corresponding RNAs) from the test set of SMRTnet if their Tanimoto similarity with any training set small molecules exceeded the specified threshold. On the RNA side, we generated a multistrand-binding-site exclusion test set of SMRTnet by removing RNAs (along with their corresponding small molecules) that shared identical multistrand binding sites with any RNA in the training set across fivefold CV.
To investigate potential biases of quantitative structure–activity relationship in SMRTnet, we constructed a modified test set of SMRTnet, termed the RNA′ test set of SMRTnet. Specifically, during each round of fivefold CV, each ligand in the positive samples in the test set was replaced with a different ligand randomly sampled from the test set, while keeping the corresponding all fragments within an RNA pocket and label unchanged. In a further variant, instead of preserving the positive label of the permuted samples, we modified them as negative and combined it with the original test set of SMRTnet, termed the RNA″ test set of SMRTnet.
SMRTnet-benchmark dataset collection and processing
To evaluate SMRTnet, we constructed a benchmark dataset comprising 2,011 experimentally validated SRIs and noninteraction pairs, including 1,665 interaction pairs and 346 noninteraction pairs. This dataset, referred to as the SMRTnet-benchmark dataset, excludes any small molecule–RNA pairs present in the SMRTnet dataset (deduplicated with 100% identity). Specifically, we obtained the RNA sequences and their secondary structure from the corresponding publications, manually drew the chemical structures of the small molecules on the basis of these publications and converted them into canonical SMILES using the Open Babel92 web server. In total, we collected 1,795 interaction and noninteraction pairs from 178 papers across four published databases: R-BIND45,46, R-SIM47, SMMRNA48 and NALDB49. Additionally, we collected 216 interaction and noninteraction pairs not included in any of these databases from 22 new publications9,60,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112, which we refer to as the ‘NewPub’ subset. Details of the SMRTnet-benchmark dataset are provided in Supplementary Table 2.
Drug-screening dataset collection and processing
To evaluate SMRTnet’s capability in drug discovery, we constructed datasets for identifying SRIs, referred to as the drug-screening datasets. These datasets comprised ten disease-associated RNA structural elements and a curated library containing 7,350 compounds of natural products and metabolites. Specifically, the RNA structural elements included MYC IRES60, pre-miR-155 (ref. 60), HOTAIR helix 7 (ref. 65), CAG repeat expansion in HTT gene113, HIV RRE IIb (ref. 114) and five elements in SARS-CoV-2 5′ UTR regions68 (including SL1, SL2/3, SL4, SL5a and SL5b). We truncated each full-length RNA to 31 nt and predicted its secondary structure from sequence using RNAstructure54, ensuring that the predicted secondary structures were consistent with experimentally validated RNA secondary structures reported in previous studies. Additionally, we collected several natural product libraries from the in-house chemical library of the Center of Pharmaceutical Technology, Tsinghua University (http://cpt.tsinghua.edu.cn/hts/), including the natural product library for high-throughput screening (n = 4,160), the BBP natural product library (n = 3,200), the TargetMol natural compound library (n = 409), the MCE natural product library (n = 1,384) and the Pharmacodia natural product library (n = 935). From these collections, we constructed a natural product library comprising 7,350 unique compounds with distinct CAS numbers and obtained their canonical SMILES using RDKit91. The compounds used in the experiments were purchased from Topscience. Details of the drug-screening datasets are provided in Supplementary Tables 3 and 4.
We applied a stratified sampling approach to generate a downsampled subset of compounds for experimental validation of MYC IRES binding on the basis of the full natural product library. Specifically, we performed binding predictions for MYC IRES using SMRTnet against the full natural product library (n = 7,350) and then divided the predicted binding scores into ten intervals with increments of 0.1. From each interval, we randomly selected 5% of the compounds, except for the 0.9–1.0 interval, from which 100% of compounds (n = 7) were included because of the limited number in this interval. This stratified sampling approach resulted in a downsampled screening library of 376 compounds for experimental validation. Details of the a downsampled drug-screening library are provided in Supplementary Table 5.
MYC IRES mutation dataset construction
To evaluate model interpretability and identify the small-molecule-binding sites on MYC IRES, we constructed an MYC IRES mutation dataset comprising 20 mutant RNAs. Specifically, we generated five distinct types of RNA-binding sites by altering the nucleotides within the predicted small-molecule-binding region, each type consisting of four RNA variants. The original 2 × 2 internal loop of MYC IRES was altered to form two types of 1 × 1 internal loops, 3 × 3 internal loops and fully complementary base-paired structures. We also preserved the original 2 × 2 internal loop by modifying only its sequence. All 20 mutant RNAs were folded to satisfy minimum free energy criteria, ensuring that they adopted the expected conformations consistent with in vitro experiments. Details of the MYC IRES mutation dataset are provided in Supplementary Table 6.
Architecture of SMRTnet
SMRTnet is a binary classification model with three inputs (RNA sequence, RNA secondary structure and small-molecule SMILES) and one output (binding score). Specifically, each RNA sequence is a 31-nt sequence composed of {A, U, C, G}. The RNA secondary structure in each sample is represented by 31-nt dot-bracket notations using {‘(‘, ‘.’, ‘)’}. The small molecule is encoded using canonical SMILES, processed by RDKit91. Labels in each sample are binary with two symbols (‘1’ for positive samples and ‘0’ for negative samples). During inference, SMRTnet processes input RNAs (≥31 nt, regardless of the presence of a known binding site) with a sliding-window approach. The binding score is computed as follows:
[where x is the RNA sequence, y is the RNA secondary structure with dot-bracket notation, z is the canonical SMILES of small molecule and z′ is the two-dimensional (2D) molecular graph of the small molecule derived from z using RDKit91. σ is the sigmoid activation function, \({f}_{\mathrm{FC}}\) is a fully connected layer, \({f}_{\mathrm{MDF}}\) is the MDF module, \({f}_{\mathrm{CNN}-\mathrm{Res}}\) is the RNA structure encoder based on CNN with ResNet, \({f}_{\mathrm{RNASwan}-\mathrm{seq}}\) is the RNA sequence encoder based on RNA language model, \({f}_{\mathrm{MoLFormer}}\) is the drug sequence encoder based on chemical language model and \({f}_{\mathrm{GAT}}\) is the small-molecule structure encoder based on GAT115. The model output is transformed from the output value of the \({f}_{\mathrm{FC}}\) through the σ activation layer.
RNA sequence encoder
We developed an RNA language model, RNASwan-seq, for learning RNA sequence representations (Extended Data Fig. 1a). The pretraining dataset for RNASwan-seq was compiled from seven sources: the European Nucleotide Archive116, National Center for Biotechnology Information nucleotide database117, GenBank118, Ensembl119, RNAcentral120, CSCD2 (ref. 121) and GreeNC 2.0 (ref. 122), encompassing a total of 470 million RNA sequences. Redundant sequences with 100% sequence identity were removed using MMSeqs2 (ref. 123), resulting in approximately 214 million unique RNA sequences. A random splitting strategy with a 30% sequence identity threshold was applied to divide the data into training and test sets for self-supervised training.
RNASwan-seq consisted of 30 transformer encoder blocks with rotary positional embeddings (RoPEs). Each block includes a feedforward layer with a hidden size of 640 and 20 attention heads. During training, a random cropping strategy was applied to extract 1,024-nt segments from the full-length RNA sequences in each iteration and 15% of nucleotide tokens were randomly selected for potential replacement. The model was trained using masked language modeling (MLM) to recover the original masked tokens using cross-entropy loss. A flash attention mechanism was used to accelerate the training process. The training process is formulated as an objective function as follows:
where \({\mathcal{M}}\) represents indices of masked tokens randomly sampled from each input sequence x. For each masked token, given masked sequence \({x}_{/{\mathcal{M}}}\) as context, the objective function minimizes the negative log likelihood of the true nucleotides \({x}_{i}\). Finally, the pretrained model was integrated into SMRTnet as an RNA sequence encoder using a fine-tuning strategy.
RNA structure encoder
We designed an RNA structure encoder (Extended Data Fig. 1b), adapted from our previous work31. This encoder represents the RNA sequence x (four-dimensional) using one-hot encoding and incorporates RNA secondary structure y (one-dimensional, 1D), expressed in dot-bracket notation, to form a five-dimensiaonal vector \(x{\prime}\). The embedding of the RNA structure encoder is computed as follows:
Here, \({f}_{{\rm{C}}}\) denotes the convolutional block, \({f}_{\mathrm{SE}}\) denotes the squeeze–excitation (SE) block and \({f}_{{\rm{R}}}\) denotes the residual block. These three blocks are defined as follows:
In \({f}_{C}\), \(\mathrm{ReLU}\) represents the rectifier linear unit activation function, \(\mathrm{BN}\) represents the batch normalization layer and \(\mathrm{Conv}\) represents the 2D convolutional layers. This configuration ensures that the output shape of the convolution layers matches that of the input shape.
In \({f}_{\mathrm{SE}}\), the SE block functions as a channel-wise self-attention mechanism that identifies binding-site patterns through weight recalibration. \(\bigotimes\) represents channel-wise multiplication between the input and the learned vector by SE block. The SE block first compresses the global sequence context using the global average pooling function \({f}_{\mathrm{sq}}\) and then transformed it into a set of channel-wise weights, scaled between 0 and 1, through a nonlinear transformation \({f}_{\mathrm{ex}}\), which consists of two fully connection layers and a ReLU activation function.
In \({f}_{{\rm{R}}}\), \({f}_{{\rm{R}}1}\) denotes residual blocks with 1D convolutional kernels, learning combined sequence and structural patterns, while \({f}_{{\rm{R}}2}\) denotes residual blocks with 2D convolutional kernels, capturing spatial context features that localize the precise binding site. The \(\mathrm{AvgPool}\) function is an average pooling layer that convert the 2D feature maps into the 1D vectors.
Small-molecule SMILES encoder
We introduced MoLFormer28, a chemical language model, as the small-molecule sequence encoder to represent the small molecules. The model uses a linear attention mechanism with RoPE to process SMILES derived from approximately 1.1 billion unlabeled molecules in the PubChem124 and ZINC125 databases. Specifically, MoLFormer is a transformer-based encoder with linear attention, comprising 12 layers, 12 attention heads per layer and a hidden state size of 768. As a result, each SMILES is encoded into an \(L\times 768\) matrix. Finally, we fine-tuned the final checkpoint of MoLFormer (N-Step-Checkpoint_3_30000.ckpt) to integrate it into SMRTnet.
Small-molecule structure encoder
We designed a three-layer GAT block as the small-molecule structure encoder, which adaptively learns edge weights and captures node representations through message passing for small molecule (Extended Data Fig. 1c). Specifically, each canonical SMILES is converted into a 2D molecular graph \(G=(V,E)\) using RDKit, where V represents the set of atomic nodes for molecule and \(E\) represents the set of edges connecting these nodes. Each node is represented by a 74-dimensional feature vector based on the DGL-LifeSci package126. For molecules with fewer nodes, virtual nodes (zero-padded) are added to ensure dimensional consistency. The embedding of a small-molecule structure encoder is computed as follows:
Here, \(\phi\) and \(\psi\) represent the learnable aggregation and attention functions, respectively. \({{\mathscr{N}}}_{i}\) represents the set of neighbors of atom i, l indexes the graph attention layer and \(v\) represents the feature vector of each node. Masked attention is applied to restrict computation to neighboring nodes, as defined by the adjacency matrix. Attention weights are normalized across all potential neighbors using the SoftMax function to ensure comparability. The final molecular representation is derived by iteratively aggregating the bond-connected atom features along with their associated chemical bond features.
MDF module
We proposed a MDF module consisting of three layers, including one coattention layer and two self-attention layers with different parameters (Extended Data Fig. 1d). Each layer incorporates residual blocks and layer normalization blocks, with concatenation blocks integrating multimodal features for the input to the next layer. This MDF module is defined as follows:
Here, \({E}_{r}\) is the output of RNA sequence encoder, \({E}_{s}\) is the output of RNA structure encoder, \({E}_{m}\) is the output of small-molecule SMILES encoder and \({E}_{t}\) is the output of the small-molecule structure encoder. The first fusion layer \({f}_{f1}\) contains a co-attention block, while the second (\({f}_{f2}\)) and third (\({f}_{f3}\)) fusion layers contain self-attention block. The three fusion layers are constructed as follows:
For the first fusion layer \({f}_{f1}\), we implemented a co-attention mechanism based on ViLBERT29, which extends the BERT architecture to a multimodal, two-stream model.
For the second fusion layer \({f}_{f2}\) and third fusion layer \({f}_{f3}\), a self-attention mechanism is used to further enhance the integration of the sequence and structure features of small molecule and RNA, resulting in a single embedding to predict the binding score.
Here, \({f}_{\mathrm{ln}}\) is the layer normalization block, \({f}_{\mathrm{MLP}}\) is the multilayer perceptron and \([]\) is the concatenation block. The output embedding dimension c is set to 128 across all three fusion layers. The number of attention heads \(h\) is set to 2 in \({f}_{f1}\) and \({f}_{f2}\) and to 8 in \({f}_{f3}\).
Training strategy of SMRTnet
SMRTnet uses supervised learning to predict binding scores between small molecules and RNA by minimizing the error between predicted binding scores and ground-truth labels. Specifically, SMRTnet optimizes its parameters by minimizing a loss function composed of binary cross-entropy loss and L2 regularization, calculated between the target labels T and predictions y across the training set.
Here, \({t}_{i}\) is the ground-truth label, \({y}_{i}\) is the predicted binding score, \({\mathbb{W}}\) represents all parameters of SMRTnet and \(N\) is the batch size. Model parameters were optimized using the Adam optimizer, an extension of stochastic gradient descent algorithm that adaptively adjusts step sizes and requires minimal hyperparameter tuning. Additionally, a warmup scheme with a linear scaling rule was applied to adjust learning rates during training.
To mitigate overfitting, each convolutional layer was followed by a batch normalization layer, each residual block was followed by a dropout layer and the L2 normalization on all parameters acted as a weight decay term to further reduce overfitting. Early stopping was used to halt SMRTnet training automatically when the validation auROC value did not improve for 20 consecutive epochs.
Ensemble scoring strategy of SMRTnet
We implemented an ensemble scoring strategy to enhance model robustness. Specifically, this strategy is defined as the median value of five models obtained from fivefold CV and a sliding-window approach was applied to process the input RNAs longer than 31 nt (refs. 31,41). The final binding score was calculated as follows:
Here, F represents the ensemble scoring strategy, \({f}_{\mathrm{median}}\left({r}_{i},{s}_{i}\right)\) represents the median value of five models obtained from fivefold CV. Given an input RNA r and small molecule s, a sliding-window approach (window size = 31 nt, step size = 1 nt) is used to compute the binding scores for each RNA segment \({r}_{i}\) and its corresponding small molecule \({s}_{i}\). For input RNAs with lengths between 31 nt and 40 nt (\(31 < L\le 40\)), the final binding score is defined as the maximum binding score across all windows. For input RNAs longer than 40 nt (\(L > 40\)), \(\phi (\bullet )\) is applied to identify potential binding regions, which are characterized by at least four consecutive windows with binding scores greater than 0.5. The final binding score is then taken as the maximum score within the identified potential binding regions. If no such potential binding regions exist, the minimum binding score across all windows is taken as the final binding score and the small molecule–RNA pair is classified as unbound (Extended Data Fig. 2c).
Hyperparameters of SMRTnet
The hyperparameters of SMRTnet were determined through grid search. Specifically, we extensively evaluated and empirically tuned the hyperparameters of each module to achieve optimal model performance. The final hyperparameter settings of SMRTnet are described below.
Batch size
A batch size of 32 was used for all experiments.
Learning rate
Base learning rates of 1 × 10−4 and 1 × 10−5 were applied to SMRTnet and to RNASwan-seq and MoLFormer, respectively. A warmup strategy was used to scale the learning rate by a factor of 8 during initial training epochs.
Training epochs
Models were trained for up to 100 epochs, with early stopping triggered if no improvement in auROC value was observed on the validation set over 20 consecutive epochs.
Optimizer
The Adam optimizer was applied.
L 2 norm penalty
The L2 penalty weight (λ) was set to 1 × 10−6.
Loss function
Binary cross-entropy loss was used. Positive sample weights of 1, 2, 3, 4, 5 and 10 were applied on the basis of different positive-to-negative sample ratios.
Gradient clipping
Gradients were clipped using a maximum norm of 5.0.
Hyperparameters in RNA structure encoder
The optimal kernel size and padding were determined to be 7 and 3, respectively, on the basis of a grid search over (kernel size, padding) pairs: (3,1), (5,2), (7,3), (9,4) and (11,5). The optimal number of channel size was found to be 16, selected from the set {2, 4, 6, 8, 16, 32, 64}. Dropout rates of 0.5 and 0.3 were applied after each residual block.
Hyperparameters in small-molecule structure encoder
The optimal attention heads were set to 3, selected from the set {1, 2, 3, 4, 5, 6}. The optimal number of GAT layers was set to 3, selected from the set {1, 2, 3, 4, 5, 6}. The optimal output head dimension was set to 256, selected from the set {32, 64, 128, 256, 512}.
Hyperparameters in MDF module
We tested various configurations of the MDF module and fully connected decoder and identified the optimal combination of parameters. The first coattention layer used two attention heads with a dropout rate of 0.1; the second self-attention layer used two attention heads with a dropout rate of 0.1; the third self-attention layer used eight attention heads with a dropout rate of 0.3. The fully connected decoder consisted of four layers with 1,024, 1,024, 1,024 and 512 nodes, respectively.
Evaluation of SMRTnet
We used accuracy, recall, precision, F0.5 score, auROC and auPRC (area under the precision–recall curve) to assess model performance. These metrics were calculated using the Python package scikit-learn (https://scikit-learn.org/stable/). Specifically, accuracy was calculated using the accuracy_score function from sklearn.metrics, which measures the proportion of correct predictions among all predictions. Precision, quantifying the percentage of predicted positive bindings that are true positives, was calculated using the precision_score function. Recall, indicating the proportion of actual positive bindings correctly identified, was computed using recall_score function. The F0.5 score is a variant of the F1 score where precision is weighted twice as much as recall. Similarly, auROC was calculated using the roc_auc_score function, providing an aggregate measure of the model’s ability to discriminate between positive and negative samples across all classification thresholds. auPRC, precision and recall values at various thresholds were obtained using the precision_recall_curve function and these values were subsequently used to calculate the auPRC.
Computational complexity and scalability of SMRTnet
SMRTnet contains a total of 208,191,155 model parameters. We assessed its computational complexity using an NVIDIA A800 (80 GB) GPU during both training and inference. Training the model using a fivefold CV process on the SMRTnet dataset, with a positive-to-negative sample ratio of 1:2, required approximately 48 h. GPU memory usage during training was about 14 GB with a batch size of 32. For inference, SMRTnet used approximately 4 GB of GPU memory with a batch size of 1. It took approximately 25 s to predict the binding score of a small molecule–RNA pair using the ensemble scoring strategy on a single GPU. This process could be substantially accelerated using the parallel ensemble scoring strategy, where the five models from fivefold CV were distributed across multiple GPUs.
Comparison of SMRTnet to existing computational methods
We evaluated SMRTnet and RNAmigos2 on each other’s test sets using their respective evaluation strategies and used the decoy evaluation task described in RNAmigos2 to benchmark SMRTnet against current field-leading computational tools. We used the RNAmigos2 model directly from its GitHub repository (https://github.com/cgoliver/rnamigos2/tree/3dab30ed6f5f63c328f32d2c6215ec14c572c2e2) without retraining.
Following the test set of SMRTnet and its evaluation strategy, we applied the SMRTnet model trained in each fold to predict binding scores for small molecule–RNA pairs in the corresponding test set and calculated the auROC values for each fold. For RNAmigos2, we generated RNAmigos2-compatible input pockets from PDB structures corresponding to RNAs in the test set of SMRTnet and predicted binding scores for each pocket–small molecule pair using RNAmigos2 to calculate the auROC values. In parallel, following the evaluation strategy of RNAmigos2 based on the test set of RNAmigos2, we calculated the auROC values for each binding pocket by comparing predicted binding scores between the pocket and both native and decoy molecules from different libraries. For SMRTnet, the binding score of a pocket–small molecule pair was computed by averaging the scores of all fragments within the pocket against the molecule. Decoys were obtained from three libraries: (1) the ChEMBL library, consisting of selected decoys from the ChEMBL database (n = 500); (2) the PDB library, which includes all ligands from the RNAmigos2 PDB dataset (n = 264); and (3) the ChEMBL + PDB library, a combination of the previous two libraries (n = 764). For RNAmigos2, we used the publicly available results provided by the authors on the Zenodo repository (https://doi.org/10.5281/zenodo.14803961)127 directly and visualized the results using Python (version 3.8.10), matplotlib (v.3.7.5) and seaborn (v.0.13.2) packages.
For decoy evaluation task, we generated a decoy molecule library for each small molecule in the SMRTnet dataset and evaluated the model’s performance on the basis of the ranking of the target molecule within its corresponding decoy library. Here, we used DecoyFinder to retrieve up to 30 decoys from the ZINC15 bioactive small-molecule library. These decoys were selected to be physically similar (for example, molecular weight, partition coefficient, hydrogen bonds donors, hydrogen bond acceptors and number of rotatable bonds) but chemically distinct (based on Tanimoto similarity) from the target molecule. Specifically, RNAmigos2 uses the RNA tertiary structure of the binding site and the corresponding target molecule with its decoy in the test set and ranks the target molecule within the decoy library on the basis of the predicted scores. RNAmigos predicts the ligand’s fingerprint on the basis of the RNA tertiary structure of the binding site and ranks the target molecule by computing the similarity between the predicted fingerprint and those of the target molecule and its decoys. AutoDock Vina, NLDock, RLDOCK and rDock treat the RNA tertiary structure of each target molecule in the test set as the receptor and the target molecule together with its decoys as the ligands and rank them on the basis of the scores generated by their respective scoring functions. SMRTnet calculates the binding scores between each RNA fragment and the target molecule or decoys using an ensemble scoring strategy. The final score for each molecule is obtained by averaging the binding scores across all RNA fragments. The target molecule is then ranked within its decoy library accordingly. Lastly, the ranking percentage is calculated by dividing the rank of the target molecule, determined by sorting the scores, by the total number of molecules.
For comparison, we retrained SMRTnet on a much smaller RNAmigos2 subset compatible with the SMRTnet training protocol by excluding nontherapeutically relevant small molecules and structures with fewer than 31 RNA residues. To prevent data leakage, we also built a data-leakage-excluded RNAmigos2 test set by excluding samples with identical multistrand binding sites and a maximum Tainmoto similarity threshold of 0.7 relative to the SMRTnet training data.
Binding-site identification of SMRTnet
We used Grad-CAM59 and the SmoothGrad algorithm128 to quantify the contribution of each nucleotide to the SRIs through backpropagation calculations. Specifically, Grad-CAM was used to generate saliency maps on the basis of gradient signals, while the SmoothGrad algorithm reduced noise and smoothed the saliency maps by averaging gradient signals across 20 perturbation rounds of the input. Higher gradient signals in the saliency maps indicate a greater importance of individual nucleotides in the binding process.
For an input SRI pair, the combined methods compute gradient matrices, where the RNA sequence encoder yields a 1 × 31 gradient matrix and the RNA structure encoder produces a 5 × 31 gradient matrix (later averaged to 1 × 31). Then, these are combined into a final 2 × 31 gradient matrix, where the first row represents RNA sequence-based gradient signals and the second row represents RNA structure-based gradient signals. In summary, given an input \(x\in {{\mathbb{R}}}^{L\times D}\), the gradient \(g\left(x\right)\) for each encoder is calculated as follows:
Here, \({\boldsymbol{g}}\) represents the Grad-CAM algorithm, \(\hat{{\boldsymbol{M}}}\) represents the SmoothGrad algorithm, \({\boldsymbol{n}}\) is equal to 20, representing 20 rounds of minuscule Gaussian noise (\({\boldsymbol{N}}\left({\boldsymbol{0}},{{\boldsymbol{\sigma }}}^{{\boldsymbol{2}}}\right)\)) and \(\odot\) represents the operation of multiplication. To identify potential binding sites, we applied the Savitzky–Golay filter129 to smooth the discrete gradient signal and normalized the signals to the range of 0 to 1 using min–max normalization, thereby highlighting HARs in saliency maps.
Binding-site evaluation of SMRTnet
We quantitatively evaluated SMRTnet’s ability to identify RNA-binding sites by calculating the auROC value and the Pearson correlation coefficient between the gradient signals (from both RNA sequence and RNA structure) and the experimentally determined binding sites. For the auROC value calculation, nucleotides located within 10 Å of the ligand were label as 1 (positive) and all other nucleotides were labeled as 0 (negative)23; we computed the auROC value by comparing the gradient signals to these binary labels. For the Pearson correlation coefficient calculation, we measured the correlation between the gradient signals and the proximity to experimentally determined binding sites, where proximity was defined as 1 − normalized minimum distance from each nucleotide to any determined binding-site nucleotide. These evaluations were applied to five case studies, the test set of SMRTnet and two existing benchmarks (TE18 and RB19)57. For TE18, six entries were excluded: five entries involved magnesium ions (PDB 2MIS, 364D, 430D and 4PQV) or cobalt (II) ions (PDB 379D), commonly used to stabilize RNA structures, and one entry lacked a small molecule in the current PDB release (PDB 6EZ0).
Ablation study of SMRTnet
We evaluated the contribution of each encoder by setting the output embeddings of one or more encoders to zeros, thereby isolating their respective contributions to the model’s performance. Specifically, the output embedding of the following encoders was set to zero: (1) the RNA sequence encoder and small-molecule SMILES encoder; (2) the RNA structure encoder and small-molecule structure encoder; (3) the RNA sequence encoder; (4) the RNA structure encoder; (5) the small-molecule structure encoder; and (6) the small-molecule SMILES encoder. Additionally, we directly replaced the MDF module with a concatenation module to evaluate contributions of the MDF module (7). This approach enabled us to evaluate the impact of removing each component on the overall model performance.
MST assay
MST assays were performed using the Monolith NT.115 system (NanoTemper Technologies) with standard Monolith capillaries (NanoTemper Technologies, MO-K022). Cy5-labeled RNAs (10 nM), purchased from RuiBiotec (China), were folded in 1× assay buffer (8 mM Na2HPO4, 190 mM NaCl, 1 mM EDTA and 0.05%(v/v) Tween-20) by heating to 70 °C for 5 min, followed by slow cooling to room temperature at 0.1 °C s−1, as the RNA sample.
For the ‘binding check’ mode, small molecules were added into the RNA sample at the final concentration of 50 μM and incubated at room temperature for 30 min, as the small molecule–RNA complex sample. The red laser was selected on the basis of the Cy5 label of RNAs. The RNA sample and the small molecule–RNA complex sample were loaded into standard Monolith capillaries in four replicates and measured at room temperature by using 20% excitation power and medium MST power.
For the ‘binding affinity’ mode, small molecules were added into the RNA sample at the final concentration of 100 μM and then serially diluted twofold (15 times) with RNA sample. After the incubation at room temperature for 30 min, the 16 samples were loaded into standard Monolith capillaries and measured at room temperature using 20% excitation power and medium MST power.
For fluorescence intensity change after adding the small molecule into the RNA sample, SD-test was used to distinguish between small molecules as a function of changing fluorescence because of nonspecific effects or interaction with the RNA. The samples were mixed 1:1 with SD-mix (4% SDS, 40 mM DTT) and incubated at 95°C for 5 min, followed by detection of fluorescence intensity using 20% excitation power and medium MST power.
Data from the ‘binding check’ mode and SD-test were analyzed using MO.Control software (v.2.3) and data from the ‘binding affinity’ mode were analyzed using MO.Affinity Analysis software (v.2.3).
Cell culture
HEK293T and HeLa cells were purchased from Cell Bank, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences. Raji and Jurkat cells were purchased from Procell Life Science and Technology. HEK293T and HeLa cells were cultured in DMEM (Gibco) supplemented with 10% FBS (PAN-Biotech) and penicillin–streptomycin (Gibco) at 37 °C in 5% CO2. Raji and Jurkat cells were cultured in RPMI 1640 medium (Gibco) supplemented with 10% FBS (PAN-Biotech) and penicillin–streptomycin (Gibco) at 37 °C in 5% CO2. Mycoplasma contamination in all cultures was routinely checked using a Mycoplasma detection kit (Vazyme).
qPCR analysis
HeLa cells were seeded in six-well plates (200,000 cells per well) and treated with 0.1% (v/v) DMSO or candidate compounds (10 μM) for 48 h. Total RNA was extracted using the HiPure total RNA kit (Vazyme Biotech). RNA concentration and purity were assessed using a NanoDrop 2000 (Thermo Fisher Scientific), with an A260:A280 ratio between 1.8 and 2.0. For complementary DNA synthesis, 100 ng of RNA was reverse-transcribed using HiScript II Q RT SuperMix for qPCR (+gDNA wiper) (Vazyme, R223-01). qPCR was performed using ChamQ Universal SYBR qPCR master mix (Vazyme, Q711-02) on a QuantStudio 3 Flex real-time PCR system (Thermo Fisher Scientific), following the manufacturer’s instructions. Relative mRNA levels were calculated using the \(\varDelta \varDelta {C}_{t}\) method. qPCR primers and processed results are provided in Supplementary Table 7.
Western blotting analysis
HeLa cells were seeded in six-well plates (200,000 cells per well) and treated with 0.1% (v/v) DMSO or candidate compounds (10 μM) for 48 h. Total protein was extracted with RIPA buffer (100 μl per 200,000 cells). Approximately 20 μg of protein was separated by 12.5% SDS–PAGE, transferred to a PVDF membrane and blocked with 1× TBST containing 5% skim milk for 2 h. The membrane was incubated overnight at 4 °C with primary anti-MYC antibody (ABclonal, A19032, lot 3523042615; 1:1,500), followed by incubation at room temperature for 1.5 h with anti-rabbit IgG horseradish peroxidase secondary antibody (EASYBIO, BE0101, lot 80861011; 1:10,000). Alternatively, for GAPDH detection, the membrane was incubated overnight at 4 °C with 1× TBST containing 5% skim milk and then incubated at room temperature for 1.5 h with anti-GAPDH antibody (EASYBIO, BE0034, lot 80790311; 1:5,000). The membranes were detected using the enhanced chemiluminescence kit (New Cell & Molecular Biotech) and quantified using Fiji software. Processed statistical results are provided in Supplementary Table 8.
Cell proliferation assay
Cells were seeded in 96-well plates (2,000 cells per well) and treated with 0.1% (v/v) DMSO or candidate compounds (10 μM) for 48 h. For HeLa cells, cell proliferation was assessed using the MTT cell proliferation and cytotoxicity assay kit (Solarbio, M1020), following the manufacturer’s protocol. Briefly, after 48 h of incubation, the medium was replaced with 90 μl of fresh medium and 10 μl of MTT solution. The cells were incubated at 37 °C in 5% CO2 for 4 h. Subsequently, the solution was removed, 100 µl of DMSO was added and the plates were shaken at room temperature for 30 min. Absorbance at 490 nm was measured using a Spark microplate reader (Tecan). For Raji and Jurkat cells, cell proliferation was assessed using the cell viability/toxicity assay kit (CCK-8 kit) (FeiMoBio, FB29236-500), following the manufacturer’s protocol. Briefly, after 48 h of incubation, 10 μl of CCK-8 solution was added. The cells were incubated at 37 °C in 5% CO2 for 1 h. Absorbance at 450 nm was measured using a Spark microplate reader (Tecan). The relative cell proliferation was normalized to the absorbance of cells treated with 0.1% (v/v) DMSO. Processed statistical results are provided in Supplementary Table 9.
Cell apoptosis assay
Cells were seeded in 96-well plates (2,000 cells per well) and treated with 0.1% (v/v) DMSO or candidate compounds (10 μM) for 48 h. Cell apoptosis was assessed using the Caspase-Glo 3/7 assay system (Promega, G8091), following the manufacturer’s protocol. Briefly, after 48 h of incubation, 100 μl of Caspase-Glo 3/7 reagent was added. The cells were incubated at room temperature for 30 min. Luminescence was measured using a Spark microplate reader (Tecan). The relative cell apoptosis was normalized to the luminescence of cells treated with 0.1% (v/v) DMSO. Processed statistical results are provided in Supplementary Table 10.
MYC IRES luciferase reporter assay
The MYC IRES luciferase plasmid incorporates the MYC IRES sequence upstream of the Renilla luciferase gene in the psiCHECK-2 vector. HEK293T cells (approximately 80% confluent) were seeded in 60-mm dishes and transfected with 2 μg of the MYC IRES luciferase plasmid using jetPRIME (Polyplus, 101000027), following the manufacturer’s protocol. Cells were trypsinized and seeded into 96-well plates (2,000 cells per well) and treated with either 0.1% (v/v) DMSO or candidate compounds (10 μM) for 48 h. Renilla luciferase and firefly luciferase expression were measured using the Dual-Glo luciferase assay system (Promega, E2920), following the manufacturer’s instructions, and Renilla luciferase expression was normalized to firefly luciferase activity. Processed results are provided in Supplementary Table 11.
Quantification and statistical analysis
Statistical analysis was conducted using GraphPad Prism 8 and Python (v.3.8.10), while visualizations, including bar plots, scatter plots, violin plots, radar plots and heat maps were generated with Python (v.3.8.10), matplotlib (v.3.7.5) and seaborn (v.0.13.2) packages. Statistical details are provided in the figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All processed data used in this study are available from GitHub (https://github.com/Yuhan-Fei/SMRTnet/tree/main/data) and in Supplementary Tables 1–6. The raw data for the SMRTnet dataset were obtained from the PDB and can be downloaded from Zenodo (https://doi.org/10.5281/zenodo.14986116)130. Source data are provided with this paper.
Code availability
All source code used in this study is publicly available from GitHub (https://github.com/Yuhan-Fei/SMRTnet) and all pretrained models can be downloaded from Zenodo (https://doi.org/10.5281/zenodo.14715564)131. SMRTnet can be installed through PyPI (https://pypi.org/project/smrtnet/ or https://pypi.org/project/smrtnet-latest/). Additionally, an installation-free online service is available through Google Colab (https://drive.google.com/drive/folders/1HQo3o2saY5U9vPqebz4ZdpCVVQXqw0q_?usp=sharing).
Change history
07 January 2026
In the version of the article initially published, the text “This work has also been supported by the New Cornerstone Science Foundation through the XPLORER PRIZE” was missing from the Acknowledgements section and has now been added to the HTML and PDF versions of the article.
References
Zheng, S. et al. Structural basis for stabilization of the tau pre-mRNA splicing regulatory element by novantrone (mitoxantrone). Chem. Biol. 16, 557–566 (2009).
Fedorova, O. et al. Small molecules that target group II introns are potent antifungal agents. Nat. Chem. Biol. 14, 1073–1078 (2018).
Zhang, P. et al. Translation of the intrinsically disordered protein α-synuclein is inhibited by a small molecule targeting its structured mRNA. Proc. Natl Acad. Sci. USA 117, 1457–1467 (2020).
Tong, Y. et al. Decreasing the intrinsically disordered protein α-synuclein levels by targeting its structured mRNA with a ribonuclease-targeting chimera. Proc. Natl Acad. Sci. USA 121, e2306682120 (2024).
Childs-Disney, J. L. et al. Rationally designed small molecules targeting the RNA that causes myotonic dystrophy type 1 are potently bioactive. ACS Chem. Biol. 7, 856–862 (2012).
Kumar, A. et al. Chemical correction of pre-mRNA splicing defects associated with sequestration of muscleblind-like 1 protein by expanded r(CAG)-containing transcripts. ACS Chem. Biol. 7, 496–505 (2012).
Suresh, B. M. et al. A general fragment-based approach to identify and optimize bioactive ligands targeting RNA. Proc. Natl Acad. Sci. USA 117, 33197–33203 (2020).
Becquart, C. et al. Exploring heterocycle-spermine conjugates as modulators of oncogenic microRNAs biogenesis. ACS Omega 3, 16500–16508 (2018).
Haniff, H. S. et al. Targeting the SARS-CoV-2 RNA genome with small molecule binders and ribonuclease targeting chimera (RiboTAC) degraders. ACS Cent. Sci. 6, 1713–1721 (2020).
Palacino, J. et al. SMN2 splice modulators enhance U1-pre-mRNA association and rescue SMA mice. Nat. Chem. Biol. 11, 511–517 (2015).
Ratni, H. et al. Discovery of risdiplam, a selective survival of motor neuron-2 (SMN2) gene splicing modifier for the treatment of spinal muscular atrophy (SMA). J. Med. Chem. 61, 6501–6517 (2018).
Aguilar, R. et al. Targeting Xist with compounds that disrupt RNA structure and X inactivation. Nature 604, 160–166 (2022).
Yazdani, K. et al. Machine learning informs RNA-binding chemical space. Angew. Chem. Int. Ed. Engl. 62, e202211358 (2023).
Lipinski, C. & Hopkins, A. Navigating chemical space for biology and medicine. Nature 432, 855–861 (2004).
Cherkasov, A. The ‘Big Bang’ of the chemical universe. Nat. Chem. Biol. 19, 667–668 (2023).
Manigrasso, J., Marcia, M. & De Vivo, M. Computer-aided design of RNA-targeted small molecules: a growing need in drug discovery. Chem 7, 2965–2988 (2021).
Fedorova, O. et al. In silico discovery of group II Intron RNA splicing inhibitors. ACS Chem. Biol. 18, 1968–1975 (2023).
Eberhardt, J. et al. AutoDock Vina 1.2.0: new docking methods, expanded force field, and Python bindings. J. Chem. Inf. Model. 61, 3891–3898 (2021).
Sun, L. Z. et al. RLDOCK: a new method for predicting RNA–ligand interactions. J. Chem. Theory Comput. 16, 7173–7183 (2020).
Feng, Y. et al. NLDock: a fast nucleic acid–ligand docking algorithm for modeling RNA/DNA–ligand complexes. J. Chem. Inf. Model. 61, 4771–4782 (2021).
Ruiz-Carmona, S. et al. rDock: a fast, versatile and open source program for docking ligands to proteins and nucleic acids. PLoS Comput Biol. 10, e1003571 (2014).
Zhou, Y., Jiang, Y. & Chen, S. J. RNA-ligand molecular docking: advances and challenges. Wiley Interdiscip. Rev. Comput. Mol. Sci. 12, e1571 (2022).
Oliver, C. et al. Augmented base pairing networks encode RNA–small molecule binding preferences. Nucleic Acids Res. 48, 7690–7699 (2020).
Carvajal-Patino, J. G. et al. RNAmigos2: accelerated structure-based RNA virtual screening with deep graph learning. Nat. Commun. 16, 2799–2811 (2025).
Sun, S. & Gao, L. Contrastive pre-training and 3D convolution neural network for RNA and small molecule binding affinity prediction. Bioinformatics 40, btae155 (2024).
Jerabek-Willemsen, M. et al. Microscale thermophoresis: interaction analysis and beyond. J. Mol. Struct. 1077, 101–113 (2014).
Stoneley, M. et al. Analysis of the c-myc IRES; a potential role for cell-type specific trans-acting factors and the nuclear compartment. Nucleic Acids Res. 28, 687–694 (2000).
Ross, J. et al. Large-scale chemical language representations capture molecular structure and properties. Nat. Mach. Intell. 4, 1256–1264 (2022).
Lu, J. et al. Vilbert: pretraining task-agnostic visiolinguistic representations for vision-and-language tasks. In Proc. 33rd International Conference on Neural Information Processing Systems 13–23 (Curran Associates, 2019).
Vaswani, A. et al. Attention is all you need. In Proc. 31st International Conference on Neural Information Processing Systems 6000–6010 (Curran Associates, 2017).
Sun, L. et al. Predicting dynamic cellular protein–RNA interactions by deep learning using in vivo RNA structures. Cell Res. 31, 495–516 (2021).
Playe, B. & Stoven, V. Evaluation of deep and shallow learning methods in chemogenomics for the prediction of drugs specificity. J. Cheminform. 12, 11–29 (2020).
Zabihian, A. et al. DEDTI versus IEDTI: efficient and predictive models of drug–target interactions. Sci. Rep. 13, 9238–9256 (2023).
Chen, H. et al. Drug target prediction through deep learning functional representation of gene signatures. Nat. Commun. 15, 1853–1868 (2024).
Zhao, Q. et al. HyperAttentionDTI: improving drug–protein interaction prediction by sequence-based deep learning with attention mechanism. Bioinformatics 38, 655–662 (2022).
He, C. et al. Exploring drug–target interaction prediction on cold-start scenarios via meta-learning-based graph transformer. Methods 234, 10–20 (2025).
Arevalo, J. et al. MOTIVE: a drug–target interaction graph for inductive link prediction. In Proc. 38th International Conference on Neural Information Processing System 140320–140333 (Curran Associates, 2024).
Yang, Z. et al. ML-DTI: mutual learning mechanism for interpretable drug–target interaction prediction. J. Phys. Chem. Lett. 12, 4247–4261 (2021).
Ye, Q. et al. A unified drug–target interaction prediction framework based on knowledge graph and recommendation system. Nat. Commun. 12, 6775–6787 (2021).
Lu, Z. et al. DTIAM: a unified framework for predicting drug–target interactions, binding affinities and drug mechanisms. Nat. Commun. 16, 2548–2565 (2025).
Singh, J. et al. RNA secondary structure prediction using an ensemble of two-dimensional deep neural networks and transfer learning. Nat. Commun. 10, 5407–5420 (2019).
Soufan, O. et al. DRABAL: novel method to mine large high-throughput screening assays using Bayesian active learning. J. Cheminform. 8, 64–78 (2016).
Carrell, D. S. et al. Improving methods of identifying anaphylaxis for medical product safety surveillance using natural language processing and machine learning. Am. J. Epidemiol. 192, 283–295 (2023).
Rifaioglu, A. S. et al. Recent applications of deep learning and machine intelligence on in silico drug discovery: methods, tools and databases. Brief. Bioinform. 20, 1878–1912 (2019).
Morgan, B. S. et al. R-BIND: an interactive database for exploring and developing RNA-targeted chemical probes. ACS Chem. Biol. 14, 2691–2700 (2019).
Donlic, A. et al. R-BIND 2.0: an updated database of bioactive RNA-targeting small molecules and associated RNA secondary structures. ACS Chem. Biol. 17, 1556–1566 (2022).
Krishnan, S. R., Roy, A. & Gromiha, M. M. R-SIM: a database of binding affinities for RNA–small molecule interactions. J. Mol. Biol. 435, 167914–167922 (2023).
Mehta, A. et al. SMMRNA: a database of small molecule modulators of RNA. Nucleic Acids Res. 42, D132–D141 (2014).
Kumar Mishra, S. & Kumar, A. NALDB: nucleic acid ligand database for small molecules targeting nucleic acid. Database 2016, baw002 (2016).
Deng, Z. et al. Predicting ligand–RNA binding using E3-equivariant network and pretraining. In Proc. Machine Learning for Structural Biology Workshop (NeurIPS, 2022).
Cereto-Massague, A. et al. DecoyFinder: an easy-to-use Python GUI application for building target-specific decoy sets. Bioinformatics 28, 1661–1662 (2012).
Sterling, T. & Irwin, J. J. ZINC 15—ligand discovery for everyone. J. Chem. Inf. Model. 55, 2324–2337 (2015).
Ding, J. et al. Vina-GPU 2.0: further accelerating AutoDock Vina and its derivatives with graphics processing units. J. Chem. Inf. Model. 63, 1982–1998 (2023).
Reuter, J. S. & Mathews, D. H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010).
Shen, T. et al. Accurate RNA 3D structure prediction using a language model-based deep learning approach. Nat. Methods 21, 2287–2298 (2024).
Wang, N. et al. Multi-purpose RNA language modelling with motif-aware pretraining and type-guided fine-tuning. Nat. Mach. Intell. 6, 548–557 (2024).
Su, H., Peng, Z. & Yang, J. Recognition of small molecule-RNA binding sites using RNA sequence and structure. Bioinformatics 37, 36–42 (2021).
Gao, J. et al. Predicting small molecule binding nucleotides in RNA structures using RNA surface topography. J. Chem. Inf. Model. 64, 6979–6992 (2024).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. Int. J. Comput. Vis. 128, 336–359 (2020).
Tong, Y. et al. Programming inactive RNA-binding small molecules into bioactive degraders. Nature 618, 169–179 (2023).
Li, X. D., Liu, L. & Cheng, L. Identification of thienopyridine carboxamides as selective binders of HIV-1 trans activation response (TAR) and Rev response element (RRE) RNAs. Org. Biomol. Chem. 16, 9191–9196 (2018).
Hoskins, J. W. et al. Lomofungin and dilomofungin: inhibitors of MBNL1–CUG RNA binding with distinct cellular effects. Nucleic Acids Res. 42, 6591–6602 (2014).
Liu, X. et al. Targeted degradation of the oncogenic microRNA 17-92 cluster by structure-targeting ligands. J. Am. Chem. Soc. 142, 6970–6982 (2020).
Krajewski, S. S., Isoz, I. & Johansson, J. Antibacterial and antivirulence effect of 6-N-hydroxylaminopurine in Listeria monocytogenes. Nucleic Acids Res. 45, 1914–1924 (2017).
Somarowthu, S. et al. HOTAIR forms an intricate and modular secondary structure. Mol. Cell 58, 353–361 (2015).
Prado, S. et al. A small-molecule inhibitor of HIV-1 Rev function detected by a diversity screen based on RRE–Rev interference. Biochem. Pharm. 156, 68–77 (2018).
Khan, E. et al. Discovery of a potent small molecule inhibiting Huntington’s disease (HD) pathogenesis via targeting CAG repeats RNA and poly Q protein. Sci. Rep. 9, 16872–16887 (2019).
Sun, L. et al. In vivo structural characterization of the SARS-CoV-2 RNA genome identifies host proteins vulnerable to repurposed drugs. Cell 184, 1865–1883 (2021).
Li, L. et al. Structural lability in stem-loop 1 drives a 5′ UTR–3′ UTR interaction in coronavirus replication. J. Mol. Biol. 377, 790–803 (2008).
Vora, S. M. et al. Targeting stem-loop 1 of the SARS-CoV-2 5′ UTR to suppress viral translation and Nsp1 evasion. Proc. Natl Acad. Sci. USA 119, e2117198119 (2022).
Liu, P. et al. A U-turn motif-containing stem-loop in the coronavirus 5′ untranslated region plays a functional role in replication. RNA 13, 763–780 (2007).
Yang, D. & Leibowitz, J. L. The structure and functions of coronavirus genomic 3′ and 5′ ends. Virus Res. 206, 120–133 (2015).
Yang, D. et al. Mouse hepatitis virus stem-loop 4 functions as a spacer element required to drive subgenomic RNA synthesis. J. Virol. 85, 9199–9209 (2011).
Rangan, R. et al. RNA genome conservation and secondary structure in SARS-CoV-2 and SARS-related viruses: a first look. RNA 26, 937–959 (2020).
Kretsch, R. C. et al. Tertiary folds of the SL5 RNA from the 5′ proximal region of SARS-CoV-2 and related coronaviruses. Proc. Natl Acad. Sci. USA 121, e2320493121 (2024).
Chen, H., Liu, H. & Qing, G. Targeting oncogenic Myc as a strategy for cancer treatment. Signal. Transduct. Target. Ther. 3, 5–12 (2018).
Fu, L. et al. ADMETlab 3.0: an updated comprehensive online ADMET prediction platform enhanced with broader coverage, improved performance, API functionality and decision support. Nucleic Acids Res. 52, W422–W431 (2024).
Ruiz-Ruiz, M. C. et al. Activation-induced apoptosis in Jurkat cells through a Myc-independent mechanism. Mol. Immunol. 32, 947–955 (1995).
Molina, E. et al. MYC directly transactivates CR2/CD21, the receptor of the Epstein–Barr virus, enhancing the viral infection of Burkitt lymphoma cells. Oncogene 42, 3358–3370 (2023).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Luo, Q. J. et al. RNA structure probing reveals the structural basis of Dicer binding and cleavage. Nat. Commun. 12, 3397–3409 (2021).
Childs-Disney, J. L. et al. Targeting RNA structures with small molecules. Nat. Rev. Drug Discov. 21, 736–762 (2022).
Kovachka, S. et al. Small molecule approaches to targeting RNA. Nat. Rev. Chem. 8, 120–135 (2024).
Scott, D. E. et al. Small molecules, big targets: drug discovery faces the protein–protein interaction challenge. Nat. Rev. Drug Discov. 15, 533–550 (2016).
Xie, X. et al. Recent advances in targeting the ‘undruggable’ proteins: from drug discovery to clinical trials. Signal. Transduct. Target. Ther. 8, 335–406 (2023).
Du, X. et al. Insights into protein–ligand interactions: mechanisms, models, and methods. Int. J. Mol. Sci. 17, 144–178 (2016).
Boniecki, M. J. et al. SimRNA: a coarse-grained method for RNA folding simulations and 3D structure prediction. Nucleic Acids Res. 44, e63 (2016).
Meng, E. C. et al. UCSF ChimeraX: tools for structure building and analysis. Protein Sci. 32, e4792 (2023).
Ireland, S. M. & Martin, A. C. R. atomium—a Python structure parser. Bioinformatics 36, 2750–2754 (2020).
Lu, X. J., Bussemaker, H. J. & Olson, W. K. DSSR: an integrated software tool for dissecting the spatial structure of RNA. Nucleic Acids Res. 43, e142 (2015).
Landrum, G. RDKit: open-source cheminformatics software. rdkit.org https://www.rdkit.org (2016).
O’Boyle, N. M. et al. Open Babel: an open chemical toolbox. J. Cheminform. 3, 33–47 (2011).
Yang, W. Y. et al. Inhibition of non-ATG translational events in cells via covalent small molecules targeting RNA. J. Am. Chem. Soc. 137, 5336–5345 (2015).
Moon, M. H. et al. Measuring RNA–ligand interactions with microscale thermophoresis. Biochemistry 57, 4638–4643 (2018).
Velagapudi, S. P., Li, Y. & Disney, M. D. A cross-linking approach to map small molecule–RNA binding sites in cells. Bioorg. Med Chem. Lett. 29, 1532–1536 (2019).
Wang, Z. F. et al. The hairpin form of r(G4C2)exp in c9ALS/FTD is repeat-associated non-ATG translated and a target for bioactive small molecules. Cell Chem. Biol. 26, 179–190 (2019).
Ursu, A. et al. Structural features of small molecules targeting the RNA repeat expansion that causes genetically defined ALS/FTD. ACS Chem. Biol. 15, 3112–3123 (2020).
Angelbello, A. J. et al. A small molecule that binds an RNA repeat expansion stimulates its decay via the exosome complex. Cell Chem. Biol. 28, 34–45 (2021).
Bush, J. A. et al. A blood–brain penetrant RNA-targeted small molecule triggers elimination of r(G4C2)exp in c9ALS/FTD via the nuclear RNA exosome. Proc. Natl Acad. Sci. USA 119, e2210532119 (2022).
Menichelli, E. et al. Discovery of small molecules that target a tertiary-structured RNA. Proc. Natl Acad. Sci. USA 119, e2213117119 (2022).
Tong, Y. et al. Transcriptome-wide mapping of small-molecule RNA-binding sites in cells informs an isoform-specific degrader of QSOX1 mRNA. J. Am. Chem. Soc. 144, 11620–11625 (2022).
Fukunaga, K. et al. Small-molecule aptamer for regulating RNA functions in mammalian cells and animals. J. Am. Chem. Soc. 145, 7820–7828 (2023).
Abulwerdi, F. A. et al. Development of small molecules with a noncanonical binding mode to HIV-1 trans activation response (TAR) RNA. J. Med Chem. 59, 11148–11160 (2016).
Bandyopadhyay, S. et al. Novel 5′ untranslated region directed blockers of iron-regulatory protein-1 dependent amyloid precursor protein translation: implications for down syndrome and Alzheimer’s disease. PLoS ONE 8, e65978 (2013).
Blount, K. F. & Breaker, R. R. Riboswitches as antibacterial drug targets. Nat. Biotechnol. 24, 1558–1564 (2006).
Connelly, C. M. et al. Synthetic ligands for PreQ1 riboswitches provide structural and mechanistic insights into targeting RNA tertiary structure. Nat. Commun. 10, 1501–1513 (2019).
Neupane, K. et al. Anti-frameshifting ligand active against SARS coronavirus-2 is resistant to natural mutations of the frameshift-stimulatory pseudoknot. J. Mol. Biol. 432, 5843–5847 (2020).
Patwardhan, N. N. et al. Amiloride as a new RNA-binding scaffold with activity against HIV-1 TAR. Medchemcomm 8, 1022–1036 (2017).
Shortridge, M. D. et al. Drug-like small molecules that inhibit expression of the oncogenic microRNA-21. ACS Chem. Biol. 18, 237–250 (2023).
Tibodeau, J. D. et al. The up-regulation of ferritin expression using a small-molecule ligand to the native mRNA. Proc. Natl Acad. Sci. USA 103, 253–257 (2006).
Vo, D. D. et al. Building of neomycin–nucleobase–amino acid conjugates for the inhibition of oncogenic miRNAs biogenesis. Org. Biomol. Chem. 16, 6262–6274 (2018).
Zhou, Z. D. & Tan, E. K. Iron regulatory protein (IRP)–iron responsive element (IRE) signaling pathway in human neurodegenerative diseases. Mol. Neurodegener. 12, 75–87 (2017).
de Mezer, M. et al. Mutant CAG repeats of Huntingtin transcript fold into hairpins, form nuclear foci and are targets for RNA interference. Nucleic Acids Res. 39, 3852–3863 (2011).
Schneeberger, E. M. et al. Native mass spectrometry reveals the initial binding events of HIV-1 rev to RRE stem II RNA. Nat. Commun. 11, 5750–5760 (2020).
Veličković, P. et al. Graph attention networks. In Proc. International Conference on Learning Representations (ICLR, 2018).
Yuan, D. et al. The European Nucleotide Archive in 2023. Nucleic Acids Res. 52, D92–D97 (2024).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 50, D20–D26 (2022).
Sayers, E. W. et al. GenBank 2024 update. Nucleic Acids Res. 52, D134–D137 (2024).
Harrison, P. W. et al. Ensembl 2024. Nucleic Acids Res. 52, D891–D899 (2024).
The RNAcentral Consortium RNAcentral: a hub of information for non-coding RNA sequences. Nucleic Acids Res. 47, D1250–D1251 (2019).
Feng, J. et al. CSCD2: an integrated interactional database of cancer-specific circular RNAs. Nucleic Acids Res. 50, D1179–D1183 (2022).
Di Marsico, M. et al. GreeNC 2.0: a comprehensive database of plant long non-coding RNAs. Nucleic Acids Res. 50, D1442–D1447 (2022).
Steinegger, M. & Soding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Kim, S. et al. PubChem substance and compound databases. Nucleic Acids Res. 44, D1202–D1213 (2016).
Irwin, J. J. et al. ZINC20—a free ultralarge-scale chemical database for ligand discovery. J. Chem. Inf. Model 60, 6065–6073 (2020).
Li, M. et al. DGL-LifeSci: an open-source toolkit for deep learning on graphs in life science. ACS Omega 6, 27233–27238 (2021).
Oliver, C. RNAmigos2: fast and accurate structure-based RNA virtual screening with semi-supervised graph learning and large-scale docking data. Zenodo https://doi.org/10.5281/zenodo.14803961 (2025)
Smilkov, D. et al. SmoothGrad: removing noise by adding noise. In Proc. Visualization for Deep Learning Workshop (ICML, 2017).
Luo, J., Ying, K. & Bai, J. Savitzky–Golay smoothing and differentiation filter for even number data. Signal Process. 85, 1429–1434 (2005).
Fei, Y. Predicting small molecule–RNA interactions without RNA tertiary structures [dataset]. Zenodo https://doi.org/10.5281/zenodo.14986116 (2025).
Fei, Y. Predicting small molecule–RNA interactions without RNA tertiary structures [model]. Zenodo https://doi.org/10.5281/zenodo.14715564 (2025).
Acknowledgements
We thank J.Z. (School of Life Sciences, Tsinghua University) for assistance with cell-based assays. We thank K.X. (State Key Laboratory of Membrane Biology, Beijing Tsinghua Institute for Frontier Interdisciplinary Innovation) for assistance with model testing. We thank J.W. (College of Future Technology, Peking University) for assistance with reproducing MST assays. We thank X.W. (Xungene Therapeutics) and Y.S. (Xungene Therapeutics) for their assistance in experimental design. We thank P.L. (Technology Center for Protein Sciences, Tsinghua University) and B.L. (NanoTemper Technologies) for technical assistance with MST experiments. We thank P.L. (Technology Center for Protein Sciences, Tsinghua University), Y.C. and B.Z. (Core Facility for Protein Research, Institute of Biophysics) for providing facility support for MST experiments. We thank J.R. and T.Z. (Center of Pharmaceutical Technology, Tsinghua University) for assisting with providing compounds from the natural product library. We thank G.X. (Academy for Advanced Interdisciplinary Studies, Peking University), Z.Z. (School of Life Sciences, Tsinghua University) and Z.L. (School of Life Sciences, Tsinghua University) for assistance with testing the installation and execution of SMRTnet. This work was supported by the National Key Research and Development Project of China (2022YFF1203100 to Q.C.Z. and 2021YFA1100200 to Y.W.), the National Natural Science Foundation of China (32230018 and 32125007 to Q.C.Z., 32100504 to Y.F., and 32025007 and 32130017 to Y.W.), the Beijing Natural Science Foundation (M23011 to Y.F.), the Postdoctoral Foundation of Tsinghua-Peking Center for Life Sciences (to Y.F.), the Beijing Advanced Innovation Center for Structural Biology, the Tsinghua-Peking Center for Life Sciences and the Beijing Advanced Center of RNA Biology. This work has also been supported by the New Cornerstone Science Foundation through the XPLORER PRIZE.
Author information
Authors and Affiliations
Contributions
Q.C.Z. and Y.W. supervised the project. Y.F. designed and developed the deep learning models. Y.F. and J.Z. analyzed the results. Y.F., Z.C. and J.Z. conducted the model benchmarking. J.M. and Y.F built the RNA language model. Y.F and J.Z maintained the GitHub repository. P.W., Y.F. and J.Z. performed the MST assays. P.W., J.Z. and X.S. performed the cell-based assays. Y.F., Y.W. and Q.C.Z. wrote the paper with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
Q.C.Z., Y.F. and J.Z. have filed a patent application (CN202510263239.6) related to this work. Q.C.Z and Y.W. are the co-founders of Xungene Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Carlos Oliver and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Architecture of SMRTnet components.
a. The RNA sequence encoder. This encoder accepts RNA sequences as input and generates corresponding embeddings as output. It was pre-trained on RNA sequences from various databases using the transformer model, and fine-tuned with the SMRTnet dataset. b. The RNA structure encoder. This encoder accepts RNA sequences and their secondary structure as input and generates corresponding embeddings as output. It applies one-hot encoding to RNA sequences and labeling encoding to RNA structures. The combined encodings are then processed by convolutional neural networks with residual neural networks. c. The small-molecule structure encoder. The encoder accepts the chemical structure of small molecules as input, including both structural information and atomic types. The input is then processed by graph attention networks. The output of the encoder is an embedding representing the structure of the small molecule, including its atomic and structural characteristics. d. The multimodal data fusion module. The module takes inputs from RNA and drug encoders. This module comprises three layers, including a co-attention layer and two self-attention layers with different parameters, which progressively integrate pairwise binding information from different encoders.
Extended Data Fig. 2 Dataset construction and model evaluation.
a. Bar plot shows the number of different structures in the PDB. b. Flowchart of the SMRTnet dataset construction. The overall data process comprises four main steps, as shown in subpanels i-iv: (i) Structures of SRIs were retrieved from the PDB; (ii) 1,061 high-quality paired SRIs were filtered; (iii) RNA binding positions within a 10 Å radius of the small molecule were selected and extended; (iv) RNA structures were converted into 31-nt fragments with RNA secondary structures. c. Ensemble scoring strategy of SMRTnet. The overall of this strategy comprises four main steps, as depicted in subpanels i-iv: (i) RNA input longer than 31-nt were processed into multiple fragments using a sliding window; (ii) Each fragment was predicted using five models from fivefold CV; (iii) The median of predictions based on five models was taken as the final binding score for each fragment; (iv) The final binding score and binding region are determined based on the input RNA length. d. Precision and recall curves, along with the corresponding F0.5 score curve of SMRTnet based on fivefold CV. The red dot indicates maximum F0.5 score, and the black dashed line marks the threshold corresponding to the maximum F0.5 score. e. Performance of SMRTnet and RNAmigos2 based on the SMRTnet dataset under different positive-to-negative sample ratios. Data are presented as mean ± s.d. based on fivefold CV (n = 5). f. Performance of SMRTnet on the SMRTnet dataset under different positive-to-negative sample ratios, and with different negative samples generated by varying random seeds. Data are presented as mean ± s.d. based on fivefold CV (n = 5). g. Performance comparison between SMRTnet and RNAmigos2 on the data leakage excluded test set of RNAmigos2 using decoys from different libraries, including ChEMBL (n = 500), PDB (n = 264), and ChEMBL+PDB (n = 764). Data are represented as box and violin plots with underlaid dot plots (n = 22), where the middle line is the median; the lower and upper edges of the rectangle are the first and third quartiles; the lower and upper whiskers of the box plot represent the interquartile range (IQR) × 1.5; mean values for each distribution are shown as numbers. h. Distribution of Tanimoto similarity between training and test compounds for the SMRTnet dataset across fivefold CV (n = 49, 40, 42, 44, and 32). Labels above the violin plots represent the average Tanimoto similarity between training and test sets. i. Performance comparison of SMRTnet on the test set of SMRTnet using different maximum Tanimoto similarity thresholds. Left: Average Tanimoto similarity between the training and test compounds. Right: Performance of SMRTnet on the test set using the indicated Tanimoto similarity thresholds. Data are represented as mean ± s.d. based on fivefold CV (n = 5). j. Performance comparison of SMRTnet on the original test set of SMRTnet and the multi-strand binding site exclusion test set of SMRTnet. Data are presented as mean ± s.d. based on fivefold CV (n = 5). k. Performance comparison of SMRTnet using a ligand-based data splitting strategy or an RNAmigos2-based data splitting strategy on the original test set of SMRTnet. Data are presented as mean ± s.d. based on fivefold CV (n = 5). l. Performance comparison of SMRTnet across the original, RNA’, and RNA’’ test set of SMRTnet. Data are presented as mean ± s.d. based on fivefold CV (n = 5). m. Comparison of inference time between SMRTnet and Vina-GPU 2.0 across different scales of small molecule–RNA pairs. n. Bar plots show the performance of SMRTnet with different modules in the ablation study. The upset plot with bar plots represents the ablation study of SMRTnet. Data are presented as mean ± s.d. based on fivefold CV (n = 5). Black dots indicate the inclusion of a module, while gray dots indicate the exclusion of a module. For models lacking the multimodal data fusion module, a concatenation module was used as a substitute. o. Performance comparison between SMRTnet on the original SMRTnet dataset and SMRTnet-seq on the modified SMRTnet dataset (using only RNA sequence information). Data are presented as mean ± s.d. based on fivefold CV (n = 5). p. Performance comparison of SMRTnet on the SMRTnet-benchmark dataset with varying RNA secondary structural information. Bar plots represent the auROC values between RNA secondary structure from publications (where, for more than 80% cases, structures are experimentally derived), only RNA sequence without structural information, and predicted RNA secondary structures using RNAstructure on the SMRTnet-benchmark dataset, respectively. Data are presented as mean ± s.d. based on fivefold CV (n = 5). q. Performance comparison between RNASwan-seq and two other RNA language models on the test set of SMRTnet. Bar plots represent auROC values when using RNASwan-seq, RNA-FM, and RNAErnie as the RNA sequence encoder on the test set of SMRTnet. Data are presented as mean ± s.d. based on fivefold CV (n = 5).
Extended Data Fig. 3 Case study of small molecule–RNA interactions using different computational methods.
a-f. Heatmap of structural similarity (measured by the Tanimoto similarity) between theophylline and its 30 decoys across different computational methods, including SMRTnet (a), RNAmigos2 (b), AutoDock Vina (c), NLDock (d), RLDOCK (e), and rDock (f). The red dashed box in (a) represents two groups clustered by ranking score. g-h. Chemical structure of theophylline and its six top-ranked decoys (g) and ten bottom-ranked decoys (h) predicted by SMRTnet. SMRTnet predicted binding scores and names of decoys are listed below. The red dashed box highlights the pyrimidine dione-like structure, where the triazole in decoy_13 serves as a chemical equivalent of pyrimidine.
Extended Data Fig. 4 MST validation of disease-associated RNA targets and corresponding compounds.
MST validation results for 40 compounds targeting ten disease-associated RNA targets. The colors of the outer frame represent various RNA types, with the response amplitude labeled in the frame, and the CAS number, predicted binding score, and signal-to-noise (S/N) ratio listed below. Results are presented using normalized fluorescence, except for those marked by black dashed boxes, which highlight interactions that altered initial fluorescence but passed the SDS denaturation test (SD-test), with results shown using initial fluorescence.
Extended Data Fig. 5 Binding affinity of disease-associated RNA targets and corresponding compounds.
Binding affinity results for ten disease-associated RNA targets and corresponding compounds along with their chemical structures. The colors of the outer frame represent various RNA types.
Extended Data Fig. 6 MST validation and binding affinity of the MYC IRES and corresponding compounds.
a. MST validation for the binding of 15 compounds with the MYC IRES across different prediction intervals. The colors of the outer frame represent various prediction intervals, with the response amplitude labeled in the frame. The CAS number, predicted binding score, and S/N ratio are listed below. Results are presented using normalized fluorescence, except for those marked by black dashed boxes, which highlight interactions that altered initial fluorescence but passed the SDS denaturation test (SD-test), with results shown using initial fluorescence. b. Binding affinity of 15 compounds across different prediction intervals and the MYC IRES. The colors of the outer frame represent various prediction intervals.
Extended Data Fig. 7 Physicochemical and medicinal chemistry profiling of MYC IRES-targeting compounds.
a-e. Physicochemical properties of 15 MYC IRES-targeting compounds were analyzed using ADMETlab 3.0, including the stereo centers (SC) (a), the topological polar surface area (TPSA) (b), the logarithm of aqueous solubility value (logS) (c), the logarithm of the n-octanol/water distribution coefficient (logP) (d), and the logarithm of n-octanol/water distribution coefficients at pH = 7.4 (logD7.4) (e). Based on ADMETlab 3.0 criteria, compounds highlighted with a green background and labeled are considered desirable. f-g. Medicinal chemistry characteristics of 15 MYC IRES-targeting compounds were analyzed using ADMETlab 3.0, including the quantitative estimation of drug-likeness (QED) (f), the ratio of sp3 hybridized carbons/total carbon count (FSP3) (g), the medicinal chemistry evolution in 2018 (MCE-18) (h), violations of Lipinski’s rule (i), and satisfactions of Pfizer’s rule (j). Based on ADMETlab 3.0 criteria, compounds highlighted with a green background and labeled are considered desirable.
Extended Data Fig. 8 MST validation of the mutant MYC IRES against IHT.
a-e. MST validation results for IHT targeting the fully base-paired (a), upper 1×1 internal loop (b), lower 1×1 internal loop (c), 2×2 internal loop (d), and 3×3 internal loop (e) structures of the mutant MYC IRES. The response amplitude is labeled within the frame, while the predicted binding score and S/N ratio are listed below.
Supplementary information
Supplementary Table 1 (download XLSX )
The SMRTnet dataset.
Supplementary Table 2 (download XLSX )
The SMRTnet-benchmark dataset.
Supplementary Table 3 (download XLSX )
CAS numbers and SMILES of the natural product library.
Supplementary Table 4 (download XLSX )
Sequence and secondary structure of ten disease-associated RNAs.
Supplementary Table 5 (download XLSX )
CAS numbers and SMILES of the downsampled subset library.
Supplementary Table 6 (download XLSX )
Sequence and secondary structure of MYC IRES mutants.
Supplementary Table 7 (download XLSX )
qPCR analysis of HeLa cells treated with IHT.
Supplementary Table 8 (download XLSX )
Western blotting analysis of HeLa cells treated with IHT.
Supplementary Table 9 (download XLSX )
Cell proliferation results in three cell lines treated with IHT.
Supplementary Table 10 (download XLSX )
Cell apoptosis results in three cell lines treated with IHT.
Supplementary Table 11 (download XLSX )
Luciferase reporter assay results in 293T cells.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Unprocessed scans of gels.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fei, Y., Wang, P., Zhang, J. et al. Predicting small molecule–RNA interactions without RNA tertiary structures. Nat Biotechnol (2026). https://doi.org/10.1038/s41587-025-02942-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-025-02942-z
This article is cited by
-
Predicting small-molecule–RNA interactions
Nature Reviews Drug Discovery (2026)