Abstract
To maintain cell fitness, deleterious genetic alterations are buffered by compensatory changes in additional genes. In cancer, buffering processes could be targeted by synthetic lethality. However, despite the large-scale identification of synthetic lethal effects in preclinical models, evidence that these operate clinically is limited. This impedes the application of synthetic lethal approaches. By integrating molecular profiling data from >9,000 cancers with synthetic lethal screens, we show that transcriptomic buffering of tumor suppressor gene (TSG) loss by hyperexpression of synthetic lethal partners is a common phenomenon, extending to multiple TSGs and histotypes. Transcriptomic buffering is also notable in cancers that phenocopy TSG loss, such as BRCAness cancers, where expression of BRCA1/2 synthetic lethal genes correlates with clinical outcome. Synthetic lethal genes that exhibit transcriptomic buffering also represent more robust synthetic lethal effects. These observations have implications for understanding how tumor cells tolerate TSG loss, in part explain transcriptomic architectures in cancer and provide insight into target selection.
Similar content being viewed by others
Main
Loss of tumor suppressor genes (TSGs) is critical to oncogenic transformation1. Although providing fitness advantages to cells, TSG loss can also impart deleterious phenotypes, which, if not buffered, impair cell fitness. Buffering processes that compensate for TSG loss are sometimes unveiled as synthetic lethal effects with TSGs. For example, although loss-of-function mutations in BRCA1 or BRCA2 impair homologous recombination (HR) and foster genomic instability and mutagenesis, they also, in certain circumstances, invoke a proliferative defect2. The negative impact of BRCA1 or BRCA2 mutation on tumor cells is in part ameliorated by buffering from other DNA-repair and genome stability mechanisms such as Polθ-mediated end-joining3,4,5,6 and CIP2A-mediated processes that maintain the integrity of mitotic chromosomes7. The importance of these two buffering processes was recently confirmed by the identification of synthetic lethal interactions between BRCA1 or BRCA2 (BRCA1/2) and either Polθ3,4,5,6 or CIP2A7. On a wider scale, the mapping of synthetic lethal relationships, often using genetic perturbation screens, has identified buffering processes that compensate for gene loss in tumor cell lines and model organisms such as yeast8,9,10,11,12,13. However, to date, a systematic assessment of whether synthetic lethal effects identified by experimental perturbation screens operate in human cancers has been missing. We reasoned that, if evidence of buffering relationships between TSGs and synthetic lethal genes could be established in human tumors, this could potentially explain the molecular make-up of human cancers and also possibly identify subsets of the disease where synthetic lethal buffering relationships could be exploited therapeutically.
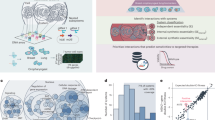
We developed an analytical approach, SYLVER (SYnthetic Lethal Vulnerabilities Exhibiting Reciprocation), that integrates CRISPR–Cas9 perturbation screen data with genome-wide molecular profiles from 9,316 tumours representing 32 tumor subtypes to identify buffering processes associated with TSG dysfunction in human cancers (Fig. 1). We first classified >1,000 tumor cell lines according to the presence or absence of an obvious defect (gene deletion, deleterious mutation or low messenger RNA abundance) in each of 193 TSGs14 (Supplementary Table 1). Using these classifications, we then interrogated genome-wide DepMap CRISPR–Cas9 screen data for each of the tumor cell lines12 and used a linear model to identify candidate synthetic lethal effects associated with each TSG defect. SYLVER was then used to identify buffering relationships between TSGs and synthetic lethal gene partners by interrogating genome-wide transcriptomic profiles of 9,316 TCGA (The Cancer Genome Atlas) tumours spanning 32 cancer subtypes (Supplementary Table 2). We used the same criteria as for the cell lines to classify cohorts of clinical samples with or without a TSG defect15 and used linear regression modeling of gene expression data from the same tumors to identify hyperexpression of synthetic lethal gene partners in the presence of specific TSG defects. Groups of synthetic lethal genes that were hyperexpressed alongside a specific TSG defect were termed ‘synthetic lethal metagenes’ (SLMs; Fig. 1). For example, the collection of synthetic lethal genes associated with RB1 loss, which were also hyperexpressed in RB1 defective cancers, comprised an RB1 SLM. As a final step in the analytical workflow of SYLVER, SLMs were assessed for their prognostic and predictive potential.
Schematic illustrating five steps of SYLVER enabling integration of CRISPR–Cas9 screens with molecular profiles of clinical samples. Integration of these data highlights hyperexpression of pan-cancer synthetic lethal genes as a form of compensating mechanism for the loss of a TSG(s). The resulting hyperexpressed SLMs can be used to develop biomarkers of risk prediction and response to therapy. In step 4, the asterisk indicates exclusion of one TSG (PTK6) that did not reveal any synthetic lethal effects in step 2.
For the 193 TSGs analyzed, SYLVER identified 45,213 candidate synthetic lethal gene pairs for 192 TSGs (Fig. 2a). Interrogation of TCGA transcriptomic data, showed that 57% (25,698) of the pan-cancer candidate synthetic lethal pairs were represented by hyperexpression of the synthetic lethal gene in cancers where the relevant TSG was defective (Fig. 2a). When we considered the synthetic lethal genes associated with 192 TSGs and assessed whether these were hyperexpressed in 32 cancer types where the relevant TSG loss was observed, we identified 4,564 SLMs, of which 312 (representing 72 TSGs) exhibited significant overlap between synthetic lethal and hyperexpressed genes (SLM enrichment P < 0.05, Fisher’s exact test; Extended Data Fig. 1a). Overall, 52.5% of the SLMs contained ≥5 genes, with 25% containing ≥25 genes (Fig. 2b). In many cases, large numbers of synthetic lethal genes were upregulated in multiple cancer histologies where the TSG was defective, as illustrated for synthetic lethal effects associated with either RB1 (RB transcriptional corepressor 1), CDH1 (E-cadherin), PTEN (phosphatase and tensin homolog), CDKN2A (cyclin-dependent kinase inhibitor) or MAP3K13 (mitogen-activated protein kinase) defects (Fig. 2c–e and Supplementary Figs. 1 and 2). The fact that this reciprocal relationship between the loss of specific TSGs and hyperexpression of specific synthetic lethal partners was a widespread phenomenon suggested that distinctive, transcriptionally regulated processes are employed by tumor cells to maintain fitness in the setting of the specific biological challenges imposed by an individual TSG defect.
a, Bar plot showing the number of candidate synthetic lethal genes identified from CRISPR–Cas9 screens and the number of hyperexpressed synthetic lethal genes associated with specific TSG loss from TCGA tumor transcriptomes. b, Summary of observed SLM sizes expressed as empirical cumulative density function. Cumulative probabilty of observing SLM size ≤5, ≤25 and ≤50 are highlighted with dashed gray lines. c, Box plots illustrating hyperexpression (effect size >0) in RB1-defective patients in different TCGA cancer types. Dots indicate genes in cancer-specific SLMRB1 (hyperexpression effect size ≥0.25, FDR-adjusted P < 0.05 and synthetic lethal effect size ≤−0.25 and P < 0.01, with P values calculated using unpaired, two-sided, moderated Student’s t-test). The size of the dot shows the magnitude of the synthetic lethal effect size. Example RB1 synthetic lethal effects are highlighted. The number of genes in SLMs are shown in parentheses. Box plots show the first quartile (Q1), median and third quartile (Q3), with whiskers extended to points closest to Q1 − 1.5× the IQR (interquartile range) and Q3 + 1.5× the IQR. d, Box plots illustrating hyperexpressed genes in CDH1-defective patients in different TCGA cancer types. Dots indicate genes in cancer-specific SLMCDH1. Example CDH1 synthetic lethal effects are highlighted. Statistical thresholds are the same as in c. e, Box plots illustrating hyperexpressed genes in PTEN-defective patients in different TCGA cancer types. Dots indicate genes in cancer-specific SLMPTEN. Example PTEN synthetic lethal effects are highlighted. The statistical thresholds are the same as in c. f, Scatter plot illustrating RB1 synthetic lethal effects identified using an isogenic RB1 CRISPR–Cas9 screen in MCF10ATP53−/− cells. Delta (Δ) represents the difference in viability (normalized z-score) between MCF10AIsogenic and MCF10AParental cells. Key RB1 synthetic lethal genes are highlighted. g, Top, Venn diagram showing the overlap between DepMap-identified RB1 synthetic lethal genes and RB1 synthetic lethal genes identified using the MCF10A isogenic RB1 CRISPR–Cas9 screens. Bottom, Venn diagrams showing the overlap between RB1 synthetic lethal genes found in both DepMap and isogenic RB1 screens (74 genes) and hyperexpressed genes in RB1-defective cancers from SCAN-B and TCGA TNBC datasets. Over-representation P values are reported for all Venn diagrams (Fisher’s exact test).
Although nonisogenic CRISPR screens (for example, DepMap) allow the penetrance of synthetic lethal effects to be estimated, they do have the potential to identify false-positive effects, including gene dependencies that are not TSG synthetic lethal effects, but are dependencies associated with other features that correlate with TSG dysfunction16. Conversely, isogenic screens establish the causal relationship between TSG loss and synthetic lethal gene, but do not determine synthetic lethal penetrance16 and hence the combination of isogenic and nonisogenic screens is commonly used to identify real, highly penetrant, synthetic lethal effects17. For RB1, we confirmed the validity of the synthetic lethal genes identified via nonisogenic screens (DepMap) by carrying out a CRISPR–Cas9 screen in isogenic RB1 wild-type and knockout cells (Fig. 2f). When we focused on RB1 synthetic lethal genes identified by both nonisogenic and isogenic screens, we found that these were also hyperexpressed in RB1-defective breast cancers (Fig. 2g and Supplementary Table 3).
We also acknowledged that some TSGs operate in discrete cancer histologies18 and thus carried out analyses where we considered only tumor cell lines (for DepMap analysis) and tumors (for transcriptomic analysis) from specific cancer histotypes. When we linked these histotype-specific synthetic lethal signatures to matched TCGA hyperexpression analysis, we identified 1,659 SLMs, of which 88 (representing 57 TSGs) exhibited significant overlap between synthetic lethal and hyperexpressed genes in histotype-matched cancers where the appropriate TSG was defective (SLM enrichment P < 0.05, Fisher’s exact test; Supplementary Fig. 3 and Supplementary Table 4). We did note that, similar to prior histotype-specific analyses of cancer dependencies19, these histotype-specific analyses used fewer tumor cell lines and tumors than a pan-cancer analysis, and thus were likely to be less statistically powered.
We also acknowledged that some TSG defects phenocopy others and, in some cases, TSG dysfunction is not evident by mutation or loss of expression of the TSG, but detected by other molecular features. For example, BRCA1 or BRCA2 deleterious mutations or BRCA1 promoter hypermethylation partially phenocopy each other and cause a homologous recombination defect (HRD) and other shared phenotypes, collectively termed BRCAness20. HRD in cancer can also be estimated in the absence of BRCA1 or BRCA2 defects by the presence of a genomic mutational signature or scar that reflects the mutagenic processes in play when HR is defective20,21. We therefore assessed whether synthetic lethal genes associated with either BRCA1 or BRCA2 loss were also hyperexpressed in HRD cancers22, regardless of the cause of HRD. To do this we used transcriptomic data from the SCAN-B breast cancer study where whole-genome sequencing (WGS) of 254 triple-negative breast cancers (TNBCs) has enabled the classification of each cancer by mutational scar with the HRDetect algorithm, as either HR-repair defective (59%, termed ‘HRDetect-high’, HRD+) or HR-repair proficient (‘HRDetect-low’, HRD−)21. In SCAN-B, the HRD+ group was additionally subclassified into cohorts with a deleterious BRCA1, BRCA2, PALB2, RAD51C or RAD51D mutation (HRD mutant), those with BRCA1 or RAD51C promoter hypermethylation (HRD methylated) or those HRD+ cancers that are HRDetecthigh but have neither HR gene mutation nor promoter hypermethylation (HRD other). From the 234 candidate BRCA1/2 synthetic lethal genes identified by DepMap CRISPR–Cas9 screens, 23% (n = 53, P = 0.027, Fisher’s exact test) showed elevated expression in HRD+ SCAN-B TNBCs (Fig. 3a and Extended Data Fig. 2a). Of BRCA1/2 synthetic lethal genes (n = 52), 22% also exhibited elevated expression in at least one of the three HRD+ subclassified groups (enrichment: PHRD mutant = 0.4, PHRD methylated = 0.07, PHRD other = 0.019, Fisher’s exact test; Fig. 3a). The hyperexpression of BRCA1/2 synthetic lethal genes in ‘HRD other’ cancers suggested that BRCA1/2 synthetic lethal interactions might also operate in HRD+ cancers in the absence of a known HR gene defect. Hyperexpression of BRCA1/2 synthetic lethal genes was also seen in HRD mutant, HRD methylated and HRD other subsets of the TCGA TNBC cohort, prostate cancer and high-grade serous ovarian cancer (Fig. 3a and Extended Data Fig. 2b). Conscious of the reliance on the DepMap dataset as the sole source of our synthetic lethal gene list, we corroborated the list of BRCA1/2 synthetic lethal genes by carrying out new synthetic lethal CRISPR–Cas9 screens in two isogenic systems with or without HR dysfunction: (1) Capan1 pancreatic tumor cells with either mutated or dysfunctional BRCA2 (Capan1Mutant) or a reverted or functional BRCA2 (Capan1Revertant); and (2) SUM149 breast tumor cells with either mutated or dysfunctional BRCA1 (SUM149Mutant) or a reverted or functional form (SUM149Revertant)23. Both BRCA1 and BRCA2 dysfunctional cell lines were sensitive to two different poly(ADP-ribose) polymerase (PARP) inhibitors, whereas their revertant versions were not (P < 0.001, two-way analysis of variance (ANOVA); Extended Data Fig. 2c–f). CRISPR–Cas9 screens in these cells reproduced the BRCA1/2 synthetic lethal effects identified from DepMap, including CIP2A, REV3L, APEX2, XRCC1, PARP1, POLQ, LIG1 and RHNO1 and Fanconi’s family genes (Fig. 3b,c, Extended Data Fig. 2g,h, Supplementary Fig. 4a–f and Supplementary Table 5), as did reanalysis of CRISPR–Cas9 screens in other BRCA1 or -2 isogenic models7 (Fig. 3d,e, Supplementary Fig. 4g–j and Supplementary Table 5). The combined list of BRCA1/2 synthetic lethal genes identified by both nonisogenic and isogenic screens were also significantly enriched in hyperexpressed genes in breast cancers with HR defects (Fig. 3f).
a, Box plots illustrating hyperexpressed genes (effect size >0) in different subgroups of patients with HRD. The dots indicate genes in corresponding SLMs (hyperexpression effect size ≥0.25 and FDR-adjusted P < 0.05, and synthetic lethal effect size ≤−0.25 and P < 0.01; P values calculated using unpaired, two-sided, moderated Student’s t-test) across SCAN-B TNBC, TCGA breast (BRCA) TNBC, TCGA prostate (PRAD) and TCGA ovarian (OV) cancer datasets. Example BRCA1/2 synthetic lethal effects are highlighted. The number of genes in SLMs and P value of significance for overlap between synthetic lethal and hyperexpressed genes are shown. Box plots show the first quartile, median and third quartile, with whiskers extended to points closest to Q1 − 1.5× the IQR and Q3 + 1.5× the IQR. b, Scatter plot illustrating BRCA1 synthetic lethal effects identified via isogenic SUM149 screen. Delta (Δ) represents the difference in viability (normalized z-score) between SUM149Mutant and SUM149Revertant cells. Key BRCA1/2 synthetic lethal genes are highlighted. c, Scatter plot illustrating BRCA2 synthetic lethal effects identified in isogenic Capan1 screen (details as in b). d,e, Scatter plots illustrating BRCA1 and BRCA2 synthetic lethal effects identified in two previously published7 BRCA1/2 isogenic screens in RPE1 (d) and DLD1 (e) cells. For visualization purposes, published CRISPRCount Analysis (CCA) scores had the penalty constant (10,000) added back to negative scores <−9,000 and the resulting CCA scores were transformed to z-scores and multiplied by −1. Key synthetic lethal genes are highlighted. f, Top, Venn diagram showing the overlap between DepMap-identified BRCA1/2 synthetic lethal genes and BRCA1/2 synthetic lethal genes identified in both in-house (either SUM149 or Capan1) and published (either RPE1 or DLD1) isogenic CRISPR–Cas9 screens. Bottom, Venn diagrams showing the overlap between BRCA1/2 synthetic lethal genes found in both DepMap and BRCA1/2 isogenic screens (18 genes) and hyperexpressed genes in HRD+ cancers from SCAN-B and TCGA TNBC datasets. Over-representation P values are reported for all Venn diagrams (Fisher’s exact test).
We also assessed whether candidate synthetic lethal genes that show compensatory upregulation in tumors are significantly enriched for penetrant synthetic lethal interactions. To understand this, we defined two SLMs: (1) BRCA1/2 synthetic lethal genes identified from DepMap that were also upregulated in cancers with HRD (‘SLM-up’) and (2) BRCA1/2 synthetic lethal genes from DepMap that were not upregulated in HRD cancers (‘SLM-not up’). Using gene set enrichment analysis (GSEA), we assessed the enrichment of these two SLMs against the ranked list of synthetic lethal genes identified from four different BRCA1/2 isogenic screens (Fig. 3b–e). SLM-up genes were significantly enriched in the list of synthetic lethal genes identified in the isogenic screens, whereas DepMap-identified BRCA1/2 synthetic lethal genes that were not upregulated (SLM-not up) were not (Extended Data Fig. 3a,b). We replicated the same phenomenon for RB1 synthetic lethal genes, using both DepMap and RB1 isogenic screens (Extended Data Fig. 3c). Consistent with this, the proportion of synthetic lethal genes hyperexpressed in HRD+ or RB1-defective cancers was higher for synthetic lethal genes identified by both isogenic and nonisogenic screens than it was for synthetic lethal genes identified by isogenic screens alone (Extended Data Fig. 3d–g).
Although HRD-related genomic scars reflect a history of HRD and predict response to DNA-damaging treatments21, they do not guarantee that a HR defect exists at the time of treatment24,25. We hypothesized that the elevated expression of BRCA1/2 synthetic lethal genes might exist because HRD is still in place; if this were true, a SLM consisting of genes synthetic lethal with BRCA1/2 should predict a better response to DNA-damaging therapy. Consistent with our hypothesis, a SYLVER-derived SLM consisting of 73 BRCA1/2 synthetic lethal genes identified from DepMap and our isogenic screens, which were hyperexpressed in HRDetect-high cancers (SLMHRD, Supplementary Table 6), was predominantly downregulated in HR-proficient Capan1 and SUM149 BRCA1/2-revertant cell lines when compared with their BRCA1/2-mutated, HR-defective, parental clones (P = 0.026, Wilcoxon’s rank-sum test; Fig. 4a and Supplementary Fig. 5a). Using SLMHRD and tumor transcriptomic profiles of adjuvant chemotherapy-treated (ACT) TNBC patients from the SCAN-B cohort, we also estimated SLMHRD scores and tertile trichotomized these into SLMHRD-low, SLMHRD-intermediate (Int.) and SLMHRD-high groups. The SLMHRD-high group was associated with better overall survival (OS), longer invasive disease-free survival (IDFS) and a greater distant relapse-free interval (DRFI) (OS: HRlow versus high = 0.17, 95% CI = 0.05–0.59, P = 0.005, Ptrend = 0.002; IDFS: HRlow versus high = 0.27, 95% CI = 0.11–0.67, P = 0.005, Ptrend = 0.003; and DRFI: HRlow versus high = 0.28, 95% CI = 0.09–0.86, P = 0.026, Ptrend = 0.016; Fig. 4b–d). SLMHRD remained an independent predictor of patient outcome when adjusted for HRDetect status (OS: Ptrend = 0.006; IDFS: Ptrend = 0.015, DRFS: Ptrend = 0.05) highlighting the added prognostic value of SLMHRD for identifying patients at highest risk of relapse. When we tested SLMHRD in an independent TNBC cohort (KCL26), SLMHRD was predictive of OS and distance metastasis-free survival (DMFS) (OS: HRlow versus high = 0.5, 95% CI = 0.25–0.98, P = 0.044, Ptrend = 0.039; and DMFS: HRlow versus high = 0.42, 95% CI = 0.18–0.99, P = 0.047, Ptrend = 0.042; Supplementary Fig. 6a,b). Next, we tested the predictive value of SLMHRD using TNBC gene expression profiles from the BrighTNess phase III clinical trial, where the addition of the HRD-targeting platinum salt carboplatin and the PARP inhibitor veliparib to standard neoadjuvant chemotherapy (NAC, paclitaxel) benefits a subset of patients27. SLMHRD scores were consistently higher in patients with pathological complete response (pCR) compared with those with residual disease burden (RD) across all three arms of the trial (Ppaclitaxel = 0.013, Ppaclitaxel + carbopltin = 0.023, Ppaclitaxel + carbopltin + veliparib = 8.6 × 10−4, Wilcoxon’s rank-sum test; Fig. 4e). We also performed additional evaluation of SLMHRD in an adaptive phase II neoadjuvant trial, I-SPY2 (ref. 28), focusing on the standard-of-care treatment arms of TNBC. In this trial, SLMHRD was associated with higher pCR rates when patients were treated with either paclitaxel alone or in combination with platinum and veliparib (Ppaclitaxel = 0.041, Ppaclitaxel + carbopltin + veliparib = 0.055, generalized linear model (GLM) adjusted for hormone receptor status; Fig. 4f). SLMHRD was also associated with a higher pCR rate in a contemporary setting where standard-of-care NAC (paclitaxel) was combined with the anti-programmed cell death protein 1 (PD-1) immunotherapy agent pembrolizumab (P = 0.02, GLM adjusted for hormone receptor status; Fig. 4f).
We extended the predictive assessment of SLMHRD to two additional breast cancer datasets treated with chemotherapy containing sequential taxane- and anthracycline-based regimen29. In both datasets, SLMHRD was associated with pCR (PHatzis-1 = 3.4 × 10−6, PHatzis-2 = 4.3 × 10−6, GLM adjusted for hormone receptor status; Fig. 4g). Finally, we tested SLMHRD in the paclitaxel arm of a phase II clinical trial where patients with breast cancer received neoadjuvant doxorubicin/cyclophosphamide, followed by chemotherapy agents paclitaxel or ixabepilone30. SLMHRD showed a trend of positive association with pCR (P = 0.103, GLM adjusted for hormone receptor status; Fig. 4h). In summary, these data not only confirm the prognostic and predictive power of SLMHRD, especially in the contemporary NAC setting, but also highlight crucial information possessed in the transcriptional state of HR-deficient breast cancers at the time of biopsy, irrespective of the historical state of a HRD scar. While SLMHRD represents a contemporary definition of HR deficiency, earlier studies relied on deleterious mutations in HR genes. We also therefore assessed a SLM consisting of 16 BRCA1/2 synthetic lethal genes identified from either DepMap or isogenic screens (SUM149 and Capan1) that were also hyperexpressed in cancers with alterations in HR genes (SLMBRCA1/2). SLMBRCA1/2 showed similar clinical performance to SLMHRD (Supplementary Fig. 7), highlighting the robustness of SLMHRD toward new datasets with all-encompassing drivers of HR defects such as BRCA1-promoter hypermethylation (seen in 40% of HR-defective TNBCs21). To benchmark the performance of SLMHRD systematically, we also compared its prognostic and predictive ability with three other gene expression-based DNA-damage response (DDR) deficiency signatures (CIN70 (ref. 31), BRCA1ness32 and RPS33). SLMHRD outperformed these signatures in 12 of 14 datasets assessed and was the best performing method overall (Extended Data Fig. 4a).
a, Evaluation of SLMHRD (73 genes) in RNA-seq of HR-deficient BRCA1/2 mutant and revertant clones of SUM149 and Capan1 cell lines. The SLMHRD score was estimated using GSVA. The difference between mutant and revertant clones was tested using Wilcoxon’s rank-sum test. b, Kaplan–Meier survival curves (OS) for SCAN-B TNBC patients treated with ACT. Groups were based on tertiles of the SLMHRD score. SLMHRD Int. and SLMHRD high groups were compared with the SLMHRD low group and Wald’s test P values are reported. The trend test was used for estimating global P values. The trend P value for a multivariable model adjusted for HRDetect prediction (low, Int., high) is also reported. c,d, Same as b but for IDFS (c) and DRFI (d). e, Evaluation of SLMHRD in a phase III clinical trial: BrighTNess. The difference in SLMHRD scores between patients with RD and pCR was tested using Wilcoxon’s rank-sum test. f, Evaluation of SLMHRD in I-SPY2, an adaptive phase II neoadjuvant clinical trial for breast cancer. The difference in SLMHRD of patients with RD and pCR was tested using GLMs adjusted for hormone receptor status with Wald’s test P values reported. g, Evaluation of SLMHRD in two neoadjuvant taxane-anthracycline chemotherapy-treated HER2− breast cancer cohorts: Hatzis-1 and Hatzis-2. The statistics are the same as f. h, Evaluating SLMHRD in the paclitaxel arm of a phase II clinical trial30. The statistics are the same as f. i, Heatmap illustrating the association of the SLMHRD score with HRDetect status and recurrent genomic driver alterations in the SCAN-B TNBC dataset, with a bar plot showing statistical significance of association (Kruskal–Wallis rank test or Wilcoxon’s rank-sum test). Amp , amplifications; MT, mutations. j, Box plots illustrating elevated SLMHRD in HRD+ patients harboring RB1 mutations in the SCAN-B TNBC dataset when compared with HRD+ patients without RB1 mutations (Wilcoxon’s rank-sum test). For all panels, the per-patient SLMHRD score was estimated using GSVA. All box plots show the first quartile, median and third quartile, with whiskers extended to points closest to Q1 − 1.5× the IQR and Q3 + 1.5× the IQR. The number of samples is reported in parentheses. All tests were two sided.
To delineate potential genomic correlates of SLMHRD, we examined its association with key driver mutations in TNBC. Given that the derivation of SLMHRD was in part influenced by HRDetect predictions, as expected, in SCAN-B TNBCs, SLMHRD correlated with HRDetect status (P = 1.6 × 10−9, Kruskal–Wallis rank test) and known genomic correlates of HRDetect21 (MYC and MCL1 amplifications and PTEN, TP53 and RB1 mutations, P < 0.05; Fig. 4i). Of these, RB1 mutations exhibited a strong correlation with SLMHRD (P = 3.7 × 10−11, Wilcoxon’s rank-sum test) and so we asked whether combined SLMHRD and RB1 status improved precision stratification of patients who are HRDetect-high (HRD+). SLMHRD in patients who are HRD+ was strongly associated with RB1 status (P = 7.3 × 10−7, Wilcoxon’s rank-sum test; Fig. 4j), suggesting that transcriptional activity measured by SLMHRD alone is predictive of RB1 mutant HRDetect-high status in the absence of WGS data.
Although synthetic lethality offers a promising framework for targeting TSG defects, its potential in cancer treatment has not yet been fully realized17. This could be because not all synthetic lethal interactions identified in preclinical models operate in the clinical disease, as either predictive biomarkers identifying those who would most benefit from a synthetic lethal treatment have not been developed or differentiating between highly and less penetrant synthetic lethal effects is sometimes difficult, making the selection of synthetic lethal targets for drug discovery or development an imprecise process. We show here that there is widespread evidence across multiple TSGs and cancer subtypes (including phenocopies) that the transcriptomic make-up of human cancers is partially shaped by patterns of TSG loss and reciprocal upregulation of synthetic lethal genes. This core biological observation could have several real-world implications. For example, TSG and synthetic lethal gene-buffering relationships that are evident in human cancers might be those that should be fast-tracked for drug discovery or development, especially as, in some contexts, these synthetic lethal interactions represent more penetrant effects. Our observations also suggest that predictive biomarkers of synthetic lethal treatments might be improved by integrating an assessment of the TSG defect with a measure of synthetic lethal gene hyperexpression or other signs of a buffering effect; such biomarkers would probably provide evidence that the synthetic lethal effect to be targeted is still operating in the tumor and that an anti-tumor response to synthetic lethal treatment is more likely. Our observations also imply that the hyperexpression of a set of synthetic lethal genes that are normally associated with a TSG deletion or deleterious mutation, in cancers where this TSG is neither deleted nor mutated, might indicate a functional defect in the TSG and the underlying biology of the disease. Although defining the functional status of a TSG as a predictive biomarker is often complex, defining the expression of synthetic lethal genes as a way of predicting sensitivity to their targeting in these ‘phenocopy’ cancers might be a much simpler and more robust way of extending the use of synthetic lethal treatments in cancer.
Methods
Preprocessing of molecular datasets
SCAN-B TNBC dataset
Normalized RNA sequencing (RNA-seq) data were downloaded from the Gene Expression Omnibus (GE), accession no. GSE96058. Data were exponentiated, a prior of 0.1 was removed and replaced with 1 and, subsequently, log2(transformed). For the SCAN-B TNBC subset, genes for which >75% of samples had zero mRNA abundance were removed from the dataset. HR status (defective, proficient), somatic and germline mutation status for BRCA1, BRCA2, PALB2, RAD51C and RAD51D and promoter methylation status for BRCA1 and RAD51C were curated from Staaf et al.21. For HRD analysis, we did not include an analysis of transcriptomes from patients with an intermediate HRDetect status (described as ‘HRDetect-Int.’21). A RB1 defective group was determined using RB1 mutation status or RB1 deep deletion curated from ref. 21 or RB1 gene expression z-score <−0.5. The remaining samples were classed as RB1 proficient.
TCGA datasets
Data for 32 TCGA cancer types—clinical, RNA-seq RSEM-normalized data (Illumina HiSeq rnaseqv2 level 3 RSEM), SNP6 copy number data (GISTIC level 4 log2(ratio)) and DNA methylation data (level 3 β values)—were downloaded from http://gdac.broadinstitute.org (release 28 January 2016). As a result of the absence of mutation data for the TCGA mesothelioma (MESO) dataset, this cancer type was not considered in the present study. The mRNA abundance data was log2(transformed) after adding a prior of 1 and quantile normalized. For each dataset, genes for which >75% of samples had zero mRNA abundance were removed from the dataset. Gene-level deletions were identified as those with log2(ratio) < −1. Gene-level damaging mutations were curated using the cBioPortal’s QQL: TRUNC_GERMLINE TRUNC NONSENSE NONSTART NONSTOP FRAMESHIFT, dated: 1 June 2023 from cbioportal.org. HR status (defective, proficient) for TCGA BRCA and OV samples was curated from Takaya et al.34 whereas, for TCGA PRAD and PAAD samples, HR status was curated from Knijnenburg et al.35. The HRD scar score for TCGA dataset was dichotomized using published thresholds: ≥42 = HR defective, <42 = HR proficient, except for TCGA OV where a threshold of ≥63 was used, as justified in a previously published study34. Germline BRCA1, BRCA2, PALB2, RAD51C and RAD51D status was curated from Huang et al.36, limiting to pathogenic and probably pathogenic variants. Promoter methylation status for BRCA1 and RAD51C was inferred from processed data available at http://gdac.broadinstitute.org. The probe in the promoter region with the strongest inverse correlation with gene expression was selected as the representative probe for methylation status. Samples with methylation β > 0.3 were considered to be hypermethylated.
BrighTNess dataset
Normalized RNA-seq and patient annotation data were downloaded from the GEO, accession no. GSE164458.
KCL, Hatzis and Horak datasets
Normalized microarray dataset and clinical information from the KCL dataset was requested from the authors of ref. 26. Raw microarray dataset and clinical information for Hatzis-1, Hatzis-2 and Horak datasets were downloaded from the GEO, accession nos. GSE25055, GSE25065 and GSE41998, respectively. Raw data were normalized using justRMA from R package affy (v.1.64.0), followed by probe-to-gene annotation mapping using the brain-array customized CDF R packages hsentrezgcdf (v.18 for Hatzis-1 and Hatzis-2 and v.25 for Horak)37.
I-SPY2 trial data
Normalized mRNA abundance data was downloaded from the GEO, accession no. GSE194040. Clinical data were downloaded from Wolf et al.28.
All patient studies have been previously published and their respective ethical approval is described in cited publications.
SYLVER
SYLVER was implemented in R programming language. The methodology and all associated data are freely available at https://software.icr.ac.uk/app/sylver. Data and analyses performed by SYLVER (Fig. 1) are described below.
SYLVER step 1 (cancer gene census, TSGs): TSGs were identified using the COSMIC CGC portal (https://cancer.sanger.ac.uk/census), data downloaded on 16 May 2023. We limited the list to 193 TSGs that were annotated as Tier = 1, Hallmark = Yes and role in cancer defined as: ‘TSG’, ‘TSG, fusion’, ‘oncogene, TSG’ and ‘oncogene, TSG, fusion’.
SYLVER step 2 (identification of synthetic lethal genes for TSGs by integrating CRISPR–Cas9 and molecular profiles of cancer cell lines): genome-wide CRISPR–Cas9 screen data from tumor cell lines were downloaded from the DepMap portal (depmap.org), DepMap v.23Q2. Mutational, copy number and gene expression profiles were also downloaded from the DepMap portal. In addition, copy number profiles for cell lines were also downloaded from the cBioPortal (cbioportal.org), data downloaded on 1 June 2023. For each TSG, the defective group was defined as cell lines harboring damaging mutation or copy number deletion or a structural defect (fusion) or gene expression z-score <−2. The remainder of the cell lines were assigned to a proficient group. For each TSG, two-class comparison of the genome-wide CRISPR–Cas9 screen data was performed between these two groups (defective and proficient) using linear regression (function: cdsrmodels::run_lm_stats_limma) as implemented by the Broad Institute’s Cancer Data Science team for these datasets (R package: cdsrmodels, v.0.1.0; R v.4.1.0), where the continuous gene effect viability scores were treated as the dependent variable. Synthetic lethal effects were identified as those with standardized effect size (Cohen’s d with Hedges’ g correction for unbalanced group size) ≤−0.25 and nominal P < 0.01. Of the 193 TSGs, PTK6 did not yield any synthetic lethal genes (P < 0.01) and therefore this gene was not considered for subsequent analyses by SYLVER. For CDH1 as a representative TSG operating in a selected group of cancer histologies, CDH1-defective and -proficient groups were defined in cell lines from urinary tract, breast, cervix, bile duct, gastric, lung, ovary, adrenal cortex, central nervous system and uterus. To represent multiple routes to a common phenotype use case of HRD+ cancers, BRCA1/2-defective group was curated as CAPAN1, COV362, DOTC24510, HCC1395, ICC15, JHOS2, JHOS4, MDAMB436, SUM1315MO2, SUM149PT and UWB1.289, whereas the remaining cell lines were assigned to a BRCA1/2-proficient group. Lineage-specific analyses were performed using the cell line to patient cancer-type map described in Supplementary Table 4.
SYLVER step 3 (Identification of hyperexpressed genes of TSGs by integrating molecular profiles of tumors): for each TSG, in each TCGA cancer type, patients were assigned to defective and proficient groups. For a given TSG in a given cancer type, the defective group comprised patients harboring a damaging mutation (curated using the cBioPortal’s QQL: TRUNC_GERMLINE TRUNC NONSENSE NONSTART NONSTOP FRAMESHIFT, dated 1 June 2023) or copy number deletion or gene expression z-score <−2. The remaining patients were assigned to the proficient group. Two-class comparison of gene expression profiles between the defective and proficient groups was performed using R package limma (v.3.42.2; R v.3.6.0), where the continuous gene expression profile was treated as the dependent variable. Hyperexpressed genes were identified as those with standardized effect size (Cohen’s d with Hedges’ g correction for unbalanced group size) ≥0.25 and false discovery rate (FDR)-adjusted P < 0.05. For TCGA TNBC, the RB1-defective group was determined using RB1 mutation status (damaging) or RB1 deletion or RB1 gene expression z-score <−0.5. Remaining samples were classed as RB1 proficient. This exception in difference in RB1 gene expression z-score threshold between TCGA TNBC (−0.5) and other TCGA datasets (−2) was motivated by previously reported fractions of RB1-defective TNBCs (15–40%). For BRCA1/2 and RB1 analyses in the SCAN-B TNBC dataset, the definition of defective and proficient groups is explained in the data-processing section for the SCAN-B TNBC dataset. Similarly, for BRCA1/2 in selected TCGA datasets (TNBC, PRAD, PAAD and OV), the definition of HR-based defective and proficient groups is explained in the data-processing section of TCGA datasets.
SYLVER step 4 (over-representation analysis of CRISPR–Cas9-based synthetic lethal genes in hyperexpressed genes from patient studies): enrichment analysis of CRISPR–Cas9-derived synthetic lethal genes in hyperexpressed genes from human cancers was performed using R function phyper() testing for over-representation (Fisher’s exact test). The overlap between the synthetic lethal and hyperexpressed genes was referred to as SLM.
SYLVER step 5 (assessment of SLM in patient studies): gene expression profiles of SLM genes in patient studies were summarized using gene set variation analysis (GSVA). The resulting scores (or binned groups) were tested for association with patient outcome using Cox’s proportional hazards model, Wilcoxon’s rank-sum test or GLMs adjusted for hormone receptor status, as appropriate.
Statistics and reproducibility
All analyses were performed in R statistical programming environment (v.3.6.0, except where stated as v.4.1.0). No data were excluded from the analyses unless stated otherwise.
GSEA
Enrichment of gene sets in isogenic RB1 and BRCA1/2 CRISPR–Cas9 screens was tested by ranking the genes on signed δ (isogenic—proficient) of normZ viability scores and subsequently analyzed with Fast Gene Set Enrichment Analysis (FGSEA) R package fgsea (v.1.12.0), R (v.3.6.0). For published isogenic BRCA1/2 screens7, processed CRISPRCount Analysis (CCA) scores were transformed by adding back the penalty constant of 10,000 to negative scores <−9,000, and the resulting CCA scores were transformed to z-scores and multiplied by −1. This transformation ensured that the direction of ranks was comparable to z-scores used for in-house isogenic screens. Enrichment P values were estimated using 10,000 permutations. ‘SLM-not up’ gene sets were defined as DepMap-defined synthetic lethal genes with hyperexpression effect size <0 and FDR-adjusted P < 0.05.
SUM149 and Capan1 genome-wide CRISPR–Cas9 screens
SUM149Mutant and SUM149Revertant cells23 were maintained in Ham’s F-12 medium supplemented with 5% fetal bovine serum (FBS), 10 μg ml−1 of insulin and 1 μg ml−1 of hydrocortisone. Capan1Mutant and Capan1Revertant cells23 were maintained in Iscove’s modified Dulbecco’s medium supplemented with 20% FBS. To confirm PARP inhibitor (PARPi) sensitivity, SUM149 or CAPAN1 cells were seeded into a 384-well plate at a density of 500 cells per well. After 24 h, cells were exposed to increasing concentrations of olaparib (Selleckchem) or AZD5305 (MedChemExpress), diluted in dimethyl sulfoxide, using an Echo 550 liquid handler (Labcyte). Cells were incubated with the inhibitor for 7 d, after which cell viability was estimated using CellTitre-Glo reagent (Promega). Dose–response survival curves illustrating the cell inhibitory effects of olaparib and AZD5305 were compared using two-way ANOVA in GraphPad Prism (v.10.3.0). Genome-wide CRISPR–Cas9 screens were carried out in Cas9+ (SUM149) or dCas9-KRAB+ (Capan1) derivatives using previously described single guide (sg)RNA libraries (SUM149: Yusa, Capan1: Weissman), as previously described38. Preprocessing and quantification of CRISPR screen data were performed as previously detailed38. For prioritization of genes, we used normZ, which aggregates sgRNA z-scores to gene level by normalizing to the number of sgRNAs as detailed here39. To prioritize high-confidence synthetic lethal genes in SUM149 and Capan1 screens, the following filters were applied:
MCF10A genome-wide CRISPR–Cas9 screens
Genome-wide CRISPR–Cas9 screens of isogenic MCF10ATP53−/− breast epithelial cell line with and without RB1−/− were performed using a previously published sgRNA library (Yusa, Sanger), as previously described38. To generate RB1 mutant cells, MCF10ATP53−/− cells (Horizon Discovery) were transfected with recombinant Cas9 complexed with a synthetic sgRNA (Supplementary Table 7) as described previously40. After single-cell isolation and expansion, mutant clones were identified by Sanger sequencing using primers specific for RB1 exon 3 (Supplementary Table 7). Western blot detection of RB1 was carried out to confirm loss of RB1 expression in an RB1 mutant daughter clone MCF10ATP53−/−RB1−/− clone 2.5 (RB1 c.333_336delTGAG;c.333delT). Preprocessing and quantification of CRISPR screen data was performed as previously described38. For the prioritization of candidate genes, gene-level sgRNA z-scores were subjected to the following filters, which were guided by the rank of key synthetic lethal vulnerabilities of RB1, that is, CDK2 and SKP2:
SUM149 and Capan1 RNA-seq
BRCA1 mutant and revertant clones from SUM149 and BRCA2 mutant and revertant clones from Capan1 were isolated. Three biological replicates for each condition (12 samples in total) were generated and strand-specific libraries were created using the NEBNext Ultra Directional RNA library prep kit (total RNA). Sequencing was performed on an Illumina HiSeq 2500 using PE 75× cycles v.1.5 chemistry, to achieve a minimum coverage of 30 million reads per sample. RNA-seq profiling generated 35.4–49.5 million paired-end reads per sample. Library quality was assessed using FastQC (v.0.11.9), FastQ Screen (v.0.14.0)41 and MultiQC (v.1.9)42. Reads were trimmed using Trim Galore (v.0.6.6). Paired-end reads were aligned to the human reference genome GRCh38, using STAR (v.2.7.6a)43 with --quantMode GeneCounts and --twopassMode Basic alignment settings. GENCODE (v.22) was used for gene feature annotations. Genes with low expression were filtered out using edgeR’s function filterByExpr(). ENSEMBL gene identifiers were annotated with HUGO gene symbols using the R package org.Hs.eg.db (v.3.10.0).
All cell lines were authenticated using STR profiling with the Geneprint10 Kit (Promega) and cell lines were tested monthly for Mycoplasma infection.
Benchmarking SLMHRD
Prognostic and predictive performance of SLMHRD was compared with three previously published RNA-based signatures of DDR deficiency: CIN70 (ref. 31), BRCA1ness32 and RPS33. CIN70 was implemented as the average of standardized gene expression profiles of signature genes. BRCA1ness was implemented using the singular value decomposition (SVD), where standardized gene expression profiles were used to calculate the first eigenvector for the signature genes and, subsequently, used to calculate SVD. The ranked probability score (RPS) was calculated using the median centered gene expression profiles followed by the sum of RPS signature genes, multiplied by −1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
TSG-specific synthetic lethal gene signatures, hyperexpressed gene signatures and SLMs are available to browse via the R shiny web application freely available at https://software.icr.ac.uk/app/sylver. These data are also available via Zenodo at https://doi.org/10.5281/zenodo.14447480 (ref. 44). Processed data (gene-level normZ viability scores) of RB1 and BRCA1/2 isogenic screens are available via Zenodo at https://doi.org/10.5281/zenodo.14178444 (ref. 45). Raw RNA-seq data for Capan1 and SUM149 BRCA1/2 mutant and revertant clones have been deposited in the Sequence Read Archive (SRA) under accession no. PRJNA1098629. SCAN-B RNA-seq data were downloaded from the GEO, accession no. GSE96058 (ref. 21). TCGA data were downloaded from http://gdac.broadinstitute.org (release 28 January 2016) and cBioPortal. BrighTNess RNA-seq data were downloaded from the GEO, accession no. GSE164458 (ref. 27). Hatzis datasets were downloaded from the GEO, accession nos. GSE25055 and GSE25065 (ref. 29). The Horak dataset was downloaded from the GEO, accession no. GSE41998 (ref. 30). The I-SPY2 dataset was downloaded from the GEO, accession no. GSE194040 (ref. 28). DepMap cell lines datasets were downloaded from depmap.org (v.23Q2) and cBioPortal. Source data are provided with this paper.
Code availability
SYLVER and analysis code was implemented with basic R (v.3.6.0) functionality using R packages affy (v.1.64.0), hsentrezgcdf (v.18), cdsrmodels (v.0.1.0; R v.4.1.0), limma (v.3.42.2), effsize (v.0.8.1), magrittr (v.2.0.1), tidyverse (v.1.3.1), fgsea (v.1.12.0), ggplots (v.2_3.3.5), ggrepel (v.0.9.1), org.Hs.eg.db (v.3.10.0), VennDiagram (v.1.6.20), survival (v.3.2-11) and GSVA (v.1.34.0). SYLVER’s customized code is freely available via Zenodo at https://doi.org/10.5281/zenodo.14685952 (ref. 46). For details, see relevant sections in Methods.
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Findlay, G. M. et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature 562, 217–222 (2018).
Ceccaldi, R. et al. Homologous-recombination-deficient tumours are dependent on Poltheta-mediated repair. Nature 518, 258–262 (2015).
Mateos-Gomez, P. A. et al. Mammalian polymerase theta promotes alternative NHEJ and suppresses recombination. Nature 518, 254–257 (2015).
Zatreanu, D. et al. Poltheta inhibitors elicit BRCA-gene synthetic lethality and target PARP inhibitor resistance. Nat. Commun. 12, 3636 (2021).
Zhou, J. et al. A first-in-class polymerase theta inhibitor selectively targets homologous-recombination-deficient tumors. Nat. Cancer 2, 598–610 (2021).
Adam, S. et al. The CIP2A-TOPBP1 axis safeguards chromosome stability and is a synthetic lethal target for BRCA-mutated cancer. Nat. Cancer 2, 1357–1371 (2021).
Behan, F. M. et al. Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens. Nature 568, 511–516 (2019).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Giaever, G. & Nislow, C. The yeast deletion collection: a decade of functional genomics. Genetics 197, 451–465 (2014).
Hart, G. T., Lee, I. & Marcotte, E. R. A high-accuracy consensus map of yeast protein complexes reveals modular nature of gene essentiality. BMC Bioinf. 8, 236 (2007).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR-Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Wang, T. et al. Gene essentiality profiling reveals gene networks and synthetic lethal interactions with oncogenic ras. Cell 168, 890–903 e15 (2017).
Sondka, Z. et al. The COSMIC Cancer Gene Census: describing genetic dysfunction across all human cancers. Nat. Rev. Cancer 18, 696–705 (2018).
Pacheco-Barcia, V. et al. The homologous recombination deficiency scar in advanced cancer: agnostic targeting of damaged DNA repair. Cancers (Basel) 14, 2950 (2022).
Ryan, C. J., Bajrami, I. & Lord, C. J. Synthetic lethality and cancer—penetrance as the major barrier. Trends Cancer 4, 671–683 (2018).
Ryan, C. J., Devakumar, L. P. S., Pettitt, S. J. & Lord, C. J. Complex synthetic lethality in cancer. Nat. Genet. 55, 2039–2048 (2023).
Burandt, E. et al. E-Cadherin expression in human tumors: a tissue microarray study on 10,851 tumors. Biomark Res 9, 44 (2021).
Campbell, J. et al. Large-scale profiling of kinase dependencies in cancer cell lines. Cell Rep. 14, 2490–2501 (2016).
Lord, C. J. & Ashworth, A. BRCAness revisited. Nat. Rev. Cancer 16, 110–120 (2016).
Staaf, J. et al. Whole-genome sequencing of triple-negative breast cancers in a population-based clinical study. Nat. Med. 25, 1526–1533 (2019).
Lord, C. J. & Ashworth, A. PARP inhibitors: synthetic lethality in the clinic. Science 355, 1152–1158 (2017).
Drean, A. et al. Modeling therapy resistance in BRCA1/2-mutant cancers. Mol. Cancer Ther. 16, 2022–2034 (2017).
Baxter, J. S., Zatreanu, D., Pettitt, S. J. & Lord, C. J. Resistance to DNA repair inhibitors in cancer. Mol. Oncol. 16, 3811–3827 (2022).
Harvey-Jones, E. et al. Longitudinal profiling identifies co-occurring BRCA1/2 reversions, TP53BP1, RIF1 and PAXIP1 mutations in PARP inhibitor-resistant advanced breast cancer. Ann. Oncol. 35, 364–380 (2024).
Watkins, J. et al. Genomic complexity profiling reveals that HORMAD1 overexpression contributes to homologous recombination deficiency in triple-negative breast cancers. Cancer Discov. 5, 488–505 (2015).
Filho, O. M. et al. Association of immunophenotype with pathologic complete response to neoadjuvant chemotherapy for triple-negative breast cancer: a secondary analysis of the BrighTNess Phase 3 randomized clinical trial. JAMA Oncol. 7, 603–608 (2021).
Wolf, D. M. et al. Redefining breast cancer subtypes to guide treatment prioritization and maximize response: predictive biomarkers across 10 cancer therapies. Cancer Cell 40, 609–623 e6 (2022).
Hatzis, C. et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA 305, 1873–1881 (2011).
Horak, C. E. et al. Biomarker analysis of neoadjuvant doxorubicin/cyclophosphamide followed by ixabepilone or paclitaxel in early-stage breast cancer. Clin. Cancer Res. 19, 1587–1595 (2013).
Carter, S. L., Eklund, A. C., Kohane, I. S., Harris, L. N. & Szallasi, Z. A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat. Genet. 38, 1043–1048 (2006).
Severson, T. M. et al. The BRCA1ness signature is associated significantly with response to PARP inhibitor treatment versus control in the I-SPY 2 randomized neoadjuvant setting. Breast Cancer Res. 19, 99 (2017).
Pitroda, S. P. et al. DNA repair pathway gene expression score correlates with repair proficiency and tumor sensitivity to chemotherapy. Sci. Transl. Med. 6, 229ra42 (2014).
Takaya, H., Nakai, H., Takamatsu, S., Mandai, M. & Matsumura, N. Homologous recombination deficiency status-based classification of high-grade serous ovarian carcinoma. Sci. Rep. 10, 2757 (2020).
Knijnenburg, T. A. et al. Genomic and molecular landscape of DNA damage repair deficiency across the cancer genome atlas. Cell Rep. 23, 239–254.e6 (2018).
Huang, K. L. et al. Pathogenic germline variants in 10,389 adult cancers. Cell 173, 355–370.e14 (2018).
Dai, M. et al. Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res. 33, e175 (2005).
Llorca-Cardenosa, M. J. et al. SMG8/SMG9 heterodimer loss modulates SMG1 kinase to drive ATR inhibitor resistance. Cancer Res. 82, 3962–3973 (2022).
Colic, M. et al. Identifying chemogenetic interactions from CRISPR screens with drugZ. Genome Med. 11, 52 (2019).
Kim, S., Kim, D., Cho, S. W., Kim, J. & Kim, J. S. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 24, 1012–1019 (2014).
Wingett, S. W. & Andrews, S. FastQ screen: a tool for multi-genome mapping and quality control. F1000Research 7, 1338 (2018).
Ewels, P., Magnusson, M., Lundin, S. & Kaller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Brough, R., Madera, S., Lord, C. & Haider, S. Dataset for study: The transcriptomic architecture of common cancers reflects synthetic lethal interactions. Zenodo https://doi.org/10.5281/zenodo.14447480 (2024).
Brough, R., Lord, C. & Haider, S. CRISPR-Cas9 dataset of isogenic cell lines. Zenodo https://doi.org/10.5281/zenodo.14178444 (2024).
Haider, S. & Lord, C. Custom code for study: the transcriptomic architecture of common cancers reflects synthetic lethal interactions. Zenodo https://doi.org/10.5281/zenodo.14685952 (2025).
Acknowledgements
We thank S. N. Zainal (University of Cambridge), C. Ryan (University College Dublin) and V. Serra (Valle d’Hebron Institute of Oncology) for helpful discussions. We also thank R. Alcraft from the Institute of Cancer Research’s (ICR’s) research software engineering team for supporting the deployment of SYLVER’s web app. This work was funded by Programme Funding to the Breast Cancer Now Toby Robins Research Centre (grant no. CTR-Q5-Y5 to C.J.L., S.J.P. and A.N.J.T.) and Programme Funding from CRUK (grant no. DRCRPGNov21y100001 to C.J.L., S.J.P. and A.N.J.T.). We thank Breast Cancer Now, working in partnership with Walk the Walk, who funded the work of S.H. The results shown here are in whole or part based on data generated by the TCGA research network: https://www.cancer.gov/tcga. This work represents independent research supported by the National Institute for Health Research (NIHR) Biomedical Research Centre at The Royal Marsden NHS Foundation Trust and the ICR, London. The views expressed are those of the author(s) and not necessarily those of the NIHR or the Department of Health and Social Care.
Author information
Authors and Affiliations
Contributions
S.H. and C.J.L. were responsible for designing the study, carrying out the analysis and writing the report, with input from R.B., S.J.P. and A.N.J.T. R.B. carried out isogenic CRISPR–Cas9 screens with support from A.W. S.H., A.G., R.B., J.A. and J.I. analyzed isogenic CRISPR–Cas9 screens. S.M. developed SYLVER web application. All authors contributed to the final draft and review of the report.
Corresponding authors
Ethics declarations
Competing interests
C.J.L. makes the following disclosures: he receives and/or has received research funding from AstraZeneca, Merck KGaA and Artios; received consultancy, SAB membership or honoraria payments from Syncona, Sun Pharma, Gerson Lehrman Group, Merck KGaA, Vertex, AstraZeneca, Tango, 3rd Rock, Ono Pharma, Artios, Abingworth, Tesselate, Dark Blue Therapeutics, Pontifax, Astex, Neophore, Glaxo Smith Kline Dawn Bioventures, Blacksmith Medicines and FoRx; has stock in Tango, Ovibio, Hysplex, Tesselate and Ariceum. He is also a named inventor on patents describing the use of DNA-repair inhibitors, including ATR inhibitors, and stands to gain from their development and use as part of the ICR ‘Rewards to Inventors’ scheme and also reports benefits from this scheme associated with patents for PARP inhibitors, paid into C.J.L.’s personal account and research accounts at the Institute of Cancer Research. A.N.J.T. is or has been a consultant for AstraZeneca, Merck KGaA, Artios, Pfizer, Vertex, GE Healthcare, Inbiomotion, Prime Oncology, Medscape Education, EMPartners, VJ Oncology, Gilead and MD Anderson Cancer Center, has received grant or research support from AstraZeneca, Myriad, Medivation and Merck KGaA, is a stockholder in Inbiomotion, is also a named inventor on patents describing the use of DNA-repair inhibitors, and stands to gain from their development and use as part of the ICR ‘Rewards to Inventors’ scheme and also reports benefits from this scheme associated with patents for PARP inhibitors paid to A.N.J.T.’s research accounts at the ICR. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Kris Wood and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Pan-cancer summary of synthetic lethal metagenes.
a. Heatmap indicating size of SLMs i.e overlap between the synthetic lethal genes identified from pan-cancer analysis of DepMap’s CRISPR-Cas9 cell lines and hyperexpressed genes identified from clinical samples in 32 TCGA datasets. Numbers in each cell indicate the overlap. The background colour of each cell indicates the significance of the observed overlap (P < 0.05, Fisher’s exact test) when compared to random expectation. Cells with white background indicate cancer types where analysis of hyperexpressed genes was not possible due to less than 3 samples in TSG defective or proficient groups. Of the 192 TSGs tested, heatmap shows data for those TSGs where at least one cancer type showed significant overlap (P < 0.05, red background).
Extended Data Fig. 2 Synthetic lethal metagene of HR deficiency and its validation in isogenic screens.
a, b. Venn diagrams showing the overlap between DepMap-identified BRCA1/2 synthetic lethal genes (a) and hyperexpressed genes in HRD+ cancers from SCAN-B and TCGA TNBC datasets (b) (Over-representation P values are reported, Fisher’s exact test). c, d. Dose response survival curves illustrating the cell inhibitory effects of Olaparib (c) and AZD5305 (d) in isogenic BRCA1 mutant (mutant/HR defective) and BRCA1 revertant (HR proficient) SUM149 cells. BRCA1 mutant cells demonstrated a profound sensitivity to both drugs compared to BRCA1 revertant cells (n = 9 biological replicates, two-way ANOVA). Error bars show standard error of the mean. e, f. Dose response survival curves illustrating the cell inhibitory effects of Olaparib (e) and AZD5305 (f) in isogenic BRCA2 mutant (mutant/HR defective) and BRCA2 revertant (HR proficient) Capan1 cells. BRCA2 mutant cells demonstrated a profound sensitivity to both drugs compared to BRCA2 revertant cells (n = 12 biological replicates, two-way ANOVA). Error bars show standard error of the mean. g. Scatter plot illustrating correlation between SUM149Parental/Mutant and SUM149Revertant screens. Example genes that lose BRCA1 synthetic lethality in revertant cells are highlighted. Pearson correlation coefficient and P value are shown. h. Scatter plot illustrating correlation between Capan1Parental/Mutant and Capan1Revertant screens. Example genes that lose BRCA2 synthetic lethality in revertant cells are highlighted. Pearson correlation coefficient and P value are shown.
Extended Data Fig. 3 Evaluating DepMap-identified synthetic lethal metagenes in isogenic screens.
a, b. Summary of Gene Set Enrichment Analysis (GSEA) P values for the gene sets defined as BRCA1/2 synthetic lethal in DepMap and hyperexpressed in HRD+ TNBCs (SLM-Up), and BRCA1/2 synthetic lethal in DepMap and not hyperexpressed in HRD+ TNBCs (SLM-Not Up). Two TNBC cohorts (SCAN-B and TCGA BRCA) were used to define hyperexpressed and not hyperexpressed genes. The gene sets were tested for enrichment in four isogenic BRCA1 or BRCA2 CRISPR-Cas9 screens (a: SUM149, Capan1; b: RPE1, DLD1) where viability scores were ranked from least viable to highly viable. Enrichment was considered significant (P < 0.05, dashed grey line) where the normalised enrichment scores (NES) were strongly negative as this highlighted enrichment of a given gene set amongst the less viable (i.e high-ranking) candidate synthetic lethal genes identified from isogenic screens. P values were calculated using permutation test (two-sided). c. Same as panels ‘a, b’ but for DepMap-derived RB1 synthetic lethal genes that were hyperexpressed in RB1 defective TNBCs (SLM-Up), and DepMap-derived RB1 synthetic lethal genes that were not hyperexpressed in RB1 defective TNBCs (SLM-Not Up). The gene sets were tested for enrichment in an isogenic RB1 CRISPR-Cas9 screen. d–g. Bar plots showing the proportion of synthetic lethal genes identified in both isogenic and corresponding DepMap analysis that were also hyperexpressed in patient tumours versus the proportion of synthetic lethal genes identified in isogenic screen only that were also hyperexpressed in patient tumours. Proportions were compared using proportion test with P values reported (one-sided). d. TSG = BRCA1, where isogenic screen data was from the SUM149Mutant and SUM149Revertant screen; e. TSG = BRCA2, where isogenic screen data was from the Capan1Mutant and Capan1Revertant screen; f. TSGs = BRCA1/2 where isogenic screen data and associated (previously published by Adam et al. (7)) synthetic lethal gene list was from the DLD1 BRCA2+/+ and BRCA2−/− screen, and RPE1 BRCA1+/+ and BRCA1−/− screen. g. TSG = RB1, where isogenic screen data was from a MCF10A TP53−/−, RB1+/+ and MCF10A TP53−/−, RB1−/− screen.
Extended Data Fig. 4 Benchmarking of SLMHRD.
a. Benchmarking of SLMHRD against three other RNA-based DNA damage response deficiency signatures in six independent studies (14 datasets). Heatmap showing significance of association between patients’ outcome data and signature score-derived patient risk groups (for prognostic models), or, difference in signature scores between clinical response following neoadjuvant chemotherapy (for predictive models; residual disease or pathological complete response). Significance is reported as trend P value for prognostic models and a P value from generalised linear models for predictive associations (models adjusted for hormone receptor status for I-SPY2, Hatzis and Horak datasets; two-sided Wald-test). Number in cells show the rank (across a row) of each signature compared to other signatures for each dataset. Cumulative ranking was calculated using the geometric mean of individual ranks and shown as a bar plot, with smallest geometric mean indicating best overall performance.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7 and Legends for Supplementary Tables 1–7.
Supplementary Tables 1–7 (download XLSX )
Supplementary Table 1 TSGs. List of TSGs analyzed. Supplementary Table 2 TCGA cancer types. List of TCGA cancer types analyzed. Supplementary Table 3 RB1 nonisogenic versus isogenic screens. Overlap between RB1 synthetic lethal genes identified from DepMap and RB1 isogenic CRISPR–Cas9 screen using MCF10A breast epithelial cell line. Supplementary Table 4 Cell lines to human cancers map. DepMap cell lines lineage to TCGA cancer type map. Supplementary Table 5 BRCA1/2 nonisogenic versus isogenic screens. Overlap between BRCA1/2 synthetic lethal genes identified from DepMap, and BRCA1 and BRCA2 isogenic CRISPR–Cas9 screens using SUM149, Capan1, RPE1 and DLD1 cell lines. For RPE1 and DLD1 screens, a combined listed of BRCA1 or -2 synthetic lethal genes as published in the original study7 was used. Supplementary Table 6 HRD gene signature. List of genes in SLMHRD. Supplementary Table 7 SgRNA and primer sequences. List of sgRNA and primers used in MCF10A genome-wide CRISPR–Cas9 screens.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data for all panels as tabs.
Source Data Fig. 3 (download XLSX )
Statistical source data for all panels as tabs.
Source Data Fig. 4 (download XLSX )
Statistical source data for all panels as tabs.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for all panels as tabs.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for all panels as tabs.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Haider, S., Brough, R., Madera, S. et al. The transcriptomic architecture of common cancers reflects synthetic lethal interactions. Nat Genet 57, 522–529 (2025). https://doi.org/10.1038/s41588-025-02108-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02108-2
This article is cited by
-
Death-ision: the link between cellular resilience and cancer resistance to treatments
Molecular Cancer (2025)
-
Autocrine interferon poisoning mediates ADAR1-dependent synthetic lethality in BRCA1/2-mutant cancers
Nature Communications (2025)