Abstract
Epigenetic editing holds the promise of durable therapeutic effects by silencing disease-causing genes without changing the underlying DNA sequence. In this study, we designed an epigenetic editor to target human PCSK9 and thereby induce DNA methylation at this locus. A single administration of lipid nanoparticles encapsulating mRNA encoding this epigenetic editor was sufficient to drive near-complete silencing of human PCSK9 in transgenic mice. Silencing was durable for at least 1 year and was fully maintained after partial hepatectomy–induced liver regeneration. In addition, we showed reversibility of epigenetic editing in mice with previously silenced PCSK9 upon treatment with a targeted epigenetic activator designed to demethylate the PCSK9 locus. Notably, in cynomolgus monkeys, a single administration of the epigenetic editor potently and durably decreased circulating PCSK9 protein levels by approximately 90% with concomitant reduction in low-density lipoprotein cholesterol levels by approximately 70%. These findings demonstrate the therapeutic potential of durable and reversible epigenetic editing in vivo and support the development of epigenetic editor–based treatment for hypercholesterolemia.
Similar content being viewed by others
Main
Epigenetic editing has emerged as a powerful approach to regulate gene expression in vitro and in vivo. Inducing DNA methylation at CpG dinucleotide sites located at promoter regions has proven to be effective in durably locking genes in a silenced state1,2,3. DNA methylation is a repressive mark that is durable and faithfully propagated through cell division by the activity of the DNA methyltransferase DNMT1, which recognizes the hemi-methylated state resulting from DNA replication and restores the methyl group to newly synthesized cytosines4,5. Unlike gene editing methods that rely on a single-strand DNA break (nick), such as base and prime editing, or a double-strand DNA break (cut), such as CRISPR–Cas9 nuclease editing, targeted epigenetic editing does not disrupt the integrity of the DNA sequence, thus avoiding potential genotoxic risks associated with nuclease editing6,7,8 and with much lower frequency, base and prime editing9. Epigenetic editing is, thus, a promising strategy for silencing or activating gene of interest through targeted methylation or demethylation, respectively, when the primary DNA sequence does not need to be changed. In contrast, base editing and prime editing were developed for correcting disease-associated genetic variants through single nucleotide change or targeted insertion of a new DNA sequence10,11,12. Base editing has also been used in vivo to decrease expression of gene of interest by introducing nonsense mutations or disrupting splicing site at an exon–intron boundary13,14,15.
Durable silencing using programmable epigenetic editors (EEs) was demonstrated in human cells in vitro where gene silencing was maintained for many months1,3. These epigenetic editors comprised a combination of DNA methyltransferase and a KRAB-based transcriptional repressor domains fused to a DNA-binding domain either as single or multiple fusion proteins1,3. Although initial proof of concept of in vivo epigenetic editing showed durable silencing of the endogenous mouse Pcsk9 gene in the liver2, validation of epigenetic editing as a potentially viable clinical application requires demonstration of activity, durability and safety in non-human primates (NHPs).
Additionally, although gene editing and base editing approaches for therapeutic target inhibition have been demonstrated in preclinical species14,15,16,17,18,19 and in humans20,21,22, the ability to reverse these changes in the DNA sequence in vivo has not been published and would require identification of a new DNA-targeting moiety (that is, new guide RNA (gRNA)) to account for those change(s). In contrast, an epigenetic activator designed to remove methyl marks on CpGs could theoretically be deployed in previously silenced tissue(s) in vivo at the same genomic location using the same targeting DNA-binding domain because the underlying DNA sequence after epigenetic editing has not changed.
We report here the development of an EE targeting human PCSK9 for the durable reduction of low-density lipoprotein cholesterol (LDL-C) levels in vivo. Activity, potency and durability of the PCSK9-targeting EE was measured in transgenic mice carrying the human PCSK9 genomic locus and in NHPs, whereas specificity was assessed in primary human hepatocytes.
Results
Identification of a potent PCSK9-targeting EE
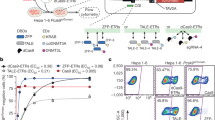
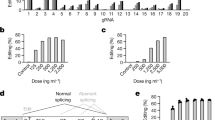
To evaluate the ability of a DNA-based methylation approach to silence the human PCSK9 gene, we designed a single EE construct composed of DNMT3A, DNMT3L and a KRAB transcriptional repressor domain fused to catalytically inactive Streptococcus pyogenes Cas9 (dCas9) (Fig. 1a and Supplementary Fig. 1). We then performed a gRNA screen by transiently co-transfecting the EE expression plasmid with DNA fragments encoding individual gRNA expression cassettes tiling the human PCSK9 locus, approximately 1 kilobase (kb) upstream and downstream of the transcription start site (TSS) in HeLa cells. These cells are amenable to high-throughput screening and show similarly low levels of CpG methylation near the PCSK9 TSS compared to human liver hepatocytes (Fig. 1b). We measured the level of PCSK9 secreted into the media 7 d after EE/gRNA transfection and found that multiple gRNAs robustly reduced PCSK9 levels (up to 80% inhibition), with the most active guides being closest to the TSS (Fig. 1b and Supplementary Table 1). We confirmed the activity of the top 40 gRNA hits from the primary screen by co-transfecting in vitro produced mRNA encoding for the EE and chemically synthesized gRNA into HeLa cells at an effector-to-gRNA mass ratio of 2:1, and we found that most gRNA hits reduced secreted PCSK9 to a level similar to that of the wild-type (WT) Cas9 nuclease positive control, which was durable up to 28 d after transfection (Fig. 1c and Supplementary Table 2). In contrast, CRISPR interference (CRISPRi) controls, which lack DNA methyltransferase domains, show robust but transient reduction of secreted PCSK9 (Fig. 1c). To evaluate the ability of the EE to silence PCSK9 in primary human hepatocytes (PHHs), we selected gRNAs that were the most active and durable in HeLa cells (Fig. 1b,c), had no perfect match anywhere in the human genome outside of the PCSK9 locus and had a perfectly homologous spacer sequence in the cynomolgus macaque PCSK9 locus to allow for testing in NHPs. Because background statin therapy would likely complement treatment with our EE and because statins have been shown to regulate PCSK9 expression at the transcriptional level23, we also evaluated the response of EE-silenced cells to an acute challenge with statins. As expected, we found that statin treatment increased secreted PCSK9 levels from cells treated with transfection reagent only (Extended Data Fig. 1). In contrast, suppression of PCSK9 secretion by any of the top five gRNAs identified in Fig. 1c was maintained after treatment with statin, suggesting that robust epigenetic editing at the PCSK9 locus is maintained in the presence of statin (Extended Data Fig. 1). Finally, to ensure sufficient expression of our construct and mimic in vivo delivery of RNA-based drug substance to the liver, EE mRNA and selected gRNAs were formulated into lipid nanoparticles (LNPs) and incubated with PHHs in the presence of APOE3 recombinant protein to facilitate uptake via an LDL receptor (LDLR)-dependent mechanism24. Treatment of PHHs with LNPs containing EE mRNA and any of the five gRNA candidates resulted in rapid and sustained suppression of PCSK9 secretion for at least 14 d (Fig. 1d). In contrast, treatment with a CRISPRi control showed a rapid but non-durable reduction of PCSK9, consistent with the transient nature of non-methylation-based silencing via CRISPRi (Fig. 1d). Together, our data demonstrate that transient application of an EE can efficiently and durably reduce PCSK9 protein in in vitro cultured PHHs.
a, Schematic outline of the architecture of PCSK9 EEs b, Primary screen evaluating 240 candidate gRNAs targeting the human PCSK9 gene using a spCas9-based EE. Each point represents the average of two independent measurements of secreted PCSK9 protein levels 7 d after transfection; the location of each point along the x axis indicates the position of the gRNA relative to the distance (in nucleotides) to the PCSK9 gene TSS. PCSK9 protein levels in cells transfected with a non-targeting (NT) gRNA or effector only (no gRNA) are shown with a dotted line. CpG location (first row); methylation percentage (0–100%) of each CpG dinucleotide at the PCSK9 locus measured by whole-genome bisulfite sequencing (WGBS) or hybrid capture in neurons (second row); HeLa cells (third row); liver hepatocytes (fourth row); and DNaseI accessibility in human liver (fifth row) are shown below the graph and mapped onto the CpG island (CGI) and PCSK9 5′ gene region. c, The top 40 gRNA were selected from b and were evaluated for their ability to potently and durably silence PCSK9 in HeLa cells for up to 28 d. Individual data points and means are shown (n = 2 replicates per experimental condition). Results are expressed as percent of secreted PCSK9 protein in cells treated with transfection (txn) reagent only. d, The top five gRNAs were selected based on their activity and durability in HeLa cells (from c) as well as having full cross-reactivity with the cynomolgus macaque PCSK9 gene. PHHs isolated from PXB mice were treated with LNPs containing selected gRNAs and EE mRNA. Results are shown as mean ± s.d. (n = 4 replicates per gRNA). For b and c, WT Cas9 served as a control for durable silencing of PCSK9; for c and d, CRISPRi served as a control for non-durable silencing of PCSK9.
To determine the in vivo activity and potency of our candidate human PCSK9-targeting gRNAs, we used mice expressing the human PCSK9 gene whose expression is driven by its own promoter (PCSK9-Tg)25,26. These PCSK9-Tg mice were previously shown to have circulating PCSK9 levels similar to those observed in humans25,26, and DNA methylation analysis of the PCSK9 transgene showed a pattern that mirrors the PCSK9 methylation pattern in PHHs (Extended Data Fig. 2a). We selected the top three candidates with non-overlapping gRNA spacer sequences (gRNA41, gRNA43 and gRNA49; Extended Data Fig. 2b) and formulated them into LNPs as either single gRNA or dual gRNA combinations, together with EE mRNA at a 1:1 gRNA/mRNA or a 0.5:0.5:1 gRNA1/gRNA2/mRNA weight ratio for single and dual gRNA evaluation, respectively. We identified a single gRNA (gRNA41) and a gRNA pair (gRNA41 + gRNA49) that robustly decreased circulating PCSK9 in PCSK9-Tg after a single administration of LNPs at a near-saturating dose of 0.75 mg kg−1 (Extended Data Fig. 2c). Interestingly, we observed that gRNA43, which produced durable silencing in vitro (Fig. 1c,d), showed only transient knockdown in vivo (Extended Data Fig. 2c). It is possible that the potency threshold needed to achieve methylation-based silencing in vivo is higher than that of in vitro cell cultures but that engagement of KRAB-based transcriptional repression (CRISPRi) has a lower threshold, which was met when gRNA43 was tested in PCSK9-Tg mice at near-saturating dose. To further evaluate the potency of PCSK9-targeting EEs, we tested LNPs formulated with the best single or dual gRNA at a sub-saturating dose of 0.2 mg kg−1. Mice treated with the dual gRNA combination (gRNA41 + gRNA9) showed more robust silencing than those receiving the single gRNA (gRNA41), indicating that a dual gRNA combination was more effective at achieving epigenetic silencing of the PCSK9 locus (Extended Data Fig. 2d). On the basis of these studies, this EE mRNA/dual gRNA combination (hereafter referred to as PCSK9-EE; Supplementary Fig. 2) was selected for additional studies.
Specificity of PCSK9-targeting EE
We next evaluated potential off-target epigenetic editing in PHHs treated with PCSK9-EE by measuring global changes in gene expression and DNA methylation. Efficient on-target activity was evidenced by reduced PCSK9 protein (Fig. 2a) and transcript levels (Fig. 2b). To detect changes in gene expression that may result from off-target methylation in regulatory elements controlling gene transcription, such as promoter and enhancer regions, we performed deep-read RNA sequencing (RNA-seq) (Fig. 2c). The most significantly affected gene after PCSK9-EE treatment was PCSK9 itself as compared to cells treated with EE alone. No other genes, except for ENSG00000285976, a gene with low expression (<4 transcripts per million (TPM)) and no change in methylation in hepatocytes (Extended Data Fig. 3a), were significantly changed by PCSK9-EE (log2 fold change (FC) ≤ 2 and adjusted P ≥ 1 × 10−5). We then assessed methylation by hybrid capture of approximately 4 million CpGs distributed across the genome. Treatment with PCSK9-EE led to a large increase in methylated CpGs at the PCSK9 locus compared to cells treated with EE only (beta value difference > 0.2 and P < 1 × 10−10; Fig. 2d). Smaller increases in CpG methylation (beta value difference > 0.2, P < 1 × 10−10) were detected at potential off-target sites (Fig. 2d), but these increases did not result in significant changes in gene expression as measured by RNA-seq (Fig. 2c and Extended Data Fig. 3b). To further evaluate potential off-target CpG methylation by PCSK9-EE, we performed whole-genome methylation sequencing (WGMS) across more than 30 million genomic CpGs. As shown in Fig. 2e, changes in CpG methylation with the most significant and profound effect size were specific to PCSK9. No other genes within 20 kb of any differentially methylated region (DMR) as determined by WGMS (P < 1 × 10−10) had both a significant change in methylation at the DMR (>0.2) and a significant change in gene expression (FC > 2) (Fig. 2f). CpG methylation of DMRs with either a significant change in methylation in at least one of the methylation assays or within 20 kb of a TSS of a gene with a significant change in gene expression and TPM plots for the nearby genes (TSS within 20 kb) are shown in Extended Data Fig. 3c–i. All genes near these DMRs have low expression or do not show a change in gene expression, further confirming the specificity of the PCSK9-EE with minimal off-target methylation and no functional consequences as measured by gene expression.
a. Activity of the LNP formulation with the top-ranked PCSK9 EE (PCSK9-EE) in PHHs isolated from chimeric mice with a humanized liver. Each point represents the average of three independent measurements of secreted PCSK9 protein levels at baseline and at 6 d and 15 d after treatment. Results are shown as mean ± s.d. PHHs treated with APOE only or LNP formulation containing the effector without gRNA (effector only) served as negative controls. b–f, Assessment of the specificity of PCSK9-EE was performed using PHHs obtained at 15 d after treatment. b, Specificity testing was assessed using RNA-seq on three independent replicates of each control condition (ApoE Only and Effector Only) and PCSK9-EE. On-target PCSK9 TPM from RNA-seq for each replicate are shown in the dot plot. c, Volcano plot of RNA-seq data comparing PCSK9-EE versus Effector Only control. Thresholds for differential expression: adjusted P value (DEseq2 Wald test, two-sided, Benjamini–Hochberg multiple comparisons adjustment) < 1×10−5, log2FC > 1 or log2FC < −1. PCSK9 is shown as a yellow circle; off-target (upregulated) DEG is shown as a navy blue circle; all other genes below the thresholds are shown as light blue circles. d, Specificity of methylation at CpG-enriched sites was measured using a Twist Human Methylome Hybrid Capture Methylation Sequencing assay. Volcano plot of CpG methylation comparing PCSK9-EE versus Effector Only control. Individual CpGs are colored according to whether they were called as a DMR at the PCSK9 locus (yellow), at an off-target genomic region (navy blue) or were not part of a DMR (light blue). DMR thresholds were set as P value (DSS Wald test, two-sided, unadjusted) < 1 × 10−10, beta value difference < −0.2 or beta value difference > 0.2. e, Manhattan plot of genome-wide methylation, as determined by a WGMS assay, comparing PCSK9-EE versus Effector Only control. Benjamini–Hochberg (false discovery rate (FDR)) adjusted P values for each CpG (DSS Wald test, two-sided) are plotted versus genomic coordinate for each CpG. Differentially methylated CpGs within the PCSK9 DMR are shown in yellow. The DMR threshold was set as P value (DSS Wald test, two-sided, unadjusted) < 1 × 10−10. f, Scatterplot showing methylation difference of DMRs from WGMS (y axis) versus log2FC from RNA-seq (x axis) of all genes within 20 kb of each DMR for the PCSK9-EE versus Effector Only control comparison. PCSK9 gene is shown in yellow. Thresholds (gray dashed lines) are set as methylation (beta value) difference > 0.2 or < −0.2, RNA-seq log2FC > 1 or < −1. DEG, differentially expressed gene.
Effects of PCSK9-targeting EE in mice
We next sought to evaluate the PCSK9-EE as a potential one-and-done approach to reduce PCSK9 in vivo. We formulated PCSK9-EE in LNPs and administered it at a pharmacologically saturating dose (3 mg kg−1) in PCSK9-Tg mice (Fig. 3a). We also formulated in LNPs and administered to PCSK9-Tg mice CRISPRi and WT Cas9 targeting PCSK9 as controls for transient and durable silencing, respectively (Fig. 3a). As expected, treatment with CRISPRi led to a rapid and robust reduction in circulating PCSK9, with levels returning to baseline within 7 d (Fig. 3b). Consistent with previously reported observations17,18, treatment with WT Cas9 nuclease led to approximately 90% reduction in circulating PCSK9, which was maintained for the duration of the study (1 year; Fig. 3b). The PCSK9-EE achieved near-complete (>98%) reduction of circulating PCSK9 that, similar to WT Cas9, was durable for at least 1 year in mice (Fig. 3b). Hepatic PCSK9 expression was evaluated in a separate cohort of animals and showed a large reduction in transcript levels (Fig. 3c) that was associated with reduced plasma cholesterol levels (Fig. 3d). To further establish a mechanistic link between PCSK9-EE’s molecular mechanism of action and the robust suppression of PCSK9 in vivo, we measured CpG methylation at the PCSK9 locus using a hybridization capture approach in liver samples obtained 1 month after treatment. Baseline methylation in PCSK9-Tg mice showed hypomethylation at the PCSK9 promoter region, similar to that of liver hepatocytes and consistent with active transcription of PCSK9 in hepatocytes (Fig. 3e). Transient expression of PCSK9-EE led to robust methylation of multiple CpGs across the TSS when examined 28 d after treatment. This methylation signature and abundance was very similar to that of livers isolated from mice treated with PCSK9-EE for 1 year (methylation levels at day 28: 47% ± 4 and at day 365: 43% ± 8; Fig. 3e). In contrast, treatment with CRISPRi or WT Cas9 nuclease did not modify CpG methylation at the PCSK9 locus, which is consistent with their mechanisms of action (Fig. 3e).
a, Schematic outline of the in vivo study in transgenic mice (PCSK9-Tg) carrying the human PCSK9 genomic locus. The mice were treated with LNPs formulated with the top-ranked PCSK9 EE (PCSK9-EE), CRISPRi or WT Cas9 payload. Illustration was created with BioRender. b, Circulating PCSK9 protein levels in PCSK9-Tg over a 1-year period after a single administration of an LNP formulation with PCSK9-EE (n = 6 mice per group). c–e, Effect of a single administration of an LNP formulation with PCSK9-EE on PCSK9 mRNA (c), total plasma cholesterol (d) and CpG methylation levels in liver (e) in PCSK9-Tg mice 1 month after treatment (n = 4 mice per group). For e, liver methylation data from all PCSK9-Tg mice treated with PCSK9-EE for 1 year (n = 6, from b) are also included. CpG location (first row) and the methylation percentage (0–100%) of each CpG dinucleotide at the PCSK9 locus as measured by WGBS in cells expressing PCSK9 (liver hepatocyte WGBS; second row) or not expressing PCSK9 (neuron WGBS; third row) are shown. Vehicle-treated animals received a single administration of saline solution. CRISPRi served a control for robust but transient silencing of PCSK9, whereas WT Cas9 served as a control for durable silencing of PCSK9. For b–d, results are shown as mean ± s.d. For c and d, statistical analysis was performed by one-way ANOVA followed by two-tailed Dunnett’s test. For c, ***P = 0.000893; ****P = 000005 versus vehicle-treated mice. For d, ***P = 0.000253; ****P = 0.000024 versus vehicle-treated mice. For e, CpG methylation profiles for all analyzed samples are shown.
To further confirm the durability and maintenance of epigenetic editing in rapidly dividing hepatocytes, we used a partial hepatectomy (PHx) mouse model of liver regeneration27, where mice previously treated with PCSK9-EE undergo two-thirds PHx and are subsequently monitored until full liver regeneration is achieved (Fig. 4a). Treatment with PCSK9-EE or WT Cas9 elicited robust reduction of circulating PCSK9 in PCSK9-Tg mice, which was completely maintained after the PHx surgery (Fig. 4b). Monitoring of methylation marks at the PCSK9 locus in livers from mice that underwent PHx showed a virtually identical epigenetic signature to that of sham-operated mice as well as resected liver lobes from hepatectomized mice (Fig. 4c). Together, these data indicate that epigenetic editing is durable in both the homeostatic and regenerating liver and is maintained through cell division.
a, Schematic outline of the in vivo PHx study. The timing of the PHx (or sham) procedure and the length of time allowed for full liver regeneration before liver sample collection are highlighted. Illustration was created with BioRender. b, Circulating PCSK9 protein levels after a single administration of an LNP formulation with the top-ranked PCSK9 EE (PCSK9-EE) in PCSK9-Tg mice before and after PHx (n = 6) or sham (n = 5) procedures. The PHx or sham procedure was performed on day 35. WT Cas9 served as a control for durable silencing before and after PHx (n = 6) or sham (n = 6) procedures. Control animals received saline (vehicle) and were also subjected to pre-PHx and post-PHx (n = 8) or sham (n = 3) procedures. c, Effect of a single administration of an LNP formulation with PCSK9-EE on CpG methylation levels in liver from PCSK9-Tg mice at 90 d after LNP treatment. Methylation data from the resected liver section after the PHx procedure at day 35 were also included in the analysis. CpG location (first row) and the methylation percentage (0–100%) of each CpG dinucleotide at the PCSK9 locus in cells expressing PCSK9 (liver hepatocyte; second row) or not expressing PCSK9 (neuron; third row) are shown. For b, results are shown as mean ± s.d. For c, CpG methylation profiles for all analyzed samples are shown.
Reversibility of PCSK9-targeting EE effects
To assess whether epigenetic editing of the PCSK9 locus can be reversed, we designed and tested an epigenetic activator composed of the catalytic domain of TET1 fused to S. pyogenes dCas9, termed dCas-Tet, in mice previously silenced with PCSK9-EE (Fig. 5a and Supplementary Fig. 3). Because PCSK9-EE modifies DNA methylation without altering the underlying DNA sequence, we evaluated an mRNA encoding dCas-Tet with the same gRNA combinations used to silence the PCSK9 locus (gRNA41 + gRNA49; Fig. 3 and Extended Data Fig. 2). The reversal experiment was initiated by treating a group of five mice with PCSK9-EE (1 × 1.5 mg kg−1 or 2 × 0.75 mg kg−1) to achieve robust reduction in plasma PCSK9 (Fig. 5b), similar to what we observed in our previous mouse experiments (Figs. 3b and 4b). PCSK9 suppression before reactivation was maintained for 173 d, after which mice received a single administration of an LNP formulation containing dCas-Tet and gRNA41 + gRNA49. Treatment with the PCSK9 epigenetic activator resulted in near-complete normalization of plasma PCSK9 levels (~90% of baseline) within the first 2 weeks, and this was maintained for 8 weeks, after which animals were euthanized to collect liver samples (Fig. 5b). Targeted methylation analysis revealed an overall reduction in CpG methylation at the PCSK9 locus in livers from animals treated with the epigenetic activator when compared to historical data of PCSK9-EE-treated mice but slightly higher than that of the endogenous PCSK9 locus in PHHs (Fig. 5c) or vehicle-treated PCSK9-Tg mice (Figs. 3e and 4c).
a, Schematic outline of PCSK9 silencing in mice using a PCSK9 EE (PCSK9-EE), followed at more than 100 d by treatment with a PCSK9 activator (dCas-Tet). Illustration was created with BioRender. b, Circulating PCSK9 protein levels after a single administration of an LNP formulation with dCas-Tet in PCSK9-Tg previously treated with PCSK9-EE to silence PCSK9. Results are shown as mean ± s.d. (n = 5 mice). c, Effect of a single administration of LNP formulation with dCas-Tet in PCSK9-Tg mice previously treated with PCSK9-EE to silence PCSK9 on CpG methylation levels at 56 d after dCas-Tet treatment. CpG location (first row) and the methylation percentage (0–100%) of each CpG dinucleotide at the PCSK9 locus in cells expressing PCSK9 (liver hepatocyte; second row) or not expressing PCSK9 (neuron; third row). CpG methylation profiles for all analyzed samples are shown for c.
Effects of PCSK9-targeting EE in monkeys
One of the key challenges in translating genomic medicines delivered using LNP formulations from rodents to higher species is ensuring sufficient potency of payload to achieve the desired pharmacology at a tolerated dose28,29. We designed an optimized version of our EE using the same gRNA combination at the same 0.5:0.5:1 gRNA41/gRNA49/mRNA weight ratio that was previously identified (Figs. 3 and 4 and Extended Data Fig. 2), which we refer to as PCSK9-EE-V2 (Supplementary Fig. 4). The analysis of human and cynomolgus PCSK9 loci within proximity to the TSS region (±1 kb) shows a high degree of sequence homology (93.7%) and CpG conservation (84 out of 109 total CpGs; Fig. 6a). Moreover, the targeting region for each selected gRNA for PCSK9-EE-V2 is fully conserved in cynomolgus monkeys. Although no transcription factor (TF) binding data are publicly available for cynomolgus liver tissues, it has been shown that TFs are generally conserved between humans and primates30. To confirm activity and potency of our PCSK9-targeting EE at the cynomolgus PCSK9 locus, we treated primary human and cynomolgus macaque hepatocytes (PHHs and PCHs, respectively) with LNPs formulated with PCSK9-EE-V2. We observed that an LNP formulation containing PCSK9-EE-V2 was able to fully suppress PCSK9 secretion in PCHs but was approximately threefold less potent when compared to PHHs (Fig. 6b), which is consistent with previous observations using mRNA/gRNA/LNP delivered to primary hepatocytes in vitro14. Altogether, this suggests that the mechanisms regulating PCSK9 expression are conserved between humans and cynomolgus monkeys.
a, In silico alignment of CpGs (the molecular target of the EE) between human and cynomolgus monkey around the PCSK9 TSS. Matched and unmatched CpGs are labeled in blue and yellow, respectively (84 out of 112 CpGs are matched). Note that the cynomolgus PCSK9 gene is located on the negative strand; hence, CpGs in the reverse complement (rev-comp) sequence are shown. b, Activity and potency of LNP formulation using the EE PCSK9-EE-V2 in cultured PHHs and PCHs were assessed by measuring secreted PCSK9 protein in the supernatant. IC50 values are indicated. Results are shown as mean ± s.d. (n = 4 replicates per group). c, Dose–response of a single infusion of an LNP formulation with PCSK9-EE-V2 on circulating PCSK9 protein levels (left) and LDL-C (right) in cynomolgus macaques (n = 3 per group). Vehicle-treated animals received a single infusion of saline solution (n = 4). Results are shown as mean ± s.d. Plasma samples were obtained from two of the vehicle-treated animals at days 84 and 98. These data were averaged and plotted at day 91 to better visualize the group mean. d, Effect of a single administration of PCSK9-EE-V2 on CpG methylation levels in liver biopsy samples at 24 d after treatment. CpG location (first row) and the methylation percentage (0–100%) of each CpG dinucleotide at the cynomolgus PCSK9 locus for individual animals are shown. Note that liver biopsies were obtained for only two of the vehicle-treated animals shown in c. e, Pearson’s correlation (r) comparing average cynomolgus PCSK9 (cPCSK9) TSS methylation levels (day 24) versus cynomolgus PCSK9 protein levels (day 21) for individual animals shown in c and d. Two-sided P value is shown. Cyno, cynomolgus; IC50, half-maximal inhibitory concentration.
To evaluate the activity of PCSK9-EE-V2 in NHPs, we performed a dose–response study in cynomolgus monkeys using three animals for each dose tested (0.5, 1.0 and 1.5 mg kg−1), together with four vehicle-treated animals for the control group. A single infusion of LNP containing PCSK9-EE-V2 led to a rapid reduction in circulating PCSK9 levels that was maintained for at least 3 months, resulting in a dose-dependent mean reduction of PCSK9 of 50%, 84% and 89% and LDL-C of 40%, 68% and 64%, respectively, when compared to pre-infusion baseline values (Fig. 6c). No significant difference in PCSK9 or LDL-C reduction was observed between the 1 mg kg−1 and 1.5 mg kg−1 groups, suggesting that a maximal pharmacologic response was achieved with 1 mg kg−1 PCSK9-EE-V2 in NHPs. Liver safety monitoring at 1 d, 3 d and 7 d after dose showed transient increases in alanine transaminase (ALT), aspartate transaminase (AST) and total bilirubin plasma levels that returned to baseline by day 14 (Extended Data Fig. 4). To confirm that changes in circulating PCSK9 were associated with increased CpG methylation at the PCSK9 locus, liver biopsies were performed on day 24 in all monkeys treated with PCSK9-EE-V2 together with two vehicle-treated animals. Transient application of PCSK9-EE-V2 in monkeys led to a robust increase in methylation of multiple CpGs covering a region across the TSS (Fig. 6d). Quantification of CpG methylation in a 500-bp genomic region around the gRNA binding sites showed a dose-dependent increase in average methylation of 20.0%, 30.5% and 35.0%, compared to 5.9% for vehicle-treated monkeys (Fig. 6d), and was strongly correlated with plasma PCSK9 levels (Fig. 6e). Although inter-animal variability is to be expected in such a study (Extended Data Fig. 5), one animal in the 0.5 mg kg−1 group had much lower methylation at the PCSK9 locus (13.2% versus 21.1% and 26.6%; Fig. 6e) with minimal change in plasma PCSK9 (Fig. 6e and Extended Data Fig. 5b, square symbols) and LDL-C (Extended Data Fig. 5g, square symbols). It is possible that lower hepatic uptake of the LNP drug product, reduced endosomal escape and/or translation of the epigenetic editor mRNA or decreased nuclear localization of the epigenetic editor protein may have contributed to reduced CpG methylation in that monkey.
Discussion
The studies presented here demonstrate the potential therapeutic applications of epigenetic editing to modulate gene expression in vivo. We identified a PCSK9-targeting EE that reproducibly induced a CpG methylation signature at the PCSK9 locus, resulting in near-complete suppression of circulating PCSK9 protein in mice with efficacy similar to that of genome editing approaches using adenine base editors14,15,16 and CRISPR–Cas9 (refs. 17,18) but does not require generation of single-strand or double-strand DNA breaks. Although the general architecture of our EE is similar to that described by Cappelluti et al.2, PCSK9-EE-V2 was developed to be active against the human PCSK9 gene and extensively optimized for in vivo potency. Notably, our findings in rodents were extended to NHPs where our PCSK9-targeting EE achieved about 90% reduction in circulating PCSK9, leading to a robust and clinically meaningful reduction of plasma LDL-C (around 70%). Our mouse studies also suggest that epigenetic editing of the human PCSK9 gene is durable for at least 1 year. These data support the development of a single-dose treatment of hypercholesterolemia with reduction in LDL-C on par or better than approved treatments that require chronic dosing, such PCSK9 monoclonal antibodies31,32 and small interfering RNA (siRNA)33. Current treatment algorithms recommend addition of PCSK9 inhibitors for patients not at LDL-C goal on maximally tolerated statin therapy34,35. Importantly, the ability of PCSK9 inhibitors to further reduce LDL-C when added to statin therapy was demonstrated in preclinical experiments including mice36 and NHPs37 and confirmed in multiple clinical settings, irrespective of the mechanism by which reduction of plasma PCSK9 is achieved, such as monoclonal antibodies32,38, RNA interference (RNAi) therapeutics33 or adenine base editor20. This suggests that inhibition of PCSK9 using epigenetic editing has the potential to lower LDL-C as an add-on therapy to statin.
A key aspect of developing our PCSK9-targeting EE was to ensure that our payload was sufficiently potent to retain activity as we progressed preclinical testing from mice to NHPs. In fact, studies using siRNA or mRNA payloads have shown large reductions in potency (~3–10-fold) during LNP drug product testing in monkeys when compared to rodents14,15,29,39. Our data indicate that we can reach maximal pharmacological effect at approximately 1 mg kg−1, a dose that was well tolerated in monkeys. Thus, achieving sufficient potency is critically important as LNP-based therapeutics have a narrow therapeutic index, which can hinder clinical development when there is little separation between pharmacologically active doses and those at which adverse events are observed.
Durable changes in gene expression without modifications of the underlying DNA sequence1,3 is a potential safety advantage of epigenetic editing when compared to other genome editing technologies. Introduction of double-strand DNA breaks using traditional gene editing approaches, such as CRISPR–Cas9, can result in chromosomal aberrations, including large deletions, rearrangements or chromothripsis6,7,8. Furthermore, recent editing technologies using a DNA-binding domain that relies on a single-stand break (or nick) to ultimately modulate gene expression, such as base editing and prime editing, are not completely exempt from potential genotoxic effects, which can be further compounded when two or more genes are simultaneously targeted9. In contrast, our PCSK9-targeting EE uses a catalytically inactive form of Cas9 protein incapable of creating DNA breaks40,41,42, thus avoiding any therapeutic-related risk of chromosomal abnormalities. Although the specificity assessment of our EE showed that a limited number of genomic regions outside of the PCSK9 locus were differentially methylated, these changes were not associated with significant alterations in gene expression. In addition, on-target methylation at the PCSK9 locus by our EE was highly reproducible, constrained within an approximately 1.5-kb window at the promoter region and faithfully maintained during cell division, as demonstrated in mice undergoing PHx where mature hepatocytes re-enter the cell cycle to regenerate the liver to its normal size.
Another advantage of preserving the DNA sequence after targeted epigenetic editing is the potential for reversibility using that same targeting agent fused to a demethylation domain comprising a Tet enzyme, a methylcytosine dioxygenase that facilitates demethylation of methylated CpGs43. As an initial proof of concept, we were able to rapidly re-activate the PCSK9 locus and restore normal plasma PCSK9 levels in mice that had previously been treated with our PCSK9-targeting EE. This was achieved using a construct designed to remove methyl marks at CpG dinucleotides paired with the same PCSK9-targeting gRNAs that were used to methylate the PCSK9 locus in the first place. Although less relevant for clinically validated targets such as PCSK9 (refs. 31,32,33,44,45), such an approach could have utility when the safety profile associated with durable silencing of a specific target is not fully elucidated or when target inhibition is no longer therapeutically required.
In summary, we described the development of an EE targeting PCSK9 for the treatment of hypercholesterolemia that is durable, specific and reversible and, notably, has sufficient potency to enable continued development toward clinical evaluation. Further assessment of the efficacy, tolerability and safety of our PCSK9-targeting EE will be needed before initiation of clinical testing, including additional in vivo studies to evaluate the pharmacodynamic and pharmacokinetic properties of our drug product as well as its biodistribution and potential toxicological effects. Ultimately, our PCSK9-targeting EE holds the potential for a one-and-done treatment that would disrupt the current treatment paradigm for reducing LDL-C, ensuring lifelong cardiovascular risk reduction in patient populations where treatment adherence with chronic therapy is extremely low. Epigenetic editing thus represents a promising therapeutic approach for gene regulation in vivo.
Methods
Cell culture and transfection
HeLa (American Type Culture Collection, CCL-2) cells were maintained in DMEM (Life Technologies) and supplemented with 10% FBS, 1× penicillin–streptomycin (Gibco) and 1× GlutaMAX (Gibco). Cells were kept at 37 °C in a 5% CO2 incubator. For DNA delivery, cells were seeded at a density of 12,000 cells per well in 96-well plates. After 24 h, cells were transfected in duplicate using Mirus TransIT-LT1 reagent with 50 ng of an EE expression plasmid, 25 ng of a gRNA expression DNA fragment and 10 ng of a puromycin resistance plasmid. After 24 h, 1 µg ml−1 puromycin was added to all transfected wells. For RNA delivery, Mirus TransIT-mRNA reagent with 25 ng of an EE mRNA and 12.5 ng of a gRNA was added to wells, and 24,000 cells per well were added afterwards. At noted timepoints, cell viability was determined using CellTiter-Glo (Promega) and used to normalize PCSK9 protein measured in the cell supernatant via a LEGEND MAX Human PCSK9 ELISA kit (BioLegend). GraphPad Prism (version 10.2.2) software was used to visualize results.
RNA production
First, 100-mer gRNAs were chemically synthesized by commercial suppliers Integrated DNA Technologies and BioSpring. For in vitro screening experiments, gRNAs were modified with 2′-O-methylation and phosphorothioate linkages at the three terminal nucleotides of both the 5′ and 3′ ends (protospacer sequences of all gRNA used in these studies are shown in Supplementary Table 1). Subsequent experiments made use of gRNAs containing these end-modifications as well as extensive internal inclusion of 2′-O-methylation.
mRNA was produced via in vitro transcription and purification. Plasmids containing an EE or a control construct were linearized by restriction digest and used as template with T7RNA polymerase, NTPs (including N1-methylpseudouridine) and cap analogue. mRNAs either underwent enzymatic A-tailing or contained an encoded polyA in the template (amino acid sequences of all constructs used in these studies are shown in Supplementary Figs. 1–4).
LNP formulations
Small-scale LNP formulations of EE mRNA and gRNA for initial PHH experiments were done using the pre-made lipid mix GenVoy-ILM kit and prepared on the NanoAssemblr Spark (Cytiva). Mouse- and NHP-scale LNP formulations were prepared as previously described29. Formulations were optimized with an aim to attain LNPs of uniform size and high encapsulation efficiency. The ratio of lipids to the total RNA cargo, comprising mRNA and gRNA, was calculated based on the molar weights of lipids and RNAs. These optimized formulations were evaluated in in vivo studies. For the in vivo scale LNP formulations (mouse and NHP), the RNAs were mixed and encapsulated in LNPs using a self-assembly process. In brief, an aqueous buffered solution of mRNA and gRNA at pH 4.0 was rapidly mixed with a solution of lipids dissolved in ethanol29. LNPs used in this study contained an ionizable cationic lipid (proprietary to Acuitas Therapeutics)/phosphatidylcholine/cholesterol/PEG-lipid. The proprietary lipid and LNP composition are described in patent application WO2020146805A1. The LNPs from the upstream particle formation process were buffer exchanged to remove the ethanol and concentrate the LNPs to desired concentration. The resulting LNPs had a diameter of approximately 80 nm as measured by dynamic light scattering using a Zetasizer Nano ZS (Malvern Instruments) instrument and encapsulation efficiencies of more than 90% as measured by Quant-iT RiboGreen Assay (Life Technologies).
Primary hepatocyte studies
PHHs isolated from mice with a chimeric humanized liver (PXB cells) were obtained from Phoenix Bio. Pre-plated cells were kept in hepatocyte growth medium as previously described46. Cryopreserved PHHs and PCHs were obtained from Lonza and Bio-IVT, respectively. In brief, cells were thawed, combined with Human Cryopreserved Hepatocyte Thawing Medium (Lonza) and centrifuged at 50g for 8 min. Cells were resuspended in Hepatocyte Plating Medium (Lonza) and plated at a density of 80,000 cells per well (Collagen I–coated 96-well plate; STEMCELL Technologies). LNP formulations were mixed with recombinant human ApoE3 (R&D Systems) and added onto cells at the desired concentration. The next day, Matrigel Basement Membrane Matrix (Corning) was overlayed on hepatocytes to a final concentration of 0.25 mg ml−1. PCSK9 protein was measured in supernatant as described above.
Animal studies
LNP treatment in human PCSK9 transgenic mice
All mouse studies were conducted at Charles River Accelerator and Development Lab (CRADL), CBSET research institute or Chroma Medicine animal care facility according to Institutional Animal Care and Use Committee (IACUC) guidelines. Human PCSK9 transgenic (PCSK9-Tg) mice were generated as previously described25,26. For silencing experiments, PCSK9-Tg male and female mice aged between 8 weeks and 16 weeks were randomized to different experimental groups based on baseline plasma PCSK9 protein levels and treated with either vehicle control or various LNP drug products containing genetic or epigenetic editors via intravenous injection. In the PHx experiments, PCSK9-Tg mice were first epigenetically silenced for 8 weeks using PCSK9-EE, after which mice were assigned to undergo either a two-thirds PHx or sham surgery as control. After the surgery, mice underwent 2 weeks of recovery when body weight and health conditions were closely monitored. In the experiment where PCSK9 was reactivated, PCSK9-Tg mice were epigenetically silenced and were then administered an LNP containing dCas-Tet. During the studies, the plasma of animals was collected periodically for analysis of circulating PCSK9 protein using the Simple Plex Human PCSK9 assay kit on Ella instrument (R&D Systems). The data are presented as percentage of baseline with the average of two baseline values before LNP administration. At euthanasia or at surgery, the liver biopsies were collected and snap frozen for DNA or RNA analyses.
LNP treatment in cynomolgus monkeys
The cynomolgus monkey studies were conducted at Altasciences according to IACUC guidelines. Two- to four-year-old male monkeys were genotyped at the PCSK9 locus to ensure that the DNA sequence at the sites where gRNAs bind have no mutations. Monkeys were randomized to various experimental groups based on body weight and established social unit. Pretreatment medications of diphenhydramine (5 mg kg−1), dexamethasone (1 mg kg−1) and famotidine (0.5 mg kg−1) were administrated to all monkeys via intramuscular injection 1 day and 30–60 min before the start of infusion. The LNP formulations or saline vehicle were administered by intravenous infusion for 1 h. Animals were fasted for at least 12 h before blood collection. Blood was collected from peripheral vein, and serum and plasma samples were obtained for clinical chemistry and cynomolgus PCSK9 biomarker analyses, respectively. Serum LDL-C levels were measured using a clinical analyzer, and plasma PCSK9 was measured using the Simple Plex Human PCSK9 assay kit on Ella instrument (R&D Systems), which is 100% cross-reactive against cynomolgus macaque PCSK9 protein. For each analyte, the data are presented as percentage of baseline with the average of at least two baseline values before LNP administration. During the study, liver biopsy collections were performed under general anesthesia by ultrasonographic-guided percutaneous biopsy using a 16-gauge biopsy needle. Samples were flash frozen in liquid nitrogen, maintained on dry ice and then stored in a freezer set to maintain at −80 °C.
RNA-seq
Total RNA was extracted from primary liver cells using simultaneous RNA and DNA extraction from cultured cells without splitting lysate protocol and reagents (Beckman Coulter). Total RNA-seq libraries were constructed using an Illumina stranded Total RNA Prep Ligation with Ribo-Zero Plus kit. Libraries were sequenced on a NextSeq 2000 using NextSeq 1000/2000 Control Software version 1.5.0.42699 to a read depth of approximately 50 million read pairs per sample. Sequencing reads were aligned to the human reference genome (GRCh38), and gene and transcript counts and normalized counts (TPM) were calculated using the nf-core RNA-seq pipeline (version 3.6). Differential gene expression was detected using DESeq2 (version 1.38.0), with a threshold of two-fold change (log2FC = 1) and adjusted P < 1 × 10−5. TPM and volcano plots were visualized using the Python (version 3.10.1), matplotlib (version 3.7.2) and seaborn (version 0.13.0) libraries.
Twist Human Methylome Hybrid Capture Sequencing
Genomic DNA (gDNA) was extracted from primary liver cells using simultaneous RNA and DNA extraction from cultured cells without splitting lysate protocol and reagents (Beckman Coulter). gDNA was sheared to a target size of 350 bp using a Covaris 96 AFA-Tube TPX plate. Sequencing libraries were then constructed from the sheared gDNA using an NEBNext Enzymatic Methyl-seq (EM-seq) kit plus NEBNext Multiplex Oligos for Enzymatic Methyl-seq. After methylation conversion, the Twist Human Methylome Hybrid Capture panel was used to enrich for regions of interest, following the recommended Twist protocol, and libraries were then amplified and sequenced using an Illumina NovaSeq 6000 instrument to a read depth of approximately 70 million read pairs per sample, using NovaSeq 6000 Control Software 1.7.0. We used the nf-core Methyl-seq pipeline (version 2.3.0dev) (https://zenodo.org/badge/latestdoi/124913037) to map sequencing reads to the reference genome (GRCh38) using Bismark version 0.24.0 (ref. 47), with the following parameters: igenomes_ignore = true, clip_r1 = 10, clip_r2 = 10, three_prime_clip_r1 = 10, three_prime_clip_r2 = 10 and cytosine_report = true. MultiQC (version 1.13) (https://doi.org/10.1093/bioinformatics/btw354) reports were then generated from the stats files resulting from the Methyl-seq pipeline. Next, we used the R (version 4.3.1) packages Methrix (version 1.16.0) (https://doi.org/10.18129/B9.bioc.methrix) and DSS (version 2.48.0) (https://doi.org/10.18129/B9.bioc.DSS) to filter the results from the Methyl-seq pipeline and to call DMRs. We loaded the Bismark coverage files into Methrix objects using methrix::read_bedgraphs with the following parameters: pipeline = ‘bismark_cov’, stranded = FALSE, zero_based = FALSE and collapse_strands = FALSE and filtered to keep only CpGs that meet a minimum 5× coverage threshold, across all libraries in the dataset. CpGs that are in the top 99.9% of coverage were also filtered out of the dataset, as these may represent mapping artifacts or repetitive regions. Next, we used DSS::DMLTest to call significantly differentially methylated CpGs between the experimental and reference conditions, with the following parameter: smoothing = true. Then, we ran DSS::callDMR to aggregate nearby differentially methylated CpGs into DMRs, with the following parameters: p_threshold = 1 × 10−10 and minCG = 10. DMRs were further filtered by the average methylation difference between control and experimental conditions, with >20% average difference DMRs retained.
Targeted Hybrid Capture Methyl-seq (custom panel)
gDNA was extracted from mouse liver tissue samples or HeLa cells using simultaneous RNA and DNA extraction from cultured cells without splitting lysate protocol and reagents or DNAdvance kit (Beckman Coulter). EM-seq libraries were prepared as above followed by hybridization capture target enrichment using a custom-designed panel and manufacturer recommendations (Twist Biosciences). Libraries were sequenced using an Illumina MiSeq or NextSeq 2000 (using NextSeq 1000/2000 Control Software version 1.5.0.42699) and 150-bp paired-end reads. We used the Nextflow Methyl-seq (version 2.3.0dev) pipeline (https://zenodo.org/badge/latestdoi/124913037) to map sequencing reads to the human reference genome (GRCh38) using Bismark version 0.24.0 (ref. 47), with the following parameters: igenomes_ignore = true, clip_r1 = 10, clip_r2 = 10, three_prime_clip_r1 = 10, three_prime_clip_r2 = 10 and cytosine_report = true. MultiQC (version 1.16.0) (https://doi.org/10.1093/bioinformatics/btw354) reports were then generated from the stats files resulting from the Methyl-seq pipeline. Integrative Genomics Viewer (IGV) (version 2.17.0) was used to visualize methylation in the region of the PCSK9 TSS.
Targeted amplicon bisulfite sequencing
gDNA was extracted from NHP liver biopsies using a DNAdvance Kit (Beckman Coulter). gDNA was denatured and treated with sodium bisulfite and heated to deaminate unmethylated cytosines to uracil using the EZ-96 Methylation-Gold MagPrep bisulfite conversion kit (Zymo Research). Primers were designed to amplify the region around the NHP PCSK9 TSS, as seven short, partially overlapping, amplicons. Amplicons were generated using Platinum Taq polymerase (Invitrogen) and pooled for each condition so that each sample contained all associated amplicons and was used as input for NEBNext Ultra II DNA library prep, amplified with unique dual indexes to allow for multiplexing. Libraries were sequenced using an Illumina NextSeq 2000 instrument (2 × 150-bp paired-end reads) to a read depth of at least 9 million read pairs per sample using NextSeq 1000/2000 Control Software version 1.5.0.42699. An nf-core Methyl-seq pipeline (version 2.3.0dev) was used for initial read alignment and estimation of methylation calls using Bismark, aligned to the Ensembl Cyno_6.0 reference genome without deduplication. IGV (version 2.17.0) was used to visualize the methylation bedgraphs in the region surrounding the PCSK9 TSS. Average percent methylation of CpGs with coverage in all conditions in a 500-bp region centered on the gRNA binding sites (Chr1:169045699–169046199) was calculated for each condition using R (version 4.3.1).
WGMS
gDNA was extracted from primary liver cells using simultaneous RNA and DNA extraction from cultured cells without splitting lysate protocol and reagents (Beckman Coulter). gDNA quality and quantification assessment was done using Qubit fluorometric quantification and a fragment analyzer to measure DNA fragment lengths. Sequencing library was prepared using NEBNext Enzymatic Methyl-seq kit. Input gDNA (200 ng) was sheared using Bioruptor Pico to an average fragment size of approximately 300 bp. Sequencing was performed on an Illumina NovaSeq 6000 with NovaSeq 6000 Control Software 1.7.0 using 150 bp paired-end reads, resulting in at least 600 million read pairs per library. A nf-core Methyl-seq pipeline (version 2.3.0dev) was used for initial read alignment and estimation of methylation calls using Bismark (version 0.24.0). Initial quality control and low coverage filtering was performed using Methrix (version 1.16.0) (https://doi.org/10.18129/B9.bioc.methrix). Here, we assessed genome-wide coverage for each sequencing sample and replicate concordance. At least two samples were included in each condition, which were then used for downstream differential CpG and methylated region analysis. DSS (version 2.48.0) (https://doi.org/10.18129/B9.bioc.DSS) was run to aggregate nearby differentially methylated CpGs into DMRs, with the following parameters: p_threshold = 1 × 10−10 and minCG = 3. IGV (version 2.17.0) was used to visualize methylation bedgraphs in the region surrounding selected DMRs. The genome-wide Manhattan plot visualizing CpG methylation was generated using R (version 4.4.1), libraries CMplot (version 4.4.1) and data.table (version 1.14.8).
PCSK9 TSS CpG conservation analysis
Human and cynomolgus PCSK9 TSS regions (±1 kb from Ensembl reference genome canonical TSS annotations) were aligned by performing a pairwise sequence alignment using Geneious (version 2024.0.5) (https://www.geneious.com) with the human TSS region as the reference sequence. Matching CpGs in the pairwise alignment were manually annotated.
Data analysis
RNA-seq and methylation sequencing data were analyzed as described above. All other data were graphed and analyzed using GraphPad Prism (version 10.2.2).
Public epigenomic data
WGMS48 and DNaseI data49 were used in analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq and methylation sequencing data have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (GEO) and are accessible through GEO Series accession number GSE282522. Publicly available datasets GSM5652237, GSM5652231 and ENCSR802ZYE and Ensembl reference genomes GRCh38.p13 and Macaca_fascicularis_6.0 were used in this study.
References
Amabile, A. et al. Inheritable silencing of endogenous genes by hit-and-run targeted epigenetic editing. Cell 167, 219–232 (2016).
Cappelluti, M. A. et al. Durable and efficient gene silencing in vivo by hit-and-run epigenome editing. Nature 627, 416–423 (2024).
Nuñez, J. K. et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184, 2503–2519 (2021).
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Leibowitz, M. L. et al. Chromothripsis as an on-target consequence of CRISPR–Cas9 genome editing. Nat. Genet. 53, 895–905 (2021).
Nahmad, A. D. et al. Frequent aneuploidy in primary human T cells after CRISPR–Cas9 cleavage. Nat. Biotechnol. 40, 1807–1813 (2022).
Turchiano, G. et al. Quantitative evaluation of chromosomal rearrangements in gene-edited human stem cells by CAST-Seq. Cell Stem Cell 28, 1136–1147 (2021).
Fiumara, M. et al. Genotoxic effects of base and prime editing in human hematopoietic stem cells. Nat. Biotechnol. 42, 877–891 (2024).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Chadwick, A. C., Wang, X. & Musunuru, K. In vivo base editing of PCSK9 (proprotein convertase subtilisin/kexin type 9) as a therapeutic alternative to genome editing. Arter. Thromb. Vasc. Biol. 37, 1741–1747 (2017).
Musunuru, K. et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates. Nature 593, 429–434 (2021).
Rothgangl, T. et al. In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels. Nat. Biotechnol. 39, 949–957 (2021).
Carreras, A. et al. In vivo genome and base editing of a human PCSK9 knock-in hypercholesterolemic mouse model. BMC Biol. 17, 4 (2019).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Thakore, P. I. et al. RNA-guided transcriptional silencing in vivo with S. aureus CRISPR–Cas9 repressors. Nat. Commun. 9, 1674 (2018).
Lee, R. G. et al. Efficacy and safety of an investigational single-course CRISPR base-editing therapy targeting PCSK9 in nonhuman primate and mouse models. Circulation 147, 242–253 (2023).
Horie, T. & Ono, K. VERVE-101: a promising CRISPR-based gene editing therapy that reduces LDL-C and PCSK9 levels in HeFH patients. Eur. Heart J. Cardiovasc. Pharmacother. 10, 89–90 (2023).
Longhurst, H. J. et al. CRISPR–Cas9 in vivo gene editing of KLKB1 for hereditary angioedema. N. Engl. J. Med. 390, 432–441 (2024).
Gillmore, J. D. et al. CRISPR–Cas9 in vivo gene editing for transthyretin amyloidosis. N. Engl. J. Med. 385, 493–502 (2021).
Dubuc, G. et al. Statins upregulate PCSK9, the gene encoding the proprotein convertase neural apoptosis-regulated convertase-1 implicated in familial hypercholesterolemia. Arter. Thromb. Vasc. Biol. 24, 1454–1459 (2004).
Akinc, A. et al. Targeted delivery of RNAi therapeutics with endogenous and exogenous ligand-based mechanisms. Mol. Ther. 18, 1357–1364 (2010).
Essalmani, R. et al. A single domain antibody against the Cys- and His-rich domain of PCSK9 and evolocumab exhibit different inhibition mechanisms in humanized PCSK9 mice. Biol. Chem. 399, 1363–1374 (2018).
Weider, E. et al. Proprotein convertase subtilisin/kexin type 9 (PCSK9) single domain antibodies are potent inhibitors of low density lipoprotein receptor degradation. J. Biol. Chem. 291, 16659–16671 (2016).
Mitchell, C. & Willenbring, H. A reproducible and well-tolerated method for 2/3 partial hepatectomy in mice. Nat. Protoc. 3, 1167–1170 (2008).
Hatit, M. Z. C. et al. Species-dependent in vivo mRNA delivery and cellular responses to nanoparticles. Nat. Nanotechnol. 17, 310–318 (2022).
Maier, M. A. et al. Biodegradable lipids enabling rapidly eliminated lipid nanoparticles for systemic delivery of RNAi therapeutics. Mol. Ther. 21, 1570–1578 (2013).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Schwartz, G. G. et al. Alirocumab and cardiovascular outcomes after acute coronary syndrome. N. Engl. J. Med. 379, 2097–2107 (2018).
Sabatine, M. S. et al. Evolocumab and clinical outcomes in patients with cardiovascular disease. N. Engl. J. Med. 376, 1713–1722 (2017).
Ray, K. K. et al. Two phase 3 trials of inclisiran in patients with elevated LDL cholesterol. N. Engl. J. Med. 382, 1507–1519 (2020).
Grundy, S. M. et al. 2018 AHA/ACC/AACVPR/AAPA/ABC/ACPM/ADA/AGS/APhA/ASPC/NLA/PCNA guideline on the management of blood cholesterol. J. Am. Coll. Cardiol. 73, e285–e350 (2019).
Lloyd-Jones, D. M. et al. 2022 ACC expert consensus decision pathway on the role of nonstatin therapies for LDL-cholesterol lowering in the management of atherosclerotic cardiovascular disease risk: a report of the American College of Cardiology Solution Set Oversight Committee. J. Am. Coll. Cardiol. 80, 1366–1418 (2022).
Pouwer, M. G. et al. Alirocumab, evinacumab, and atorvastatin triple therapy regresses plaque lesions and improves lesion composition in mice. J. Lipid Res. 61, 365–375 (2020).
Lehoux, D., Kallend, D., Wijngaard, P. L. J., Brown, A. P. & Zerler, B. The N‐Acetylgalactosamine‐conjugated small interfering RNA inclisiran can be coadministered safely with atorvastatin in cynomolgus monkeys resulting in additive low‐density lipoprotein cholesterol reductions. Pharmacol. Res. Perspect. 11, e01080 (2023).
Robinson, J. G. et al. Efficacy and safety of alirocumab in reducing lipids and cardiovascular events. N. Engl. J. Med. 372, 1489–1499 (2015).
Lam, K. et al. Optimizing lipid nanoparticles for delivery in primates. Adv. Mater. 35, e2211420 (2023).
Jinek, M. et al. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Wu, X. & Zhang, Y. TET-mediated active DNA demethylation: mechanism, function and beyond. Nat. Rev. Genet. 18, 517–534 (2017).
Cohen, J. C., Boerwinkle, E., Mosley, T. H. & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Zhao, Z. et al. Molecular characterization of loss-of-function mutations in PCSK9 and identification of a compound heterozygote. Am. J. Hum. Genet. 79, 514–523 (2006).
Yamasaki, C. et al. Growth and differentiation of colony-forming human hepatocytes in vitro. J. Hepatol. 44, 749–757 (2006).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Loyfer, N. et al. A DNA methylation atlas of normal human cell types. Nature 613, 355–364 (2023).
Vierstra, J. et al. Global reference mapping of human transcription factor footprints. Nature 583, 729–736 (2020).
Acknowledgements
We thank J. Chen and W. Yang for their contributions to bioinformatic analysis and pipeline/software development; N. Wong, P. Koppana, J. Labonne and Z. Hoque for their contributions to next-generation sequencing data acquisition; A. Zhai and S. Abraham for their contributions to analysis and code for plots generation; D. Litvak for his supervision of external gRNA and LNP formulation productions; C. Fernandez, S. Alexander and P. Spinelli for program management support; and M. A. Cappelluti for providing reagents and technical expertise. We are grateful to the entire team at Chroma Medicine for their help and support throughout execution of these studies. We also thank the team at Acuitas Therapeutics who supported the work by manufacturing LNP formulations for mouse and NHP studies. We are grateful to N. Goodman, J. Marlowe and C. Stehman-Breen for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
A.E.F., M.S.M. and T.E. conceived and directed the epigenetic editor design and testing. M.L.M., S.J.L. and S.L. conceived and directed the dCas-Tet design and testing. E.M.H., R.N.R. and S.A. conceived the specificity workflow and directed and analyzed cell-based studies for off-target assessment. C.-W.K., Q.X. and S.B.V. conceived, designed and directed in vivo studies in mice and cynomolgus monkeys. C.-W.K., C.G., C.W. and G.K.E. contributed to in vivo studies and data analysis. K.K., L.L., S.H.W., S.C. and S.S.S. contributed to wet lab experiments and data analysis. P.M. and S.P. conceived and directed mRNA production and purification workflow. S.M. and Y.W. contributed to gRNA, mRNA and LNP analytical testing. F.T. wrote the manuscript and supervised the work, with oversight from A.B.J. and advisory input from A.L. and V.E.M. All authors reviewed and edited the manuscript and approved the final version for submission.
Corresponding author
Ethics declarations
Competing interests
F.T., Q.X., S.S.S., C.-W.K., K.K., M.S.S., C.G., R.N.R., E.M.H., S.B.V., G.K.E., S.H.W., L.L., S.C., C.W., S.J.L., S.L., T.E., S.P., Y.W., S.M., P.M., S.A., A.E.F., M.L.M., V.E.M. and A.B.J. were employees of Chroma Medicine and held an equity interest in Chroma Medicine at the time the work was conducted. A.L. is a founder of and holds an equity interest in Chroma Medicine. Chroma Medicine has filed for patent protection related to various aspects of epigenetic editing of PCSK9, with S.A., A.E.F., M.L.M., V.E.M., F.T. and M.S.S. as the inventors.
Peer review
Peer review information
Nature Medicine thanks Albert Jeltsch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Michael Basson, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effect of statin treatment on in vitro activity of top-ranked human PCSK9 epigenetic editors in immortalized cells.
Top 5 gRNA selected based on their activity and durability in HeLa cells (from Fig. 1c), as well as having full cross-reactivity with the cynomolgus macaque PCSK9 gene were evaluated for their ability to maintain PCSK9 protein reduction in the presence of 1 mM simvastatin or DMSO (Vehicle). Simvastatin or DMSO was added in the last 24 hrs of incubation. Individual data points and means are shown, n = 2 replicates per experimental condition. Results are expressed as % of secreted PCSK9 protein in cells treated with transfection (txn) reagent only. WT Cas9 serves as a control for durable silencing of PCSK9.
Extended Data Fig. 2 In vivo activity of human PCSK9 epigenetic editors in mice.
a. CpG methylation levels in livers from untreated transgenic mice carrying the human PCSK9 genomic locus (PCSK9-Tg, in blue) and primary human hepatocytes (in red). b. Location of top 5 gRNAs is shown relative to the distance (in nucleotides) to the PCSK9 gene transcription start site (TSS). c. Effect of a single administration at near-saturating dose (0.75 mg/kg) of an LNP formulation evaluating an epigenetic editor with individual or combination of two gRNA candidates on circulating PCSK9 protein levels in PCSK9-Tg. Results are shown as mean ± s.d. (n = 6 mice/group). d. Effect of a single administration at a sub-saturating dose (0.2 mg/kg) of an LNP formulation evaluating epigenetic editor with best individual or combination of two gRNA candidates on circulating PCSK9 protein levels in PCSK9-Tg mice. Results are shown as mean ± s.d. (n = 4 mice in gRNA41 group and n = 5 mice in gRNA41 + 49 group).
Extended Data Fig. 3 Specificity follow-up of top human PCSK9 epigenetic editor in primary human hepatocytes.
a. WGMS CpG methylation for Effector Only (grey) and PCSK9-EE (blue) in the genomic region surrounding off-target DEG ENSG00000285976 (boxed gene) (left) and low level of baseline expression (TPM) of off-target DEG ENSG00000285976 from RNA-Seq is shown in the dot plot (n = 3) (right). b. Scatterplot of RNA-Seq Log2 fold-change of all genes within 20 kb of any DMR called in Twist Human Methylome Hybrid Capture Methylation Sequencing assay (PCSK9-EE vs Effector Only) vs methylation difference of the associated DMR for each gene (PCSK9-EE vs Effector Only). PCSK9 gene/DMR is shown in yellow. Thresholds (grey dashed lines) set as methylation (beta-value) difference > 0.2 or < -0.2, RNA-Seq Log2FC > 1 or < -1. c-i. For all DMRs meeting either the methylation or gene expression change thresholds in Fig. 2f and Extended Data Fig. 3b, WGMS CpG methylation for Effector Only (grey) and PCSK9-EE (blue) in the genomic region surrounding each off-target DMR (left) and expression (TPM) of all off-target genes with transcription start sites within 20 kb of each DMR from RNA-Seq is shown in the dot plot (n = 3) (right). RNA-seq padj values are from a DEseq2 Wald test, two-sided, with Benjamani-Hotchburg multiple comparison correction. c. DMR near WDR81/MIR22HG/SERPINF2, average methylation difference within DMR by Twist Human Methylome assay: 0.24. Average methylation difference within DMR by WGMS: 0.18. WDR81 RNA-Seq padj = 9.998728e-01, MIR22HG RNA-Seq padj = 9.998728e-01, SERPINF2 RNA-Seq padj = 5.354105e-02, d. DMR near RB1-DT/PPP1R26P1, average methylation difference within DMR by WGMS: 0.14. RB1-DT RNA-Seq padj = 9.998728e-01, PPP1R26P1 RNA-Seq padj = 9.998728e-01. e. DMR near MCF2, average methylation difference within DMR by Twist Human Methylome assay: 0.07, MCF2 RNA-Seq padj = 9.998728e-01, f. DMR near TCEA1P3, average methylation difference within DMR by Twist Human Methylome assay: -0.16, TCEA1P3 RNA-Seq padj = 9.998728e-01, g. DMR near SLC47A2, average methylation difference within DMR by WGMS: 0.13, SLC47A2 RNA-Seq padj = 9.998728e-01. h. DMR near L3MBTL1, average methylation difference within DMR by Twist Human Methylome assay: −0.22, L3MBTL1 RNA-Seq padj = 9.998728e-01, i. DMR near Y_RNA, average methylation difference within DMR by WGMS: 0.11, Y_RNA RNA-Seq padj = 9.998728e-01.
Extended Data Fig. 4 Liver safety monitoring of a human PCSK9 epigenetic editor in cynomolgus monkeys.
Serial measurements of blood alanine transaminase (a), aspartate transaminase (b), and total bilirubin (c) were performed in non-human primates following dosing with either a vehicle control or PCSK9-EE-V2. Results are shown as mean ± s.d. (n = 4 animals in vehicle group and n = 3 animals in each experimental group receiving PCSK9-EE-V2). Plasma samples were obtained from two of the vehicle-treated animals at day 84 and 98. These data have been averaged and plotted at day 91 to better visualize the group mean.
Extended Data Fig. 5 Activity of a human PCSK9 epigenetic editor in individual cynomolgus monkeys in vivo.
Time-course and dose-response effect of a single infusion of an LNP formulation with PCSK9-EE-V2 on circulating PCSK9 protein levels (b-d) and LDL-cholesterol (g-i) in individual cynomolgus macaque. Vehicle-treated animals receive a single infusion of saline solution (a and f). Average cPCSK9 (e) and LDL-cholesterol (j) change at day 91 post dose from baseline. For each experimental group, individual animals are represented by a different symbol (a-j). For e and j, results are shown as mean ± s.d. (n = 4 animals in vehicle group and n = 3 animals in each experimental group receiving PCSK9-EE-V2).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–4 and Supplementary Tables 1 and 2.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tremblay, F., Xiong, Q., Shah, S.S. et al. A potent epigenetic editor targeting human PCSK9 for durable reduction of low-density lipoprotein cholesterol levels. Nat Med 31, 1329–1338 (2025). https://doi.org/10.1038/s41591-025-03508-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41591-025-03508-x
This article is cited by
-
The promise of epigenetic editing strategies in functionally curing chronic hepatitis B virus infections
Epigenetics Communications (2026)
-
Epigenetic Editing in Neurological and Neuropsychiatric Disorders: Pioneering Next-Gen Therapeutics for Precision Gene Control
Molecular Neurobiology (2026)
-
Proprotein Convertase Subtilisin/Kexin Type 9 (PCSK9) in Alzheimer’s Disease: Recent Advances and Controversies
Molecular Neurobiology (2026)
-
Clinical epigenetics: recent advances and opportunities
Epigenetics Communications (2025)
-
Epigenetic editing and epi-drugs: a combination strategy to simultaneously target KDM4 as a novel anticancer approach
Clinical Epigenetics (2025)