Abstract
Natural killer (NK) cells are innate immune effectors, but their cytotoxic potential can be compromised within the immunosuppressive tumor microenvironment. Identifying molecular mechanisms that underly this dysfunction is essential for advances in cancer immunotherapy. Here we show that CLEC12B, a C-type lectin receptor, functions as an inhibitory checkpoint that restricts NK cell-mediated antitumor immunity. High expression of CLEC12B by tumor-infiltrating NK cells correlates with poor clinical prognosis in patients with hepatocellular carcinoma. We identify lipoprotein lipase as a functional ligand for CLEC12B, triggering inhibitory signaling that suppresses NK cell activation. We developed a high-affinity nanobody as a potential therapeutic that disrupts the CLEC12B–lipoprotein lipase axis, thereby revitalizing NK cell activity and suppressing tumor progression in preclinical models. Furthermore, this nanobody has potent synergistic efficacy when combined with PD-1 blockade. These findings establish CLEC12B as a promising therapeutic target for rearming the immune system against solid malignancies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
scRNA-seq data from tumor-infiltrating immune cells from Hepa1-6 mouse models generated in this study have been deposited at the National Genomics Data Center under accession code CRA023427. Bulk RNA-seq data for NK cells with electroporation-mediated knockout of CLEC12B are available in the National Genomics Data Center under accession code HRA010572. All other data supporting the findings of this study are available within the article and Supplementary Information. Source data are provided with this paper.
Code availability
No unique or original code was developed in this study. All bioinformatic analyses, including scRNA-seq clustering and bulk RNA-seq differential expression analysis, were performed using standard, publicly available software and packages with default or previously described parameters. Detailed descriptions of the workflows and parameters used are provided in the Methods.
References
Lanier, L. L. Five decades of natural killer cell discovery. J. Exp. Med. 221, e20231222 (2024).
Narni-Mancinelli, E. & Vivier, E. Advancing natural killer therapies against cancer. Cell 185, 1451–1454 (2022).
Kyrysyuk, O. & Wucherpfennig, K. W. Designing cancer immunotherapies that engage T cells and NK cells. Annu. Rev. Immunol. 41, 17–38 (2023).
Sun, J. C. & Lanier, L. L. NK cell development, homeostasis and function: parallels with CD8+ T cells. Nat. Rev. Immunol. 11, 645–657 (2011).
Dean, I. et al. Rapid functional impairment of natural killer cells following tumor entry limits anti-tumor immunity. Nat. Commun. 15, 683 (2024).
Zhang, X. et al. IRF4 expression by NK precursors predetermines exhaustion of NK cells during tumor metastasis. Nat. Immunol. 26, 1062–1073 (2025).
Lupo, K. B. & Matosevic, S. CD155 immunoregulation as a target for natural killer cell immunotherapy in glioblastoma. J. Hematol. Oncol. 13, 76 (2020).
Sivori, S. et al. Human NK cells: surface receptors, inhibitory checkpoints, and translational applications. Cell. Mol. Immunol. 16, 430–441 (2019).
Wolf, N. K., Kissiov, D. U. & Raulet, D. H. Roles of natural killer cells in immunity to cancer, and applications to immunotherapy. Nat. Rev. Immunol. 23, 90–105 (2023).
Cao, Y. et al. Immune checkpoint molecules in natural killer cells as potential targets for cancer immunotherapy. Signal Transduct. Target. Ther. 5, 250 (2020).
Reis, E. S. C., Yamasaki, S. & Brown, G. D. Myeloid C-type lectin receptors in innate immune recognition. Immunity 57, 700–717 (2024).
Brown, G. D., Willment, J. A. & Whitehead, L. C-type lectins in immunity and homeostasis. Nat. Rev. Immunol. 18, 374–389 (2018).
Piersma, S. J. et al. Expression of a single inhibitory member of the Ly49 receptor family is sufficient to license NK cells for effector functions. Elife https://doi.org/10.7554/eLife.100218.3 (2025).
Babic, M. et al. NK cell receptor NKG2D enforces proinflammatory features and pathogenicity of Th1 and Th17 cells. J. Exp. Med. 217, e20190133 (2020).
Li, K. et al. The uric acid crystal receptor Clec12A potentiates type I interferon responses. Proc. Natl Acad. Sci. USA 116, 18544–18549 (2019).
Neumann, K. et al. Clec12a is an inhibitory receptor for uric acid crystals that regulates inflammation in response to cell death. Immunity 40, 389–399 (2014).
Hoffmann, S. C. et al. Identification of CLEC12B, an inhibitory receptor on myeloid cells. J. Biol. Chem. 282, 22370–22375 (2007).
Montaudie, H. et al. CLEC12B decreases melanoma proliferation by repressing signal transducer and activator of transcription 3. J. Invest. Dermatol. 142, 425–434 (2022).
Chi, D. et al. CLEC12B suppresses lung cancer progression by inducing SHP-1 expression and inactivating the PI3K/AKT signaling pathway. Exp. Cell. Res. 409, 112914 (2021).
Chen, C. H. et al. Dendritic-cell-associated C-type lectin 2 (DCAL-2) alters dendritic-cell maturation and cytokine production. Blood 107, 1459–1467 (2006).
Strack, R. Nanobodies made versatile. Nat. Methods 20, 37 (2023).
Ding, Z. et al. Nanobody-based trispecific T cell engager (Nb-TriTE) enhances therapeutic efficacy by overcoming tumor-mediated immunosuppression. J. Hematol. Oncol. 16, 115 (2023).
Zhang, F. et al. Structural basis of a novel PD-L1 nanobody for immune checkpoint blockade. Cell Discov. 3, 17004 (2017).
Bernard, N. J. CTLA-4 nanobody stops colitis. Nat. Immunol. 25, 190 (2024).
Blot, L. et al. CLEC12B regulates melanocyte immunity and homeostasis in the skin through the STAT1/IRF1 axis. J. Invest. Dermatol. 145, 2007–2020 (2025).
Tang, F. et al. A pan-cancer single-cell panorama of human natural killer cells. Cell 186, 4235–4251 (2023).
Ohs, I. et al. Restoration of natural killer cell antimetastatic activity by IL12 and checkpoint blockade. Cancer Res. 77, 7059–7071 (2017).
Liu, W. N. et al. Single-cell RNA sequencing reveals anti-tumor potency of CD56+ NK cells and CD8+ T cells in humanized mice via PD-1 and TIGIT co-targeting. Mol. Ther. 32, 3895–3914 (2024).
Hockemeyer, K. et al. The stress response regulator HSF1 modulates natural killer cell anti-tumour immunity. Nat. Cell Biol. 26, 1734–1744 (2024).
Goh, W. et al. IKAROS and AIOLOS directly regulate AP-1 transcriptional complexes and are essential for NK cell development. Nat. Immunol. 25, 240–255 (2024).
Duan, S. & Paulson, J. C. Siglecs as immune cell checkpoints in disease. Annu. Rev. Immunol. 38, 365–395 (2020).
Awwad, M. H. S. et al. Selective elimination of immunosuppressive T cells in patients with multiple myeloma. Leukemia 35, 2602–2615 (2021).
Zhang, Q. et al. Blockade of the checkpoint receptor TIGIT prevents NK cell exhaustion and elicits potent anti-tumor immunity. Nat. Immunol. 19, 723–732 (2018).
Zhang, H., Yang, L., Wang, T. & Li, Z. NK cell-based tumor immunotherapy. Bioact. Mater. 31, 63–86 (2024).
Chan, W. K. et al. Antibody-dependent cell-mediated cytotoxicity overcomes NK cell resistance in MLL-rearranged leukemia expressing inhibitory KIR ligands but not activating ligands. Clin. Cancer Res. 18, 6296–6305 (2012).
Kim, N., Yi, E., Lee, E., Park, H. J. & Kim, H. S. Interleukin-2 is required for NKp30-dependent NK cell cytotoxicity by preferentially regulating NKp30 expression. Front Immunol. 15, 1388018 (2024).
Deuse, T. et al. The SIRPα–CD47 immune checkpoint in NK cells. J. Exp. Med. 218, e20200839 (2021).
Porgador, A. Unveiling the relevance of immune checkpoints for innate and adaptive response to hepatocellular carcinoma using improved model of humanized mice. Mol. Ther. 32, 3761–3762 (2024).
Zhang, H. et al. Silencing of SIRPα enhances the antitumor efficacy of CAR-M in solid tumors. Cell. Mol. Immunol. 21, 1335–1349 (2024).
Ma, S. et al. Identification of a small-molecule Tim-3 inhibitor to potentiate T cell-mediated antitumor immunotherapy in preclinical mouse models. Sci. Transl. Med. 15, eadg6752 (2023).
Anania, C. Reprogramming-based gene therapy promotes anti-tumor immunity in vivo. Nat. Genet. 56, 1999 (2024).
Verhaar, E. R., Woodham, A. W. & Ploegh, H. L. Nanobodies in cancer. Semin. Immunol. 52, 101425 (2021).
Wu, S. Y., Wu, F. G. & Chen, X. Antibody-incorporated nanomedicines for cancer therapy. Adv. Mater. 34, e2109210 (2022).
Jiang, C. et al. Engineering a smart agent for enhanced immunotherapy effect by simultaneously blocking PD-L1 and CTLA-4. Adv. Sci. 8, e2102500 (2021).
Boukhaled, G. M. et al. Pre-encoded responsiveness to type I interferon in the peripheral immune system defines outcome of PD-1 blockade therapy. Nat. Immunol. 23, 1273–1283 (2022).
Chen, G. et al. Chemotherapy-induced neoantigen nanovaccines enhance checkpoint blockade cancer immunotherapy. ACS Nano 17, 18818–18831 (2023).
Tsao, L. C., Force, J. & Hartman, Z. C. Mechanisms of therapeutic antitumor monoclonal antibodies. Cancer Res. 81, 4641–4651 (2021).
Ma, L. et al. Multiregional single-cell dissection of tumor and immune cells reveals stable lock-and-key features in liver cancer. Nat. Commun. 13, 7533 (2022).
Ma, L. et al. Single-cell atlas of tumor cell evolution in response to therapy in hepatocellular carcinoma and intrahepatic cholangiocarcinoma. J. Hepatol. 75, 1397–1408 (2021).
Ma, L. et al. Tumor cell biodiversity drives microenvironmental reprogramming in liver cancer. Cancer Cell 36, 418–430 (2019).
Sharma, A. et al. Onco-fetal reprogramming of endothelial cells drives immunosuppressive macrophages in hepatocellular carcinoma. Cell 183, 377–394 (2020).
Losic, B. et al. Intratumoral heterogeneity and clonal evolution in liver cancer. Nat. Commun. 11, 291 (2020).
Zhu, G. Q. et al. CD36+ cancer-associated fibroblasts provide immunosuppressive microenvironment for hepatocellular carcinoma via secretion of macrophage migration inhibitory factor. Cell Discov. 9, 25 (2023).
Giraud, J. et al. THBS1(+) myeloid cells expand in SLD hepatocellular carcinoma and contribute to immunosuppression and unfavorable prognosis through TREM1. Cell Rep 43, 113773 (2024).
Liu, Z. et al. Viral-Track integrated single-cell RNA-sequencing reveals HBV lymphotropism and immunosuppressive microenvironment in HBV-associated hepatocellular carcinoma. Commun. Biol. 8, 1030 (2025).
Alvarez, M. et al. Human liver single nucleus and single cell RNA sequencing identify a hepatocellular carcinoma-associated cell-type affecting survival. Genome Med. 14, 50 (2022).
Darci-Maher, N. et al. Cross-tissue omics analysis discovers ten adipose genes encoding secreted proteins in obesity-related non-alcoholic fatty liver disease. EBioMedicine 92, 104620 (2023).
Lee, S. H. T. et al. Single nucleus RNA-sequencing integrated into risk variant colocalization discovers 17 cell-type-specific abdominal obesity genes for metabolic dysfunction-associated steatotic liver disease. EBioMedicine 106, 105232 (2024).
Netskar, H. et al. Pan-cancer profiling of tumor-infiltrating natural killer cells through transcriptional reference mapping. Nat. Immunol. 25, 1445–1459 (2024).
Bi, J. et al. Checkpoint TIPE2 limits the helper functions of NK cells in supporting antitumor CD8+ T cells. Adv. Sci. 10, e2207499 (2023).
Ma, H. et al. Hetero-bivalent nanobodies provide broad-spectrum protection against SARS-CoV-2 variants of concern including Omicron. Cell Res. 32, 831–842 (2022).
Yakkala, C. et al. Cryoablation does not significantly contribute to systemic effector immune responses in a poorly immunogenic B16F10 melanoma model. Clin. Cancer Res. 30, 4190–4200 (2024).
Wu, L. et al. Camouflaging attenuated Salmonella by cryo-shocked macrophages for tumor-targeted therapy. Signal Transduct. Target. Ther. 9, 14 (2024).
Zhao, L. et al. Antitumor efficacy and potential mechanism of FAP-targeted radioligand therapy combined with immune checkpoint blockade. Signal Transduct. Target. Ther. 9, 142 (2024).
Acknowledgements
This work was supported by the National Key R&D Program of China (2022YFA1303200), National Natural Science Foundation of China (82394452), Anhui Provincial Natural Science Foundation (2408085JX013), CAS Project for Young Scientists in Basic Research (grant no. YSBR-068), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0940000) and USTC Research Funds of the Double First-Class Initiative (YD9100002506).
Author information
Authors and Affiliations
Contributions
C.S. designed the research and wrote the manuscript. P.S. and X.X. contributed to manuscript preparation. P.S., X.X., B.H., K.Z., A.K., H.C., H.L., S.K. and Q.T. performed the experiments and analyzed the data. B.N. provided clinical expertise and guidance. C.S. supervised the project. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
P.S. and C.S. are coinventors on a pending patent application (CN117304319A) submitted by the USTC that covers the development of nanobodies targeting CLEC12B. The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Nick Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
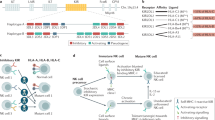
Extended Data Fig. 1 CLEC12B is predominantly expressed in NK and myeloid lineages and is selectively upregulated in tumor-infiltrating NK cells in HCC.
a, b, Single-cell transcriptomic landscape of the multicellular ecosystem in HCC and iCCA tumors (n = 189 patients). UMAP plot showing 7 major cell clusters (a) and heat map displaying the average expression of CLEC12B and PTPRC (CD45) across these clusters (b). c, Representative gating plots and expression of CLEC12B in CD45⁻ and CD45⁺ populations from subcutaneous Hepa1-6 tumors in wild-type mice (n = 7). d, Genotyping of the Clec12b−/− murine model by PCR analysis. The mutant allele (503 bp) and WT allele (824 bp) are indicated. e, f, Terminal tumor weights of Hepa1-6 (e) and MC38 (f) subcutaneous tumors in WT and Clec12b−/− mice at day 21 (n = 5). g, Gating strategy for CLEC12B expression in NK cells from murine splenic lymphocytes and tumor-infiltrating lymphocytes (left). Antibody specificity was validated using NK-specific knockout (Clec12bΔNK/ΔNK) mice, with WT mice as controls (n = 4; right). Full characterization of the Clec12bΔNK/ΔNK system is provided in Fig. 4 and Extended Data Fig. 3f. h–k, Comparison of CLEC12B expression in myeloid lineages from WT mice bearing Hepa1-6 tumors (n = 6). Representative plots (h, j) and statistical quantification of CLEC12B expression in macrophages (i) and neutrophils (k) from spleen and TILs are shown. l–q, High-resolution re-annotation of the HCC scRNA-seq dataset. UMAP visualization of major immune lineages (l), dot plot of lineage-specific markers (m), and feature plots showing the distribution of key marker genes (n). UMAP projection highlights CLEC12B expression, with the NK cell cluster marked in red (o). Bar graph quantifying the composition of the CLEC12B⁺ population (p) and violin plots comparing CLEC12B expression levels between myeloid and NK cells (q) are presented. Data in e–g, i, k and q are presented as mean ± s.e.m. Statistical significance was determined by unpaired two-tailed Student’s t-test for pairwise comparisons (e–g, i, k, q). Dates are representative of three independent experiments (c–k). Exact P values are indicated in the figures.
Extended Data Fig. 2 CLEC12B is selectively upregulated in tumor-infiltrating NK cells and positively correlates with immune checkpoint receptor expression in HCC.
a-d, Single-cell RNA-seq analysis showing that CLEC12B expression in NK cells is significantly higher in HCC tumor-infiltrating (IT, n = 17) compared to peritumoral tissue (PT, n = 17) unpaired samples with CLEC12B⁺ NK cells (a). Gene Ontology (GO) enrichment of pathways down-regulated vs. up-regulated in CLEC12B⁺ NK cells (from IT NK cell subset), derived by comparison of CLEC12B⁺ vs. CLEC12B⁻ NK cells (b). Analysis of TCGA HCC tumors (n = 374) and peritumoral tissues (n = 50) showing CLEC12B/CD56 transcript ratio is significantly higher in tumors compared to peritumor tissues (c). Correlation in the same TCGA HCC tumor cohort (n = 374) between normalized CLEC12B/CD56 expression and various immune-checkpoint receptor/CD56 ratios (d): TIGIT, PDCD1, HAVCR2, LAG3. e-g, Mass cytometry analysis of HCC patients (n = 19). Gating strategy (e). Numbers in the drawn gates indicate the cell percentages. Representative flow cytometry plots (f) and quantification (g) of CELC12B expression in CD3+ cells. h, Correlation between CLEC12B and the expression of LAG-3 inhibitory receptors was analyzed in the PT and IT NK cells. i, Heat map represented correlation between the expression level of CLEC12B and the expression levels of HACVR2, TIGIT, PDCD1, and LAG3 in LIHC (n = 371); Data in a, c, g are presented as mean ± s.e.m. Data in (d, h) is shown with shaded areas representing 95% confidence intervals. Box-and-whisker plots show the median (centre line), the interquartile range (25th–75th percentiles), and whiskers extending from the minimum to the maximum values (a), all individual data points are shown. Statistical significance was determined by unpaired two-tailed Student’s t-test (a, c) or paired two-tailed Student’s t-test (g) two-sided correlation analysis (d, h).Exact P values are indicated in the figures.
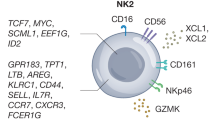
Extended Data Fig. 3 Enrichment analysis of genes and pathways related to NK cell effector function, cell migration, and adhesion in CLEC12B knockout NK.
a, Correlation analysis between CLEC12B expression levels and the infiltration magnitude of NK cells in LIHC (n = 371), as quantified by the MCP-counter algorithm. b-d, Single-cell sequencing of CD45+ TILs in subcutaneous Hepa1-6 tumors of WT and Clec12b⁻/⁻ mice. Mean average plot of DEGs (P < 0.05) the NK cells in the scRNA-seq dataset (b) (graph shows fold change in expression (log2 (Clec12b⁻/⁻/WT)). Distribution of two types NK cells (NK1 and NK2) over the UMAP space (c). Systematic expression comparison (d) of top DEGs (avg_log2FC > 0.5, P < 0.05) among NK1 and NK2 clusters.e, PCR genotyping for the Clec12bΔNK/ΔNK mice. Successful construction was confirmed by the detection of homozygous Clec12bflox/flox (434 bp) and Ncr1iCre/+ (230 bp) alleles. f, Representative images of MC38 tumors harvested from Clec12bWT/WT and Clec12bΔNK/ΔNK mice at the experimental endpoint (n = 5 per group). g, Bone marrow cells were harvested from Clec12bWT/WT and Clec12bΔNK/ΔN (n = 6 per group) under naïve conditions or following subcutaneous Hepa1-6 tumor challenge (5 × 10⁶ cells; tumors grown to ~100–150 mm³). NK cells (CD3⁻NK1.1⁺) were stratified into maturation subsets based on the CD11b/CD27 axis (CD11b-CD27 + , CD11b + CD27 + , and CD11b + CD27-). The frequency of NK cells expressing education markers (Ly49C/H/I, CD43) and inhibitory checkpoints (NKG2A, KLRG1, TIGIT, and PD-1) was quantified by flow cytometry. h, Pathway enrichment analysis of DEGs between CLEC12B-knockout and control NK cells following electroporation-mediated CRISPR/Cas9 editing. Data in g are presented as mean ± s.e.m. Statistical significance was determined by Wilcoxon rank-sum test for single-cell differential expression analysis (b) two-way ANOVA followed by Sidak’s multiple comparisons test (g) or simple linear regression for correlation analysis (a). Dates are representative of three independent experiments (e-h). Exact P values are indicated in the figures. Illustrations in g created in BioRender; Sun, P. https://biorender.com/6odbjft (2026).
Extended Data Fig. 4 Purification of hCLEC12B antigen and H6-Fc nanobody.
a, Representative chromatogram of the purification of hCLEC12B extracellular domain (ECD) fused with an Fc tag, expressed in the HEK293F mammalian system and purified via the AKTA-pure system. b, SDS-PAGE analysis of the hCLEC12B-Fc protein before and after enzymatic digestion. The hCLEC12B-Fc fusion protein (lane 2) and the cleaved hCLEC12B protein (lane 3) migrated at approximately 70 kDa and 50 kDa, respectively. c, Size-exclusion chromatography (SEC) profile to verify the purity and monomeric state of the enzymatically digested hCLEC12B protein. d, Purification profile of the H6-Fc nanobody from the HEK293F system using the AKTA-pure system.e, Flow cytometry gating strategy for assessing the binding affinity of H6-Fc nanobody to the surface of CLEC12B-overexpressing HEK293T cells. f, Dose–response binding curves of nanobody clones (A1-Fc, E2-Fc, and H6-Fc) to immobilized mCLEC12B-ECD. EC50 values were determined using a four-parameter logistic regression model. g, ELISA analysis assessing the cross-reactivity of indicated nanobodies with hCLEC12B and hCLEC12A. h, Molecular weight determination of purified CLEC12B via SEC. A calibration curve was generated using standard markers, including tyrosine (181 Da), cytochrome c (12,365 Da), BSA (66,430 Da), and IgG (150,000 Da), with detection at UV 280 nm. Data in f, g is presented as mean ± s.e.m. Dates are representative of three independent experiments (a-h). Exact P values are indicated in the figures.
Extended Data Fig. 5 Anti-CLEC12B nanobody promotes production of granzyme B and perforin, and enhances NK cell proliferation.
a-d, NK cells were co-cultured with K562 target cells at 1:1 and 1:2 E:T ratios for 4 hours in the presence of H6-Fc, H6, or IgG control at 0, 5, 10, or 100 nM. Target cell death was assessed by CFSE/7-AAD staining and flow cytometry. Representative flow cytometry plots (a, c) and quantification (b, d). Dates are representative of the mean ± SEM. from triplicate wells in three independent experiments. e, f, Functional evaluation of murine splenic NK cells co-cultured with Hepa1-6 tumor cells (1:1 ratio) for 12 h. Cells were treated with H6-Fc (0.1 or 1 µg/mL) or control. Representative flow cytometry plots (e) and quantification (f) of CD107a, Granzyme B, and IFN-γ are presented.g-h, Proliferation analysis of human PBMC-derived NK cells following 24 h stimulation with indicated concentrations of H6-Fc (0, 10, or 100 nM; n = 3). Representative CFSE histograms (g) and statistical analysis (h) are shown. i-j, Cytotoxicity assay of MV4-11 cells co-cultured with human NK cells (1:1 ratio) for 4 h in the presence of H6-Fc (0, 10, or 100 nM; n = 3). Representative plots (i) and quantification of cell lysis percentage (j) are presented. Data in b, d, f, h, j are presented as mean ± s.e.m. Statistical significance was determined by two-way ANOVA followed by Sidak’s multiple comparisons test (b, d) or ordinary one-way ANOVA followed by Tukey’s multiple comparisons test (f, h, j). Dates are representative of three independent experiments (a–j). Exact P values are indicated in the figures.
Extended Data Fig. 6 Anti-CLEC12B treatment alone prolong the survival of tumor-bearing mice.
a, Survival analysis of C57BL/6 J mice bearing Hepa1-6 tumors. C57BL/6 J mice were subcutaneously injected with Hepa1-6 cells and received intraperitoneal injections of H6-Fc nanobody (10 mg/kg) every 2 days for five treatments (n = 10). b–e, Individual tumor growth curves in Hape1-6 tumor model. (n = 6) Vehicle (b), H6-Fc (c), αPD-1 (d), and αPD-1 + H6-Fc combination (e) f–i, BALB/c mice were subcutaneously injected with H22 cells and treated with Vehicle, H6-Fc (10 mg/kg), anti-PD-1 (10 mg/kg), or a combination (5 mg/kg each) every 2 days for five treatments (n = 6). Schematic of the therapeutic protocol (f), representative tumor images (g, n = 4), terminal tumor weight statistics (h), and tumor growth curves (i) are presented. j, Survival analysis of BALB/c mice bearing H22 tumors. BALB/c mice were subcutaneously injected with H22 cells and received intraperitoneal injections of H6-Fc nanobody (10 mg/kg) every 2 days for five treatments (n = 10). k–n, Individual tumor growth curves (n = 6) in H22 tumor model. Vehicle (b), H6-Fc (c), αPD-1 (d), and αPD-1 + H6-Fc combination (e). o, Survival analysis of C57BL/6 J mice bearing MC38 tumors. C57BL/6 J mice were subcutaneously injected with MC38 cells and received intraperitoneal injections of H6-Fc nanobody (10 mg/kg) every 2 days for five treatments (n = 10). p–q, C57BL/6 J mice were subcutaneously injected with MC38 cells and treated with Vehicle, H6-Fc (10 mg/kg), anti-PD-1 (10 mg/kg), or a combination (5 mg/kg each) every 2 days for five treatments (n = 6). Schematic of the therapeutic protocol (p) and tumor growth curves (q) are presented. r–u, Individual tumor growth curves in MC38 tumor model. (n = 6) Vehicle (r), H6-Fc (s), αPD-1 (t), and αPD-1 + H6-Fc combination (u). v, Survival analysis of C57BL/6 J mice bearing C1498 tumors. C57BL/6 J mice were subcutaneously injected with Hepa1-6 cells and received intraperitoneal injections of H6-Fc nanobody (10 mg/kg) every 2 days for five treatments (n = 10). w–z, Individual tumor growth curves in C1498 tumor model. (n = 6) Vehicle (w), Panobinostat (x), H6-Fc (y), and Panobinostat + H6-Fc combination (z). Data in h, i, q are presented as mean ± s.e.m. Data in (a, j, o, v) is shown with shaded areas representing 95% confidence intervals Statistical significance was determined by log-rank (Mantel-Cox) test (a, j, o, v); two-way ANOVA followed by Sidak’s multiple comparisons test (i, q); or ordinary one-way ANOVA followed by Tukey’s multiple comparisons test (h). Dates are representative of two (a, j, o, v). or three independent experiments (b–e, g–i, k–n, p–u, w–z). Exact P values are indicated in the figures. Illustrations in f and p created in BioRender; Sun, P. https://biorender.com/5ulyhnb (2026).
Extended Data Fig. 7 Low, medium, and high doses of CLEC12B nanobody do not induce adverse events in mice.
a, Comparison of surface (non-permeabilized) versus total (permeabilized) LPL expression across multiple cell lines to confirm specific surface presentation measurements. b, Quantification of surface LPL expression across multiple tumor cell lines (Hepa1-6, MC38, B16-F10, HepG2, K562, THP-1) with heparin displacement validation. c, Surface plasmon resonance analysis of the binding kinetics between immobilized CLEC12B-Fc and purified human GPC1 protein. Kinetic constants and equilibrium dissociation constant (K_D) were determined using a 1:1 Langmuir binding model. d, Representative flow cytometry plots showing the purity of NK cells expanded from human PBMCs. e–h, In vivo toxicology evaluation of H6-Fc nanobody in mice (n = 5). Serum levels of ALT (e) and AST (f) were measured at day 28 post-treatment. Weights of major organs, including heart, liver, spleen, lung, and kidneys, were recorded at the experimental endpoint (g). Representative H&E staining images of these organs harvested from Vehicle and H6-Fc treated mice (h). i, j, Immunohistochemical analysis of CLEC12B expression on tumor tissue microarrays from Thyroid cancer (i; n = 26) and Endometrial cancer (j; PT n = 26, IT n = 34). Representative images show lymphocyte aggregation areas at 20× (left) and 40× (right) magnification, with positive staining indicated by brown coloration. Statistical analysis of CLEC12B positivity in random fields of view is provided. Data in a, b, e–g, i, j are presented as mean ± s.e.m. Statistical significance was determined by ordinary one-way ANOVA followed by Tukey’s multiple comparisons test (e, f, g), or unpaired two-tailed Student’s t-test for pairwise comparisons between two independent groups (a, b, i, j). Dates are representative of two (d) or three independent experiments (a–c, e–h). Exact P values are indicated in the figures.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–6 and Methods.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download PDF )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, P., Xu, X., Hu, B. et al. Targeting NK cell CLEC12B enhances cancer immunotherapy. Nat Immunol (2026). https://doi.org/10.1038/s41590-026-02471-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41590-026-02471-0