Abstract
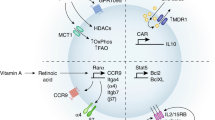
Recent studies have highlighted the presence of memory T cells in human brains, some of which are specific for peripheral infections. To address their potential origins, we used two models of polymicrobial exposure to ‘normalize’ the immune systems of specific pathogen-free mice and queried the impact on brain T cell biology. Here, we show that cohousing and sequential infection induce marked enhancement of memory T cells in the brain tissue of mice. These resident and circulating memory T cells localized to diverse brain regions where dynamic interactions with myeloid cells occurred. Following an induced seizure, brain-localized memory T cells were functionally altered in microbe-experienced mice. Microbial exposure also induced T cell-dependent changes in seizure duration. These data not only suggest a potential origin for memory T cells in human brains but also reveal the ability of these cells to modulate brain biology, prompting the future utilization of microbe-experienced mice in studies of neurological health and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data associated with this study can be found in the main text or the Supplementary Information. The raw flow cytometric data, tissue imaging files or seizure video recordings that support the data in this manuscript are available from the corresponding author upon request. Source data are provided with this paper.
References
Svenningsson, A., Andersen, O., Edsbagge, M. & Stemme, S. Lymphocyte phenotype and subset distribution in normal cerebrospinal fluid. J. Neuroimmunol. 63, 39–46 (1995).
Smolders, J. et al. Tissue-resident memory T cells populate the human brain. Nat. Commun. 9, 4593 (2018).
Pappalardo, J. L. et al. Transcriptomic and clonal characterization of T cells in the human central nervous system. Sci. Immunol. 5, eabb8786 (2020).
Piehl, N. et al. Cerebrospinal fluid immune dysregulation during healthy brain aging and cognitive impairment. Cell 185, 5028–5039 (2022).
Smolders, J. et al. Characteristics of differentiated CD8+ and CD4+ T cells present in the human brain. Acta Neuropathol. 126, 525–535 (2013).
Mix, M. R. & Harty, J. T. Keeping T cell memories in mind. Trends Immunol. 43, 1018–1031 (2022).
Ayasoufi, K. et al. Brain resident memory T cells rapidly expand and initiate neuroinflammatory responses following CNS viral infection. Brain Behav. Immun. 112, 51–76 (2023).
Pasciuto, E. et al. Microglia require CD4 T cells to complete the fetal-to-adult transition. Cell 182, 625–640 (2020).
Wakim, L. M., Woodward-Davis, A. & Bevan, M. J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. Proc. Natl Acad. Sci. USA 107, 17872–17879 (2010).
Wakim, L. M. et al. The molecular signature of tissue resident memory CD8 T cells isolated from the brain. J. Immunol. 189, 3462–3471 (2012).
Landrith, T. A. et al. CD103+ CD8 T cells in the Toxoplasma-infected brain exhibit a tissue-resident memory transcriptional profile. Front. Immunol. 8, 335 (2017).
Garber, C. et al. T cells promote microglia-mediated synaptic elimination and cognitive dysfunction during recovery from neuropathogenic flaviviruses. Nat. Neurosci. 22, 1276–1288 (2019).
Vincenti, I. et al. Tissue-resident memory CD8+ T cells cooperate with CD4+ T cells to drive compartmentalized immunopathology in the CNS. Sci. Transl. Med. 14, eabl6058 (2022).
Fransen, N. L. et al. Tissue-resident memory T cells invade the brain parenchyma in multiple sclerosis white matter lesions. Brain 143, 1714–1730 (2020).
Bernard-Valnet, R. et al. Influenza vaccination induces autoimmunity against orexinergic neurons in a mouse model for narcolepsy. Brain 145, 2018–2030 (2022).
Urban, S. L. et al. Peripherally induced brain tissue-resident memory CD8+ T cells mediate protection against CNS infection. Nat. Immunol. 21, 938–949 (2020).
Dulken, B. W. et al. Single-cell analysis reveals T cell infiltration in old neurogenic niches. Nature 571, 205–210 (2019).
Cassidy, B. R., Sonntag, W. E., Leenen, P. J. M. & Drevets, D. A. Systemic Listeria monocytogenes infection in aged mice induces long-term neuroinflammation: the role of miR-155. Immun. Ageing 19, 25 (2022).
Wang, X. et al. CD8+ T cells exacerbate AD-like symptoms in mouse model of amyloidosis. Brain Behav. Immun. 122, 444–455 (2024).
Masopust, D. & Soerens, A. G. Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37, 521–546 (2019).
Martin, M. D. & Badovinac, V. P. Defining memory CD8 T cell. Front. Immunol. 9, 2692 (2018).
Frieser, D. et al. Tissue-resident CD8+ T cells drive compartmentalized and chronic autoimmune damage against CNS neurons. Sci. Transl. Med. 14, eabl6157 (2022).
Chen, X. et al. Microglia-mediated T cell infiltration drives neurodegeneration in tauopathy. Nature 615, 668–677 (2023).
Ning, J. et al. Functional virus-specific memory T cells survey glioblastoma. Cancer Immunol. Immunother. 71, 1863–1875 (2022).
Ghazanfari, N. et al. CD8+ and CD4+ T cells infiltrate into the brain during Plasmodium berghei ANKA infection and form long-term resident memory. J. Immunol. 207, 1578–1590 (2021).
Casey, K. A. et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 188, 4866–4875 (2012).
Mix, M. R. et al. Repetitive antigen stimulation in the periphery dictates the composition and recall responses of brain-resident memory CD8+ T cells. Cell Rep. 44, 115247 (2025).
Gate, D. et al. Clonally expanded CD8 T cells patrol the cerebrospinal fluid in Alzheimer’s disease. Nature 577, 399–404 (2020).
Chang, J. W. et al. Clonally focused public and private T cells in resected brain tissue from surgeries to treat children with intractable seizures. Front. Immunol. 12, 664344 (2021).
Hamilton, S. E. et al. New insights into the immune system using dirty mice. J. Immunol. 205, 3–11 (2020).
Rehermann, B., Graham, A.L., Masopust, D. & Hamilton, S.E. Integrating natural commensals and pathogens into preclinical mouse models. Nat. Rev. Immunol. 25, 385–397 (2024).
Bruno, P., Schüler, T. & Rosshart, S.P. Born to be wild: utilizing natural microbiota for reliable biomedical research. Trends Immunol. 46, 17–28 (2025).
Beura, L. K. et al. Normalizing the environment recapitulates adult human immune traits in laboratory mice. Nature 532, 512–516 (2016).
Reese, T. A. et al. Sequential infection with common pathogens promotes human-like immune gene expression and altered vaccine response. Cell Host Microbe 19, 713–719 (2016).
Rosshart, S. P. et al. Laboratory mice born to wild mice have natural microbiota and model human immune responses. Science 365, eaaw4361 (2019).
Huggins, M. A. et al. Microbial exposure enhances immunity to pathogens recognized by TLR2 but increases susceptibility to cytokine storm through TLR4 sensitization. Cell Rep. 28, 1729–1743 (2019).
Fiege, J. K. et al. Mice with diverse microbial exposure histories as a model for preclinical vaccine testing. Cell Host Microbe 29, 1815–1827 (2021).
Block, K. E. et al. Physiological microbial exposure transiently inhibits mouse lung ILC2 responses to allergens. Nat. Immunol. 23, 1703–1713 (2022).
Berton, R. R., Jensen, I. J., Harty, J. T., Griffith, T. S. & Badovinac, V. P. Inflammation controls susceptibility of immune-experienced mice to sepsis. Immunohorizons 6, 528–542 (2022).
Sjaastad, F. V. et al. Reduced T cell priming in microbially experienced ‘dirty’ mice results from limited IL-27 production by XCR1+ dendritic cells. J. Immunol. 209, 2149–2159 (2022).
Burger, S. et al. Natural microbial exposure from the earliest natural time point enhances immune development by expanding immune cell progenitors and mature immune cells. J. Immunol. 210, 1740–1751 (2023).
Martin, M. D. et al. CD115+ monocytes protect microbially experienced mice against E. coli-induced sepsis. Cell Rep. 42, 113345 (2023).
Li, Y. et al. Sequential early-life viral infections modulate the microbiota and adaptive immune responses to systemic and mucosal vaccination. PLoS Pathog. 20, e1012557 (2024).
Anderson, K. G. et al. Intravascular staining for discrimination of vascular and tissue leukocytes. Nat. Protoc. 9, 209–222 (2014).
DuPage, M. & Bluestone, J. A. Harnessing the plasticity of CD4+ T cells to treat immune-mediated disease. Nat. Rev. Immunol. 16, 149–163 (2016).
Wang, H., Gavil, N. V., Koewler, N., Masopust, D. & Jameson, S. C. Parabiosis in mice to study tissue residency of immune cells. Curr. Protoc. 2, e446 (2022).
Hayward, S. L. et al. Environmental cues regulate epigenetic reprogramming of airway-resident memory CD8+ T cells. Nat. Immunol. 21, 309–320 (2020).
Watanabe, R. et al. Human skin is protected by four functionally and phenotypically discrete populations of resident and recirculating memory T cells. Sci. Transl. Med. 7, 279ra239 (2015).
Bartolome-Casado, R. et al. Resident memory CD8 T cells persist for years in human small intestine. J. Exp. Med. 216, 2412–2426 (2019).
Christo, S. N. et al. Discrete tissue microenvironments instruct diversity in resident memory T cell function and plasticity. Nat. Immunol. 22, 1140–1151 (2021).
Fonseca, R. et al. RUNX3 drives a CD8+ T cell tissue residency program that is absent in CD4+ T cells. Nat. Immunol. 23, 1236–1245 (2022).
Dani, N. et al. A cellular and spatial map of the choroid plexus across brain ventricles and ages. Cell 184, 3056–3074 (2021).
Constant, O. et al. Role of dendritic cells in viral brain infections. Front. Immunol. 13, 862053 (2022).
Manglani, M. & McGavern, D. B. Intravital imaging of neuroimmune interactions through a thinned skull. Curr. Protoc. Immunol. 120, 24.2.1–24.2.12 (2018).
GBD 2016 Neurology Collaborators.Global, regional, and national burden of neurological disorders, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Neurol. 18, 459–480 (2019).
GBD 2021 Nervous System Disorders Collaborators. Global, regional, and national burden of disorders affecting the nervous system, 1990–2021: a systematic analysis for the Global Burden of Disease Study 2021. Lancet Neurol. 23, 344–381 (2024).
Grone, B. P. & Baraban, S. C. Animal models in epilepsy research: legacies and new directions. Nat. Neurosci. 18, 339–343 (2015).
Buchanan, G. F., Murray, N. M., Hajek, M. A. & Richerson, G. B. Serotonin neurones have anti-convulsant effects and reduce seizure-induced mortality. J. Physiol. 592, 4395–4410 (2014).
Rogawski, M. A. Molecular targets versus models for new antiepileptic drug discovery. Epilepsy Res. 68, 22–28 (2006).
Toman, J. E., Swinyard, E. A. & Goodman, L. S. Properties of maximal seizures, and their alteration by anticonvulant drugs and other agents. J. Neurophysiol. 9, 231–239 (1946).
Narasimhan, H. et al. An aberrant immune–epithelial progenitor niche drives viral lung sequelae. Nature 634, 961–969 (2024).
Evrard, M. et al. Single-cell protein expression profiling resolves circulating and resident memory T cell diversity across tissues and infection contexts. Immunity 56, 1664–1680 (2023).
Su, W. et al. CXCR6 orchestrates brain CD8+ T cell residency and limits mouse Alzheimer’s disease pathology. Nat. Immunol. 24, 1735–1747 (2023).
Leitner, D. et al. Similar brain proteomic signatures in Alzheimer’s disease and epilepsy. Acta Neuropathol. 147, 27 (2024).
Kumar, P. et al. Single-cell transcriptomics and surface epitope detection in human brain epileptic lesions identifies pro-inflammatory signaling. Nat. Neurosci. 25, 956–966 (2022).
Gate, D. et al. CD4+ T cells contribute to neurodegeneration in Lewy body dementia. Science 374, 868–874 (2021).
Galiano-Landeira, J., Torra, A., Vila, M. & Bove, J. CD8 T cell nigral infiltration precedes synucleinopathy in early stages of Parkinson’s disease. Brain 143, 3717–3733 (2020).
Kedia, S. et al. T cell-mediated microglial activation triggers myelin pathology in a mouse model of amyloidosis. Nat. Neurosci. 27, 1468–1474 (2024).
Sun, E.D. et al. Spatial transcriptomic clocks reveal cell proximity effects in brain ageing. Nature 638, 160–171 (2024).
Badovinac, V. P., Messingham, K. A., Jabbari, A., Haring, J. S. & Harty, J. T. Accelerated CD8+ T-cell memory and prime-boost response after dendritic-cell vaccination. Nat. Med. 11, 748–756 (2005).
Van Braeckel-Budimir, N., Varga, S. M., Badovinac, V. P. & Harty, J. T. Repeated antigen exposure extends the durability of influenza-specific lung-resident memory CD8+ T cells and heterosubtypic immunity. Cell Rep. 24, 3374–3382 (2018).
Anthony, S. M. et al. Protective function and durability of mouse lymph node-resident memory CD8+ T cells. eLife 10, e68662 (2021).
Hickman, H. D. Imaging CD8+ T cells during diverse viral infections. Intravital 4, e1055425 (2015).
Acknowledgements
We thank members of the laboratories of J.T.H., V.P.B., T.S.G. and G.F.B. for valuable discussions. We thank I. Badovinac, C. Sievers, J. Harty (no relation), Z. Darr, C. Mills and A. Tillford for maintaining laboratory solutions and equipment. We recognize the laboratory of A. Bassuk for sharing their slide-scanning microscope for image procurement. We acknowledge the Iowa NeuroBank Core for the procurement of human choroid plexus specimens with help from Q. Lin. Human biobanking services in the Iowa NeuroBank Core are supported by the Iowa Neuroscience Institute, Department of Pathology at the University of Iowa Roy J. and Lucille A Carver College of Medicine and the Roy J. Carver Charitable Trust. We also acknowledge the Iowa Comparative Pathology Laboratory with help from M. Leidinger. We thank E. Kozurek for assistance in procuring tissues from SPF and CoH mice. Tetramers were provided by the NIH Tetramer Core. Graphical illustrations from figures were created with BioRender.com. This work was supported by National Institutes of Health grants R01AI042767, R01AI167847, R21AI178159, R21AI185067 (J.T.H.), R01AI114543 (J.T.H. and V.P.B.), R35GM134880 (V.P.B.), R21AI154527, R35GM140881 (T.S.G.), R01NS129722 (G.F.B.), T32GM139776 (M.R.M., B.L.K. and S.A.A.), T32AI007485 (R.R.B.), T32AI007260 (M.A.H. and C.E.F.) and F32AI174382 (M.A.H.); Veterans Administration grants I01BX001324 and I21BX005679 (T.S.G.); a Joanna Sophia Grant from CURE Epilepsy (G.F.B.); the Beth L. Tross Epilepsy Professorship (G.F.B.); the University of Iowa Graduate College Post-Comprehensive Research Fellowship (M.R.M., B.L.K. and S.K.K.) and the Howard Hughes Medical Institute Hanna H. Gray Fellows Program (C.E.F.).
Author information
Authors and Affiliations
Contributions
Conceptualization: M.R.M., G.F.B., T.S.G., V.P.B. and J.T.H. Methodology: M.R.M., B.L.K., R.R.B., G.F.B., T.S.G., V.P.B. and J.T.H. Investigation: M.R.M., B.L.K., R.R.B., J.X., C.E.F., S.v.d.W., L.L.P., L.S.H., M.A.H., S.K.K., S.A.A. and C.M.S. Visualization: M.R.M. and J.T.H. Funding acquisition: G.F.B., T.S.G., V.P.B. and J.T.H. Project administration: G.F.B., T.S.G., V.P.B. and J.T.H. Supervision: G.F.B., T.S.G., V.P.B. and J.T.H. Writing, original draft: M.R.M., V.P.B. and J.T.H. Writing, review and editing: M.R.M., B.L.K., R.R.B., J.X., C.E.F., S.v.d.W., L.L.P., L.S.H., M.A.H., S.K.K., S.A.A., C.M.S., G.F.B., T.S.G., V.P.B. and J.T.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Thomas Korn, Aron Lukacher and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: L. A. Dempsey, in collaboration with the rest of the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Representative gating strategy.
Representative gating strategy of immune cells in the brain including microglia, monocytes, neutrophils, dendritic cells (DC), NK cells, B cells, CD4 T cells, and CD8 T cells. Intravenous exclusion (IV-) was only performed among SPF and SPExp mice.
Extended Data Fig. 2 Brain immune cell enumeration in SPF, CoH, and SPExp mice.
(a) Numbers of brain-localized immune cells between SPF (n = 6) and CoH (n = 6) mice. (b) Numbers of IV- immune cells in SPF (n = 3) and SPExp (n = 4) mice. Experiments in (a,b) show data from 1 of 2 independent experiments. Experiments comparing SPF vs. CoH mice and SPF vs. SPExp mice were conducted independently due to housing at separate academic institutions. Statistical significance was determined by two-sided unpaired Student’s t-test. Graphs show the mean ± s.e.m. Individual P values are noted as the following: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 3 Splenic memory T cell populations are proportionally enhanced in microbe-experienced mice.
(a) Representative flow plot, frequency, and numbers of antigen-experienced (Ag-Exp, CD11ahi) CD8+ T cells in the spleens of SPF (n = 6) and CoH (n = 6) mice at D60. (b) Representative flow plot, frequency, and numbers of antigen-experienced (Ag-Exp, CD11ahiCD44hi) CD4+ T cells in the spleens of SPF and CoH mice. (c,d) Same as for (a,b) but for SPF (n = 5) and SPExp (n = 5) mice. Experiments in (a-d) show data from 1 of 3 independent experiments. Statistical significance was determined by two-sided unpaired Student’s t-test. Graphs show the mean ± s.e.m. with each symbol representing one mouse. Individual P values are noted on respective graphs.
Extended Data Fig. 4 Microbe-experienced mice harbor brain-localized memory T cells with TRM phenotypes.
(a) Representative histograms and (b) gMFI of TRM-associated marker expression (that is CD69, CD49a, CD103, PD-1, CXCR6, and CXCR3) and TCIRCM-associated marker expression (that is CX3CR1, CD62L) among Ag-Exp (CD11ahi) CD8+ memory T cells derived from the spleen and brain of CoH mice (n = 6) at D60. (c, d) Same as for (a,b) but for Ag-Exp (CD11ahiCD44hi) CD4+ T cells. (e-h) Same as for (a-d) but in SPExp mice (n = 5). Experiments in (a-h) show data from 1 of 2 independent experiments. Statistical significance was determined by two-sided unpaired Student’s t-test for each marker. Graphs show the mean ± s.e.m. with each symbol representing one mouse. Individual P values are noted as the following: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Extended Data Fig. 5 The human choroid plexus harbors CD8+ and CD4+ T cells.
Lateral ventricle-derived choroid plexus was procured from three healthy human donors and stained for CD8α, CD4, or isotype control (cyan) and DAPI (magenta). Experiments show one representative image from n = 2 replicate sections from each human donor. Scale bar = 20 μm. Graphical illustrations were created using BioRender (https://biorender.com).
Extended Data Fig. 6 Gray matter regions contain increased CD8+ and CD4+ T cells in microbially-experienced mice.
a, Schematic of the imaged locations. b, Representative immunofluorescent images of CD8α+ T cells (green) and CD31+ vasculature (magenta) across gray matter regions (cerebral cortex, hypothalamus, brainstem, and cerebellum) of n = 3 mice per group after D60 + . c, Same as b but for CD4+ T cells (green). Experiments in b and c show one representative image from n = 3-4 replicate mice at every gray matter region from 2 independent experiments. Scale bar = 200 μm. Graphical illustrations were created using BioRender (https://biorender.com).
Extended Data Fig. 7 Neurogenic niches in microbe-experienced mice have increased presence of T cells.
(a) Representative immunofluorescent images of CD8α+ T cells (green) and CD31+ vasculature (magenta) across neurogenic niches (subventricular zone and dentate gyrus) of n = 3 mice per group after D60 + . (b) Same as A but for CD4+ T cells (green). Experiments in (a,b) show one representative image from n = 3-4 replicate mice at every neurogenic niche region from 2 independent experiments. Scale bar = 200 μm. Graphical illustrations were created using BioRender (https://biorender.com).
Extended Data Fig. 8 Memory T cells in the spleen are not functionally altered after seizure induction.
(a) Absolute numbers of Ag-Exp (CD11ahi) CD8+ or (b) (CD11ahiCD44hi) CD4+ T cells in the spleens of SPF mice 2 days after sham (n = 4) or seizure (Sz) induction (n = 4) or SPExp mice 2 day after sham (n = 10) or seizure induction (n = 10). (c) Representative flow plot and (d) proportions of splenic IFNγ+ TNF+ CD8+ or (e) CD4+ T cells from SPExp mice after PMA/ionomycin stimulation. (f) Frequency of CD107a+ degranulated splenic CD8+ T cells from SPExp mice after PMA/ionomycin stimulation. (g) Frequency of Ki-67+ CD8+ and (h) CD4+ T cells from SPExp spleens following sham (n = 5) or seizure induction (n = 5). (i) gMFI of MitoTrackerTM Green (MTG) in CD8+ and (j) CD4+ T cells from SPExp spleens following sham (n = 4) or seizure induction (n = 3). Experiments in (a-f) show concatenated data from 3 independent experiments. Experiments in (g-j) show representative data from 1 of 2 independent experiments. Statistical significance was determined by two-sided unpaired Student’s t-test. Graphs show the mean ± s.e.m. with each symbol representing one mouse. Individual P values are noted on respective graphs.
Extended Data Fig. 9 Validation of antibody depletion strategies and CXCR6 signaling abrogation.
(a) Experimental design. C57BL/6 N SPExp mice were treated with high-dose isotype antibody, anti-CD4 antibody, anti-CD8α antibody, or a combination of both antibodies to deplete CD4+ and CD8+ T cells respectively on D30 and D36 post-SPExp infection initiation. On D40, peripheral blood and brain tissue was isolated to determine depletion efficiency. (b) Numbers of CD4+ and (c) CD8+ T cells in the peripheral blood after isotype (n = 11), anti-CD4/CD8α (n = 7), anti-CD4 (n = 9) or anti-CD8α (n = 10) antibody treatments. (d) Numbers of CD4+ and (e) CD8+ T cells in the IV- brain after isotype (n = 5), anti-CD4/CD8α (n = 3), anti-CD4 (n = 3) or anti-CD8α (n = 3) antibody treatment. (f) Experimental design. WT or Cxcr6KO C57BL/6 J mice underwent SPExp infections. On D60, spleen and brain tissue were isolated to determine effects on T cell numbers and phenotypes. (g) Numbers of CD4+ and (h) CD8+ T cells in the peripheral blood of WT (n = 4) and Cxcr6KO (n = 7) mice. (i) Numbers of CD4+ and (j) CD8+ T cells in the IV- brain of WT and Cxcr6KO mice. (k) Representative histogram of CXCR6 expression among Ag-Exp CD4+ and CD8+ T cells derived from the brains of WT and Cxcr6KO mice. Experiments in (b,c) show concatenated data from 2 independent experiments. Experiments in (g-k) show representative data from 1 of 2 independent experiments. Statistical significance was determined by two-sided unpaired Student’s t-test or two-sided one-way ANOVA with Tukey’s multiple comparisons test. Graphs show the mean ± s.e.m. with each symbol representing one mouse. Individual P values are noted on respective graphs. Graphical illustrations were created using BioRender (https://biorender.com).
Supplementary information
Supplementary Video 1 (download MP4 )
Dynamic interactions between memory OT-I and CD11c+ cells in the brain. Representative time-lapse video shows dynamics of memory CD8+ T cells and CD11c+ cells through a thinned-skull window by two-photon laser-scanning microscopy of DC–rLM-OVA-immunized CD11c–eYFP mice at a memory time point. Memory eGFP OT-I T cells (green), CD11c+ cells (yellow) and blood vessels (magenta) were imaged over a 25-min period; scale bar, 20 μm.
Supplementary Video 2 (download MP4 )
Dynamic interactions between memory OT-I and CD11c+ cells in the brain. Representative time-lapse video shows dynamics of memory CD8+ T cells and CD11c+ cells through a thinned-skull window by two-photon laser-scanning microscopy of DC–rLM-OVA-immunized CD11c–eYFP mice at a memory time point. Memory eGFP OT-I T cells (green), CD11c+ cells (yellow) and blood vessels (magenta) were imaged over a 15-min period; scale bar, 20 μm.
Supplementary Video 3 (download MP4 )
Dynamic interactions between memory OT-I and CX3CR1+ cells in the brain. Representative time-lapse video shows dynamics of memory CD8+ T cells and CX3CR1+ cells through a thinned-skull window by two-photon laser-scanning microscopy of DC–rLM-OVA-immunized CX3CR1–TdTomato mice at a memory time point. Memory eGFP OT-I T cells (green), CX3CR1+ cells (indigo) and blood vessels (magenta) were imaged over an 18-min period; scale bar, 20 μm.
Supplementary Video 4 (download MP4 )
Dynamic interactions between memory OT-I and CX3CR1+ cells in the brain. Representative time-lapse video shows dynamics of memory CD8+ T cells and CX3CR1+ cells through a thinned-skull window by two-photon laser-scanning microscopy of DC–rLM-OVA-immunized CX3CR1–TdTomato mice at a memory time point. Memory eGFP OT-I T cells (green), CX3CR1+ cells (indigo) and blood vessels (magenta) were imaged over a 17-min period; scale bar, 15 μm.
Supplementary Table 1 (download PDF )
Choroid plexus donor demographics.
Supplementary Table 2 (download PDF )
Antibodies and tetramers used for flow cytometry and tissue staining.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mix, M.R., Kreitlow, B.L., Berton, R.R. et al. Physiological microbial exposure normalizes memory T cell surveillance of the brain and modifies host seizure outcomes. Nat Immunol 26, 1087–1098 (2025). https://doi.org/10.1038/s41590-025-02174-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02174-y
This article is cited by
-
Peripheral infection wires T cells in the brain
Nature Immunology (2025)