Abstract
Polycystic ovary syndrome (PCOS), the leading endocrine disorder in women of reproductive age, is highly heritable, yet its polygenic architecture remains poorly understood. Here we conducted a genome-wide association study on 12,419 Chinese women with PCOS and 34,235 controls, followed by a multi-ancestry meta-analysis with up to 13,773 European cases and 411,088 controls, identifying 94 independent loci, 73 of which were previously unreported. Despite different evolutionary pressures, Chinese and European ancestries showed substantial genetic overlap. Integrative functional analyses prioritized regulatory variants controlling gene activity in specific tissues, disease-causing genes including anti-Müllerian hormone (AMH), and biological pathways involving ligand-binding domain interactions and peroxisome proliferator-activated receptor gamma (PPARG) signaling. We identified granulosa cells as particularly important in PCOS development. Our genetics-driven drug discovery approach revealed multiple drug targets and repurposing opportunities, enabling personalized treatment strategies. These results enhance our understanding of the molecular basis of PCOS, paving the way for precision medicine.
This is a preview of subscription content, access via your institution
Access options








Similar content being viewed by others
Data availability
Summary statistics for the meta-analyses presented in this paper have been deposited in OMIX at the National Genomics Data Center (NGDC) (https://ngdc.cncb.ac.cn/omix; accession number OMIX010148, https://ngdc.cncb.ac.cn/omix/release/OMIX010148) and are freely and openly accessible for research activities. We used publicly available data from Apollo–University of Cambridge Repository (https://doi.org/10.17863/CAM.36024), the FinnGen Freeze 10 cohort (https://www.finngen.fi/en/access_results), the NHGRI-EBI GWAS Catalog (https://www.ebi.ac.uk/gwas/studies/GCST90044902), the IEU OpenGWAS project (https://gwas.mrcieu.ac.uk/) and the BioBank Japan PheWeb (https://pheweb.jp/). RNA-sequencing data of adult human ovarian tissue are derived from Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/) with accession id GSE118127. Source data are provided with this paper.
Code availability
The following computer code was used in this paper: RICOPILI (v.2019_Jun_25.001, https://github.com/Ripkelab/ricopili/wiki)56 for quality control, principal-component analysis, pre-phasing, imputation, association testing and meta-analysis, with several embedded tools including EIGENSOFT (v.6.1.4, https://github.com/DReichLab/EIG)96, Eagle (v.2.3.5, https://github.com/poruloh/Eagle)58, Minimac3 (v.2.0.1, https://github.com/Santy-8128/Minimac3)59, PLINK (v.1.90b6.10, https://www.cog-genomics.org/plink/1.9/)97 and METAL (v.2011-03-25, https://github.com/statgen/METAL)61; LDSC (v.1.0.1, https://github.com/bulik/ldsc)18 for heritability and genetic correlation estimations; Popcorn (v.0.9.9, https://github.com/brielin/Popcorn)47 for trans-ancestry genetic correlation analysis; MAGMA (v.1.10, https://cncr.nl/research/magma/)27 and VEGAS2 (v.2.01.17, https://github.com/raydai/VEGAS2)28 for pathway analyses; DEPICT (v.1rel173, https://github.com/perslab/depict)29 for post GWAS analyses. Seurat (v.4.3.0, https://github.com/satijalab/seurat)71 for single-cell analysis; and SuSiE (v.0.14.2, https://stephenslab.github.io/susieR)78, SuSiEx (v.1.1.2, https://github.com/getian107/SuSiEx)33 and echolocatoR (v.2.0.3, https://github.com/RajLabMSSM/echolocatoR)79 for fine-mapping.
We also used PWCoCo (v.1.1.1, https://github.com/jwr-git/pwcoco)73 for pairwise conditional and colocalization analysis, LocusCompareR (v.1.0.0, https://github.com/boxiangliu/locuscomparer)77 for visualization of colocalization events, Metascape (v.3.5, https://metascape.org/)83 for gene annotation and analysis, PRSice (v.2.3.5, https://choishingwan.github.io/PRSice/)85 for PRS analyses, TwoSampleMR (v.0.6.3, https://mrcieu.github.io/TwoSampleMR/)95 for Mendelian randomization analyses and GREP (v.1.0.0, https://github.com/saorisakaue/GREP)35 and DGIdb (v.5.0.7, https://github.com/dgidb/dgidb-v5)38 for drug analyses.
References
Joham, A. E. et al. Polycystic ovary syndrome. Lancet Diabetes Endocrinol. 10, 668–680 (2022).
Azziz, R. et al. Polycystic ovary syndrome. Nat. Rev. Dis. Prim. 2, 16057 (2016).
Stener-Victorin, E. et al. Polycystic ovary syndrome. Nat. Rev. Dis. Prim. 10, 27 (2024).
Liu, X., Zhang, J. & Wang, S. Global, regional, and national burden of infertility attributable to PCOS, 1990-2019. Hum. Reprod. 39, 108–118 (2024).
Chai, O. et al. Global and regional burden of polycystic ovary syndrome, 1990–2021: a systematic analysis of the global burden of disease study 2021. BMC Women’s Health 25, 437 (2025).
Dapas, M. & Dunaif, A. Deconstructing a syndrome: genomic insights into PCOS causal mechanisms and classification. Endocr. Rev. 43, 927–965 (2022).
Franks, S., Stark, J. & Hardy, K. Follicle dynamics and anovulation in polycystic ovary syndrome. Hum. Reprod. Update 14, 367–378 (2008).
Rosenfield, R. L. & Ehrmann, D. A. The pathogenesis of polycystic ovary syndrome (PCOS): the hypothesis of PCOS as functional ovarian hyperandrogenism revisited. Endocr. Rev. 37, 467–520 (2016).
Chen, Z.-J. et al. Genome-wide association study identifies susceptibility loci for polycystic ovary syndrome on chromosome 2p16.3, 2p21 and 9q33.3. Nat. Genet. 43, 55–59 (2011).
Shi, Y. et al. Genome-wide association study identifies eight new risk loci for polycystic ovary syndrome. Nat. Genet. 44, 1020–1025 (2012).
Dapas, M. et al. Distinct subtypes of polycystic ovary syndrome with novel genetic associations: an unsupervised, phenotypic clustering analysis. PLoS Med. 17, e1003132 (2020).
Hayes, M. G. et al. Genome-wide association of polycystic ovary syndrome implicates alterations in gonadotropin secretion in European ancestry populations. Nat. Commun. 6, 7502 (2015).
Lee, H. et al. Genome-wide association study identified new susceptibility loci for polycystic ovary syndrome. Hum. Reprod. 30, 723–731 (2015).
Tyrmi, J. S. et al. Leveraging Northern European population history: novel low-frequency variants for polycystic ovary syndrome. Hum. Reprod. 37, 352–365 (2022).
Dou, Y. et al. DENND1A desensitizes granulosa cells to FSH by arresting intracellular FSHR transportation. Sci. China Life Sci. 67, 1620–1634 (2024).
McAllister, J. M. et al. Overexpression of a DENND1A isoform produces a polycystic ovary syndrome theca phenotype. Proc. Natl Acad. Sci. USA 111, E1519–E1527 (2014).
Zhang, Y. et al. THADA inhibition in mice protects against type 2 diabetes mellitus by improving pancreatic β-cell function and preserving β-cell mass. Nat. Commun. 14, 1020 (2023).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Day, F. et al. Large-scale genome-wide meta-analysis of polycystic ovary syndrome suggests shared genetic architecture for different diagnosis criteria. PLoS Genet. 14, e1007813 (2018).
Dong, S. et al. Annotating and prioritizing human non-coding variants with RegulomeDB v.2. Nat. Genet. 55, 724–726 (2023).
Dodé, C. et al. Loss-of-function mutations in FGFR1 cause autosomal dominant Kallmann syndrome. Nat. Genet. 33, 463–465 (2003).
Gill, J. C., Moenter, S. M. & Tsai, P.-S. Developmental regulation of gonadotropin-releasing hormone neurons by fibroblast growth factor signaling. Endocrinology 145, 3830–3839 (2004).
Naguib, A. et al. SUPT4H1 Depletion Leads to a Global Reduction in RNA. Cell Rep. 26, 45–53 (2019).
Heikinheimo, M. et al. Expression and hormonal regulation of transcription factors GATA-4 and GATA-6 in the mouse ovary. Endocrinology 138, 3505–3514 (1997).
Hammond, G. L. Diverse roles for sex hormone-binding globulin in reproduction. Biol. Reprod. 85, 431–441 (2011).
Levine, A. J., Tomasini, R., McKeon, F. D., Mak, T. W. & Melino, G. The p53 family: guardians of maternal reproduction. Nat. Rev. Mol. Cell Biol. 12, 259–265 (2011).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 11, e1004219 (2015).
Mishra, A. & Macgregor, S. VEGAS2: software for more flexible gene-based testing. Twin Res Hum. Genet 18, 86–91 (2015).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Fan, X. et al. Single-cell reconstruction of follicular remodeling in the human adult ovary. Nat. Commun. 10, 3164 (2019).
Yang, R. et al. Changes in the prevalence of polycystic ovary syndrome in China over the past decade. Lancet Reg. Health West Pac. 25, 100494 (2022).
Chiaffarino, F. et al. Prevalence of polycystic ovary syndrome in European countries and USA: a systematic review and meta-analysis. Eur. J. Obstet. Gynecol. Reprod. Biol. 279, 159–170 (2022).
Yuan, K. et al. Fine-mapping across diverse ancestries drives the discovery of putative causal variants underlying human complex traits and diseases. Nat. Genet. 56, 1841–1850 (2024).
Sendur, S. N. & Yildiz, B. O. Influence of ethnicity on different aspects of polycystic ovary syndrome: a systematic review. Reprod. Biomed. Online 42, 799–818 (2021).
Sakaue, S. & Okada, Y. GREP: genome for REPositioning drugs. Bioinformatics 35, 3821–3823 (2019).
Franik, S., Eltrop, S. M., Kremer, J. A., Kiesel, L. & Farquhar, C. Aromatase inhibitors (letrozole) for subfertile women with polycystic ovary syndrome. Cochrane Database Syst. Rev. 5, CD010287 (2018).
Xu, X.-L., Deng, S.-L., Lian, Z.-X. & Yu, K. Estrogen receptors in polycystic ovary syndrome. Cells 10, 459 (2021).
Cannon, M. et al. DGIdb 5.0: rebuilding the drug-gene interaction database for precision medicine and drug discovery platforms. Nucleic Acids Res. 52, D1227–D1235 (2024).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Teede, H. J. et al. Recommendations from the 2023 international evidence-based guideline for the assessment and management of polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 108, 2447–2469 (2023).
Cataldo, N. A. et al. Metabolic and ovarian effects of rosiglitazone treatment for 12 weeks in insulin-resistant women with polycystic ovary syndrome. Hum. Reprod. 21, 109–120 (2006).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Kevenaar, M. E. et al. A functional anti-mullerian hormone gene polymorphism is associated with follicle number and androgen levels in polycystic ovary syndrome patients. J. Clin. Endocrinol. Metab. 93, 1310–1316 (2008).
Dharani, V., Nishu, S. & Hariprasath, L. PCOS and genetics: exploring the heterogeneous role of potential genes in ovarian dysfunction, a hallmark of PCOS—a review. Reprod. Biol. 25, 101017 (2025).
Speed, D., Kaphle, A. & Balding, D. J. SNP-based heritability and selection analyses: Improved models and new results. Bioessays 44, e2100170 (2022).
Browning, S. R. & Browning, B. L. Population structure can inflate SNP-based heritability estimates. Am. J. Hum. Genet. 89, 191–193 (2011).
Brown, B. C., Ye, C. J., Price, A. L. & Zaitlen, N. Transethnic genetic-correlation estimates from summary statistics. Am. J. Hum. Genet. 99, 76–88 (2016).
Lam, M. et al. Comparative genetic architectures of schizophrenia in East Asian and European populations. Nat. Genet. 51, 1670–1678 (2019).
Duncan, L. et al. Analysis of polygenic risk score usage and performance in diverse human populations. Nat. Commun. 10, 3328 (2019).
Liu, Q. et al. A genome-wide cross-trait analysis identifies shared loci and causal relationships of type 2 diabetes and glycaemic traits with polycystic ovary syndrome. Diabetologia 65, 1483–1494 (2022).
Casarini, L. & Brigante, G. The polycystic ovary syndrome evolutionary paradox: a genome-wide association studies-based, in silico, evolutionary explanation. J. Clin. Endocrinol. Metab. 99, E2412–E2420 (2014).
Sova, H. et al. Hormone profiling, including anti-Müllerian hormone (AMH), for the diagnosis of polycystic ovary syndrome (PCOS) and characterization of PCOS phenotypes. Gynecol. Endocrinol. 35, 595–600 (2019).
VanHise, K. et al. Racial and ethnic disparities in polycystic ovary syndrome. Fertil. Steril. 119, 348–354 (2023).
Affimetrix. Axiom Genotyping Solution Data Analysis Guide (Affimetrix, 2016).
Illumina. GenomeStudio Genotyping Module v2.0 Software Guide (Illumina, 2016).
Lam, M. et al. RICOPILI: rapid imputation for consortias pipeline. Bioinformatics 36, 930–933 (2020).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Loh, P.-R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet 48, 1443–1448 (2016).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet 48, 1284–1287 (2016).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Ghoussaini, M. et al. Open Targets Genetics: systematic identification of trait-associated genes using large-scale genetics and functional genomics. Nucleic Acids Res. 49, D1311–D1320 (2021).
Mountjoy, E. et al. An open approach to systematically prioritize causal variants and genes at all published human GWAS trait-associated loci. Nat. Genet. 53, 1527–1533 (2021).
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
Durbin, R. M. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Liu, J. Z. et al. A versatile gene-based test for genome-wide association studies. Am. J. Hum. Genet. 87, 139–145 (2010).
Tanaka, T. et al. Cellular senescence of granulosa cells in the pathogenesis of polycystic ovary syndrome. Mol. Hum. Reprod. 30, gaae015 (2024).
Li, S. et al. Deconvolution at the single-cell level reveals ovarian cell-type-specific transcriptomic changes in PCOS. Reprod. Biol. Endocrinol. 22, 24 (2024).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Robinson, J. W., et al. An efficient and robust tool for colocalisation: Pair-wise Conditional and Colocalisation (PWCoCo). Preprint at bioRxiv https://doi.org/10.1101/2022.08.08.503158 (2022).
Zheng, J. et al. Phenome-wide Mendelian randomization mapping the influence of the plasma proteome on complex diseases. Nat. Genet. 52, 1122–1131 (2020).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Liu, B., Gloudemans, M. J., Rao, A. S., Ingelsson, E. & Montgomery, S. B. Abundant associations with gene expression complicate GWAS follow-up. Nat. Genet. 51, 768–769 (2019).
Wang, G., Sarkar, A., Carbonetto, P. & Stephens, M. A simple new approach to variable selection in regression, with application to genetic fine mapping. J. R. Stat. Soc. Ser. B 82, 1273–1300 (2020).
Schilder, B. M., Humphrey, J. & Raj, T. echolocatoR: an automated end-to-end statistical and functional genomic fine-mapping pipeline. Bioinformatics 38, 536–539 (2022).
Wright, S. The genetical structure of populations. Ann. Eugen. 15, 323–354 (1951).
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Zhang, M. et al. Genetic variants underlying differences in facial morphology in East Asian and European populations. Nat. Genet. 54, 403–411 (2022).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Choi, S. W. & O'Reilly, P. F. PRSice-2: polygenic risk score software for biobank-scale data. Gigascience 8, giz082 (2019).
He, Y., Koido, M., Shimmori, Y. & Kamatani, Y. GWASLab: a Python package for processing and visualizing GWAS summary statistics. Preprint at Jxiv, https://doi.org/10.51094/jxiv.370 (2023).
Burgess, S., Butterworth, A. & Thompson, S. G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 37, 658–665 (2013).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Akiyama, M. et al. Genome-wide association study identifies 112 new loci for body mass index in the Japanese population. Nat. Genet. 49, 1458–1467 (2017).
Pulit, S. L. et al. Meta-analysis of genome-wide association studies for body fat distribution in 694 649 individuals of European ancestry. Hum. Mol. Genet 28, 166–174 (2019).
Suzuki, K. et al. Genetic drivers of heterogeneity in type 2 diabetes pathophysiology. Nature 627, 347–357 (2024).
Yengo, L. et al. A saturated map of common genetic variants associated with human height. Nature 610, 704–712 (2022).
Horikoshi, M. et al. Elucidating the genetic architecture of reproductive ageing in the Japanese population. Nat. Commun. 9, 1977 (2018).
Graham, S. E. et al. The power of genetic diversity in genome-wide association studies of lipids. Nature 600, 675–679 (2021).
Hemani, G., Tilling, K. & Davey Smith, G. Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet. 13, e1007081 (2017).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Acknowledgements
We thank the reviewers for their insightful and constructive comments, which substantially improved the paper. We thank all participants and staff of the contributing studies. In particular, we acknowledge the participants and investigators of the FinnGen study. This study was supported by 973 Program (2015CB559100 to Y. Shi), National Key Research and Development Program of China (2021YFC2700400 to H.Z., 2024YFC2707300 to S.Z. and 2023YFA0913804 to H.Z.), National Natural Science Foundation of China (82421004 to H.Z., 32588201 to Z.C., 32370916 to S.Z., 82271540 to Z.L., 82101707 to X.G. and 82071606 to S.Z.), Shandong Provincial Key Research and Development Program (2024CXPT087 to H.Z., 2021ZDSYS06 to Z.L. and 2020ZLYS02 to Z.C.), Ningxia Hui Autonomous Region Key Research and Development Program (2024BEG02019 to H.Z.), Natural Science Foundation of Shandong Province (ZR2023YQ061 to S.Z. and YDZX2021009 to Z.L.), Shanghai Municipal Science and Technology Major Project (2019SHZDZX02 and 2017SHZDZX01 to Y. Shi), MOE Key Laboratory of Population Health Across Life Cycle (JK20232 to Y.C.), CAMS Innovation Fund for Medical Sciences (2021-I2M-5-001 to Z.C.), Taishan Scholars Program of Shandong Province (tstp20240526 to Z.L. and ts20190988 to H.Z.), Fundamental Research Funds for the Central Universities (YG2021ZD2020 to Y. Shi), Medicine Plus Interdisciplinary Cluster Joint Exploration Project of Qingdao Medical University (RZ2400001468) and Fundamental Research Funds of Shandong University (2023QNTD004 to S.Z.).
Author information
Authors and Affiliations
Contributions
Y. Shi, Z.L., Y.C. and H.Z. jointly supervised the research. Z.L., Y. Shi, H.Z. and S.Z. designed and conceived the study. Z.L., B.X., S.Z., M.Z., C.G. and S.L. performed bioinformatics analyses. H.Z., S.Z., Y.C., Y.X., X.W., T.W., S.L., Z.Y., X.G., Z. Wang, C.Z., X.Z., T.P., C.G., C.W., L.P., Y. Sun, Y.D. and Z.C. contributed samples and performed phenotyping. Y.W., Q.Z., H.S., Y.L., X.J., B.W., L.W., Y.H., D.X., Z. Wu, Q.Y., S.D., G.X., Y.J., H.X., W.S., J.L. and L.H. performed genotyping. Z.L., B.X., S.Z., M.Z., H.Z., Y. Shi, S.L. and Y.X. wrote and edited the paper. All authors critically reviewed the paper and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Elisabet Stener-Victorin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Workflow for polycystic ovary syndrome genome-wide association study.
Schematic diagram showing the study design, sample collection, and analysis pipeline used for identifying genetic variants associated with polycystic ovary syndrome (PCOS).
Extended Data Fig. 2 Distribution of genome-wide significant loci across ancestry groups.
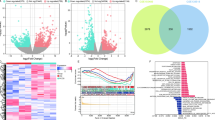
a, Discovery of genome-wide significant loci across analyses. Bar chart showing the number of genome-wide significant loci (P < 5 × 10−8) identified in each analysis, with numbers above bars indicating exact loci counts. b, Overlap of the susceptibility loci between ancestries. Venn diagram illustrating the overlap of significant loci across Chinese (CHN), European (EUR/EUR2), and trans-ancestry (MIX/MIX2) analyses. Analyses included: CHN, Chinese cohort GWAS (12,419 cases; 34,235 controls); EUR, European cohort meta-analysis (8,589 cases; 328,329 controls); EUR2, European cohort meta-analysis focusing on 10 K array variants (13,773 cases; 411,088 controls); MIX, trans-ancestry meta-analysis of Chinese and European cohorts (21,008 cases; 362,564 controls); MIX2, trans-ancestry meta-analysis focusing on 10 K array variants (26,192 cases; 445,323 controls).
Extended Data Fig. 3 Tissue-specific regulatory scores for polycystic ovary syndrome associated lead variants.
Heatmap displaying RegulomeDB tissue-specific regulatory scores for lead variants, where each row represents a lead variant and each column represents a specific tissue. Color intensity indicates regulatory score strength (see color key within figure). Tissues are grouped by biological systems with system-specific colors indicated in the top sidebar. Left sidebars show overall RegulomeDB scores (1a-7, where lower scores indicate stronger regulatory evidence) and variant probability values from Supplementary Table 4.
Extended Data Fig. 4 Enriched biological processes in polycystic ovary syndrome associated genes.
a, Biological process enrichment in the associated gene sets. Heatmap showing enrichment of biological processes across polycystic ovary syndrome (PCOS) associated gene sets from East Asian (EAS) and European (EUR) populations, where color intensity represents -log₁₀(P-value), with darker orange indicating stronger enrichment (see color key within figure). Each row represents a biological process term, and columns represent different gene sets. b, Hierarchical clustering of enriched functional terms. Hierarchical organization of enriched terms according to Gene Ontology (GO) parent-child relationships, with color coding following panel (a). Enrichment analysis employed a one-sided hypergeometric test to identify over-represented biological processes. Multiple testing correction was applied using the Benjamini-Hochberg method with FDR < 0.05 as the significance threshold. Heatmap colors represent −log₁₀(adjusted P-values).
Extended Data Fig. 5 Polygenic risk score model performance across P-value thresholds.
Bar graph showing the variance explained (Nagelkerke’s pseudo-R²) by polygenic risk scores (PRS) constructed using variants meeting different P-value thresholds (x-axis: 5e-08, 1e-05, 0.001, 0.05, 0.1, 0.2, 0.3, 0.4, 0.5, 1). Y-axis shows R2 values (0 to maximum observed value). Bars are grouped by target dataset and colored by ancestry cohort (see color key within figure). Four independent Chinese cohorts include ASA (7,891 cases/12,001 controls), CHB (2,406 cases/4,863 controls), ASI (1,362 cases/10,501 controls), and SNP6 (760 cases/6,870 controls). PRS were constructed using summary statistics from an inverse-variance weighted (IVW) fixed-effects meta-analysis (CHN = Chinese-specific, EUR = European-specific, CHN + EUR = cross-ancestry mixed). All statistical tests were two-sided, with no multiple testing correction applied.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figures 1–5, and Supplementary Note.
Supplementary Table (download XLSX )
Supplementary Tables 1–22.
Supplementary Data (download XLSX )
Source data for supplementary figures.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, H., Xu, Y., Xue, B. et al. Multi-ancestry genome-wide association analyses of polycystic ovary syndrome. Nat Genet 57, 2669–2681 (2025). https://doi.org/10.1038/s41588-025-02393-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02393-x
This article is cited by
-
Genetic polymorphism of long non-coding RNA TUG1 and susceptibility to polycystic ovary syndrome: a case-control study
Hereditas (2026)
-
A novel PCOS susceptibility gene, cathepsin B, might be likely to contribute to the pyroptosis of ovarian granulosa cells
Journal of Assisted Reproduction and Genetics (2026)