Abstract
Natural killer (NK) cells and T cells need to infiltrate solid tumors to eradicate them. Here we show programmable mechanisms that can mobilize NK and T cells to solid tumors using metabolite-sensing receptors. In vivo and in vitro CRISPR activation screens using NK-92 cells identified GPR183, GPR84, GPR34 and GPR18 as top enhancers of infiltration and chemotaxis to breast and ovarian cancers. While endogenously expressed in restricted cellular contexts, expressing these receptors in NK and T cells drives migration to factors released by cancer cells and alters the NK cell transcriptome in a ligand-dependent manner. Expressing GPR183 in NK, chimeric antigen receptor (CAR) NK and CAR T cells increased tumor infiltration and control. Likewise, expressing GPR183 in mouse T cells increased tumor eradication in immunocompetent mice. These data show that metabolite sensing can be rewired to obtain biochemically guided spatially targeted cells, creating new possibilities for therapeutic intervention.
Similar content being viewed by others
Main
Mapping the logic of immune cell migration and identifying mechanisms to redirect immune cells to specific sites in the body and within a target tissue are critical objectives, as the location of immune cells dictates whether and how these cells elicit their effects. Although immunotherapies and cell therapies can be exceptionally effective1, insufficient recruitment and infiltration of cytotoxic lymphocytes to solid tumors still limit the efficacy of these treatments2,3, underscoring the need and potential of ‘cell delivery’ solutions.
Immune cell migration and infiltration rely on chemical, mechanical and electrical signals4. The main genes thought to orchestrate immune cell recruitment to solid tumors are chemokines and their cognate receptors expressed by immune cells, as well as adhesion proteins, such as selectins and integrins, that mediate transmigration and infiltration5,6. Yet, while these protein–protein interactions have been extensively studied, our ability to mobilize immune cells to specific sites and into solid tumors remains limited.
Forward genetic screens offer a powerful framework to optimize different cell functions of interest via directed evolution, such that genetic variation is induced (for example, via CRISPR or other tools) and a pool of genetically modified cells undergoes selection to identify the cells that are optimal at performing a ‘task’ of interest. Gain-of-function screens are particularly useful for studying and leveraging gene functions outside of their endogenous cellular contexts. Yet, while CRISPR knockout7,8, interference and activation9 as well as open reading frame (ORF)10 and short hairpin RNA11 screens have been performed to study immune cell functions, to date, gain-of-function screens have not been used to study tumor infiltration.
Here, we report programmable mechanisms to mobilize cytotoxic lymphocytes to solid tumors based on the activation of metabolite sensing and orphan G-protein-coupled receptors (GPCRs), identified in an array of in vivo and in vitro gain-of-function screens. As we show, this information can be used to design more efficacious biochemically guided therapeutic cells.
Results
Metabolite-sensing GPCRs are tumor infiltration enhancers
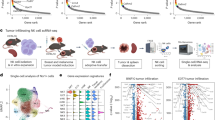
To identify mechanisms to mobilize natural killer (NK) cells to solid tumors, we conducted tumor infiltration in vivo CRISPR activation (CRISPRa) screens in NK cells (Fig. 1a) using breast and ovarian cancer in vivo models and a clinically applicable human NK cell line12 (NK-92) that has been widely used for chimeric antigen receptor (CAR) NK studies and in clinical trials13,14. To power the screens, we first identified candidate tumor infiltration enhancers based on single-cell RNA-sequencing (scRNA-seq) data from hundreds of individuals with cancer15,16.
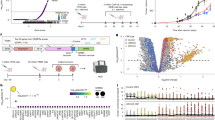
a, Experimental scheme of in vivo CRISPRa screens; i.v., intravenous; i.p., intraperitoneal; s.c., subcutaneous. b, Uniform manifold approximation and projection (UMAP) of scRNA-seq data from individuals with breast cancer. Each dot corresponds to a cell, colored by cell type (left), sample type (middle) and the overall expression of the genes identified as overexpressed in the immune cells residing in the tumor compared to blood samples (right; Methods); ILC, innate lymphoid cell; DC, dendritic cell. c, UMAP of NK scRNA-seq data from individuals with breast cancer16. Each dot corresponds to an NK cell, colored by NK subtype (left), sample type (middle) and overall expression of the NK pan-cancer tumor infiltration signature (right; Methods); Br, CD56bright; Di, CD56dim; Lo, CD16low; Hi, CD16high. d, CRISPRa breast cancer tumor infiltration screen results are shown as the significance (MAGeCK P values combined via Fisher’s method with BH correction for multiple hypothesis testing, y axis, Methods) and log-transformed fold change (FC, x axis) of each target gene (dot) in the tumor compared to the lung samples obtained from the same mouse based on MAGeCK22. Target genes whose sgRNAs were significantly or not significantly enriched in the tumor compared to lung samples are colored in red and gray, respectively; FC, fold change; NS, not significant. e, CRISPRa ovarian cancer tumor infiltration screen results shown as in d, here showing animal-matched comparison of ovarian cancer tumors to ascites samples. f, Top, log-transformed fold change of sgRNAs in breast cancer tumor versus animal-matched lung samples shown for sgRNAs targeting the indicated genes in the breast cancer in vivo screen shown in d. Bottom, log-transformed fold change of sgRNAs in ovarian cancer tumor versus animal-matched ascites samples shown for sgRNAs targeting indicated genes in the ovarian cancer in vivo screen shown in e. g, Large-scale (more than 5,500 genes) CRISPRa ovarian cancer tumor infiltration screen, showing enrichment in the tumor (MAGeCK P values, y axis) when comparing tumor to animal-matched ascites samples (left, matched per mouse) or tumor to the NK-92 cell library used (right). Panel a created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
First, we sought to identify genes that facilitate or drive the recruitment of other (non-NK) immune cells to solid tumors, postulating that such genes may elicit similar effects in NK cells if ectopically expressed. We analyzed scRNA-seq data collected from 22 individuals with breast cancer with matched tumor and peripheral blood samples15. For each immune cell type with sufficient representation in the data, we identified genes that were significantly overexpressed in the tumor compared to blood samples (Benjamini–Hochberg (BH) false discovery rate (FDR) < 0.05, log fold change > 0.1, model-based analysis of single-cell transcriptomics (MAST) test17; Fig. 1b, Extended Data Fig. 1a and Supplementary Table 1a).
Second, we analyzed a pan-cancer NK cell atlas from over 700 individuals and 24 cancer types16 and identified tumor-enriched genes per cancer type (that is, differentially expressed in the tumor compared to blood samples per cancer type; FDR < 0.01, edgeR18 differential expression test; Supplementary Table 1b). We defined a pan-cancer tumor infiltration NK gene signature, consisting of 320 genes that are tumor enriched in at least ten cancer types (Fig. 1c, Methods and Supplementary Information), showing a significant yet partial overlap with the tumor infiltration genes identified in other immune cell types.
Using this information, we generated a library of 1,070 CRISPRa single guide RNAs (sgRNAs) targeting a total of 256 genes (4 sgRNAs per target gene and 46 nontargeting control (NTC) sgRNAs), including both candidate genes identified in patients as well as genes with well-established roles in NK biology or immune chemoattraction and invasion, as proteases, adhesion proteins, integrins and chemokine receptors (Extended Data Fig. 1a and Supplementary Table 1). We transduced NK-92 cells to stably express the CRISPR–dCas9-synergistic activation mediator (SAM) system19, confirmed sgRNA-based gene overexpression at the protein level via flow cytometry (Extended Data Fig. 1b) and transduced CRISPR–dCas9-SAM NK-92 cells with the sgRNA library (Supplementary Table 1c).
In the breast cancer setting, we intravenously administered the NK-92 cell library to female mice with subcutaneous or orthotopic triple-negative breast cancer xenografts (MDA-MB-231). Mice were either non-obese diabetic scid gamma (NSG) mice or transgenic NOG mice that expressed the human IL2 gene (hIL-2-NOG), which better retain NK cells. Twenty four hours after the NK cell injections, we collected both tumors and lungs, as transferred tumor-reactive lymphocytes often accumulate in the lungs, decreasing tumor control and resulting in severe lung toxicities20,21. We processed and sequenced all samples together, including the tumors, lungs and NK-92 cell library used for the injections (Fig. 1a).
In the ovarian cancer setting, we regenerated the NK-92 cell library and intraperitoneally injected it into NSG female mice carrying intraperitoneal ovarian cancer TYK-nu xenografts. Intraperitoneal injections are a standard administration route in ovarian cancer clinical management adopted for ovarian cancer CAR T cell clinical trials (for example, NCT06646627). In this model, cancer cells spread and form tumors at multiple sites in the abdominal cavity, which are identified and collected using fluorescence-guided tumor resection. One to 3 weeks after NK cell injections, we collected tumors and intraperitoneal ascites fluid, which we then processed and sequenced together with the NK-92 cell library used for the injections (Fig. 1a).
Animal-matched statistical analyses identified gene activations that resulted in significant and reproducible tumor enrichment in each model, both compared to the NK-92 cell library and compared to the reference site (lung and ascites for breast and ovarian cancer, respectively, using MAGeCK22; Methods and Fig. 1d–f). Eight genes (GPR183, GPR84, GPR34, GPR18, LPAR2, FPR3, C5AR1 and CXCR2) were consistently ranked as the top hits (BH FDR < 1 × 10−7, Fisher test) in all breast cancer models (Fig. 1d,f, Extended Data Fig. 2 and Supplementary Table 2) and are hence referred to as tumor-homing GPCRs (thGPRs). Seven genes were identified as tumor infiltration hits in the ovarian cancer screen, including the integrin ITGAX and six of the eight thGPRs identified in the breast cancer screens (GPR183, GPR84, GPR34, GPR18, FPR3 and C5AR1; Fig. 1e,f and Supplementary Table 3a–d).
To further examine these hits, we performed a larger tumor infiltration CRISPRa NK-92 screen spanning more than 5,500 genes in the ovarian cancer model (Fig. 1a,g and Supplementary Table 3e–g). Once again GPR183 and GPR84 were identified as top hits, together with other genes with a well-established role in immune cell migration (for example, ITGB1, EPHA6 and SLMAF1; Fig. 1g). While the focused screens included only 12 chemokine receptors, this screen included 21, yet still only CXCR2 was identified as an enhancer of tumor infiltration, indicating that chemokine receptors may already be sufficiently active to support these processes.
Collectively, the screens repeatedly identified a unique subset of GPCRs as enhancers of NK-92 cell migration to breast and ovarian cancer tumors in mice, demonstrating the role of bioactive metabolites in immune cell recruitment and the potential of harnessing the cognate receptors to mobilize immune cells to solid tumors.
thGPRs enhance NK cell migration to factors released by cancer cells
thGPRs are GPCRs coupled to Gi/Go proteins23,24,25,26,27,28,29,30,31,32, a primary signal transducer for most conventional chemokine receptors, with GPR18 also reported to be Gq/G11 and Gs coupled23,33 and LPAR2 also shown to be Gq/G11 and G12/G13 coupled34,35 (Supplementary Table 4). Yet, unlike chemokine receptors, GPR183, GPR84, GPR34, GPR18 and LPAR2 have been reported to bind and respond to bioactive lipids, including phospholipids, fatty acids and cholesterol derivatives (Supplementary Table 4)23,24,25,26,27,28.
Using the PRESTO-Tango36 reporter lines, where the conformational change of a GPCR in response to its agonist ligand is converted to a bioluminescence signal (Extended Data Fig. 3a), we confirmed that 7α,25-dihydroxycholesterol (7α,25-OHC)25, lysophosphatidylserine (LysoPS)27,37 and 6-OAU26 are the agonists of GPR183, GPR34 and GPR84, respectively (Extended Data Fig. 3b). Isogenic NK-92 lines that constitutively express top hits via CRISPRa or ORFs show GPR183-, GPR34-, GPR84-, C5AR1- and CXCR2-dependent migration to 7α,25-OHC, LysoPS, 6-OAU, C5a and interleukin-8 (IL-8), respectively (Extended Data Fig. 3c–f). These isogenic lines also confirm CRISPRa specificity, such that, although GPR183 and GPR18 are located less than 40 kilobases (kb) apart, GPR183 is overexpressed in GPR183A+ (that is, cells with GPR183 CRISPRa) but not GPR18A+ cells, whereas GPR18 is overexpressed in GPR18A+ but not GPR183A+ cells (Extended Data Fig. 3d,e). Combinatorial expression of GPR183 and CXCR2 led to a threefold increase in NK-92 cell migration to a combination of their ligands compared to expression of only GPR183 and only CXCR2, indicative of synergistic effects (Extended Data Fig. 3g,h).
thGPRs highlight the importance of immune cell tumor recruitment via chemoattracting metabolites. To examine if these metabolites are released from cancer cells, we performed two additional in vitro screens (Fig. 2a). In the first screen, we used a transwell assay to identify perturbations that enhance chemotaxis to factors released from breast cancer cells (MCF7 and MDA-MB-231; Fig. 2a,b). In the second screen, we cocultured NK cells with breast cancer (MCF7) spheroids and selected for NK cells that both migrated to and infiltrated the spheroids within 6, 24 and 48 h (Fig. 2c and Extended Data Fig. 4a). All eight thGPRs were among the top ten hits in both the chemotaxis and spheroid in vitro screens (Fig. 2b,c, Extended Data Fig. 4b–d and Supplementary Table 5).
a–c, Chemotaxis and spheroid infiltration CRISPRa screens in NK-92 cells. a, Experimental scheme. b,c, Screen results shown as the significance (MAGeCK P values combined via Fisher’s method with BH correction for multiple hypothesis testing, y axis) and fold change (x axis) of each target gene (dot). Positive and negative values denote enrichment and depletion, respectively, in the NK cell population that migrated to the breast cancer supernatant (b) and infiltrated into breast cancer spheroids (c) compared to the NK-92 cell library. d, Differential expression of top hits (rows) in different immune cell types (columns) when comparing tumor and blood samples of individuals with breast cancer based on scRNA-seq data15 (MAST17; Methods). e, Differential expression of top hits (rows) in NK cells when comparing tumor and blood samples in different cancer types (column) based on the pan-cancer NK single-cell atlas16 (edgeR18 with BH correction for multiple hypothesis testing; Methods). f, UMAP of NK cells in tumor and blood samples from individuals with breast cancer (matching Fig. 1c), with cells colored based on the expression of GPR183 and genes positively (IL7R and CCR7) and negatively (PRF1 and FCGR3A encoding CD16) correlated with GPR183, marking early and late stages of NK cell maturation, respectively. Expression values are shown as log1p -transformed transcripts per million (TPM). g, Expression levels of top hits and top GPR183 program genes across NK cells from breast cancer tumor and blood samples, stratified by NK cell subtype. h, GPR183 is positively and negatively associated with early and late stages of lymphocyte maturation from precursor/naive to cytotoxic effector cells both in NK and CD8+ T cells. Spearman correlation coefficients (rs) of the expression of single genes (dots) with the expression of GPR183 in NK16 (x axis) and CD8+ T cells15 in breast cancer tumors are shown. GPR183 coexpression patterns in the two cell types were correlated (rs = 0.29, P < 1 × 10−20); cDC2, conventional type 2 dendritic cells; pDC, plasmacytoid DC; ALL, acute lymphoblastic leukemia; BRCA, breast cancer; CLL, chronic lymphocytic leukemia; CRC, colorectal cancer; ESCA, esophageal cancer; GC, gastric cancer; HCC, hepatocellular carcinoma; HNSCC, head and neck squamous cell carcinoma; LC, lung cancer; MELA, melanoma; MM, multiple myeloma; NB, neuroblastoma; NPC, nasopharyngeal cancer; OV, ovarian cancer; PACA, pancreatic cancer; PRAD, prostate cancer; RC, renal carcinoma; THCA, thyroid carcinoma; UCEC, uterine corpus endometrial carcinoma. Panel a created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
The chemotaxis screen identified several other hits (Fig. 2b), including the chemokine receptor CXCR3, EPHA1 (known for its role in NK cell migration38) and P2RY10, which was also identified in the ovarian cancer in vivo screen (Fig. 1e) and encodes another LysoPS receptor, similar to the thGPR GPR34. Although conducted with chemically inert transwell inserts, not coated with any integrin ligands, the chemotaxis screen also identified multiple integrins (ITGAD, ITGAL, ITGAX and ITGAM) as hits, potentially due to integrin ligands expressed by NK-92 cells (for example, ICAM1 and ICAM2, which are expressed by >40% of NK-92 cells based on scRNA-seq data). The only other hits identified in the spheroid screen were ICOS and ITGAL.
thGPRs mark specific lineages of tumor-infiltrating cells
In humans, thGPRs are associated with tumor infiltration (Fig. 2d,e, Extended Data Fig. 5 and Supplementary Table 1a,b), but their expression is restricted to specific cell types, lineages or maturation stages. Four of the eight thGPRs (GPR84, GPR34, FPR3 and C5AR1) are primarily expressed by myeloid cells and especially macrophages (Extended Data Fig. 5a,c,d,g,h), potentially explaining the relatively high abundance of these cells in solid tumors39. GPR183 and GPR18 are included in the NK pan-cancer tumor infiltration signature (Fig. 2e and Supplementary Table 1b), and GPR34 is tumor enriched in nine cancer types. CXCR2, however, is tumor depleted in NK cells (that is, significantly underexpressed in the tumor compared to blood) in nine different cancer types (FDR < 1 × 10−10, edgeR; Fig. 2e and Supplementary Table 1b) but is associated with tumor infiltration in monocytes (Fig. 2d and Supplementary Table 1a).
Based on the NK pan-cancer atlas16, GPR34, GPR84, LPAR2 and FPR3 are lowly expressed in primary NK cells (detected in <2% of NK cells; Extended Data Fig. 6a,b), GPR18 is expressed sporadically, and CXCR2 is lowly expressed in early stages of NK cell maturation (Extended Data Fig. 6a). By contrast, GPR183 is primarily expressed in immature NK cells (CD56brightCD16lo c4-IL-7R, c2-IL-7R-RGS1lo and CD56brightCD16lo-c5-CREM cell clusters; Figs. 1c, 2f,g). Moreover, in both NK cells and CD8+ T cells, GPR183 is upregulated in tumor-infiltrating lymphocytes and early developmental stages and downregulated in cytotoxic stages. In both cell types, it is coexpressed with IL7R, CCR7, GZMK, SELL and CD44 and inversely correlated with PRF1, FCGR3A (CD16), NKG7 and granzymes (Fig. 2g,h). GPR183 is also strongly associated with CXCR4, which may sequester GPR183hi T/NK cells to CXCL12-producing cancer associated fibroblasts, as previously reported in ovarian cancer40, further underscoring the potential of decoupling GPR183 from its endogenous regulation.
GPR183 is an oxysterol-responsive receptor broadly expressed across immune cells, including naive B cells25, naive CD4+41 cells, innate lymphoid cells42 and dendritic cells43. As it was identified as a tumor infiltration hit in all of our in vivo and in vitro screens and ranked among the top five most tumor-enriched GPCRs in our pan-cancer scRNA-seq analyses, we focused on GPR183 to further investigate thGPR-based mechanisms and their therapeutic potential.
GPR183 affects the NK cell transcriptome in a ligand-dependent manner
GPCR signaling is known to impact transcriptional regulation. To investigate how thGPRs impact the NK cell transcriptome, we performed a Perturb-seq44,45 screen, combining scRNA-seq with matched sgRNA sequencing in NK-92 cells with activation of one of seven thGPRs (C5AR1, CXCR2, GPR18, GPR183, GPR34, GPR84 and LPAR2; three sgRNAs per hit and five NTCs; Fig. 3a). To test for ligand-dependent GPCR transcriptional effects, we also profiled the cells after a 4-h exposure to the GPR183 ligand 7α,25-OHC (100 nM).
a–d, NK-92 Perturb-seq screen of top hits. a, Schematics of Perturb-seq workflow. b, UMAP of Perturb-seq data: each dot corresponds to an NK-92 cell colored by the sgRNA target gene (left) and cell culture conditions (right); n denotes the number of cells. c, Number of differentially expressed genes (DEGs) identified in response to different CRISPRa perturbations (x axis) in NK-92 cells with and without 7α,25-OHC treatment. d, Overall expression of the GPR183 CRISPRa signature across NK-92 cells stratified by CRISPRa target gene and culture conditions; n denotes the number of cells in each category; for the box plots, the middle lines indicate the median, box edges indicate the 25th and 75th percentiles, whiskers denote the most extreme points that do not exceed ±IQR ×1.5, and further outliers are marked individually with circles (minima/maxima). e, Expression of the top 100 genes in the GPR183 CRISPRa signature (rows) across NK-92 cells stratified by CRISPRa target gene and culture conditions (columns). f, Expression of genes involved in steroid and cholesterol synthesis and regulation that are differentially expressed in NK-92 cells in response to 7α,25-OHC treatment. These effects are independent of GPR183. Panel a created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
The Perturb-seq data confirmed on-target gene activation and demonstrated GPR183-ligand-dependent transcriptional remodeling (Fig. 3b–e and Supplementary Table 6). The level of on-target gene activation varied by sgRNA and showed a high degree of specificity, such that sgRNA targeting one hit did not activate other hits (Extended Data Fig. 7a,b) or any other gene located within 200 kb of the sgRNA target gene. With 7α,25-OHC treatment, GPR183 CRISPRa had a significant impact on the cell transcriptome, resulting in differential expression of 393 genes compared to control cells (Fig. 3c–e; defined as the ‘GPR183 CRISPRa signature’). These effects were diminished in the absence of 7α,25-OHC treatment, with only 13 differentially expressed genes (Fig. 3c). The GPR183 CRISPRa signature was not triggered by other perturbations (Fig. 3d,e) and was enriched with up- and downregulation of genes positively and negatively correlated with GPR183 in the pan-cancer NK cell atlas, respectively (P < 8 × 10−5, hypergeometric test; Supplementary Table 6a,i,j), including upregulation of CD44 and CD82 and downregulation of NKG7 and CST7 (Figs. 2h and 3e), indicating that GPR183 might regulate these central genes. The signature is also enriched for upregulation of adhesion (for example, ADK, CCDC88B, CD74, CLASP2 and ECM2) and membrane organization genes (for example, DOCK2 and ELMO1) and downregulation of metabolic genes (for example, GLUL, H2AFZ and LGALS1; P < 1 × 10−5, hypergeometric test; Supplementary Table 6a,i,j).
C5AR1 CRISPRa had a significant impact on the NK-92 cell transcriptome with and without 7α,25-OHC treatment. C5a, the ligand of C5AR1, is formed from the cleavage of the complement protein C5, which is expressed by 7.29% of NK-92 cells based on our Perturb-seq data. The C5AR1 CRISPRa effect was more pronounced with 7α,25-OHC treatment (Fig. 3c and Extended Data Fig. 7c), involving upregulation of antiapoptotic and immune defense response genes (P < 1 × 10−5, hypergeometric test; Supplementary Table 6a,m,n). 7α,25-OHC is known to trigger reactive oxygen species formation46 and cell stress, which can activate complement46,47,48,49 and may also synergize with C5a intracellular signaling, as both C5a and 7α,25-OHC are proinflammatory. Other perturbations, aside from C5AR1, did not show this trend and had only moderate (GPR18, GPR34 and GPR84) to minimal (CXCR2 and LPAR2) transcriptional remodeling effects under both conditions (Fig. 3c and Supplementary Table 6).
7α,25-OHC can readily travel in and out of cells in a GPR183-independent manner50. In accordance with this, 7α,25-OHC also altered the NK-92 cell transcriptome in a GPR183-independent manner, including a significant downregulation of genes involved in cholesterol and sterol synthesis and lipid transport (Fig. 3f and Supplementary Table 6b).
GPR183 mobilizes NK cells to breast cancer cells and tumors
The most potent ligand for GPR183 is 7α,25-OHC, generated from cholesterol through the sequential action of the enzyme cholesterol 25-hydroxylase (CH25H) and the cytochrome P450, family 7, subfamily b, polypeptide 1 (CYP7B1; Fig. 4a)25,50. 7α,27-OHC is another potent GPR183 ligand, generated from cholesterol through the sequential action of sterol 27-hydroxylase (CYP27A1) and CYP7B1 (Fig. 4a)25,50. Thus, CYP7B1 is required for the synthesis of both ligands, whereas CH25H is required for 7α,25-OHC, but not 7α,27-OHC, synthesis.
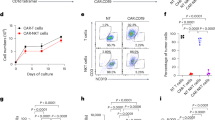
a, Enzymatic oxysterol synthesis pathway. Sequential cholesterol degradation by CH25H and CYP7B1 enzymes generates the GPR183 ligand 7α,25-OHC, and sequential cholesterol degradation by CYP27A1 and CYP7B1 generates the GPR183 ligand 7α,27-OHC. b, Expression levels of oxysterol-synthesizing enzymes in MDA-MB-231 cells in monoculture with and without IFNγ and TNF treatment (n = 3 biological replicates per group). Expression levels were measured via quantitative real-time PCR (qPCR; Supplementary Information) and normalized to GAPDH. Data are representative of two independent experiments, are presented as the mean ± s.e.m. and were analyzed by two-tailed unpaired Student’s t-test. c, CH25H- and CYP7B1-deficient MDA-MB-231 cells were generated using CRISPR knockout. Knockout efficiency was validated using the Synthego ICE analysis tool (v3). Control cells (transduced with NTC sgRNA) are shown in gray. The dashed black bar indicates the sequence alignment window, the solid black bar indicates the ICE interference window, and the gray dotted line marks the start of the guide sequence; KO, knockout. d,e, Transwell assay with flow cytometry-based quantification of NK-92 cell migration to CM from isogenic MDA-MB-231 cells with and without knockout of oxysterol synthesis enzymes (d; n = 3 technical replicates per group) and to tumor and lung lysates (e; n = 9 technical replicates per group). f, GPR183 cell surface expression in GPR183OE and control primary human NK (pNK) cells. g,h, Transwell assay with flow cytometry-based quantification of control and GPR183OE primary NK cell migration to 7α,25-OHC (g; n = 3 technical replicates per group) and to tumor and lung lysates (h; n = 6 technical replicates per group). i, In vivo co-transfer experiment. NK-92 cells were transduced with either control–GFP or GPR183–mCherry. GFP+ and mCherry+ cells were mixed at a 1:1 ratio and intravenously transferred into MDA-MB-231-bearing NSG mice. Spleens, lungs and tumors were dissected, and tissue-infiltrating cells were assessed by flow cytometry. The GPR183:control ratio was calculated across tissues (n = 7 mice). Experimental scheme (left), representative flow cytometry plots (middle) and a bar plot with each dot depicting the ratio per mouse (right) are shown. j, Proliferation of GPR183A+ cells and control NK-92 cells quantified via decay of cell-trace dye. Top, histogram plots. Bottom, relative mean fluorescence intensity (MFI) quantification (n = 3 technical replicates per group). Data are representative of two independent experiments. Two independent experiments were pooled for e, h and i. Data are representative of two independent experiments for d and g. Data are presented as mean values ± s.e.m. for d, e and g–j. Data analysis was performed using one-way analysis of variance (ANOVA) with a Dunnett’s multiple comparisons test for d, g and i and two-way ANOVA with a Tukey’s multiple comparisons test for e and h. Panel i scheme created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
Oxysterol synthesis is transcriptionally active in breast cancer cell lines and is associated with advanced stage, invasiveness and expression of immune activation genes. More specifically, the GPR183 ligand synthesis genes CYP27A1, CYP7B1 and CH25H are broadly expressed in breast cancer cell lines51 (n = 69) and are overexpressed in invasive breast carcinoma lines (n = 64) compared to noninvasive breast ductal carcinoma in situ (n = 5) and noncancerous (n = 1) lines (Extended Data Fig. 8a). CYP27A1, CYP7B1 and CH25H are coexpressed across breast cancer cell lines (BH FDR < 0.05, Spearman correlation). They are also coexpressed with a shared set of 208 genes (a significantly large overlap, empirical P = 0.0055; Methods and Supplementary Table 7a), which is enriched for genes involved in immune responses, the interferon-γ (IFNγ) response, viral defense mechanisms and so on (for example, CD14, C3, CHST3, CMPK2, CXCL1, CXCL16, DDX60, GBP1, GBP3, GBP4, HERC5, IFI44, IFI44L, IFIT2, IFIT3, IL22RA1, IRAK2, IRAK3, IRF1, LYN, MX1, OASL, OPTN, PML, PSMB9, PTGFR, RSAD2, TBX21, TLR1, TLR2, TLR6 and TNFAIP3; Supplementary Table 7b). MDA-MB-231 cells used in our screens express CYP27A1, CYP7B1 and CH25H and other genes involved in oxysterol and 7α,25-OHC synthesis and overexpress them in response to IFNγ and TNF (Fig. 4b).
Indicating that GPR183-dependent migration to breast cancer cells is driven by 7α,25-OHC synthesis, it depends on CYP7B1 expression in the cancer cells. More specifically, relative to NTC NK-92 cells, GPR183A+ NK-92 cells showed a significant increase in cell migration toward conditioned medium (CM) from NTC MDA-MB-231 cells. This enhanced migration was abolished when using CM from CYP7B1-knockout MDA-MB-231 cells and was more moderately decreased when using CM from CH25H-knockout MDA-MB-231 cells (Fig. 4c,d and Supplementary Information).
Further investigation shows the tumor specificity of GPR183-driven migration in NK-92 and primary NK cells. GPR183A⁺ NK-92 cells showed enhanced migration toward tumor lysate compared to NTC NK-92 cells, with a more subtle increase to lung lysates derived from the same NSG mice (Fig. 4e). To test GPR183 in primary NK cells, we isolated and expanded primary NK cells from human peripheral blood mononuclear cells (PBMCs) using bead-based isolation and a feeder-cell-free expansion method, resulting in >92% CD56+CD3− NK cell products (Extended Data Fig. 8b). We then transduced primary NK cells using high-titer spin infection with either GPR183 ORF or control retrovirus (Fig. 4f, Extended Data Fig. 8c and Supplementary Information). Compared to donor-matched control NK cells, GPR183-overexpressing (GPR183OE) primary NK cells exhibited increased migration to 7α,25-OHC (Fig. 4g) and tumor lysate compared to lung lysate (Fig. 4h).
To test GPR183-driven migration in vivo, we transduced NK-92 cells with either control–green fluorescent protein (control–GFP) or GPR183–mCherry ORF vectors, mixed them at a 1:1 ratio, injected the pool into MDA-MB-231 tumor-bearing NSG mice and quantified relative abundance in the tumor, lung and spleen by flow cytometry (Fig. 4i and Extended Data Fig. 8d). GPR183OE NK-92 cells were more than fourfold more abundant in the tumors than control NK-92 cells, with only a modest increase or no increase in NK cell abundance in the lung and spleen, respectively (Fig. 4i). These effects are likely to be primarily due to GPR183-driven migration as GPR183 ectopic expression had no impact on NK-92 proliferation (Fig. 4j) and showed variable effects on NK-92 cell and primary NK cell cytotoxicity in coculture, as well as CD107a surface expression and IFNγ production following phorbol 12-myristate 13-acetate (PMA)/ionomycin stimulation and at baseline (Extended Data Fig. 8e and Supplementary Information).
GPR183 enhances NK-92 and CAR NK-92 cell efficacy in vivo
Expressing GPR183 in NK-92 and CAR NK-92 cells significantly improves the ability of these cells to control breast cancer tumors in NSG mice (Fig. 5a). Intravenous transfer of GPR183A+ NK-92 cells to NSG female mice bearing MDA-MB-231 xenografts significantly delayed tumor growth compared to both untreated mice (that is, treated with cell-free medium) and mice treated with NTC NK-92 cells (Fig. 5b and Extended Data Fig. 9a). To test whether GPR183 can enhance the in vivo efficacy of CAR NK cells, we cloned a CAR construct that targets epithelial cell adhesion molecule (EpCAM) on the surface of breast cancer cells (Fig. 5c and Extended Data Fig. 9b) and co-introduced the construct (anti-human EpCAM-CAR–GFP) with GPR183 ORF–mCherry or control–mCherry into NK-92 cells (Extended Data Fig. 9c), resulting in antigen-specific cell cytotoxicity (Extended Data Fig. 9d,e). Intravenous transfer of GPR183OE CAR NK-92 cells to tumor-bearing mice showed significantly better control of MDA-MB-231 tumor growth than both the untreated group (that is, treated with cell-free medium) and the group treated with control–mCherry CAR NK-92 cells (Fig. 5d and Extended Data Fig. 9f).
a, Disease outcome study design. b, Growth curve of MDA-MD-231 breast cancer tumors in NSG mice treated with cell-free medium (n = 3 mice), control NK-92 cells (n = 6 mice) or GPR183A+ NK-92 cells (n = 6 mice). c, EpCAM CAR construct; LTR, long terminal repeat; EFS, EF1α-short promoter; SP, signal peptide; scFC, single-chain variable fragment; TM, transmembrane. d, Growth curve of MDA-MD-231 breast cancer tumors in NSG mice treated with cell-free medium (n = 3 mice), anti-EpCAM-CAR NK-92 (n = 6 mice) or GPR183OE anti-EpCAM-CAR NK-92 (n = 6 mice) cells. The results from a biological replicate of b and d are depicted in Extended Data Fig. 9a,f. In b and d, data are shown as the mean ± s.e.m. (left) and individual growth curves per mouse (right); statistical significance was evaluated by two-way ANOVA with a Sidak correction for multiple hypothesis testing; ****P < 0.0001. Illustrations in a and c created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
GPR183 enhances T cell tumor infiltration and in vivo efficacy
Given the biochemical and biophysical properties of GPR183, its association with tumor infiltration in multiple cell types, including CD8+ T cells (Fig. 2d and Extended Data Fig. 5a,b), and the similarity in its transcriptional regulation in NK and CD8+ T cells (Fig. 2h), we hypothesized that expressing GPR183 in CD8+ T cells could mobilize T cells to solid tumors and hence improve in vivo efficacy. To test this, we isolated CD8+ T cells from human PBMCs (Extended Data Fig. 8b) and transduced them with either GPR183 ORF or control lentivirus (Fig. 6a). Consistent with previous NK-92 and primary NK cell migration data, GPR183OE primary T cells displayed increased migration to 7α,25-OHC (Fig. 6b), MDA-MB-231 supernatant (Fig. 6c) and tumor versus lung lysate (Fig. 6d) compared to control CD8+ T cells from the same donor.
a, GPR183 cell surface expression in GPR183OE and control primary human CD8+ T cells. b–d, Transwell assay with flow cytometry-based quantification of control and GPR183OE primary T cell migration to 7α,25-OHC (b), MDA-MB-231 cell CM (c) and tumor and lung lysates (d); n = 6 technical replicates per group in b and n = 5 technical replicates per group in c and d. e–g, In vivo CAR T cell quantification and functional test. e, Experimental scheme. f, GPR183OE EpCAM-targeting CAR T cells accumulate in the tumor more than control CAR T cells, whereas their infiltration remains comparable in the lung and spleen. Representative flow cytometry plots (left) and bar plots show T cell number and relative abundance (right; n = 9 mice); m, mouse; h, human. g, IFNγ production and CD107a surface expression were quantified via flow cytometry in tumor-infiltrating CAR T cells isolated from nine mice at baseline and following ex vivo PMA/ionomycin stimulation. The data in f and g depict n = 4 mice treated with CAR T cells and n = 5 mice treated with GPR183OE EpCAM-targeting CAR T cells. h,i, In vivo antitumor efficacy of CAR T cells in NSG mice with EpCAM-overexpressing MDA-MB-231 xenografts. Tumor growth (h) and survival (i) curves of mice treated with control anti-EpCAM-CAR T cells (n = 6 mice) or GPR183OE anti-EpCAM-CAR T cells (n = 7 mice). Data are representative of two independent experiments for b–d. In vivo CAR T cell experiments were repeated with T cells generated from an independent donor (Extended Data Fig. 10c–e). Data are presented as the mean ± s.e.m. for b,d, f and g. In h, data are shown as the mean ± s.e.m. (left) and individual growth curves per mouse (right). Data analysis was performed using a one-way ANOVA with a Dunnett’s multiple comparisons test for b, two-way ANOVA with a Tukey’s multiple comparisons test for d, two-tailed unpaired Student’s t-test for c, f and g, two-way ANOVA with a Sidak correction for multiple hypothesis testing for h and log-rank test for i. Panel e created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
GPR183OE EpCAM CAR T cells (Extended Data Fig. 10a) transferred into NSG mice bearing established EpCAMOE MDA-MB-231 tumors were significantly more abundant in the tumor than control CAR T cells, whereas their abundance in the lung and spleen remained comparable (Fig. 6e,f and Extended Data Fig. 10b,c). Tumor-infiltrating GPR183OE and control CAR T cells isolated from the tumors expressed similar levels of effector molecules (that is, IFNγ and CD107a) at baseline and following PMA/ionomycin stimulation (Fig. 6g and Extended Data Fig. 10d), as also observed in vitro (Extended Data Fig. 8e). GPR183-dependent tumor infiltration translated to significantly better in vivo efficacy of CAR T cells in NSG mice. Treatment with GPR183OE CAR T cells resulted in significantly better control of tumor growth and prolonged survival (Fig. 6h,i and Extended Data Fig. 10e).
GPR183 enhances T cell in vivo efficacy in immunocompetent mice
Our data demonstrate that expressing GPR183 in NK cells and T cells preferentially redirects them into the tumor, resulting in improved tumor control. Because both the immune cells and cancer cells used here were of human origin, we conducted the in vivo experiments in immune-compromised NSG mice, which severely lack endogenous mouse immune cells.
To study GPR183 in a fully immunocompetent model, where diverse immune responses are orchestrated, we expressed the mouse homolog Gpr183 or a control vector in ovalbumin-specific OT-1 CD8+ T cells (Fig. 7a) and confirmed that GPR183OE OT-1 T cells show increased migration to 7α,25-OHC in vitro (Fig. 7b). To establish a breast tumor model in C57BL/6 mice, we transduced a mouse syngeneic triple-negative breast cancer line (E0771) to stably express ovalbumin (E0771-OVA; Fig. 7c), enabling antigen-specific recognition and killing by OT-1 T cells (Fig. 7d).
a, Retroviral constructs for mouse Gpr183 ORF (top) and representative flow cytometry plots showing transduction efficiency in mouse OT-1 T cells (bottom). b, Transwell flow cytometry-based quantification of control and GPR183OE OT-1 T cell migration to 7α,25-OHC (n = 3 technical replicates per group). c, Ovalbumin overexpression in the E0771 C57BL/6 syngeneic breast cancer cell line. Surface expression of SIINFEKL-peptide-loaded H-2Kb was assessed by flow cytometry; WT, wild-type. d, OT-1 antigen-specific cytotoxicity measured in coculture with E0771-wild-type and E0771-OVA cells at varying effector:target (E:T) ratios for 24 h (n = 3 technical replicates per group). e-g, In vivo antitumor efficacy of OT-1 T cells. e, Experimental workflow: E0771-OVA cells were orthotopically engrafted to the mammary fat pad of C57BL/6 mice, followed by OT-1 cell intravenous injection 8 days after engraftment. Tumor growth (f) and survival (g) curves of mice treated with control OT-1 or GPR183OE OT-1 cells (n = 10 per group); CR, complete response. Data are representative of two independent experiments and are presented as the mean ± s.e.m. for b and d. Two independent experiments were pooled for f and g. Data analysis was performed using a one-way ANOVA with a Dunnett’s multiple comparisons test for b, two-tailed unpaired Student’s t-test for d, two-sided mixed-effects model with Sidak correction for multiple hypothesis testing at day 43 after tumor engraftment for f and one-sided log-rank test for g. Illustrations in a and e created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
For in vivo efficacy testing, we orthotopically engrafted E0771-OVA cells into the fourth mammary fat pad and, after tumors formed, treated the tumor-bearing C57BL/6 mice with either control OT-1 or GPR183OE OT-1 T cells (Fig. 7e). GPR183OE OT-1 T cells were significantly more effective than control OT-1 T cells at delaying tumor growth (Fig. 7f) and prolonging survival (Fig. 7g), resulting in a complete response in 70% of the mice compared to a 30% complete response rate in the control OT-1 treatment group.
Discussion
Altered metabolism has been widely recognized as a cancer hallmark52, enabling cancer cells to generate biomass and sustain anabolic processes through the Warburg effect53 and altered lipid metabolism54. Aberrant metabolism in cancer has long been exploited for cancer detection via positron emission tomography imaging55 (primarily using radiolabeled glucose analogs and, more recently, choline56 and acetate57 analogs). Here, through a combination of gain-of-function screens, mechanistic and preclinical in vivo studies, we show that metabolite-sensing GPCRs provide programmable mechanisms to mobilize cytotoxic lymphocytes to breast and ovarian cancer tumors, significantly improving the efficacy of T and NK cell therapies.
thGPRs have been studied for their role in modulating immune cell function and migration. GPR84 drives cell migration to medium-chain fatty acids and is known to support phagocytosis, inflammation and metabolic sensing58. GPR18 is important for directional microglial migration in the central nervous system24, sperm motility59 and the development of CD8αα intestinal intraepithelial lymphocytes60 and CD8+KLRG1+ effector memory T cells61. GPR34 is highly expressed in microglia, plays a protective role against pathogen infection in the central nervous system62, facilitates tissue repair by promoting innate lymphoid cell recruitment to areas of damage-induced cell death63 and supports immune cell accumulation in the peritoneal cavity and omentum64.
GPR183 is known to orchestrate immune cell positioning within lymphoid and mucosal tissues25,28. Inflammation upregulates oxysterol-synthesizing enzymes, thereby enhancing the recruitment of GPR183-expressing cells to inflamed sites50. GPR183 has also been linked to NK cell expansion and lymphocyte function in diverse tissue contexts42,50,65, and its knockout has been shown to impair the homing of naive B cells and a subset of memory T cells into inflamed lymph nodes and inflamed nonlymphoid tumor endothelium in an oxysterol-dependent manner66. Direct quantitation of GPR183 ligands via mass spectrometry (MS) has been shown to be confounded due to their low abundance relative to cholesterol and overlaps in MS/MS transition and retention times67,68. Thus, studies in the field have established the GPR183-dependent migration transwell assay (which was also used here) to detect GPR183 ligands with subnanomolar sensitivity50,67,68. Given these technical limitations, further investigation is needed to examine additional factors that may explain the effects observed here, including potential synergies of GPR183 with other receptors or other metabolites other than 7α,25-OHC and 7α,27-OHC, which may act as GPR183 ligands, but are yet to be identified.
GPR183 selectively enhanced NK migration to breast and ovarian cancer tumors in a series of independent CRISPRa screens and in a suite of follow-up in vitro, ex vivo and in vivo experiments in NK and T cells, making GPR183 T cell/NK cell therapies a promising modality to pursue in preclinical and potentially clinical investigations. Both GPR183 and the other thGPRs identified here merit further investigation to elucidate their underlying mechanism of action and therapeutic potential in breast, ovarian and other cancer types. thGPR combinations, synthetic thGPR variants and circuits can provide an array of modular modalities to mobilize and reprogram immune cells in ways that diverge from and extend upon current cell and protein engineering efforts. The results presented here provide the rationale to pursue these directions and develop new tools to do so at scale.
While cell engineering has primarily focused on protein–protein interactions, our work demonstrates the rich biochemical interface between immune cells and their environment and, through systematic in vivo gain-of-function screens, highlights the potential of bioactive metabolites and their cognate receptors as programmable levers to control immune cell localization. By converting tumor-derived biochemical cues into chemoattractant signals sensed by engineered cytotoxic lymphocytes, our study shows how we can turn the tumor’s own biochemical signals against it, opening new avenues for the design and development of spatially targeted biochemically guided cell therapies.
Methods
Study approval and regulations
The study received institutional regulatory approval. All experiments involving recombinant DNA and other biosafety considerations were performed under the guidelines and with the approval of Stanford University Environment, Health and Safety Committee (APB-3910-L.J.0921 and APB-5416-L.J.0724). All experiments involving materials from human samples were performed under the guidelines and with the approval of the Stanford University Institutional Review Board (protocol IRB-13942). All animal experiments were performed in mice with the approval of the Stanford University Institutional Animal Care and Use Committee (IACUC; APLAC-34218). IACUC guidelines and protocols were strictly followed. Mice were closely monitored, and approved tumor burden limits were not exceeded. The maximal permitted tumor burden was defined as a tumor diameter of 17 mm in adult mice, with earlier euthanasia required if animals showed signs of predefined clinical and morbidity criteria. Immunocompromised mice were housed in a pathogen-free facility. All mice were maintained on 12-h light/12-h dark cycles at room temperature ranging between 20 °C and 26 °C and humidities between 30 and 70%. Mice were fed ad libitum with 2018 Teklad 18% protein rodent diet (Envigo) and were provided ad libitum access to water.
Cell lines
NK-92 (CRL-2407), MDA-MB-231 (HTB-26), MCF7 (HTB-22) and E0771 (CRL-3461) cells were purchased from ATCC. Lenti-X 293T cells (632180) were purchased from Takara Bio. TYK-nu ovarian cancer cells (JCRB0234.0) were purchased from JRCB Cell Bank. NK-92 cells were cultured with RPMI-1640 medium with GlutaMAX supplement (Gibco, 61870036) containing 5 mM MEM nonessential amino acids solution (Cytiva, SH30598.01), 5 mM sodium pyruvate 100 mM solution (Cytiva, SH30239.01), 10% fetal bovine serum (FBS; Gibco, A3840102), 1% penicillin/streptomycin (referred to here as NK base medium) and 200 U ml−1 IL-2 (Frederick National Laboratory). MDA-MB-231, MCF7, Lenti-X 293T and E0771 cells were cultured with DMEM containing 5 mM MEM nonessential amino acids solution, 5 mM sodium pyruvate 100 mM solution, 10% FBS and 1% penicillin/streptomycin. TYK-nu ovarian cancer cells (JRCB Cell Bank, JCRB0234.0) were cultured in EMEM (ATCC, 30-2003) with 10% FBS. All cell lines were maintained at 37 °C and 5% CO2 and tested for mycoplasma by PCR. NK-92, MDA-MB-231 and TYK-nu cell lines were authenticated through ATCC using short tandem repeat profiling.
CRISPRa NK-92 cell libraries
The screens were conducted with both a customized sgRNA library and a large-scale sgRNA library. The customized library consisted of 1,070 sgRNAs targeting 256 genes with four guides per gene and 46 NTCs (Supplementary Table 1c), using protospacer sequences from the Human Genome-Wide CRISPRa-v2 Libraries69 and Human CRISPRa Library19 cloned into pKVL2-U6gRNA_SAM(BbsI)-PGKpuroBFP-W vector (Addgene, 112925) by Genscript. The large-scale library was a human membrane protein activation library that was purchased from Addgene (113345).
The CRISPR SAM system was used to generate the NK-92 cell libraries. NK-92 cells were sequentially transduced with lentivirus encoding dCas9-VP64–GFP (Addgene, 61422) and lentiMPH V2 (Addgene, 89308) at a multiplicity of infection (MOI) of <0.3. The lentivirus preparation procedure is described in the Supplementary Information. Transduced NK-92 cells were selected by GFP-based cell sorting and 500 μg ml−1 hygromycin B Gold (InvivoGen, ant-hg-1). dCas9-MPH NK-92 cells were transduced with the lentiviral custom or large-scale sgRNA library at an MOI of <0.3 and selected with 1 μg ml−1 puromycin for 7 days. Descriptions of the genomic DNA extraction, sgRNA amplification and sequencing procedures conducted as a part of the CRISPRa screens are available in the Supplementary Information.
In vivo NK CRISPRa tumor infiltration screens
Tumor infiltration NK-92 CRISPRa screens were conducted in breast and ovarian cancer tumors using a total of 36 female mice. NK-92 CRISPRa screens for infiltration to breast cancer tumors were performed with the customized sgRNA library using the following models (n = 10 mice): (1) NOD.Cg-Prkdcscid Il2rgtm1Sug Tg(CMV-IL2)4-2Jic/JicTac (Taconic Biosciences, hIL-2-NOG, 13440-F) mice with subcutaneous breast cancer xenografts (n = 4), (2) NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ (The Jackson Laboratory, NSG, 005557) mice with subcutaneous breast cancer xenografts (n = 3) and (3) NSG mice with orthotropic mammary fat pad breast cancer xenografts (n = 3). Seven- to 8-week-old female NSG and hIL-2-NOG mice were purchased from The Jackson Laboratory and Taconic Biosciences, respectively. In total, 5 × 106 MDA-MB-231 cells were engrafted (subcutaneously into the right flank or orthotopically into the left inguinal mammary fat pad) in a 1:1 mix of serum-free DMEM and Matrigel (Corning, 354483). Two weeks after engraftment, when the tumor size reached ~100 mm3, 107 library NK cells (that is, dCas9-MPH NK-92 cells transduced with the custom sgRNA library) were injected intravenously via tail vein injection. For the experiment with NSG mice, human IL-2 (5,000 U) was injected intraperitoneally immediately before NK cell transfer. Twenty four hours after NK infusion, mice were killed, and tumors and lungs were collected, immediately processed for genomic DNA (gDNA) extraction and processed together with the matched library NK-92 cells.
NK-92 CRISPRa screens for infiltration to ovarian cancer tumors were conducted with both the customized and large-scale sgRNA libraries in 6- to 7-week-old female NSG mice bearing intraperitoneal TYK-nu tumors (n = 10 mice for the customized screen and n = 16 mice for the large-scale screen). TYK-nu cells were lentivirally transduced to express luciferase (a plasmid provided as a gift from the Wucherpfennig lab (Dana–Farber Cancer Institute)) and an mScarlet reporter (a gift from the Hernandez-Lopez lab (Stanford University)). Transduced cells were enriched by fluorescence-activated cell sorting (FACS). For preliminary assessment of ovarian cancer engraftment, cells were injected intraperitoneally into NSG mice, and bioluminescent signals were monitored following intraperitoneal administration of 150 mg per kg (body weight) D-luciferin (Abcam, ab143655) under isoflurane–oxygen anesthesia using an in vivo imaging system. For in vivo screens, 3 × 106 TYK-nu-mScarlet cells were injected intraperitoneally and were allowed to engraft for 3 weeks. Mice were treated intraperitoneally with 1.5 × 107 library NK cells and supplemented with 5 μg of IL-15 (Frederick National Laboratory) intraperitoneally every 2 days for the duration of the experiment. One to 3 weeks after NK cell injections, mice were killed, and ascites fluid was collected by injecting 5 ml of PBS into the peritoneal cavity, followed by aspiration through a small abdominal incision. mScarlet+ tumor nodules were identified and dissected under a fluorescence microscopy. Tumor nodules were pooled on a per-mouse basis and immediately processed for gDNA extraction together with the animal-matched ascites fluid and matched library NK cell samples.
Chemoattraction NK CRISPRa screen
Customized library NK cells (that is, dCas9-MPH NK-92 cells transduced with the customized sgRNA library) were collected and washed with serum-free RPMI medium. Cells were resuspended in RPMI containing 1% FBS and counted using a Countess 3 Automated Cell Counter (Invitrogen, AMQAX2000). In total, 2.5 × 106 cells were added on a 24-mm transwell with an 8-µm pore (Corning, 3428) in 2 ml of RPMI containing 1% FBS. Next, 2.7 ml of MDA-MB-231 and MCF7 cancer cell conditioned media was added to the bottom plate and incubated at 37 °C and 5% CO2 for 4 h. After incubation, the NK cells that reached the bottom plate were collected and immediately processed for gDNA extraction together with the matched library NK-92 cells. Cells from six wells were pooled for a single experimental replicate, and three replicates were collected per condition.
Spheroid NK cell CRISPRa screen
For scalable spheroid generation, MCF7 cells were seeded at 1 × 106 cells per ml in 10 ml of DMEM containing 1% methylcellulose (Sigma-Aldrich, H7509) in a 90-mm ultra-low attachment dish (Sbio, MS-90900Z) and incubated for 2 days for spheroid formation. MCF7 spheroids were collected, diluted with PBS, centrifuged and resuspended in NK base medium. A small fraction of cells was taken, digested with TrypLE (Gibco, 12605010) and counted. In total, 5 × 106 custom library NK cells and 2.5 × 106 MCF spheroids were seeded at a 2:1 E:T ratio in 12.5 ml of RPMI + IL-2 (200 U ml−1) in 90-mm ultra-low attachment dishes. The cocultures were collected after 6, 24 and 48 h and filtered through a 70-μm strainer to separate noninfiltrated NK cells and spheroids. The filter was washed with PBS five times to remove residual noninfiltrated single cells. The remaining spheroids on top of the filter were collected to profile spheroid-infiltrating NK cells. The matching NK library cells were also in vitro cultured and collected at the same time points to obtain control reference samples. gDNA from spheroids and control samples were immediately processed. Cells from six dishes were pooled for a single experimental replicate, and three replicates were collected per condition.
PRESTO-Tango assay
HTLA cells, an HEK293 cell line stably expressing a tTA-dependent luciferase reporter and a β-arrestin2-TEV fusion gene, previously developed by R. Axel70 and used in the PRESTO-Tango assay36, were provided as a gift from the laboratory of M. Bassik (Stanford University). HTLA cells were maintained in DMEM supplemented with 10% FBS, 1% penicillin/streptomycin, 2 μg ml−1 puromycin (InvivoGen, ant-pr-1) and 100 μg ml−1 hygromycin B at 37 °C and 5% CO2. For GPCR cloning, gene fragments encoding ORFs of GPCR hits (CXCR2, C5AR1, GPR34, GPR84 and GPR183; sequences are provided in Supplementary Table 8b) and the TEV-V2 tail (encoding the TEV-cleavage site and tTA transcription factor) were synthesized from Twist Bioscience, cloned into pLV-EF1a-IRES-Blast vector (Addgene, 85133) and digested with BamHI-HF (New England Biolabs, R3136) and EcoRI-HF (New England Biolabs, R3101), with Gibson assembly. Cloned GPCR constructs were packaged with lentivirus and used to transduce HTLA cells. Transduced cells were selected with 8 μg ml−1 blasticidin (InvivoGen, ant-bl-05). For the PRESTO-Tango assay, 4 × 104 of each GPCR reporter and control HTLA cells were seeded in a white-walled, clear-bottom 96-well plate (Corning, 354651). One day after cell seeding, varying doses of relevant ligands, as indicated in the figures and figure legends, were added to each well and incubated at 37 °C with 5% CO2 for 24 h. The next day, Bright-Glo solution (Promega, E2610) was added and incubated for 5 min at room temperature. Luminescence was measured using a Tecan Infinite M1000 plate reader.
CAR construct design, cloning and assessment of surface expression
Anti-hEpCAM CAR (Fig. 5c) was cloned and used as follows. Lentiviral CAR backbone plasmid encoding anti-CD19-41bb-CD3ζ–eGFP was purchased from Addgene (135992). The original CD19 CAR construct was digested with FastDigest Bpil to replace the anti-CD19-scFv-CD8 hinge region with the anti-hEpCAM-scFv-CD28 hinge. A codon-optimized gene fragment encoding anti-hEpCAM-scFv-CD28 hinge was synthesized by Twist Bioscience (gene fragment sequences are provided in Supplementary Table 8e) and cloned into a digested backbone with Gibson assembly. A high-titer lentivirus encoding an anti-hEpCAM CAR was generated and used to transduce NK-92 and human primary T cells, as described in ‘Human primary CD8+ T cell isolation, expansion and engineering’. Transduction efficiency was measured by flow cytometric detection of GFP. The surface expression of anti-hEpCAM CAR was assessed with Alexa Fluor 647-AffiniPure Goat Anti-Human IgG, F(ab′)2 Fragment Specific (Jackson ImmunoResearch Labs, 109605006; 1:100) via flow cytometry (Extended Data Figs. 9c and 10a).
Human primary CD8+ T cell isolation, expansion and engineering
Primary human CD8+ T cells were isolated from human whole-blood buffy coats obtained from the Stanford Blood Center. First, PBMCs were purified using Ficoll-Paque Premium Medium (Cytova, 17-5442-02) and Sepmate tubes (STEMCELL Technologies, 85450). To isolate CD8+ T cells, PBMCs were further processed with an EasySep Human CD8+ T cell isolation kit (STEMCELL Technologies, 17953). Enriched CD8+ T cells were cultured in T cell medium consisting of RPMI-1640 with GlutaMAX and HEPES medium (Gibco, 72400047) containing 10% heat-inactivated FBS, 1% heat-inactivated human AB serum (MilliporeSigma, H4522), 5 mM sodium pyruvate, 5 mM nonessential amino acids and 50 µM β-mercaptoethanol (Sigma, M6250) at 1 × 106 cells per ml in a nontreated 24-well plate. On day 0, T cells were activated with Dynabeads (Gibco, 11131D) at a 1:1 ratio for 3 days, followed by incubation with 100 U ml−1 IL-2. For T cell transduction with the control or GPR183 ORF vector, concentrated lentivirus was directly added to each well containing CD8+ T cells and mixed by gentle pipetting on day 1 after Dynabead activation. On day 3, beads were removed, and cells were split at 1 × 106 cells per ml with fresh medium and IL-2. On day 5, T cells were collected and sorted based on mCherry using a Sony Biotechnology SH800S Cell Sorter. Sorted mCherry+ T cells were replated at 1 × 106 cells per ml with fresh medium and IL-2 for 5 more days. For CAR T cell generation, ORF-expressing T cells were reactivated with Dynabeads and transduced with lentivirus encoding anti-hEpCAM CAR. Beads were removed 2 days after the second activation, and cells were sorted for mCherry and GFP expression using a cell sorter. The sorted cells were further expanded for 4–5 days before being used for in vivo experiments.
Human primary NK cell isolation, expansion and engineering
Primary human NK cells were isolated from human whole-blood buffy coats obtained from the Stanford Blood Center and expanded via feeder-cell-free procedures71. First, PBMCs were purified using Ficoll-Paque Premium Medium (Cytova, 17-5442-02) and Sepmate tubes (STEMCELL Technologies, 85450). To isolate primary NK cells, PBMCs were further processed with an EasySep Human NK cell isolation kit (STEMCELL Technologies, 17955). Enriched NK cells were cultured in NK MACS medium (Miltenyi, 130-114-429) supplemented with 5% EliteGro-Adv Human Platelet Lysate (Elite Cell, EPA-050) and fresh recombinant human IL-2 (1,000 U ml−1). On the day of isolation (day 0), NK cells were supplemented with washed MACSi-Beads conjugated with anti-human NKp46 and anti-CD2 made from an NK Cell Activation/Expansion kit (human; Miltenyi Biotec, 130-094-483) at a bead:cell ratio of 1:2 and seeded at 1 × 106 cells per ml in a nontreated 24-well plate. On day 5, the medium was replaced with fresh medium and IL-2. One day before transduction, a nontreated 24-well plate was coated with retronectin (Takara, T100A) diluted to 20 μg ml−1 in PBS and stored overnight at 4 °C. On day 7, NK cells were collected, resuspended at 3 × 106 cells per ml and mixed with retrovirus. NK cell/virus mixture was then plated in the retronectin-coated plates and left at room temperature for 5 min and 37 °C for 10 min. The cells were then centrifuged at 2,000g for 1 h at 30 °C. Cells were incubated for 48 h at 37 °C, and the virus was removed. NK cells were collected and sorted based on mCherry using a Sony Biotechnology SH800S Cell Sorter. Sorted mCherry+ NK cells were replated at 1 × 105 cells per ml with fresh medium containing IL-2 for 4–5 days and used for the in vitro migration assay.
Mouse T cell isolation, expansion and engineering
Spleens and peripheral lymph nodes from C57BL/6Tg(TcraTcrb)-1100Mjb/J (The Jackson Laboratory, OT-1, 003831) mice were mechanically dissociated through a 100-μm cell strainer and treated with red blood cell lysis buffer. The single-cell suspension was further processed using an EasySep Mouse CD8+ T cell isolation kit (STEMCELL Technologies, 19853). Enriched OT-1 T cells were cultured in T cell medium in a 24-well plate precoated with anti-mouse CD3e/CD28 (BD Biosciences, 550275 and 553294) at 1 × 106 cells per ml (day 0). On day 1, highly concentrated control or mouse Gpr183 retrovirus was added directly to the cells along with 4 µg ml−1 polybrene (Sigma, 107689), followed by centrifugation at 2,000g for 1 h at 30 °C. The cells were incubated for 48 h at 37 °C and removed from the virus and anti-mouse CD3e/CD28 stimulation. GFP⁺ cells were sorted using a cell sorter and expanded in T cell medium supplemented with 100 U ml−1 IL-2 for 5–7 days before use. For the mouse Gpr183 construct, the gene fragment (Supplementary Table 8d) was synthesized by Twist Bioscience and cloned into pBMN-mGFP vector (Addgene, 188643), which was digested with BamHI-HF and EcoRI-HF, using Gibson assembly. Retroviruses carrying control or mouse Gpr183 constructs were generated via Plat-E cells.
Quantification of in vivo NK or CAR T cell infiltration
For in vivo NK-92 co-transfer experiments, 7- to 8-week-old female NSG mice (n = 7) were subcutaneously injected with 5 × 106 MDA-MB-231 cells in a 1:1 mix of serum-free DMEM and Matrigel. One week after engraftment, when the tumor size reached ~50 mm3, the tumor-bearing mice were randomized, and 107 GFP–control and GPR183–mCherry NK cells, mixed at a 1:1 ratio, were transferred into the mice by intravenous tail vein injection once a week for 4 weeks. Human IL-2 (5,000 U) was intraperitoneally administered daily for 4 consecutive days. Two days after the fourth NK cell injection, lungs, spleens and tumors were dissected.
For in vivo CAR T cell infiltration experiments, 7- to 8-week-old female NSG mice (n = 20 in total) were subcutaneously injected with 5 × 106 EpCAM-overexpressing MDA-MB-231 cells in a 1:1 mix of serum-free DMEM and Matrigel. At day 13 after tumor engraftment, tumor-bearing mice were randomized, and 4 × 106 control (n = 9) or GPR183 CAR T cells (n = 11) were transferred into tumor-bearing mice by intravenous tail vein injection. Six days after T cell injection, lungs, spleens and tumors were dissected. These experiments were conducted twice with T cells isolated from two donors: 9 mice were used in the experiment shown in Fig. 6e,f (n = 4 and 5 mice treated with control CAR T and GPR183 CAR T cells, respectively, both derived from the same donor), and 11 mice were used in the experiment shown in Extended Data Fig. 10c (n = 5 and 6 mice treated with control CAR T and GPR183 CAR T cells, respectively, both derived from the same donor).
In both NK-92 and CAR T cell experiments, lungs and tumors were minced into small pieces, digested with collagenase type IV and DNase I for 45 min at 37 °C with agitation and filtered through a 100-μm cell strainer. Spleens were mechanically dissociated through a 100-μm cell strainer. Red blood cells were removed by using Red Blood Cell Lysing Buffer (Sigma, R7757) for 5 min at room temperature. The cell suspension was then diluted with 10 ml of PBS containing 2% FBS, passed through a 100-μm cell strainer, pelleted and resuspended in NK base medium. The single-cell suspension was further processed for flow cytometry staining to quantify tissue-infiltrating NK and CAR T cells, as described in ‘Flow cytometry and cell sorting’. 7-AAD−mouse CD45−human CD45+ cells were identified as injected lymphocytes. Cell counting beads were used for precise cell counts.
Disease outcome experiments
For in vivo NK-92 and CAR NK-92 efficacy experiments (Fig. 5 and Extended Data Figs. 9a,f), 7- to 8-week-old female NSG mice (n = 53 in total) were subcutaneously engrafted with 5 × 106 MDA-MB-231 cells in a 1:1 mix of serum-free DMEM and Matrigel. One week later, when the tumor size reached ~50 mm3, the tumor-bearing mice were randomized, and 107 NK cells were adoptively transferred via intravenous tail vein injection once a week for 4 weeks (n = 5–6 mice per group) or injected with cell-free medium (n = 3 mice). The experiments (Fig. 5) were repeated with five mice per group (Extended Data Fig. 9a,f). Human IL-2 (5,000 U) was intraperitoneally administered daily for 4 consecutive days.
For the in vivo CAR T cell efficacy experiment (Fig. 6h,i and Extended Data Fig. 10e), 7- to 8-week-old female NSG mice (n = 26 in total) were subcutaneously engrafted with 5 × 106 EPCAMOE MDA-MB-231 cells in a 1:1 mix of serum-free DMEM and Matrigel. At day 13 after tumor implantation, tumor-bearing mice were randomized, and 4 × 106 control (n = 6) or GPR183 CAR T cells (n = 7) generated from the same human donor were adoptively transferred into tumor-bearing mice by intravenous tail vein injection. The experiments were conducted twice using T cells isolated from two different human donors.
For the in vivo OT-1 T cell efficacy experiment (Fig. 7e–g), 2.5 × 106 E0771-OVA cells were orthotopically engrafted into the left inguinal mammary fat pad of 7- to 8-week-old female C57BL/6J (The Jackson Laboratory, 000664) mice. On day 8 after implantation, tumor-bearing mice were randomized, and 105 OT-1 T cells expressing either the control vector or mouse Gpr183 were intravenously injected into the tail vein (n = 10 mice per group).
Tumor volumes were measured by mechanical calipers every 3 or 4 days and calculated with the following formula: volume = 0.5 × length × width2. Mice were killed when tumor size reached the humane endpoint (>17 mm in diameter) or when mice were diagnosed with ulceration, according to institutional guidelines. None of the tumors exceeded the permitted tumor volume.
Flow cytometry and cell sorting
Cells were collected and washed with FACS buffer (PBS + 2% FBS). Human TruStain FcX (Fc receptor blocker; Biolegend, 422302; 1:50) was used to minimize nonspecific binding of antibodies. For live/dead cell stain, 7-AAD (Sigma-Aldrich, SML1633; 1:1,000) was used. Cells were then stained for surface or intracellular markers at 4 °C in the dark for 20 min unless stated otherwise. For detecting surface levels of GPCRs, staining was performed at 37 °C in the dark for 30 min. The following antibodies were used at a dilution of 1:200: anti-human CD182 (CXCR2)–Alexa Fluor 647 (Biolegend, 320714, clone 5E8), anti-human CD88 (C5AR1)–PE (Biolegend, 344303, clone S5/1), anti-human GPR183–Alexa Fluor 647 (Biolegend, 368903, clone SA313E4), anti-human CD326 (EpCAM)–APC (Biolegend, 324207, clone 9C4), anti-human CD16–APC (Biolegend, 980104, clone 3G8), anti-human CXCR6–APC (Biolegend, 356005, clone K041E5), anti-human CD3–VioGreen (Miltenyi Biotech, 130-113-134, clone BW264/56), anti-human CD8a–VioBlue (Miltenyi Biotech, 130-113-162, clone BW135/80), anti-human TCRα/TCRβ–APC (Biolegend, 306717, clone IP26), anti-human CD56–APC (Miltenyi Biotech, 130-113-305, clone AF12-7H3), anti-mouse CD45–APC/Cyanine7 (Biolegend, 103116, clone 30-F11) and antibody to mouse H-2Kb bound to SIINFEKL (Biolegend, 141603, clone 25-D1.16). Anti-human CD45 (Biolegend, 368510, clone 30-F11) was used at a 1:50 dilution. Flow cytometry was performed on a Quanteon, Penteon (NovoCyte) or Sony Biotechnology SH800S Cell Sorter. Cell sorting was performed using a Sony Biotechnology SH800S Cell Sorter. All flow cytometry data were analyzed using FlowJo version 10.10.0.
Perturb-seq library and sequencing
Protospacer sequences of five NTC sgRNAs and three sgRNAs targeting the top seven hits from the CRISPRa screens were used. The oligonucleotides were synthesized from Integrated DNA Technologies DNA, annealed, cloned into pKVL2-U6gRNA_SAM(BbsI)-PGKpuroBFP-W vector and digested with FastDigest Bpil. sgRNA vectors targeting each GPCR or NTC were pooled at equal amounts and used for transfecting Lenti-X 293T cells to generate lentivirus. CRISPR/dCas9-MPH-expressing NK-92 cells were transduced with a pooled sgRNA at an MOI of <0.2 and selected with 1 μg ml−1 puromycin. Cells were passaged five times after puromycin removal. Cells were either treated or not treated with 100 nM 7α,25-OHC for 4 h, prepared according to the 10x ‘Single Cell Suspensions from Cultured Cell Lines for Single Cell RNA Sequencing’ protocol (10x Genomics, CG00054 Rev B) and processed on a 10x Chromium controller instrument. The libraries were prepared according to the ‘Chromium Next GEM Single Cell 5′ Reagent Kits v2 (Dual Index) with Feature Barcode technology for CRISPR Screening’ protocol (10x Genomics, CG000510). Equimolar amounts of indexed libraries were pooled and sequenced on a NovaSeq X 1.5B in a paired-end run.
scRNA-seq immune cell data analyses
scRNA-seq data previously generated from tumor and blood samples obtained from individuals with breast cancer were downloaded from the Gene Expression Omnibus (GEO; accession number GSE169246)15. For each cell type, differentially expressed genes (FDR < 0.01, log fold change > 0.1) were identified by comparing the tumor and blood samples using the MAST test17 (a hurdle model tailored to scRNA-seq data; MAST R package, version 1.36.0). Cell types represented by fewer than 500 cells in either tumor or blood samples were excluded from these analyses. To examine GPR183 expression patterns in CD8+ T cells, Spearman correlation coefficients and P values were computed when examining the TPM of GPR183 and all other genes in CD8+ T cells (Fig. 2h). In all scRNA-seq data analyses, the overall expression of gene signatures was computed using a previously established normalization scheme to filter out technical variation40,72.
scRNA-seq pan-cancer NK cell data analyses
A pan-cancer NK cell scRNA-seq atlas16 spanning 24 cancer types and more than 700 participants was analyzed to identify genes specific to tumor-infiltrating versus blood-circulating NK cells in different cancer types and to map thGPR expression patterns.
Differential gene expression analyses were conducted for each cancer type, comparing tumor samples with blood samples from individuals with the same cancer type. In cases where cancer-type-matched blood samples were unavailable or included fewer than 200 cells, blood samples from all other individuals with cancer were used. To control for interparticipant variation, differential gene expression analyses were performed using edgeR18,73 (BH FDR < 0.01, edgeR, log fold change > 0.1). edgeR was applied to pseudobulk samples generated by aggregating the expression per sample. Using the edgeR R package (version 3.40.2), a quasi-likelihood negative binomial generalized log-linear model was fit to the pseudobulk count data. Cancer types with less than 200 tumor-infiltrating NK cells were not considered in these analyses. Tumor-enriched genes were identified per cancer type (BH FDR < 0.01, edgeR, log fold change > 0.1). Genes identified as tumor enriched in at least nine cancer types comprise the NK pan-cancer tumor infiltration signature (Supplementary Table 1b).
NK cell subtypes previously defined by Tang et al.16 were used to map the hits to specific subtypes. Spearman correlation analyses were performed between the TPM of GPR183 and those of all other genes for each cancer type, using only tumor samples.
CRISPRa screen data analyses
Data from the CRISPRa screens were analyzed using MAGeCK (version 0.5.9.4)22,74. Fastq files were processed to align the reads to the sgRNA spacer sequences, generating a counts matrix of sgRNA by sample. Using MAGeCK, the variance of read counts was estimated by sharing information across the different sgRNAs and by using the NTC sgRNAs to adjust and control for technical variation across the samples. Effect size and P values were computed based on a negative binomial statistical model for every target gene by testing whether the abundance of sgRNAs targeting the pertaining genes differs significantly between two groups of samples. All P values were corrected for multiple hypothesis testing using BH correction, and FDRs are reported (Supplementary Tables 2–3 and 5). In the in vivo screens, NK cells in the tumor were compared to NK cells at the animal-matched reference site (lung in the breast cancer screens and ascites in the ovarian cancer screens) and with the NK library cells. The results were combined via a Fisher’s combined probability test to obtain summary statistics for each of the three in vivo screens. The results are provided both separately and as summary statistics (Supplementary Tables 2 and 3).
In the in vitro chemotaxis screens, NK cells collected from the bottom insert of the transwell (4 h after placement at the top insert) were compared to the initial NK cell pool (cultured for 4 h). The screen was conducted twice, once with MDA-MD-231 supernatant and once with MCF7 supernatant as the chemoattractant. Fisher’s method P values were computed as summary statistics combining the results from the two types of chemotaxis screens. The results are provided both separately and as summary statistics (Supplementary Table 5). In the spheroid screens, NK cells residing within MCF7 spheroids after 6, 24 and 48 h were compared to NK cells cultured for 6, 24 and 48 h, respectively. Fisher’s combined probability test P values were computed as summary statistics to combine the results from all three time points. The results are provided both separately and as summary statistics (Supplementary Table 5).
Perturb-seq data analyses
Raw fastq files were processed using the cellranger pipeline (10x Genomics Cell Ranger 7.1.0) to align reads to the human genome (GRCh38) and to the sgRNA spacer sequences to generate two count matrices, one of gene expression (genes by cells) and another of sgRNA detection (sgRNA by cells). Gene expression counts were converted to TPM values and log1p-transformed. The Seurat R package was used to cluster the TPM gene expression profiles using the shared nearest neighbor clustering algorithm75 (Seurat package version 5.3.1 and SeuratObject package version 5.2.0). As the screen was conducted at a low MOI, each cell was assigned to a single sgRNA, the one it expressed at the highest level. For each of the seven genes targeted in the screen, a MAST analysis was performed to identify differentially expressed genes in cells with the corresponding CRISPRa gene activation compared to control cells (that is, cells carrying an NTC sgRNA; absolute log fold change > 0.1 and BH FDR < 0.1). Here, a cell was defined as subject to CRISPRa gene activation if it was assigned to one of its sgRNAs and expressed the target gene. The analyses were performed separately in each of the two culture conditions (that is, with and without 7α,25-OHC treatment).
Analysis of breast cancer cell line bulk gene expression data
RNA-seq (log1p-transformed TPM) data of 69 breast cancer cell lines and 1 noncancerous immortalized breast cell line were downloaded from DepMap51. Spearman correlation analysis was performed to identify coexpressed genes (BH FDR > 0.05, Spearman correlation). In total, 5,000 sets of three genes were used to sample the null distribution to examine if CYP27A1, CYP7B1 and CH25H show a significantly large overlap in the genes they are coexpressed with.
General study design
Each of the screens and experiments performed here were designed with sample sizes similar to those used in previous statistically robust studies7,8,28,36. No statistical method was used to predetermine sample size. Analyses were performed using statistical tests suitable for the data distribution. Analyses of CRISPR sgRNA count, scRNA-seq and Perturb-seq data were performed using methods (MAGeCK22,74, MAST17 and edgeR18) specifically designed for these types of data and their distributions or via nonparametric tests (for example, Spearman correlation). In cases where parametric tests (for example, t-tests and ANOVAs) were used, data distribution was assumed to be normal (for example, signature overall expression values in scRNA-seq data), but this was not formally tested. No animals or data were excluded from the analyses. Investigators were not blinded to experimental group allocation in experiments validating or investigating specific hits or treatments. In all in vivo experiments, mice were randomly allocated to the different treatment and control groups. CRISPRa and Perturb-seq screens are inherently randomized and blinded as investigators do not control allocation of cells to experimental groups (that is, perturbations).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data collected in this study are accessible via figShare at https://doi.org/10.6084/m9.figshare.31302178 (ref. 76). scRNA-seq immune cell data from individuals with breast cancer were accessed in GEO under accession code GSE169246 (ref. 15). Pan-cancer NK cell scRNA-seq data16 were downloaded from GEO using the accession code GSE212890 and from Zenodo at https://zenodo.org/records/8275845 (ref. 77). Breast cancer cell line RNA-seq data were downloaded from the DepMap51 portal (https://depmap.org/portal). Source data are provided with this paper.
Code availability
Code developed here for data analyses is available as the GitHub repository at https://github.com/Jerby-Lab/TNK_redirect.
References
Tan, S., Li, D. & Zhu, X. Cancer immunotherapy: pros, cons and beyond. Biomed. Pharmacother. 124, 109821 (2020).
Marofi, F. et al. CAR T cells in solid tumors: challenges and opportunities. Stem Cell Res. Ther. 12, 81 (2021).
Liu, D. et al. Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat. Med. 25, 1916–1927 (2019).
Shellard, A. & Mayor, R. All roads lead to directional cell migration. Trends Cell Biol. 30, 852–868 (2020).
Fowell, D. J. & Kim, M. The spatio-temporal control of effector T cell migration. Nat. Rev. Immunol. 21, 582–596 (2021).
Krummel, M. F., Bartumeus, F. & Gérard, A. T cell migration, search strategies and mechanisms. Nat. Rev. Immunol. 16, 193–201 (2016).
Dong, M. B. et al. Systematic immunotherapy target discovery using genome-scale in vivo CRISPR screens in CD8+ T cells. Cell 178, 1189–1204 (2019).
Peng, L. et al. In vivo AAV–SB-CRISPR screens of tumor-infiltrating primary NK cells identify genetic checkpoints of CAR-NK therapy. Nat. Biotechnol.43, 752–761 (2025).
Schmidt, R. et al. CRISPR activation and interference screens decode stimulation responses in primary human T cells. Science 375, eabj4008 (2022).
Legut, M. et al. A genome-scale screen for synthetic drivers of T cell proliferation. Nature 603, 728–735 (2022).
Zhou, P. et al. In vivo discovery of immunotherapy targets in the tumour microenvironment. Nature 506, 52–57 (2014).
Klingemann, H., Boissel, L. & Toneguzzo, F. Natural killer cells for immunotherapy—advantages of the NK-92 cell line over blood NK cells. Front. Immunol. 7, 91 (2016).
Boyiadzis, M. et al. Phase 1 clinical trial of adoptive immunotherapy using ‘off-the-shelf’ activated natural killer cells in patients with refractory and relapsed acute myeloid leukemia. Cytotherapy 19, 1225–1232 (2017).
Williams, B. A. et al. A phase I trial of NK-92 cells for refractory hematological malignancies relapsing after autologous hematopoietic cell transplantation shows safety and evidence of efficacy. Oncotarget 8, 89256–89268 (2017).
Zhang, Y. et al. Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39, 1578–1593 (2021).
Tang, F. et al. A pan-cancer single-cell panorama of human natural killer cells. Cell 186, 4235–4251 (2023).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Qin, D. et al. Potential lung attack and lethality generated by EpCAM-specific CAR-T cells in immunocompetent mouse models. Oncoimmunology 9, 1806009 (2020).
Morgan, R. A. et al. Case report of a serious adverse event following the administration of T cells transduced with a chimeric antigen receptor recognizing ERBB2. Mol. Ther. 18, 843–851 (2010).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Harding, S. D. et al. The IUPHAR/BPS Guide to Pharmacology in 2024. Nucleic Acids Res. 52, D1438–D1449 (2024).
McHugh, D. et al. N-Arachidonoyl glycine, an abundant endogenous lipid, potently drives directed cellular migration through GPR18, the putative abnormal cannabidiol receptor. BMC Neurosci. 11, 44 (2010).
Liu, C. et al. Oxysterols direct B-cell migration through EBI2. Nature 475, 519–523 (2011).
Suzuki, M. et al. Medium-chain fatty acid-sensing receptor, GPR84, is a proinflammatory receptor. J. Biol. Chem. 288, 10684–10691 (2013).
Kitamura, H. et al. GPR34 is a receptor for lysophosphatidylserine with a fatty acid at the sn-2 position. J. Biochem. 151, 511–518 (2012).
Hannedouche, S. et al. Oxysterols direct immune cell migration via EBI2. Nature 475, 524–527 (2011).
Schraufstatter, I. U., DiScipio, R. G., Zhao, M. & Khaldoyanidi, S. K. C3a and C5a are chemotactic factors for human mesenchymal stem cells, which cause prolonged ERK1/2 phosphorylation. J. Immunol. 182, 3827–3836 (2009).
Kelly, L. M., Pereira, J. P., Yi, T., Xu, Y. & Cyster, J. G. EBI2 guides serial movements of activated B cells and ligand activity is detectable in lymphoid and nonlymphoid tissues. J. Immunol. 187, 3026–3032 (2011).
Migeotte, I. et al. Identification and characterization of an endogenous chemotactic ligand specific for FPRL2. J. Exp. Med. 201, 83–93 (2005).
Sugo, T. et al. Identification of a lysophosphatidylserine receptor on mast cells. Biochem. Biophys. Res. Commun. 341, 1078–1087 (2006).
Chiang, N., Dalli, J., Colas, R. A. & Serhan, C. N. Identification of resolvin D2 receptor mediating resolution of infections and organ protection. J. Exp. Med. 212, 1203–1217 (2015).
An, S., Bleu, T., Hallmark, O. G. & Goetzl, E. J. Characterization of a novel subtype of human G-protein-coupled receptor for lysophosphatidic acid. J. Biol. Chem. 273, 7906–7910 (1998).
Moolenaar, W. H., Kranenburg, O., Postma, F. R. & Zondag, G. C. Lysophosphatidic acid: G-protein signalling and cellular responses. Curr. Opin. Cell Biol. 9, 168–173 (1997).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Ritscher, L. et al. The ligand specificity of the G-protein-coupled receptor GPR34. Biochem. J. 443, 841–850 (2012).
Darling, T. K. & Lamb, T. J. Emerging roles for Eph receptors and ephrin ligands in immunity. Front. Immunol. 10, 1473 (2019).
Christofides, A. et al. The complex role of tumor-infiltrating macrophages. Nat. Immunol. 23, 1148–1156 (2022).
Yeh, C. Y. et al. Mapping spatial organization and genetic cell-state regulators to target immune evasion in ovarian cancer. Nat. Immunol.25, 1943–1958 (2024).
Baptista, A. P. et al. The chemoattractant receptor EBI2 drives intranodal naive CD4+ T cell peripheralization to promote effective adaptive immunity. Immunity 50, 1188–1201 (2019).
Emgård, J. et al. Oxysterol sensing through the receptor GPR183 promotes the lymphoid-tissue-inducing function of innate lymphoid cells and colonic inflammation. Immunity 48, 120–132 (2018).
Lu, E., Dang, E. V., McDonald, J. G. & Cyster, J. G. Distinct oxysterol requirements for positioning naive and activated dendritic cells in the spleen. Sci. Immunol. 2, eaal5237 (2017).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866 (2016).
Replogle, J. M. et al. Combinatorial single-cell CRISPR screens by direct guide RNA capture and targeted sequencing. Nat. Biotechnol. 38, 954–961 (2020).
Kim, J.-S.et al. GPR183 regulates 7α,25-dihydroxycholesterol-induced oxiapoptophagy in L929 mouse fibroblast cell. Molecules 27, 4798 (2022).
Trakkides, T.-O. et al. Oxidative stress increases endogenous complement-dependent inflammatory and angiogenic responses in retinal pigment epithelial cells independently of exogenous complement sources. Antioxidants 8, 548 (2019).
Vogt, W., Damerau, B., von Zabern, I., Nolte, R. & Brunahl, D. Non-enzymic activation of the fifth component of human complement, by oxygen radicals. Some properties of the activation product, C5b-like C5. Mol. Immunol. 26, 1133–1142 (1989).
Tezel, G. et al. Oxidative stress and the regulation of complement activation in human glaucoma. Invest. Ophthalmol. Vis. Sci. 51, 5071–5082 (2010).
Cyster, J. G., Dang, E. V., Reboldi, A. & Yi, T. 25-Hydroxycholesterols in innate and adaptive immunity. Nat. Rev. Immunol. 14, 731–743 (2014).
Arafeh, R., Shibue, T., Dempster, J. M., Hahn, W. C. & Vazquez, F. The present and future of the Cancer Dependency Map. Nat. Rev. Cancer 25, 59–73 (2025).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Warburg, O. On respiratory impairment in cancer cells. Science 124, 269–270 (1956).
Snaebjornsson, M. T., Janaki-Raman, S. & Schulze, A. Greasing the wheels of the cancer machine: the role of lipid metabolism in cancer. Cell Metab. 31, 62–76 (2020).
Schwenck, J. et al. Advances in PET imaging of cancer. Nat. Rev. Cancer 23, 474–490 (2023).
Hara, T. 18F-Fluorocholine: a new oncologic PET tracer. J. Nucl. Med. 42, 1815 (2001).
Deford-Watts, L. M., Mintz, A. & Kridel, S. J. The potential of 11C-acetate PET for monitoring the fatty acid synthesis pathway in tumors. Curr. Pharm. Biotechnol. 14, 300–312 (2013).
Wojciechowicz, M. L. & Ma’ayan, A. GPR84: an immune response dial? Nat. Rev. Drug Discov. 19, 374 (2020).
Flegel, C. et al. Characterization of non-olfactory GPCRs in human sperm with a focus on GPR18. Sci. Rep. 6, 32255 (2016).
Wang, X., Sumida, H. & Cyster, J. G. GPR18 is required for a normal CD8αα intestinal intraepithelial lymphocyte compartment. J. Exp. Med. 211, 2351–2359 (2014).
Sumida, H. & Cyster, J. G. G-protein coupled receptor 18 contributes to establishment of the CD8+ effector T cell compartment. Front. Immunol. 9, 660 (2018).
Lou, L. et al. Mafba and Mafbb regulate microglial colonization of zebrafish brain via controlling chemotaxis receptor expression. Proc. Natl Acad. Sci. USA 119, e2203273119 (2022).
Wang, X. et al. GPR34-mediated sensing of lysophosphatidylserine released by apoptotic neutrophils activates type 3 innate lymphoid cells to mediate tissue repair. Immunity 54, 1123–1136 (2021).
Tam, H. et al. Phosphatidylserine phospholipase A1 enables GPR34-dependent immune cell accumulation in the peritoneal cavity. J. Exp. Med. 221, e20240992 (2024).
Brownlie, D. et al. Expansions of adaptive-like NK cells with a tissue-resident phenotype in human lung and blood. Proc. Natl Acad. Sci. USA 118, e2016580118 (2021).
Chen, K. Y. et al. Inflammation switches the chemoattractant requirements for naive lymphocyte entry into lymph nodes. Cell 188, 1019–1035 (2025).
Ceglia, S. et al. An epithelial cell-derived metabolite tunes immunoglobulin A secretion by gut-resident plasma cells. Nat. Immunol. 24, 531–544 (2023).
Frascoli, M. et al. Skin γδ T cell inflammatory responses are hardwired in the thymus by oxysterol sensing via GPR183 and calibrated by dietary cholesterol. Immunity 56, 562–575 (2023).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. eLife 5, e19760 (2016).
Barnea, G. et al. The genetic design of signaling cascades to record receptor activation. Proc. Natl Acad. Sci. USA 105, 64–69 (2008).
Huang, R.-S., Lai, M.-C., Shih, H.-A. & Lin, S. A robust platform for expansion and genome editing of primary human natural killer cells. J. Exp. Med. 218, e20201529 (2021).
Jerby-Arnon, L. et al. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 175, 984–997 (2018).
Chen, Y., Lun, A. T. L. & Smyth, G. K. From reads to genes to pathways: differential expression analysis of RNA-seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Res. 5, 1438 (2016).
Li, W. et al. Quality control, modeling, and visualization of CRISPR screens with MAGeCK-VISPR. Genome Biol. 16, 281 (2015).
Waltman, L. & van Eck, N. J. A smart local moving algorithm for large-scale modularity-based community detection. Eur. Phys. J. B 86, 471 (2013).
Kim, Y.-M., Tsai, M. & Jerby, L. Redirecting NK and T cells to breast and ovarian cancer tumors via metabolite-sensing receptors. figshare https://doi.org/10.6084/m9.figshare.31302178 (2026).
Tang, F. & Li, J. Datasets for the paper ‘A pan-cancer single cell panorama of human natural killer cells’. Zenodo https://doi.org/10.5281/zenodo.8275844 (2023).
Acknowledgements
We thank M. C. Bassik (Stanford University) for providing the HTLA cell line and K. Wucherpfennig (Dana–Farber Cancer Institute) and R. Hernandez-Lopez (Stanford University) for providing plasmids. Y.-M.K. is a recipient of the Stanford School of Medicine Dean’s Postdoctoral Fellowship and of the Basic Science Research Program fellowship (RS-2023-00244834) through the National Research Foundation of Korea, funded by the Ministry of Education. L.J. is a Chan Zuckerberg Biohub Investigator and an Allen Distinguished Investigator. This study was supported by the Burroughs Wellcome Fund (1019508.01; to L.J.), Liz Tilberis Early Career Award from the Ovarian Cancer Research Alliance (889076; to L.J.), the Alba Tull Molecular Therapeutics Award for Innovative Medicines (to L.J.), Department of Defense, Office of the Congressionally Directed Medical Research Programs (HT94252510234; to L.J.), Under One Umbrella, Stanford Women’s Cancer Center, Stanford Cancer Institute, a National Cancer Institute-designated Comprehensive Cancer Center (311982; to L.J.), Discovery Innovation Funds (to L.J.) and funds from the Department of Genetics at Stanford University (to L.J.).
Author information
Authors and Affiliations
Contributions
Y.-M.K. and L.J. designed the study. Y.-M.K., M.K.T., R.V.A., C.S., K.F. and O.L. performed the experiments under the supervision of L.J. Y.-M.K. performed the breast cancer screens and follow-up experiments. M.K.T. performed the ovarian cancer screens. Y.-M.K. and L.J. visualized and analyzed data. Y.-M.K. and L.J. wrote the paper. L.J. obtained funding. L.J. supervised the study. All authors reviewed and approved the paper.
Corresponding author
Ethics declarations
Competing interests
Stanford University has filed a patent application (PCT/US2025/023673) on the basis of this work, and Y.-M.K. and L.J. are named as inventors. The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Minsoo Kim, Lili Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Nick Bernard, in collaboration with the rest of the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental design of CRISPRa screens in NK-92 cells.
a, CRISPRa sgRNA library design based on scRNA-Seq data. Top: Genes stratified by the number of cell subtypes in which they are significantly overexpressed in the tumor compared to blood samples of breast cancer patients (Zhang et al., 2021), shown for the genes targeted (left) or not (right) in the customized sgRNA library. Bottom, NK pan-cancer atlas analyses: Genes stratified by the number of cancer types in which they are significantly overexpressed in the NK cells from the tumor compared to blood samples, shown for the genes targeted (left) or not (right) in the customized sgRNA library. b, Validations of CRISPR/dCas9-SAM system in NK-92 via flow cytometry of target gene protein expression on the cell surface for two different sgRNAs targeting FCGR3A (encoding for CD16; top) and CXCR6 (bottom).
Extended Data Fig. 2 Tumor infiltration hits identified in vivo across different breast cancer models.
(a-g) Fold change of sgRNAs in tumor vs. lung samples or NK cell library shown for sgRNAs targeting indicated genes. Combined analysis of all in vivo breast cancer models when comparing tumors to NK-92 cell library samples (a). In vivo screen in hIL-2-NOG mice (b,c) and NSG mice with subcutaneous MDA-MB-231 tumors (d,e), and in vivo screen in NSG mouse with orthotopic MDA-MB-231 tumors (f,g).
Extended Data Fig. 3 Isogenic NK-92 and PRESTO-tango reporter lines confirm thGPR-dependent sensing of and migration to chemoattracting metabolites.
a, Schematics of PRESTO-tango reporter cell lines where GPCR conformational changes in response to a ligand result in luciferase transcription for bioluminescence readouts. b, PRESTO-tango reporter cell lines generated for CXCR2, C5AR1, GPR34, GPR84, and GPR183 confirm IL8, C5a, LysoPS, 6-OAU, and 7α,25-OHC as the respective GPCR agonists based on the bioluminescence readouts (y axis, fold change; n = 3 technical replicates per group). c, Transwell flow cytometry based quantification of isogenic NK-92 cell migration, confirming GPCR-dependent migration to the respective ligand (n = 4 technical replicates for NTC, CXCR2, C5AR1, GPR84, and GPR183, and n = 5 technical replicates for ORF control and GPR34). (b-c) Data are representative of two independent experiments and presented as the mean ± SEM; ****P < 0.0001, ***P < 0.001, **P < 0.01, and *P < 0.05, one-way ANOVA with Dunnett’s multiple comparisons. d, qPCR quantification of GPR183, GPR84, GPR34, CXCR2, and C5AR1 expression in control NK-92 cells and upon CRISPRa of the target GPCRs (mean ± SEM; n = 3 technical replicates per group). mRNA levels are normalized to GAPDH expression in each cell line. ****P < 0.0001 and ***P < 0.001, two-tailed unpaired Student’s t-test. e, qPCR quantification of GPR18 (left) and GPR183 (right) expression in isogenic NK-92 cells with CRISPRa of GPR18, C5AR1, and GPR183 (n = 1 per group). mRNA levels are normalized to GAPDH expression in each cell line. f, Cell surface expression levels of CXCR2, GPR183, and C5AR1 in NK-92 cells post-transduction with a NTC sgRNA (left) or with the target gene sgRNAs (middle) and 5 days after fluorescence-activated cell sorting (FACS) of successfully transduced NK-92 cells (right). g-h, Combinatorial migratory effects of CXCR2 and GPR183. g, Surface expression of CXCR2 and GPR183 assessed by flow cytometry in NK-92 cells transduced with control, CXCR2, GPR183, or both CXCR2 and GPR183 ORF vectors. h, Transwell flow cytometry based quantification of NK-92 cells in the presence of 100 ng/ml CXCL8 or 100 nM 7α,25-OHC (mean ± SEM; n = 3 technical replicates per group). ****P < 0.0001, one-way ANOVA with Dunnett’s multiple comparisons.
Extended Data Fig. 4 In vitro CRISPRa screens.
a, Experimental scheme and image of spheroid-NK coculture system. b, Chemotaxis NK-92 CRISPRa screens: Significance (MAGeCK p-values combined via Fisher’s method with BH correction for multiple hypothesis testing, Methods, y axis) and log-transformed fold change (x axis) of each target gene in NK-92 cells that have migrated to MCF7 (left) and MDA-MB-231 (right) supernatant compared to the NK-92 cell library. c-d, Log-transformed fold change of sgRNAs targeting top hits in chemotaxis (c) and spheroids (d) screens, shown separately for the two breast cancer cell lines (c) and across time points (d). Illustrations in a created in BioRender; Jerby Lab https://biorender.com/c1gnk9g (2026).
Extended Data Fig. 5 thGPR expression in scRNA-seq data of immune cells of breast cancer patients15.
a, Expression of hits in the tumor and blood samples per cell subtypes. (b-i) GPR183 (b), GPR84 (c), GPR34 (d), GPR18 (e), LPAR2 (f), FPR3 (g), C5AR1 (h), and CXCR2 (i) expression, stratified by cell type (row), patient and site (blood or tumor; column). cDC1, conventional type 1 dendritic cells; cDC2, conventional type 2 dendritic cells;mDC, mature DC; pDC, plasmacytoid DC.
Extended Data Fig. 6 thGPR expression in NK cell pan-cancer atlas16.
a-b, Expression (log1p TPM) of thGPRs and top GPR183 signature genes in NK cells shown: (a) in the UMAP of NK cell scRNA-seq data from breast cancer patients, matching Fig. 1c, and (b) across tumor and blood samples from 22 cancer types.
Extended Data Fig. 7 NK-92 Perturb-seq screen confirms CRISPRa specificity and depicts the transcriptional impact of thGPR genetic and molecular perturbations.
a-b, examining off-target and on-target CRISPRa effects in Perturb-seq data. Expression of the 8 thGPRs (columns) in the NK-92 Perturb-seq data: Average expression (color) and percentage of expressing cells (dot size) stratified based on the sgRNA detected and conditions (rows), where (+) and (-) denote that the cells were cultured with and without 7α,25-OHC treatment, respectively. In (a), all cells are shown. In (b), only cells that express the pertaining sgRNA target gene and passed the functional RNA-based filter are shown. c, Intersection size (y axis) of genes differentially expressed under different conditions (bottom rows), that is, in GPR183A+ and C5AR1A+ NK-92 cells cultured with (+) and without (-) 7α,25-OHC treatment compared to control NK-92 cells under the same conditions, and in control NK-92 cells in response to 7α,25-OHC treatment. The dots represent the gene sets considered. The top bars denote the size of the intersection between the different sets (2-3 dots) or the size of the set that does not overlap any other set (single dot). The left horizontal bars denote the size of the differentially expressed gene (DEG) sets for each perturbation.
Extended Data Fig. 8 Models and readouts of GPR183 function in the cancer-lymphocyte interplay.
a, Expression of GPR183 ligand synthesis genes in a collection of 70 breast cancer cell lines. Left: The top bars stratify breast cancer cell lines by the GPR183 ligand synthesis genes they express (bottom rows). The number of breast cancer cell lines expressing the pertinent gene is shown in the bottom-left bars. Right: Expression (log1p TPM) of the GPR183 ligand synthesis genes CYP27A1, CYP7B1, and CH25H, shown separately for the DCIS/non-cancerous (n = 6) and invasive breast carcinoma cell lines (n = 64). P-values were quantified via a one-tailed unpaired Student’s t-test, comparing the expression of the gene (log1p-TPM) in the two groups. Boxplots middle line: median; box edges: 25th and 75th percentiles; whiskers: most extreme points that do not exceed ± IQR x 1.5; further outliers are marked individually with circles (minima/maxima). b, Flow cytometry plots of primary human NK and CD8 T cells isolated from PBMCs. Left: NK cell purity assessed based on the T cell marker CD3 and NK cell marker CD56. Right: CD8 T cell purity assessed based on CD8a and human TCRab. c, Retroviral vectors used to express either control-GFP or human GPR183-GFP constructs (top). Representative flow cytometry analysis of primary NK cells after transduction (bottom). d, Flow cytometry plots show the gating strategy to detect and quantify the relative abundance of injected NK-92 cells in the tumor. Only live (7-AAD negative) cells were considered. mCD45−hCD45+ were identified as injected NK-92 cells in the NSG mice. mCherry+/GFP+ ratio was calculated. mCD45, mouse CD45; hCD45, human CD45; GFP, green fluorescent protein. e, Fold increases in IFNγ (left) and CD107a (middle) expression at baseline and upon PMA/Ionomycin stimulation, and cytotoxicity against breast cancer cells, in GPR183OE CAR T cells (right), primary NK cells (pNK), and NK-92 cells relative to their respective controls (n = 3 CAR T cell donors per group, n = 2 pNK cell donors per group, and n = 6 technical replicates per group for NK-92 cells). Statistical significance was determined via paired two-sided t-tests for CAR T and pNK cells (pairing donors), two-sided unpaired t-tests for NK-92 cytotoxicity, and two-sided mixed effect tests for NK-92 IFNγ and CD107a production, testing whether GPR183OE cells significantly diverge from the matching control ones.
Extended Data Fig. 9 Generating and validating CAR NK-92 cells.
a, GPR183 impact on NK-92 in vivo efficacy (biological replicate of the experiment depicted in Fig. 5b). Left: MDA-MD-231 tumor growth (mean ± SEM) in NSG mice treated with NTC NK-92 (n = 5 mice), or GPR183A+ NK-92 (n = 5 mice); two-way ANOVA with Sidak correction. Right: Individual tumor growth plots. b, EpCAM surface levels in MDA-MD-231 and MCF7 cells quantified via flow cytometry. c, FACS of anti-EpCAM-CAR+ and GPR183OE NK-92 cells via mCherry and GFP, respectively (left), followed by confirmation of GPR183 surface expression via an anti-GPR183 antibody (middle) and confirmation of CAR expression via anti-human IgG (right). d, EpCAM surface levels in control (NTC) and EpCAM KO MDA-MB-231 cells quantified via flow cytometry. e, Antigen-dependent cytotoxicity of control and GPR183OE anti-EpCAM-CAR NK-92 cells in coculture with control (NTC, EpCAM wildtype) and EpCAM KO MDA-MB-231 cells at 1:1 E:T ratio for 24 h (n = 3 technical replicates per group). f, GPR183 impact on CAR NK-92 cell in vivo efficacy (biological replicate of the experiment shown in Fig. 5d). NSG mice were treated with cell-free media (n = 3 mice), anti-EpCAM-CAR NK-92 (n = 5 mice), or GPR183OE-anti-EpCAM-CAR NK-92 (n = 5 mice). Left: MDA-MD-231 tumor growth (mean ± SEM); ****P < 0.0001, two-way ANOVA with Sidak correction for multiple hypotheses testing. Right: Tumor weight (mean ± SEM) after tissue dissection at day 35; one-way ANOVA was performed with Dunnett’s multiple comparisons.
Extended Data Fig. 10 CAR T cell generation and GPR183-dependent in vivo infiltration and in vivo efficacy.
a, Surface expression of anti-EpCAM-CAR was assessed by flow cytometry. GFP+CAR+ T cells were sorted (left). CAR T cells were transduced with either control or GPR183-ORF vectors and surface GPR183 expression was assessed by flow cytometry (right). b, A gating strategy for in vivo CAR T cell experiments. Only live (7-AAD negative) cells were considered. mCD45−hCD45+hCD8+ cells were identified as injected CAR T cells. IFNg and CD107a expression were further assessed at baseline and upon PMA/ionomycin stimulation. c, GPR183 increases CAR T cell tumor infiltration (biological replicate of the experiment shown in Fig. 6f with T cells derived from another human donor). Flow cytometry-based quantification of in vivo accumulation of CAR T cells in lung, spleen, and tumor. d, IFNγ and CD107a expression in tumor-infiltrating CAR T cells. (c-d) Data are presented as mean values ± SEM of GPR183OE-CAR T cells (n = 6 mice) and control-CAR T (n = 5 mice) compared via two-tailed unpaired Student’s t-tests. e, GPR183 increases CAR T cell in vivo efficacy (biological replicate of the experiment shown in Fig. 6h,i with T cells derived from another human donor). Growth and survival curve of EpCAMOE MDA-MD-231 breast cancer tumors in NSG mice treated with anti-EpCAM-CAR T (n = 6 mice), or GPR183OE anti-EpCAM-CAR T (n = 7 mice). In (e) data are shown as the mean ± SEM (left) and individual growth curves per mouse (middle). Data analysis was performed using two-way ANOVA with Sidak correction for multiple hypotheses testing for growth curve and log-rank test for survival curve.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, table captions and references.
Supplementary Tables 1–8 (download XLSX )
Supplementary Table 1. a–c, sgRNA library information: target gene annotation and tumor enrichment in the breast cancer scRNA-seq data (a), tumor enrichment in the primary NK cell pan-cancer atlas (b) and spacer sequence per sgRNA (c). Supplementary Table 2. a–g, In vivo NK-92 CRISPRa screens for infiltration to breast cancer tumors: MAGeCK summary statistics (a) and output per screen (b–g). Supplementary Table 3. a–g, In vivo NK-92 CRISPRa screens for infiltration to ovarian cancer tumors: MAGeCK summary statistics and outputs from analyses of the customized (a–d) and large-scale (e–g) screens. Supplementary Table 4. thGPR information including G-protein and ligands previously reported. Supplementary Table 5. a–h, In vitro CRISPRa screen MAGeCK summary statistics (a) and outputs per screen (b–h). Supplementary Table 6. a–q, Perturb-seq screen results: differentially expressed genes identified per target gene per condition (a), 7α,25-OHC differentially expressed genes in control and GPR183A+ cells (b), MAST differential gene expression analysis output per target gene and condition (c–p) and differential expression of NK and T cell signatures (q)30,31. Supplementary Table 7. a and b, Expression patterns of GPR183 ligand synthesis genes (CYP27A1, CYP7B1 and CH25H) in a collection of 70 breast cancer cell lines: gene-level coexpression (a) and gene set enrichment analysis (b) results. Supplementary Table 8. a–g, Gene sequences for Illumina sequencing primers (a), ORFs (b, d and e), CRISPRa or knockout sgRNA (c and f) and qPCR primers (g).
Source data
Source Data Figs. 4–7 (download XLSX )
Statistical source data.
Source Data Extended Data Figs. 3 and 8–10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kim, YM., Tsai, M.K., Sun, C. et al. Engineering NK and T cells with metabolite-sensing receptors to target solid tumors. Nat Immunol (2026). https://doi.org/10.1038/s41590-026-02473-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41590-026-02473-y