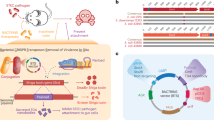
Abstract
Studying the pathogenesis of human-adapted microorganisms is challenging, since small animal models often fail to recapitulate human physiology. Hence, the comprehensive genetic and regulatory circuits driving the infection process of principal human pathogens such as Shigella flexneri remain to be defined. We combined large-scale Shigella infections of enteroids and colonoids with transposon-directed insertion sequencing and Bayesian statistical modeling to address infection bottlenecks, thereby establishing the comprehensive genome-wide map of Shigella genes required to infect human intestinal epithelium. This revealed the Shigella virulence effectors essential for epithelial cell colonization across geometries and intestinal segments, identified over 100 chromosomal genes involved in the process and uncovered a post-transcriptional mechanism whereby tRNA-modification enzymes and differential codon usage exert global control of a bacterial virulence program. Our findings provide a broadly applicable framework for combining advanced organotypic tissue culture with functional genomics and computational tools to map human–microorganism interactions at scale.
Similar content being viewed by others
Main
Shigella flexneri is a principal human-/primate-restricted pathogen responsible for bacillary dysentery (reviewed in refs. 1,2). As in other human-adapted microorganisms, in vivo molecular studies addressing Shigella pathogenesis have been challenging due to the shortage of animal models that mimic human physiology (reviewed in refs. 3,4). Previous work has identified Shigella virulence factors promoting invasion, cytosolic colonization and spread within nonphagocytic cells, reliant on a type-3-secretion-system (T3SS) and cognate effectors (reviewed in refs. 5,6). Most of these virulence factors are encoded on a large virulence plasmid (pINV) and their expression is tightly controlled by environmental stimuli7,8. Whereas tumor-derived cell lines have been used to map contributions of Shigella T3SS effectors and select chromosomal loci to invasive behavior (reviewed in refs. 9,10), these models show transformed features (for example, ref. 11), having altered morphology, metabolism, signaling and cell death pathways compared to native epithelia. Limitations of animal models have also hampered the application of in vivo high-throughput forward genetics screens. Therefore, the contribution of most of Shigella’s genome to colonization of a physiologically arranged gut epithelium remains undetermined.
Intestinal epithelial organoids (for example, enteroids and colonoids)—three-dimensional (3D) gut epithelial assemblies derived from stem cells12—have become promising tools for host–pathogen interaction studies13,14,15,16,17,18,19,20. Shigella preferentially infect intestinal epithelial cells (IECs) from the basolateral side following epithelial traversal21. Therefore, we developed a large-scale infection protocol for 3D-suspension-grown human enteroids with basal-out (BO) polarity16,22 and combined this with transposon-directed insertion sequencing (TraDIS)23—a functional genomics approach coupling transposon mutagenesis with next-generation sequencing to link effects of gene disruptions to specific phenotypes.
A key challenge in applying genome-scale screening to an infection assay is the presence of population bottlenecks23. Bottlenecks occur in infection models when factors such as barrier defences, nutritional restriction or competition for colonization sites lead to transient population size reduction irrespective of the fitness of individual mutants24. This causes stochastic loss of mutants without a true virulence phenotype, thereby distorting results. To overcome this challenge, we systematically optimized the experimental procedures and developed a Bayesian statistical framework25 based on the zero-inflated negative binomial (ZINB) distribution to extract mutant fitness parameters. We applied this to analyze 43 parallel enteroid bulk infections with Shigella Tn5 mutant sub-libraries, establishing a deep genome-wide map of Shigella genes required to colonize nontransformed gut epithelium. Informed by the TraDIS results, we mapped and quantified differences in colonization efficiency of Shigella wild-type and virulence gene mutants across cell geometries and intestinal segments, performing barcoded infections in enteroids and colonoids exposing the basal or apical surface outwards26,27. Finally, this work identified a post-transcriptional mechanism, linking MnmE/G U34-dependent tRNA modifications to global control of the Shigella T3SS virulence cascade. This study expands our understanding of enterobacterial pathogenesis and provides a generalizable framework for genome-wide forward genetics screens in complex infection models.
Results
A scalable human gut epithelium model for Shigella infection
To establish a large-scale human intestinal epithelial model for Shigella infection, we developed a high-throughput gentamicin protection assay, using suspension cultures of enteroids with BO polarity16,22 (Extended Data Fig. 1a). To explore Shigella infection dynamics, we infected the enteroids with wild-type Shigella flexneri M90T harboring the intracellular reporter puhpT-GFP13,28, or a constitutive pmCherry reporter29. At 3 h post-infection (h.p.i.) Shigella infection foci appeared in IECs. Between 3 and 9 h.p.i., foci increased in size and cell-to-cell spread was also evident (Fig. 1a,b and Supplementary Videos 1 and 2).
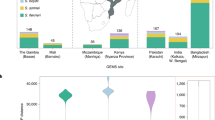
a,b, Representative time-lapse series of a BO enteroid infected with wild-type Shigella harboring cytosolic reporter puhpT-GFP (a) or constitutive reporter p-mCherry (b) at MOI 40. See also Supplementary Videos 1 and 2. Cartoon depicts the imaging plane. Experiment repeated twice with imaging of at least ten enteroids per experiment. Lu., lumen. c, Shigella CFU counts (top) and percentage of inoculum (bottom) upon coinfection of BO enteroids with wild-type (WT) and ΔmxiD (noninvasive) strains for 6 h at MOI 40; n = 7 biological replicates pooled from three independent experiments. Data shown as mean ± s.d. Significance determined by two-sided Mann–Whitney U test; ***P < 0.001. d, BO enteroids were infected with a Shigella Tn5 random mutant library containing ~10,000 or ~30,000 mutants. Shown is the number of unique TIS mapped in input (dark gray) or intracellular population output libraries (red). e, Simulations of the number of unique TIS in output samples for bottleneck sizes up to 6,000 bacteria (solid lines) and the measured number of unique TIS (dashed lines). Gray area indicates the plausible range of bottleneck sizes based on the simulations. f, Shigella Tn5 random mutant library 1 (~130,000 mutants) was grown overnight at 30 °C or 37 °C and the mutant abundance compared by TraDIS with respect to the subculture (Methods). Shown is the number of significant advantageous (Adv) (blue; gene presence favors growth) or detrimental (Detr) (orange; gene presence supresses growth) genes; log2FC ≥ 1, FDR ≤ 0.01. g, Venn diagram of Shigella differentially expressed genes in the following comparisons: wild-type, 37 °C versus 30 °C (temperature-regulated genes), wild-type versus ΔvirF 37 °C (VirF-regulated genes) and wild-type versus ΔvirF 30 °C; log2FC ≥ 1, FDR ≤ 0.01. h, Graph showing relative mutant fitness for pINV-located genes (log2FC as noted in Extended Data Fig. 2d; Enr 37 °C versus Enr 30 °C) versus the respective expression changes (RNA-seq; log2FC between wild-type Shigella 37 °C versus 30 °C; Extended Data Fig. 2e). Significant pINV advantageous (dark blue; gene presence favors growth) or detrimental (dark orange; gene presence supresses growth) genes are shown. Two selected chromosomal virF regulator genes, cpxR and fis, also indicated in light orange; log2FC ≥ 1; FDR ≤ 0.01. i, Barcoded competition assay with a consortium comprising three Shigella wild-type (tagA, tagB, tagC), two ΔmxiD (tagD, tagE) and two ΔvirF (tagF, tagG) strains grown overnight at 30 °C or 37 °C. Left, percentage of each strain in the input populations. Right, quantification of relative strain abundances in the Shigella barcoded consortia cultures grown at 30° or 37 °C. Relative abundances normalized to the input inoculum. Data for three independently generated consortia. NS, not significant. j, As in i, but using a consortium comprising three Shigella wild-type (tagA, tagB, tagC), two ΔmxiD (tagD, tagE) and two ΔvirB (tagF, tagG) strains. Significance in i and j determined by two-sided paired t-test between normalized output and normalized input abundances (Methods). *P < 0.05. Panels a and b partially created using BioRender.com.
Using TraDIS to map the Shigella geneset necessary for enteroid colonization requires a well-controlled experimental design. We systematically addressed the following critical parameters: (1) the size of the intracellular bacterial population, (2) the impact of infection bottlenecks and (3) the effect of bacterial growth enrichment steps following infection23,30. To estimate the size of the intracellular bacterial population, ~800 enteroids were infected with a mix of wild-type and ΔmxiD (noninvasive; lacks a T3SS structural component) strains at multiplicity of infection (MOI) 40. At 6 h.p.i., we consistently retrieved ~100,000 colony-forming units (CFUs) per infection for the wild-type Shigella strain and >60-fold fewer CFUs for the ΔmxiD strain, providing a wide dynamic range for the assay (Fig. 1c). In this window of early epithelial colonization, the intracellular Shigella population was the product of active T3SS-invasion and around four rounds of replication, with only limited IEC death elicited (Extended Data Fig. 1b,e).
Infection bottlenecks can confound genome-wide screens and generate false positive results23,24,31. To quantify the infection bottleneck size, we generated a random Shigella Tn5 mutant library with, in total, ~130,000 mutants, covering both the chromosome and pINV (Extended Data Fig. 2a). Sub-libraries containing 10,000 or 30,000 mutants were used to infect single wells of enteroids in triplicate. Because of the limited size of the intracellular Shigella population, input and output libraries were enriched before sequencing. This revealed the presence of approximately 1,500–4,000 unique Tn5 mutants within the intracellular bacterial population retrieved at 6 h.p.i. (output), indicating a tight bottleneck (Fig. 1d). We next simulated output mutant library compositions for bottleneck sizes up to 6,000 bacteria by randomly selecting bacteria from the input libraries (Methods), and comparing this to the number of unique mutants recovered in the output libraries (Fig. 1e). Assuming no selection, this indicated a bottleneck size of ~2,000 bacteria at this experimental scale.
Shigella expresses its T3SS at 37 °C (refs. 1,8,32). This comes at a growth cost, evident from the enhanced growth of Congo Red negative (T3SS off) Shigella clones33, and dissected at the molecular level in other bacteria34,35,36. Therefore, the enrichment temperature could potentially affect the mutant composition of input and output libraries due to selective growth effects. To test the global impact of enrichment temperature, we pooled the 130,000 Shigella Tn5 mutants and grew the resulting cultures at 30° (T3SS cascade off) or 37 °C (T3SS cascade on). Transposon insertions in a large number of genes (~1,200) led to differential fitness during enrichment at 37 °C, while approximately tenfold fewer gene disruptions (~130) affected fitness at 30 °C (Fig. 1f, Extended Data Fig. 2b–d and Supplementary Data 1). To further link bacterial fitness and T3SS virulence gene expression, we performed RNA-sequencing (RNA-seq) in a wild-type Shigella strain grown at 30° or 37 °C (temperature-regulated genes) and a wild-type Shigella strain versus a ΔvirF mutant (lacking the master regulator of the T3SS cascade), grown at 37 °C (VirF-regulated genes) (Fig. 1g, Extended Data Fig. 2e–g and Supplementary Data 2). By combining RNA-seq (gene expression, wild-type, 37° versus 30 °C) and TraDIS (fitness, enrichment at 37° versus 30 °C) data, we quantified the genome-wide cost of virulence gene expression in Shigella (Fig. 1h and Extended Data Fig. 2h). Among the T3SS-associated mutants, two—namely those with disrupted virF and virB—showed significantly better growth at 37 °C. This is particularly relevant since virF and virB are master regulators of the T3SS cascade and, at 37 °C, activate the structural T3SS apparatus genes and those encoding secreted effectors32. By contrast, little or no cost was associated with individual T3SS structure or effector genes (Fig. 1h). Other mutants with increased fitness at 37 °C exhibited disruption of virF transcriptional activators (for example, Δfis and ΔcpxR mutants)37,38,39 (Fig. 1h and Extended Data Fig. 2d–h).
For validation, we developed an internally controlled competition assay, encompassing defined consortia of genetically barcoded wild-type and mutant Shigella strains and quantification by quantitative PCR (qPCR; Extended Data Fig. 2i,j) (informed by refs. 26,40). Here, we mixed 1:1:1:1:1:1:1 consortia, containing seven strains: three wild-type (tagA, tagB, tagC), two ΔmxiD (tagD, tagE) and two ΔvirF (tagF, tagG) or two ΔvirB (tagF, tagG) strains. All strains were represented equally in the input consortia (Fig. 1i,j). We then grew these overnight at either 30° or 37 °C. Again the ΔvirF and ΔvirB mutants were markedly overrepresented at 37 °C compared to wild-type and ΔmxiD strains, whereas no difference in relative strain abundance was observed at 30 °C (Fig. 1i,j). Altogether, we successfully mapped and quantified the cost of temperature-induced T3SS gene expression in Shigella, expanding previous gene essentiality studies41. Moreover, performing the bacterial input and output enrichment at 30 °C seems critical to avoid biased results from TraDIS experiments in Shigella-infected enteroids, or other infection models.
Shigella genome-wide map of enteroid colonization genes
To define the Shigella factors required to colonize human epithelium, we scaled the enteroid infection assay to a genome-wide screen using Tn5 mutant libraries. We optimized the infection assay (Fig. 2a) based on pilot experiments (Fig. 1) to balance cost, experimental throughput and ease of analysis. Given the infection bottleneck of ~2,000 bacteria with 1,000–2,000 unique insertion sites recovered after infection (Fig. 1d,e), we expected sub-libraries with 2,000–3,000 unique Tn5 mutants to limit stochastic loss to about 50%. Hence, approximately 40 sub-libraries would result in coverage of at least 80% of the genome (Extended Data Fig. 3a,b and Methods) (43 sub-libraries opted for). Each sub-library was used to infect a single batch of ~800 enteroids (Fig. 2a and Extended Data Fig. 3c). At 6 h.p.i. we recovered 40,000–200,000 intracellular bacteria per infection (Extended Data Fig. 3d). We then enriched intracellular (output) and inoculum (input) bacteria for 14 h at 30 °C to minimize selection bias. As expected, the unique insertion sites of any single sub-library did not cover the genome evenly (Fig. 2b, blue), but a combination of all input sub-libraries provided complete coverage of the chromosome and pINV (Fig. 2b, gray), with sequencing reads distributed evenly across 42 of the 43 samples (Fig. 2c). This approach allowed screening of ~112,700 mutant clones (Extended Data Fig. 3c), and quantification of ~85,000 unique Shigella Tn5 mutants for colonization defects (by TraDIS), covering 86% of annotated genes with at least one insertion site (Extended Data Fig. 3e).
a, Schematic of the optimized protocol used for the Shigella TraDIS screen in human BO enteroids. b, Shigella M90T chromosome and pINV maps showing the distribution of TIS across the 43 input sub-libraries (gray) versus input sub-library 1 only (blue); ori, origin of replication for the chromosome and pINV; ER, entry region for the pINV. c, Number of unique TIS and total number of sequencing read counts across all input and output sub-libraries. Sub-library 11 was removed in the subsequent analysis due to the low number of sequencing read counts in the input sample. d, Schematic of the ZINB model developed to map Shigella genome-wide colonization factors in the presence of infection bottlenecks. Workflow shows the TraDIS raw dataset used for the sequential fitting of ZINB model parameters. The ZINB model was first applied to TIS in pseudogenes only to fit technical parameters (pairwise normalization factors and the dependence of stochastic mutant loss on input abundance αrgk,in and the fraction of zeros in the output sample fz,r). These fitted parameters were then used in a second ZINB model applied to all ~85,000 TIS to extract gene-wise log2FC values and determine significance. e,f, MA plots showing chromosome-located (e) or pINV-located (f) Shigella gene mutant relative abundances upon infection of enteroids, as quantified by TraDIS. Shown is the log2FC in the output library on the y axis and the log2CPM (average log2-transformed counts per million) on the x axis as detailed in d; each dot represents a gene. CDS, coding sequence; Pseudo, yellow, pseudogene. Panel a partially created using BioRender.com.
To enable robust statistical analysis, we devised a Bayesian model tailored to our experimental design (Fig. 2d, Extended Data Fig. 3f, Supplementary Note 1 and Methods). Each sub-library has a unique mutant composition, yielding a pairwise structure (Extended Data Fig. 3g) that requires normalization. We estimated gene-wise log2-transformed fold change (log2FC) across all insertion sites in a gene, assuming that all Tn5 mutants have similar effects on colonization. To handle stochastic mutant loss, we used a ZINB distribution combining the negative binomial model for count data with a zero-inflation component to capture excess zeros (Extended Data Fig. 3h). Observed zero rates (25–75%, Extended Data Fig. 3i), matched expectations from bottleneck size estimates, and increased with lower mutant abundance (Extended Data Fig. 3j). We followed a two-step process to extract global gene fitness measurements (Fig. 2d). Initially, pairwise normalization factors and technical parameters, such as the abundance-dependence of mutant loss (Extended Data Fig. 3f), were derived by fitting a ZINB model to 769 pseudogenes assumed to be neutral. Subsequently, these parameters informed a second ZINB model applied to all genes to extract gene-wise log2FCs, representing the differential abundance in the output samples adjusted for stochastic loss (Extended Data Fig. 3j). We calculated z values from the log2FC posterior distributions (Fig. 2d) and assigned a false discovery rate (FDR) to correct for multiple hypothesis testing42 (Extended Data Fig. 3k,l, Supplementary Note 1 and Methods). This allowed us to systematically evaluate the effect of gene disruption on enteroid colonization genome-wide, accounting for infection bottlenecks and maintaining a stringent FDR.
Applying this framework, we found 105 chromosomal genes to be advantageous for Shigella enteroid colonization (that is, mutants less invasive; 62 genes with FDR < 0.1 and 43 additional genes with FDR < 0.2) (Fig. 2e and Supplementary Data 3 and 4), a large fraction of which not previously linked to Shigella epithelial colonization (>70; Supplementary Data 4). Among the advantageous genes previously unreported were those encoding the stress regulator RpoS and anti-adapter protein IraM, the polyribonucleotide nucleotidyltransferase PNPase, several transporters (NikB, FruB, NagE, ArgO, YfeH, BrnQ, YhjX, TsgA and CusA), the beta-ketoacyl-ACP synthase II FabF, tRNA-modification enzymes (MnmE and MnmG) and sulfur metabolism-associated genes (yftE, cysI, cysQ and yeeE). Seven mutants belonging to transmembrane transporter complexes (ΔmlaC, ΔmlaE, ΔmlaF, ΔpbgA, ΔacrA, ΔacrB and ΔtolC), were essentially lost from the output (intracellular bacteria) population. Follow-up barcoded consortium infections showed that the ΔacrA, ΔacrB, ΔacrAB and ΔtolC mutants were attenuated for enteroid colonization and also sensitive to several infection assay conditions, extending previous observations43 (Extended Data Fig. 4a–g).
Moreover, 61 chromosomal genes decreased the colonization ability of Shigella (that is, mutants enriched in intracellular population; 49 with FDR < 0.1; 12 with FDR < 0.2; Fig. 2e). Among these, we identified LPS and O-antigen biosynthesis genes (waaO, rfbD, galU, waaJ, rfbB, waaD, rfaL, rfbA and rfbC) (Supplementary Data 4). These mutants probably harbor a shorter LPS, favoring T3SS docking to the IEC membrane44,45,46, although LPS-affecting mutants may be attenuated in vivo owing to poor cell envelope protection44,45,46,47.
On pINV, 38 genes scored as required or advantageous for enteroid colonization (34 with FDR < 0.1; 4 with FDR < 0.2) (Fig. 2f). Among them were the virF and virB transcriptional regulator genes, most genes encoding T3SS structural components (including mxiD, deleted in our noninvasive control strain in Fig. 1) and icsA, involved in actin-based intracellular spread. This comprehensive coverage of the core T3SS machinery demonstrates the high level of saturation attained in the screen. In conclusion, we successfully combined TraDIS with high-throughput enteroid infections to generate, to our knowledge, the first genome-wide map of Shigella genes required to colonize nontransformed human gut epithelium.
Shigella T3SS effector requirements across epithelial contexts
The TraDIS screen revealed Shigella genes required for enteroid colonization from the basal side (Fig. 2e,f). For T3SS-related components, in addition to the IpaBCD translocon, we found a set of secreted effectors (ipgB1, ipgD, icsB, ipaA and virA) to promote colonization (Figs. 2f and 3a). Previous studies have explored the dependence of Shigella on T3SS effectors for epithelial cell colonization, typically using tumor-derived cell lines and sometimes with contrasting results (reviewed in ref. 9). Moreover, Shigella invasion seems less efficient from the apical than from the basal side of the epithelium14,15,21. In follow-up experiments, we found Shigella intracellularly in infected human 3D enteroids and colonoids showing either BO or AO polarity (Fig. 3b–e). A similar number of wild-type Shigella could be recovered from BO enteroids and BO colonoids when coinfected with a mix of wild-type and ΔmxiD strains (Fig. 3f–h). Shigella colonization of AO enteroids was around fivefold less efficient, and in AO colonoids ~15-fold less efficient than in the corresponding BO counterparts (Fig. 3f–i). In all cases, a consistent and large (≥56-fold) difference in CFUs was evident between the wild-type and the ΔmxiD noninvasive strain (Fig. 3f–i).
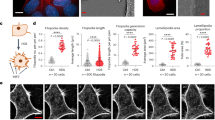
a, Heat map showing log2FC for T3SS effector gene mutants, as informed by the TraDIS screen in Fig. 2f; **FDR < 0.1; *FDR < 0.2. b–e, Representative confocal fluorescent microscopy images of a BO enteroid (b), AO enteroid (c), BO colonoid (d) and AO colonoid (e) infected with wild-type Shigella harboring cytosolic reporter puhpT-GFP at MOI 40. Experiments were repeated at least twice with imaging of at least ten enteroids or colonoids per experiment. f–i, Shigella CFU counts (left) and percentage of inoculum (right) upon coinfection with wild-type and ΔmxiD (noninvasive) strains for 6 h at MOI 40 of BO enteroids (f, same plots as in c), AO enteroids (g), BO colonoids (h) and AO colonoids (i); n = 7 biological replicates pooled from two to three independent experiments. Data shown as mean ± s.d. Significance determined by two-sided Mann–Whitney U-test; ***P < 0.001. j–m, BO and AO enteroids (j) and BO and AO colonoids (l) were infected with a barcoded consortium comprising two wild-type (tagA, tagB), two ΔmxiD (tagC, tagD) and two mutant (tagE, tagF) strains at MOI 40 for 6 h. Shown is the quantification of relative tag abundance, normalized against the corresponding input (Extended Data Fig. 5a,b). Data represent two independently generated consortia for each infection. Significance determined by two-sided paired t-test for each specific mutant between normalized output and normalized input abundances (Methods). *P < 0.05, **P < 0.01, ***P < 0.001. k–m, Bargraph showing the colonization index for each mutant in BO (dark gray) and AO (light gray) enteroid infections (k, derived from data in j and calculated as 1 – (WT/mutant); n as in j) and BO (dark gray) and AO (light gray) colonoid infections (m, derived from data in l; n as in l). Data shown as mean ± s.d. Wild-type shown as 0.
To substantiate and extend the TraDIS results (Figs. 2 and 3a), we quantified differences in IEC colonization efficiency of wild-type Shigella and clean T3SS effector mutants by barcoded consortium infections in BO enteroids (same condition as the TraDIS screen), as well as AO enteroids, BO colonoids and AO colonoids. For each infection, an equally mixed (1:1:1:1:1:1) inoculum of six strains, two wild-type (tagA, tagB), two ΔmxiD (tagC, tagD) and two mutant strains for a given effector (tagE, tagF) was prepared. This setup thus used two biological replicates for each T3SS genotype, allowing powerful internally controlled comparisons. We first infected BO and AO enteroids. All strains were represented equally in the input inocula (Extended Data Fig. 5a). In the intracellular population, the two noninvasive ΔmxiD control strains were approximately 100-fold to 1,000-fold less abundant than the two wild-type strains (Fig. 3j). To visualize colonization differences between strains, we calculated a colonization index for each mutant (1 − (mean relative abundanceWT/mean relative abundancemutant)) in relation to wild-type Shigella (value 0). Deletion of ipgD and icsB resulted in a minimal but significant colonization reduction in BO enteroids. In AO enteroids, deletion of ipgD did not cause significant reduction of colonization, whereas icsB deletion again showed a minor contribution (Fig. 3j,k). By contrast, deletion of ipgB1 attenuated colonization (around twofold in BO and around fourfold in AO enteroids), and the combined deletion of IpgB1 and IpgB2 (ΔipgB1/B2) even more so (around fourfold in BO and greater than tenfold in AO enteroids). Deletion of IpaA (ΔipaA) had a minor impact in BO enteroids (mutant ~30% less invasive than wt), but caused an attenuation in AO enteroids of about fivefold. Moreover, deletion of VirA caused an attenuation in both BO and AO enteroids of approximately twofold. We next used the same consortia to infect BO and AO colonoids (Fig. 3l,m and Extended Data Fig. 5b). Here, deletion of ipgB1 and ipgB1/B2 had an even bigger effect on Shigella colonization. A negligible contribution could again be observed for IpgD and IcsB and a modestly stronger one for VirA (Fig. 3l,m). Finally, deletion of IpaA caused markedly attenuated colonization, particularly in AO colonoids, similar to the enteroid infections (compare Fig. 3l,m with Fig. 3j,k). This phenotype was also validated in additional enteroid and colonoid lines from independent donors (Extended Data Fig. 5c–f).
We conclude that the TraDIS screen captured even minor contributions of Shigella genetic elements to the colonization process (compare Figs. 2f and 3a to Fig. 3j; BO enteroids). Moreover, barcoded infections reveal that a minimal set of T3SS effectors required for early Shigella colonization of a nontransformed human intestinal epithelium includes four main effectors: IpgB1 and IpgB2 (Rho GTPase GEFs; see refs. 48,49,50), IpaA (a vinculin-binding and actin-depolymerizing protein; see refs. 51,52) and VirA (a Rab GAP; see refs. 53,54). We found no obvious difference in the overall effector requirement for colonization of jejunum- or colon-derived IECs. However, our results identify a context-dependent requirement for IpaA specifically during apical epithelial invasion.
MnmE/G tRNA-modification enzymes exert global virulence control
We next scrutinized the TraDIS results with a focus on chromosomal genes (Fig. 2e and Supplementary Data 3 and 4). Clean barcoded Shigella mutants for select genes were constructed and BO enteroid infections with mixed consortia conducted as above, which replicated weak hits (for example fabF, yfeH), and nonsignificant findings from the TraDIS (for example, for cysK, fliM, flgA and fliH) (Extended Data Fig. 6a–c). Most interestingly, the TraDIS results suggested the tRNA-modification enzymes MnmE and MnmG to promote Shigella enteroid colonization (Fig. 2e and Supplementary Data 3 and 4). The MnmEG complex modifies position 5 of the wobble uridine (U34) in the anticodon of specific tRNAs, namely tRNAArg-mnm5UCU, tRNAGly-mnm5UCC, tRNALeu-cmnm5UmAA, tRNAGln-cmnm5s2UUG, tRNAGlu-mnm5s2UUC and tRNALys-mnm5s2UUU55,56 (Fig. 4a). The related enzymes MnmA and MnmC drive further U34 modifications55,56. Tn5 insertions in mnmA were rare in the input, whereas mnmC insertions did not affect Shigella colonization (Supplementary Data 3). By contrast, the ΔmnmE and ΔmnmG mutants showed a slight growth defect (Extended Data Fig. 6d) and a striking six- to tenfold lower colonization capacity than wild-type strains in barcoded consortium infections, and in CFU plating assays (Fig. 4b and Extended Data Fig. 6e).
a, Schematic of MnmE/G-dependent tRNA modifications. b, BO enteroids were infected with barcoded consortia comprising two wild-type (tagA, tagB), two ΔmxiD (tagC, tagD) and two mutant (tagE, tagF) strains at MOI 40 for 6 h. Left, quantification of relative tag abundance, normalized against the corresponding input (Extended Data Fig. 6d). Data for three independently generated consortia for each infection. Significance determined by two-sided paired t-test between normalized output and normalized input abundances. **P < 0.01. Right, colonization index for the ΔmnmE and ΔmnmG mutants (derived from data in left panel). Data shown as mean ± s.d. c,d, Volcano plots showing differentially abundant proteins in Shigella ΔmnmE versus wild-type (c) or ΔmnmG versus wild-type (d) strains. Each dot represents a protein. Differentially abundant proteins encoded on pINV shown in pink, on chromosome in black and all nonsignificant differentially abundant proteins in gray. Significance determined by two-sided limma analysis with Benjamini–Hochberg multiple comparison correction; log2FC ≥ 0.5; adjusted P value ≤ 0.01. e,f, Plots showing relative protein levels (log2FC as in c, ΔmnmE versus wild-type) versus AGA codon usage ratios for Arg (e) and GGA codon usage ratios for Gly (f) per protein. Differentially abundant proteins encoded on pINV shown in pink, on chromosome in black, and nonsignificant differentially abundant proteins in gray; log2FC ≥ 0.5; adjusted P value ≤ 0.01. Dark and light green dashed lines specify mean AGA/GGA codon usage ratios for all open reading frames on pINV or chromosome, respectively. g–i, ipaA (g), mnmE (h) and mnmG (i) mRNA levels (2–ΔΔCt) upon induction with 0.25 mM IPTG in the indicated strains carrying an inducible ipaA-3xFT plasmid, p-Empty or p-mnmE or p-mnmG, normalized to the corresponding mRNA levels in wild-type Shigella. j, Relative IpaA-3xFT protein levels (0.25 mM IPTG) in wild-type Shigella, ΔmnmE or ΔmnmG strains carrying plasmids as in g–i. Quantification by western blot of serially diluted samples with wt/ipaA-3xFT protein levels set as 1. Data shown as mean ± s.d. of three independent experiments (Extended Data Fig. 7a,b). k,l, BO enteroids were infected with barcoded consortia comprising the indicated strains at MOI 40 for 6 h. Shown is the quantification of relative tag abundance, normalized on the corresponding input (Extended Data Fig. 7c,d). Data for two independently generated consortia per infection. Significance determined by two-sided paired t-test between normalized output and normalized input abundances. Only consortia including the two mutant strains carrying the p-Empty or p-mnmE (k) p-mnmG (l) plasmids were used in the analysis. *P < 0.05, **P < 0.01, ***P < 0.001. m,n, mnmE (m) and mnmG (n) mRNA expression (2–ΔΔCt) in the indicated strains, normalized to mnmE (m) and mnmG (n) expression in wild-type Shigella. For g,h,i,m and n, n = 3 biological replicates. Data shown as mean ± s.d. Panel a partially created using BioRender.com.
Suspecting that MnmE and MnmG act through translation effects, we conducted proteome profiling. For the 2,153 identified proteins in the ΔmnmE mutant, 154 proteins were significantly downregulated, and 71 significantly upregulated in comparison to wild-type strains (Fig. 4c and Supplementary Data 5; log2FC ≥ 0.5; adjusted P value ≤ 0.01). Essentially identical results were obtained for the ΔmnmG proteome (158 downregulated; 72 upregulated; Fig. 4d, Extended Data Fig. 6f,g and Supplementary Data 5). Among the downregulated proteins, 25 were virulence proteins encoded on pINV, including the VirB transcriptional regulator, T3SS structural components (for example, IpaB, IpaC, MxiA, MxiC, MxiH, MxiN, SpaM, Spa33 and Spa47), early effectors (for example, IpaA and IpgB2) and late effectors (for example, OspC2, OspC3, OspF and PhoN2) (Fig. 4c, d). This offers a compelling explanation for the reduced colonization capacity of ΔmnmE and ΔmnmG strains.
The MnmE/G U34-dependent tRNA modifications xm5U ensure translation accuracy and fidelity. They stabilize base pairing between codons ending in A or G (NNA and NNG) with the corresponding anticodons, while hampering base pairing with codons ending in C and U57. This led us to ask whether codon usage correlates with Shigella virulence protein expression and its dependence on mnmE and mnmG. Analysis across the Shigella genome revealed that pINV genes showed higher frequency for AGA-Arg, AGG-Arg, GGA-Gly and AUA-Ile codons when compared to chromosomal genes (Extended Data Fig. 6h). Notably, most of the downregulated virulence proteins in the tRNA modification mutants (21 out of 25) showed an even higher AGA-Arg codon usage ratio than the average ratio (0.19) of all pINV genes. This is particularly interesting since the arginine AGA codon-reading anticodon is modified by MnmE/G and AGA is among the rarest codons in Escherichia coli58. Indeed, this difference was even more evident when comparing the AGA codon usage ratio for virulence proteins with the average ratio (0.04) for genes on the Shigella chromosome (Fig. 4e and Extended Data Fig. 6i). Similar observations were true for the GGA-Gly codon usage ratio (Fig. 4f and Extended Data Fig. 6j). Among other codons modified by the MnmE/G complex, we also observed a mild enrichment for UUA-Leu and CAA-Gln codon usage ratio for downregulated virulence proteins, but no evident enrichment for GAA-Glu and AAA-Lys (Extended Data Fig. 6k–n). Moreover, AGG-Arg and GGG-Gly codons, which can potentially pair with MnmE/MnmG-modified U34 of tRNAArg-mnm5UCU and tRNAGly-mnm5UCC, were also slightly enriched among the downregulated virulence proteins (Extended Data Fig. 6o,p).
To formally test the link between MnmE/G U34 tRNA modifications and Shigella virulence protein expression, we selected IpaA as a prototypical T3SS protein with a high AGA and GGA codon usage (Fig. 4e,f). We assessed ipaA/IpaA mRNA and protein production following inducible induction with isopropyl β-d-1-thiogalactopyranoside (IPTG), finding comparable mRNA levels across all strains tested (Fig. 4g). However, IpaA protein levels were decreased strongly in the ΔmnmE and ΔmnmG mutants (Fig. 4j and Extended Data Fig. 7a,b). This indicates a direct MnmE/G effect on translation of AGA- and GGA-enriched transcripts. Complementation of mnmE/G restored IpaA protein to wild-type levels in the ΔmnmE mutant and partially so in the ΔmnmG mutant (Fig. 4j,h,i and Extended Data Fig. 7a,b). Complementation also completely restored the enteroid colonization defect of the ΔmnmE mutant, and to a high extent for the ΔmnmG mutant (Fig. 4k–n and Extended Data Fig. 7c,d; input). Hence, the combination of enteroid infections with a Shigella genome-wide TraDIS screen allowed us to identify a global mode of Shigella virulence control governed by selective codon usage and the MnmE/MnmG tRNA-modification enzymes.
Discussion
Historically, infection experiments aspiring to mimic host physiology have been possible using in vivo models or short-lived explants. Despite much recent progress59,60, no small animal model exists that fully recapitulates human shigellosis3. Organotypic tissue culture has opened new possibilities to explore microorganisms–host interactions while capturing central aspects of host cell and tissue physiology13,14,15,16,17,18,19,20,61,62,63. Here, we combined TraDIS with high-throughput infection assays in 3D enteroid/colonoid suspension cultures16,22 to map the comprehensive geneset Shigella uses to colonize human intestinal epithelium. Among limitations, it should be noted that the enteroids/colonoids still lack innervation, blood-derived cells and vasculature, and that the short infection window of the screen limits the capture of phenotypes linked to immune evasion and cell death suppression. Nevertheless, our effort provides a key proof-of-principle for functional genomics approaches in organotypic infection models that future studies can build on for diverse pathogens and time-scales.
Common concerns when using TraDIS and similar screening techniques in an infection setting are: (1) the impact of enrichment of bacterial input and output populations and (2) the consequences of infection bottlenecks, which result in stochastically reduced genetic diversity24,64. We addressed these concerns to ensure the success of the screen. Combining RNA-seq (gene expression) and TraDIS (fitness), we quantified the condition-specific cost of virulence gene expression in Shigella. Besides providing a framework for measuring the effect of virulence gene expression on bacterial fitness in a genome-wide fashion, this analysis provided a key premise for the main TraDIS screen: performing enrichment growth step(s) at 30 °C instead of 37 °C leads to minimal bias in defining the Shigella geneset requirement for epithelial colonization.
Infection bottlenecks limit forward genetic screens in complex models24,65,66,67,68,69, including a previous screen using an infant rabbit shigellosis model64. To overcome this obstacle, we assessed bottlenecks experimentally and using computational simulations. Performing about 40 infections with Shigella Tn5 mutant sub-libraries with 2,000–3,000 mutants each achieved a balance between reducing bottleneck effects and experimental feasibility. Previous ZINB models to describe stochastic loss in single-cell RNA-seq70,71 and Tn-seq experiments72 fit a gene-dependent zero-inflation parameter, conflating stochastic loss with biological zeros. Using nonfunctional pseudogenes, we showed that stochastic loss depends on the input abundance of each mutant and varies between replicates. Our Bayesian model incorporates these gene-independent effects, enabling integration across experiments and identification of genuine phenotypes. The combination of experimental and computational optimization hence defined a Shigella epithelial infectome of 105 chromosomal and 38 pINV genes, a large fraction of which not previously linked to virulence.
Key findings from the TraDIS screen were validated and expanded upon by our Shigella barcoded consortium infection technology26,27, applied to BO and AO enteroids and colonoids16,22. This highlighted, for example, an impact of four main T3SS effectors: the actin ruffle-inducers IpgB1 and IpgB2 (refs. 48,49,50), the actin-depolymerization protein IpaA51,52 and the Rab inhibitor VirA53,54 for early IEC colonization across segments and geometries. Moreover, we observed a context-dependent requirement for IpaA specifically for Shigella apical invasion of enterocytes and colonocytes. Follow-up studies may decipher how these T3SS activities are coordinated, and how the IpaA apical invasion phenotype relates to the vinculin-binding property of this effector51,52.
Among the >100 chromosomal Shigella genes required for enteroid colonization, we focused on mnmE and mnmG, two tRNA-modification enzymes modifying U34 of various tRNA species55,56. Previous reports have linked virulence phenotypes to MnmE/G tRNA modifications in other bacteria73,74. In addition, MnmA contributed to Mycobacterium tuberculosis intracellular growth in macrophages75, while MiaA favored extraintestinal pathogenic E. coli (Ex-PEC) in vivo infections76. In Shigella, Tgt and MiaA have been shown to impact VirF translation77,78, and TruB slightly affected IpaB expression79. However, global effects of tRNA modifications on virulence gene expression have remained elusive. We show that pINV-encoded virulence proteins are globally downregulated in MnmE/G mutants. This reduction is associated with an increased fraction of AGA and GGA codons in the corresponding transcripts. Both AGA and GGA codons are translated by tRNAs carrying MnmE/G-dependent modifications80. This suggests direct MnmE/G-dependent global translation control of AGA and GGA codon-enriched transcripts encoded on pINV. Such stringent virulence expression control is crucial to balance virulence potential with the fitness cost of expressing these genes. The nucleoid associated protein H-NS silences transcription of horizontally acquired AT-rich sequences in bacterial pathogens37,81,82, and in Salmonella H-NS deletion results in severe fitness defects owing to aberrant T3SS expression83. In Yersinia, T3SS expression leads to growth arrest, with control mediated by virulence plasmid copy number regulation84,85. As shown here, expression of the Shigella virulence machinery causes growth defects at the inducing temperature in vitro, again highlighting the need for fine-tuned regulation. The MnmE/G-dependent regulation uncovered in Shigella represents a third globally acting mechanism for virulence gene expression control in an enterobacterium, made possible by differential codon usage between chromosomal genes and pINV-located virulence genes. Whether, and how, this regulatory circuit is modulated by environmental cues remains an intriguing topic for future studies.
Resolving molecular infection mechanisms is crucial to eradicate bacterial pathogens. Here, we derived a genome-wide map of Shigella genes required for colonization of a species-specific intestinal epithelial model by combining enteroid/colonoid infections, TraDIS, computational modeling, barcoded consortium infections and proteomics. This offers a generalizable framework to conduct genome-wide screens in complex infection models.
Methods
Ethics statement
Human adult stem cell-derived enteroids/colonoids were established from jejunal tissue resected during bariatric surgery (enteroids), or morphologically normal nontumor colon tissue resected during elective colon cancer surgery (colonoids), in all cases following the subject’s informed written consent. To ensure anonymity of tissue donors, samples were pseudonymized. All procedures were approved by the Swedish National Governing Body (Etikprövningsmyndigheten) under license/protocol 2010-157, 2010-157-1, 2020-05754 and 2023-01524-01.
Bacterial strains, plasmids and general procedures
Bacterial strains are listed in Supplementary Table 1. M90T, a Shigella flexneri serotype 5 strain7 was used as the wild-type strain and for all mutant construction. For construction of wild-type and ΔmxiD Shigella barcoded strains, TagA-G were integrated individually into an inert spot of the M90T chromosome, between genes ybhC and ybhB. A fragment containing an individual 40-bp tag and the chloramphenicol resistance cassette was amplified from Salmonella Typhimurium barcoded strains40 and used for chromosomal integration by a modified Lambda Red Recombination protocol. For the construction of ΔvirF, ΔvirB, ΔyfeH and ΔyejB barcoded strains, tagF and tagG were transferred by P1 transduction from donor wild-type Shigella barcoded strains, followed by selection on LB agar containing 12.5 µg ml–1 chloramphenicol. For all other Shigella barcoded mutants, the desired deletion was introduced by Lambda Red recombination in the relevant wild-type Shigella barcoded strains86. p-Empty is a de novo synthetized cloning vector carrying p15A ori and AmpR. In vitro synthesized DNA fragments containing mnmE and mnmG including their native promoters were cloned in the BamHI site of the p-Empty vector, obtaining p-mnmE and p-mnmG. pTac-28 is a de novo synthesized IPTG-inducible expression vector carrying a Ptac promoter and a lacI gene, together with a pMB1 ori and CmI resistance. An in vitro synthesized ipaA-3xFT DNA fragment was cloned into the BamHI site of the pTac-28 vector, resulting in the pTac-ipaA-3xFT plasmid. Bacteria were grown in Luria Bertani (LB) broth at 37 °C, unless otherwise specified. When necessary, antibiotics were supplemented at the following concentrations: chloramphenicol, 12.5 μg ml–1; kanamycin, 50 μg ml–1; ampicillin, 50 μg ml–1. All plasmids and oligonucleotide primers used in this study are listed in Supplementary Tables 2 and 3, respectively.
Shigella Transposon 5 library construction
Two Shigella Tn5-libraries were generated using EZ-Tn5 <KAN-2>Tnp Transposome kit (Lucigen, TSM99K2). In brief, 50 µl aliquots of Shigella M90T wt electrocompetent cells were electroporated with 1 µl of EZ-Tn5 <KAN-2>Tnp Transposome. Each reaction was recovered in 1 ml SOC (2% tryptone, 0.5% yeast extract, 10 mM NaCl, 2.5 mM KCl, 10 mM MgCl2, 10 mM MgSO4, 20 mM glucose) for 1 h at 37 °C, and plated on LB agar plates containing 50 µg ml–1 kanamycin. For library 1 (used in the pilot screen and under different growth conditions), on the following day transformants were scraped off the LB plates and pooled into 23 sub-libraries of approximately 4,000–7,000 transformants for a total of approximately 130,000 mutants. For library 2 (used in the main TraDIS screen), transformants scraped off the LB plates were pooled into 43 sub-libraries of approximately 2,000–3,000 transformants for a total of ~112,700 mutants (Extended Data Fig. 3c).
Enteroid and colonoid maintenance culture
Human jejunal enteroids (pseudonym ID, 18-9jej; 22-2jej) and human colonoids (pseudonym ID, 21-9col; 22-4col) were used in this study and handled as previously described13,62,63. For maintenance cultures, human enteroids/colonoids embedded into Matrigel domes (Corning, 356230) were subcultured weekly by mechanical dissociation and incubation with gentle cell dissociation reagent (StemCell, 07174). The resulting fragments were washed with Dulbecco’s modified Eagle’s medium (DMEM-F12; Gibco, 11039021) containing 0.25% bovine serum albumin (BSA; Gibco; 15260-037) and re-embedded into Matrigel domes at 1:4–1:10 splitting ratio. Cultures were kept at 37 °C in 5% CO2, and OGM growth medium (StemCell, 06010) exchanged every 2–4 days.
Enteroid and colonoid suspension cultures
Medium sized enteroids/colonoids were extracted by gentle dislodgment of the Matrigel domes and incubation in Cell Recovery Solution (Corning, 354253) for minimum 1 h on ice on a rotating table. Subsequently, enteroids/colonoids were allowed to sediment by gravity and the supernatant removed. After washing with DMEM-F12/0.25% BSA, the pellet was resuspended in OGM growth medium containing 8% cold Matrigel to maintain basolateral polarity (for BO) or without Matrigel to promote eversion (for AO) and aliquoted in ultra-low attachment 24-well tissue culture plates (Corning Costar, CLS3473-24EA). BO and AO suspension cultures were incubated at 37 °C with 5% CO2 for 1 day or 3–4 days before infection, respectively.
Enteroid and colonoid bulk infections
Shortly before infection, BO or AO enteroids/colonoids were transferred into 40 µm or 25 µm mini-cell strainers (Funakoshi, HT-AMS-12502, HT-AMS-14002) and washed three times with DMEM-F12/0.25% BSA. Enteroids/colonoids were resuspended in OGM and aliquoted in ultra-low attachment 24-well tissue culture plates. The indicated Shigella strains or Tn5-sub-libraries were grown overnight in LB broth containing appropriate antibiotics at 30 °C, diluted 1:50, and subcultured for 2 h at 37 °C without antibiotics. Strains were diluted in OGM medium (unless otherwise indicated) to achieve the desired MOI. To generate barcoded consortia, the indicated tagged strains were mixed in equal ratios. Bacteria were added to each well and spun down at 300g for 10 min. At 1 h.p.i., enteroids/colonoids were transferred into 25 µm mini-strainers, washed four times with DMEM-F12/0.25% BSA and incubated with medium containing 200 µg ml–1 gentamicin (Sigma, G1914) for 2 h, unless otherwise indicated. For infections extending 3 h incubation, medium was replaced with 20 µg ml–1 gentamicin medium up to 6 h.p.i. Infected enteroids/colonoids were washed six times with DMEM-F12/0.25% BSA, recovered from the strainers and lysed in 0.1% Na-deoxycholate (unless otherwise indicated) by vigorous pipetting or by homogenization with a Tissue Lyser (Qiagen). For live imaging, BO organoids were transferred to eight-chamber slides (Cellvis, C8-1.5H-N) and when indicated 1.5 µM DRAQ7 (Invitrogen, D15106) and/or 0.2% Saponin (Calbiochem, 558255-100) was supplemented to visualize dead or permeabilized cells. For CFU counting, intracellular bacterial populations were serially diluted and plated on LB plates containing appropriate antibiotics. For the TraDIS screen infections and barcoded infection assays, intracellular bacterial populations were enriched overnight at 30 °C in 2 ml LB broth (unless otherwise indicated). A diluted culture of the inoculum was enriched in parallel and used as the input reference.
Caco-2 cell culture and infections
Caco-2 cell culture (ATCC HTB-37) and infections were performed as in Supplementary Note 2.
Barcoded assays—tag quantification
For barcoded competition assays and barcoded infections, the indicated tagged strains were mixed in equal ratios. Genomic DNA from the ‘input consortium’ and the ‘output consortium’ (following growth under the indicated conditions, or following an infection assay) was extracted using the GenElute Bacterial Genomic DNA kit (Sigma, NA2110-1KT) or the DNeasy Blood and Tissue Kit (Qiagen, 69504). For tag quantification, qPCR was performed using Maxima SYBR green/ROX qPCR master mix (2×) (ThermoFisher Scientific, K0222) on a CFX384 Touch Real-Time PCR Detection System (Bio-Rad), using 9 ng gDNA and tag-specific primers (Supplementary Table 3 and Extended Data Fig. 2i). Relative abundance of each strain was normalized to abundance in the inoculum. Standard curves were generated using gDNA from each tagged wild-type Shigella strain (Extended Data Fig. 2j). For barcoded infection assays, a colonization index was calculated as 1 – (mean relative abundanceWT/mean relative abundancemutant). To determine statistical significance, we applied a two-sided paired t-test to the relative input and output abundances of each strain normalized by the respective mean wild-type abundance across all barcoded consortium infections or competition assays.
Live-cell microscopy
BO enteroids infected with Shigella harboring the puhpT-GFP or the pmCherry plasmid, were imaged on a custom-built microscope, based on a Nikon Eclipse Ti2 body fitted with either ×40/0.6, or ×60/0.7, PlanApo air objectives (Nikon) and a Prime 95B 25 mm camera (Photometrics). The microscope chamber was maintained at 37 °C with 5% CO2. Bright-field images were acquired using differential interference contrast, and fluorescence was imaged using the 475/34, 575/35 or 648/20 excitation channels of light engine Spectra-X (Lumencor) and emission collected through quadruple bandpass filters (Chroma, 89402 and 89403). Focus was maintained over time using the Perfect Focus System (PFS). Imaging started 2–2.5 h.p.i. and images were acquired every 5 min.
TraDIS pilot screen in BO enteroids
Sub-libraries from the Shigella Tn5 library 1 were pooled to generate three replicate sub-libraries containing approximately 10,000 or 30,000 mutants each. The resulting sub-libraries were used to infect replicate wells containing ~500 BO enteroids at MOI 200 as described above. After infection, enteroids were lysed in 0.1% Na-deoxycholate by extensive pipetting. Intracellular bacterial populations were enriched overnight at 37 °C in 2 ml LB broth. A diluted culture of the inoculum was also enriched and used as the input reference; 2 ml of the bacterial culture was used to extract genomic DNA. A fraction of the pooled libraries, undergoing no further growth, was used to extract gDNA for TraDIS sequencing and used as Minimal Libraries reference; 2 µg of gDNA were used to prepare single 5′ sequencing libraries primed off the 5′ end of Tn5 for each sample. gDNA shearing, clean up, end repair, ‘A’ tailing, adapter ligation and PCR amplification were performed as detailed in ref. 87. Two pools of eight libraries were sequenced on a MiSeq instrument using v.3 150-cycle kits, run as 1 × 142 bp using a custom recipe with ten dark cycles as detailed in ref. 87.
TraDIS under different growth conditions
The 23 sub-libraries from the Shigella Tn5 library 1 were pooled to generate a dense library containing ~130.000 mutants; 200 µl of the pooled library, undergoing no further growth, were used to extract gDNA for TraDIS sequencing and used as Minimal Library reference. Then, 13 µl from the pooled library (~109 bacteria) was inoculated into 200 ml LB broth with 50 µg ml–1kanamycin and cultures were incubated at 30 °C, 180 rpm, for 16 h. The following day, the overnight culture was subcultured 1:50 in 10 ml LB broth without antibiotics (~2 × 108 bacteria) at 37 °C for 2 h. Subsequently, 50 µl aliquots of the subculture (~1.25 × 107 bacteria) were grown overnight (12 h) in LB broth without antibiotics at either 30 °C or 37 °C, to mimic the growth enrichment step that bacteria would undergo following infection experiments. Each passage was done in triplicate. At each step, 1.5 ml culture fractions were collected and used for gDNA extraction. gDNA isolation was performed using the GenElute Bacterial gDNA kit; 2 µg of gDNA was used to prepare single 5′ sequencing libraries primed off the 5′ end of Tn5 for each sample, as in ref. 87. gDNA shearing, clean up, end repair, ‘A’ tailing, adapter ligation and PCR amplification were performed as in ref. 87. A single pool of all libraries was sequenced on a NextSeq 500 instrument with a 150-cycle Mid Output kit and 25% PhiX spike-in.
RNA-seq and analysis
Shigella flexneri wild-type M90T and M90T ΔvirF strains were grown in LB broth at 30 °C or 37 °C to an OD600 of ~0.7, in triplicate. Total RNA was extracted with acid phenol as detailed in Supplementary Note 3. Total RNA was DnaseI-treated for 30 min at 37 °C and ribosomal RNA were depleted. An oligonucleotide adapter was ligated to the 3′ end of the RNA molecules. First-strand cDNA synthesis was performed using M-MLV reverse transcriptase and the 3′ adapter as primer. A 5′ Illumina TruSeq sequencing adapter was ligated to the 3′ end of the antisense cDNA. The resulting cDNA was PCR-amplified to about 10–20 ng μl–1 using a high-fidelity DNA polymerase. The cDNA was purified using the Agencourt AMPure XP kit (Beckman Coulter Genomics). For Illumina NextSeq sequencing, the samples were pooled in approximately equimolar amounts. The cDNA pool was size fractionated in the size range of 250–600 bp using a preparative agarose gel. The cDNA pool was sequenced on an Illumina NextSeq 500 system using 1 × 150 bp read length. Library preparation and sequencing was performed by vertis Biotechnologie AG. The raw sequencing reads were demultiplexed and adapter trimming was performed with Cutadapt (v.4.1). Subsequently, we ran FastQC (v.0.11.8) for quality control. The reads were aligned and quantified with STAR (v.2.6.0a), using manually curated annotations for Shigella flexneri serotype 5a M90T based on NCBI accessions CP037923.1 and CP037924.1 (ref. 88). EdgeR (v.4.0.1) was used to analyze differential gene expression for each comparison89. Only genes with at least ten reads in at least three samples were included. Data were normalized using trimmed-mean of M-values (TMM) with default settings. log2-FC was calculated with glmQLFit. Subsequently, a cutoff of ±1 was imposed on the log2-FC and an FDR of 0.01 was selected to obtain a list of genes with significant differences in gene expression.
In silico determination of bottleneck size
We estimated the size of the infection bottleneck from the pilot BO enteroid infections with sub-libraries containing 10,000 and 30,000 mutants each. The minimally enriched input was downsampled to infection bottleneck sizes between 0 and 6,000 bacteria with a stepsize of 100 (Fig. 1e), using the R base function sample. The median counts of the minimally enriched input libraries were used as weights. For every bottleneck size, we calculated the number of unique insertion sites. Comparing this number to the number of unique insertion sites in the output libraries (Fig. 1e), we estimated the size of the infection bottleneck to be about 2,000 bacteria. This results in 1,000 to 2,500 unique insertion sites per output library.
In silico determination of number of replicates
To determine the number of sequencing libraries necessary to cover the Shigella genome based on the number of mutants per input sample, we conducted an in silico analysis based on the pilot data. Using the Minimal Library reference containing 130,000 Shigella mutants (including mutants in intergenic regions), we downsampled them to libraries with 1,000, 2,000 or 3,000 mutants each. Subsequently, the number of genes featuring at least three insertion sites across all libraries was calculated (Extended Data Fig. 3a) estimating that this threshold represents the minimum required for statistical significance. The analysis revealed that for library sizes ranging from 2,000 to 3,000 mutants, approximately 3,400 to 3,700 genes achieve a coverage of three insertion sites per gene when using 40 replicates. By contrast, achieving a comparable coverage with only 1,000 mutants per input sample would require more than 100 replicates. Additionally, the number of insertion sites per gene for 40 input libraries with 3,000 mutants each was calculated (Extended Data Fig. 3b). A total of 4,032 genes (83%, mean value across 100 runs) had at least one insertion site, 1,371 genes had 20 and more insertion sites. Consequently, we opted for 43 replicates with input library sizes between 2,000 and 3,000 mutants, anticipating that this would yield technical stochastic mutant loss rates of approximately 50%, based on the number of unique insertion sites per output sample in the pilot infection assay (1,000–2,000) (Fig. 1d).
Main TraDIS screen in BO enteroid infections
The Shigella Tn5 library 2 comprising 43 sub-libraries was used to infect 43 separate wells, each containing ~800 BO enteroids at MOI 40 for 6 h as described above. After infection, enteroids were lysed in 0.1% Na-deoxycholate by extensive pipetting. Intracellular bacterial populations were enriched ON at 30 °C in 2 ml LB broth. A diluted culture of the inoculum was also enriched and used as the input reference. A 1.5 ml portion of the bacterial culture was used to extract genomic DNA using the GenElute Bacterial gDNA kit. gDNA (1.5 µg) was cleaned with Ampure XP beads to remove EDTA. Cleaned gDNA (500 ng) was used as input for the sequencing library preparations. The NEXTFLEX Rapid XP library prep kit was used for fragmentation (300–400 bp), end repair and A-tailing. An Ampure XP bead clean-up was performed followed by splinkerette ligation as previously described87. Size selection was performed with the beads included in the NEXTFLEX Rapid XP kit. PCR amplification was performed as previously described87 followed by a clean-up with the NEXTFLEX Cleanup Beads XP included in the kit. A single pool of the 43 input and the 43 output libraries was sequenced on a NextSeq 500 instrument over three runs using the 75-cycle High Output kit with a custom dark cycle recipe, similar to the recipe used on the above MiSeq runs, run as 1 × 60 bp.
TraDIS read quantification, analysis and data preparation
The raw sequencing reads were demultiplexed and filtered by transposon tag allowing one mismatch. Sequencing adapters were removed with BBDuk (BBMap v.38.94) and quality control was performed using FastQC (v.0.11.8). The reads were aligned and quantified with the Bio-TraDIS pipeline (v.1.4.5), using manually curated annotations for Shigella flexneri serotype 5a M90T (NCBI accessions CP037923.1 and CP037924.1). The read counts for the insertion sites across all samples were concatenated and genomic positions without transposon insertion sites (TIS) removed using a custom Bash script. The unique insertion sites were assigned to genetic features and intergenic regions in R. For the TraDIS under different growth conditions, EdgeR89 (v.4.0.1) was used to analyze significant changes in mutant fitness between the selected conditions. Only genes with at least ten reads in at least three samples were included. Data were normalized using TMM with default settings. log2-FC values were calculated with glmQLFit. Subsequently, a cutoff of 1 was imposed on the log2-transformed fold changes and an FDR of 0.01 was selected to obtain a list of genes with significant differences in gene expression. For the TraDIS screen in BO enteroid infections, insertion sites with fewer than 20 counts in the input sample were removed from the respective replicate. The input sample of sub-library 11 had only 300,000 aligned reads, about 5% of the average number of reads across all input samples (Fig. 2c) and was hence removed from subsequent analyses. This resulted in 85,464 nonzero TIS across 42 replicates, which were used as input to the Bayesian ZINB model.
ZINB model for TraDIS screen
We developed a Bayesian model to extract genome-wide gene-wise fitness scores in the presence of experimental bottlenecks (Fig. 2d and Extended Data Fig. 3f). For a full model description including priors and determination of statistical significance, see Supplementary Note 1. In brief, we modeled the output counts for a single insertion site \({Y}_{{\rm{rgk}},\text{out}}\) with the ZINB distribution, where the probability of a stochastic (technical) zero due to experimental bottlenecks is given by the mixing coefficient \({\theta }_{{\rm{rgk}}}\) and the expected mean \({\mu }_{{\rm{rgk}}}\) depends on the corresponding input count \({Y}_{{\rm{rgk}},\text{in}}\), a gene-wise logFC \({\log \text{FC}}_{g}\) (fitness score) and a normalization constant nr between the input and output counts of an individual sub-library
We determined the technical parameters (\({\theta }_{{\rm{rgk}}}\), nr) by fitting the ZINB model to 12,573 insertion sites located within pseudogenes, assuming that mutations in these nonfunctional genes should not affect fitness, that is, \({\log \text{FC}}_{g}=0\). Subsequently, we used these technical parameters to extract the fitness scores (\({\log \text{FC}}_{g}\)) from 85,464 insertion sites with the full ZINB model.
To derive the posterior distributions of the model parameters, we used the probabilistic programming language Stan90 (v.2.31.0). The statistical models were fitted to the TraDIS screen data running two chains of 1,000 Markov Chain Monte Carlo samples each. We assessed statistical significance and controlled the FDR by comparing z values \({z}_{g}=\frac{{\log \text{FC}}_{g}}{\varDelta {\log \text{FC}}_{g}}\) (with the posterior s.d. \(\varDelta {\log \text{FC}}_{g}\)) to the standard normal distribution42, assuming that at least 10% of genes have fitness effects during infection (Extended Data Fig. 3k,i).
Immunostaining and confocal microscopy
After infection with Shigella puhpT-GFP, BO or AO enteroids/colonoids were fixed with 2% paraformaldehyde for 30 min at room temperature and washed twice with PBS/0.25% BSA. Subsequently, enteroids/colonoids were permeabilized ON with 3% BSA, 0.1% saponin, followed by 4′,6-diamidino-2-phenylindole (DAPI; 1:1,000) (Sigma, D9542) and F-actin (1:400) (phalloidin-Alexa Fluor 647, Molecular Probes, A22287) staining for 1 h at room temperature. Samples were imaged at the BioVis platform of Uppsala University, using a LSM 700 AxioObserver (Zeiss) with a ×40/0.95 PlanApo objective, 405/488/639 diode lasers for excitation and the pinhole set to 1 a.u. for each channel.
Mass spectrometry-based proteomics
The indicated wild-type Shigella and mutant strains were grown at 37 °C until OD600 reached ~0.7. Bacterial cultures (2 ml) were spun down and washed once in PBS. Bacterial pellets were lysed in 50 µl 2% SDS and boiling at 98 °C for 10 min. Samples (5 μg) were denatured with a final concentration of 2% SDS and 20 mM Tris (2-carboxyethyl) phosphine. Samples were digested with a modified sp3 protocol91 as previously described92. In brief, samples were added to a bead suspension (10 μg of beads (Sera-Mag Speed Beads, 4515-2105-050250 and 6515-2105-050250) in 10 μl 15% formic acid and 30 μl ethanol) and incubated shaking for 15 min at room temperature. Beads were then washed four times with 70% ethanol. Proteins were digested overnight by adding 40 μl of 5 mM chloroacetamide, 1.25 mM Tris (2-carboxyethyl) phosphine and 200 ng trypsin in 100 mM HEPES pH 8.5. Peptides were eluted from the beads and dried under vacuum. Peptides were then labeled with TMTpro (ThermoFisher Scientific), pooled and desalted with solid-phase extraction using a Waters OASIS HLB μElution Plate (30 μm). Samples were fractionated onto 48 fractions on a reversed-phase C18 system running under high pH conditions, with every sixth fraction being pooled together. Samples were analyzed by liquid chromatography with tandem mass spectrometry, using a data-dependent acquisition strategy on a ThermoFisher Scientific Vanquish Neo LC coupled with a ThermoFisher Scientific Orbitrap Exploris 480. Raw files were processed with MSFragger93 (v.3.0) against the NCBI Shigella flexneri 5a strain M90T genome (CP037923 and CP037924) using standard settings for TMT. Data were normalized using vsn94 (v.3.74.0) and statistical significance was determined using limma95 (v.3.62.2).
Western blots
To evaluate IpaA-3xFT translation, equal amounts of total protein was extracted from the indicated strains following growth at 37 °C until OD600 ~0.3 and induction by 0.25 mM IPTG for 30 min. Proteins were separated using Any kD Mini-PROTEAN TGX Stain-Free Protein Gels (Bio-Rad, 4568126) and transferred onto Trans-Blot Turbo Mini 0.2 µm PVDF Transfer Packs (Bio-Rad, 1704156). Loading controls were obtained by the stain-free method, imaging each gel upon exposure to UV-light for 5 min96. Immunodetection was performed using an anti-Flag antibody (1:10,000) (Sigma, F1804), as described previously97. PageRuler Plus Prestained Protein Ladder (ThermoFisher Scientific, 26619) was used as weight marker. To quantify western blots, protein extracts were serially diluted and relative protein amounts calculated from a standard curve.
Quantitative PCR with reverse transcription
Total RNA purification and cDNA synthesis were performed as previously described98. RT–qPCR was performed using Maxima SYBR green/ROX qPCR master mix (2×) (ThermoFisher Scientific, K0222) on a CFX384 Touch Real-Time PCR Detection System (Bio-Rad). The levels of ipaA, mnmE and mnmG transcripts were analyzed using the 2−ΔΔCt (cycle threshold) method and results are reported as the fold increase relative to the reference. The housekeeping gene nusA was used for normalization. Oligonucleotide primers used are listed in Supplementary Table 3.
Codon usage analysis
Codon usage was analyzed using a Python (v.3.9.16) script and the BioPython (v.1.81) library. In brief, the occurrences of each codon for each nonpseudogene coding sequence was counted on the chromosome and virulence plasmid (accessions CP037923.1 and CP037924). The relative frequencies for each synonymous codon were calculated globally and for each nonpseudogene coding sequence.
Statistics and reproducibility
Sample sizes were not predetermined, except for the Main TraDIS screen in BO enteroid infections (43 biological replicate sub-libraries; see above). For the main TraDIS screen in BO enteroid infections, sub-library 11 was excluded due to low sequencing read counts. TraDIS pilot experiments in BO enteroids were done in biological triplicate. TraDIS under different growth conditions and RNA-seq experiments were performed in biological triplicate. Proteomic profiling was done using four biological replicates per strain. Barcoded assays were performed at least in duplicate (in this setup each replicate used two biological replicates for each genotype, allowing for a powerful and internally controlled comparison). All other experiments were performed with at least three biological independent replicates. Exact sample sizes indicated in each figure legend. All experiments were highly reproducible and our sample sizes were similar to previous related publications. The experiments were not randomized. Data collection and analysis was not blinded. Statistical comparisons were done with two-sided t-tests, or two-sided Mann–Whitney U-test, as described in each figure legend.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Transcriptomic data and TraDIS data have been deposited in the Gene Expression Omnibus (GEO) database, with the SuperSeries no. GSE267520. The mass spectrometry proteomics data have been deposited at the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD046629. Source data are provided with this paper.
Code availability
For TraDIS under different growth conditions we used the Bio-TraDIS pipeline87 (v.1.4.5) to align and quantify reads and EdgeR89 (v.4.0.1) to analyze significant changes in fitness. For differential gene expression we used STAR (v.2.6.0a) and EdgeR (v.4.0.1). For mass spectrometry-based proteomics MSFragger was used93, data were normalized using vsn94 and statistical significance was determined using limma95. For the main TraDIS screen in BO enteroid infections we developed a custom Bayesian ZINB model (Supplementary Note 1; Methods). For codon usage analysis we used a custom Python (v.3.9.16) script and the BioPython (v.1.81) library. All custom code was deposited to Zenodo at the following links: https://doi.org/10.5281/zenodo.15096674 (ref. 99) and https://zenodo.org/records/15100662 (ref. 100).
References
Schroeder, G. N. & Hilbi, H. Molecular pathogenesis of Shigella spp.: Controlling host cell signaling, invasion, and death by type III secretion. Clin. Microbiol. Rev. 21, 134–156 (2008).
Kotloff, K. L., Riddle, M. S., Platts-Mills, J. A., Pavlinac, P. & Zaidi, A. K. M. Shigellosis. Lancet 391, 801–812 (2018).
Alphonse, N. & Odendall, C. Animal models of shigellosis: a historical overview. Curr. Opin. Immunol. 85, 102399 (2023).
Marteyn, B., Gazi, A. & Sansonetti, P. Shigella: a model of virulence regulation in vivo. Gut Microbes 3, 104–120 (2012).
Bajunaid, W. et al. The T3SS of Shigella: expression, structure, function, and role in vacuole escape. Microorganisms 8, 1933 (2020).
Mellouk, N. & Enninga, J. Cytosolic access of intracellular bacterial pathogens: the shigella paradigm. Front. Cell. Infect. Microbiol. 6, 35 (2016).
Sansonetti, P. J., Kopecko, D. J. & Formal, S. B. Involvement of a plasmid in the invasive ability of Shigella flexneri. Infect. Immun. 35, 852–860 (1982).
Le Gall, T. et al. Analysis of virulence plasmid gene expression defines three classes of effectors in the type III secretion system of Shigella flexneri. Microbiology 151, 951–962 (2005).
Mattock, E. & Blocker, A. J. How do the virulence factors of shigella work together to cause disease? Front. Cell. Infect. Microbiol. 7, 64 (2017).
Schnupf, P. & Sansonetti, P. J. Shigella pathogenesis: new insights through advanced methodologies. Microbiol. Spectr. 7, https://doi.org/10.1128/microbiolspec.bai-0023-2019 (2019).
Landry, J. J. M. et al. The genomic and transcriptomic landscape of a HeLa cell line. G3 (Bethesda) 3, 1213–1224 (2013).
Sato, T. & Clevers, H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science 340, 1190–1194 (2013).
Geiser, P. et al. Salmonella enterica serovar typhimurium exploits cycling through epithelial cells to colonize human and murine enteroids. MBio 12, e02684–20 (2021).
Koestler, B. J. et al. Human intestinal enteroids as a model system of Shigella pathogenesis. Infect. Immun. 87, e00733–18 (2019).
Ranganathan, S. et al. Evaluating Shigella flexneri pathogenesis in the human enteroid model. Infect. Immun. 87, e00740–18 (2019).
Co, J. Y. et al. Controlling epithelial polarity: a human enteroid model for host-pathogen interactions. Cell Rep. 26, 2509–2520.e4 (2019).
Däullary, T. et al. A primary cell-based in vitro model of the human small intestine reveals host olfactomedin 4 induction in response to Salmonella Typhimurium infection. Gut Microbes 15, 2186109 (2023).
Nickerson, K. P. et al. A versatile human intestinal organoid-derived epithelial monolayer model for the study of enteric pathogens. Microbiol. Spectr. 9, e0000321 (2021).
Heo, I. et al. Modelling Cryptosporidium infection in human small intestinal and lung organoids. Nat. Microbiol. 3, 814–823 (2018).
Noel, G. et al. A primary human macrophage-enteroid co-culture model to investigate mucosal gut physiology and host-pathogen interactions. Sci. Rep. 7, 45270 (2017).
Mounier, J., Vasselon, T., Hellio, R., Lesourd, M. & Sansonetti, P. J. Shigella flexneri enters human colonic Caco-2 epithelial cells through the basolateral pole. Infect. Immun. 237, 248 (1992).
Co, J. Y., Margalef-Català, M., Monack, D. M. & Amieva, M. R. Controlling the polarity of human gastrointestinal organoids to investigate epithelial biology and infectious diseases. Nat. Protoc. 16, 5171–5192 (2021).
Cain, A. K. et al. A decade of advances in transposon-insertion sequencing. Nat. Rev. Genet. 21, 526–540 (2020).
Abel, S., Abel zur Wiesch, P., Davis, B. M. & Waldor, M. K. Analysis of bottlenecks in experimental models of infection. PLoS Pathog. 11, e1004823 (2015).
Jenniches, L. et al. Improved RNA stability estimation through Bayesian modeling reveals most Salmonella transcripts have subminute half-lives. Proc. Natl Acad. Sci. USA. 121, e2308814121 (2024).
Di Martino, M. L., Ek, V., Hardt, W.-D., Eriksson, J. & Sellin, M. E. Barcoded consortium infections resolve cell type-dependent Salmonella enterica serovar typhimurium entry mechanisms. MBio 10, e00603–e00619 (2019).
Di Martino, M. L. & Sellin, M. E. Barcoded consortium infections: a scalable, internally controlled method to study host cell binding and invasion by pathogenic bacteria. Methods Mol. Biol. 2674, 295–311 (2023).
Hausmann, A. et al. Intestinal epithelial NAIP/NLRC4 restricts systemic dissemination of the adapted pathogen Salmonella Typhimurium due to site-specific bacterial PAMP expression. Mucosal Immunol. 13, 530–544 (2020).
Drecktrah, D. et al. Dynamic behavior of Salmonella-induced membrane tubules in epithelial cells. Traffic 9, 2117–2129 (2008).
Chao, M. C., Abel, S., Davis, B. M. & Waldor, M. K. The design and analysis of transposon insertion sequencing experiments. Nat. Rev. Microbiol. 14, 119–128 (2016).
Abel, S. et al. Sequence tag–based analysis of microbial population dynamics. Nat. Methods 12, 223–226 (2015).
Di Martino, M. L., Falconi, M., Micheli, G., Colonna, B. & Prosseda, G. The multifaceted activity of the VirF regulatory protein in the Shigella lifestyle. Front. Mol. Biosci. 3, 61 (2016).
Schuch, R. & Maurelli, A. T. Virulence plasmid instability in Shigella flexneri 2a is induced by virulence gene expression. Infect. Immun. 65, 3686–3692 (1997).
Sturm, A. et al. The cost of virulence: retarded growth of Salmonella Typhimurium cells expressing type III secretion system 1. PLoS Pathog. 7, e1002143 (2011).
Vasanthakrishnan, R. B. et al. PrfA regulation offsets the cost of Listeria virulence outside the host. Environ. Microbiol. 17, 4566–4579 (2015).
Diard, M. & Hardt, W. D. Evolution of bacterial virulence. FEMS Microbiol. Rev. 41, 679–697 (2017).
Falconi, M., Prosseda, G., Giangrossi, M., Beghetto, E. & Colonna, B. Involvement of FIS in the H-NS-mediated regulation of virF gene of Shigella and enteroinvasive Escherichia coli. Mol. Microbiol. 42, 439–452 (2001).
Nakayama, S. & Watanabe, H. Involvement of cpxA, a sensor of a two-component regulatory system, in the pH-dependent regulation of expression of Shigella sonnei virF gene. J. Bacteriol. 177, 5062–5069 (1995).
Nakayama, S. I., & Watanabe, H. Identification of cpxR as a positive regulator essential for expression of the Shigella sonnei virF gene. J. Bacteriol. 180, 3522–3528 (1998).
Grant, A. J. et al. Modelling within-host spatiotemporal dynamics of invasive bacterial disease. PLoS Biol. 6, e74 (2008).
Freed, N. E., Bumann, D. & Silander, O. K. Combining Shigella Tn-seq data with gold-standard E. coli gene deletion data suggests rare transitions between essential and non-essential gene functionality. BMC Microbiol. 16, 203 (2016).
Efron, B. Microarrays, empirical Bayes and the two-groups model. Statist. Sci. 23, 1–22 (2008).
Coluccia, M. et al. Role of the MDR efflux pump AcrAB in epithelial cell invasion by Shigella flexneri. Biomolecules 13, 823 (2023).
West, N. P. et al. Optimization of virulence functions through glucosylation of Shigella LPS. Science 307, 1313–1317 (2005).
Caboni, M. et al. An O antigen capsule modulates bacterial pathogenesis in Shigella sonnei. PLoS Pathog. 11, e1004749 (2015).
Watson, J. L. et al. Shigella sonnei O-antigen inhibits internalization, vacuole escape, and inflammasome activation. MBio 10, e02654–19 (2019).
Hong, M. & Payne, S. M. Effect of mutations in Shigella flexneri chromosomal and plasmid-encoded lipopolysaccharide genes on invasion and serum resistance. Mol. Microbiol. 24, 779–791 (1997).
Ohya, K., Handa, Y., Ogawa, M., Suzuki, M. & Sasakawa, C. IpgB1 is a novel Shigella effector protein involved in bacterial invasion of host cells. Its activity to promote membrane ruffling via Rac1 and Cdc42 activation. J. Biol. Chem. 280, 24022–24034 (2005).
Hachani, A. et al. IpgB1 and IpgB2, two homologous effectors secreted via the Mxi-Spa type III secretion apparatus, cooperate to mediate polarized cell invasion and inflammatory potential of Shigella flexenri. Microbes Infect. 10, 260–268 (2008).
Weddle, E. A., Köseoğlu, V. K., DeVasure, B. A. & Agaisse, H. F. The type three secretion system effector protein IpgB1 promotes Shigella flexneri cell-to-cell spread through double-membrane vacuole escape. PLoS Pathog. 18, e1010380 (2022).
Bourdet-Sicard, R. et al. Binding of the Shigella protein IpaA to vinculin induces F-actin depolymerization. EMBO J. 18, 5853–5862 (1999).
Tran Van Nhieu, G., Ben-Ze’ev, A. & Sansonetti, P. J. Modulation of bacterial entry into epithelial cells by association between vinculin and the Shigella IpaA invasin. EMBO J. 16, 2717–2729 (1997).
Uchiya, K. I. et al. Identification of a novel virulence gene, virA, on the large plasmid of Shigella, involved in invasion and intercellular spreading. Mol. Microbiol. 17, 241–250 (1995).
Dong, N. et al. Structurally distinct bacterial TBC-like GAPs link Arf GTPase to Rab1 inactivation to counteract host defenses. Cell 150, 1029–1041 (2012).
Armengod, M. E. et al. Enzymology of tRNA modification in the bacterial MnmEG pathway. Biochimie 94, 1510–1520 (2012).
Moukadiri, I., Garzón, M. J., Björk, G. R. & Armengod, M. E. The output of the tRNA modification pathways controlled by the Escherichia coli MnmEG and MnmC enzymes depends on the growth conditions and the tRNA species. Nucleic Acids Res 42, 2602–2623 (2014).
Armengod, M. E. et al. Modification of the wobble uridine in bacterial and mitochondrial tRNAs reading NNA/NNG triplets of 2-codon boxes. RNA Biol 11, 1495–1507 (2014).
Chen, G. F. T. & Inouye, M. Role of the AGA/AGG codons, the rarest codons in global gene expression in Escherichia coli. Genes Dev 8, 2641–2652 (1994).
Mitchell, P. S. et al. NAIP-NLRC4-deficient mice are susceptible to shigellosis. eLife 9, 1e59022 (2020).
Miles, S. L., Holt, K. E. & Mostowy, S. Recent advances in modelling Shigella infection. Trends Microbiol. 32, 917–924 (2024).
Perlman, M., Senger, S., Verma, S., Carey, J. & Faherty, C. S. A foundational approach to culture and analyze malnourished organoids. Gut Microbes 15, 2248713 (2023).
Ventayol, P. S. et al. Bacterial detection by NAIP/NLRC4 elicits prompt contractions of intestinal epithelial cell layers. Proc. Natl Acad. Sci. USA 118, e2013963118 (2021).
van Rijn, J. M. et al. High-definition DIC imaging uncovers transient stages of pathogen infection cycles on the surface of human adult stem cell-derived intestinal epithelium. MBio 13, e0002222 (2021).
Kuehl, C. J., D’gama, J. D., Warr, A. R. & Waldor, M. K. An oral inoculation infant rabbit model for Shigella infection. MBio 11, e03105–e03119 (2020).
Warr, A. R. et al. Transposon-insertion sequencing screens unveil requirements for EHEC growth and intestinal colonization. PLoS Pathog. 15, e1007652 (2019).
Ghany et al. Functional analysis of colonization factor antigen I positive enterotoxigenic Escherichia coli identifies genes implicated in survival in water and host colonization. Microb. Genomics 7, 000554 (2021).
Paczosa, M. K. et al. Transposon mutagenesis screen of Klebsiella pneumoniae identifies multiple genes important for resisting antimicrobial activities of neutrophils in mice. Infect. Immun. 88, e00034–20 (2020).
Armbruster, C. E. et al. Twin arginine translocation, ammonia incorporation, and polyamine biosynthesis are crucial for Proteus mirabilis fitness during bloodstream infection. PLoS Pathog. 15, e1007653 (2019).
McCarthy, A. J., Stabler, R. A. & Taylor, P. W. Genome-wide identification by transposon insertion sequencing of Escherichia coli K1 genes essential for in vitro growth, gastrointestinal colonizing capacity, and survival in serum. J. Bacteriol. 200, e00698–17 (2018).
Risso, D., Perraudeau, F., Gribkova, S., Dudoit, S. & Vert, J. P. A general and flexible method for signal extraction from single-cell RNA-seq data. Nat. Commun. 9, 284 (2018).
Choi, K., Chen, Y., Skelly, D. A. & Churchill, G. A. Bayesian model selection reveals biological origins of zero inflation in single-cell transcriptomics. Genome Biol 21, 1–16 (2020).
Subramaniyam, S. et al. Statistical analysis of variability in TnSeq data across conditions using zero-inflated negative binomial regression. BMC Bioinformatics 20, 603 (2019).
Shippy, D. C., Eakley, N. M., Lauhon, C. T., Bochsler, P. N. & Fadl, A. A. Virulence characteristics of Salmonella following deletion of genes encoding the tRNA modification enzymes GidA and MnmE. Microb. Pathog. 57, 1–9 (2013).
Krueger, J. et al. tRNA epitranscriptome determines pathogenicity of the opportunistic pathogen Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 121, e2312874121 (2024).
Tomasi, F. G., Kimura, S., Rubin, E. J. & Waldor, M. K. A tRNA modification in Mycobacterium tuberculosis facilitates optimal intracellular growth. eLife 12, RP87146 (2023).
Fleming, B. A. et al. A tRNA modifying enzyme as a tunable regulatory nexus for bacterial stress responses and virulence. Nucleic Acids Res 50, 7570–7590 (2022).
Durand, J. M. B., Dagberg, B., Uhlin, B. E. & Björk, G. R. Transfer RNA modification, temperature and DNA superhelicity have a common target in the regulatory network of the virulence of Shigella flexneri: the expression of the virF gene. Mol. Microbiol. 35, 924–935 (2000).
Durand, J. M. B., Björk, G. R., Kuwae, A., Yoshikawa, M. & Sasakawa, C. The modified nucleoside 2-methylthio-N6-isopentenyladenosine in tRNA of Shigella flexneri is required for expression of virulence genes. J. Bacteriol. 179, 5777–5782 (1997).
Urbonavičius, J., Durand, J. M. B. & Björk, G. R. Three modifications in the D and T arms of tRNA influence translation in Escherichia coli and expression of virulence genes in Shigella flexneri. J. Bacteriol. 184, 5348–5357 (2002).
de Crécy-Lagard, V. & Jaroch, M. Functions of bacterial tRNA modifications: from ubiquity to diversity. Trends Microbiol. 29, 41–53 (2021).
Prosseda, G. et al. The virF promoter in Shigella: more than just a curved DNA stretch. Mol. Microbiol. 51, 523–537 (2004).
Lucchini, S. et al. H-NS mediates the silencing of laterally acquired genes in bacteria. PLoS Pathog. 2, e81 (2006).
Ali, S. S. et al. Silencing by H-NS potentiated the evolution of Salmonella. PLoS Pathog. 10, e1004500 (2014).
Wang, H. et al. Increased plasmid copy number is essential for Yersinia T3SS function and virulence. Science 353, 492–495 (2016).
Schneiders, S. et al. Spatiotemporal variations in growth rate and virulence plasmid copy number during yersinia pseudotuberculosis infection. Infect. Immun. 89, e00710–e00720 (2021).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Barquist, L. et al. The TraDIS toolkit: sequencing and analysis for dense transposon mutant libraries. Bioinformatics 32, 1109–1111 (2016).
Cervantes-Rivera, R., Tronnet, S. & Puhar, A. Complete genome sequence and annotation of the laboratory reference strain Shigella flexneri serotype 5a M90T and genome-wide transcriptional start site determination. BMC Genomics 21, 285 (2020).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Carpenter, B. et al. Stan: a probabilistic programming language. J. Stat. Softw. 76, 1 (2017).
Hughes, C. S. et al. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 10, 757 (2014).
Mateus, A. et al. The functional proteome landscape of Escherichia coli. Nature 588, 473–478 (2020).
Kong, A. T., Leprevost, F. V., Avtonomov, D. M., Mellacheruvu, D. & Nesvizhskii, A. I. MSFragger: ultrafast and comprehensive peptide identification in mass spectrometry-based proteomics. Nat. Methods 14, 513–520 (2017).
Huber, W., Von Heydebreck, A., Sültmann, H., Poustka, A. & Vingron, M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18, S96–S104 (2002).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Colella, A. D. et al. Comparison of stain-free gels with traditional immunoblot loading control methodology. Anal. Biochem. 430, 108–110 (2012).
Skovajsová, E., Colonna, B., Prosseda, G., Sellin, M. E. & Di Martino, M. L. The VirF21:VirF30 protein ratio is affected by temperature and impacts Shigella flexneri host cell invasion. FEMS Microbiol. Lett. 369, fnac043 (2022).
Di Martino, M. L., Romilly, C., Wagner, E. G. H., Colonna, B. & Prosseda, G. One gene and two proteins: a leaderless mRNA supports the translation of a shorter form of the Shigella VirF regulator. MBio 7, e01860–16 (2016).
Jenniches, L. ZINB model for TraDIS infection screens. Zenodo https://doi.org/10.5281/zenodo.15096674 (2025).
Eriksson, J. Codon occurrence counter for TraDIS project. Zenodo https://doi.org/10.5281/zenodo.15100662 (2025).
Acknowledgements
We are grateful to members of the Sellin laboratory and to J. Näsvall for helpful discussions, and to the staff at our surgical units for assistance with human intestinal tissue sampling for enteroid and colonoid establishment. We acknowledge the Ramaciotti Center for Genomics, Sydney, Australia for TraDIS library preparation and sequencing. This work was supported by grants from ESCMID (Research Grant 2020 to M.L.D.M.), the Carl Trygger Foundation (CTS 22:1915 to M.L.D.M.), the Clas Groschinsky Memorial Foundation (M21112 to M.L.D.M.), the Bavarian State Ministry for Science and the Arts through the research network bayresq.net (to L.B.), the Swedish Research Council (2018-02223, 2022-01590 to M.E.S.), the Swedish Foundation for Strategic Research (FFL18-0165 to M.E.S.) and the SciLifeLab Fellows program (to M.E.S.). The proteomics analysis was enabled by a grant from Kempestiftelserna (JCK3126 to A.M.). M.P. acknowledges support by a 6-month international visiting scholarship from Sapienza University of Rome.
Funding
Open access funding provided by Uppsala University.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.L.D.M., M.E.S., L.J. and L.B. Validation: M.L.D.M. and M.E.S. Investigation: M.L.D.M., A.B., J.E., A.N., A.C.C.L., M.P. and A.M. Methodology: M.L.D.M., L.J., A.B., M.P., A.M., L.B. and M.E.S. Formal analysis: M.L.D.M., L.J., A.B., J.E. and A.M. Resources: M.L.D.M., M. Sundbom, M. Skogar, W.G., D.-W.L., P.M.H., A.M., LB. and M.E.S. Software: L.J., J.E. and L.B. Data curation: M.L.D.M. Visualization: M.L.D.M., L.J. Project administration: M.L.D.M., M. Sundbom, M. Skogar, W.G., D.-L.W., P.M.H. and M.E.S. Supervision: M.L.D.M., L.B. and M.E.S. Funding acquisition: M.L.D.M., L.B. and M.E.S. Writing—original draft: M.L.D.M., L.J., L.B. and M.E.S. Writing–review and editing: all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Jens Puschhof and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characteristics of the basal-out enteroid infection model for early Shigella colonization of human intestinal epithelium.
(Extended Data Fig) Representative image showing a BO enteroid suspension culture as it appears before infection. (b) Shigella CFU counts upon coinfection of BO enteroids with a mix of wt and ΔmxiD (non-invasive) strains for 2 h and 6 h at MOI 40. CFU counts come from 5 biological replicates pooled from 2 independent experiments. Data shown as Mean +/- SD. Significance determined by two-sided Mann-Whitney U-test; **p < 0.01. (c) Shigella CFU counts upon coinfection of Caco-2 cells with a mix of wt and ΔmxiD (non-invasive) strains for 2 h and 6 h at MOI 40. CFU counts come from 7 biological replicates pooled from 2 independent experiments. Data shown as Mean +/- SD. Significance determined by two-sided Mann-Whitney U-test; **p < 0.01; ***p < 0.001. (d) Graph showing the number of generations (left panel) and the generation time (right panel) for Shigella intracellular populations in BO enteroids and Caco-2 cells between 2 and 6 h pi. Number of generations was calculated using CFUs counts from Extended Data Fig. 1b-c. Data shown as Mean +/- SD. Significance determined by two-sided Mann-Whitney U-test; **p < 0.01. (e) Representative time-lapse series of BO enteroids stained with DRAQ7 and treated as following: uninfected (first row); uninfected with addition of saponin after 4 h (second row) and, infected with Shigella wt containing the cytosolic reporter puhpT-GFP at MOI 40 (third row). Experiments were repeated 4 times with imaging of at least 10 enteroids/experiment. Roman numbers represent five different examples of infection foci. Scale bar: 50 µm.
Extended Data Fig. 2 Optimization of experimental design for combining TraDIS with Shigella enteroid infections.
(a) Shigella M90T chromosome and pINV maps showing the TIS distribution across the Tn5 Library #1 (grey). Origin of replication (ori) for chromosome and pINV and entry region (ER) for pINV in magenta. (b–d) MA plots of relative mutant fitness expressed as Log2FC between Enr 30 °C vs Sub (b), Enr 37 °C vs Sub (c) and Enr 37 °C vs Enr 30 °C (d). Each dot represents a gene. Detrimental genes located on pINV shown in dark orange, detrimental genes on the chromosome in light orange, advantageous genes on the pINV in dark blue, and advantageous genes on the chromosome in light blue. Significance determined by two-sided F-test with Benjamini-Hochberg correction for multiple hypothesis testing. Log2FC ≥ 1; FDR ≤ 0.01. (e–g) Volcano plots showing differentially expressed genes in Shigella wt grown at 37 °C vs 30 °C (e), Shigella wt vs ΔvirF mutant grown at 37 °C (f) and Shigella wt vs ΔvirF mutant grown at 30 °C (g). Each dot represents a gene. Differentially expressed genes located on the pINV shown in pink, on the chromosome in black, and all non-significant differentially expressed genes in grey. Significance determined by two-sided F-test with Benjamini-Hochberg correction for multiple hypothesis testing. Log2FC ≥ 1; FDR ≤ 0.01. (h) Plot showing relative mutant fitness for both pINV- and chromosome-located genes on the y-axis (Log2FC as noted in Extended Data Fig. 2d – Enr 37 °C vs Enr 30 °C) versus the respective expression changes on the x-axis (RNA-Seq – Log2FC as noted in Extended Data Fig. 2e for Shigella wt grown at 37 °C vs 30 °C). Significant pINV-encoded advantageous (dark blue) or detrimental (dark orange) and chromosomal advantageous (light blue) or detrimental (light orange) genes are shown. Log2FC ≥ 1; FDR ≤ 0.01. (i) Schematic representation of the tag-containing Shigella chromosomal locus. Primer binding sites for qPCR tag quantification indicated. (j) Standard curves for qPCR detection of tags A-G, using specific primer pairs (see Supplementary Table 3). gDNA from pure monocultures of each of seven Shigella wt tagA-G barcoded strains were diluted (ten-fold dilutions up to 10−6) and used to generate standard curves for each primer pair. R2, slope (Y) and efficiency (E) values indicated in each panel.
Extended Data Fig. 3 Shigella mutant library characteristics and modelling p rameters.
(a) In silico estimation of the number of genes with at least 3 TIS for input sub-libraries with 1,000-3,000 mutants. (b) Pie chart showing expected gene-wise TIS number across input sub-libraries of 40 replicates with 3,000 mutants each, based on the simulation in Extended Data Fig. 3a. 17% of genes had no TIS, ~1,371 ≥ 20 TIS. (c) The Shigella Tn5 mutant library #2 for enteroid infections pooled into 43 sub-libraries. Shown is the number of mutants/sub-library determined by CFU plating. (d) Total Shigella intracellular population sizes upon enteroid infection with sub-libraries in c, determined by CFU plating. No data exist for sub-libraries 4-9. (e) Pie chart showing the gene-wise TIS number across samples of the 42 Shigella Tn5 mutant input sub-libraries. Sub-library 11 removed due to low input sequencing read counts (see Fig. 2c). 14% of genes had no TIS, ~1,219 genes ≥20 TIS. (f) Plate diagram depicting the ZINB model to map Shigella invasion factors (see also Fig. 2d). Technical parameters (normalization nr, dependence of mutant loss on abundance αrgk,in and fraction of zeros in the output sample fz,r) estimated from pseudogenes in grey, gene-fitness scores (logFCg) and disperson (Φg) estimated from all genes in red. (g) PCA plot of TraDIS data from the Shigella enteroid infections. Input-output sub-libraries cluster by replicate. (h) Histogram showing that distribution of read counts of input TIS can be approximated by a negative binomial distribution (red line). Infection bottleneck leads to zero-inflated count distribution in output samples. (i) Fraction of TIS with zero counts in output (and non-zero counts in the respective input) varies between ~25 and ~75% across sub-libraries. (j) Fraction of zero counts in the output depends on the input TIS abundance. Y-axis shows fraction of zero counts, x-axis TIS input abundance normalized by average TIS input abundance. (k) Histogram showing the distribution of z-values approximated by a normal distribution around zero with a standard deviation of 0.9 (see Methods, Supplementary Note 1). (l) The FDR corresponds to the cumulative fraction of z-values above and below the normal distribution (see Methods). Dashed line indicates an FDR cut-off of 0.01.
Extended Data Fig. 4 Contribution of the AcrABTolC efflux pump system to Shigella colonization of enteroids, and resilience to environmental conditions relevant to the infection assay.
(a) BO enteroids were infected with barcoded consortia comprising two Shigella wt (tagA, tagB), two ΔmxiD (tagC, tagD) and two mutant (tagE, tagF) strains at MOI 40 for 6 h in Organoid growth medium (OGM). Mut refers to mutant specified on x-axis. Left panel, percentage of each strain in the input population. Middle panel, relative tag abundance in the intracellular population, normalized against the corresponding input (see Left panel). Right panel, Colonization Index for the indicated mutants (derived from data in the middle panel and calculated as 1-(wt/mut)). Data in right panel shown as Mean +/- SD. Data for two independently generated consortia per infection. Grey shading indicates detection limit. Note that relative abundances of ΔacrA, ΔacrB, ΔacrAB, and ΔtolC mutants fall under the detection limit and are ~1000-fold lower than the non-invasive ΔmxiD strain. (b–f) Barcoded competition assay with a consortium comprising two wt (tagA, tagB), two ΔmxiD (tagC, tagD) and two ΔacrAB or ΔtolC mutant (tagF, tagG) strains grown in different media (LB broth, Organoid growth medium (OGM) or DMEM-F12/0.25%BSA) (b), or LB broth with increasing concentration of Na-deoxycholate (c), Bile salts (d), gentamicin (e), or LL-37 (f). Shown is the relative tag abundance under each condition. Relative tag abundances normalized to the consortia grown in LB. (g) BO enteroids infected with barcoded consortia as in a, but in DMEM-F12/0.25%BSA medium. Data for three independently generated consortia per infection. Significance determined by two-sided paired t-test between normalized output and input abundances (see Methods). **p < 0.01. Data in right panel shown as Mean +/- SD. Interpretation: ΔacrA, ΔacrB, ΔacrAB and ΔtolC mutants were not only sensitive to bile salts/deoxycholate, but also to the OGM medium used for infections (see b-d). Therefore, the low Log2FC value observed for these mutants in the TraDIS screen does not exclusively reflect decreased enteroid colonization, but also generalized sensitivity to multiple infection condition stressors (a–f). Nevertheless, an epithelial colonization defect was validated in the absence of OGM and Na-deoxycholate. Here, the Colonization Indexes for ΔacrA, ΔacrB, ΔacrAB mutants were ~10-fold lower and for ΔtolC mutant ~20-30-fold lower than the wt (g; compare to a).
Extended Data Fig. 5 Input compositions and phenotypic validations for barcoded consortium infections with strains carrying mutations in Shigella T3SS effecto s.
(a, b) BO and AO enteroids (a) and BO and AO colonoids (b) were infected with a mixed barcoded consortium comprising two wt (tagA, tagB), two ΔmxiD (tagC, tagD) and two mutant (tagE, tagF) strains at MOI 40 for 6 h. Mut refers to the specific mutant as indicated on x-axis. Graphs depict the composition of barcoded Shigella consortia used as input inoculums for experiments in Fig. 3j–m. The relative abundance of each tag in the input consortia is plotted as percentage of the total population. Note that no strain is consistently over- or underrepresented. Shown are data for two independently generated consortia for each infection. See Fig. 3j–m for the corresponding relative tag abundances in the intracellular population and Colonization Index scores. (c–f) BO and AO enteroids (ID 22-2jej) (c) and BO and AO colonoids (ID 22-4col) (e) were infected with a mixed barcoded consortium comprising two wt (tagA, tagB), two ΔmxiD (tagC, tagD) and two ΔipaA (tagE, tagF) strains at MOI 40 for 6 h. Shown is the quantification of relative tag abundance, normalized against the corresponding input. For each geometry, infections were executed with four independently generated consortia. Significance determined by two-sided paired t-test for each specific mutant between the normalized output and the normalized input abundances (see Methods). ***p < 0.001. (d–f) Bar graph showing the Colonization Index for each mutant in BO (dark grey) and AO (light grey) enteroid 22-2jej infections (derived from data in c and calculated as 1-(wt/mut); sample size as in c) and BO (dark grey) and AO (light grey) colonoid 22-4col infections (derived from data in e and calculated as 1-(wt/mut); sample size as in e). wt is shown as 0. Data shown as Mean +/- SD.
Extended Data Fig. 6 Enteroid colonization phenotypes for chromosomal Shigella genes, and analysis of codon usage b.
(a–c) BO enteroids infected with barcoded consortia comprising two wt (tagA, tagB), two ΔmxiD (tagC, tagD) and two mutant (tagE, tagF) strains at MOI 40 for 6 h. Mut refers to the mutant specified on x-axis. (a) Percentage of each strain in the input and (b) relative tag abundance in the intracellular population, normalized against the corresponding input. Data for two independent consortia per infection. Significance determined by two-sided paired t-test (see Methods). ns - non-significant; *p < 0.05. (c) Colonization Indexes for the indicated mutants (Derived from b and calculated as 1-(wt/mut)). Data shown as Mean +/- SD. (d) Percentage of each strain in the input for barcoded consortia used for infections in Fig. 4b. (e) Intracellular bacteria in BO enteroids infected with Shigella wt, ΔmxiD, ΔmnmE or ΔmnmG for 3 h at MOI 40. CFU counts from 6 (wt), 5 (ΔmxiD) and 7 (ΔmnmE/ ΔmnmG) biological replicates pooled from 2 independent experiments, normalized to wt. Data shown as Mean +/− SD. Significance determined by two-tailed Mann-Whitney U-test; ns - non-significant; *p < 0.05; **p < 0.01. (f) Venn-diagram of Shigella differentially expressed proteins in ΔmnmE vs wt (MnmE regulated proteins), and ΔmnmG vs wt (MnmG regulated proteins); Log2FC ≥ 0.5, adj_pvalue ≤ 0.01. (g) Linear regression between pINV differentially expressed proteins in ΔmnmE vs wt (MnmE regulated proteins), and ΔmnmG vs wt (MnmG regulated proteins); Log2FC ≥ 0.5, adj_pvalue ≤ 0.01. (h) Codon usage frequency in pINV-located Shigella genes compared to chromosomal genes. Dashed line indicates no enrichment. (i, j) Relative protein levels (Log2FC as in Fig. 4d – ΔmnmG vs wt) versus AGA codon usage ratios for Arg (i) and GGA for Gly (j) per protein. (k–p) Relative protein levels (Log2FC as in Fig. 4c – ΔmnmE vs wt) versus UUA codon usage ratios for Leu (k), CAA for Gln (l), GAA for Glu (m), AAA for Lys (n), AGG for Arg (o) and GGG for Gly (p) per protein. For i-p, differentially expressed proteins on pINV shown in pink, on chromosome in black, and all non-significant differentially expressed proteins in grey. Log2FC ≥ 0.5; adj_pvalue ≤ 0.01. Dark and light green dashed lines specify mean codon usage for all open reading frames on pINV, or chromosome, respectively.
Extended Data Fig. 7 Effects of mnmE and mnmG complementation on IpaA-3xFT protein translation and Shigella fitness during inoculum growth.
(a) Relative IpaA-3xFT protein content in Shigella wt and ΔmnmE strains carrying an inducible pTac-ipaA-3xFT plasmid and the p-Empty or p-mnmE plasmids, determined by quantification of western blots of serially diluted samples. Shown are 3 independent western blots of serially diluted samples for each strain, used for generating standard curves and calculating the relative protein abundance. See Fig. 4j for the corresponding IpaA-3xFT protein content quantification. (b) Relative IpaA-3xFT protein content in Shigella wt and ΔmnmG strains carrying an inducible pTac-ipaA-3xFT plasmid and the p-Empty or p-mnmG plasmids, determined by quantification of western blots of serially diluted samples. Shown are 3 independent western blots of serially diluted samples for each strain, used for generating standard curves and calculating the relative protein abundance. See Fig. 4j for the corresponding IpaA-3xFT protein content quantification. (c) BO enteroids were infected with mixed barcoded consortia comprising the indicated strains at MOI 40 for 6 h, as shown in Fig. 4k. The relative abundance of each tag in the input consortia is plotted as percentage of the total population. Shown are data for two independently generated consortia for each infection. (d) BO enteroids were infected with mixed barcoded consortia comprising the indicated strains at MOI 40 for 6 h, as shown in Fig. 4l. The relative abundance of each tag in the input consortia is plotted as percentage of the total input population. Shown are data for two independently generated consortia for each infection.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–3.
Supplementary Data 1 (download XLSX )
Genome-wide relative mutant fitness as scored by TraDIS using a Shigella Tn5 library grown in different broth conditions. log2FC, log2CPM and FDR values for the comparative analysis of each coding gene and pseudogene. Supporting data for Fig. 1.
Supplementary Data 2 (download XLSX )
Gene expression data of Shigella wt grown at 30° or 37 °C (Temperature-regulated genes), Shigella wt versus a virF mutant grown at 37 °C (VirF-regulated genes) or Shigella wt versus a ΔvirF mutant grown at 30 °C. log2FC, log2CPM and FDR values for the comparative analysis of each coding gene and pseudogene. Supporting data for Fig. 1.
Supplementary Data 3 (download XLSX )
Shigella TraDIS screen in BO enteroids. log2FC, log2CPM and FDR values for the comparative analysis of each coding gene and pseudogene. Supporting data for Fig. 2.
Supplementary Data 4 (download XLSX )
Shigella TraDIS screen in BO enteroids. log2FC, log2CPM and FDR values for the comparative analysis of the 105 significant advantageous chromosomal genes (mutant less invasive) and the 61 detrimental chromosomal genes (mutant more invasive) for enteroid colonization. References with comments for Shigella genes previously implicated in virulence or altered during epithelial colonization are included. Supporting data for Fig. 2.
Supplementary Data 5 (download XLSX )
Proteomic profiling of Shigella ΔmnmE and ΔmnmG mutants. log2FC and adjusted P values for the comparative analysis of each coding gene. Supporting data for Fig. 4.
Supplementary Tables 1–3 (download XLSX )
Table 1, Bacterial strains used in this study. Table 2, Plasmids used in this study. Table 3, Oligonucleotides used in this study.
Supplementary Video 1 (download MP4 )
Representative time-lapse video of a BO enteroid infected with the Shigella wt strain containing the cytosolic reporter puhpT-GFP at MOI 40. Supporting video for Fig. 1a.
Supplementary Video 2 (download MP4 )
Representative time-lapse video of a BO enteroid infected with the Shigella wt strain containing the constitutive reporter p-mCherry at MOI 40. White arrows, Shigella spread between neighboring IEC. Supporting video for Fig. 1b.
Source data
Source Data Figs. 1, 3 and 4, Extended Data Figs. 1–7 (download XLSX )
Source data for plotted values and exact P values.
Source Data Fig. 4j and Extended Data Fig. 7a,b (download PDF )
Unprocessed western blots/gels.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Di Martino, M.L., Jenniches, L., Bhetwal, A. et al. A scalable gut epithelial organoid model reveals the genome-wide colonization landscape of a human-adapted pathogen. Nat Genet 57, 1730–1741 (2025). https://doi.org/10.1038/s41588-025-02218-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02218-x