Abstract
In isocitrate dehydrogenase wildtype glioblastoma (GBM), cellular heterogeneity across and within tumors may drive therapeutic resistance. Here we analyzed 121 primary and recurrent GBM samples from 59 patients using single-nucleus RNA sequencing and bulk tumor DNA sequencing to characterize GBM transcriptional heterogeneity. First, GBMs can be classified by their broad cellular composition, encompassing malignant and nonmalignant cell types. Second, in each cell type we describe the diversity of cellular states and their pathway activation, particularly an expanded set of malignant cell states, including glial progenitor cell-like, neuronal-like and cilia-like. Third, the remaining variation between GBMs highlights three baseline gene expression programs. These three layers of heterogeneity are interrelated and partially associated with specific genetic aberrations, thereby defining three stereotypic GBM ecosystems. This work provides an unparalleled view of the multilayered transcriptional architecture of GBM. How this architecture evolves during disease progression is addressed in the companion manuscript by Spitzer et al.
Similar content being viewed by others
Main
Tumor heterogeneity is a hallmark of isocitrate dehydrogenase (IDH) wildtype (WT) glioblastoma (GBM)1. Different GBM samples have distinct genetic and transcriptomic profiles (intertumor heterogeneity)2 and even a single GBM specimen contains diverse malignant and nonmalignant cells (intratumor heterogeneity)3,4. This diversity endows malignant GBM cells with distinct functional properties5 and is thought to enable them to evade therapies, leading to progression and relapse6. Nonmalignant cells in the tumor microenvironment (TME) establish an ecosystem with complex interactions that also plays critical roles in shaping GBM biology and treatment response5,7,8,9.
Previous studies used single-cell RNA sequencing (scRNA-seq) to uncover gene expression heterogeneity within human gliomas3,4,10,11,12,13,14,15,16,17. These studies revealed that GBM malignant cells exhibit a variety of cellular states, including those resembling neurodevelopmental cell types. The frequency of these states in a particular tumor is influenced by genomic alterations and the TME7. However, previous studies had several limitations: (1) they were based on limited number of samples and cells and could still miss rare cellular states; (2) they relied on enzymatic tumor dissociation, which may deplete certain cell types and states, as evident by the scarcity of detected astrocytes, neurons and endothelial cells; (2) they addressed either heterogeneity across tumors (that is, tumor subtypes) or within tumors (that is, cell states) but did not develop an integrated model that explains tumor heterogeneity by a combination of tumor-level and cell-level features; (4) they lacked in-depth genetic characterization of the tumors; (5) they had limited clinical metadata; and (6) they were published by different groups, and leveraged diverse pipelines, analytics and classification schemes4,10,12,15,16.
To address these challenges, we created the GBM Cellular Analysis of Resistance and Evolution (CARE) consortium and profiled 121 IDH WT GBM samples using single-nucleus RNA sequencing (snRNA-seq) and bulk DNA sequencing, thereby establishing a multi-omic dataset of unprecedented size while eliminating the confounding effects of enzymatic tumor dissociation. This dataset is richly annotated with clinical data and contains primary and recurrent matched pairs. In this article, we describe the identification of new GBM cellular states, for both the malignant and TME compartments. Furthermore, we describe new analyses that distinguish the contributions of intratumor and intertumor heterogeneity, toward a more complete understanding of the multilayered transcriptional architecture of GBM. The main conclusions of this article are independent of the timing of sampling (primary or recurrent) and can be reached by only analyzing primary GBM specimens, yet we include the recurrent samples for more robust statistical analysis and to provide a broad view of the range of GBM tumors. In the accompanying study by Spitzer et al.18, we focus on characterizing the longitudinal evolution of GBM after standard-of-care therapy.
Results
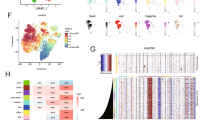
Our groups jointly generated an snRNA-seq dataset of 121 tumor samples from 59 patients spanning initial disease and recurrence (Fig. 1a; see Supplementary Table 1 for detailed clinical information). All tumors were diagnosed as ‘Glioblastoma, IDH WT’ based on the World Health Organization (WHO) 2021 classification, which was further confirmed using bulk DNA sequencing. We profiled all 121 samples with the droplet-based 10x Genomics platform, and also profiled a large subset of tumors (54 of 121) using plate-based full-length Smart-seq2 (ref. 19). Most samples (116 of 121) were also profiled using bulk whole-exome sequencing (WES) or whole-genome sequencing (WGS), enabling the integration of genetic information with single-nucleus transcriptomics (Fig. 1b and Supplementary Table 1). The frequency of common genetic alterations was consistent with previous studies of GBM20.
a, Scheme describing the workflow of our study. b, The proportion (%) of patients (n = 46) in the dataset in which a genetic aberration was detected is shown. Arm-level refers to whole-chromosome arm amplification or deletion. CNA refers to gene-level amplification or deletion. Single-nucleotide variant (SNV) refers to single-nucleotide mutations. c, Uniform manifold approximation and projection (UMAP) for dimension reduction based on gene expression of the 429,305 cells in our cohort colored according to the inferred cell type. d, Pie chart demonstrating the proportion (%) of each cell type in this dataset. amp, amplification; del, deletion.
In the 10x snRNA-seq dataset, 429,305 of 752,458 cells passed our stringent quality control (Methods) with 2,318 genes detected per cell on average (Extended Data Fig. 1a). Cells were classified as malignant (246,408 cells) or nonmalignant (182,897 cells) by inferring chromosomal copy number aberrations (CNAs) from gene expression data, as described previously13,21, but with additional steps to improve classification accuracy (Extended Data Fig. 1b–d). Clustering of all cells highlighted the diversity of malignant cells, including many small patient-specific clusters, while nonmalignant cells were intermixed across patients without apparent batch effects (Fig. 1c and Extended Data Fig. 2a,b). Nonmalignant cells were assigned to cell types using clustering and well-established signatures (Extended Data Fig. 2c)22,23,24,25,26. Notably, we detected neurons and astrocytes (Fig. 1c,d) that were typically not captured in scRNA-seq studies with enzymatic tissue dissociation. In the Smart-seq2 dataset, 10,062 of 11,136 cells passed our quality control and were similarly classified to malignant and nonmalignant cell types (Extended Data Fig. 2d). Overall, the fractions of cell types were similar between 10x Genomics and Smart-seq2 datasets (Extended Data Fig. 2e–g).
Revisiting malignant cellular states
We previously defined four principal cellular states in GBM based on 28 samples (7,930 cells) obtained at initial diagnosis4: neural-progenitor-like cells (NPC-like); oligodendrocyte-progenitor-like cells (OPC-like); astrocyte-like cells (AC-like); and mesenchymal-like cells (MES-like). While this robust classification scheme has been broadly adopted and validated by the community5,27,28,29, we sought to extend the atlas of GBM malignant cellular states by leveraging our much larger snRNA-seq dataset. First, nonnegative matrix factorization (NMF) was used to derive gene expression programs that vary within each tumor30. After removing programs suspected of low technical quality (for example, enrichment of ribosomal proteins or pseudogenes; Methods), we found ten groups of similar programs and defined a consensus meta-program (MP) for each of them, representing the malignant cellular states in our dataset (Fig. 2a, Extended Data Fig. 3a and Supplementary Table 2).
a, Jaccard similarity indices of the robust NMF programs (n = 707) based on their top 50 genes. Programs are ordered according to clustering and grouped into MPs (Methods). MP10 (stress1) and MP15 (stress2) were coalesced into a single stress state. MP9 (excitatory neuron (ExN)) and MP14 (neurogranin neuron) were coalesced into a single NEU-like state. The ‘cycling’ MP is separated to emphasize it as a feature that cells can have on top of their neurodevelopmental identity (and not as an independent state; Extended Data Fig. 3a). b, Gene expression scores of each cell for the different cellular states and the cell cycle. Cells are aggregated according to the state to which they were classified and are ordered within each group according to their cell cycle score. ‘Unresolved’ represents cells that could not be confidently classified to a particular state. c, Relative expression (centered log2) of the three new MPs discovered in this work. Each row represents a gene and each column a cell. d, Association between the malignant MPs (rows) and gene sets reflecting normal brain development and adult brain cell types (columns). Each cell in the heatmap represents the Pearson correlation coefficient between the gene expression scores of an MP and a normal brain gene set across the malignant cells in the dataset. e, Association between pathway-based (rows) and gene-based (columns) MPs. Each cell in the heatmap represents the Pearson correlation coefficient between the gene expression scores of the pathway-based and gene-based MPs. The colored squares represent the seven groups of PMPs and their associated MPs. f, Demonstration of the hybrid state concept. The rows represent genes, which are aggregated according to the MP to which they belong. The columns represent cells, which are aggregated according to their singular (AC, GPC) or hybrid (AC-GPC) state. The heatmap of genes’ centered log2 expression shows that hybrids express two MPs to the same extent, whereas singulars highly express a single MP. g, GBM cellular hierarchy model stemming from the high-frequency hybrid pairs. Each vertex represents a state; vertices are connected when they represent a high-frequency hybrid state. h, Two-dimensional representation of cellular states. Each quadrant reflects a cellular state and each dot represents a malignant cell; exact cell positions reflect their relative scores for the MPs, and their colors reflect the proportion of cells around them classified to the GPC state.
Overall, seven of the ten MPs largely recapitulate the patterns from ref. 4. One of these MPs represented the cell cycle and another five represented the previously defined cellular states: AC-like; OPC-like; NPC-like; MES-like; and hypoxia (previously denoted as MES2)4 (Fig. 2b). Another MP (stress) included genes associated with heat shock and unfolded protein response, which were previously included in the MES-like MP and associated with disordered DNA methylation12. Thus, high-resolution analysis divides MES-like states into three MPs that we termed MES-like, hypoxia and stress.
Although these seven MPs produce highly correlated cell scores as those from ref. 4 (Extended Data Fig. 3b), it is important to note that the exact sets of genes that define the MPs here and in ref. 4 differ (Extended Data Fig. 3c), suggesting platform-dependent effects such that distinct genes are highlighted by scRNA-seq versus snRNA-seq (Extended Data Fig. 3d). We also compared these MPs to prior GBM cell-state classifications, further corroborating their identities (Extended Data Fig. 3e)31,32.
The first new MP contained NPC-like genes but was mostly associated with markers of more differentiated neurons (NRG1, NRG3, NRXN3, CNTNAP2) and synaptic activity (SYN3, SYT1; Fig. 2c) and was similar to the NEU (neuronal) pathway-based program reported previously15. Consistent with a differentiated NEU state, cycling cells were depleted in this program relative to NPC-like cells (Extended Data Fig. 4a). Comparison to the signatures of neural cell types in development and in adult human brains showed the highest similarity to L2–L3 excitatory neurons (P = 7.2−5, hypergeometric test; Fig. 2d); thus, we named this state NEU-like. This state is depleted in single-cell relative to single-nucleus data4, suggesting that it may be poorly captured by enzymatic tumor dissociation, possibly because of complex cellular morphology.
The second new MP included the tyrosine kinase genes EGFR and ALK, the transcription factor genes MEIS1, MEOX2, ETV1, ELOVL2 and KLHDC8, which were previously linked to glioma stemness33,34,35. Comparison to normal brain signatures showed the strongest association with a recently described signature of glial progenitor cells (GPCs)36; hence, we denoted it as GPC-like. Consistent with a progenitor state, the GPC-like MP was enriched with cycling cells (Extended Data Fig. 4a). Like the NEU-like state, GPC-like cells were depleted in single-cell relative to single-nucleus data4.
The third new MP mostly contained cilia genes, including 12 of 50 DNAH and CFAP family members; therefore, it was named cilia-like. This MP was specific to a small fraction of cells (~1.6%), underscoring the utility of our large cohort for identifying rare cellular states. Notably, all of those cells also expressed the AC-like MP, indicating that a subset of AC-like cells activate a cilia program. As expected, this MP was most enriched with external cilia signatures. Therefore, all MPs are enriched with corresponding cell-type-signatures from developing or adult human brains24,26,36 (Fig. 2d), supporting our annotations and suggesting that GBM states mirror developmental trajectories32.
Collectively, our MPs are reflective of cell identity (for example, GPC-like, AC-like, NEU-like) or cell activity (for example, response to hypoxia or stress). We also scored malignant cells for signatures of functional attributes5,37,38, which suggested that AC-like and MES-like cells had greater connectivity, while OPC-like, NPC-like and NEU-like cells had greater motility and invasion scores, while cilia-like and GPC-like cells were intermediate (Extended Data Fig. 4b).
The new MPs defined from 10x data were validated in Smart-seq2; they were still detected when analyzing only primary samples and were detected independently in specimens profiled by different labs and obtained from different medical centers (Extended Data Fig. 4c–f). Overall, our analysis expands our original malignant GBM MPs to ten, some of which represent more granular subsets of the states in ref. 4.
Consistent classification by pathway-based and gene-based analysis
As an orthogonal description of cellular states, we defined pathway-based MPs (PMPs) (Fig. 2e). We previously defined four PMPs, namely NEU, glycolytic/pluri-metabolic (GPM), mitochondrial (MTC) and proliferative/progenitor (PPR)15,39. To preserve a similar computational approach to the gene-expression-based MPs, we adapted the NMF-based algorithm to consider pathways instead of genes, including 3,015 pathway gene sets from the Molecular Signatures Database Hallmark, Gene Ontology (GO) and Reactome. This resulted in seven PMPs that recapitulated and extended the previously defined PMPs (Extended Data Fig. 5a–c).
We confirmed the existence of the NEU PMP, which correlated with the OPC-like, NPC-like and NEU-like MPs (Fig. 2e). We identified two distinct PMPs related to the previous GPM, one linked to glycolysis and hypoxia, and another closely resembling the MES-like state (denoted as GPM-MES). MTC-related and PPR-related PMPs also emerged from this analysis. We additionally identified two new PMPs, one similar to the AC-like and cilia-like MPs (AC-CIL), and another that is highly correlated with the GPC-like state, denoted as GPC/radial glia-like (RGL) (GPC-RGL). Overall, the association of PMPs and MPs provides further functional characterization to each state, supporting the identification and biological interpretation of the new malignant cell states.
Hybrid states reveal putative state transitions
Most malignant cells in our dataset were classified into a single state, yet roughly 20% of the cells could be classified into multiple states, with a limited difference between the two highest state scores (for an example, see Fig. 2f). These hybrid cells did not have a high number of detected genes as would be expected from technical doublets (two nuclei occupying the same droplet; Extended Data Fig. 5d). Moreover, hybrid cells were not distributed as expected across all state pairs but rather were highly enriched for certain state pairs (Extended Data Fig. 5e). Thus, we reasoned that these may represent cells in transition between two states, and constructed a GBM model where each node represents a cellular state and each edge reflects a high-frequency hybrid state (Fig. 2g).
Although this analysis lacks evidence for directionality, the differentiated cellular states (NEU-like, AC-like, cilia-like, MES-like and hypoxia) were all at the edges, while the three progenitor-like and stem-like states (NPC-like, OPC-like and GPC-like) were all in the center of the model. Specifically, the GPC-like state was positioned at the center of three trajectories (NEU, AC-cilia, hypoxia-MES), perhaps suggesting that this population represents an early progenitor state with differentiation potential along multiple trajectories. Consistent with this possibility, cells having high GPC-like scores tended to exhibit intermediate expression levels of programs for the other cellular states (Fig. 2h).
Baseline expression patterns of intertumor heterogeneity
When assessing differences between samples, a dominant signal reflects variations in the abundance of shared cellular states. To highlight additional patterns of intertumor differences, we developed a cell-state-controlled approach in which we compared tumors only based on cells in corresponding states. First, each tumor sample was decomposed into subpopulations of cells representing seven common malignant states. For each subpopulation with a minimum of 25 cells in a particular sample, we then defined a state-specific pseudo-bulk (average) expression profile. Then, for each state, the pseudo-bulk profiles were compared across tumors, thereby eliminating state-specific signals and highlighting tumor-specific differences in that state. Principal component analysis (PCA) was applied to the pseudo-bulk profiles of each cell state, defining high and low signatures for the top three principal components (Fig. 3a).
a, Schematic of state-controlled analysis to define the BPs. b, Intertumor and intratumor variability of the BPs (n = 3) and MPs (that is, cell states, n = 7). Intertumor variability reflects the average variability measured between scores of different samples (defined as the average s.d. of program scores in each state across all states). Intratumor variability reflects the average variability measured between scores of same samples (defined as the average s.d. of program scores in each sample across all samples). Intertumor variability was significantly larger in BPs relative to MPs (mean score = 1.05 versus 0.65, P = 0.0005, one-sided t-test) whereas intratumor variability was significantly higher in MPs relative to BPs (mean score = 0.76 versus 0.62, P = 0.028, one-sided t-test). P values were not corrected for multiple testing. c, Intra-sample variability in BP scores across state-specific pseudo-bulk profiles. Each dot reflects a state-specific pseudo-bulk profile, is colored according to state and aggregated horizontally according to sample. The black horizontal lines connect pseudo-bulk profiles from the same sample as a visualization aid and have no other meaning. The black dots reflect the mean BP score of each sample and are connected using a line for visualization purposes. The dashed lines mark y = −1 and y = 1 to aid in visualization. The y axis shows the difference between the ECM and NEU BPs. d, Top, two-dimensional representation of the BPs. Each dot represents a specific sample (the mean score across the pseudo-bulks for each BP) and are colored according to the maximal score. The y axis reflect the BP-glial score, the x axis reflects the difference between the NEU and ECM BPs as follows: sign(NEU-ECM) × max(NEU, ECM). Bottom, limited intratumor variability in BP scores. Cells are colored according to the maximal BP score. Cellular states that reflects each quadrant are indicated on the right.
We found highly consistent signatures of intertumor variation across PCA analyses of different states. For example, signatures of variation across AC-like pseudo-bulk profiles were highly similar to signatures of variation across NPC-like pseudo-bulk profiles. Three sets of signatures were each identified in analyses of most states, reflecting patterns of intertumor heterogeneity that commonly influences several malignant states (Extended Data Fig. 6a). We denoted their consensus signatures as state-controlled baseline profiles (BPs), that is, consistent malignant gene expression differences between tumors that are not accounted by differences in tumor cell-state distribution.
We annotated BPs based on functional enrichment as NEU, glial and extracellular matrix (ECM) (Extended Data Fig. 6b and Supplementary Table 3). While BP annotations are reminiscent of cell-state MPs (that is, BP-NEU and NEU-like MP; BP-ECM and MES-like MP), we note that BPs and cell-state MPs reflected distinct sets of genes, such that MPs preferentially varied within tumors while BPs preferentially varied between tumors (Fig. 3b,c and Extended Data Fig. 6c) and remain consistent within a tumor (P = 2.3 × 10−9, Wilcoxon rank-sum test; Extended Data Fig. 6d). Accordingly, when we assigned tumors to BPs by averaging across the respective pseudo-bulks (Fig. 3d top; Extended Data Fig. 6e), we found that this classification tended to be consistent across different cellular states in the tumor (Fig. 3d bottom). In a tumor with high ECM BP, cells existed in multiple states but most tended to express high ECM, while in a tumor with high NEU BP most cells tended to express high NEU BP, even those in a MES-like state.
To examine whether BPs are consistent across regions of the same tumor, we leveraged a GBM spatial transcriptomics dataset that includes multi-sampled tumors37. We derived de novo BPs from the spatial transcriptomics data, resulting in three BPs that were highly correlated with the snRNA-seq-derived original BPs (Extended Data Fig. 7a,b). Sample pairs from the same tumor had significantly higher similarity in BP scores than sample pairs from different tumors (Extended Data Fig. 7c), supporting the consistency of BPs within tumors. Finally, we leveraged a subset of samples with both bulk RNA-seq and snRNA-seq data to compare the newly defined BPs to The Cancer Genome Atlas (TCGA) expression subtypes17,40. We found a significant association (Fisher’s exact test, P = 0.02) with the greatest overlap between the BP-ECM and the MES TCGA subtype (Extended Data Fig. 7d). Taken together, the three BPs reflect core intertumor transcriptional variation (that is, tumor-level phenotypes) that are partially independent of the variation in cellular states within tumors.
Interplay between TME, malignant states and BPs
We next investigated how TME heterogeneity may influence malignant BPs and cell-state abundance. To accomplish this, we applied our NMF framework to derive TME MPs for each nonmalignant cell type (n = 62 total MPs; Fig. 4a). We annotated TME MPs based on pathway enrichment and by scoring cells for published signatures (Extended Data Fig. 8a–d, Supplementary Table 4 and Methods). TME MPs were linked to diverse functions, for example, reflecting cortical layers (among excitatory neurons), immune phenotypes (microglia versus bone marrow-derived macrophages) and differentiation status (immature versus mature oligodendrocytes). A subset of TME MPs had high overlap with malignant cell MPs (for example, hypoxia and MES-like), suggesting a similar effect of microenvironmental niches on multiple cell types30 (Extended Data Fig. 8e).
a, Donut chart representing the number of cells per cell type analyzed to identify robust NMF MPs, labeled with the number of cells and MPs found in each cell type. The size of the donut represents the relative proportion of all TME cells. b, Selected TME NMF state composition of tumors assigned to each BPs. The statistical significance of TME composition between samples with assigned BP to ECM (n = 23), glial (n = 39) or NEU (n = 35) was computed using a two-sided Kruskal–Wallis test (n = 97). c, Interaction graph between malignant cell states and TME cell type abundance. Each node represents malignant cell-state (red) or TME cell type nodes (colored according to cell type as presented). The thickness and color of the edge indicate the strength of the Pearson correlation. d, Correlation heatmap for the Pearson correlation between the relative malignant cell-state proportions (rows) and relative TME cell-state abundance (columns), which show representative associations for the hypoxic environment. e, Ligand–receptor cross-talk graph. Each green circled node represents a malignant cell state versus nonmalignant cell type pair, while each squared node represents a ligand–receptor interaction; there is an edge if that cross-talk is present. The thickness and color of the edge indicate the percentage of tumors in which both malignant cell state and nonmalignant cell type are present and the cross-talk exist. Open ‘(’ and closed ‘)’ parentheses reflect multiple ligand or receptor interaction from the same family. Specifically, closed parentheses ‘)’ indicated multiple ligands interacting with the same receptor, and viceversa for closed ‘)’ parentheses, multiple receptors interacting with the same ligand. UPR, unfolded protein response.
We assigned all TME cells to their highest-scoring TME MP (Methods). TME MP abundances differed significantly across GBMs mapping to the three BPs, suggesting an association between TME and BPs (Fig. 4b). Similarly, TME MP abundances were associated with malignant MP abundances (Extended Data Fig. 9). Abundance of MES-like cells was associated with increased tumor-associated myeloid cells (TAMs) and monocyte-derived macrophages (MDMs)4,7,9, but also with endothelial cells (Fig. 4c). Abundance of NEU-like malignant cells was associated with neurons, suggesting that nonmalignant neurons may interact with malignant cells and shape their states. Abundance of GPC-like cells was positively correlated with TAM-microglia; abundance of OPC-like malignant cells correlated with nonmalignant OPCs.
Collectively, these results indicate that malignant state abundance is associated with diverse TME cell types. Clustering of all states revealed two major TME-malignant compartments that probably reflect tumor periphery and core regions41 (Extended Data Fig. 9). In particular, malignant hypoxia, stress and MES-like states were associated with inflammatory and hypoxic macrophages, reactive astrocytes, and inflammatory and angiogenic endothelial states (Fig. 4d). Thus, beyond shaping glioma cell identity3,4, hypoxia may cultivate an immunosuppressive ecosystem and induce astrocyte reactivity.
Finally, we used CellChat42 to quantitatively infer ligand–receptor interactions between malignant cell states and nonmalignant cell types. This resulted in two separate subnetworks, supporting the division into tumor core regions (Fig. 4e, left) versus periphery (Fig. 4e, right). Among those, we found (1) IGF1 and IGF1R cross-talk between TAM-MDM and MES-like cells, which is consistent with a proposed resistance mechanism43; (2) PTN-PTPRZ1 OPC cross-talk, which may promote tumor progression and invasion17,31; and (3) NLGN-NRXN and NRG-ERBB4 interactions between neurons and NEU-like cells, reported as potential therapeutic target in glioma44,45,46 (Fig. 4e). Taken together, GBM microenvironmental niches such as hypoxia, glial and NEU TME may serve as determinants of malignant and nonmalignant states.
Genetic associations with GBM transcriptional states
Malignant gene expression profiles may also be driven by genetic alterations. Previously, we found that the CNAs of CDK4, PDGFRA and EGFR and mutations in NF1 were associated with inferred abundance of cellular states in bulk GBM data from TCGA4. To examine these associations in our snRNA-seq dataset, we analyzed malignant state abundance in samples with matched DNA profiling and sufficient malignant cells (n = 108 samples). As expected, we found associations between CDK4 amplification and NPC-like abundance, PDGFRA amplification and OPC-like abundance, and NF1 mutations and MES-like abundance (Fig. 5a; Wilcoxon rank-sum test, P < 0.05). In contrast, we did not observe an association between EGFR amplifications and AC-like abundance.
a, Association between genetic aberrations and malignant cell-state abundance. Two-sided Wilcoxon rank-sum tests are presented. P value adjustments for multiple hypothesis testing were not performed. b, Cell type composition of each of the tumors (n = 121) in the dataset, aggregated based on their composition group. c, Malignant cell-state proportions (rows) in each tumor sample (n = 111 with at least 50 malignant cells; columns). Tumors are aggregated according to the dominant malignant state (at least 30% of cells). d, Interaction graph between the different layers of GBM (composition groups, malignant state groups, BPs, gene-level CNAs and single-nucleotide mutations). Each node represents a layer; nodes are connected with an edge if a statistically significant association exists between the nodes. mut, mutation.
We next tested whether EGFR amplifications, or other genetic alterations, might be associated with the newly identified states (Fig. 1b). The strongest associations were between canonical genetic alterations of GBM (that is, chromosome 7 gain, chromosome 10p loss, EGFR amplification and TERT promoter mutations) and increased GPC-like state abundance (Fig. 5a and Extended Data Fig. 10a). These canonical genetic alterations typically co-occur in GBM, so we compared samples with at least one of these alterations to those without. Tumors with canonical alterations had, on average, 12% GPC-like malignant cells, compared with less than 1% of those without (Wilcoxon rank-sum test, P = 0.003; Extended Data Fig. 10b), suggesting that the GPC-like state is rare in GBMs lacking canonical alterations.
Other associations included TP53 mutations linked to NEU-like abundance and chromosome 7 amplifications linked to hypoxia abundance, which is consistent with a recent study28. Similar analyses of BPs found that glial BP is more closely aligned with canonical GBM genetic alterations compared to ECM or NEU BPs (Extended Data Fig. 10c). Together, these results indicate an interplay between genetic alterations and specific cellular states, although our dataset may still be underpowered to detect additional associations given the low frequencies of GBM genetic alterations.
From transcriptional layers to integrated GBM ecosystems
We defined three distinct layers of GBM transcriptional patterns: cell type composition; malignant cellular states; and BPs. To examine the associations across these layers, we first classified all samples according to each layer (Supplementary Table 5). Regarding cell type composition, tumors were divided into three main groups according to malignant fraction (high, intermediate and low); the latter group was further subdivided according to the dominant TME cell type (Fig. 5b). Regarding malignant state, most tumors were dominated by one state, thereby defining six groups (Fig. 5c). Finally, BPs were used to define three groups of tumors as described above (Fig. 3d).
Next, we investigated associations across layers (Fig. 5d). Each of the three BPs was significantly associated with classifications according to cell type composition and malignant state, as well as specific genetic events. BP-NEU was associated with the NEU-like and OPC and NPC-like states, with a glio-neural (GN)-rich (neurons, astrocytes, oligodendrocytes) TME and with TP53 and RB1 mutations. BP-ECM was associated with the MES-like and hypoxia states, with macrophage-rich TME and NF1 mutations. Finally, BP-glial was associated with the GPC-like state, with a highly malignant fraction and with EGFR and MDM2 amplifications (Fig. 5d).
These results define three multilayer ecosystems, each including correlated features among the three layers (Fig. 6a). Overall, 43% (52 of 121) of the samples could be robustly assigned to one of these three multilayer ecosystems, based on either two or three of its layers (Fig. 6b and Supplementary Table 6), which is significantly more than expected by chance (P < 0.01 for each ecosystem according to a binomial test). The remaining samples could be described according to individual layers but did not correspond to a stereotypic association between these layers (Extended Data Fig. 10d), highlighting the complexity and diversity of GBM ecosystems.
a, Layer map of the 54 samples that could be classified to one of the multilayer groups. b, Model for stereotype architecture in GBM, with exemplar tumors for each ecosystem and their compositional, cell-state and BPs. Multiple intercorrelated transcriptomic layers make up the GBM ecosystems.
Discussion
After the first report of scRNA-seq for 430 cells derived from five GBM samples3, technical progress has dramatically expanded the scale of single-cell profiling. In this study, we leveraged a large-scale single-nucleus dataset (~430,000 nuclei, 121 samples) to comprehensively redefine the GBM cellular architecture. We discovered three new cellular states that were not captured in previous glioma studies, including a differentiated NEU state (NEU-like), a new progenitor state (GPC-like) and a cilia-related state. The capacity to identify these new states may be due to the high number of samples and cells profiled in this study. Yet this explanation is insufficient given that two of the new states are relatively common in our data (6.7% NEU-like and 14.2% GPC-like cells). Thus, we speculate that these states were depleted by single-cell isolation protocols that require enzymatic digestion and may poorly capture specific cell states (for example, those with complex morphology). Single-nucleus protocols isolate nuclei using tissue homogenization in cold buffer, thus capturing a wider variety of cell types23,47. Accordingly, the newly described malignant states may resemble normal cells with complex cellular processes (for example, neurons, ciliated cells).
GBM cellular states resemble the programs observed in normal glial or NEU cell types. This finding is further supported by analysis of potential cellular transitions. Transitions between GBM states was demonstrated previously4, but the rate of specific transitions is unknown. We explored this question through quantification of hybrid states, based on the assumption that cells mapping to two states often reflect a transition. Only eight of the potential 38 hybrid states accounted for 71% of hybrids, highlighting the enrichment of specific hybrids and presumably the identity of common transitions. These putative transitions also mirror normal brain development, supporting the view that many aspects of GBM biology are inherited from the cell of origin.
Previous work noted that much of the diversity between GBMs is tightly linked to diversity within GBMs, such that tumor subtypes can be partially explained by the abundance of recurring cellular states. However, not all intertumor variation may be explained in this way; to our knowledge, no previous studies attempted to define the residual signal of intertumor heterogeneity. In this study, we addressed this issue by analyzing intertumor heterogeneity only among cells of the same cellular state, thereby controlling for the diversity of states within tumors. We found that the main patterns that vary between tumors in one state also vary in other states, thus defining a state-independent pattern of intertumor variability that may be thought of as reflecting a tumor baseline profile.
Notably, the three BPs we identified—NEU, ECM and glial—are functionally reminiscent of some of the malignant states that vary within tumors (NEU-like, MES-like and GPC-like), although they include distinct sets of genes. Moreover, these features are correlated, such that tumors with one BP also tend to be enriched with the related malignant state. We speculate that certain genetic and environmental features give rise to two related effects: a BP that influences all cells in the tumor and a higher fraction of cells with a certain expression program (that is, state). Thus, tumors with an NEU BP (represented by one set of NEU-related genes) also tend to have an abundance of cells that activate another set of genes that corresponds to the NEU-like MP. Similarly, tumors with an ECM BP also have an abundance of cells that activate genes of the MES-like MP. At bulk, the two phenomena of BP and frequency of certain MPs coalesce into the signal of GBM subtypes, while single-cell analysis allows us to dissect these distinct effects.
What might be the signals that concomitantly activate BP and frequencies of MPs? Our analysis shows a correlation of both of these malignant features with the TME. NEU malignant features are associated with low-purity tumors enriched with glial and neural TME cells. Mesenchymal and ECM malignant features are associated with low-purity tumors enriched with macrophages. Finally, glial malignant features are associated with high purity. In addition to the TME, we also found genetic events associated with a combination of tumor features: NF1 mutations, TP53 and RB1 mutations, and EGFR and MDM2 amplifications are linked to the ECM, NEU and glial ecosystems, respectively. Thus, we speculate that tumor genetics along with tumor location, which influences the tumor TME, define the propensity of each GBM toward a particular BP and enriched MP, culminating in three stereotypic ecosystems, each associated with multiple correlated components—tumor genetics, nonmalignant TME and two malignant features (BP and MP frequency). Nevertheless, the coupling between these components is only partial, such that around half of the tumors cannot be assigned to a stereotypical ecosystem but rather reflect unique combinations of features, highlighting the diversity and complexity of GBM.
Methods
Participants and ethical approval
Frozen GBM specimens, diagnosed as ‘glioblastoma, IDH WT’ according to the WHO 2021 classification, were collected at seven institutions, as outlined in the sections below. Collection and genomic profiling were approved by the institutional review board (IRB) of each institution; all participants provided written informed consent accordingly. Cohorts were added to the IRB of the Dana-Farber/Harvard Cancer Center (no. 10-417). Patients were male and female. Clinical information of the cohorts is summarized in Supplementary Table 1. Additional clinical characteristics related to the longitudinal analyses are shown in the companion study by Spitzer et al.18.
MD Anderson Cancer Center cohort
For this cohort (n = 4), tumors were collected from MD Anderson Cancer Center. Collection was approved by the Institutional Review Board of MD Anderson Cancer Center under the protocol number 2012-0441.
Duke University cohort
For this cohort (n = 12), tumors were collected from the Preston Robert Tisch Brain Tumor Center Biorepository, Duke University Hospital. Collection was approved by the IRB of Duke University under protocol no. Pro0007434.
Tokyo University cohort
For this cohort (n = 30), tumors were collected from the Department of Neurosurgery, Tokyo University Hospital. Collection was approval by the IRB of Tokyo University Hospital under protocol no. G10028.
Pitié-Salpêtrière Hospital cohort
For this cohort (n = 28), tumors were collected from the Pitié-Salpêtrière Hospital and were approved by the Onconeurotek Brain Tumor bank of the hospital Pitié-Salpêtrière under certification no. 96-900.
St. Michael’s Hospital cohort
For this cohort (n = 14), tumors were collected from the Division of Neurosurgery, St. Michael’s Hospital, Unity Health Toronto. Collection was approved by the Research Ethics Board of St. Michael’s Hospital, Unity Health Toronto, under protocol no. REB 13-141; all patients provided signed informed consent accordingly.
Seoul National University cohort
For this cohort (n = 20), tumors were collected from the Department of Neurosurgery, Seoul National University Hospital, Seoul. Collection was approval by the IRB of Seoul National University Hospital (approval no. H-2004-049-1116).
NORLUX Neuro-oncology Laboratory
For this cohort (n = 13), tumors were collected from Centre Hospitalier de Luxembourg (Neurosurgical Department). Collection was approved by the National Committee for Ethics in Research Luxembourg, under protocol no. 201201/06. Cohorts were added to IRB protocol Dana-Farber/Harvard Cancer Center 10-417. Patients were male and female. Clinical information of the cohorts is summarized in Supplementary Table 1. Additional clinical characteristics related to longitudinal analyses are shown in the companion paper by Spitzer et al.18.
Nuclei isolation from frozen tissue
In protocol 1 (laboratory 1 and laboratory 2 for the cohorts from Duke University, MD Anderson Cancer Center, Tokyo University Hospital, St. Michael’s Hospital and Pitié-Salpêtrière Hospital), nuclei from frozen tumor tissue were isolated. Tissue was thawed and mechanically dissociated in ST buffer (10 mM Tris-HCl, pH 7.5, 146 mM NaCl, 1 mM CaCl2, 21 mM MgCl2) with 0.49% CHAPS (no. 28300, Merck Millipore). Single-nucleus suspensions were filtered using a 40-μm strainer, centrifuged at 500g for 5 min and resuspended in ST buffer supplemented with 0.01% BSA (cat. no. B9000S, New England Biolabs). Nuclear suspensions were inspected by microscope, counted using a hemocytometer and used for 10x Genomics workflow or fluorescence-activated cell sorting (FACS) for the Smart-seq2 workflow.
In protocol 2 (laboratory 3 for the cohorts from NORLUX and Seoul National (SNU) cohorts), tissue samples were thawed and mechanically dissociated in Nuclei EZ Lysis Buffer (NUC101, Sigma-Aldrich) using dounce homogenization. Solutions were incubated on ice for 5 min and mixed 1–2 times during incubation. Single-nucleus suspensions were filtered through a 70-μm strainer and centrifuged at 500g for 5 min at 4 °C, resuspended in Nuclei EZ Lysis Buffer and incubated on ice for 5 min. Solutions were centrifuged at 500g for 5 min at 4 °C and resuspended in 1% BSA, 0.2 U μl−1 RNase inhibitor and PBS buffer (three times). For the final resuspension, 4′,6-diamidino-2-phenylindole was added to the buffer; the solution was filtered through a 40-μm strainer, cells were counted on a Countess II Automated Cell Counter (Thermo Fisher Scientific) and nuclei were taken into the 10x Genomics workflow.
10x Genomics for single-nucleus sequencing
The Chromium Single Cell 3′ Reagent Kit Vv3 (cat. no. PN1000128, 10x Genomics) was used according to the manufacturer’s protocol. Briefly, nuclei were loaded on the Chromium Chip (cat. no. PN12000120, 10x Genomics) with a target cell recovery of 6,000–8,000 nuclei and processed in the Chromium Controller. Single nuclei were partitioned into Gel Beads-in-emulsion, followed by RNA reverse transcription with barcoding. Libraries were created by breaking the Gel Beads-in-emulsion and pooling barcoded fractions, complementary DNA amplification, fragmentation and attachment of sample index and adapter, and sequenced on the NextSeq 500 or NovaSeq (Illumina) platform, with paired-end 28–91 bp reads. For the NORLUX and SNU cohorts, nuclei were loaded on a Chromium Chip with a target cell recovery of 6,000 nuclei for single-cell multi-omic assay for transposase-accessible chromatin and gene expression according to the manufacturer’s protocol. The gene expression chemistry for the 10x multi-ome possesses the same chemistry as the 10x v3 3′ snRNA-seq.
Nucleus sorting for Smart-seq2
The nuclear suspension was stained using the Vybrant DyeCycle Ruby Stain (cat. no. V10309, Thermo Fisher Scientific) immediately before FACS. Single-nucleus sorting was performed on a FACSAria Fusion Sorter (Becton Dickinson) using a 640-nm laser (Ruby, 670/14 filter) (Extended Data Fig. 2d). After doublet discrimination, intact nuclei were selected with ruby+ and were sorted into 96-well plates containing TCL buffer (cat. no. 1031576, QIAGEN) with 1% 2-mercaptoethanol (cat. no. M6250, Sigma-Aldrich). Plates were frozen on dry ice immediately after sorting and stored at −80 °C before whole-transcriptome amplification, library preparation and sequencing.
Smart-seq2
Whole-transcriptome amplification, library construction and sequencing were performed as previously published with slight modification for the nucleus as follows4,19. Briefly, on 96-well plates, RNA derived from single nuclei was first purified with Agencourt RNAClean XP beads (cat. no. A66514, Beckman Coulter). Then, Oligo(dT)-primed reverse transcription was performed using Maxima H Minus Reverse Transcriptase (cat. no. EP0753, Thermo Fisher Scientific) and locked TSO oligonucleotide (5′-AAGCAGTGGTATCAACGCAGAGTACATrGrG+G-3′, QIAGEN). This was followed by PCR amplification (22 cycles for snRNA-seq) using the KAPA HiFi HotStart ReadyMix (cat. no. K2602, KAPA Biosystems) with subsequent Agencourt AMPure XP Bead (cat. no. A63882, Beckman Coulter) purification. Libraries were tagmented using the Nextera XT Library Prep Kit (cat. no. FC-131-1096, Illumina) with custom barcode adapters. Libraries from 768 cells with unique barcodes were pooled and sequenced on a NextSeq 500 or NovaSeq 6000 sequencer (Illumina), with paired-end 38 bp reads.
WES
WES for the samples from Duke University, the MD Anderson Cancer Center, Tokyo University Hospital and St. Michael’s Hospital was performed as follows. DNA was extracted from each frozen tumor sample and blood samples corresponding to the patients using the DNeasy Blood & Tissue Kit (cat. no. 69504, QIAGEN). Genomic DNA (100–250 ng) was acoustically sheared using an ultrasonicator (Covaris), targeting 150-bp fragments. Library preparation was performed using the KAPA HyperPrep Kit (cat. no. KK8504, KAPA Biosystems) followed by enzymatic cleanup using AMPure XP beads. Exome capture was performed using a custom exome bait (manufactured by Twist Bioscience to Broad Institute’s specifications). Captured libraries were sequenced with 150 bp paired-end sequencing on a NovaSeq 6000. For the samples from Pitié-Salpêtrière Hospital, after DNA was fragmented by a LE220 ultrasonicator (Covaris) and size-selected, library preparation and capture were performed using Twist Human Core Exome kit (Twist Bioscience) according to manufacturer’s protocol. Sequencing was performed on a NovaSeq 6000, with 200-bp paired-end sequencing.
WGS
Newly generated whole-genome DNA sequencing data were collected for a cohort of frozen samples from the NORLUX Neuro-oncology Laboratory. Briefly, DNA was extracted from each tumor sample using the AllPrep DNA/RNA Mini Kit (cat. no. 80204, QIAGEN) for samples with sufficient tumor tissue and matched normal blood when it was available. Selected tissue for bulk DNA isolation was adjacent to tissue used for single-nucleus data generation. Briefly, DNA was sheared to 400 bp using an LE220 ultrasonicator (Covaris) and size-selected using AMPure XP beads. Whole-genome libraries were prepared and sequenced with 150 bp paired-end sequencing on a NovaSeq 6000 platform. WGS data for the SNU cohort was prepared in an identical manner and was previously reported in ref. 9. Data for the SNU cohort are available on Synapse (https://www.synapse.org/glass).
Statistics and reproducibility
No statistical method was used to predetermine sample size. All available samples were provided by the aforementioned centers and no data were excluded from the analyses. For figures containing box plots, the box spans from the first to third quartiles, median values are indicated by a horizontal line and the whiskers show 1.5× the interquartile range; individual sample points are presented. The statistical analysis described in this work was done using R v.4.0.1 and above.
Somatic variant detection and copy number calling
DNA sequencing alignment, fingerprinting, somatic variant detection (Mutect2) and copy number segmentation was performed in accordance with the Genome Analysis Toolkit best practices using the Genome Analysis Toolkit v.4.0.10.1, as described previously9,48. Briefly, whole-exome and whole-genome reads were aligned to the b37 genome (human_g1k_v37_decoy) using the Burrows–Wheeler Aligner MEM v.0.7.17. DNA fingerprint analysis using ‘CrosscheckFingerprints’ (Picard) confirmed that all samples belonging to a patient came from the same individual, indicating that there were no sample mismatches in this study. Somatic mutations were detected using Mutect2 (v.4.1.0.0) for tumor samples with a matched normal blood sample. A panel of normals was constructed for each sequencing batch to account for differences in DNA library preparation (for example, whole-genome versus whole-exome) and used to filter out common artifactual and germline variants. Patient tumor samples without matched normal blood were analyzed for copy number alterations but not for SNV detection.
Single-cell and single-nucleus data processing of human glioma samples
For Smart-seq2 sequencing data, the reads were mapped to the UCSC hg19 human transcriptome using Bowtie with the following parameters:-q-phred33-quals -n -e 99999999 -l 25 -I 1 -X 2000 -a -m 15–best -S -p 6. For droplet-based sequencing data, the Cellranger v.3.1.0 pipeline provided by 10x Genomics was used for alignment (GRCh38 release 93) and to generate count matrices. Smart-seq2 gene expression levels were quantified as \({E}_{i,\,j}=\log 2(\frac{{{\mathrm{TPM}}}_{i,\,j}}{10}+1)\) where TPMi,j refers to transcripts per million for gene i in sample j, as calculated using RSEM. For droplet-based sequencing data, we quantified the gene expression levels as \({E}_{i,\,j}=\log 2(\frac{{{\mathrm{CPM}}}_{i,\,j}}{10}+1)\), where CPMi,j refers to counts per million for gene i in sample j. We divided the TPM and CPM values by ten as the size of single-cell libraries is estimated to be in the order of 100,000 transcripts; therefore, we wanted to avoid inflating the expression levels by counting each transcript approximately ten times. After these normalization steps, initial filtering of low-quality cells was done based on the low number of detected genes (fewer than 200 genes) or the high expression of mitochondrially encoded genes (more than 3%). Most low-quality cells were removed downstream in case they could not be robustly classified as malignant or nonmalignant (see the later sections on the CNA analysis). Lastly, we computed the average expression of each gene i as \(\log 2[(\frac{1}{n}{\sum }_{j=1}^{n}{{\mathrm{TPM|CPM}}}_{i,\,j})+1]\) and filtered out lowly expressed genes by limiting the analyzed genes to the top 3,000 most highly expressed genes (using the Seurat v.4.0.4 (ref. 49) function CreateSeuratObject). For the cells and genes that passed these quality control filters, we defined relative expression levels by centering the expression levels for each gene across all cells in the dataset as follows: \({{Er}}_{i,\,j}={E}_{i,\,j}-\,\frac{1}{N}{\sum }_{k=1}^{N}{E}_{i,k}\) where N is the number of cells in the dataset.
Clustering and cell type annotation
To facilitate clustering, cell type annotation and scoring of droplet-based data, we used the R package Seurat v.4.0.4 (ref. 49). Unique molecular identifier count data were normalized and scaled (using the NormalizeData and ScaleData functions) and then clustered by projecting the data to a lower dimensional space using PCA and then running the FindNeighbors and FindClusters functions. Initial classification of tumor-associated macrophages, T cells, oligodendrocytes and endothelial cells was based on the expression of known marker genes22,23,24,25,26, whereas the rest of the cells were annotated as presumed malignant. After classification of cells as malignant or nonmalignant (see the following two sections), cells classified as nonmalignant were scored (using the AddModuleScore function) for known cell-type-signatures and assigned a cell type using the method explained in the section ‘Assignment of cells to states’. Finally, heterotypic doublets were filtered our using the R package scDblFinder50.
Inferring CNAs from gene expression data
CNAs were estimated as described previously4,13,14,21 using the function infercna from the R package infercna (an implementation of the original method presented in ref. 3, which is available at https://github.com/jlaffy/infercna) with default parameters. Briefly, the algorithm sorts the analyzed genes in each sample according to their chromosomal location and applies a moving average with a sliding window of 100 genes in each chromosome to the relative expression values. The scores computed for the cells classified as nonmalignant (oligodendrocytes, macrophages and endothelial cells) define the baseline of the normal karyotype and their average CNA values are used to center the values of all cells.
Classification of cells into malignant or nonmalignant cell types
In cells classified using single-nucleus droplet-based sequencing, we generally encountered a lower CNA signal and continuous signal distributions related to (1) lower data quality relative to that observed in single-cell data, and (2) inclusion of nonmalignant cell types, such as astrocytes and neurons, in the presumed malignant group as these cell types could not be classified a priori as nonmalignant because of transcriptional similarity with malignant cells.
Therefore, we used a multi-step classification scheme for the droplet-based, single-nucleus sequencing data.
Step 1: cell classification using detection of copy number events
For each cell j, we computed the CNA signal on each chromosome separately as \({{\mathrm{CNA}}}_{j}^{C}=\frac{1}{n}{\sum }_{i=1}^{n}{{\mathrm{CNA}}}_{i,\,j}^{C}\) where n is the number of genes on chromosome C included in the analysis. We fitted a normal distribution (that is, a classifier) to the signal distributions of the reference cells for each chromosome (using the function fitdistr from the R package MASS v.7.3-56). This enabled assigning each cell with a P value for each chromosome using a two-sided z-test (function pnorm from the R package stats) reflecting whether the cell had a CNA signal that was significantly different than expected from nonmalignant cells. These P values were further corrected for multiple testing (for each chromosome) using the Benjamini–Hochberg method; cells with Benjamini–Hochberg-adjusted P < 0.05 for a particular chromosome were considered as having a copy number event on that chromosome (either amplification or deletion based on the sign of the CNA signal). Cells with an unadjusted P < 0.05 for a particular chromosome that failed to achieve statistical significance after adjustment were considered to have a ‘suspicious event’ on that chromosome. Then, inferred copy number events were cross-referenced with copy number calls from the WES and WGS data; only events detected in both methods were considered for classification and were termed ‘real’ events. For a subset of samples, this procedure was not possible because of (1) lack of WES or WGS data, or (2) inability to call CNA events from WES or WGS because of low purity or quality. For this subset of cases, we defined a panel of highly recurring CNA events (intersection of highly recurring events inferred from the snRNA-seq and WES and WGS data), which were used as a short-list of potential events. Cells classified a priori as nonmalignant were reclassified as ‘unresolved’ in case a ‘real’ or ‘suspicious’ CNA event was detected and ‘nonmalignant’ otherwise. Presumed malignant cells with at least one ‘real’ CNA event were classified as ‘malignant’; those with a ‘suspicious’ event were classified as ‘unresolved’ and those with no events at all (‘real’ or ‘suspicious’) were classified as ‘nonmalignant’.
Step 2: exclusion of questionable cells based on CNA correlation
We defined the CNA correlation as the Pearson correlation between each cell and the average CNA profile of the tumor sample the cell originated from. We limited the computation to the chromosomes on which ‘real’ events were detected and a control panel of ~1,000 genes from the other chromosomes. Based on the distribution of CNA correlations across the cells classified as malignant, we defined 0.5 as the lower threshold of malignancy. To determine the upper threshold for nonmalignancy, we performed SNV calling from the snRNA-seq data to estimate the algorithm misclassification rate. Briefly, for each sample with available WES data, we called SNVs for each cell using the Vartrix tool (https://github.com/10XGenomics/vartrix) from the FASTQ files. Misclassified cells were defined as cells harboring a malignant mutation that were classified as nonmalignant in step 1 of the algorithm. After a cost-effectiveness analysis (misclassification rate versus percentage of excluded cells), we set 0.35 as the upper threshold for nonmalignancy. Finally, cells classified as malignant with CNAcor < 0.5 or cells classified as nonmalignant with CNAcor > 0.35 were reclassified as ‘unresolved’ and excluded from the downstream analysis.
Step 3: refining classification by setting confidence levels
To refine the classification, we defined confidence levels. Confidence level 1 includes cells classified as malignant that harbored a malignant SNV, cells in which at least 50% of the expected CNA events were detected including chromosome 7 amplification or chromosome 10 deletion, and cells classified as nonmalignant with CNAcor < 0.25. Confidence level 2 includes cells classified as malignant in which at least 50% of the expected CNA events were detected that do not include chromosome 7 amplification or chromosome 10 deletion, cells classified as malignant in which less than 50% of the expected CNA events were detected, including chromosome 7 amplification or chromosome 10 deletion, and cells classified as nonmalignant with CNAcor < 0.5. The rest of the cells classified as malignant or nonmalignant were assigned confidence level 3. In the analysis presented in this work, we used only cells in confidence levels 1 and 2.
Deriving MPs from gene expression data
To capture the heterogeneity between cells from the same cell type, we leveraged the NMF51. NMF was performed on the relative expression values of each sample independently after setting negative values to zero. The NMF algorithm requires defining a priori the ‘k’ parameter, reflecting the expected number of latent features in the data. Since k varies between samples and is largely unknown, we ran the NMF algorithm on each sample using different values (3–10), thereby generating 52 programs for each sample. Each of these NMF programs was summarized using the top 50 genes based on the NMF coefficients. Derivation of the MPs from the NMF programs was described thoroughly by in ref. 30 and is described briefly here. The method first filters out NMF programs that are not robust (do not recur in the sample they originated from or across several samples) or are redundant in the sample (that is, they highly overlap other NMF programs in that sample). Then, robust NMF programs are clustered according to Jaccard similarity using a customized clustering algorithm that iteratively considers the degree of overlap between robust NMF programs and combines highly overlapping ones to form a cluster. The R 50 most-recurring genes in each cluster define an MP.
Overall, the algorithm yielded 16 MPs that were annotated based on functional enrichment analysis (using GO terms, Molecular Signatures Database Hallmark gene sets and gene sets derived from normal brain development datasets). MP16 included genes from several cell types resulting in an MP that fitted almost all cells and thereby did not reflect any heterogeneity; thus, it was excluded from the analysis. This same procedure was independently repeated for each nonmalignant cell type.
Assignment of cells to states
Malignant cells were scored for the NMF MPs using the Seurat function AddModuleScore. To facilitate cell classification, we generated 20 shuffled expression matrices by sampling each time 5,000 cells and shuffling the expression values for each gene. We then scored each shuffled matrix for the NMF MPs, thereby yielding 100,000 normally distributed scores for each expression program. These served as null distributions for cell-state classification. For each original cell, we computed a P value for each of the expression programs with a z-test (using the R pnorm function) using the statistics of the null distributions that we generated previously. We adjusted all P values for multiple testing using the Benjamini–Hochberg method. Each cell was classified into a specific state if the adjusted P value computed for that state was smaller than 0.05. Cells that achieved Padj < 0.05 for multiple states were assigned to the state for which they scored maximally. Cells that did not achieve Padj < 0.05 for any of the states were assigned an ‘unresolved’ state.
Cells were assigned a ‘cycling’ state on top of their cellular state if they achieved Padj < 0.05 for the cell cycle MP and ‘non-cycling’ otherwise.
Hybrids detection and classification
We define hybrids as cells scoring significantly (that is, with Padj < 0.05) for at least two cellular states with a difference between the top two significant scores smaller than expected by chance. To quantify the score difference expected by chance, we leveraged on the null distribution generated for the state classification algorithm and computed for each artificial cell the difference between the top two states. The 95 quantiles of this score difference distribution (equal to a difference of 0.24) was set as the threshold for determining if a state was singular or hybrid.
To estimate the expected frequency of technical hybrids, we first computed for each pair of states (A, B) the expected frequency of hybrids, defined as \({\mathrm{Exp}}\left({\mathrm{A,B}}\right)={\mathrm{Freq}}\left({\mathrm{A}}\right)\times {\mathrm{Freq}}\left({\mathrm{B}}\right)\) and the observed/expected ratio defined as \({\log }_{2}(\frac{{\mathrm{Obs}}({\mathrm{A,{B}}})}{{\mathrm{Exp}}({\mathrm{A,B}})})\). As this is an overestimation of the expected frequency attributed to technical effects, we estimated this technical factor (TF) by averaging across observed/expected ratios of hybrid pairs detected less than expected by chance (that is, observed/expected < 0). Then, we computed the expected frequency of technical hybrids for each hybrid pair by multiplying the expected frequency with the technical factor and defined the observed/expected ratio as \({\log }_{2}(\frac{{\mathrm{Obs}}({\mathrm{A,{B}}})}{{{\mathrm{TF}}}\times {\mathrm{Exp}}({\mathrm{A,B}})})\). Finally, we filtered out insignificant hybrid pairs with large observed/expected ratio (attributed to small numbers) using a one-sided Fisher’s test. The hybrid lineage model was generated using the R package igraph, with cell states as nodes and edges connecting the states that formed significant hybrid pairs. We did not include the ‘cycling’ state in this analysis because we did not view ‘cycling’ as an independent state but rather as an additional feature that cells can have on top of their neurodevelopmental identity (that is, AC-like, OPC-like, and so on).
BP derivation
Each tumor sample was decomposed to up to seven pseudo-bulk profiles (one for each of the common states—AC-like, MES-like, hypoxia, GPC-like, OPC-like, NPC-like, NEU-like—given at least 25 cells classified to that particular state) by averaging across the normalized unique molecular identifier counts and log2-transforming. Genes were included if their mean log2 expression value across all pseudo-bulks was at least 1 and if their median variance (computed separately within state) was at least 2.5. Overall, 1,005 genes passed these filters. Then, pseudo-bulks were analyzed within state to remove the state-specific signal using PCA. We then derived six gene programs from each state (top and bottom 50 genes of the first three principal components), computed the Jaccard similarity index between each pair of programs and hierarchically clustered the similarity matrix using the ConsensusClusterPlus package (v.1.56.0). Analysis of the cumulative distribution function revealed minor increases in area under the curve for k > 5 (where k is the number of clusters); therefore, k = 5 was chosen as the number of clusters. Then, we derived five consensus signatures by including genes that recurred in at least 25% of programs in each of the clusters (with a hard minimum of at least three times). Using manual inspection and GO enrichment, we annotated the consensus signatures. Two signatures were excluded because of short length or high suspicion of reflecting a technical artifact. To exclude the possibility that ambient RNA had a role in driving the BPs, we estimated the contamination by ambient RNA per sample using the R package SoupX (v.1.6.2). There was no difference in the estimated contamination level across the different BPs and no significant overlap between the top most contaminated genes per sample and the MP or BP signatures (data not shown).
Assigning tumors to composition groups
To measure the differences in tumor composition, we generated for each tumor sample a composition vector reflecting the proportion of each cell type in the tumor sample. This enabled assigning tumors to three main composition groups: high malignant fraction (percentage of malignant cells > 75%); intermediate malignant fraction (percentage of malignant cells 50–75%); and low malignant fraction (percentage of malignant cells < 50%), where the low malignant fraction group could be further subdivided according to the dominant TME cell types (percentage of TAM, oligodendrocyte, GN > 40%, or mixed in case of no dominant TME cell type). Similarly, we defined the proportion vectors for the malignant cell states and assigned tumors to six groups according to the dominant cell state (at least 25% of cells assigned to the particular state)—AC, MES/hypoxia, GPC, OPC/NPC, NEU and mixed (in case of no dominant state). Tumors were classified according to the BP with maximal score and to the mixed category if the maximal score was less than 0.25.
Multilayer group definition
The association between each pair of features (A, B) was computed using a binomial test with the number of times the pair was observed as the number of successes, the number of tumor samples as the number of experiments and the frequency of feature A times the frequency of feature B as the expected probability of success (under the assumption that the features were independent). We then generated an undirected graph by defining each feature as a node and connecting the nodes with edges when a statistically significant association (P < 0.05, unadjusted for multiple testing) between the two features was observed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Processed snRNA-seq data generated for this study are available at the Gene Expression Omnibus under accession no. GSE274546 (10x Genomics) and GSE274548 (Smart-seq2). All de-identified somatic mutation and copy number alteration calls, and genomic analysis tables, are available via Synapse (https://www.synapse.org/care_glioblastoma). Raw sequencing data are available with limitations in accordance with the consent forms from the Data Use Oversight System (DUOS) at https://duos.boardinstitute.org under the following IDs: DUOS-000475; DUOS-000476; DUOS-000477; DUOS-000478; DUOS-000479; and DUOS-000480.
Code availability
The analysis scripts used in this study are available via GitHub at https://github.com/dravishays/GBM-CARE-WT and via Zenodo at https://doi.org/10.5281/zenodo.14966015(ref. 52). Scripts for processing the DNA sequencing data can be accessed at https://github.com/Kcjohnson/care-glass.
Change history
26 June 2025
Following publication of this article, text citations to Extended Data Fig. 9a and 9b have been amended to cite "Extended Data Fig. 9" in the HTML and PDF versions of the article.
References
Louis, D. N. et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol. 23, 1231–1251 (2021).
Sottoriva, A. et al. Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc. Natl Acad. Sci. USA 110, 4009–4014 (2013).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Neftel, C. et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178, 835–849 (2019).
Venkataramani, V. et al. Glioblastoma hijacks neuronal mechanisms for brain invasion. Cell 185, 2899–2917 (2022).
Meacham, C. E. & Morrison, S. J. Tumour heterogeneity and cancer cell plasticity. Nature 501, 328–337 (2013).
Hara, T. et al. Interactions between cancer cells and immune cells drive transitions to mesenchymal-like states in glioblastoma. Cancer Cell 39, 779–792 (2021).
Venkatesh, H. S. et al. Electrical and synaptic integration of glioma into neural circuits. Nature 573, 539–545 (2019).
Varn, F. S. et al. Glioma progression is shaped by genetic evolution and microenvironment interactions. Cell 185, 2184–2199 (2022).
Couturier, C. P. et al. Single-cell RNA-seq reveals that glioblastoma recapitulates a normal neurodevelopmental hierarchy. Nat. Commun. 11, 3406 (2020).
Filbin, M. G. et al. Developmental and oncogenic programs in H3K27M gliomas dissected by single-cell RNA-seq. Science 360, 331–335 (2018).
Johnson, K. C. et al. Single-cell multimodal glioma analyses identify epigenetic regulators of cellular plasticity and environmental stress response. Nat. Genet. 53, 1456–1468 (2021).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Venteicher, A. S. et al. Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq. Science 355, eaai8478 (2017).
Garofano, L. et al. Pathway-based classification of glioblastoma uncovers a mitochondrial subtype with therapeutic vulnerabilities. Nat. Cancer 2, 141–156 (2021).
Wang, L. et al. The phenotypes of proliferating glioblastoma cells reside on a single axis of variation. Cancer Discov. 9, 1708–1719 (2019).
Wang, Q. et al. Tumor evolution of glioma-intrinsic gene expression subtypes associates with immunological changes in the microenvironment. Cancer Cell 32, 42–56 (2017).
Spitzer, A. et al. Deciphering the longitudinal trajectories of glioblastoma ecosystems by integrative single-cell genomics. Nat. Genet. https://doi.org/10.1038/s41588-025-02168-4 (2025).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Ceccarelli, M. et al. Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164, 550–563 (2016).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Campbell, J. N. et al. A molecular census of arcuate hypothalamus and median eminence cell types. Nat. Neurosci. 20, 484–496 (2017).
Habib, N. et al. Div-Seq: single-nucleus RNA-Seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Polioudakis, D. et al. A single-cell transcriptomic atlas of human neocortical development during mid-gestation. Neuron 103, 785–801 (2019).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Coy, S. et al. Single cell spatial analysis reveals the topology of immunomodulatory purinergic signaling in glioblastoma. Nat. Commun. 13, 4814 (2022).
Ravi, V. M. et al. Spatially resolved multi-omics deciphers bidirectional tumor–host interdependence in glioblastoma. Cancer Cell 40, 639–655 (2022).
Schmitt, M. J. et al. Phenotypic mapping of pathologic cross-talk between glioblastoma and innate immune cells by synthetic genetic tracing. Cancer Discov. 11, 754–777 (2021).
Gavish, A. et al. Hallmarks of transcriptional intratumour heterogeneity across a thousand tumours. Nature 618, 598–606 (2023).
Bhaduri, A. et al. Outer radial glia-like cancer stem cells contribute to heterogeneity of glioblastoma. Cell Stem Cell 26, 48–63 (2020).
Mathur, R. Glioblastoma evolution and heterogeneity from a 3D whole-tumor perspective. Cell 187, 446–463 (2024).
Gimple, R. C. et al. Glioma stem cell-specific superenhancer promotes polyunsaturated fatty-acid synthesis to support EGFR signaling. Cancer Discov. 9, 1248–1267 (2019).
Mukasa, A. et al. Mutant EGFR is required for maintenance of glioma growth in vivo, and its ablation leads to escape from receptor dependence. Proc. Natl Acad. Sci. USA 107, 2616–2621 (2010).
Schönrock, A. et al. MEOX2 homeobox gene promotes growth of malignant gliomas. Neuro Oncol. 24, 1911–1924 (2022).
Liu, D. D. et al. Purification and characterization of human neural stem and progenitor cells. Cell 186, 1179–1194 (2023).
Greenwald, A. C. et al. Integrative spatial analysis reveals a multi-layered organization of glioblastoma. Cell 187, 2485–2501 (2024).
Hai, L. et al. A clinically applicable connectivity signature for glioblastoma includes the tumor network driver CHI3L1. Nat. Commun. 15, 968 (2024).
Migliozzi, S. et al. Integrative multi-omics networks identify PKCδ and DNA-PK as master kinases of glioblastoma subtypes and guide targeted cancer therapy. Nat. Cancer 4, 181–202 (2023).
Verhaak, R. G. W. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Puchalski, R. B. et al. An anatomic transcriptional atlas of human glioblastoma. Science 360, 660–663 (2018).
Jin, S. et al. Inference and analysis of cell–cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Quail, D. F. et al. The tumor microenvironment underlies acquired resistance to CSF-1R inhibition in gliomas. Science 352, aad3018 (2016).
Gjørlund, M. D. et al. Neuroligin-1 induces neurite outgrowth through interaction with neurexin-1β and activation of fibroblast growth factor receptor-1. FASEB J. 26, 4174–4186 (2012).
Müller, T. et al. Neuregulin 3 promotes excitatory synapse formation on hippocampal interneurons. EMBO J. 37, e98858 (2018).
Venkatesh, H. S. et al. Targeting neuronal activity-regulated neuroligin-3 dependency in high-grade glioma. Nature 549, 533–537 (2017).
Krishnaswami, S. R. et al. Using single nuclei for RNA-seq to capture the transcriptome of postmortem neurons. Nat. Protoc. 11, 499–524 (2016).
Barthel, F. P. et al. Longitudinal molecular trajectories of diffuse glioma in adults. Nature 576, 112–120 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Germain, P.-L., Lun, A., Meixide, C. G., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res. 10, 979 (2021).
Gaujoux, R. & Seoighe, C. A flexible R package for nonnegative matrix factorization. BMC Bioinformatics 11, 367 (2010).
Spitzer, A., Johnson, K., Nomura, M. & Garofano, L. The multi-layered transcriptional architecture of glioblastoma ecosystems. Zenodo https://doi.org/10.5281/zenodo.14966015 (2025).
Acknowledgements
We thank R. A. DePinho, D. Arons and K. Knight for support with the GBM CARE initiative. This work was supported by a GBM CARE grant (supplemented to no. P30CA006516-56S6 (to M.L.S.)); M.L.S. is also supported by National Institutes of Health (NIH)-National Cancer Institute (NCI) grant nos. R01CA276765, R37CA245523, R01CA258763 and P50CA165962, the Mark Foundation Emerging Leader Award, the MGH Research Scholars Award, the Sontag Foundation Distinguished Scientist Alumni Award and the National Foundation for Cancer Research. I.T. is supported by a European Research Council consolidator grant no. 101044318, the Zuckerman STEM Leadership Program and a Mexican Friends New Generation grant, and he is the incumbent of the Dr. Celia Zwillenberg-Fridman and Dr. Lutz Zwillenberg Career Development Chair. A.I. is supported by NIH grant nos. R35CA253183, P30CA013696-46S1 and P30CA240139. R.G.W.V. is supported by NIH grant nos. R01CA237208, R21NS114873, R21CA256575 and P30CA034196, and by the Luxembourg National Research Fund. A. Lipsa, A.G. and S.P.N. are supported by C20/BM/14646004/GLASSLUX. A. Lasorella is supported by NIH grant nos. R01CA268592, R01CA280560 and R01CA239721. M.N. is supported by the JSPS Overseas Research Fellowship from the Japan Society for the Promotion of Science and the Overseas Research Fellowship from Uehara Memorial Foundation. A.S. is partially supported by the Israeli Council for Higher Education via the Weizmann Data Science Research Center. A. Lipsa, A.G. and S.P.N. are supported by the Luxembourg National Research Fund and TRANSCAN3 grants (C20/BM/14646004/GLASSLUX, INTER/TRANSCAN22/17612718/PLASTIG). L.N.G.C. has received honoraria from Elsevier, BMJ Best Practice, Oakstone Publishing and Servier, as well as research support from Merck, Conquer Cancer (The ASCO Foundation), the Robert Wood Johnson Foundation and the NCI. We thank the PROACTIVE team at MD Anderson for their assistance in procuring patient material. We gratefully acknowledge the contribution of The Jackson Laboratory Single Cell Biology service, S. Bessonett in particular, the Genome Technologies service and the Cyberinfrastructure high performance computing resources at The Jackson Laboratory for expert assistance related to processing the NORLUX/SNU samples and data analysis. These shared services are supported in part by The Jackson Laboratory Cancer Center (no. P30CA034196). We thank R. Matsuura in the neurosurgery laboratory, Tokyo University Hospital, for the technical assistance in preparing the clinical samples. We thank SciStories for the figure schematics. We thank the patients and their families for their generous donation to biomedical research.
Author information
Authors and Affiliations
Contributions
M.N., A.S., K.C.J., L.G., A. Lasorella, R.G.W.V., A.I., M.L.S. and I.T. conceived and designed the study. M.N., F.S.V., K.J.A. and Y.T.O. led the generation of data. A.S., K.C.J., L.G. and I.T. performed the computational analyses. D.N., N.G.D., A.C.G., L.B., F.S.V., F.D., S.G., K.J.A., S.M., F.D., L.N.G.C., T.C., N.R., C.R. and J.B.P. provided experimental and analytical support. A. Lipsa, F.H., A.G., S.P.N., L.N., S.K., S.D., H.E.M., S.H.P., F.B., A. Laurenge, A.L.D., B.M., A.P., M.S., S.T., N.S., D.M.A., S.T.K., J.T.H., W.K.A.Y. and K.L.L. provided the patient samples and curated the clinical annotation. M.N., A.S., K.C.J., L.G., A. Lasorella, R.G.W.V., A.I., M.L.S. and I.T. wrote the manuscript, with feedback from all authors. R.G.W.V., A. Lasorella, A.I., M.L.S. and I.T. jointly supervised the study and interpreted the results.
Corresponding authors
Ethics declarations
Competing interests
M.L.S. is equity holder, scientific cofounder and advisory board member of Immunitas Therapeutics. I.T. is advisory board member of Immunitas Therapeutics and scientific cofounder of Cellyrix. R.G.W.V. is a cofounder and equity holder of Boundless Bio. The authors declare that such activities have no relationship to the present study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Richard Gilbertson, Wolfgang Wick and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Study workflow and dataset overview.
a, Total counts (left), number of detected genes (middle), and mitochondrial expression proportion (right) per cell across the whole dataset. The vertical lines represent the 10% and 90% quantiles. b, Flow diagram of the CNA calling algorithm used to classify cells as malignant or non-malignant. CNA signal reflects the extent of CNAs on each chromosome per cell. CNA correlation reflects the similarity between the CNA pattern of each cell and the overall CNA pattern of the tumor. SNV - single nucleotide variation. c, Sample-level copy number aberrations. Rows represent samples, columns represent chromosomes, values reflect the average CNA signal across all cells classified as malignant in the tumor. d, Cost-effectiveness of the 2nd step of classification. Top panel shows the % of misclassified cells (that is cells harboring a malignant SNV that were classified as non-malignant in the 1st step) that remain for each CNA correlation threshold. Bottom panel shows the % discarded cells for each CNA correlation threshold. Dashed line marks the upper threshold for non-malignancy (0.35).
Extended Data Fig. 2 GBM cell type expression patterns and abundance.
a, UMAP colored by the lab that processed the tumor specimens and generated the 10x snRNA-seq data. b, Proportion (%) of samples contributing to each cluster of cells. Each dot represents a cluster and is colored by the dominant cell type in the cluster (n = 195, 11, 6, 6, 43, 5, 45, 6 for Malignant, Excitatory neuron, Endothelial, Astrocyte, Oligodendrocyte, Inhibitory neuron, TAM and Lymphocyte cell types respectively). Y axis reflects the proportion (%) of samples (out of all samples in which the cell type was detected) that contribute to the cluster. c, Heatmaps representing the average gene expression level (log2 relative expression) of cell type marker genes within each TME cell type for the 10x dataset. d, Gating strategy for single nuclei sorting for Smart-seq2. First, ruby positive proportion (P1, top) was selected by 570 nm laser. Then, doublet discrimination was performed by strict forward scatter height (FSC-H) versus area (FSC-A) criteria (P2, bottom left). It was further gated by removing 480 nm laser positive fraction was to exclude false positive proportion (P3, bottom right), and was sorted into 96 well plates for Smart-seq2. e, Heatmaps representing the average gene expression level (log2 relative expression) of cell type marker genes within each TME cell type for the Smart-seq2 dataset. Data generated at the two different labs was analyzed separately. f, Pie chart demonstrating the proportion (%) of each cell type in the Smart-seq2 dataset (two labs data combined, left), and 10x dataset considering only the same samples profiled also by Smart-seq2 (right). g, Boxplots of cell type proportions (%) in each tumor (n = 44) in both 10x (red) and Smart-seq2 (SS2, blue). Pearson’s correlations of the proportions of each cell type are reported below each panel. Lines connect the cell type proportions of the patient’s tumor profiled with both 10x and SS2.
Extended Data Fig. 3 Revisiting malignant state definition, and contextualization with state-of-the-art classification schemes.
a, Jaccard similarity indices of the robust NMF programs (n = 835) based on their top 50 genes. Programs are ordered by clustering and grouped into meta-programs (Methods). MP10 (Stress1) and MP15 (Stress2) were coalesced into a single Stress state. MP9 (ExN) and MP14 (NRGN neuron, NRGN) were coalesced into a single NEU-like state. MP1 (Ribosomal protein, RP), MP11 (Doublet), MP12 (Low quality, LQ) reflect low quality and are not considered a state. b, Correlation between cell state scores of meta-programs derived from this dataset (columns) and those derived by Neftel et al. (rows). Each cell was scored within tumor-of-origin for both sets of meta-program signatures. Pearson’s correlation coefficient was computed across all cells for each pair of meta-programs. c, Jaccard similarity between meta-programs derived from this dataset (columns) and those derived by Neftel et al. (rows). d, Distribution of scores in this dataset (upper panel) and Neftel et al. (lower panel) dataset of cells that scored maximally for both corresponding states (for example to both AC-like signatures derived from this dataset and from Neftel et al.) across all corresponding states. Dashed vertical lines mark the mean score of each distribution. e, Association between the malignant MPs from our datasets (rows) and malignant signatures from other studies (columns). Each cell in the heatmap represents the Pearson correlation coefficient between the gene expression scores of an MP and other signature scores across the malignant cells in the dataset.
Extended Data Fig. 4 Consistency of meta-programs across platforms and technical covariates.
a, Percent cycling cells in each subpopulation across samples with at least 50 cells in any of the states (n = 84, 16, 51, 67, 60, 65, 61, 28, 70 for AC-like, Cilia-like, Hypoxia, MES-like, Stress, NEU-like, GPC-like, NPC-like and OPC-like states respectively). NPC-like (13.6%) vs. NEU-like (9.5%) p = 0.012, GPC-like (12.5%) vs. AC-like (1%) p < 2.2−16 and vs. MES-like (6.6%) p = 0.0001 (one-sided Wilcoxon rank sum test for all comparisons). b, Pearson’s correlation coefficient heatmap between GBM state signature scores defined in this dataset and published functional gene signature scores. The published signature name is followed by each study’s lead author in parentheses. c, Pearson’s correlation of the proportions (%) of cells classified in each MP per tumor in both 10x (rows) and Smart-seq2 (columns). d, Similarity of MPs derived using all samples (x-axis) and MPs derived using primary (untreated) samples. e, Proportion (%) of contributing samples to each MP (relative to the number of samples profiled by each lab). f, Proportion (%) of contributing samples to each MP (relative to the number of samples provided from each sample source).
Extended Data Fig. 5 Pathway-based meta-programs and putative cellular hierarchy.
a, Jaccard similarity indices of the robust NMF programs based on their top 50 pathways. Programs are ordered by clustering and grouped into Pathway-based metaprograms (PMPs) according to their functional activities. b, Clustering of the PMPs based on their pairwise Pearson correlations reveal functional activity groups. c, Scores of each cell for the different PMPs. Cells are aggregated by the PMP to which they were classified. “Unresolved” represents cells that could not be classified to a PMP. d, Average number of detected genes per sample (n = 121) for singular and hybrid cells. Two-sided t-test was used to test for statistical significance. e, Relative frequency (in log2 scale) of observed hybrids vs. expected technical hybrids (Methods) for each of the possible state pairs (order within a pair is meaningless). Dashed line marks log2(Fold Change) = 2 to aid in visualization. Statistical significance was assessed for each hybrid state pair using a one-sided Fisher’s exact test. P values were corrected for multiple testing using the Benjamini-Hochberg method. The bar plots were colored according to the BH-adjusted p-value.
Extended Data Fig. 6 BP of inter-tumor heterogeneity.
a, Jaccard similarity matrix of the PCA-derived gene programs colored by cluster (see Methods). b, GO enrichment results for C1 (BP-ECM), C3 (BP-Neuronal) and C5 (BP-Glial) program clusters. Nodes represent enriched terms. Node color represents the extent of statistical significance and size represents the number of overlapping genes. Edges connect terms with a Jaccard similarity above 0.2 c, Same as Fig. 3c but here shown are the three BP scores. d, Observed vs. expected within-sample standard deviation (SD) of BP scores. Observed SD was defined as the SD across state-specific pseudo-bulk profiles in each tumor, shown are n = 98 samples with at least 3 pseudo-bulk profiles. Expected SD was estimated by randomly sampling n = 100 times 3 pseudo-bulk profiles and computing the SD across these PBs. Statistical significance was assessed with a two-sided Wilcoxon rank sum test. Each box plot represents a specific group (that is observed or expected) that spans from the first to third quartiles, median values are indicated by a horizontal line, whiskers show 1.5× interquartile range, and each dot reflects a specific standard deviation value. e, Same as Fig. 3d but here shown are the individual state scores in the 3-axis plane.
Extended Data Fig. 7 De-novo BP of inter-tumor heterogeneity in spatial transcriptomic data.
a, Same as Fig. 3c for the BP derived de-novo from the spatial transcriptomics data. For each tumor sample on the x-axis, the points represent the scores of the pseudobulk of that sample for a specific spatial equivalent of a state (Methods). b, Pearson’s correlation between scores of de-novo spatial BPs (rows) and those derive from the snRNAseq data (columns) with p-values (unadjusted for multiple testing) for each comparison in the tile. c, Euclidean distance between BP scores of samples resected from the same tumor (n = 6 distances from 4 tumor samples) and samples resected from different tumors (n = 225 distances from 22 tumor samples). Each point represents a Euclidian distance between BP scores of two samples. Statistical significance was estimated using a two-sided t-test. d, TCGA subtype and the association with BP assignment. The samples profiled by both bulk RNA-seq and snRNA-seq were used (n = 28).
Extended Data Fig. 8 Heterogeneity within the microenvironmental cell states.
a-b, Gene ontology term (rows) enrichment (log10 of adjusted p-values) for each metaprogram’s 50 gene list (columns) against background for a, TAMs and b, astrocytes. Statistical significance was assessed with a two-sided Fisher’s exact test and Benjamini-Hochberg p-value adjustment was performed. c-d, Pairwise Pearson correlation heatmap for all c, CARE TAMs and d, astrocytes scored for both this dataset’s MPs as well as publicly available cell type-specific gene signatures. e, Jaccard similarity index representing the intersection of genes for both malignant and non-malignant (astrocytes and TAMs) metaprograms. 50 genes per metaprogram.
Extended Data Fig. 9 Co-occurrence of TME cell state abundance.
Correlation heatmap for the Pearson correlation between the relative TME cell state abundances (n = 121 samples). Only correlations with unadjusted P < 0.05 are shown and increasing size of circle reflects increasing absolute value of correlation coefficient.
Extended Data Fig. 10 Genetic associations with transcriptional states and GBM multi-layered ecosystems, and stereotype transcriptomic architectures in GBM.
a, Association between genetic aberrations and cell state abundances. Color intensity reflects the statistical significance from two-sided Wilcoxon rank sum tests comparing state abundances in tumors with and without each genetic alteration (log10 of adjusted p-values). b, Association between abundance of GPC-like cells and the representative genomic alterations in GBM (either of chr.7 gain, 10p loss, EGFR amplification, and TERT promoter mutations, n = 87, P = 0.003, two-sided Wilcoxon rank sum test). c, The difference of the BP scores between samples with and without the representative genomic alterations in GBM (either of chr.7 gain, 10p loss, EGFR amplification, and TERT promoter mutations). Two-sided Wilcoxon rank sum test (n = 95 samples with sufficient mutation, copy number, and malignant cell data) were not corrected for multiple hypothesis testing. Boxplots span from the first to third quartiles, median values are indicated by a horizontal line, whiskers show 1.5× interquartile range, and outlier points are not shown. The violin plots surrounding the boxplots represent the continuous distribution of the data. d, Same as in Fig. 6a but for samples that could not be classified to an ecosystem.
Supplementary information
Supplementary Tables (download XLSX )
Supplementary Table 1. Sample, sequencing and clinical metadata; Supplementary Table 2. Malignant MP gene lists; Supplementary Table 3. BP gene lists; Supplementary Table 4. MP gene lists for nonmalignant cells; Supplementary Table 5. Sample-level composition cluster, malignant state cluster and BPs; Supplementary Table 6. Ecosystem assignment.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nomura, M., Spitzer, A., Johnson, K.C. et al. The multilayered transcriptional architecture of glioblastoma ecosystems. Nat Genet 57, 1155–1167 (2025). https://doi.org/10.1038/s41588-025-02167-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02167-5
This article is cited by
-
Immunosuppressive mechanisms and therapeutic interventions shaping glioblastoma immunity
Nature Cancer (2026)
-
Increased neural excitability and glioma synaptic activity drives glioma proliferation in human cortex
Nature Neuroscience (2026)
-
A spatially resolved human glioblastoma atlas reveals distinct cellular and molecular patterns of anatomical niches
Nature Communications (2026)
-
Genomic heterogeneity drives distinct infiltration patterns in glioblastoma
Acta Neuropathologica Communications (2025)
-
Spatial transcriptomic analysis reveals lack of response to PD-1 blockade in recurrent glioblastoma
Acta Neuropathologica (2025)