Abstract
Conventional dendritic cells (cDCs) are a heterogeneous population of professional antigen-presenting cells that bridge innate and adaptive immunity. Many studies in mice have identified various populations of cDCs whose inter-relationships and discrete identities, as well as their link to plasmacytoid DCs (pDCs), have not been cohesively addressed. Here, by combining single-cell sequencing, transcription factor fate-mapping models, conditional knockout models and adoptive transfer, we show that Klf4 expression clearly separates cDC lineage from the pDC lineage, and defined two pre-DC2 subsets: Siglec-H+CD115− pre-DC2s and Siglec-HloCD115+ pre-DC2s. While Siglec-H+CD115− pre-DC2s represent the pDC-like cells that give rise to CD7+CD11blo DC2As in a TCF4-dependent manner, Siglec-HloCD115+ pre-DC2s give rise to CD7−CD11bhi DC2Bs in a KLF4-dependent manner. These data reveal the transcriptional basis of two pre-DC2 subsets and present a firm framework for mouse cDC classification, paving the way for a better understanding of these cells in tissues and in disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The scRNA-seq datasets used in this study are from Liu et al.11.
Code availability
This study did not report original code.
References
Steinman, R. M. & Idoyaga, J. Features of the dendritic cell lineage. Immunol. Rev. 234, 5–17 (2010).
Guilliams, M. et al. Unsupervised high-dimensional analysis aligns dendritic cells across tissues and species. Immunity 45, 669–684 (2016).
Anderson, D. A. 3rd, Dutertre, C. A., Ginhoux, F. & Murphy, K. M. Genetic models of human and mouse dendritic cell development and function. Nat. Rev. Immunol. 21, 101–115 (2021).
Guilliams, M. et al. Dendritic cells, monocytes and macrophages: a unified nomenclature based on ontogeny. Nat. Rev. Immunol. 14, 571–578 (2014).
Siegal, F. P. et al. The nature of the principal type 1 interferon-producing cells in human blood. Science 284, 1835–1837 (1999).
Cella, M. et al. Plasmacytoid monocytes migrate to inflamed lymph nodes and produce large amounts of type I interferon. Nat. Med. 5, 919–923 (1999).
Eisenbarth, S. C. Dendritic cell subsets in T cell programming: location dictates function. Nat. Rev. Immunol. 19, 89–103 (2019).
Cabeza-Cabrerizo, M., Cardoso, A., Minutti, C. M., Pereira da Costa, M. & Reis e Sousa, C. Dendritic cells revisited. Annu. Rev. Immunol. 39, 131–166 (2021).
Ginhoux, F., Guilliams, M. & Merad, M. Expanding dendritic cell nomenclature in the single-cell era. Nat. Rev. Immunol. 22, 67–68 (2022).
Brown, C. C. et al. Transcriptional basis of mouse and human dendritic cell heterogeneity. Cell 179, 846–863 (2019).
Liu, Z. et al. Dendritic cell type 3 arises from Ly6C+ monocyte-dendritic cell progenitors. Immunity 56, 1761–1777 (2023).
Rodrigues, P. F. et al. Distinct progenitor lineages contribute to the heterogeneity of plasmacytoid dendritic cells. Nat. Immunol. 19, 711–722 (2018).
Leylek, R. et al. Integrated cross-species analysis identifies a conserved transitional dendritic cell population. Cell Rep. 29, 3736–3750 (2019).
Villani, A. C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017).
Rodrigues, P. F. et al. pDC-like cells are pre-DC2 and require KLF4 to control homeostatic CD4 T cells. Sci. Immunol. 8, eadd4132 (2023).
Sulczewski, F. B. et al. Transitional dendritic cells are distinct from conventional DC2 precursors and mediate proinflammatory antiviral responses. Nat. Immunol. 24, 1265–1280 (2023).
Fogg, D. K. et al. A clonogenic bone marrow progenitor specific for macrophages and dendritic cells. Science 311, 83–87 (2006).
Auffray, C. et al. CX3CR1+ CD115+ CD135+ common macrophage/DC precursors and the role of CX3CR1 in their response to inflammation. J. Exp. Med. 206, 595–606 (2009).
Onai, N., Manz, M. G. & Schmid, M. A. Isolation of common dendritic cell progenitors (CDP) from mouse bone marrow. Methods Mol. Biol. 595, 195–203 (2010).
Onai, N. et al. Identification of clonogenic common Flt3+M-CSFR+ plasmacytoid and conventional dendritic cell progenitors in mouse bone marrow. Nat. Immunol. 8, 1207–1216 (2007).
Naik, S. H. et al. Development of plasmacytoid and conventional dendritic cell subtypes from single precursor cells derived in vitro and in vivo. Nat. Immunol. 8, 1217–1226 (2007).
Schlitzer, A. et al. Identification of cDC1- and cDC2-committed DC progenitors reveals early lineage priming at the common DC progenitor stage in the bone marrow. Nat. Immunol. 16, 718–728 (2015).
Grajales-Reyes, G. E. et al. Batf3 maintains autoactivation of Irf8 for commitment of a CD8ɑ+ conventional DC clonogenic progenitor. Nat. Immunol. 16, 708–717 (2015).
Liu, K. et al. In vivo analysis of dendritic cell development and homeostasis. Science 324, 392–397 (2009).
Lin, D. S. et al. DiSNE movie visualization and assessment of clonal kinetics reveal multiple trajectories of dendritic cell development. Cell Rep. 22, 2557–2566 (2018).
Dress, R. J. et al. Plasmacytoid dendritic cells develop from Ly6D+ lymphoid progenitors distinct from the myeloid lineage. Nat. Immunol. 20, 852–864 (2019).
Schlitzer, A. et al. Identification of CCR9− murine plasmacytoid DC precursors with plasticity to differentiate into conventional DCs. Blood 117, 6562–6570 (2011).
Lutz, K. et al. Ly6D+Siglec-H+ precursors contribute to conventional dendritic cells via a Zbtb46+Ly6D+ intermediary stage. Nat. Commun. 13, 3456 (2022).
See, P. et al. Mapping the human DC lineage through the integration of high-dimensional techniques. Science 356, eaag3009 (2017).
Tussiwand, R. et al. Klf4 expression in conventional dendritic cells is required for T helper 2 cell responses. Immunity 42, 916–928 (2015).
Minutti, C. M. et al. Distinct ontogenetic lineages dictate cDC2 heterogeneity. Nat. Immunol. 25, 448–461 (2024).
Rodrigues, P. F. et al. Progenitors of distinct lineages shape the diversity of mature type 2 conventional dendritic cells. Immunity 57, 1567–1585 (2024).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Kamimoto, K. et al. Dissecting cell identity via network inference and in silico gene perturbation. Nature 614, 742–751 (2023).
Cisse, B. et al. Transcription factor E2-2 is an essential and specific regulator of plasmacytoid dendritic cell development. Cell 135, 37–48 (2008).
Grajkowska, L. T. et al. Isoform-specific expression and feedback regulation of E protein TCF4 control dendritic cell lineage specification. Immunity 46, 65–77 (2017).
Ghosh, H. S., Cisse, B., Bunin, A., Lewis, K. L. & Reizis, B. Continuous expression of the transcription factor E2-2 maintains the cell fate of mature plasmacytoid dendritic cells. Immunity 33, 905–916 (2010).
Scott, C. L. et al. The transcription factor Zeb2 regulates development of conventional and plasmacytoid DCs by repressing Id2. J. Exp. Med. 213, 897–911 (2016).
Wu, X. et al. Transcription factor Zeb2 regulates commitment to plasmacytoid dendritic cell and monocyte fate. Proc. Natl Acad. Sci. USA 113, 14775–14780 (2016).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Satpathy, A. T. et al. Zbtb46 expression distinguishes classical dendritic cells and their committed progenitors from other immune lineages. J. Exp. Med. 209, 1135–1152 (2012).
Tomura, M. et al. Contrasting quiescent G0 phase with mitotic cell cycling in the mouse immune system. PLoS ONE 8, e73801 (2013).
Goodwin, L. O. et al. Large-scale discovery of mouse transgenic integration sites reveals frequent structural variation and insertional mutagenesis. Genome Res. 29, 494–505 (2019).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (NSFC) (grants 32270916 and 32070880 to Z. Liu) and Shanghai Jiao Tong University 2030 Initiative (grant WH510363001-16 to Z. Liu). We thank the flow cytometry team, sequencing core and imaging core at Shanghai Institute of Immunology, and the Core Facility of Basic Medical Sciences, Shanghai Jiao Tong University School of Medicine, for their support.
Author information
Authors and Affiliations
Contributions
Y. Zhu, H.W., S.Z., P.C., F. Gao, W.T.K. and J.Q. conducted the experiments; Z. Li, Y. Zeng, Z. Liu and F. Ginhoux analyzed the data; Z. Liu and F. Ginhoux wrote the paper; B.S. provided intellectual input; Z. Liu and F. Ginhoux conceptualized and supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Stephanie Houston in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Pre-DC2s contain cells with a pDC-like phenotype.
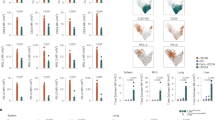
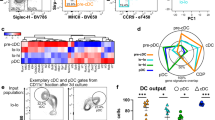
a, Dot plot showing the expression of top 10 most differentationally expressed genes across clusters. Colors indicate the average expression of each gene. Spot sizes represent the proportion of gene-expressing cells. b, UMAP plots showing clusters in Siglec-H+ fraction in BM and spleen, data from Rodrigues et al., 2018. c, Stacked violin plots showing the expression of selected genes across clusters. d, Volcano plot showing the DEGs between BM pDC-like cells and pre-pDCs. Genes in brown are upregulated in pDC-like cells; genes in pink are upregulated in pre-pDCs. Horizontal dashed line indicates p-value = 0.05; vertical dashed lines indicate fold change = 1.5 or 0.67 (log2FC = 0.3 or -0.3). e, Stacked violin plot showing the expression of selected genes identified in (d). f, Venn plots showing the overlap of top 25 genes in pre-DC2s (upper panel) or pDCs (lower panel) in BM, blood, and spleen. g, Stacked violin plot showing the expression of genes identified in (f).
Extended Data Fig. 2 KLF4 expression distinguishes cDCs from pDCs.
a, Schemic showing the construct of Klf4EGFP-CreERT2 model, an IRES-EGFP-2A-CreERT2-Wpre-pA cassette were inserted into the 3’ UTR region of Klf4 gene. b, t-distributed Stochastic Neighbor Embedding (tSNE) plots and histograms showing KLF4-GFP expression in the blood of Klf4EGFP-CreERT2 mice. c,d, Flow cytometry plots showing the gating strategy for monocyte and DC subsets in the blood (c), and spleen (d) of Klf4EGFP-CreERT2 mice.
Extended Data Fig. 3 KLF4 expression distinguishes cDCs from pDCs.
a, Flow cytometry plots showing the gating strategy for monocyte and DC subsets in the BM of Klf4EGFP-CreERT2 mice. b, tSNE plots showing KLF4-GFP+ and GFP− populations in Lin−FLT3+ cells separately and merged from BM of Klf4EGFP-CreERT2 mice. c, Quantification of cells generated per 105 FLT3+KLF4-GFP+ or FLT3+c-Kit-KLF4-GFP− cells in in vitro culture (n = 4), data are presented as mean ± s.d. d, Histogram showing the expression of KLF4-GFP and LGALS3 in populations in the BM of Klf4EGFP-CreERT2 mice. e, Flow cytometry plots showing the expression of KLF4-GFP and LGALS3.
Extended Data Fig. 4 Pre-DC2s contain two subpopulations.
a,b, Gating strategy for pre-DC2 subsets in the blood (a) and spleen (b) of Klf4EGFP-CreERT2 mice. c, Giemsa staining image showing the morphology of indicated cells sorted from the BM of Klf4EGFP-CreERT2 mice. Scale bar = 10 μm. d, Quantification of cells generated per 105 Siglec-H+ pre-DC2 or CD115+ pre-DC2 in in vitro culture (n = 4), data are presented as mean ± s.d. e, Gating strategy for DC populations in the spleen of Klf4EGFP-CreERT2 mice. f, Histogram plots showing the expression of CD7-EGFP and CD45RB in DC and monocyte populations in the spleen of Cd7EGFP mice. g, Quantification of cells generated per 105 Siglec-H+ pre-DC2 or CD115+ pre-DC2 in in vivo adoptive transfer (n = 3), data are presented as mean ± s.d.
Extended Data Fig. 5 Siglec-H+ pre-DC2s can be aligned to pDC-like cells and tDCs.
a, Overlay of pre-DC2 subsets defined with our gating strategy to the gating of pDC-like cells in Rodrigues et al., 2018. b, Overlaying pre-DC2 subsets defined with our gating strategy to theirs. c, Overlay of CD11chi or CD11clo tDCs defined by Leylek et al., 2019 to our gating strategy. d, Percentages of proliferating cell indicated by Fucci expression in populations in the BM, blood, and spleen of Fucci mice (n = 4), data are presented as mean ± s.d.
Extended Data Fig. 6 Siglec-H+ pre-DC2s can be aligned to pDC-like cells and tDCs.
a, Overlay of pre-DC2 subsets defined with our gating strategy to pre-DC2A and pre-DC2B defined by Minutti et al., 2024 to our gating strategy, b, Overlay of tDC, DC2B, early DC2A, and DC2A defined by Minutti et al., 2024 to our gating strategy.
Extended Data Fig. 7 Siglec-H+ pre-DC2s can be aligned to pDC-like cells and tDCs.
a, Overlay of Siglec-H+Zbtb46-GFP+Ly6D+ cells defined by Lutz et al., 2022 to our gating strategy. b, Overlay of Siglec-H+B220+ pDCs and pre-pDCs defined with our gating strategy to populations defined by Lutz et al., 2022. c, Flow cytometry plot showing the overlay of pre-pDCs (cyan) defined with our gating strategy with B220loCCR9lo cells (red) defined by Lutz et al., 2022. d, Schemic showing the relationship between populations in DC2 lineage.
Extended Data Fig. 8 KLF4 and TCF4 control the development of pre-DC2 populations.
a, Schemic showing the construct of Tcf4tdTomato-LSL-DTA model, an tdTomato-LoxP-stop-LoxP-IRES-DTA-Wpre-pA cassette was inserted into the 3’ UTR region of Tcf4 gene. b, Flow cytometry (left panel) and histogram (right panel) showing the pDCs and cDCs in Klf4fl/fl and Klf4fl/fl;Vav1iCre spleen (n = 3). c, Schemic showing the construct of Tcf4flox model. d, Flow cytometry (left panel) and histogram (right panel) showing the pDCs and cDCs in Tcf4fl/fl and Tcf4fl/fl;Vav1iCre spleen (n = 5). Data in this figure are presented as mean ± s.d.; statistics were calculated by unpaired two-sided Student’s t-test; ns, not significant, *P < 0.05, ****P < 0.0001.
Extended Data Fig. 9 Proposed model for DC lineage development.
Schemic showing the proposed model for DC lineage development.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhu, Y., Cai, P., Li, Z. et al. Transcription factors TCF4 and KLF4 respectively control the development of the DC2A and DC2B lineages. Nat Immunol 26, 1275–1286 (2025). https://doi.org/10.1038/s41590-025-02208-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02208-5
This article is cited by
-
2B or not 2B is the question in DC ontogeny
Nature Immunology (2025)
-
Crosstalk between the microbiota and intestinal dendritic cells in IBD
Seminars in Immunopathology (2025)
-
Sequential lymphotoxin-β receptor and retinoic acid receptor signals regulate cDC2 fate
Nature Immunology (2025)