Abstract
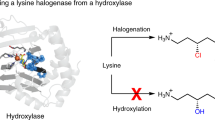
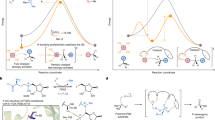
The activation of inert C(sp3)–H bonds by nonheme Fe enzymes provides a powerful biocatalytic platform for the chemical synthesis of molecules with increased sp3 complexity. In this context, FeII/α-ketoglutarate-dependent radical halogenases are uniquely capable of carrying out transfer of a diverse array of bound anions following C–H activation. Here, we provide experimental evidence that bifurcation of radical rebound after H-atom abstraction can be driven both by the ability of a dynamic metal coordination sphere to reorganize and by a second-sphere hydrogen-bonding network where only two residues are sufficient. In addition, we present crystallographic data supporting the existence of an early peroxyhemiketal intermediate in the O2 activation pathway of FeII/α-ketoglutarate-dependent enzymes. These data provide a paradigm for understanding the evolution of catalytic plasticity in these enzymes and yields insight into the design principles by which to expand their reaction scope.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper and its Supplementary Information. Should any raw data files be needed in another format, they are available from the corresponding author upon request. Crystallographic data for structures were deposited to the Protein Data Bank (PDB) under accession numbers 9OER, 9OEU, 9OES, 9OET, 9OEW and 9OEV. Source data are provided with this paper.
Code availability
The code used to produce the TD-DFT fits of the XAS data is provided as Supplementary Data 2.
References
Islam, M.dS., Leissing, T. M., Chowdhury, R., Hopkinson, R. J. & Schofield, C. J. 2-Oxoglutarate-dependent oxygenases. Annu. Rev. Biochem. 87, 585–620 (2018).
Zwick, C. R. I. & Renata, H. Overview of amino acid modifications by iron- and α-ketoglutarate-dependent enzymes. ACS Catal. 13, 4853–4865 (2023).
Gao, S.-S., Naowarojna, N., Cheng, R., Liu, X. & Liu, P. Recent examples of α-ketoglutarate-dependent mononuclear non-haem iron enzymes in natural product biosyntheses. Nat. Prod. Rep. 35, 792–837 (2018).
Ushimaru, R. & Abe, I. Unusual dioxygen-dependent reactions catalyzed by nonheme iron enzymes in natural product biosynthesis. ACS Catal. 13, 1045–1076 (2023).
Chen, K. & Arnold, F. H. Engineering new catalytic activities in enzymes. Nat. Catal. 3, 203–213 (2020).
Goldberg, N. W., Knight, A. M., Zhang, R. K. & Arnold, F. H. Nitrene transfer catalyzed by a non-heme iron enzyme and enhanced by non-native small-molecule ligands. J. Am. Chem. Soc. 141, 19585–19588 (2019).
Zhang, X. et al. Divergent synthesis of complex diterpenes through a hybrid oxidative approach. Science 369, 799–806 (2020).
Amatuni, A., Shuster, A., Adibekian, A. & Renata, H. Concise chemoenzymatic total synthesis and identification of cellular targets of cepafungin I. Cell Chem. Biol. 27, 1318–1326(2020).
Bollinger Jr., J. M. et al. Mechanisms of 2-oxoglutarate-dependent oxygenases: the hydroxylation paradigm and beyond. In 2-Oxoglutarate-Dependent Oxygenases (eds Schofield, C. J. & Hausinger, R. P.) (The Royal Society of Chemistry, 2015).
Martinez, S. & Hausinger, R. P. Catalytic mechanisms of Fe(II)- and 2-oxoglutarate-dependent oxygenases. J. Biol. Chem. 290, 20702–20711 (2015).
Solomon, E. I., DeWeese, D. E. & Babicz, J. T. Jr. Mechanisms of O2 activation by mononuclear non-heme iron enzymes. Biochemistry 60, 3497–3506 (2021).
Vaillancourt, F. H., Yin, J. & Walsh, C. T. SyrB2 in syringomycin E biosynthesis is a nonheme FeII α-ketoglutarate- and O2-dependent halogenase. Proc. Natl Acad. Sci. USA 102, 10111–10116 (2005).
Blasiak, L. C., Vaillancourt, F. H., Walsh, C. T. & Drennan, C. L. Crystal structure of the non-haem iron halogenase SyrB2 in syringomycin biosynthesis. Nature 440, 368–371 (2006).
Agarwal, V. et al. Enzymatic halogenation and dehalogenation reactions: pervasive and mechanistically diverse. Chem. Rev. 117, 5619–5674 (2017).
Matthews, M. L. et al. Direct nitration and azidation of aliphatic carbons by an iron-dependent halogenase. Nat. Chem. Biol. 10, 209–215 (2014).
Neugebauer, M. E. et al. A family of radical halogenases for the engineering of amino-acid-based products. Nat. Chem. Biol. 18, 1–8 (2019).
Chan, N. H. et al. Non-native anionic ligand binding and reactivity in engineered variants of the Fe(II)- and α-ketoglutarate-dependent oxygenase SadA. Inorg. Chem. 61, 14477–14485 (2022).
Matthews, M. L. et al. Substrate positioning controls the partition between halogenation and hydroxylation in the aliphatic halogenase, SyrB2. Proc. Natl Acad. Sci. USA 106, 17723–17728 (2009).
Mitchell, A. J. et al. Structural basis for halogenation by iron- and 2-oxo-glutarate-dependent enzyme WelO5. Nat. Chem. Biol. 12, 636–640 (2016).
Papadopoulou, A., Meyer, F. & Buller, R. M. Engineering Fe(II)/α-ketoglutarate-dependent halogenases and desaturases. Biochemistry 62, 229–240 (2023).
Gomez, C. A., Mondal, D., Du, Q., Chan, N. & Lewis, J. C. Directed evolution of an iron(II)- and α-ketoglutarate-dependent dioxygenase for site-selective azidation of unactivated aliphatic C–H bonds. Angew. Chem. Int. Ed. 62, e202301370 (2023).
Duewel, S. et al. Directed evolution of an FeII-dependent halogenase for asymmetric C(sp3)–H chlorination. ACS Catal. 10, 1272–1277 (2020).
Hayashi, T. et al. Evolved aliphatic halogenases enable regiocomplementary C–H functionalization of a pharmaceutically relevant compound. Angew. Chem. Int. Ed. Engl. 58, 18535–18539 (2019).
Büchler, J. et al. Algorithm-aided engineering of aliphatic halogenase WelO5* for the asymmetric late-stage functionalization of soraphens. Nat. Commun. 13, 371 (2022).
Martinie, R. J. et al. Experimental correlation of substrate position with reaction outcome in the aliphatic halogenase, SyrB2. J. Am. Chem. Soc. 137, 6912–6919 (2015).
Wong, S. D. et al. Elucidation of the Fe(IV)=O intermediate in the catalytic cycle of the halogenase SyrB2. Nature 499, 320–323 (2013).
Srnec, M. & Solomon, E. I. Frontier molecular orbital Contributions to chlorination versus hydroxylation selectivity in the non-heme iron halogenase SyrB2. J. Am. Chem. Soc. 139, 2396–2407 (2017).
Srnec, M. et al. Electronic structure of the ferryl intermediate in the α-ketoglutarate dependent non-heme iron halogenase SyrB2: contributions to H atom abstraction reactivity. J. Am. Chem. Soc. 138, 5110–5122 (2016).
Costas, M., Mehn, M. P., Jensen, M. P. & Que, L. Dioxygen activation at mononuclear nonheme iron active sites: enzymes, models, and intermediates. Chem. Rev. 104, 939–986 (2004).
Martinie, R. J. et al. Vanadyl as a stable structural mimic of reactive ferryl intermediates in mononuclear nonheme-iron enzymes. Inorg. Chem. 56, 13382–13389 (2017).
Borowski, T., Noack, H., Radoń, M., Zych, K. & Siegbahn, P. E. M. Mechanism of selective halogenation by SyrB2: a computational study. J. Am. Chem. Soc. 132, 12887–12898 (2010).
Huang, J. et al. Selective chlorination of substrates by the halogenase SyrB2 is controlled by the protein according to a combined quantum mechanics/molecular mechanics and molecular dynamics study. ACS Catal. 6, 2694–2704 (2016).
Zhang, J. et al. Conformational isomerization of the Fe(III)–OH species enables selective halogenation in carrier-protein-independent halogenase BesD and hydroxylase-evolved halogenase. ACS Catal. 14, 9342–9353 (2024).
Vennelakanti, V., Mehmood, R. & Kulik, H. J. Are vanadium intermediates suitable mimics in non-heme iron enzymes? An electronic structure analysis. ACS Catal. 12, 5489–5501 (2022).
Brown, C. A. et al. Spectroscopic and theoretical description of the electronic structure of S = 3/2 iron-nitrosyl complexes and their relation to O2 activation by non-heme iron enzyme active sites. J. Am. Chem. Soc. 117, 715–732 (1995).
Yoshizawa, K., Kamachi, T. & Shiota, Y. A theoretical study of the dynamic behavior of alkane hydroxylation by a compound I model of cytochrome P450. J. Am. Chem. Soc. 123, 9806–9816 (2001).
Bathelt, C. M., Ridder, L., Mulholland, A. J. & Harvey, J. N. Mechanism and structure–reactivity relationships for aromatic hydroxylation by cytochrome P450. Org. Biomol. Chem. 2, 2998–3005 (2004).
Mitchell, A. J. et al. Visualizing the reaction cycle in an iron(II)- and 2-(oxo)-glutarate-dependent hydroxylase. J. Am. Chem. Soc. 139, 13830–13836 (2017).
Kovaleva, E. G. & Lipscomb, J. D. Crystal structures of Fe2+ dioxygenase superoxo, alkylperoxo, and bound product intermediates. Science 316, 453–457 (2007).
Iyer, S. R. et al. Direct coordination of pterin to FeII enables neurotransmitter biosynthesis in the pterin-dependent hydroxylases. Proc. Natl Acad. Sci. USA 118, e2022379118 (2021).
Zhang, Z. et al. Crystal structure of a clavaminate synthase–Fe(II)–2-oxoglutarate–substrate–NO complex: evidence for metal centered rearrangements. FEBS Lett. 517, 7–12 (2002).
Chekan, J. R. et al. Molecular basis for enantioselective herbicide degradation imparted by aryloxyalkanoate dioxygenases in transgenic plants. Proc. Natl Acad. Sci. USA 116, 13299–13304 (2019).
Vennelakanti, V., Jeon, M. & Kulik, H. J. How do differences in electronic structure affect the use of vanadium intermediates as mimics in nonheme iron hydroxylases? Inorg. Chem. 63, 4997–5011 (2024).
King, A. E. et al. A well-defined terminal vanadium(III) oxo complex. Inorg. Chem. 53, 11388–11395 (2014).
Slater, J. W. et al. Synergistic binding of the halide and cationic prime substrate of l-lysine 4-chlorinase, BesD, in both ferrous and ferryl states. Biochemistry 62, 2480–2491 (2023).
Neugebauer, M. E. et al. Reaction pathway engineering converts a radical hydroxylase into a halogenase. Nat. Chem. Biol. 18, 171–179 (2022).
Levina, A., McLeod, A. I. & Lay, P. A. Vanadium speciation by XANES spectroscopy: a three-dimensional approach. Chem. Eur. J. 20, 12056–12060 (2014).
Rees, J. A. et al. Experimental and theoretical correlations between vanadium K-edge X-ray absorption and Kβ emission spectra. J. Biol. Inorg. Chem. 21, 793–805 (2016).
Ehudin, M. A. et al. Tuning the geometric and electronic structure of synthetic high-valent heme iron(IV)–oxo models in the presence of a Lewis acid and various axial ligands. J. Am. Chem. Soc. 141, 5942–5960 (2019).
Price, J. C., Barr, E. W., Glass, T. E., Krebs, C. & Bollinger, J. M. Evidence for hydrogen abstraction from C1 of taurine by the high-spin Fe(IV) intermediate detected during oxygen activation by taurine:α-ketoglutarate dioxygenase (TauD). J. Am. Chem. Soc. 125, 13008–13009 (2003).
Matthews, M. L. et al. Substrate-triggered formation and remarkable stability of the C−H bond-cleaving chloroferryl intermediate in the aliphatic halogenase, SyrB2. Biochemistry 48, 4331–4343 (2009).
Smithwick, E. R. et al. Electrostatically regulated active site assembly governs reactivity in nonheme iron halogenases. ACS Catal. 13, 13743–13755 (2023).
Kissman, E. N. et al. Biocatalytic control of site-selectivity and chain length-selectivity in radical amino acid halogenases. Proc. Natl Acad. Sci. USA 120, e2214512120 (2023).
Arndtsen, B. A., Bergman, R. G., Mobley, T. A. & Peterson, T. H. Selective intermolecular carbon–hydrogen bond activation by synthetic metal complexes in homogeneous solution. Acc. Chem. Res. 28, 154–162 (1995).
Colby, D. A., Bergman, R. G. & Ellman, J. A. Rhodium-catalyzed C–C bond formation via heteroatom-directed C–H bond activation. Chem. Rev. 110, 624–655 (2010).
Hartwig, J. F. & Larsen, M. A. Undirected, homogeneous C–H bond functionalization: challenges and opportunities. ACS Cent. Sci. 2, 281–292 (2016).
Hartwig, J. F. Evolution of C–H bond functionalization from methane to methodology. J. Am. Chem. Soc. 138, 2–24 (2016).
Saint-Denis, T. G., Zhu, R.-Y., Chen, G., Wu, Q.-F. & Yu, J.-Q. Enantioselective C(sp3)‒H bond activation by chiral transition metal catalysts. Science 359, eaao4798 (2018).
Gérard, E. F., Yadav, V., Goldberg, D. P. & de Visser, S. P. What drives radical halogenation versus hydroxylation in mononuclear nonheme iron complexes? A combined experimental and computational study. J. Am. Chem. Soc. 144, 10752–10767 (2022).
Feig, A. L. & Lippard, S. J. Reactions of non-heme iron(II) centers with dioxygen in biology and chemistry. Chem. Rev. 94, 759–805 (1994).
MacBeth, C. E. et al. O2 activation by nonheme iron complexes: a monomeric Fe(III)–oxo complex derived from O2. Science 289, 938–941 (2000).
Kovaleva, E. G. & Lipscomb, J. D. Versatility of biological non-heme Fe(II) centers in oxygen activation reactions. Nat. Chem. Biol. 4, 186–193 (2008).
Sahu, S. & Goldberg, D. P. Activation of dioxygen by iron and manganese complexes: a heme and nonheme perspective. J. Am. Chem. Soc. 138, 11410–11428 (2016).
Kastner, D. W., Nandy, A., Mehmood, R. & Kulik, H. J. Mechanistic insights into substrate positioning that distinguish non-heme Fe(II)/α-ketoglutarate-dependent halogenases and hydroxylases. ACS Catal. 13, 2489–2501 (2023).
Wenger, E. S. et al. Optimized substrate positioning enables switches in the C–H cleavage site and reaction outcome in the hydroxylation–epoxidation sequence catalyzed by hyoscyamine 6β-hydroxylase. J. Am. Chem. Soc. 146, 24271–24287 (2024).
Borowski, T., Bassan, A. & Siegbahn, P. E. M. Mechanism of dioxygen activation in 2-oxoglutarate-dependent enzymes: a hybrid DFT study. Chem. Eur. J. 10, 1031–1041 (2004).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Ravel, B. & Newville, M. ATHENA, ARTEMIS, HEPHAESTUS: data analysis for X-ray absorption spectroscopy using IFEFFIT. J. Synchr. Radiat. 12, 537–541 (2005).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Luo, L. et al. An assay for Fe(II)/2-oxoglutarate-dependent dioxygenases by enzyme-coupled detection of succinate formation. Anal. Biochem. 353, 69–74 (2006).
Hoops, S. et al. COPASI—a complex pathway simulator. Bioinformatics 22, 3067–3074 (2006).
Stoll, S. & Schweiger, A. EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J. Mag. Res. 178, 42–55 (2006).
Becke, A. D. Density-functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
Stephens, P. J., Devlin, F. J., Chabalowski, C. F. & Frisch, M. J. Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. J. Phys. Chem. 98, 11623–11627 (1994).
Grimme, S., Antony, J., Ehrlich, S. & Krieg, H. A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H–Pu. J. Chem. Phys. 132, 154104 (2010).
Weigend, F. & Ahlrichs, R. Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: design and assessment of accuracy. Phys. Chem. Chem. Phys. 7, 3297–3305 (2005).
Neese, F. The ORCA program system. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2, 73–78 (2012).
Neese, F. Prediction and interpretation of the 57Fe isomer shift in Mössbauer spectra by density functional theory. Inorg. Chim. Acta 337, 181–192 (2002).
Weigend, F. Accurate Coulomb-fitting basis sets for H to Rn. Phys. Chem. Chem. Phys. 8, 1057–1065 (2006).
Römelt, M., Ye, S. & Neese, F. Calibration of modern density functional theory methods for the prediction of 57Fe Mössbauer isomer shifts: meta-GGA and double-hybrid functionals. Inorg. Chem. 48, 784–785 (2009).
Roemelt, M. et al. Manganese K-edge X-ray absorption spectroscopy as a probe of the metal–ligand interactions in coordination compounds. Inorg. Chem. 51, 680–687 (2012).
Acknowledgements
This work was funded by generous support from the National Institutes of Health (NIH; R01 GM134271 to M.C.Y.C. and R01 GM138580 to J.M.B.) and by DOE/LBL DEAC02-05CH11231 FWP CH030201. E.N.K. acknowledges the support of an NIH National Research Service Award training grant (T32 GM066698). I.K. is supported by the Miller Institute for Basic Research in Science (University of California, Berkeley). E.A.S. acknowledges support from the Jane Coffin Childs Fund for Medical Research. J.W.S. acknowledges support of the National Institute of General Medical Sciences of the NIH (F32 GM136156). J.Y. acknowledges support of the NIH (R01 GM110501) for the XAS data collection. We thank E. I. Solomon and C. Krebs for helpful discussions and access to their EPR and Mössbauer spectrometers, respectively. We also thank P. Jeffrey for his helpful feedback regarding the analysis of crystallography data. We acknowledge L. Nocka and the J. Kuriyan laboratory for assistance using their size-exclusion chromatography multiangle light scattering instrument. X-ray diffraction data were collected at the Advanced Light Source beamline 8.3.1, which is operated by the University of California Office of the President, Multicampus Research Programs and Initiatives (MR-15-328599), the NIH (R01 GM124149 and P30 GM124169), Plexxikon and the Integrated Diffraction Analysis Technologies program of the US Department of Energy Office of Biological and Environmental Research. The Advanced Light Source is a national user facility operated by Lawrence Berkeley National Laboratory on behalf of the US Department of Energy under contract number DEAC02-05CH11231. XAS data were collected at SSRL beamline 9-3. Use of the SSRL, Stanford Linear Accelerator Center National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under contract no. DEAC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the Department of Energy Office of Biological and Environmental Research and by the NIH National Institute of General Medical Sciences (P30 GM133894).
Author information
Authors and Affiliations
Contributions
E.N.K. designed the study, performed the enzyme characterization experiments, led the protein crystallography studies and analyzed the SF-Abs kinetics. I.K. designed the study, performed the DFT calculations and contributed to the analysis of spectroscopic data. J.W.S. assisted with the enzyme kinetics, EPR and Mössbauer data collection and analysis. E.A.S. assisted with the protein crystallography. A.Y.Y. and A.R.E. assisted with the enzyme characterization and protein crystallography studies. A.B. assisted with the spectroscopic data analysis. A.M.W. assisted with the protein crystallography. K.C., I.B. and J.Y. performed the XAS experiments. J.M.B. assisted with the design and analysis of SF-Abs, EPR and Mössbauer experiments. M.C.Y.C. designed the study, assisted with the data analysis and managed the project. E.N.K., I.K. and M.C.Y.C. wrote the paper with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Jared Lewis, Francesco Stellato, Binju Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Crystal structure of HalA with the peroxyhemiketal intermediate bound.
Analysis of the assignment of the electron density found in the in crystallo reaction of anaerobic HalA with O2 as the peroxyhemiketal intermediate. See Supplementary Table 15 for additional information.
Extended Data Fig. 3 Crystal structure of Hydrox with VIV-oxo bound.
A crystal structure of Hydrox substituted with VIV-oxo was also solved to compare to HalA. See Supplementary Table 15 for additional information.
Extended Data Fig. 4 EPR spectroscopy of HalA and Hydrox with VIV-oxo.
Details of the analysis of VIV-oxo speciation in HalA and Hydrox based on EPR spectroscopy. See Supplementary Table 15 for additional information.
Extended Data Fig. 5 DFT-optimized structures and relative energies for VIV-oxo and FeIV-oxo isomers of HalA.
DFT was used to analyze the energetics of various configurational isomers of VIV-oxo and FeIV-oxo intermediates for HalA with varying coordination numbers (5-coordinate with succinate monodentate vs 6-coordinate with succinate bidentate), oxo/Cl isomerism (in-line vs off-line), and axial metal ligands (that is trans to the oxo ligand; succinate vs histidine). See Supplementary Table 15 for additional information.
Extended Data Fig. 6 Mössbauer spectroscopy of HalA.
Mössbauer spectroscopy of HalA shows evidence of only one FeIV-oxo species, however, the in-line and off-line isomers of FeIV-oxo HalA are pseudo-enantiomers with indistinguishable Mössbauer parameters. See Supplementary Table 15 for additional information.
Extended Data Fig. 7 SF-Abs kinetics of wild-type HalA and proposed model.
Pre-steady state kinetic analysis of HalA was carried out to measure the rate of H-atom abstraction by rapid mixing of anaerobic HalA samples with O2 and measurement of the rate of decay of the signature FeIV-oxo peak. See Supplementary Table 15 for additional information.
Extended Data Fig. 8 Thr226 tunes the active site H-bonding network.
Analysis of the role of Thr226 in altering the partitioning between halogenation and hydroxylation. Although no effect is observed in the T226 single mutants, mutation in a HalA N224V background does show an effect on this ratio. See Supplementary Table 15 for additional information.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Supplementary Figs. 1–9, Supplementary Tables 1–15 and Supplementary References.
Supplementary Data 1 (download XLSX )
Analytical size-exclusion chromatography data for Supplementary Fig. 1.
Supplementary Data 2 (download XLSX )
XAS and TD-DFT data for Supplementary Fig. 4.
Supplementary Data 3 (download XLSX )
Chloride-dependent kinetics data for Supplementary Fig. 6.
Supplementary Data 4 (download XLSX )
Steady-state kinetics of HalA variants for Supplementary Fig. 7.
Supplementary Data 5 (download XLSX )
Stopped-flow kinetics of HalA variants for Supplementary Fig. 8.
Supplementary Data 6 (download TXT )
DFT model coordinates.
Supplementary Code 1 (download TXT )
Code to generate TD-DFT fit.
Source data
Source Data Fig. 3 (download XLSX )
XAS and EPR data.
Source Data Extended Data Fig. 6 (download XLSX )
Mössbauer data.
Source Data Extended Data Fig. 7 (download XLSX )
Stopped-flow kinetics data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kissman, E.N., Kipouros, I., Slater, J.W. et al. Dynamic metal coordination controls chemoselectivity in a radical halogenase. Nat Chem Biol 22, 491–500 (2026). https://doi.org/10.1038/s41589-025-02077-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-025-02077-x