Abstract
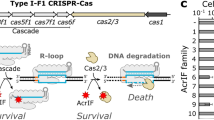
Prokaryotic clustered regularly interspaced short palindromic repeat (CRISPR)–Cas systems are highly vulnerable to phage-encoded anti-CRISPR (Acr) factors. How CRISPR–Cas systems protect themselves remains unclear. Here we uncovered a broad-spectrum anti-anti-CRISPR strategy involving a phage-derived toxic protein. Transcription of this toxin is normally repressed by the CRISPR–Cas effector but is activated to halt cell division when the effector is inhibited by any anti-CRISPR proteins or RNAs. We showed that this abortive infection-like effect efficiently expels Acr elements from bacterial population. Furthermore, we exploited this anti-anti-CRISPR mechanism to develop a screening method for specific Acr candidates for a CRISPR–Cas system and successfully identified two distinct Acr proteins that enhance the binding of CRISPR effector to nontarget DNA. Our data highlight the broad-spectrum role of CRISPR-repressed toxins in counteracting various types of Acr factors. We propose that the regulatory function of CRISPR–Cas confers host cells herd immunity against Acr-encoding genetic invaders whether they are CRISPR targeted or not.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data are included in the article and/or Supplementary Information. The coordinates and structure factors for AcrIF25 were deposited to the PDB under accession number 8X24. The raw data for the RNAseq experiments in Fig. 2 were deposited to the NCBI with the BioProject accession number PRJNA1100312. All strains and plasmids are available from the corresponding author upon request; requests will be answered within 2 weeks. Source data are provided with this paper.
References
Barrangou, R. & Horvath, P. A decade of discovery: CRISPR functions and applications. Nat. Microbiol. 2, 17092 (2017).
Wiedenheft, B., Sternberg, S. H. & Doudna, J. A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 482, 331–338 (2012).
Hille, F. et al. The biology of CRISPR–Cas: backward and forward. Cell 172, 1239–1259 (2018).
Nussenzweig, P. M. & Marraffini, L. A. Molecular mechanisms of CRISPR–Cas immunity in bacteria. Annu. Rev. Genet. 54, 93–120 (2020).
Jackson, S. A. et al. CRISPR–Cas: adapting to change. Science 356, eaal5056 (2017).
Makarova, K. S. et al. Evolutionary classification of CRISPR–Cas systems: a burst of class 2 and derived variants. Nat. Rev. Microbiol. 18, 67–83 (2020).
Sternberg, S. H., Richter, H., Charpentier, E. & Qimron, U. Adaptation in CRISPR–Cas systems. Mol. Cell 61, 797–808 (2016).
Semenova, E. et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc. Natl Acad. Sci. USA 108, 10098–10103 (2011).
Wiedenheft, B. et al. RNA-guided complex from a bacterial immune system enhances target recognition through seed sequence interactions. Proc. Natl Acad. Sci. USA 108, 10092–10097 (2011).
Bondy-Denomy, J. et al. Multiple mechanisms for CRISPR–Cas inhibition by anti-CRISPR proteins. Nature 526, 136–139 (2015).
Borges, A. L., Davidson, A. R. & Bondy-Denomy, J. The discovery, mechanisms, and evolutionary impact of anti-CRISPRs. Annu. Rev. Virol. 4, 37–59 (2017).
Jia, N. & Patel, D. J. Structure-based functional mechanisms and biotechnology applications of anti-CRISPR proteins. Nat. Rev. Mol. Cell Biol. 22, 563–579 (2021).
Yin, P., Zhang, Y., Yang, L. & Feng, Y. Non-canonical inhibition strategies and structural basis of anti-CRISPR proteins targeting type I CRISPR–Cas systems. J. Mol. Biol. 435, 167996 (2023).
Camara-Wilpert, S. et al. Bacteriophages suppress CRISPR–Cas immunity using RNA-based anti-CRISPRs. Nature 623, 601–607 (2023).
Faure, G. et al. CRISPR–Cas in mobile genetic elements: counter-defence and beyond. Nat. Rev. Microbiol. 17, 513–525 (2019).
Li, M. et al. Toxin–antitoxin RNA pairs safeguard CRISPR–Cas systems. Science 372, eabe5601 (2021).
Liu, C. et al. Widespread RNA-based cas regulation monitors crRNA abundance and anti-CRISPR proteins. Cell Host Microbe 31, 1481–1493 (2023).
Shmakov, S. A. et al. Widespread CRISPR-derived RNA regulatory elements in CRISPR–Cas systems. Nucleic Acids Res. 51, 8150–8168 (2023).
Workman, R. et al. A natural single-guide RNA repurposes Cas9 to autoregulate CRISPR–Cas expression. Cell 184, 675–688 (2021).
Ratner, H. K. et al. Catalytically active Cas9 mediates transcriptional interference to facilitate bacterial virulence. Mol. Cell 75, 498–510 (2019).
Cheng, F. et al. Divergent degeneration of creA antitoxin genes from minimal CRISPRs and the convergent strategy of tRNA-sequestering CreT toxins. Nucleic Acids Res. 49, 10677–10688 (2021).
Wang, R. et al. Associate toxin–antitoxin with CRISPR–Cas to kill multidrug-resistant pathogens. Nat. Commun. 14, 2078 (2023).
Richter, C. et al. Priming in the type IF CRISPR–Cas system triggers strand-independent spacer acquisition, bi-directionally from the primed protospacer. Nucleic Acids Res. 42, 8516–8526 (2014).
Iyer, L. M., Koonin, E. V. & Aravind, L. Extensive domain shuffling in transcription regulators of DNA viruses and implications for the origin of fungal APSES transcription factors. Genome Biol. 3, 1–11 (2002).
Hansen, E. B. Structure and regulation of the lytic replicon of phage P1. J. Mol. Biol. 207, 135–149 (1989).
Das, A., Biswas, S. & Biswas, M. Expression of Phi11 Gp07 causes filamentation in Escherichia coli. Open Microbiol. J. 12, 107 (2018).
Cech, G. M., Szalewska-Pałasz, A., Potrykus, K. & Kloska, A. Virus–host interaction gets curiouser and curiouser. Part II: functional transcriptomics of the E. coli DksA-deficient cell upon phage P1 vir infection. Int. J. Mol. Sci. 22, 6159 (2021).
de Boer, P. A. J. Advances in understanding E. coli cell fission. Curr. Opin. Microbiol. 13, 730–737 (2010).
Richter, C., Gristwood, T., Clulow, J. S. & Fineran, P. C. In vivo protein interactions and complex formation in the Pectobacterium atrosepticum subtype IF CRISPR/Cas system. PLoS ONE 7, e49549 (2012).
Cheng, F. et al. The toxin–antitoxin RNA guards of CRISPR–Cas evolved high specificity through repeat degeneration. Nucleic Acids Res. 50, 9442–9452 (2022).
Xie, Y. et al. AcrIF5 specifically targets DNA-bound CRISPR–Cas surveillance complex for inhibition. Nat. Chem. Biol. 18, 670–677 (2022).
Gao, Z. et al. Anti-CRISPR protein AcrIF4 inhibits the type IF CRISPR–Cas surveillance complex by blocking nuclease recruitment and DNA cleavage. J. Biol. Chem. 298, 102575 (2022).
Yang, L. et al. Insights into the inhibition of type IF CRISPR–Cas system by a multifunctional anti-CRISPR protein AcrIF24. Nat. Commun. 13, 1931 (2022).
Mukherjee, I. A., Gabel, C., Noinaj, N., Bondy-Denomy, J. & Chang, L. Structural basis of AcrIF24 as an anti-CRISPR protein and transcriptional suppressor. Nat. Chem. Biol. 18, 1417–1424 (2022).
Wu, S. et al. A pyrF-based efficient genetic manipulation platform in Acinetobacter baumannii to explore the vital DNA components of adaptive immunity for IF CRISPR–Cas. Microbiol. Spectr. 10, e01957-22 (2022).
Jia, J. et al. Post-transcriptional regulation is involved in the cold-active methanol-based methanogenic pathway of a psychrophilic methanogen. Environ. Microbiol. 23, 3773–3788 (2021).
Acknowledgements
This work was supported by the Science and Technology Fundamental Resources Investigation Program (2022FY101100 to H.Z.), the National Natural Science Foundation of China (32150020 to M.L., 82225028 to S.O., 32270092 to R.W., 32022003 to M.L., 32370090 to H.Z., 32200057 to F.C. and 82172287 to S.O.), the National Key Research and Development Program of China (2021YFC2301403 to S.O.), the Youth Innovation Promotion Association of the Chinese Academy of Sciences (2020090 to M.L.), the China National Postdoctoral Program for Innovative Talents (BX20220331 to F.C.), the China Postdoctoral Science Foundation (2022M720160 to F.C.) and the Special Research Assistant Program of the Chinese Academy of Sciences (2023000056 to F.C.).
Author information
Authors and Affiliations
Contributions
M.L. and R.W. conceptualized this study. M.L., R.W., X.S., J. Li and S.O. designed the experiments, with valuable suggestions from C.H. X.S., R.W. and Q.X. constructed the plasmids and mutant strains with the assistance of F.C., C.L. and H.Z. X.S. and R.W. conducted the dilution plating assay, SEM investigation, fluorescence measurement and bacteria transformation assays. X.S., R.W. and H.Z. performed RNAseq and primer extension. R.W., X.S. and Q.X. performed the bioinformatic analyses of CrePA and Acrs. Z.L. and X.S. conducted protein purification, western blot, SEC, TAP and EMSAs with the assistance of J. Li and J. Liu. X.S. conducted the DNA competition experiments and cell population assays. J.W. conducted crystallization and X-ray diffraction data collection of AcrIF25. Formal analysis of the results was performed by X.S., R.W., Z.L., Q.X., J. Li and S.O. M.L., J. Li, S.O. and R.W. analyzed the data and supervised the project. M.L. wrote the paper, which was edited and approved by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Jennifer Doudna, Yue Feng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1 and Figs. 1–10.
Supplementary Data 1 (download XLSX )
A. baylyi ADP1 crePA and its homologs.
Supplementary Data 2 (download XLSX )
Strains, chemicals, plasmids, oligonucleotides and synthetic genes.
Supplementary Data 3 (download XLSX )
Source data for supplementary figures.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shu, X., Wang, R., Li, Z. et al. CRISPR-repressed toxin–antitoxin provides herd immunity against anti-CRISPR elements. Nat Chem Biol 21, 337–347 (2025). https://doi.org/10.1038/s41589-024-01693-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-024-01693-3
This article is cited by
-
Phage-associated Cas12p nucleases require binding to bacterial thioredoxin for activation and cleavage of target DNA
Nature Microbiology (2026)
-
A broad anti-anti-CRISPR strategy
Nature Chemical Biology (2025)
-
Inducible transposon mutagenesis identifies bacterial fitness determinants during infection in mice
Nature Microbiology (2025)