Abstract
Granulocyte-macrophage colony-stimulating factor (GM-CSF, Csf2) is a potent proinflammatory cytokine1. At steady state, however, GM-CSF has a distinct homeostatic function, being essential for the differentiation and maintenance of alveolar macrophages2,3,4,5. Whether macrophage development in nonpulmonary tissues is similarly dependent on GM-CSF is unclear. Here we examine developing tissues of GM-CSF fate-mapping and reporter mice6. We show that type 2 innate lymphoid cells (ILC2s) in the salivary glands produce GM-CSF and identify a macrophage subset that we refer to as ‘adenophages’. Adenophages are noncanonical macrophages that are derived from fetal monocytes and are progressively replaced by monocyte–dendritic cell progenitor-derived monocytes. These cells form a spatial niche triad with GM-CSF-producing ILC2s and myoepithelial cells and are required for efficient secretion of saliva. Importantly, adenophages are present throughout exocrine glands, including lacrimal glands and mammary glands, and are also present in humans, indicating a conserved role in exocrine glands across species.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data have been deposited in the Gene Expression Omnibus under accession no. GSE280844.
References
Becher, B., Tugues, S. & Greter, M. GM-CSF: from growth factor to central mediator of tissue inflammation. Immunity 45, 963–973 (2016).
Schneider, C. et al. Induction of the nuclear receptor PPAR-γ by the cytokine GM-CSF is critical for the differentiation of fetal monocytes into alveolar macrophages. Nat. Immunol. 15, 1026–1037 (2014).
Gschwend, J. et al. Alveolar macrophages rely on GM-CSF from alveolar epithelial type 2 cells before and after birth. J. Exp. Med. https://doi.org/10.1084/JEM.20210745/21260 (2001).
Guilliams, M. et al. Alveolar macrophages develop from fetal monocytes that differentiate into long-lived cells in the first week of life via GM-CSF. J. Exp. Med. 210, 1977–1992 (2013).
Bain, C. C. & MacDonald, A. S. The impact of the lung environment on macrophage development, activation and function: diversity in the face of adversity. Mucosal Immunol. 15, 223–234 (2022).
Komuczki, J. et al. Fate-mapping of GM-CSF Expression identifies a discrete subset of inflammation-driving T helper cells regulated by cytokines IL-23 and IL-1β. Immunity 50, 1289–1304 (2019).
McKendrick, J. G. et al. CSF1R-dependent macrophages in the salivary gland are essential for epithelial regeneration after radiation-induced injury. Sci. Immunol. 8, eadd4374 (2023).
Lu, L. et al. Differential expression of CD11c defines two types of tissue-resident macrophages with different origins in steady-state salivary glands. Sci. Rep. 12, 1–14 (2022).
de Boer, J. et al. Transgenic mice with hematopoietic and lymphoid specific expression of Cre. Eur. J. Immunol. 33, 314–325 (2003).
Tsou, A. M. et al. Neuropeptide regulation of non-redundant ILC2 responses at barrier surfaces. Nature 611, 787–793 (2022).
Jarick, K. J. et al. Non-redundant functions of group 2 innate lymphoid cells. Nature 611, 794–800 (2022).
Van Hove, H. et al. Interleukin-34-dependent perivascular macrophages promote vascular function in the brain. Immunity 58, 1289–1305 (2025).
Zubeidat, K. et al. Microbiota-dependent and -independent postnatal development of salivary immunity. Cell Rep. 42, 111981 (2023).
Koren, N. et al. Maturation of the neonatal oral mucosa involves unique epithelium-microbiota interactions. Cell Host Microbe 29, 197–209 (2021).
Croxford, A. L. et al. The cytokine GM-CSF drives the inflammatory signature of CCR2+ monocytes and licenses autoimmunity. Immunity 43, 502–514 (2015).
Liu, Z. et al. Fate mapping via Ms4a3-expression history traces monocyte-derived cells. Cell 178, 1509–1525 (2019).
Trzebanski, S. et al. Classical monocyte ontogeny dictates their functions and fates as tissue macrophages. Immunity 57, 1225–1242 (2024).
Schraml, B. U. et al. Genetic tracing via DNGR-1 expression history defines dendritic cells as a hematopoietic lineage. Cell 154, 843–858 (2013).
Liu, Z. et al. Dendritic cell type 3 arises from Ly6C+ monocyte-dendritic cell progenitors. Immunity 56, 1761–1777 (2023).
Guilliams, M. & Scott, C. L. Liver macrophages in health and disease. Immunity 55, 1515–1529 (2022).
Amorim, A. et al. IFNγ and GM-CSF control complementary differentiation programs in the monocyte-to-phagocyte transition during neuroinflammation. Nat. Immunol. 23, 217–228 (2022).
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Miller, J. C. et al. Deciphering the transcriptional network of the dendritic cell lineage. Nat. Immunol. 13, 888–899 (2012).
Black, S. et al. CODEX multiplexed tissue imaging with DNA-conjugated antibodies. Nat. Protoc. 16, 3802–3835 (2021).
Gordon, S. & Plüddemann, A. Tissue macrophages: heterogeneity and functions. BMC Biol. 15, 53 (2017).
Mass, E., Nimmerjahn, F., Kierdorf, K. & Schlitzer, A. Tissue-specific macrophages: how they develop and choreograph tissue biology. Nat. Rev. Immunol. 23, 563–579 (2023).
Bijnen, M. & Bajénoff, M. Gland macrophages: reciprocal control and function within their niche. Trends Immunol. 42, 120–136 (2021).
Emmelin, N. & Gjörstrup, P. On the function of myoepithelial cells in salivary glands. J. Physiol. 230, 185–198 (1973).
Hawley, D. et al. Myoepithelial cell-driven acini contraction in response to oxytocin receptor stimulation is impaired in lacrimal glands of Sjögren’s syndrome animal models. Sci. Rep. 8, 9919 (2018).
Lung, M. A. Autonomic nervous control of myoepithelial cells and secretion in submandibular gland of anaesthetized dogs. J. Physiol. 546, 837–850 (2003).
Hauser, B. R., Aure, M. H., Kelly, M. C., Hoffman, M. P. & Chibly, A. M. Generation of a single-cell RNAseq atlas of murine salivary gland development. iScience 23, 101838 (2020).
Lu, C. P. et al. Identification of stem cell populations in sweat glands and ducts reveals roles in homeostasis and wound repair. Cell 150, 136–150 (2012).
Mauduit, O. et al. A closer look into the cellular and molecular biology of myoepithelial cells across various exocrine glands. Ocul. Surf. 31, 63–80 (2024).
Thiemann, R. F. et al. Regulation of myoepithelial differentiation. PLoS ONE 17, e0268668 (2022).
Zubeidat, K., Saba, Y., Barel, O., Shoukair, F. L. & Hovav, A. H. Protocol for parotidectomy and saliva analysis in mice. STAR Protoc. 3, 101048 (2022).
Jäppinen, N. et al. Fetal-derived macrophages dominate in adult mammary glands. Nat. Commun. 10, 281 (2019).
Dawson, C. A. et al. Tissue-resident ductal macrophages survey the mammary epithelium and facilitate tissue remodelling. Nat. Cell Biol. 22, 546–558 (2020).
Cansever, D. et al. Lactation-associated macrophages exist in murine mammary tissue and human milk. Nat. Immunol. 24, 1098–1109 (2023).
Fan, Q. et al. Exploring immune cell diversity in the lacrimal glands of healthy mice: a single-cell RNA-sequencing atlas. Int. J. Mol. Sci. https://doi.org/10.3390/ijms25021208 (2024).
Gallerand, A. et al. CD226+ adipose tissue macrophages arise from MDP-derived monocytes and regulate lipid metabolism. Preprint at bioRxiv https://doi.org/10.1101/2024.12.03.626330 (2024).
Kim, K. W. et al. MHC II+ resident peritoneal and pleural macrophages rely on IRF4 for development from circulating monocytes. J. Exp. Med. https://doi.org/10.1084/jem.20160486 (2016).
Ivan, B. et al. Gain of virulence caused by loss of a gene in murine cytomegalovirus. J. Virol. 78, 7536–7544 (2004).
Lemmermann, N. A. W. et al. CD8 T-cell immunotherapy of cytomegalovirus disease in the murine model. Methods Microbiol. 37, 369–420 (2010).
Solis, N. V. & Filler, S. G. Mouse model of oropharyngeal candidiasis. Nat. Protoc. 7, 637–642 (2012).
Bayerl, F. et al. Guidelines for visualization and analysis of DC in tissues using multiparameter fluorescence microscopy imaging methods. Eur. J. Immunol. 53, e2249923 (2023).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Cotet, T. S. et al. ePlatypus: an ecosystem for computational analysis of immunogenomics data. Bioinformatics 39, btad553 (2023).
Andreatta, M. & Carmona, S. J. STACAS: Sub-Type Anchor Correction for Alignment in Seurat to integrate single-cell RNA-seq data. Bioinformatics 37, 882–884 (2021).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Wright, K. pals: color palettes, colormaps, and tools to evaluate them. R package v1.10 https://kwstat.github.io/pals/ (2025).
Mills, B. R. MetBrewer: color palettes inspired by works at The Metropolitan Museum of Art. R package v0.2.0 https://CRAN.R-project.org/package=MetBrewer (2022).
Yasuhara, R., Kang, S., Tokumasu, R. & Mishima, K. in Isolation and Functional Analysis of Myoepithelial Cells from Adult Mouse Submandibular Glands BT—Stem Cells and Lineage Commitment: Methods and Protocols (ed. Turksen, K.) 53–64 (Springer, 2024).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences [version 2; peer review: 2 approved]. F1000Research https://doi.org/10.12688/f1000research.7563.2 (2016).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Chen, Y., Chen, L., Lun, A. T. L., Baldoni, P. L. & Smyth, G. K. edgeR v4: powerful differential analysis of sequencing data with expanded functionality and improved support for small counts and larger datasets. Nucleic Acids Res. 53, gkaf018 (2025).
Decoene, I., Herpelinck, T., Geris, L., Luyten, F. P. & Papantoniou, I. Engineering bone-forming callus organoid implants in a xenogeneic-free differentiation medium. Front. Chem. Engin. https://www.frontiersin.org/journals/chemical-engineering/articles/10.3389/fceng.2022.892190 (2022).
Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41–44 (2020).
Wolski, W. E. et al. prolfqua: a comprehensive R-package for proteomics differential expression analysis. J. Proteome Res. 22, 1092–1104 (2023).
Acknowledgements
We thank the University of Zürich Center for Microscopy and Image Analysis for help with SEM; the Functional Genomics Center (University of Zürich) for technical support with the scRNA-seq, snRNA-seq and bulk RNA-seq experiments; and the University of Zürich Laboratory Animal Service Center for animal mouse husbandry. We also thank S. Mundt for sharing mice; L. Casty, M. Lutz and P. Häne for technical support; and C. Ulutekin for providing data analysis scripts (all at the University of Zurich). This work was supported by funds from the European Research Council under the European Union’s Horizon 2020 research and innovation program (grant no. 882424), the Swiss National Science Foundation (grant nos. 170320, 188450, 219287, CRSII5_183478 and CRSII–222718 to B.B.), Swiss Cancer Research (KFS-6022-02-2024-R to B.B.) and UK Research and Innovation (grant no. MR/X018733/1 to E.E.).
Author information
Authors and Affiliations
Contributions
F.W., S. Tuzlak and B.B. conceived of the study. F.W. and S. Tuzlak designed, performed and analyzed most experiments, apart from specific contributions outlined here. V.K. performed computational data analysis of the sequencing experiments. D.B. and A.S. performed the CODEX experiments and analysis. E.E. performed the immunofluorescent immunology on mouse and human tissue. A.I. performed the adenophage analysis in germ-free mice. M.B., V.C., H.v.H., H.W., G.L., C. Sparano, E.R., R.L., M.A., R.F.-M. and L.O. supported the mouse experiments. I.N. provided the human SMG samples. M.G., D.d.F., S. Tuques, C.C.B., A.K., K.M., A.G., S.L.-L. and S.I. were involved in data discussion. C. Schneider, Z.L. and F.G. provided the mice. B.B. supervised the study. F.W., S. Tuzlak and B.B. wrote the paper with contributions from S. Tuques, E.E. and all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Mona Sadeghalvad and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Nick Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
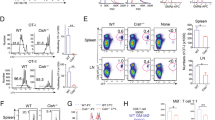
Extended Data Fig. 1 Lymphocytes are the exclusive source of GM-CSF for adenophages.
A) Genetic construct of the FROGxAi14 model. B) Representative FACS plot for tdTomato and eGFP expression in the SG at P20. C) Marker expressions in lymphoid populations in the SG at P20 D) Mosaic plot showing the fraction of ex-GM-CSF producing (red) and GM-CSF producing (green) cells in lymphoid populations in the SG of FROGxAi14 mice at P20. The width of the columns represents the abundance of the indicated populations. Gated on liveCD45+CD11b−. E) snRNaseq data showing expression of Csf1 and Csf2 within the CD45-negative population of the SG of Csf2+/− mice at P20 SG. F) Frequencies of myeloid cells (CD11b+), ILC2s, ILC1s, cNKcells, trNKcells and T cells in the SG of Csf2+/− mice over time. Data represented as mean and error±SD. n = 3-6 per timepoint. G) Frequencies of MHCIIlo MACS and MHCII+ MACS throughout postnatal development in the SG of Csf2+/− and Csf2−/− mice. Pre-gated on live CD45+Lin−Ly6G−SiglecF−Ly6C−CD11b+F4/80+ cells. Data are represented as mean ± SD. Statistical test: 2-way ANOVA. N = 6-12 mice per group and timepoint and data from 2-3 independent experiments per timepoint. H) Frequencies of DCs throughout postnatal development in the SG of Csf2+/− and Csf2−/− mice. Pre-gated on live CD45+Lin−Ly6G−SiglecF−Ly6C−F4/80−CD11c+MHCII+ cells. Statistical test: 2-way ANOVA. N = 6-12 mice per group and timepoint and data is shown from 2-3 independent experiments per timepoint. I) Frequencies of AMs of Csf2fl/fl and Csf2Vav1iCre mice at P20. Data are pooled from 2 independent experiments and are represented as mean ± SD. Statistical test: t-test. J) Frequencies of adenophages and MHCII+ MACS in Csf1rfl/fl mice and Csf1rMrc1Cre mice at P20 in the SG. Data represents mean ± SD of two independent experiments. K) Frequency (right) and counts (left) of MHCII+ MACS in the SG of germfree (purple) and SPF (blue) mice at indicated time-points. Statistical test: 2-way ANOVA; only significant p-values are displayed. Data is from 2 independent experiments per timepoint and represents n = 6-12 mice per group and timepoint. Data are represented as mean ± SEM. AM = alveolar macrophages; SPF = specific-pathogen-free. Statistical values: ns > 0.05; * ≤ 0.05; ** ≤ 0.01; *** ≤ 0.005; **** ≤ 0.001.
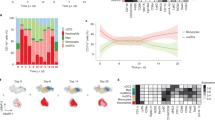
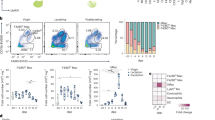
Extended Data Fig. 2 Adenophages arise from fetal monocytes and have a transcriptional signature imprinted by GM-CSF.
A) Experimental scheme showing the procedure to label fetal monocyctes in Ccr2CreERT2xAi14 mice. Graphics in A created using BioRender.com. B) Frequencies of tdTomato+ cells within MHCIIlo MACS (green), MHCII+MACS (blue) and AM (red) at indicated timepoints after Tamoxifen application at P0/P1 in Ccr2CreERT2xAi14 mice. Data represents 1-2 independent experiments per timepoint. N = 2-6 mice per timepoint. C) Total counts of tdTomato+ adenophages in the SG at indicated timepoints after Tamoxifen application at P0/P1 in Ccr2CreERT2xAi14 mice. Data represents 1-2 independent experiments per timepoint. N = 4-6 mice per timepoint. D) left: Frequency of tdTomato+ MHCIIlo MACS (green) and MHCII+ MACS (blue) in the SG of Ms4a3CrexAi14 mice at indicated timepoints. right: Frequency of tdTomato+ neutrophils, eosinophils and monocytes in the SG of Ms4a3CrexAi14 mice at indicated timepoints. Data represents 1-2 independent experiments per timepoint. N = 2-4 per timepoint. E) Frequency of tdTomato+ MHCII + MACS (blue), MHCIIloMACS (green) and DCs (brown) in Clec9aCrexAi14 mice. Data represents 1-2 independent experiments per timepoint. N = 2-5 per timepoint. B-E) Bar plots show frequency at indicated postnatal timepoints. Data are represented as mean±SEM. F) Comparison of gene expression in adenophages isolated from the SG and splenic DC3s (Liu et al., 2023). G) Probability prediction of pro-DC3s falling into one of the myeloid clusters identified in the SG of control mice at P20. H) UMAP visualisation of scRNAseq data on all CD45+ cells in the SG at P20. Bar diagram visualizes the abundance of the identified immune cell subsets in the SG of Csf2+/− and Csf2−/− mice at P20. I) Dot plot showing Csf2ra, Csf2rb and Csf1r expression across indicated myeloid subsets. Colors indicate the average expression of each gene, circle sizes represent percentage of cells within a cluster expressing the indicated gene. J) Violin plots showing expression of Irf4, Pparg, Cebpb and Batf in macrophage subsets, adenophages and cDC2 in the SG of Csf2+/− mice at P20.
Extended Data Fig. 3 Adenophages share functional and morphological features with DCs and macrophages.
A) FACS sorting strategy for adenophages, MHCII+MACS and DCs for SEM imaging. 2100 adenophages, 8800 MHCII+MACS and 3500 DCs were sorted from C57BL/6 mice at P20. B) Additional SEM images of FACS sorted adenophages. Scale = 2 µm. Representative images of 2 independent experiments with 5-10 images per cell type. C) Giemsa staining micrographs of sorted adenophages, MHCII+ MACS and DCs from the SG of Control mice at P20, representative images taken from two independent experiments. D) MFI of pHrodo Red+ adenophages, MHCII+ MACS, and DCs from the P20 SG of Csf2+/− mice after in vitro exposure to S.aureus pHrodo bioparticles for 180 min. Data is pooled from 3 independent experiments. Statistics: Kruskal-Wallis test with Dunn’s multiple comparisons test. Data in bar diagram is represented as mean±SD. N = 7. E) CellTraceViolet (CTV) expression in CD4+ OT-II transgenic T cells after co-culture with adenophages, MHCII+ MACS or DCs isolated from the SG of Csf2+/− mice at P20 and pulsed with OVA323–339 for 1 hour. 8-10 mice were pooled per experiment. Data is representative for two independent experiments. F) Candida albicans fungal load on the tongue of Csf2+/− and Csf2−/− mice at 3 days post infection. Mice were infected at P20 and data pooled from two independent experiments with total n = 4-5 mice per group and experiment. Statistical test: two-tailed Welch’s t-test. Data in bar diagram is represented as mean±SD. G) CODEX analysis showing the different cellular subsets by indicated color coding. Data is representative for one CODEX experiment with analysis of 6 independent sections. H) Immunofluorescent histology image of SMG of FROGxAi14 mice at P20. Green arrow heads indicate adenophages, magenta arrow heads indicate tdTomato+ cells (indicating GM-CSF production). Green = CD226, Blue = MHCII, Red = tdTomato, White = DAPI. Scale: 20 µm. MFI = mean fluorescent intensity. CFU = colony forming units. Statistical values: ns > 0.05; * ≤ 0.05; ** ≤ 0.01; *** ≤ 0.005; **** ≤ 0.001.
Extended Data Fig. 4 Adenophages have a relationship with ILC2s and MECs.
A) Immunofluorescent histology image of the SMG of wildtype mice at P20. Images are representative for the imaging used to define location of adenophages within the tissue (see Fig. 4b). Left image shows staining for αSMA (green); right image shows staining for PDGFRα (green); both images: Magenta = CD226, Blue = MHCII and White = DAPI. Scale: 20 µm. B) Gating strategy for FACS-isolation of MECs. Images are representative for two independent experiments with two scanned sections each. C) Heatmap showing gene expression obtained by RNAseq of the indicated MEC samples Ctrl = Csf2+/−; KO = Csf2−/−. D) Volcanoplot showing DEGs between MECs isolated from the SG of Csf2+/− and Csf2−/− mice at P20. Statistical testing: Generalized Linear Models. E) Volcanoplots show identity and functionality signature genes (see Supplementary Table 2) in MECs isolated from the SG of Csf2+/− and Csf2−/− mice at P20. Statistical testing: Generalized Linear Models. F) PAS staining on SG of Csf2+/− and Csf2−/− mice at P20. G) Bar diagram shows the overall volume of saliva collected in 20 min from P20 Control and Csf2−/− mice. Data shows five independent experiments and is represented as mean ± SEM. H) Representative FACS plots showing the gating strategy of for adenophages in the LG (left) and MG (right). Pre-gated on singlets live CD45+CD90−Ly6G−SiglecF−Ly6C−CD11b+F4/80+. I) Immunofluorescent histology image of the P20 LG of wildtype mice. Green = αSMA, Magenta = CD226, Blue = MHCII, White = DAPI. Scale: 20 µm. Images are representative for one experiment with three scanned sections each. J) Reanalysis of the scRNAseq dataset on naïve mammary gland (Cansever, Petrova et al., 2023) highlighting the myeloid populations. The dot plot shows the expression of the adenophage signature genes across the myeloid populations. Colors indicate the average expression of each gene, circle sizes represent percentage of cells within a cluster expressing a gene. PAS = Periodic acid-Schiff stain; MEC = myoepithelial cell; DEGs = differentially expressed genes.
Extended Data Fig. 5 Adenophages are a conserved cell population across glands and species.
A) Representative FACS plot showing the expression of RELMα and CD226 in adenophages (CD45+CD90−SiglecF−Ly6G−Ly6C−CD11b+F4/80 + CX3CR1-MHCII + CD226 + ), MHCII+ macrophages (CD45+CD90−SiglecF−Ly6G−Ly6C−CD11b+F4/80 + CX3CR1 + MHCII), and DCs (CD45+CD90−SiglecF−Ly6G−Ly6C−CD11b+F4/80-CD11c+MHCII + ) in the SG of Csf2+/− control mice at P20. B) Heatmap showing marker expression of indicated cell populations in LG, MG and SG of Csf2+/− control mice at P20 measured by HDCyto. C) representative FACS plot showing expression of RELMα and CD226 in indicated cell populations of the MG of Csf2+/− control mice at P20. D) Gating for ILC2s in P20 SG of Csf2+/− mice. Pregated on live CD45 + CD11b− cells. E) Representative FACS plots for tdTomato+ and tdTomato+eGFP+ cells in LG (left) and MG (right) of FROGxAi14 mice at P20. Pre-gated on singlet live CD45+CD11b− cells. F) Immunofluorescent histology image of the P20 LG of FROGxAi14 mice. Green arrow heads indicate adenophages and magenta arrow heads indicate tdTomato+ cells indicating GM-CSF expression. Green = CD226, Blue = MHCII, Red = tdTomato, White = DAPI. Scale: 20 µm. Images are representative for one experiment with three scanned sections each. G) Proximity measurement of IBA1+CD226+ cells and IBA1+CD226− cells to acini (left) or ducts (right) as indicated in Fig. 4j in human SMG. N = 3; data are represented as mean + SD. Statistical test: unpaired t-test H) Graphical depiction of the niche-triad consisting of adenophages, GM-CSF producing ILC2s and MECs throughout exocrine glands. Graphics in H created using BioRender.com. LG = lacrimal gland; MG = mammary gland; SMG = submandibular salivary gland; MEC = myoepithelial cell. Statistical values: ns > 0.05; * ≤ 0.05; ** ≤ 0.01; *** ≤ 0.005; **** ≤ 0.001.
Supplementary information
Supplementary Table 4 (download XLSX )
Supplementary Table 4.
Supplementary Table 5 (download XLSX )
Supplementary Table 5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Westermann, F., Tuzlak, S., Kreiner, V. et al. Adenophages are an atypical macrophage population in exocrine glands sustained by ILC2-derived GM-CSF. Nat Immunol 27, 26–34 (2026). https://doi.org/10.1038/s41590-025-02356-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02356-8
This article is cited by
-
Macrophages conquer the glands
Nature Immunology (2026)