Abstract
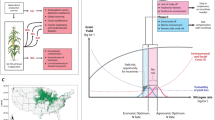
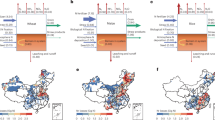
Although nitrogen fertilizer use has boosted crop yields, excessive application diminishes crop nitrogen use efficiency (NUE) and causes environmental problems. Therefore, increasing crop NUE is urgently needed for agricultural sustainability. Through a genome-wide association study, we identified a locus, NCR1 (Nitrate Concentration Regulator 1), that correlates with nitrate concentrations in maize root xylem. NCR1 encodes a MYB transcription factor that positively regulates the transcription of nitrate transporter NRT2.3 expressed predominantly in root xylem parenchyma cells. The NCR1–NRT2.3 transcription module responds to external nitrogen and controls nitrate translocation from roots to shoots. The superior NCR1−In allele with a 123-bp promoter deletion has decreased in frequency as nitrogen fertilizer use in China has increased. Overexpression of NCR1 or NRT2.3, or introgression of NCR1−In, increases grain yield and nitrogen content in the shoot and seed. This study uncovers a crucial genetic module for improving grain yield and NUE in maize.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its Supplementary Information. A reporting summary for this article is available in Supplementary Information. The B73 reference genomic sequences are collected from the MaizeGDB (https://www.maizegdb.org/). RNA-sequencing data and CUT&Tag data can be found in the National Center for Biotechnology Information under accession number PRJNA1117020. Source data are provided with this paper.
Code availability
No custom code was generated. All code used to analyze the sequence data is publicly available in the SAMtools section of GitHub63 (https://github.com/samtools/samtools).
Change history
23 March 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41588-026-02575-1
References
Guo, J. H. et al. Significant acidification in major Chinese croplands. Science 327, 1008–1010 (2010).
Zhang, X. et al. Managing nitrogen for sustainable development. Nature 528, 7580 (2015).
Xu, G., Fan, X. & Miller, A. J. Plant nitrogen assimilation and use efficiency. Annu. Rev. Plant Biol. 63, 153–182 (2012).
Wang, Q. et al. Genetic variations in ARE1 mediate grain yield by modulating nitrogen utilization in rice. Nat. Commun. 9, 735 (2018).
Yan, X. et al. Plant nutriomics in China: an overview. Ann. Bot. 98, 473–482 (2006).
Capstaff, N. M. & Miller, A. J. Improving the yield and nutritional quality of forage crops. Front. Plant Sci. 9, 535 (2018).
Vidal, E. A. et al. Nitrate in 2020: thirty years from transport to signaling networks. Plant Cell 32, 2094–2119 (2020).
Liu, Q. et al. Improving crop nitrogen use efficiency toward sustainable green revolution. Annu. Rev. Plant Biol. 73, 523–551 (2022).
Liu, Y. et al. Genomic basis of geographical adaptation to soil nitrogen in rice. Nature 590, 600–605 (2021).
Tang, W. et al. Genome-wide associated study identifies NAC42-activated nitrate transporter conferring high nitrogen use efficiency in rice. Nat. Commun. 10, 5279 (2019).
Hu, B. et al. Variation in NRT1.1B contributes to nitrate-use divergence between rice subspecies. Nat. Genet. 47, 834–838 (2015).
Sun, H. et al. Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat. Genet. 46, 652–656 (2014).
Li, S. et al. Modulating plant growth-metabolism coordination for sustainable agriculture. Nature 560, 595–560 (2018).
Wu, K. et al. Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice. Science 367, 6478 (2020).
Gao, Y. et al. MYB61 is regulated by GRF4 and promotes nitrogen utilization and biomass production in rice. Nat. Commun. 11, 5219 (2020).
Gao, Z. et al. The indica nitrate reductase gene OsNR2 allele enhances rice yield potential and nitrogen use efficiency. Nat. Commun. 10, 5207 (2019).
Wen, Z. et al. Maize NPF6 proteins are homologs of Arabidopsis CHL1 that are selective for both nitrate and chloride. Plant Cell 29, 2581–2596 (2017).
Cao, H. et al. ZmNRT1.1B (ZmNPF6.6) determines nitrogen use efficiency via regulation of nitrate transport and signalling in maize. Plant Biotechnol. J. 22, 316–329 (2024).
Huang, Y. et al. THP9 enhances seed protein content and nitrogen-use efficiency in maize. Nature 612, 292–300 (2022).
Ota, R., Ohkubo, Y., Yamashita, Y., Ogawa-Ohnishi, M. & Matsubayashi, Y. Shoot-to-root mobile CEPD-like 2 integrates shoot nitrogen status to systemically regulate nitrate uptake in Arabidopsis. Nat. Commun. 11, 641 (2020).
York, L. M., Silberbush, M. & Lynch, J. P. Spatiotemporal variation of nitrate uptake kinetics within the maize (Zea mays L.) root system is associated with greater nitrate uptake and interactions with architectural phenes. J. Exp. Bot. 67, 3763–3775 (2016).
Dubos, C. et al. MYB transcription factors in Arabidopsis. Trends Plant Sci. 10, 573–581 (2010).
Yong, Z., Kotur, Z. & Glass, A. D. Characterization of an intact two component high-affinity nitrate transporter from Arabidopsis roots. Plant J. 63, 739–748 (2010).
Kotur, Z. et al. Nitrate transport capacity of the Arabidopsis thaliana NRT2 family members and their interactions with AtNAR2.1. New Phytol. 194, 724–731 (2012).
Liu, X. et al. Identification and functional assay of the interaction motifs in the partner protein OsNAR2.1 of the two-component system for high-affinity nitrate transport. New Phytol. 204, 74–80 (2014).
Urao, T., Shinozaki, K. Y., Urao, S. & Shinozaki, K. An Arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5, 1529–1539 (1993).
Solano, R. et al. Dual DNA binding specificity of a petal epidermis-specific MYB transcription factor (MYB.Ph3) from Petunia hybrida. EMBO J. 14, 1773–1784 (1995).
Mu, R. L. et al. An R2R3-type transcription factor gene AtMYB59 regulates root growth and cell cycle progression in Arabidopsis. Cell Res. 19, 1291–1304 (2009).
Hufford, M. B. et al. Comparative population genomics of maize domestication and improvement. Nat. Genet. 44, 808–811 (2012).
Chen, W. et al. Convergent selection of a WD40 protein that enhances grain yield in maize and rice. Science 375, eabg7985 (2022).
Yang, N. et al. Two teosintes made modern maize. Science 382, eadg8940 (2023).
Bukowski, R. et al. Construction of the third-generation Zea mays haplotype map. Gigascience. 7, 1–12 (2018).
Wang, B. et al. Genome-wide selection and genetic improvement during modern maize breeding. Nat. Genet. 52, 565–571 (2020).
Tang, Z. et al. Knockdown of a rice stelar nitrate transporter alters long-distance translocation but not root influx. Plant Physiol. 160, 2052–2063 (2012).
Fan, X. et al. Overexpression of a pH-sensitive nitrate transporter in rice increases crop yields. Proc. Natl Acad. Sci. USA 113, 7118–7123 (2016).
Chen, J. et al. Co-overexpression of OsNAR2.1 and OsNRT2.3a increased agronomic nitrogen use efficiency in transgenic rice plants. Front. Plant Sci. 11, 1245 (2020).
Zhang, Z. & Chu, C. Nitrogen-use divergence between Indica and Japonica rice: variation at nitrate assimilation. Mol. Plant 13, 6–7 (2020).
Yan, Y. et al. Nitrate confers rice adaptation to high ammonium by suppressing its uptake but promoting its assimilation. Mol. Plant 16, 1871–1874 (2023).
Toledo-Ortiz, G., Huq, E. & Quail, P. H. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 15, 1749–1770 (2003).
Gao, F. & Dubos, C. The arabidopsis bHLH transcription factor family. Trends Plant Sci. 29, 668–680 (2024).
Li, B. Z., Xin, W. J., Sun, S. B., Shen, Q. R. & Xu, G. H. Physiological and molecular responses of nitrogen-starved rice plants to re-supply of different nitrogen sources. Plant Soil 287, 145–159 (2006).
Zhou, X. & Stephens, M. Genome-wide efficient mixed model analysis for association studies. Nat. Genet. 44, 821–824 (2013).
Yu, J. M. et al. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat. Genet. 38, 203–208 (2006).
Gao, L. et al. Genetic variation in a heat shock transcription factor modulates cold tolerance in maize. Mol. Plant 17, 1423–1438 (2024).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Zhang, C., Dong, S., Xu, J., He, W. & Yang, T. PopLDdecay: a fast and effective tool for linkage disequilibrium decay analysis based on variant call format files. Bioinformatics 35, 1786–1788 (2019).
Ishida, Y., Hiei, Y. & Komari, T. Agrobacterium-mediated transformation of maize. Nat. Protoc. 2, 1614–1621 (2007).
Tian, T. et al. agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 45, W122–W129 (2017).
Chen, C. J. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Munns, R. et al. Wheat grain yield on saline soils is improved by an ancestral Na⁺ transporter gene. Nat. Biotechnol. 30, 360–364 (2012).
Batistič, O., Sorek, N., Schultke, S., Yalovsky, S. & Kudla, J. Dual fatty acyl modification determines the localization and plasma membrane targeting of CBL/CIPK Ca2+ signaling complexes in Arabidopsis. Plant Cell 20, 1346–1362 (2008).
Batistič, O., Waadt, R., Steinhorst, L., Held, K. & Kudla, J. CBL-mediated targeting of CIPKs facilitates the decoding of calcium signals emanating from distinct cellular stores. Plant J. 61, 211–222 (2010).
Kaufmann, K. et al. Chromatin immunoprecipitation (ChIP) of plant transcription factors followed by sequencing (ChIP–seq) or hybridization to whole genome arrays (ChIP–ChIP). Nat. Protoc. 5, 457–472 (2010).
Haring, M. et al. Chromatin immunoprecipitation: optimization, quantitative analysis and data normalization. Plant Methods 3, 11 (2007).
Wu, L. et al. A cost-effective tsCUT&Tag method for profiling transcription factor binding landscape. J. Integr. Plant Biol. 64, 2033–2038 (2022).
Chen, S. Ultrafast one-pass FASTQ data preprocessing, quality control, and deduplication using fastp. iMeta 2, e107 (2023).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Jefferson, R. A. Assaying chimeric genes in plants: the GUS gene fusion system. Plant Mol. Biol. Rep. 5, 387–405 (1987).
Li, H. et al. NRT1.5/NPF7.3 functions as a proton-coupled H+/K+ antiporter for K+ loading into the xylem in Arabidopsis. Plant Cell 29, 2016–2026 (2017).
Tsay, Y.-F., Schroeder, J. I., Feldmann, K. A. & Crawford, N. M. The herbicide sensitivity gene CHL1 of Arabidopsis encodes an nitrate-inducible nitrate transporter. Cell 72, 705–713 (1993).
Moll, R. H., Kamprath, E. J. & Jackson, W. A. Analysis and interpretation of factors which contribute to efficiency of nitrogen utilization. Agron. J. 74, 562–564 (1982).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Acknowledgements
We thank the High-performance Computing Platform of China Agricultural University for its support of large-scale computation. We acknowledge Beijing PARATERA Tech CO., Ltd (https://paratera.com/) for providing HPC resources that have contributed to the research results reported within this study. The transgenic seeds of maize were created by the Center for Crop Functional Genomics and Molecular Breeding of China Agricultural University. This work was supported by grants from the National Key Research and Development Program of China (2021YFF1000500), National Natural Science Foundation of China (32025004, 32425041, 32302660), STI2030-Major Projects (2023ZD0406704, 2023ZD04069, 2023ZD04071), China Postdoctoral Science Foundation (2021M703535), Pinduoduo-China Agricultural University Research Fund (PC2023A01004) and Beijing Outstanding University Discipline Program.
Author information
Authors and Affiliations
Contributions
J.L. and Y.W. directed the research and supervised the project. M.Z. and Z.W. performed most of the experiments. M.Z., C.B., Y.W. and J.L. wrote the manuscript. Z.W., M.Z. and Y.Q. performed the field tests in Beijing, Sanya and Gongzhuling. M.Z., Z.W. and J.F. collected and analyzed the field data. L.H. and X.S. provided the sequencing data of inbred lines. K.W., Y.H., B.W., Y.Q., J.F., Z.L. and B.Y. contributed to the seedlings culture in greenhouse and sample collection for GWAS. X.Z., X.W., F.Q., H.Z., J.C. and W.S. contributed to the generation and collection of biological materials and participated in project discussions.
Corresponding authors
Ethics declarations
Competing interests
Two patent applications related to this work have been submitted by J.L., Y.W., M.Z., Z.W., W.S., H.Z., J.C. and C.B. (application nos. 202411076358.2 (PCT/CN2025/108660) and 202411077906.3 (PCT/CN2025/108631)). The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Hussam Nour-Eldin, Peng Yu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification and molecular characterization of NCR1.
(a) Phylogenetic tree of 181 maize inbred lines based on UPGMA. Among these lines, 104 accessions are sourced from China, 35 accessions from the United States, 42 accessions from other countries around the globe. Using a Bayesian approach and phylogenetic tree method, eight distinct subpopulations were identified and validated, comprising 19 Tropical, 19 Lancaster, 16 Reid, 12 TSPT, 11 Iodent, 11 X-group, 10 P-group, 9 A632, and 74 mixed individuals. (b) Population structure of 181 inbred lines assessed by STRUCTURE. Eight ancestral populations were identified, represented by distinct colors: tropical (red), X-group (blue), Iodent (green), TPST (brown), P-group (purple), Reid (orange), A632 (cyan), and Lancaster (pink). Each vertical line represents an inbred line; colored segment height represents membership percentage in each ancestral population. (c) Genome-wide linkage disequilibrium analysis of 181 maize inbred lines. The values 1.6 kb and 28 kb at the red dashed lines represent the distance between molecular markers when r2 decreases to 0.2 or 0.15, respectively. (d) NO3− concentration in root xylem sap under LN condition. (e) Distribution of NO3− concentration in xylem sap under LN and HN conditions. (f) Local Manhattan plot over the 127.75-127.85 Mb region on Chr 3. The SNPs exceeding the threshold are labeled red dots and located on or near the NCR1 gene. (g) Gene structure of NCR1 and mutation sites of ncr1 mutant lines. Black boxes represent exons, and lines represent introns. The sgRNA target site and the PAM motif are indicated. A single base-pair insertion (A) in ncr1−1 is shown in blue. The deleted sequences in ncr1−2 (17 bp) and ncr1−3 (14 bp) are shown using blue dashes. (h) Sequence difference between the NCR1−In and NCR1+In promoters. The NCR1−In promoter contains an E-box motif. (i) Subcellular localization of NCR1-GFP protein in maize mesophyll cell protoplasts. The nucleus was visualized using DAPI staining. (j) Transcript levels of NCR1 in the roots of NCR1 OE lines. Data are mean ± SE. n represents numbers of biologically independent samples. Two-sided Student’s t-test was used to determine P values.
Extended Data Fig. 2 RNA-seq analysis of NCR1-regulated genes in root.
(a) Volcano plots showing the differentially expressed genes (DEGs) in root regulated by NCR1 in response to LN treatment. Red and blue dots represent upregulated and downregulated genes, respectively (absolute fold change >1.5, P < 0.05). Three biological replicates were performed. (b) Hierarchical clustering and heatmap of 1,298 NCR1-regulated DEGs and key GO terms. DEGs show segregation into nine co-expressed clusters. GO enrichment analysis of heatmap DEG clusters is shown in Supplementary Table 7. (c) Venn diagram showing the overlap between NCR1-regulated genes under LN conditions and LN-responsive genes. (d) A heat map of DEGs involved in nitrate absorption and nitrogen metabolism. The color key (blue to red) represents gene expression (FPKM) as fold changes. (e) Distributions of NCR1 binding sites at the NRT1.1B, NiR1, and GS1-1 genes shown as Integrated Genome Browser windows. Notable peaks were calculated by MACS2. The bottom track indicates genes with transcription direction on the chromosome. Blue bars indicate NCR1 binding peaks in the promoter regions. (f) Transcript levels of NR4, NiR1, and GS1-1 genes in the NCR1−In (n = 14 accessions) and NCR1+In (n = 21 accessions) maize inbred lines after HN and LN treatment for 15 days. Statistical significance was determined by a two-sided t-test. The horizontal bars of boxes represent minima, 25th percentiles, medians, 75th percentiles, and maxima.
Extended Data Fig. 3 NRT2.3 expression pattern and NO3− transport analyses.
(a) NRT2.3 expression levels in different tissues (n = 3 biologically independent samples). (b) GUS staining showing transcriptional change of NRT2.3 in response to nitrogen supply using ProNRT2.3:GUS plants. Longitudinal sections (a and b) and cross sections (c to f) of the primary root are shown. (c) Subcellular localization of NRT2.3-GFP in tobacco (Nicotiana benthamiana) leaves. The AtCBL1n-OFP and AtCBL2-mCherry were used as the PM and tonoplast markers, respectively. (d) Nitrate uptake assay in Xenopus oocytes. Oocytes expressing different cRNA combinations were incubated in the solution containing 0.25 or 5 mM K15NO3 at pH 5.5 or 7.5 for 6 h. Then, the 15N content in oocytes was measured. Data are mean ± SE. Each point represents one replicate from one oocyte. n represents numbers of biologically independent samples. Different letters indicate significant differences (P < 0.05) based on one-way ANOVA (Tukey’s test). (e) Mutation sites in two nrt2.3 mutant lines. (f) Transcript levels of NRT2.3 in the roots of NRT2.3 OE lines. Data are mean ± SE. n represents numbers of biologically independent samples. Different letters indicate significant differences (P < 0.05) based on one-way ANOVA (Tukey’s test). (g) to (i) 15N content (g), shoot/root 15N ratio (h), and total N content (i) in the WT and nrt2.3 mutants. The seedlings were cultured in 0.4 mM 15NO3− solution for 1 day. Then, the 15N content in the shoot and root was measured, respectively. Data are mean ± SE. Shoot and root samples from four seedlings were mixed as one biological replicate for each material. n represents numbers of biologically independent samples. Different letters indicate significant difference (P < 0.05) based on the one-way ANOVA (Tukey’s test). (j) Positions of P1 to P4 fragments in the NRT2.3 promoter. MYB core and MRE cis-elements are shown. Gray lines represent MYB core (CNGTTR). Red lines indicate MRE (AACCaaa).
Extended Data Fig. 4 NRT2.3 is genetically epistatic to NCR1.
(a) Phenotype comparison among WT, ncr1-1, nrt2.3-1 single mutant, and ncr1-1 nrt2.3-1 double mutant after LN and HN treatment for 14 days. (b) Mutation sites of NCR1 and NRT2.3 in the double mutant. (c) to (e) Shoot dry weight (c), NO3− concentration in xylem sap (d), and shoot nitrogen content (e) of different plants after LN and HN treatment for 14 days. Data are mean ± SE. Each point represents one biological replicate from one seedling. n represents numbers of biologically independent samples. Different letters indicate significant difference (P < 0.05) based on the one-way ANOVA (Tukey’s test). (f) Phenotype comparison among WT, ncr1-1, NRT2.3 OE-2, and ncr1-1 NRT2.3 OE-2 plants after LN and HN treatment for 15 days. (g) Mutation sites of NCR1 in the ncr1-1 NRT2.3 OE-2 plants. (h) to (j) NRT2.3 transcript level (h) (n = 3 biologically independent samples), NO3− concentration in xylem sap (i), and shoot nitrogen content (j) of different plants after LN and HN treatment for 15 days. Data are mean ± SE. Each point in (h) represents one biological replicate from four seedlings. Each point in (i) and (j) represents one biological replicate from one seedling. n represents numbers of biologically independent samples. Different letters indicate significant difference (P < 0.05) based on the one-way ANOVA (Tukey’s test).
Extended Data Fig. 5 Domestication analysis of NRT2.1, NRT2.2, and NRT2.3.
(a) XP-CLR values between maize and parviglumis. The region (70-71 Mb) in Chr 5 containing the NRT2.3 gene (red) is shown. (b) and (c) Nucleotide diversity (π) of NRT2.1 (b) and NRT2.2 (c) genes among teosinte, maize landrace, and maize inbred line. Nucleotide diversity (π) values were calculated using a 1000-bp sliding window and a 100-bp step. (d) Sequence comparison of NRT2.3 genes from 11 teosinte varieties, 7 landraces, and 8 maize inbred lines. The green box indicates the position of the 536-bp Harbinger-like TE.
Extended Data Fig. 6 Ear phenotype and yield traits of ncr1 and nrt2.3 inbred lines in field test.
Ear phenotype (a and b) and kernel weight per ear (c and d) of ncr1 and nrt2.3 inbred lines under two nitrogen conditions in the field test in Sanya, China, in 2023. The data in (c) and (d) were obtained from at least 38 plants for each line. Data are mean ± SE. Different letters indicate significant difference (P < 0.05) based on the one-way ANOVA (Tukey’s test).
Extended Data Fig. 7 Ear phenotype, yield traits, and nitrogen content of NCR1 OE and NRT2.3 OE inbred lines in field test.
Ear phenotype (a and b), kernel weight per ear (c), NUE (d), leaf nitrogen content (e), and stem nitrogen content (f) of NCR1 OE and NRT2.3 OE inbred lines under two nitrogen conditions in the field test in Sanya, China, in 2023. Data are mean ± SE. The data in (c) were obtained from 60 plants for each line. The bars within violin plots represent 25th percentiles, medians, and 75th percentiles. The data in (d) were derived from at least 3 plots; each line contained 39 plants in one plot. The ear leaves and ear stems were collected for nitrogen content measurement. n represents numbers of biologically independent samples. The horizontal bars of boxes in (e) and (f) represent minima, 25th percentiles, medians, 75th percentiles, and maxima. Different letters indicate significant difference (P < 0.05) based on the one-way ANOVA (Tukey’s test).
Extended Data Fig. 8 Yield traits of NCR1 OE and NRT2.3 OE hybrid lines in the field test in Gongzhuling.
Ear weight (a), ear length (b), kernel number per row (c), kernel row number (d), ear diameter (e), and 100-kernel weight (f) of NCR1 OE and NRT2.3 OE hybrids under two nitrogen conditions in field test in Gongzhuling, China, 2023. Data are mean ± SE. The data in (a) were obtained from 60 plants for each line. In (b) to (f), n represents numbers of biologically independent samples. The horizontal bars of boxes in (a) and (f) represent minima, 25th percentiles, medians, 75th percentiles, and maxima. Different letters indicate significant difference (P < 0.05) based on the one-way ANOVA (Tukey’s test).
Extended Data Fig. 9 Yield traits of NCR1 OE and NRT2.3 OE hybrid lines in the field test in Beijing and Sanya.
Kernel yield per plot (a and e), NUE (b and f), ear weight (c and g), and ear length (d and h) of NCR1 OE and NRT2.3 OE hybrids under two nitrogen conditions in the field test. The data from (a) to (d) were obtained from the field test in Beijing, China, in 2023. The data from (e) to (h) were obtained from the field test in Sanya, China, in 2023. Data are mean ± SE. The data in (a), (b), (e), and (f) were derived from 3 plots. The data in (c) and (g) were obtained from 60 plants for each line. The bars within violin plots represent 25th percentiles, medians, and 75th percentiles. The data in (d) and (h) were obtained from 15 plants for each line. Different letters indicate significant difference (P < 0.05) based on the one-way ANOVA (Tukey’s test).
Extended Data Fig. 10 NCR1−In frequency analyses.
(a) NCR1−In frequencies in different maize populations. (b) Pearson correlation analysis of NCR1−In frequencies and nitrogen application levels for maize in 11 countries. The nitrogen application levels for maize production in various countries were obtained from the International Fertilizer Association (IFA, https://www.fertilizer.org/). P values are determined by the two-sided Pearson correlation coefficient analysis. (c) The NCR1−In frequencies of maize inbred lines from 11 countries used in (b).
Supplementary information
Supplementary Tables 1–10 (download XLS )
Table 1: The 181 maize inbred lines used in GWAS. Table 2: NCR1 resequencing data of 155 maize inbred lines. Table 3: DEGs in WT after LN treatment. Table 4: DEGs showing opposite changes in the ncr1 mutant and NCR1 OE plants under LN conditions. Table 5: DEGs showing opposite changes in the ncr1 mutant and NCR1 OE plants under HN conditions. Table 6: NRT2.3 genotypes in teosinte, landrace, and maize inbred lines. Table 7: GO enrichment analysis of heatmap DEG clusters. Table 8: NCR1 genotypes in the elite inbred lines in China. Table 9: NCR1 genotypes in two different groups of maize inbred lines in the United States. Table 10: Primer sequences used in this study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.xls.
Source Data Fig. 3 (download XLSX )
Statistical source data.xls.
Source Data Fig. 4 (download XLSX )
Statistical source data.xls.
Source Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, M., Wu, Z., Huang, L. et al. A genetic module boosts grain yield and nitrogen use efficiency by improving nitrate transport in maize. Nat Genet 58, 618–629 (2026). https://doi.org/10.1038/s41588-026-02532-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-026-02532-y