Abstract
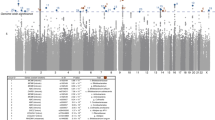
Human genetic variation affects the gut microbiota through a complex combination of environmental and host factors. Here we characterize genetic variations associated with microbial abundances in a single large-scale population-based cohort of 5,959 genotyped individuals with matched gut microbial metagenomes, and dietary and health records (prevalent and follow-up). We identified 567 independent SNP–taxon associations. Variants at the LCT locus associated with Bifidobacterium and other taxa, but they differed according to dairy intake. Furthermore, levels of Faecalicatena lactaris associated with ABO, and suggested preferential utilization of secreted blood antigens as energy source in the gut. Enterococcus faecalis levels associated with variants in the MED13L locus, which has been linked to colorectal cancer. Mendelian randomization analysis indicated a potential causal effect of Morganella on major depressive disorder, consistent with observational incident disease analysis. Overall, we identify and characterize the intricate nature of host–microbiota interactions and their association with disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Complete summary statistics of microbial taxa with genome-wide significant hits are publicly available in the NHGRI-EBI GWAS Catalog (https://www.ebi.ac.uk/gwas/) from accession GCST90032172 to GCST90032644. The metagenomic data from FINRISK 2002 samples are available from the European Genome-Phenome Archive (study ID: EGAS00001005020). The phenotype data contain sensitive information from healthcare registers and are not publicly available to avoid compromising research participant privacy/consent. They are available through the THL biobank upon submission of a research plan and signing a data transfer agreement (https://thl.fi/en/web/thl-biobank/for-researchers/application-process). Additional databases used in this work include GTDB release 89 (https://gtdb.ecogenomic.org/) and CAZy (last accessed 31 July 2019) (http://www.cazy.org/).
Code availability
Scripts used to analyze nonidentifiable data in this study have been made available on Zenodo (https://doi.org/10.5281/zenodo.5641303).
Change history
29 February 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41588-024-01693-y
References
Belizário, J. E. & Napolitano, M. Human microbiomes and their roles in dysbiosis, common diseases, and novel therapeutic approaches. Front. Microbiol. 6, 1050 (2015).
Levy, M., Kolodziejczyk, A. A., Thaiss, C. A. & Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 17, 219–232 (2017).
Blekhman, R. et al. Host genetic variation impacts microbiome composition across human body sites. Genome Biol. 16, 191 (2015).
Davenport, E. R. et al. ABO antigen and secretor statuses are not associated with gut microbiota composition in 1,500 twins. BMC Genomics 17, 941 (2016).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
Bonder, M. J. et al. The effect of host genetics on the gut microbiome. Nat. Genet. 48, 1407–1412 (2016).
Turpin, W. et al. Association of host genome with intestinal microbial composition in a large healthy cohort. Nat. Genet. 48, 1413–1417 (2016).
Wang, J. et al. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat. Genet. 48, 1396–1406 (2016).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Hughes, D. A. et al. Genome-wide associations of human gut microbiome variation and implications for causal inference analyses. Nat. Microbiol. 5, 1079–1087 (2020).
Kurilshikov, A. et al. Large-scale association analyses identify host factors influencing human gut microbiome composition. Nat. Genet. 53, 156–165 (2021).
Kolde, R. et al. Host genetic variation and its microbiome interactions within the Human Microbiome Project. Genome Med. 10, 6 (2018).
Rühlemann, M. C. et al. Application of the distance-based F test in an mGWAS investigating β diversity of intestinal microbiota identifies variants in SLC9A8 (NHE8) and 3 other loci. Gut Microbes 9, 68–75 (2018).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Xie, H. et al. Shotgun metagenomics of 250 adult twins reveals genetic and environmental impacts on the gut microbiome. Cell Syst. 3, 572–584.e3 (2016).
Lim, M. Y. et al. The effect of heritability and host genetics on the gut microbiota and metabolic syndrome. Gut 66, 1031–1038 (2017).
Le Roy, C. I. et al. Heritable components of the human fecal microbiome are associated with visceral fat. Gut Microbes 9, 61–67 (2018).
Goodrich, J. K., Davenport, E. R., Clark, A. G. & Ley, R. E. The relationship between the human genome and microbiome comes into view. Annu. Rev. Genet. 51, 413–433 (2017).
Kurilshikov, A., Wijmenga, C., Fu, J. & Zhernakova, A. Host genetics and gut microbiome: challenges and perspectives. Trends Immunol. 38, 633–647 (2017).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Zhernakova, A. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352, 565–569 (2016).
Eng, A. & Borenstein, E. Taxa-function robustness in microbial communities. Microbiome 6, 45 (2018).
Ferrer, M. et al. Microbiota from the distal guts of lean and obese adolescents exhibit partial functional redundancy besides clear differences in community structure: metaproteomic insights associated to human obesity. Environ. Microbiol. 15, 211–226 (2013).
Moya, A. & Ferrer, M. Functional redundancy-induced stability of gut microbiota subjected to disturbance. Trends Microbiol. 24, 402–413 (2016).
Louca, S. et al. Function and functional redundancy in microbial systems. Nat. Ecol. Evol. 2, 936–943 (2018).
Louca, S. et al. High taxonomic variability despite stable functional structure across microbial communities. Nat. Ecol. Evol. 1, 0015 (2017).
Banerjee, S., Schlaeppi, K. & van der Heijden, M. G. A. Keystone taxa as drivers of microbiome structure and functioning. Nat. Rev. Microbiol. 16, 567–576 (2018).
Trosvik, P. & de Muinck, E. J. Ecology of bacteria in the human gastrointestinal tract—identification of keystone and foundation taxa. Microbiome 3, 44 (2015).
Shetty, S. A., Hugenholtz, F., Lahti, L., Smidt, H. & de Vos, W. M. Intestinal microbiome landscaping: insight in community assemblage and implications for microbial modulation strategies. FEMS Microbiol. Rev. 41, 182–199 (2017).
Chia, L. W. et al. Deciphering the trophic interaction between Akkermansia muciniphila and the butyrogenic gut commensal Anaerostipes caccae using a metatranscriptomic approach. Antonie Van Leeuwenhoek 111, 859–873 (2018).
Banerjee, S., Schlaeppi, K. & van der Heijden, M. G. A. Reply to ‘Can we predict microbial keystones?’. Nat. Rev. Microbiol. 17, 194 (2019).
Röttjers, L. & Faust, K. Can we predict keystones? Nat. Rev. Microbiol. 17, 193 (2019).
Kato, K. et al. Age-related changes in the composition of gut Bifidobacterium species. Curr. Microbiol. 74, 987–995 (2017).
Engevik, M. A. et al. Bifidobacterium dentium fortifies the intestinal mucus layer via autophagy and calcium signaling pathways. mBio 10, e01087–19 (2019) .
Rahfeld, P. & Withers, S. G. Toward universal donor blood: enzymatic conversion of A and B to O type. J. Biol. Chem. 295, 325–334 (2020).
Liu, Q. P. et al. Bacterial glycosidases for the production of universal red blood cells. Nat. Biotechnol. 25, 454–464 (2007).
Arnolds, K. L., Martin, C. G. & Lozupone, C. A. Blood type and the microbiome—untangling a complex relationship with lessons from pathogens. Curr. Opin. Microbiol. 56, 59–66 (2020).
Liu, Q. P. et al. Identification of a GH110 subfamily of α1,3-galactosidases: novel enzymes for removal of the α3GAL xenotransplantation antigen. J. Biol. Chem. 283, 8545–8554 (2008).
Pichler, M. J. et al. Butyrate producing colonic Clostridiales metabolise human milk oligosaccharides and cross feed on mucin via conserved pathways. Nat. Commun. 11, 3285 (2020).
Ficko-Blean, E. & Boraston, A. B. The interaction of a carbohydrate-binding module from a Clostridium perfringens N-acetyl-β-hexosaminidase with its carbohydrate receptor. J. Biol. Chem. 281, 37748–37757 (2006).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e21 (2016).
Tailford, L. E., Crost, E. H., Kavanaugh, D. & Juge, N. Mucin glycan foraging in the human gut microbiome. Front. Genet. 6, 81 (2015).
Genome Aggregation Database Consortium et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Amarnani, R. & Rapose, A. Colon cancer and enterococcus bacteremia co-affection: a dangerous alliance. J. Infect. Public Health 10, 681–684 (2017).
Khan, Z., Siddiqui, N. & Saif, M. W. Enterococcus faecalis infective endocarditis and colorectal carcinoma: case of new association gaining ground. Gastroenterol. Res. 11, 238–240 (2018).
Huycke, M. M., Abrams, V. & Moore, D. R. Enterococcus faecalis produces extracellular superoxide and hydrogen peroxide that damages colonic epithelial cell DNA. Carcinogenesis 23, 529–536 (2002).
Allen, B. L. & Taatjes, D. J. The Mediator complex: a central integrator of transcription. Nat. Rev. Mol. Cell Biol. 16, 155–166 (2015).
Firestein, R. et al. CDK8 is a colorectal cancer oncogene that regulates β-catenin activity. Nature 455, 547–551 (2008).
Li, L., Batt, S. M., Wannemuehler, M., Dispirito, A. & Beitz, D. C. Effect of feeding of a cholesterol-reducing bacterium, Eubacterium coprostanoligenes, to germ-free mice. Lab. Anim. Sci. 48, 253–255 (1998).
Marasco, G. et al. Gut microbiota and celiac disease. Dig. Dis. Sci. 61, 1461–1472 (2016).
Lavasani, S. et al. A novel probiotic mixture exerts a therapeutic effect on experimental autoimmune encephalomyelitis mediated by IL-10 producing regulatory T cells. PLoS ONE 5, e9009 (2010).
Tomita, H. et al. G protein-linked signaling pathways in bipolar and major depressive disorders. Front. Genet. 4, 297 (2013).
Wong, M.-L. et al. Phosphodiesterase genes are associated with susceptibility to major depression and antidepressant treatment response. Proc. Natl Acad. Sci. USA 103, 15124–15129 (2006).
Schork, A. J. et al. A genome-wide association study of shared risk across psychiatric disorders implicates gene regulation during fetal neurodevelopment. Nat. Neurosci. 22, 353–361 (2019).
Burger, J. et al. Low prevalence of lactase persistence in Bronze Age Europe indicates ongoing strong selection over the last 3,000 years. Curr. Biol. https://doi.org/10.1016/j.cub.2020.08.033 (2020).
Gerbault, P. et al. Evolution of lactase persistence: an example of human niche construction. Philos. Trans. R. Soc. Lond. B Biol. Sci. 366, 863–877 (2011).
Hebert, J. R. et al. Social desirability trait influences on self-reported dietary measures among diverse participants in a multicenter multiple risk factor trial. J. Nutr. 138, 226S–234S (2008).
Schoeller, D. A. How accurate is self-reported dietary energy intake? Nutr. Rev. 48, 373–379 (2009).
Sakanaka, M. et al. Evolutionary adaptation in fucosyllactose uptake systems supports bifidobacteria-infant symbiosis. Sci. Adv. 5, eaaw7696 (2019).
Storhaug, C. L., Fosse, S. K. & Fadnes, L. T. Country, regional, and global estimates for lactose malabsorption in adults: a systematic review and meta-analysis. Lancet Gastroenterol. Hepatol. 2, 738–746 (2017).
Liu, X. et al. A genome-wide association study for gut metagenome in Chinese adults illuminates complex diseases. Cell Discov. 7, 9 (2021).
Sirugo, G., Williams, S. M. & Tishkoff, S. A. The missing diversity in human genetic studies. Cell 177, 26–31 (2019).
Szilagyi, A. Adaptation to lactose in lactase non persistent people: effects on intolerance and the relationship between dairy food consumption and evalution of diseases. Nutrients 7, 6751–6779 (2015).
Ségurel, L., Gao, Z. & Przeworski, M. Ancestry runs deeper than blood: the evolutionary history of ABO points to cryptic variation of functional importance. Bioessays https://doi.org/10.1002/bies.201300030 (2013).
Segurel, L. et al. The ABO blood group is a trans-species polymorphism in primates. Proc. Natl Acad. Sci. USA 109, 18493–18498 (2012).
Ewald, D. R. & Sumner, S. C. J. Blood type biochemistry and human disease. Wiley Interdisp. Rev. Syst. Biol. Med. 8, 517–535 (2016).
Ellinghaus, D. et al. Genomewide asociation study of severe Covid-19 with respiratory failure. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2020283 (2020).
Shelton, J. F. et al. Trans-ancestry analysis reveals genetic and nongenetic associations with COVID-19 susceptibility and severity. Nat. Genet. 53, 801–808 (2021).
Rühlemann, M. C. et al. Genome-wide association study in 8,956 German individuals identifies influence of ABO histo-blood groups on gut microbiome. Nat. Genet. 53, 147–155 (2021).
Liu, X. et al. Mendelian randomization analyses support causal relationships between blood metabolites and the gut microbiome. Preprint at bioRxiv https://doi.org/10.1101/2020.06.30.181438 (2020).
Knuesel, M. T., Meyer, K. D., Bernecky, C. & Taatjes, D. J. The human CDK8 subcomplex is a molecular switch that controls Mediator coactivator function. Genes Dev. 23, 439–451 (2009).
Tsai, K.-L. et al. A conserved Mediator–CDK8 kinase module association regulates Mediator–RNA polymerase II interaction. Nat. Struct. Mol. Biol. 20, 611–619 (2013).
De Almeida, C. et al. Differential responses of colorectal cancer cell lines to Enterococcus faecalis’ strains isolated from healthy donors and colorectal cancer patients. J. Clin. Med. 8, 388 (2019).
Marchesi, J. R. et al. The gut microbiota and host health: a new clinical frontier. Gut 65, 330–339 (2016).
Ma, Y. et al. Proposal for reunification of the genus Raoultella with the genus Klebsiella and reclassification of Raoultella electrica as Klebsiella electrica comb. nov. Res. Microbiol. https://doi.org/10.1016/j.resmic.2021.103851 (2021).
Wyres, K. L., Lam, M. M. C. & Holt, K. E. Population genomics of Klebsiella pneumoniae. Nat. Rev. Microbiol. 18, 344–359 (2020).
Jiang, H. et al. Altered fecal microbiota composition in patients with major depressive disorder. Brain Behav. Immun. 48, 186–194 (2015).
Wade, K. H. & Hall, L. J. Improving causality in microbiome research: can human genetic epidemiology help? Wellcome Open Res. 4, 199 (2020).
Foster, J. A. & McVey Neufeld, K.-A. Gut–brain axis: how the microbiome influences anxiety and depression. Trends Neurosci. 36, 305–312 (2013).
Fung, T. C., Olson, C. A. & Hsiao, E. Y. Interactions between the microbiota, immune and nervous systems in health and disease. Nat. Neurosci. 20, 145–155 (2017).
Valles-Colomer, M. et al. The neuroactive potential of the human gut microbiota in quality of life and depression. Nat. Microbiol. 4, 623–632 (2019).
Maes, M., Kubera, M. & Leunis, J.-C. The gut-brain barrier in major depression: intestinal mucosal dysfunction with an increased translocation of LPS from gram negative enterobacteria (leaky gut) plays a role in the inflammatory pathophysiology of depression. Neuro Endocrinol. Lett. 29, 117–124 (2008).
Yang, J. et al. Landscapes of bacterial and metabolic signatures and their interaction in major depressive disorders. Sci. Adv. 6, eaba8555 (2020).
Mattar, R., de Campos Mazo, D. F. & Carrilho, F. J. Lactose intolerance: diagnosis, genetic, and clinical factors. Clin. Exp. Gastroenterol. 5, 113–121 (2012).
Bodmer, W. Genetic characterization of human populations: from ABO to a genetic map of the British people. Genetics 199, 267–279 (2015).
Parks, D. H. et al. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life. Nat. Biotechnol. 36, 996–1004 (2018).
Parks, D. H. et al. A complete domain-to-species taxonomy for Bacteria and Archaea. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0501-8 (2020).
Méric, G., Wick, R. R., Watts, S. C., Holt, K. E. & Inouye, M. Correcting index databases improves metagenomic studies. Preprint at bioRxiv https://doi.org/10.1101/712166 (2019).
Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 176, 649–662.e20 (2019).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0603-3 (2020).
Borodulin, K. et al. Cohort profile: the national FINRISK study. Int. J. Epidemiol. 47, 696–696i (2018).
Borodulin, K. et al. Forty-year trends in cardiovascular risk factors in Finland. Eur. J. Public Health 25, 539–546 (2015).
Liu, Y. et al. Early prediction of liver disease using conventional risk factors and gut microbiome-augmented gradient boosting. Preprint at medRxiv https://doi.org/10.1101/2020.06.24.20138933 (2020).
Salosensaari, A. et al. Taxonomic signatures of cause-specific mortality risk in human gut microbiome. Nat. Commun. 12, 2671 (2021).
FinnGen et al. Polygenic and clinical risk scores and their impact on age at onset and prediction of cardiometabolic diseases and common cancers. Nat. Med. 26, 549–557 (2020).
Loh, P.-R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Howie, B., Fuchsberger, C., Stephens, M., Marchini, J. & Abecasis, G. R. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat. Genet. 44, 955–959 (2012).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaSci 4, 7 (2015).
Kim, D., Song, L., Breitwieser, F. P. & Salzberg, S. L. Centrifuge: rapid and sensitive classification of metagenomic sequences. Genome Res. 26, 1721–1729 (2016).
Ruuskanen, M. O. et al. Links between gut microbiome composition and fatty liver disease in a large population sample. Gut Microbes 13, 1–22 (2021).
Goodrich, J. K., Davenport, E. R., Waters, J. L., Clark, A. G. & Ley, R. E. Cross-species comparisons of host genetic associations with the microbiome. Science 352, 532–535 (2016).
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V. & Egozcue, J. J. Microbiome datasets are compositional: and this is not optional. Front. Microbiol. 8, 2224 (2017).
Aitchison, J., Barceló-Vidal, C., Martín-Fernández, J. A. & Pawlowsky-Glahn, V. Logratio Analysis and Compositional Distance. Math. Geol. 32, 271–275 (2000).
Qin, Y. et al. Genome-wide association and Mendelian randomization analysis prioritizes bioactive metabolites with putative causal effects on common diseases. Preprint at medRxiv https://doi.org/10.1101/2020.08.01.20166413 (2020).
Loh, P.-R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Abraham, G., Qiu, Y. & Inouye, M. FlashPCA2: principal component analysis of Biobank-scale genotype datasets. Bioinformatics 33, 2776–2778 (2017).
Genetic Investigation of ANthropometric Traits (GIANT) Consortium et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Li, J. & Ji, L. Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity 95, 221–227 (2005).
Nyholt, D. R. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am. J. Hum. Genet. 74, 765–769 (2004).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Davenport, E. R. et al. Seasonal variation in human gut microbiome composition. PLoS ONE 9, e90731 (2014).
Paré, G. et al. Novel association of ABO histo-blood group antigen with soluble ICAM-1: results of a genome-wide association study of 6,578 women. PLoS Genet. 4, e1000118 (2008).
Wacklin, P. et al. Secretor genotype (FUT2 gene) is strongly associated with the composition of Bifidobacteria in the human intestine. PLoS ONE 6, e20113 (2011).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Sanna, S. et al. Causal relationships among the gut microbiome, short-chain fatty acids and metabolic diseases. Nat. Genet. 51, 600–605 (2019).
Zhang, H. et al. dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 46, W95–W101 (2018).
Cantarel, B. L. et al. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res. 37, D233–D238 (2009).
Cantarel, B. L., Lombard, V. & Henrissat, B. Complex carbohydrate utilization by the healthy human microbiome. PLoS ONE 7, e28742 (2012).
The CAZypedia Consortium. Ten years of CAZypedia: a living encyclopedia of carbohydrate-active enzymes. Glycobiology 28, 3–8 (2018).
Lannelongue, L., Grealey, J. & Inouye, M. Green algorithms: quantifying the carbon footprint of computation. Adv. Sci. 8, 2100707 (2021).
Acknowledgements
The study protocol of FINRISK 2002 was approved by the Coordinating Ethical Committee of the Helsinki and Uusimaa Hospital District (Ref. 558/E3/2001). All participants signed an informed consent. The study was conducted according to the World Medical Association Declaration of Helsinki on ethical principles. All necessary patient/participant consent has been obtained and the appropriate institutional forms have been archived. We thank all participants of the FINRISK 2002 survey for their contributions to this work. The FINRISK surveys are mainly funded by budgetary funds from the Finnish Institute for Health and Welfare with additional funding from several domestic foundations. Y.Q. was partially supported by The Albert Shimmins Fund (Faculty of Science Postgraduate Writing-Up Award 2020). M.I. was supported by the Munz Chair of Cardiovascular Prediction and Prevention and the NIHR Cambridge Biomedical Research Centre (BRC-1215-20014). V.S. was supported by the Finnish Foundation for Cardiovascular Research. L.L. was supported by the Academy of Finland (decision 295741) and EU/H2020 (FindingPheno; grant 952914). T.N. was supported by the Emil Aaltonen Foundation, the Finnish Medical Foundation, the Paavo Nurmi Foundation and the Academy of Finland (grant no. 321351). A.S.H. was supported by the Academy of Finland, grant no. 321356. R.L. receives funding support from NIEHS (grant no. 5P42ES010337), NCATS (grant no. 5UL1TR001442), NIDDK (grant nos. U01DK061734, R01DK106419, P30DK120515, R01DK121378, R01DK124318) and DOD PRCRP (grant no. W81XWH-18-2-0026). S.C.R. is funded by a BHF Programme Grant (RG/18/13/33946). This study was supported by the Victorian Government’s Operational Infrastructure Support (OIS) program, and by core funding from the British Heart Foundation (grant no. RG/13/13/30194; RG/18/13/33946) and the NIHR Cambridge Biomedical Research Centre (BRC-1215-20014). National Institute for Health Research (Cambridge Biomedical Research Centre at the Cambridge University Hospitals NHS Foundation Trust) (The views expressed are those of the authors and not necessarily those of the NIHR or the Department of Health and Social Care). This work was supported by Health Data Research UK, which is funded by the UK Medical Research Council, the Engineering and Physical Sciences Research Council, the Economic and Social Research Council, the Department of Health and Social Care (England), the Chief Scientist Office of the Scottish Government Health and Social Care Directorates, the Health and Social Care Research and Development Division (Welsh Government), the Public Health Agency (Northern Ireland), the British Heart Foundation and Wellcome. We thank Dr Annalisa Buniello (EMBL-EBI, Cambridge, UK) for valuable help with GWAS Catalog submissions.
Author information
Authors and Affiliations
Contributions
Y.Q., M.I., V.S. and G.M. designed the work. A.S.H., P.J., J.G.S., L.V., M.B., Q.Z., A. Tripathi, Y.V.-B., T.N., L.L., R.K., V.S. and G.M. acquired the data. Y.Q., Y.L., S.C.R., J.G.S., L.L., A. Tokolyi and G.M. analyzed the data. R.L., S.C., M.J., T.N., L.L., R.K., V.S., M.I. and G.M. supervised the work. All authors wrote the manuscript and gave final approval of the version to be published.
Corresponding authors
Ethics declarations
Competing interests
V.S. has consulted for Novo Nordisk and Sanofi and received honoraria from these companies. He also has ongoing research collaboration with Bayer AG, all unrelated to this study. R.L. serves as a consultant or advisory board member for Anylam/Regeneron, Arrowhead Pharmaceuticals, AstraZeneca, Bird Rock Bio, Boehringer Ingelheim, Bristol-Myer Squibb, Celgene, Cirius, CohBar, Conatus, Eli Lilly, Galmed, Gemphire, Gilead, Glympse bio, GNI, GRI Bio, Inipharm, Intercept, Ionis, Janssen Inc., Merck, Metacrine, Inc., NGM Biopharmaceuticals, Novartis, Novo Nordisk, Pfizer, Prometheus, Promethera, Sanofi, Siemens and Viking Therapeutics. In addition, his institution has received grant support from Allergan, Boehringer Ingelheim, Bristol-Myers Squibb, Cirius, Eli Lilly and Company, Galectin Therapeutics, Galmed Pharmaceuticals, GE, Genfit, Gilead, Intercept, Grail, Janssen, Madrigal Pharmaceuticals, Merck, NGM Biopharmaceuticals, NuSirt, Pfizer, pH Pharma, Prometheus and Siemens. He is also cofounder of Liponexus, Inc. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Kenneth Croitoru and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
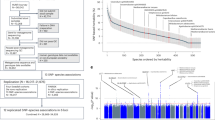
Extended Data Fig. 1
Study flowchart.
Extended Data Fig. 2 Heritability of SNPs associated with microbial taxa.
(a) Associated SNP heritability (h2) for all 2,801 taxa included in the genome-wide association analysis, grouped into their 61 corresponding GTDB phyla, and ordered by median heritability per phylum. Red denotes bacterial phyla, and purple denotes archaeal phyla. The right panel indicates the number of genome-wide significant associated taxa for each phylum. (b) Associated SNP heritability is shown for each associated taxon, grouped by its taxonomic rank. Genome-wide significance was defined as a threshold of p < 5 × 10-8 for all p-values obtained after joint analysis using GTCA-COJO in the GWAS (see Methods). For all box plots (A and B), the central line, box and whiskers represent the median, interquartile range (IQR) and 1.5 times the IQR, respectively.
Extended Data Fig. 3 LocusZoom plots for three loci with study-wide significant associations (p < 3.8 × 10-11).
Associations with top taxa are shown. Top SNPs are indicated in purple diamond. Other SNPs are coloured by their linkage disequilibrium (LD) values with the top SNPs. Genes covered by the region are indicated in the bottom and the genotyping coverage is indicated on top of the plot. (A) Associated SNPs at the LCT locus spans over a 2 Mbp genomic region, while they are grouped on a 400 kbp region for both (B) ABO and (C) the MED13L loci.
Extended Data Fig. 4 Correlation between individual baseline age and the relative abundance of bacteria from the Bifidobacterium genus in lactose intolerant individuals.
Only genetically lactose intolerance individuals (rs4988235:CC) are shown, and coloured by dietary dairy habits (blue: self-reported regular consumption of dairy, n = 763; and red: self-reported irregular dairy diet or lactose-free diet, n = 253). Best fitted lines and 95% confidence intervals are indicated. Two-sided Spearman correlation coefficients are given.
Extended Data Fig. 5 Spearman correlation of relative abundances in 4 taxa associated with the LCT locus.
Abundances of the Bifidobacterium, Negativibacillus, UBA3855 and CAG-81 genera are compared. Abundances in the entire FR02 cohort is compared to those in a subset of genetically lactose-intolerant individuals, and to a subset of genetically lactose-intolerant individuals who reported a regular dairy diet. Coloured boxes denote the strength of correlation (ranging from -1 in red to 1 in dark blue), while a white square denotes a non-significant p-value for the two-sided Spearman correlation (p > 0.05).
Extended Data Fig. 6 Co-abundance and carbohydrate-active enzymes (CAZyme) distribution patterns in 11 Bifidobacterium species harboured by > 25% of individuals in the FR02 cohort.
(a) Associations between the LCT-MCM6 locus and 11 Bifidobacterium species; (left) top association results between variation of 11 Bifidobacterium species and the LCT locus, with study-wide significant associations (with p-values from the joint analysis using GTCA-COJO below the p < 3.8 × 10-11 threshold) highlighted in bold; (middle) Two-sided Spearman coefficients calculated on CLR-transformed abundances; (right) relative abundances across the FR02 cohort, ranging from 0 (light green) to 1 (dark blue). (b) CAZyme distribution patterns in 327 previously published reference genomes from 11 Bifidobacterium GTDB species which were included in the GTDB release 89 index used to classify metagenomes in this study. The heatmap indicates abundance of corresponding CAZyme families in species, corresponding to the total count of detected families for each species divided by the number of reference genomes examined for the same species. Values <1 (white to light blue) indicate that less than one copy per genome of the corresponding CAZyme family was detected for each species, values >1 (light blue to dark blue) indicate that more than one copy per genome was detected. Preferred substrate groups are based on literature search and descriptions on CAZypedia.org. For all box plots (A), the central line, box and whiskers represent the median, interquartile range (IQR) and 1.5 times the IQR, respectively. Violin plots represent the distribution density of the data points.
Extended Data Fig. 7 Effect of ABO genotypes, blood type and secretor status on microbial diversity and gut levels of ABO-associated taxa.
(a) (left) Alpha diversity represented by Shannon indices; (right) beta diversity, represented by Bray-Curtis distances. Alpha and beta diversity were calculated from individual taxonomic profiles at the genus level. Individuals were segregated according to their predicted blood type and secretor status, both predicted from genotype data. (b) Abundances are compared across stratified groups of individuals from the FR02 cohort according to (left panel): ABO:rs545971 genotype and predicted secretor status (blue: secretor status conferred by FUT2 rs601338:GG/GA genotype; red: non-secretor status conferred by FUT2 rs601338:AA genotype) and (right panel) according to predicted A, AB, B and O blood types, and predicted secretor status. All statistical comparisons denote the p-values of Wilcoxon rank test on the distributions. (c) Effect of AB antigen secretion on gut microbial relative abundance, using the 2,801 taxa considered for GWAS in our study. Taxa with FDR adjusted p value <0.05 are highlighted in red. Red line indicates the expected distribution of p values under the null hypothesis. P values were calculated using Wilcoxon rank test. For all box plots (A and B), the central line, box and whiskers represent the median, interquartile range (IQR) and 1.5 times the IQR, respectively. Violin plots represent the distribution density of the data points.
Extended Data Fig. 8 Sequencing depth does not influence alpha diversity.
Alpha-diversity (Shannon index) were computed and plotted against the log10 (left) or the raw (right) number of sequencing reads for each 5,959 individual gut metagenome in this study. No correlation was observed between sequencing depth and Shannon diversity index (two-sided Spearman’s ⍴=-0.001598, p = 0.90). Grey shaded area corresponds to the 95% confidence interval.
Extended Data Fig. 9 Effect of geographical region of sampling, microbiome sequencing batch or antibiotic prescription on overall microbiome diversity.
Beta-diversity (Bray Curtis dissimilarity indices) was calculated using the R package vegan, and the 4 top PCoA axes (explaining a combined 25.9% of the total microbiome variation) were plotted against each other, with each individual point labelled according to geographical sampling (panel A), gut metagenomic sequencing batch (panel B), or whether antibiotics were prescribed up to 1 month (n = 250/5959) before baseline sampling.
Extended Data Fig. 10 Distribution of F. lactaris relative abundance in groups of individuals with different predicted blood types.
A beeswarm plot is used to visualise the distribution of relative abundances.
Supplementary information
Supplementary Information (download PDF )
Supplementary Note
Supplementary Tables (download XLSX )
Supplementary Tables 1–10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qin, Y., Havulinna, A.S., Liu, Y. et al. Combined effects of host genetics and diet on human gut microbiota and incident disease in a single population cohort. Nat Genet 54, 134–142 (2022). https://doi.org/10.1038/s41588-021-00991-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-021-00991-z
This article is cited by
-
Enterococcus faecalis-induced bystander effect causes epigenetic alterations leading to aberrant TGF-β signaling and epithelial-mesenchymal transition
Cell Communication and Signaling (2026)
-
Cross-ancestry genome-wide association studies of liver function biomarkers uncover pleiotropic variants, systemic disease links and therapeutic targets
Genome Medicine (2026)
-
Host m6A modifications shape microbiota that drives cell specific ferroptosis as a causal pathway to chronic respiratory diseases
Scientific Reports (2026)
-
Human and bacterial genetic variation shape oral microbiomes and health
Nature (2026)
-
Genomics of host–microbiome interactions in humans
Nature Reviews Genetics (2026)