Abstract
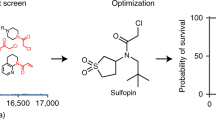
Peptidyl-prolyl cis/trans isomerase NIMA-interacting 1 (Pin1) is commonly overexpressed in human cancers, including pancreatic ductal adenocarcinoma (PDAC). While Pin1 is dispensable for viability in mice, it is required for activated Ras to induce tumorigenesis, suggesting a role for Pin1 inhibitors in Ras-driven tumors, such as PDAC. We report the development of rationally designed peptide inhibitors that covalently target Cys113, a highly conserved cysteine located in the Pin1 active site. The inhibitors were iteratively optimized for potency, selectivity and cell permeability to give BJP-06-005-3, a versatile tool compound with which to probe Pin1 biology and interrogate its role in cancer. In parallel to inhibitor development, we employed genetic and chemical-genetic strategies to assess the consequences of Pin1 loss in human PDAC cell lines. We demonstrate that Pin1 cooperates with mutant KRAS to promote transformation in PDAC, and that Pin1 inhibition impairs cell viability over time in PDAC cell lines.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
CITe-Id data for BJP-06-005-3 are provided as an excel file, Supplementary Dataset 1. RNA-sequencing data have been deposited to NCBI GEO (accession GSE147340), and the analyzed data file is provided as an excel file, Supplementary Dataset 2. All structural data have been deposited in the PDB (PDB codes 6O33 and 6O34).
References
Lu, Z. & Hunter, T. Prolyl isomerase Pin1 in cancer. Cell Res. 24, 1033–1049 (2014).
Lu, K. P., Finn, G., Lee, T. & Nicholson, L. K. Prolyl cis-trans isomerization as a molecular timer. Nat. Chem. Biol. 3, 619–629 (2007).
Yeh, E. S. & Means, A. R. Pin1, the cell cycle and cancer. Nat. Rev. 7, 381–388 (2007).
Lu, K. P. & Zhou, X. Z. The isomerase PIN1 controls numerous cancer-driving pathways and is a unique drug target. Nat. Rev. Cancer 16, 463–478 (2016).
Liang, C. et al. PIN1 maintains redox balance via the c-Myc/NRF2 axis to counteract Kras-induced mitochondrial respiratory injury in pancreatic cancer cells. Cancer Res. 79, 133–145 (2019).
Wulf, G., Garg, P., Liou, Y. C., Iglehart, D. & Lu, K. P. Modeling breast cancer in vivo and ex vivo reveals an essential role of Pin1 in tumorigenesis. EMBO J. 23, 3397–3407 (2004).
Zeitouni, D., Pylayeva-Gupta, Y., Der, C. J. & Bryant, K. L. KRAS mutant pancreatic cancer: no lone path to an effective treatment. Cancers (Basel) 8, 45 (2016).
Hanes, S. D. The Ess1 prolyl isomerase: traffic cop of the RNA polymerase II transcription cycle. Biochim. Biophys. Acta 4, 316–333 (2014).
Liou, Y. C. et al. Loss of Pin1 function in the mouse causes phenotypes resembling cyclin D1-null phenotypes. Proc. Natl Acad. Sci. USA 99, 1335–1430 (2001).
Hennig, L. et al. Selective inactivation of parvulin-like peptidyl-prolyl cis/trans isomerases by juglone. Biochemistry 37, 5953–5960 (1998).
Wei, S. et al. Active Pin1 is a key target of all-trans retinoic acid in acute promyelocytic leukemia and breast cancer. Nat. Med. 21, 457–466 (2015).
Kozono, S. et al. Arsenic targets Pin1 and cooperates with retinoic acid to inhibit cancer-driving pathways and tumor-initiating cells. Nat. Commun. 9, 3069 (2018).
Campaner, E. et al. A covalent PIN1 inhibitor selectively targets cancer cells by a dual mechanism of action. Nat. Commun. 8, 15772 (2017).
Ieda, N. et al. An irreversible inhibitor of peptidyl-prolyl cis/trans isomerase Pin1 and evaluation of cytotoxicity. Bioorg. Med. Chem. Lett. 29, 353–356 (2018).
Moore, J. D. & Potter, A. Pin1 inhibitors: pitfalls, progress and cellular pharmacology. Bioorg. Med. Chem. Lett. 23, 4283–4291 (2013).
Mah, R., Thomas, J. R. & Shafer, C. M. Drug discovery considerations in the development of covalent inhibitors. Bioorg. Med. Chem. Lett. 24, 33–39 (2014).
Liu, Q. et al. Developing irreversible inhibitors of the protein kinase cysteinome. Chem. Biol. 20, 146–159 (2013).
Chen, C. H. et al. Pin1 cysteine-113 oxidation inhibits its catalytic activity and cellular function in Alzheimer’s disease. Neurobiol. Dis. 76, 13–23 (2015).
Zhang, Y. et al. Structural basis for high-affinity peptide inhibition of human Pin1. ACS Chem. Biol. 2, 320–328 (2007).
Nabet, B. et al. The dTAG system for immediate and target-specific protein degradation. Nat. Chem. Biol. 14, 431–441 (2018).
Backus, K. M. et al. Proteome-wide covalent ligand discovery in native biological systems. Nature 534, 570–574 (2016).
Guo, C. et al. Structure-based design of novel human Pin1 inhibitors (III): optimizing affinity beyond the phosphate recognition pocket. Bioorg. Med. Chem. Lett. 24, 4187–4191 (2014).
Yang, N. J. & Hinner, M. J. Getting across the cell membrane: an overview for small molecules, peptides, and proteins. Methods Mol. Biol. 1266, 29–53 (2015).
Singh, J., Petter, R. C., Baillie, T. A. & Whitty, A. The resurgence of covalent drugs. Nat. Rev. Drug Discov. 10, 307–317 (2011).
Valley, C. C. et al. The methionine-aromatic motif plays a unique role in stabilizing protein structure. J. Biol. Chem. 287, 34979–34991 (2012).
Browne, C. M. et al. A chemoproteomic strategy for direct and proteome-wide covalent inhibitor target-site identification. J. Am. Chem. Soc. 141, 191–203 (2019).
Long, M. J., Gollapalli, D. R. & Hedstrom, L. Inhibitor mediated protein degradation. Chem. Biol. 19, 629–637 (2012).
Luo, M. L. et al. Prolyl isomerase Pin1 acts downstream of miR-200 to promote cancer stem-like cell traits in breast cancer. Cancer Res. 74, 3603–3616 (2014).
Rotem, A. et al. Alternative to the soft-agar assay that permits high-throughput drug and genetic screens for cellular transformation. Proc. Natl Acad. Sci. USA 112, 5708–5713 (2015).
Janes, M. R. et al. Targeting KRAS mutant cancers with a covalent G12C-specific inhibitor. Cell 172, 578–589 (2018).
Shi, J. et al. Discovery of cancer drug targets by CRISPR–Cas9 screening of protein domains. Nat. Biotechnol. 33, 661–667 (2015).
Erb, M. A. et al. Transcription control by the ENL YEATS domain in acute leukemia. Nature 543, 270–274 (2017).
Behrsin, C. D. et al. Functionally important residues in the peptidyl-prolyl isomerase Pin1 revealed by unigenic evolution. J. Mol. Biol. 365, 1143–1162 (2007).
An, S. & Liwu, F. Small-molecule PROTACs: an emerging and promising approach for the development of targeted therapy drugs. EBioMedicine 36, 553–562 (2018).
Tzur, A., Kafri, R., LeBleu, V. S., Lahav, G. & Kirschner, M. W. Cell growth and size homeostasis in proliferating animal cells. Science 325, 167–171 (2009).
Crenshaw, D. G., Yang, J., Means, A. R. & Kornbluth, S. The mitotic peptidyl-prolyl isomerase, Pin1, interacts with Cdc25 and Plx1. EMBO J. 17, 1315–1327 (1998).
Ryo, A., Nakamura, M., Wulf, G., Liou, Y. C. & Lu, K. P. Pin1 regulates turnover and subcellular localization of β-catenin by inhibiting its interaction with APC. Nat. Cell Biol. 9, 793–801 (2001).
Ryo, A. et al. Regulation of NF-κB signaling by Pin1-dependent prolyl isomerization and ubiquitin-mediated proteolysis of p65/RelA. Mol. Cell 12, 1413–1426 (2003).
Yeh, E. et al. A signaling pathway controlling c-Myc degradation that impacts oncogenic transformation of human cells. Nat. Cell Biol. 6, 308–318 (2004).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Farrell, A. S. et al. Pin1 regulates the dynamics of c-Myc DNA binding to facilitate target gene regulation and oncogenesis. Mol. Cell. Biol. 33, 2930–2949 (2013).
Vaseva, A. V. et al. KRAS suppression-induced degradation of MYC is antagonized by a MEK5-ERK5 compensatory mechanism. Cancer Cell 34, 807–822 (2018).
Ferguson, F. M. et al. Discovery of a selective inhibitor of doublecortin like kinase 1. Nat. Chem. Biol. https://doi.org/10.1038/s41589-020-0506-0 (2020).
Spena, C. R. et al. Liposomal delivery of a Pin1 inhibitor complexed with cyclodextrins as new therapy for high-grade serious ovarian cancer. J. Control. Release 281, 1–10 (2018).
Chao, S. H., Greenleaf, A. L. & Price, D. H. Juglone, an inhibitor of the peptidyl-prolyl isomerase Pin1, also directly blocks transcription. Nucleic Acids Res. 29, 767–773 (2001).
Auld, D. S. et al. Receptor Binding Assays for HTS and Drug Discovery. Assay Guidance Manual (eds Sittampalam, G. S. et al.) (Eli Lilly & Company and the National Center for Advancing Translational Sciences, 2004).
Yaffe, M. B. et al. Sequence-specific and phosphorylation-dependent proline isomerization: a potential mitotic regulatory mechanism. Science 278, 1957–1960 (1997).
Zhang, Z. & Marshall, A. G. A universal algorithm for fast and automated charge state deconvolution of electrospray mass-to-charge ratio spectra. J. Am. Soc. Mass Spectrom. 9, 225–233 (1998).
Ficarro, S. B., Alexander, W. M. & Marto, J. A. mzStudio: a dynamic digital canvas for user-driven interrogation of mass spectrometry data. Proteomes 5, 20 (2017).
Brinkman, E. K., Chen, T., Amendola, M. & Steensel, B. V. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Cryst. 66, 133–144 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Cryst. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adelmant, G. O. et al. in Sample Preparation in Biological Mass Spectrometry (eds Lazarev, A. V. & Ivanov, A. R.) Ch. 22 (Springer, 2015).
Zhou, F. et al. Genome-scale proteome quantification by DEEP SEQ mass spectrometry. Nat. Commun. 4, 2171 (2013).
Parikh, J. R. et al. Multiplierz: an extensible API based desktop environment for proteomics data analysis. BMC Bioinformatics 10, 364 (2009).
Ficarro, S. B. et al. Leveraging gas-phase fragmentation pathways for improved identification and selective detection of targets modified by covalent probes. Anal. Chem. 88, 12248–12254 (2016).
Wu, X. et al. The Ess1 prolyl isomerase is linked to chromatin remodeling complexes and the general transcription machinery. EMBO J. 19, 3727–3738 (2000).
Acknowledgements
We thank all members of the Gray laboratory for helpful discussions, M. Kostic for providing feedback on the manuscript and K. Westover for discussion of kinact and Ki calculations. This work was supported by the Ruth L. Kirschstein NRSA Individual Predoctoral Fellowship (grant no. F31 CA225066, B.J.P.), the Training Grant in Pharmacological Sciences (NIH grant no. T32 GM007306, B.J.P./Z.M.D), the Training Grant in Chemical Biology (NIH grant no. 5 T32 GM095450-04, B.J.P.), the Chleck Foundation (B.J.P./Z.M.D.), the American Cancer Society Postdoctoral Fellowship (grant no. PF-17-010-01-CDD, B.N.), the Claudia Adams Barr Program in Innovative Basic Cancer Research Award (B.N.), the Katherine L. and Steven C. Pinard Research Fund (N.S.G./B.N.), the Israel Science Foundation (grant no. 2462/19, N.L.), the Rising Tide Foundation (N.L.), the Israel Cancer Research Fund (N.L.), the Israeli Ministry of Science and Technology (grant no. 3-14763, N.L.), the Moross Integrated Cancer Center (N.L.), the Helen and Martin Kimmel Center for Molecular Design (N.L.), the Joel and Mady Dukler Fund for Cancer Research (N.L.), the Estate of Emile Mimran, Virgin JustGiving, the George Schwartzman Fund (N.L.), the National Institutes of Health R01s (grant no. R01 GM056663, S.B.), (grant no. R01 CA219850, grant no. R01 CA233800, J.A.M.), (grant no. R01 CA205153, N.S.G., K.P.L. and X.Z.Z.) and the Hale Center for Pancreatic Research (N.S.G.). BJP-02-118-2 and BJP-07-017-3 cocrystal structures were determined thanks to research conducted at the Advanced Photon Source on the Northeastern Collaborative Access Team beamlines (grant no. NIGMS P41 GM103403).
Author information
Authors and Affiliations
Contributions
B.J.P. performed the chemical synthesis and biological characterization and wrote the manuscript. Z.M.D. generated cell lines and performed the GFP dropout experiment and RNA-sequencing analysis. B.N. generated cell lines and designed and supervised biological studies. C.M.B., S.B.F. and J.A.M. performed CITe-Id and mass spectrometry studies. M.L.M. assisted in Pin1 mutant cellular studies. S. Kozono, S. Kibe and X.L. performed the FP assay. D.Z., D.D. and N.L. provided computational docking support. L.T. advised on compound design and T.D.M. provided support for the chemical synthesis and biological characterization. Z.C.Y., N.E.V., E.A.G. and H.-S.S. performed the protein expression and crystallography; H.-S.S. and S.D.-P. analyzed the crystallography data. Y.C. and S.B. performed the yeast complementation experiments and analysis. K.P.L. and X.Z.Z. helped supervise the project and provided experimental support. N.S.G. conceived of and led this study.
Corresponding authors
Ethics declarations
Competing interests
N.S.G. is a Scientific Founder and member of the Scientific Advisory Board (SAB) of C4 Therapeutics, Syros, Soltego, Gatekeeper and Petra Pharmaceuticals, and has received research funding from Novartis, Astellas, Taiho and Deerfield. J.A.M. serves on the SAB of 908 Devices, Boston, MA, and receives sponsored research support from Vertex and AstraZeneca. N.L. is a member of the SAB of Trilogy Sciences and has received research support from Teva. B.J.P., N.S.G., H.-S.S., S.D.-P., Z.M.D., C.M.B., J.A.M., S. Kozono, X.L., K.P.L. and X.Z.Z. are inventors on a patent application related to the Pin1 inhibitors described in this manuscript (WO/2019/241496). B.N. is an inventor on patent applications related to the dTAG system described in this manuscript (WO/2017/024318, WO/2017/024319, WO/2018/148443, WO/2018/148440). K.P.L. and X.Z.Z. are inventors on many issued patents or pending patent applications related to Pin1 inhibitors and/or biomarkers.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–4, Figs. 1–18 and Note
Supplementary Dataset 1 (download XLSX )
Raw CITe-Id data for BJP-06-005-3.
Supplementary Dataset 2 (download XLSX )
Analyzed RNAseq data for BJP-06-005-3, normalized relative to DMSO and BJP-R.
Rights and permissions
About this article
Cite this article
Pinch, B.J., Doctor, Z.M., Nabet, B. et al. Identification of a potent and selective covalent Pin1 inhibitor. Nat Chem Biol 16, 979–987 (2020). https://doi.org/10.1038/s41589-020-0550-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-020-0550-9
This article is cited by
-
GraphBAN: An inductive graph-based approach for enhanced prediction of compound-protein interactions
Nature Communications (2025)
-
Functionalizing tandem mass tags for streamlining click-based quantitative chemoproteomics
Communications Chemistry (2024)
-
Sulfopin is a covalent inhibitor of Pin1 that blocks Myc-driven tumors in vivo
Nature Chemical Biology (2021)
-
Cysteine-113 covalency inspires the development of Pin1 inhibitor
Signal Transduction and Targeted Therapy (2020)