Abstract
Linezolid, an oxazolidinone, is a cornerstone of treatment regimens for highly drug-resistant tuberculosis but cannot be used in drug-susceptible disease because of toxicity. This toxicity results from inhibition of mammalian mitochondrial protein synthesis. Here we show the development of a new oxazolidinone, MK-7762, with antitubercular activity that is better than linezolid and limited mitochondrial protein synthesis inhibition. The cryogenic electron microscopy structure of the stalled mycobacterial ribosome with MK-7762 revealed the basis for this selectivity. BALB/c mouse models of disease showed MK-7762 reduced lung bacterial burden by a 3-log-fold decrease in an acute model (N = 18) and a 2-log-fold decrease in chronically infected animals (N = 18). MK-7762 showed lesion penetration similar to linezolid in C3HeB/FeJ mice. MK-7762 had pharmacokinetic properties predicting low once-daily doses in humans and a favorable 14-day preclinical safety profile in Wistar Han rats (N = 30) and Beagle dogs (N = 6). Four-month safety studies in both rats (N = 20) and dogs (N = 24) showed no changes in hematology parameters at exposures well above the 100-mg predicted human dose. These data will enable MK-7762 to be explored as a component of new tuberculosis treatment combinations for all forms of the disease.
Similar content being viewed by others
Main
Tuberculosis (TB) is the leading cause of death due to an infectious disease globally. In 2022, there were 1.3 million deaths and 10.6 million new infections caused by Mycobacterium tuberculosis (Mtb)1. While the existing standard regimen can be highly effective in patients with drug-susceptible bacteria, this regimen requires 6 months to achieve a durable cure in most patients. Since these agents have been in use for decades in hundreds of millions of people, multidrug-resistant (MDR) and extensively drug-resistant (XDR) Mtb has evolved and spread. Over the past 10 years a variety of new agents have been introduced to treat MDR/XDR-TB2,3. These agents include the repurposed oxazolidinone antibiotic linezolid, which was first shown to lead to high rate of culture conversion (87%) and cure (71%) as functional monotherapy in patients with XDR-TB receiving salvage treatment4,5. Subsequently, linezolid in combination with other new agents, bedaquiline and pretomanid, achieved high rates of cure and shortened treatment duration for MDR-TB6. This resulted in the World Health Organization changing global recommendations for the treatment of MDR-TB and pre-XDR-TB in 2022 to a 6-month, all-oral regimen including linezolid7.
The adverse events associated with linezolid treatment, however, restrict its utility to MDR/XDR-TB given the limited treatment options for this indication8. Bone marrow suppression, peripheral neuropathy and optic neuritis are the most commonly observed adverse events and are believed to result from inhibition of human mitochondrial protein synthesis (MPS)9,10. In patients with TB, these adverse events are associated with trough concentrations of drug and low MPS as measured in patients being treated with linezolid11. Additionally, the relatively short human half-life of linezolid (5−7 h) results in a high ratio of maximum (Cmax) and trough (Ctrough) plasma concentrations observed over 24 h resulting in highly variable trough concentrations and drug exposures (area under the curve or AUC) at clinically used doses. Thus, to broaden treatment beyond drug-resistant TB and to explore the possibility of higher efficacy (including further shortening of treatment) with an oxazolidinone, we sought to discover a compound with high MPS selectivity and pharmacokinetic (PK) properties that would allow for once-daily dosing.
Results
Summary of the discovery of MK-7762 (TBD09)
The full medicinal chemistry story enabling the discovery of MK-7762 (TBD09) will be described in detail elsewhere but involved exhaustive evaluation of the molecular scaffold at the five positions (R1, R2, A, B and C) denoted in Fig. 1a. About 2,800 analogs were assessed for antimycobacterial activity (up to 50 µM) and active compounds (minimum inhibitory concentration inhibiting 99% of growth (MIC99) < 6 µM) were further evaluated for in vitro inhibition of MPS (Fig. 1b). Structure–activity relationships for the MPS activity were determined and several scaffolds were defined with good selectivity and potency (green box in Fig. 1b). Examples of potent scaffolds with selectivity for inhibition of Mtb are shown in Fig. 1c. Compounds were selected for advancement to in vivo PK studies based on Mtb-MPS selectivity (approximately >20) and a potency–PK efficiency metric (volume ligand efficiency)12,13 designed to enrich for compounds with favorable projected human doses (green box in Fig. 1d). This work led to the identification of subclasses of molecules that consistently displayed moderate-to-good levels of MPS selectivity. Modifications to the C5 position of the A ring and the C ring were most effective in conferring selectivity and showed interdependency. Further profiling and optimization of these subclasses with respect to PK properties and off-target selectivity resulted in the identification of multiple compounds for evaluation using in vivo efficacy and safety studies. The final agent (MK-7762) occupies similar (polar) physicochemical space to linezolid and was found to have similar biodistribution although with a better balance of unbound volume of distribution and unbound clearance, in addition to improved in vitro and in vivo selectivity. MK-7762 showed minimal inhibition of MPS (half-maximal effective concentration (EC50) of 98 µM) compared to linezolid (EC50 of 16 µM), suggesting a low risk for mitochondrial toxicity in humans compared to linezolid. Additionally, MK-7762, tested at 10 µM, showed no inhibition in a panel of 106 off-targets (Supplementary Table 1) except for monoamine oxidase-B (for which the half-maximal inhibitory concentration (IC50) was 6.9 µM), with no meaningful inhibition (<50%) of monoamine oxidase-A.
a, Summary of the progression from linezolid and the positions varied to identify MK-7762. R2 = C5 position. b, Plot of potency against Mtb versus MPS selectivity. Active compounds with greater than 20-fold selectivity against MPS were prioritized (green box). Arrow denotes MK-7762. Green color indicates that the MPS data were greater than the assay limit of detection (or compound solubility) and red indicates the MPS data were lower than the lowest tested concentration. Assays were performed in triplicate and average values are shown. c, Examples of selected compounds with good potency and selectivity indices (SI; see Supplementary Synthesis and Characterization of Compounds in the Supplementary Information). d, Rat volume ligand efficiency (VLE; dashed line at 5.3 is the VLE of linezolid as a benchmark) versus mean residence time (MRT) in PK studies (the vertical line at 3 roughly predicts a human half-life of 14 h). IV, intravenous.
MK-7762 is broadly active against Mtb isolates and shows a promising resistance profile
MK-7762 was evaluated against a panel of clinical isolates that included both drug-sensitive and drug-resistant strains of Mtb and one strain each of Mycobacterium bovis and Mycobacterium africanum. The minimum inhibitory concentration to cover 90% of these isolates (MIC90) was found to be 0.78 µM (0.33 µg ml−1; Fig. 2a). MK-7762 was additionally evaluated against linezolid-resistant mutants that bear binding site mutations in the bacterial ribosome from patients who developed linezolid resistance in a prior clinical trial (23S region, rrl G2814T and rrl G2685T in strains 053K113 and 28K111, respectively) and showed a similar loss of activity to linezolid for those mutants (MK-7762: 16−24×; linezolid: 18−22×), confirming similar on-target binding. MK-7762 had a more potent minimum bactericidal concentration (MBC) than that of linezolid (Fig. 2b) with an observed MBC99 of 18.1 µM compared to 52.7 µM for linezolid after a one-week exposure (although this may simply reflect the lower MIC90).
a, MIC99 for linezolid and MK-7762 against a globally diverse collection of drug-susceptible (DS) and drug-resistant (MDR and XDR, referring to strains with increasing levels of drug resistance). (Drug susceptibility testing (DST) from clinical testing against the following drugs; H, isoniazid; R, rifampicin; E, ethambutol; S, streptomycin; K, kanamycin; P, p-aminosalicylic acid; Et, ethionamide; M, moxifloxacin; Lv, levofloxacin; Cp, capreomycin; Ak, amikacin; O, ofloxacin; Rb, rifabutin; Z, pyrazinamide; LZD/L, linezolid; Cs, D-cycloserine; Pt, prothionamide; NA, not applicable). Strain lineage refers to the global phylogenetic group to which the strain belongs. b, MBC determination for linezolid and MK-7762 after 7 days of exposure of H37Rv to drug (two biological replicates shown in squares and circles). c, Sequence of Rv3160c in MK-7762-selected resistant mutants along with their cross-resistance to linezolid. d, Metabolite analysis of MK-7762 exposed to wild-type (WT) or mutant (A6) cells showing absorbance profiles at 270 nm and mass spectrometer determination of molecular weights (AMU) above select peaks (the same three mass profiles were seen in all but heat-killed organisms but not all peaks are labeled).
To determine the frequency of resistance (FOR) and confirm the mechanism of action of the compound we also selected MK-7762-resistant mutants. The FOR determinations for MK-7762 were inoculum dependent with high cell densities giving rise to lawns on plates. Observed FORs ranged from 1 in 106 to 1 in 109, and the resulting colonies required very long incubation times and appeared very tiny on plates as though cells were impaired for growth in the presence of MK-7762. When isolated, these colonies showed only relatively low levels of resistance to MK-7762 ranging from ~2–8-fold shifts in their MIC99. Surprisingly, all but one of these mutants showed no cross-resistance to linezolid (Fig. 2c). Whole-genome sequencing revealed that all these mutants shared single-nucleotide polymorphisms in a single target gene, Rv3160c, a TetR-family transcriptional regulator that has previously been shown to confer resistance to triclosan12, thioridazine13 and other preclinical candidates14. Rv3160c is directly upstream of Rv3161c, a dioxygenase presumably regulated by this transcription factor. Because triclosan and MK-7762 share a dihalogenated aromatic ring, it seemed possible that these resistance mutations resulted in an increase in metabolism of the drug mediated by this dioxygenase. To confirm this hypothesis, we incubated MK-7762 with wild-type or mutant cells and monitored drug conversion by liquid chromatography coupled to mass spectrometry (LC–MS; Fig. 2d). Wild-type cells showed little evidence of drug metabolism even after 96 h of incubation, whereas mutants showed the appearance of two new peaks, one corresponding to 435 atomic mass units (AMU; MK-7762 + OH) and a second peak at 452 AMU (MK-7762 + 2OH). These peaks appeared after only 24 h of incubation and the parent drug was completely absent. Thus, these results are consistent with increased bacterial metabolism as a resistance mechanism for this compound but the cross-resistance with linezolid-resistant mutants with alterations in the ribosome binding site confirm that MK-7762 nonetheless remains on target. To support the proposed aromatic oxidation, we performed liquid chromatography coupled to tandem mass spectrometry (LC–MS/MS) and observed that the oxygenated fragments corresponded to the aromatic ring degradation fragments (Extended Data Fig. 1). To demonstrate that this degradation was because of Rv3161c upregulation, we quantified expression of this gene by PCR with reverse transcription (RT–PCR) in both wild-type and mutant cells, with and without drug treatment. These results showed a ~250–300-fold upregulation of Rv3161c relative to sigA expression levels in the mutants (Extended Data Fig. 2). MK-7762 was also evaluated against a panel of Gram-positive and Gram-negative organisms and found to have a spectrum of activity comparable to linezolid (Supplementary Tables 2–4).
Structural basis for inhibition of the mycobacterial ribosome
To understand the structural basis for the observed mycobacterial selectivity, we purified ribosomes from Mycobacterium smegmatis and performed in vitro transcription–translation in the presence of MK-7762 and a short stalling template as previously described15,16. The stalled ribosomes were purified and subjected to cryogenic electron microscopy (cryo-EM; Extended Data Figs. 3 and 4), producing a structure with 2.0-Å resolution (Extended Data Fig. 4 and Extended Data Table 1). Well-defined densities were observed for MK-7762 at the A-site of the peptidyl transferase center and peptidyl-tRNA at the P-site (Fig. 3a). The carbonyl group of the C-5 carbamate makes a hydrogen bond with the N-6 amine of A2286, while the amide NH of that carbamate forms hydrogen bonds with the phosphate backbone of the 23S rRNA. The alanine of the nascent polypeptide is displaced by the C-5 methyl group compared to the contact made with linezolid and does not show the same CH–π interaction17. The sulfone of the thiomorpholine dioxide anchors the C-ring of the molecule with a 2.0-Å hydrogen bond to U2809 (Fig. 3b and Extended Data Fig. 4). Aligning the structures of the human mitoribosome18 (PDB 7A51) and the mycobacterial ribosome reveals that U2809 in the mycobacterial structure corresponds to U3072 in the mitochondrial system where U3072 is twisted by 45° and stabilized via a noncanonical U-U base pair with U-2993 preventing binding of the sulfone of MK-7762 (Fig. 3c). This hydrogen bond to the sulfone suggests that this feature might be responsible for the observed selectivity difference between the mycoribosome and the mitoribosome. To test this hypothesis, we also evaluated the mitotoxicity of the corresponding thioether and sulfoxide and found both to be substantially more active in inhibiting MPS (Extended Data Fig. 5).
a, Cryo-EM map at a 2.5σ level of stalled ribosomes from M. smegmatis complexed with MK-7762 (blue) (EMD-49038). Green indicates tRNA in the P-site, cyan indicates mRNA, and purple indicates nascent peptide. b, Contacts at the MK-7762 binding site of the mycobacterial ribosome. The density of MK-7762 is shown at the 3σ contour. A4 and F5 refer to the penultimate alanine and final phenylalanine of the stalled nascent peptide. MK-7762 carbon is shown in gray, mycoribosome carbon is shown in light blue, mitoribosome carbon is shown in light magenta and nascent polypeptide carbon is shown in pink. c, Superposition of the mycobacterial and mitochondrial ribosome structures near the MK-7762 binding site, highlighting the stabilizing hydrogen bond between mycoribosome U2809 and MK-7762 and the collision between mitoribosome U3072 nucleobase and MK-7762. Van der Waal surfaces are shown for MK-7762 (gray) and U3072 nucleobase (light magenta).
Preclinical PK properties and safety profile of MK-7762
PK properties of MK-7762 after IV and oral administration to mice (Supplementary Table 5a), rats (Supplementary Table 5b) and dogs (Supplementary Table 5c) showed good oral absorption (F = 90–100) with a projection of once-daily human dosing of 100 mg adequate to achieve unbound concentrations of MK-7762 above the MIC for 24 h. In 14-day nonclinical toxicity studies in rats and Beagle dogs, MK-7762 was well tolerated at doses and exposures that support a large therapeutic index (33-fold free drug, 22-fold total drug) for MK-7762 at the projected efficacious human dose (Table 1). MK-7762 was also evaluated for cardiovascular toxicity, bone marrow toxicity, drug-induced liver toxicity and genotoxicity with no evidence of safety findings. Analysis of rat and dog plasma and urine for metabolites of MK-7762 after oral administration indicated only a few minor metabolites in both species. Daily administration for 4 months in rats and dogs at exposures of ~4–11-fold the efficacious predicted human exposure with a 100-mg dose showed only minimal, reversible changes in hematological parameters with no indication of associated bone marrow toxicity, confirmed by microscopic examination (Extended Data Table 2 and Supplementary Tables 6 and 7).
MK-7762 is efficacious in acute and chronic murine models of disease
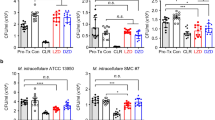
We next evaluated the in vivo efficacy of MK-7762 in mouse models of Mtb infection (Fig. 4). In an acute infection model in BALB/c mice (dosing initiated 7 days after infection with Mtb and continued for 12 days), the compound was dosed at 30 and 100 milligrams per kilogram of body weight (m.p.k.) once daily and 100 m.p.k. twice daily (six mice per dose; Fig. 4a and Supplementary Table 8). Linezolid (50 and 100 m.p.k. once daily) was also dosed in this study (controls). Treatment with MK-7762 resulted in 1.4-, 4.5- and 5.3-log reduction in lung colony-forming units (CFUs; versus untreated controls at the completion of the study) at the 30-m.p.k. once-daily, 100-m.p.k. once-daily and 100-m.p.k. twice-daily dosing groups, respectively. Treatment with linezolid resulted in similar reductions in lung CFUs as prior experiments in the same model (0.8- and 2.4-log reduction at 50 and 100 m.p.k. once daily, respectively).
a, Acute TB efficacy in BALB/c mice where treatment is initiated 7 days after infection and continued daily for 12 days. *P < 0.001, **P < 0.0001, one-way analysis of variance (ANOVA; Supplementary Table 9 contains detailed statistical group comparisons). Each group contained six animals and the individual values are shown along with the mean. b, Chronic TB efficacy study in BALB/c mice where mice were infected for 4 weeks before initiation of treatment. Treatment was daily for 1 month. **P < 0.0001, one-way ANOVA (Supplementary Table 10 contains detailed statistical group comparisons). Displayed as above with individual values for each animal and the mean. c, Lesion penetration was assessed in C3HeB/FeJ mice 10 weeks after infection, and animals were dosed for 2.5 weeks at 100 m.p.k. for both linezolid and MK-7762 (5 of 7 days per week) before euthanasia and analysis. BID, twice daily. NS, not significant. QD, once daily.
MK-7762 was further evaluated in a chronic TB infection model in BALB/c mice (Fig. 4b and Supplementary Table 9). Treatment was initiated 28 days after infection with Mtb and continued for 1 month. Similarly to the acute infection study, we also evaluated linezolid in this experiment. At 30, 100 and 200 m.p.k. of MK-7762, we observed a 0.7-, 1.9- and 2.2-log reduction in lung CFUs. Treatment with linezolid at 50 and 100 m.p.k. resulted in a 1.9- and 2.2-log reduction in lung CFUs. There was no statistically significant difference between the two highest dose levels of MK-7762 and the highest dose of linezolid. In this model the bacteria are only slowly replicating and killing is overall slower than in the acute model.
Lesion penetration of MK-7762 in C3HeB/FeJ mice, which form caseous necrotic lesions (Fig. 4c), was also evaluated19,20. Treatment was initiated 10 weeks after infection and continued for 2.5 weeks to ensure steady-state concentrations of drug in the caseous lesions. After euthanasia, the lungs were removed and the amount of drug present in a cross-section of each quantified by laser capture microdissection (LCM) LC–MS/MS (Fig. 4c). Concentrations were determined for multiple compartments/sub-compartments and included the collagen rim, cellular–caseum interface and caseum. Drug levels were also measured in the remaining lung and compared alongside drug measured in plasma before euthanasia. MK-7762 was evaluated at a dose level of 100 m.p.k. once daily. Linezolid was also evaluated in this study at 100 and 50 m.p.k. once per day. Animal cohorts were euthanized at multiple time points after the last dose (1 h, 5 h and 16 h for MK-7762, 1 h, 5 h and 16 h for linezolid). Drug levels for MK-7762 were similar across all compartments/sub-compartments and were above the unbound MIC concentration at the 16-h time point with small differences between the 1-h and 16-h time points. The biodistribution of drug into the lesions was similar to that observed for linezolid.
Discussion
When linezolid was originally developed, the precise mechanism of its antibacterial action or toxicity was not known. This lack of information made invention of an agent with the desired spectrum (coverage of Streptococcus pneumoniae, Staphylococcus aureus and Enterococcus faecium) and low toxicity particularly challenging15,16. Later work demonstrated its ability to inhibit bacterial protein synthesis via competition with endogenous substrates by binding to the A-site of the 50S ribosome subunit16,17. This mechanism of action (and other data) then suggested that the toxicity associated with agents in this class was due to inhibition of mammalian MPS9,10.
MK-7762 was active against a wide range of diverse drug-sensitive and drug-resistant clinical isolates and showed similar bactericidal activity as linezolid. Unlike linezolid it was not possible to select resistant mutants directly on drug-containing plates as there was a pronounced inoculum effect where very high concentrations of bacteria plated would overwhelm the drug and grow as a fully sensitive lawn. We did observe low-level resistance emerging at moderate frequency (1 in 106 to 109 cells) but the resistance to MK-7762 was of a low level (~2–8-fold shift in MIC). Whole-genome sequencing of these mutants revealed alterations in a transcriptional regulator (Rv3160c) that has been previously identified as a resistance determinant for triclosan, thioridazine and another preclinical series12,13,14. This low-level resistance appears to come with a substantial fitness cost as bacilli grow very slowly on MK-7762 with the observed mutations. Linezolid has a very low frequency of emergence of resistance, a phenomenon that likely allowed the success of the monotherapy trial in patients with XDR-TB4. MK-7762 may share this low level of emergence of new clinical resistance but will not be useful in combatting resistance that has already emerged to linezolid (barring demonstration of high dosing flexibility of MK-7762 in humans). The molecular basis for this selectivity was revealed by the cryo-EM structure of the mycoribosome while stalled during translation and provides insights that may be useful for other recalcitrant mycobacterial pathogens. MK-7762 showed superior activity to linezolid in acutely infected mice and equivalent activity in chronically infected mice. This is likely attributable to the slower replication of Mtb in the chronic model making them less susceptible to inhibition of protein synthesis. Because of this we performed lesion-penetration studies in C3HeB/FeJ mice but did not pursue efficacy in that model where linezolid has shown only mediocre activity21.
The improved in vitro selectivity of MK-7762 for Mtb activity versus MPS inhibition translated to improvement in the in vivo therapeutic index as determined in rat and dog safety studies with multiple week dosing. Four-month extended toxicity studies further support a likely reduction in other adverse events associated with MPS inhibition (for example, various neuropathies), although this awaits further long-term exposure in humans to confirm. As MK-7762 was observed to have a lower MIC in vitro and a longer projected human half-life than linezolid, the separation between efficacy and in vivo toxicity would be expected to be further increased in human and would represent a meaningful differentiation from linezolid with respect to therapeutic index.
Our study has important limitations. MK-7762 and linezolid have a similar spectrum of inhibition of Gram-positive and Gram-negative organisms and long-term use may still result in gastrointestinal disturbance, although the lower projected dose should make this less of a problem. Side effects that are not related to MPS inhibition may still be discovered once MK-7762 is studied in humans. The phase 1 study in humans has recently completed and will be reported in due course (NCT05824091).
Having multiple selective series allowed us to select our PK profile to better suit an agent to treat TB. Linezolid has a relatively short half-life in humans and is typically given twice daily for Gram-positive infections. Twice-daily administration for TB treatment is not considered ideal given the length of treatment and that directly observed treatment is recommended22. We selected MK-7762, in part, because the PK in two species looked very promising for once-daily administration in humans.
A wide variety of PK–pharmacodynamic models have been applied to efficacy trials in patients with TB23,24. Based on the predicted human PK, MK-7762 is projected to have trough concentrations exceeding the MIC for almost all tested clinical isolates at a low dose relative to linezolid. Assuming a similar PK–pharmacodynamic relationship for MK-7762 as for linezolid (same mechanistic and structural class), this suggests that this agent will have meaningful clinical activity at a safe and very reasonable dose. Oxazolidinones have emerged as a centerpiece of many modern regimens for improving the treatment of drug-resistant TB. MK-7762 offers the exciting possibility of further extending the utility of this class of agents into combinations designed to treat drug-sensitive TB. Further clinical data from its active development will be disclosed in due course.
Methods
Our research complies with all relevant ethical regulations and was approved by the Colorado State University Institutional Animal Care and Use Committee (ref. no. 5172; efficacy studies) or the Merck Institutional Animal Care and Use Committee (PK studies).
Antibacterial assays
Mtb-GFP was cultured in 7H9/glucose/BSA/tyloxapol media until the culture reached an OD650 of 0.3 and was then diluted in 7H9/glucose/BSA/tyloxapol media at 1,000-fold for the assay. 7H9/glucose/BSA/tyloxapol media consists of: 4.7 g Difco Middlebrook 7H9 broth, 4 g glucose, 0.81 g sodium chloride, 5 g BSA, fraction V and 0.5 ml tyloxapol, brought up to a 1-l total volume with double-distilled water. For the assays, an initial inoculum of approximately 20,000 Mtb-GFP cells was used per well. Mtb-GFP cells were then exposed to a 2-fold dilution series of MK-7762 in duplicate from 50 to 0.049 μM. MK-7762 was diluted as 2 μl of 10 mM compound stock solution in DMSO added to 200 μl 7H9/glucose/BSA/tyloxapol media to produce a 100 μM concentration (2×, before adding to cells). Using a multichannel pipettor, 50 μl was transferred to each subsequent row starting with row 1 and ending with row 11. The final extra 50 μl was discarded from row 11. MK-7762 was titrated from 50 μM to 0.049 μM. Assay plates were incubated for 7 days at 37 °C, and then evaluated for GFP signal using an Envision Multimode Plate Reader (PerkinElmer). The concentration of MK-7762 that resulted in 95% growth inhibition as measured by the fluorescence signal of the Mtb-GFP assay wells compared to the drug-free control wells was recorded as the MIC95. The concentration of MK-7762 that resulted in 95% growth inhibition as measured by the fluorescence signal of the Mtb-GFP assay wells compared to the no-drug control wells was recorded as the MIC95. There was no curve-fitting for the analysis, and only values corresponding to actual test concentrations were returned as results. The cholesterol-containing media assay was run with the same protocol but substituting cholesterol for glucose.
MBC assay
H37Rv was grown to an OD650 of 0.2 in 7H9/ADC/Tw media (composition above but with 0.5 ml Tween 80 instead of tyloxapol) The cell suspension was filtered through a 5-µm filter to create a single-cell suspension and diluted 10-fold in 7H9/ADC/Tw media. A sample of the cell suspension was appropriately diluted and plated on 7H11-OADC agar to enumerate CFUs. An equal volume of the cell dilution (0.5 ml) was added to an equal volume of 7H9/ADC/Tw media containing different concentrations of oxazolidinones (MK-7762 and linezolid at 0 µM, 6.25 µM, 12.5 µM, 25 µM, 31.3 µM, 50 µM, 62.5 µM and 125 µM) in triplicate in sterile 24-well plates. Rifampicin at 0.4 µg ml−1 was used as a positive control. Plates were incubated at 37 °C, shaking at 100 rpm for 7 days after which appropriate dilutions of cells were plated on 7H11-OADC agar for CFU enumeration. 7H11-OADC agar plates were incubated at 37 °C for 4 weeks before colonies were counted.
FOR determination
H37Rv was grown to an OD650 of 0.2 in 7H9/ADC/Tw media and cells were harvested by centrifugation. Cell suspensions of 1 × 1010, 1 × 109 and 1 × 108 CFUs per milliliter were made in 7H9/ADC/Tw media of which 0.1-ml volumes were plated on 7H11-OADC agar containing different concentrations of linezolid or MK-7762 (2.5 µM, 4 µM, 5 µM, 8 µM, 10 µM and 16 µM). Each cell suspension was plated in duplicate on each drug concentration. Appropriate dilutions of cells were plated on drug-free 7H11-OADC agar plates to enumerate inoculum cell densities for FOR calculations. 7H11-OADC agar plates were incubated at 37 °C for 4 weeks before colonies were counted. Colonies that grew up on drug-containing plates were picked and MIC determination performed as described above to determine the level of resistance.
Analysis of rv3161c mRNA expression in MK-7762-resistant mutants
Wild-type H37Rv and three MK-7762 resistant mutants with Rv3160c mutations (A1, A3 and A6) were grown in 7H9/ADC/Tw media to an OD650 of 0.2 at which stage cells were harvested, resuspended in TRIzol (Thermo Fisher) and RNA isolated and treated with DNase as previously described. This was repeated to generate independent replicates for each strain. RNA was reverse transcribed with the iScript Advanced cDNA synthesis kit for RT–qPCR (Bio-Rad) following the manufacturer’s recommendations with the additional inclusion of primers for specific reverse transcription of sigA and rv3161c (sigA GCTAGCTCGACCTCTTCC; rv3161c GAGCACGTGGTAGTTCTC). Levels of sigA and rv3161c in the cDNA samples were determined by qPCR using the iTaq Universal SYBR Green Supermix (Bio-Rad) with standard curve of Mtb genomic DNA according to the manufacturer’s instruction using the sigA (sigA-F: CGGTGATTTCGTCTGGGATG; sigA-R: TTGCCGATCTGTTTGAGGTAG) and rv3161c (rv3161c-F: TGGAATGGATTGGTGTGGATC; rv3161c-R: TTCCAATTAGCTCGCCACTC) respective primers. The gene for rv3161c expression was normalized to sigA levels for each sample.
Mitochondrial biogenesis assay
The assay was performed using the MitoBiogenesis In-Cell ELISA Kit (Abcam). HepG2 cells (American Type Culture Collection, HB-8065) cultured in DMEM, 4.5 g l−1 D-glucose, 4 mM L-glutamine 10% FBS and 1 mM sodium pyruvate from T175 flasks of approximately 60% confluency were removed from flasks using 0.05% Trypsin-EDTA and combined in 10 ml of the same medium. Cells were sieved with a 40-μm cell strainer to reduce clumps and then counted at least four times using the Cellometer Auto 2000 (Nexcelom, PerkinElmer) with the counts averaged. The HepG2 cells were then seeded in a Collagen I Coated Plate (3,500 cells per well) in 170 μl of media per well and left to adhere for 16 h to 18 h at 37 °C and 5% carbon dioxide.
Serial dilutions of compounds were prepared by pipetting 177 μl of media into column 1 of a non-assay/non-culture plate (Non-Tissue Culture Treated Plate) and 90 μl of media into column 2. Compounds were added as 12.3 μl of 40 mM solutions into column 1 for a final concentration of 0.6667 mM. Linezolid was included as a positive control. Compounds were diluted serially (1:2) by pipetting 90 μl from column 1 to column 2 through to column 10. Column 11 was designated the control without compounds, and column 12 was the control without primary antibody for background signal subtraction.
Once the cells were approximately 15% confluent (day 2), the assay was initiated. Each compound (30 μl) was added from the compound plate into 170 μl of HepG2 cells (attached for 16 h to 18 h) in duplicate. The final concentration in column 1 was, therefore, 0.1 mM for each compound. Cells were cultured for an additional 72 h, or approximately three replication cycles.
On day 5, once cells cultured without compounds were approximately 70% confluent, the assay medium was removed and 4% paraformaldehyde (100 μl per well) added. Plates were incubated for 20 min. The paraformaldehyde was removed and cells were washed three times with PBS (200 μl per well per wash). PBS was removed and 0.5% acetic acid (100 μl per well) was added. Plates were incubated for 5 min. Acetic acid was removed and cells washed once with PBS (200 μl per well). PBS was removed and 0.1% Triton X-100 (100 μl per well) was added; plates were incubated for 30 min. Triton X-100 was removed. Blocking buffer (2 ×; 200 μl per well) was added and plates were incubated for 2 h. Blocking buffer was removed. Primary antibody solution (100 μl per well) was added to columns 1–11, while 1 × blocking buffer was added to column 12. Plates were incubated for 16 h to 18 h at 4 °C.
On day 6, primary antibody was removed and cells were washed three times with PBS containing 0.1% Triton X-100 (PBS-T; 300 μl per well per wash). PBS-T was removed, secondary antibody solution (100 μl per well) was added, and plates were incubated for 1 h. Secondary antibody solution was removed and cells were washed four times with PBS-T (300 μl per well each wash). PBS-T was removed and alkaline phosphatase development solution was added (100 μl per well). Plates were incubated at room temperature for 15 min; the endpoint was immediately measured at OD405 using the FLUOstar OPTIMA plate reader (BMG LabTech).
To complete the assay, alkaline phosphatase development solution was removed and horseradish peroxidase development solution (100 μl per well) was added. Plates were incubated for 15 min; the endpoint was immediately measured at OD595 using the FLUOstar OPTIMA plate reader. Horseradish peroxidase development solution was completely removed and 1× Janus Green stain (50 μl per well) was added. Plates incubated for 5 min and then washed with five times (300 μl per well) with distilled water. The water was removed and 0.5 M hydrochloric acid added (100 μl per well) to stop the reaction. Plates were incubated for 10 min with shaking at 120 rpm. The Janus Green B endpoint was measured at OD595 using the FLUOstar OPTIMA plate reader.
COX1 signal is first subtracted by the background signal. The background signal was designated as plate column 12, which was not exposed to primary antibodies. The resulting COX1 signal is then divided by the JG background-subtracted signal. COX1/JG is then divided by the values from untreated cells and multiplied by 100 to calculate the percentage COX1/JG.
Percentage response = [concentration compound] μM COX1 ratio / 0 μM COX1 ratio × 100
Using GraphPad Prism (GraphPad), the percentage ratio against the log of the compound concentration was graphed and the EC50 concentration then extrapolated where the trend curve intersected the 50% ratio using a standard four-parameter fit.
Preparation of MK-7762 stalled ribosome complex
Two liters of M. smegmatis was cultured to an OD600 of 0.5 in 7H9 medium. Cells were washed, centrifuged and cryo-lysed using a Retsch Mixer Mill 400. Lysate was applied to a 1.1-M sucrose cushion and centrifuged for 1 h and 45 min at 310,000g using a Ti-70 rotor. The pellet was resuspended and purified on a 5−50% sucrose gradient at 164,000g for 2.5 h using a SW 41 Ti rotor. Fractions were recovered using a Beckman Coulter Fraction Recovery System, and absorbance at 260 nm was recorded for each fraction. Fractions corresponding to 70S ribosomes were combined and ultrafiltered using a 10-kDa molecular-weight cutoff an Amicon Ultra Centrifugal Filter to concentrate ribosomes and exchange buffers. Stalled ribosome–drug complexes were generated using a PURExpress ∆ Ribosome kit with MK-7762, purified ribosomes, RNase inhibitor and DNA stalling template. The ribosome–drug complex was re-isolated as previously described16 through a sucrose gradient, fractionation and ultrafiltration. Purified stalled ribosome complexes were incubated for 1 h at 4 °C with 60 µM MK-7726. Before freezing to grids, the sample was filtered through a Millipore low-binding Durapore PVDF 0.22-µm filter for 5 min at 14,000g.
Cryo-EM
A standard workflow was followed for the cryo-EM sample preparation and grid vitrification. Data were collected on a Krios G4 transmission electron microscope (Thermo Fisher Scientific) with a K3 camera and a Biocontinuum GIF and processed in cryoSPARC25. M. smegmatis ribosome structure (PDB 5O61) was docked into the cryo-EM map in ChimeraX26. Models were built in COOT27 and refined in PHENIX28. Details are provided in the Supplementary Information. Figures were made with ChimeraX and PyMOL (The PyMOL Molecular Graphics System, version 3.0 Schrödinger).
Before freezing, the stalled ribosome complexes were filtered for 5 min at 12,000 at 4 °C using a 0.22-µm low-binding Durapore PVDF filter (Millipore). Quantifoil 2/2 300 mesh (Quantifoil Micro Tools) copper grids were glow-discharged with a 50:50 oxygen–hydrogen mixture in a Solarus 950 (Gatan) for 30 s. Grids were loaded into an EM GP2 plunge freezer (Leica), which was set to 4 °C and 95% humidity, and a 3-μl droplet of the sample was added. The sample was subsequently blotted for 3 s, 3.5 s or 4 s, followed immediately by plunge freezing in liquid ethane at −184 °C. Grids were mounted in AutoGrid assemblies.
Data were collected using a Krios G4 (Thermo Fisher Scientific) transmission electron microscope operating at 300 kV equipped with a K3 (Gatan) and a Biocontinuum GIF (Gatan) operating at a slit width of 20 eV. Image acquisition was performed with a 0° or 30° stage tilt. Movies were acquired at a pixel size of 0.4131 Å per pixel with a dose of ~50 e−/Å2 per movie using EPU (Thermo Fisher Scientific). Defocus values were set to cycle between 0.5 μm and 2.5 μm. Data quality was monitored during data collection using cryoSPARC Live v4.4.
Complete data processing was performed using cryoSPARC v4.5 with only the tilted data. Patch motion correction, dose-fractionated weighting and patch contrast transfer function estimation were carried out with default parameters. Blob picking was used to pick particles for the first round. High-quality two-dimensional (2D) classes, after several rounds of 2D classification, were used as templates for template-based particle picking. To clean up the data, extracted particles were classified in 2D for as many rounds as necessary. Ab initio volumes were determined with five classes, followed by heterogeneous, homogeneous and nonuniform refinement. Two rounds of global and local contrast transfer function refinement, reference motion correction and nonuniform refinement were carried out for the best class (Extended Data Fig. 3). No symmetry was imposed during refinement. The final reconstructed map is at 2.0-Å resolution (Extended Data Fig. 4). The local resolution of the final reconstruction was calculated in cryoSPARC and displayed in ChimeraX (Extended Data Fig. 4).
Map output by cryoSPARC nonuniform refinement was sharpened by PHENIX.auto_sharpen. The M. smegmatis ribosome structure (PDB 5O61) was docked into the sharpened map in ChimeraX and then refined by molecular dynamics using the ISOLDE plugin. Cycles of manual model building in COOT v0.9, followed by real-space refinement using PHENIX.real_space_refine, were repeated many times until no noticeable improvement could be observed. MolProbity was used for model validation.
PK and metabolic identification in mouse, rat and dog
Animals and dose administration for PK studies
Eight- to ten-week-old male C57BL/6 mice (approximately 25 g) and 12- to 14-week-old male and female Wistar Han rats (272−356 g for male, 220−230 g for female) were purchased from Charles River Laboratories. Male Beagle dogs (1,015 kg) were purchased from Marshall BioResources. All animals used in the low-dose intravenous/oral dose PK studies were fasted overnight before dosing, with water provided ad libitum. Male rats used for high-dose studies were also fasted before dosing. Additional male and female rats and male dogs used for high oral dose studies were fed before dosing.
Mice
MK-7762 was administered intravenously or orally (n = 3) to fasted male mice at 2 mg per kg body weight (1 ml per kg body weight) or 10 mg per kg body weight (10 ml per kg body weight), respectively, in DMSO:propylene glycol:water (20:60:20). Blood (10 μl) was collected via tail vein before dosing and at 0.08 h, 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 7 h and 24 h after dosing for the intravenous arm, and before dosing and at 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 7 h and 24 h after dosing for the oral arm. Three volumes of sodium citrate were added to the whole blood in matrix bullet tubes, vortexed, and then frozen immediately and stored at −70 °C until analysis.
Rats
MK-7762 was administered intravenously to fasted male rats via a jugular cannula at 2 mg per kg body weight (1 ml per kg body weight) in 30% Captisol (n = 3). Blood was collected before dosing and at 0.03h, 0.13 h, 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 7 h, 12 h, 18 h and 24 h after dosing into tripotassium ethylenediaminetetraacetic acid (K3EDTA) tubes. The blood was centrifuged to obtain plasma. Plasma samples were frozen immediately and stored at −70 °C until analysis. For the low-dose oral studies, MK-7762 was administered to fasted male rats at 5 mg per kg body weight in 10% Tween 80 or at 10 mg per kg body weight in 0.5% methyl cellulose (MC) with a dosing volume of 5 ml per kg body weight (n = 3). Blood was collected before dosing and at 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 7 h, 12 h (5 mg per kg body weight arm only), 18 h and 24 h after dosing into K3EDTA tubes. The blood was centrifuged to obtain plasma and processed as described above. For the high-dose oral studies, MK-7762 was administered to fasted male rats at 100 mg per kg body weight, 500 mg per kg body weight or 1,000 mg per kg body weight in 10% Tween 80, with a dosing volume of 5 ml per kg body weight (n = 3). Blood (0.25 ml) was collected before dosing and at 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 7 h, 12 h, 18 h and 24 h after dosing into K3EDTA tubes. The blood was centrifuged to obtain plasma and processed as described above. MK-7762 was also administered to fed male and female rats at 50 mg per kg body weight or 1,000 mg per kg body weight in 10% Tween 80, with a dosing volume of 5 ml per kg body weight (n = 3). Blood was collected before the dose and at 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 7 h, 12 h, 18 h, 24 h and 48 h after dosing into K3EDTA tubes. The blood was centrifuged to obtain plasma and processed as described above. Urine samples were also collected at 0−24 h and 24−48 h after the dose and stored at −70 °C until processing.
Dogs
MK-7762 was administered intravenously to fasted male dogs (n = 3) via a saphenous/cephalic vein catheter at 1 ml per kg body weight in 30% Captisol solution to achieve the target dose of 1 mg per kg body weight. Blood was collected before dosing and at 0.03 h, 0.13 h, 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 6 h and 24 h after dosing into K3EDTA tubes. The blood was centrifuged to obtain plasma and processed as described above. For the low-dose oral studies, MK-7762 was administered to fasted male dogs at 2 mg per kg body weight in 10% Tween 80 or at 2 mg and 10 mg per kg body weight in 0.5% MC with a dosing volume of 5 ml per kg body weight (n = 3). Blood was collected before dosing and at 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 6 h and 24 h after dosing into K3EDTA tubes. The blood was centrifuged to obtain plasma and processed as described above. For the additional 10 mg per kg body weight and high-dose oral studies, MK-7762 was administered to fed male dogs at 10 mg, 30 mg and 100 mg per kg body weight, or 200 mg per kg body weight in 10% Tween 80, with a dosing volume of 5 ml per kg body weight (n = 3). Blood was collected before the dose and at 0.25 h, 0.5 h, 1 h, 2 h, 4 h, 6 h and 24 h after the dose into K3EDTA tubes. The blood was centrifuged to obtain plasma and processed as described above.
Animals and dose administration for metabolite identification
MK-7762 was administered orally to two fasted male WH rats (360−400 g) at 5 ml per kg body weight (10 mg per kg body weight) in 10% Tween 80. Blood was collected at 4 h and 24 h after dosing via cardiac puncture. Plasma was obtained by centrifugation. Urine was collected into pre-tared containers for up to 24 h after dosing from one of the rats. All samples were stored at −70 °C until processing.
MK-7762 was administered orally to a fasted male Beagle dog (11 kg) at 5 ml per kg body weight (10 mg per kg body weight) in 0.5% MC via gavage. Blood (1 ml) was collected at 4 h and 24 h after dosing into K3EDTA tubes. Plasma was obtained by centrifugation. Urine was collected into pre-tared containers for up to 24 h after the dose. All samples were stored at −70 °C until processing.
Sample processing for PK
Blood, plasma and urine samples were processed via protein precipitation extraction using a Hamilton Microlab STAR liquid handling system (Hamilton Robotics) and Microlab STAR software (version 4.1.1.3714). The supernatant was analyzed by liquid chromatography with LC–MS/MS.
Sample processing for metabolite identification
Aliquots (200 μl) of rat plasma samples at 4 h or 24 h were precipitated with an equal volume of 90% acetonitrile and 10% methanol. The samples were vortex mixed for 2 min and then centrifuged at 10,000 rpm for 10 min. The supernatants were transferred to glass vials for high-resolution mass spectrometry (HRMS) analysis. Aliquots (200 μl) of dog plasma samples at 4 h or 24 h were precipitated with two volumes of 90% acetonitrile and 10% methanol. The samples were vortex mixed for 1 min and then centrifuged at 10,000 rpm for 10 min. The supernatants were transferred to glass vials and then diluted at a 1:1 ratio with water containing 0.1% formic acid for HRMS analysis. An aliquot of the 0−24-h urine samples from a rat or a dog (200 μl) was centrifuged at 10,000 rpm for 10 min and then transferred to a glass vial for HRMS analysis.
LC–MS/MS analysis for PK
The concentrations of MK-7762 in the mouse blood, rat and dog plasma, and rat urine were determined by comparison to a calibration curve prepared in the corresponding matrix, using liquid chromatography separation followed by triple-quadrupole mass spectrometry detection. For each study and matrix, two calibration curves were prepared. The standards in the calibration curve for the PK studies had nominal values of 1 nM to 10,000 nM.
The LC–MS/MS system comprised a Thermo Transcend LX2 multiplexed UPLC, Thermo Dionez Ultimate 3000 RS pumps (Thermo Fisher Scientific) and an Applied Biosystems/MDS Sciex API4500, API5000, API 5500 or API 6500 triple-quadrupole mass spectrometer. Chromatographic separation of the analytes was achieved on a Waters XSELECT HSS TS XP column (5 cm × 2.1 mm × 2.5 μm) with gradient conditions and mobile phase A (100% water with 0.1% formic acid) and B (100% acetonitrile with 0.1% formic acid). The flow rate was held constant at 0.75 ml min−1.
Analyte response was measured by multiple reaction monitoring (MRM) of unique transitions using positive ion mode at m/z 420.1 → 388.0 or 295.9 for MK-7762, and m/z 281.3 → 193.1 for the internal standard imipramine.
LC–HRMS analysis for metabolite identification
Chromatographic separation of MK-7762 and its metabolites was conducted using an Acquity UPLC (Waters) with a Waters Acquity Xbridge BEH C18 column (2.1 mm × 10 cm, 1.7 μm) at 40 °C. The mobile phase A consisted of 0.1% formic acid in water and mobile phase B consisted of 0.1% formic acid in acetonitrile. The flow rate was held constant at 0.5 ml min−1.
The mass spectrometry analysis was conducted using an AB Sciex 6600 time of flight mass spectrometer (AB Sciex) equipped with an electrospray ionization source and operated in the positive-ionization mode. The automated calibration device system performed an external calibration every five samples. The Duo Spray ion source equipped with a stainless-steel electrode (100-μm internal diameter) was operated with the following mass spectrometry conditions: gas 1, nitrogen (50 psi); gas 2, nitrogen (55 psi); ion spray voltage, 5500; ion source temperature, 500 °C; curtain gas, nitrogen (30 psi); and collision energy, 5 eV. The mass spectrometer was operated in the SWATH acquisition mode where one complete cycle consists of a survey scan and 22 Q1 isolation scans covering a mass range of 100–1,000 m/z. The survey scan covered a mass range of 100 to 1,000 m/z with an accumulation time of 0.13 s. The Q1 isolation strategy covered a mass range of 100 to 1,000 m/z with a variable Dalton SWATH window for Q1 isolation (overlap 1 U). In each SWATH window, a collision energy of 35 eV with a spread of ±15 eV and an accumulation time of about 0.02 s in high-sensitivity mode were used. The total cycle time was 0.6 s. All mass spectrometry parameters were controlled by Analyst TF Software 1.7 from Sciex. Data processing was accomplished using Mass-MetaSite (version CLI 5.2.0-8 Mass 3.4.0 ×64) and interpretation was done in WebMetabase.
Calculation of PK parameters
PK parameters were calculated using non-compartmental methods in Watson. The area under the plasma concentration–time curve (AUC0−t) was calculated from the first time point (0 min) up to the last time point with a measurable drug concentration using the linear trapezoidal or linear/log-linear trapezoidal rule. The concentration at 0 h after intravenous administration was back-extrapolated using the first two time points. The remaining area under the plasma concentration–time curve (AUCt−∞) was estimated by dividing the observed concentration at the last time point by the elimination rate constant. This value was added to AUC0−t to estimate the AUC0−∞. The intravenous plasma clearance was calculated by dividing the dose by AUC0−∞. The terminal half-life of elimination was determined by unweighted linear regression analysis of the log-transformed data. The time points for determination of half-life were selected by visual inspection of the data. The volume of distribution (Vd,ss) was obtained from the product of plasma clearance and mean residence time (determined by dividing the area under the first moment curve by the area under the curve). The Cmax and the Tmax were obtained by inspection of the plasma concentration–time data. Absolute oral bioavailability was determined from dose-adjusted intravenous and oral AUC0−∞ ratios.
Murine efficacy models of acute and chronic TB infection
Data were analyzed in GraphPad Prism version 10.2 using a one-way ANOVA, and statistical details of these comparisons are shown in Supplementary Tables 8 and 9. Six animals per treatment group were used considering historical variability29. Mice were housed in containment caging in an Animal Biosafety Level 3 containment facility at 72 °F and 50% relative humidity with a 12-h light cycle.
BALB/c acute TB infection model
Six- to eight-week-old female specific-pathogen-free BALB/c mice (Jackson laboratories) were aerosol infected with a Glas-Col inhalation exposure system using previously calibrated frozen Mtb Erdman pFCA-LuxAB (Erdman-Lux) that was thawed and diluted before use as described30,31. The average bacterial load in lungs one day after low-dose aerosol was approximately 131 CFUs. Six infected mice were euthanized 7 days after infection as pretreatment controls. Ten mice were left untreated to serve as the study comparator. Drugs were prepared in weekly batches in 10% Tween 80/40% PEG400/50% water, and aliquots were held at 4 °C for daily use. Drugs were administered for 12 consecutive days by oral gavage in a volume of 0.2 ml per mouse once daily or twice daily, as indicated.
BALB/c chronic TB infection model
Six- to eight-week-old female specific-pathogen-free BALB/c mice (Jackson laboratories) were aerosol infected with a Glas-Col inhalation exposure system using previously calibrated frozen Mtb Erdman-Lux that was thawed and diluted before use32. The average lung bacterial load one day after low-dose aerosol was approximately 147 CFUs. Six infected mice were euthanized 28 days after infection as pretreatment controls. Eight mice were left untreated to serve as the study comparator. Drugs were prepared in weekly batches as above, and aliquots were held at 4 °C for daily use. Drugs were administered daily 7 days per week by oral gavage in a volume of 0.2 ml per mouse once daily.
Bacterial enumeration acute and chronic models
At given time points, organs were aseptically harvested from mice after a weekend drug-free holiday following the last day of dosing to allow drug clearance from tissues. Tissues were frozen in 7 ml Bertin Precellys tubes (CKmix50_7 mL P000939-LYSK0A). Tissues were thawed and homogenized (Precellys, Bertin Instruments) in 4.5 ml PBS plus 10% (wt/vol) bovine serum albumin (PBS-BSA). Portions of the homogenates from animals on treatment were serially diluted in PBS-BSA and plated for CFUs on 7H11-OADC agar (that is, Middlebrook 7H11 agar plates supplemented 0.2% (vol/vol) glycerol, 10% (vol/vol) oleic acid-albumin-dextrose-catalase (OADC) supplement, and 0.01 mg ml-1 cycloheximide, and 0.05 mg ml-1 carbenicillin) further supplemented with 0.4% (wt/vol) activated charcoal (7H11 charcoal agar) to help counteract drug carryover artifacts33,34. Plates were incubated for at least 21 days at 37 °C and for 6 weeks total to confirm all CFUs were quantified.
Lesion-penetration studies
Eight- to ten-week-old female specific-pathogen-free C3HeB/FeJ mice (Jackson laboratories) were aerosol infected with a Glas-Col inhalation exposure system to achieve approximately 75 CFUs deposited in lungs one day following infection with previously calibrated frozen Mtb Erdman (TMCC 107)34. Mice (n = 5 or 6 mice per study arm) were dosed starting 10 weeks after aerosol with MK-7762 or linezolid each at 100 mg per kg body weight, once daily. Drugs were prepared as described for the efficacy studies above. Treatment occurred on 5 of 7 days per week (Monday through Friday) for a total of 2.5 weeks to reach steady-state drug levels. Mice were euthanized, and plasma and tissues were collected at the indicated time points.
Whole blood was obtained via cardiac puncture and processed in plasma separator tubes (Becton, Dickinson and Co.) centrifuged at 3,750g for 2 min at 4 °C, aliquoted into microcentrifuge tubes and stored at −80 °C until analysis. Mice with pronounced lung pathology were selected to collect samples for spatial drug quantification by gravity-assisted LCM. Briefly, whole lung samples consisting of the cranial, medial and accessory lung lobes were collected on clear disposable base molds with the desired cutting surface in direct contact with the base of the tray (Fisher Scientific). Collection trays are then placed onto a prechilled 4-inch aluminum block in 2 inches of liquid nitrogen in a Styrofoam cooler and allow to sit covered for 10 min. Tissue trays containing frozen lobes were wrapped in foil squares, placed individually into labeled ziplock bags and immediately transferred onto dry ice. Samples were stored at −80 °C until analysis.
LCM
Twenty-five-micron-thick tissue sections were cut from C3HeB/FeJ mouse lung biopsy samples using a Leica CM 1860UV cryostat and thaw-mounted onto 1.4-µm-thick Leica PET-Membrane Frame Slides for LCM. Tissue sections were immediately stored in sealed containers at −80 °C. Adjacent 10-µm-thick tissue sections were thaw-mounted onto standard glass microscopy slides for H&E staining. Cellular, necrotic (caseum) and uninvolved lung lesion areas totaling 3 million µm2 were dissected from one to three serial lung biopsy tissue sections using a Leica LMD6 system set up within a Biosafety Level 3 facility. Areas of cellular and caseous lesions were identified optically from the brightfield image scan and by comparison to the adjacent H&E reference tissue. Pooled dissected lesion tissues were collected into 0.25-ml standard PCR tubes and immediately transferred to −80 °C. Processing of LCM samples was performed by adding 10 µl of each working standard solution to 2 µl of 1:26.7 diluted in PBS drug-free tissue homogenate to create standard curve and quality-control (QC) samples. MK-7762 was extracted from the calibration standard, QC, blank/control and LCM study samples by the addition of 50 µl of extract solvent containing imipramine. Extracts were sonicated for 10 min and centrifuged at 4,000 rpm for 5 min. Around 50 µl of supernatant was transferred to a 96-well plate and 50 µl of water was added for LC–MS/MS analysis.
Drug quantification in plasma and infected tissues by HPLC–MS/MS
Verapamil internal standard was purchased from Sigma-Aldrich. Drug-free K2EDTA plasma from BioIVT was used as a blank matrix to build standard curves. Neat 1 mg ml−1 DMSO stock for MK-7762 was serial diluted in 50/50 acetonitrile/water to create standard curves and QC spiking solutions. Spiked matrix standards and QC samples were created by adding 10 µl of spiking solutions to 90 µl of drug-free plasma and control tissue homogenate. Extraction was performed for standards, QC samples and study samples by adding 200 µl of a 1:1 acetonitrile–methanol mixture containing 10 ng ml−1 verapamil to 20 µl of plasma or homogenized tissue sample. One hundred microliters of extracts were diluted with 100 µl of water and vortexed for 2 min before injecting into the LC system.
LC–MS/MS analysis was performed on a Sciex Qtrap 6500+ triple-quadrupole mass spectrometer coupled to a Shimadzu Nexera X2 UHPLC system to quantify each drug in plasma. Chromatography was performed on an Agilent Zorbax SB-C8 column (2.1 × 30 mm; particle size, 3.5 µm) using a reverse-phase gradient elution with an aqueous mobile phase. Water with 0.1% formic acid was used for the aqueous mobile phase and 0.1% formic acid in acetonitrile for the organic mobile phase. MRM of precursor/fragment transitions in electrospray positive-ionization mode was used to quantify the analytes. MRM transition of 420.00/388.00 and 455.40/165.00 were used for MK-7762 and verapamil, respectively. Sample analysis was accepted if the concentrations of the QC samples were within 20% of the nominal concentration. Data processing was performed using Analyst software (version 1.6.3; Sciex).
Statistics and reproducibility
No statistical method was used to predetermine sample size. For the efficacy studies, sample size was chosen based on prior studies29. No data were excluded from the analyses. One-way ANOVA followed by Tukey’s multiple-comparison test was performed in GraphPad Prism (v10.4.1) for Windows (GraphPad Software; https://www.graphpad.com/). Data distribution was assumed to be normal, but this was not formally tested. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are available in the main text or the Supplementary Information except the data for Fig. 1b,d, which includes structures of all analogs synthesized and which will be disclosed in a subsequent paper focused on the medicinal chemistry program. Additional information and data can be requested from C.E.B., who will respond within 2 weeks.
References
World Health Organization. Global Tuberculosis Report 2023 (2023).
Motta, I. et al. Recent advances in the treatment of tuberculosis. Clin. Microbiol. Infect. https://doi.org/10.1016/j.cmi.2023.07.013 (2023).
Lange, C., Barry, C. E. III & Horsburgh, C. R. Jr. Treatments of multidrug-resistant tuberculosis: light at the end of the tunnel. Am. J. Respir. Crit. Care Med. 205, 1142–1144 (2022).
Lee, M. et al. Linezolid for treatment of chronic extensively drug-resistant tuberculosis. N. Engl. J. Med. 367, 1508–1518 (2012).
Lee, M. et al. Linezolid for XDR-TB–final study outcomes. N. Engl. J. Med. 373, 290–291 (2015).
Nyang’wa, B. T. et al. A 24-week, all-oral regimen for rifampin-resistant tuberculosis. N. Engl. J. Med. 387, 2331–2343 (2022).
World Health Organization. WHO consolidated guidelines on tuberculosis. Module 4: treatment - drug-resistant tuberculosis treatment, 2022 update (2022).
Conradie, F. et al. Treatment of highly drug-resistant pulmonary tuberculosis. N. Engl. J. Med. 382, 893–902 (2020).
McKee, E. E., Ferguson, M., Bentley, A. T. & Marks, T. A. Inhibition of mammalian mitochondrial protein synthesis by oxazolidinones. Antimicrob. Agents Chemother. 50, 2042–2049 (2006).
Soriano, A., Miro, O. & Mensa, J. Mitochondrial toxicity associated with linezolid. N. Engl. J. Med. 353, 2305–2306 (2005).
Song, T. et al. Linezolid trough concentrations correlate with mitochondrial toxicity-related adverse events in the treatment of chronic extensively drug-resistant tuberculosis. EBioMedicine 2, 1627–1633 (2015).
Betts, J. C. et al. Signature gene expression profiles discriminate between isoniazid-, thiolactomycin-, and triclosan-treated Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 47, 2903–2913 (2003).
Dutta, N. K., Mehra, S. & Kaushal, D. A Mycobacterium tuberculosis sigma factor network responds to cell-envelope damage by the promising anti-mycobacterial thioridazine. PLoS ONE 5, e10069 (2010).
Waddell, S. J. et al. The use of microarray analysis to determine the gene expression profiles of Mycobacterium tuberculosis in response to anti-bacterial compounds. Tuberculosis 84, 263–274 (2004).
Bibel, B. et al. Context-specific inhibition of mitochondrial ribosomes by phenicol and oxazolidinone antibiotics. Nucleic Acids Res. https://doi.org/10.1093/nar/gkaf046 (2025).
Tsai, K. et al. Structural basis for context-specific inhibition of translation by oxazolidinone antibiotics. Nat. Struct. Mol. Biol. 29, 162–171 (2022).
Wright, A. et al. Characterization of the core ribosomal binding region for the oxazolidone family of antibiotics using Cryo-EM. ACS Pharmacol. Transl. Sci. 3, 425–432 (2020).
Desai, N. et al. Elongational stalling activates mitoribosome-associated quality control. Science 370, 1105–1110 (2020).
Prideaux, B. et al. The association between sterilizing activity and drug distribution into tuberculosis lesions. Nat. Med. 21, 1223–1227 (2015).
Zimmerman, M., Blanc, L., Chen, P. Y., Dartois, V. & Prideaux, B. Spatial quantification of drugs in pulmonary tuberculosis lesions by laser capture microdissection liquid chromatography mass spectrometry (LCM-LC/MS). J Vis. Exp. https://doi.org/10.3791/57402 (2018).
Lanoix, J. P., Lenaerts, A. J. & Nuermberger, E. L. Heterogeneous disease progression and treatment response in a C3HeB/FeJ mouse model of tuberculosis. Dis. Model Mech. 8, 603–610 (2015).
Yew, W. W. Directly observed therapy, short-course: the best way to prevent multidrug-resistant tuberculosis. Chemotherapy 45, 26–33 (1999).
Simeon, S., Garcia-Cremades, M., Savic, R. & Solans, B. P. Pharmacokinetic-pharmacodynamic modeling of tuberculosis time to positivity and colony-forming unit to assess the response to dose-ranging linezolid. Antimicrob. Agents Chemother. 68, e0019024 (2024).
Alghamdi, W. A. et al. Population pharmacokinetics of linezolid in tuberculosis patients: dosing regimen simulation and target attainment analysis. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.01174-20 (2020).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Franzblau, S. G. et al. Comprehensive analysis of methods used for the evaluation of compounds against Mycobacterium tuberculosis. Tuberculosis 92, 453–488 (2012).
Akester, J. N. et al. Synthesis, structure-activity relationship, and mechanistic studies of aminoquinazolinones displaying antimycobacterial activity. ACS Infect. Dis. 6, 1951–1964 (2020).
Ray, P. C. et al. Spirocycle MmpL3 inhibitors with improved hERG and cytotoxicity profiles as inhibitors of Mycobacterium tuberculosis growth. ACS Omega 6, 2284–2311 (2021).
Lenaerts, A. J. et al. Preclinical testing of the nitroimidazopyran PA-824 for activity against Mycobacterium tuberculosis in a series of in vitro and in vivo models. Antimicrob. Agents Chemother. 49, 2294–2301 (2005).
Xu, J. et al. Contribution of pretomanid to novel regimens containing bedaquiline with either linezolid or moxifloxacin and pyrazinamide in murine models of tuberculosis. Antimicrob. Agents Chemother https://doi.org/10.1128/AAC.00021-19 (2019).
Ramey, M. E. et al. Drug distribution and efficacy of the DprE1 inhibitor BTZ-043 in the C3HeB/FeJ mouse tuberculosis model. Antimicrob. Agents Chemother. 67, e0059723 (2023).
Acknowledgements
This work was funded (in part) by the Bill and Melinda Gates Foundation (INV-001727 to C.E.B.), and in part by the Division of Intramural Research of the NIAID (to C.E.B.). V.D. acknowledges funding from the Bill and Melinda Gates Foundation (INV-004704). G.T.R. acknowledges funding from the Bill and Melinda Gates Foundation (INV-009105). Merck also financially contributed to these studies.
Author information
Authors and Affiliations
Contributions
B.M.C., H.I.B., V.Y.T., R.R.M., J.E., C.W.B., K.Y., P.G.N., L.Y., R.E.P., E.B.C., J.L.H., S.O., D.L.H., M.L., A.A., M.A.O., M.Z., F.K., L.M.M., M.E.R., A.A.B., A.J.L., G.T.R., V.D., C.E.B., C.D.W. and D.B.O. were involved in the conception of the study and design of the experiments. H.I.B., V.Y.T., A.B., J.Z., F.H., R.E.P., S.O., M.Z., F.K., L.M.M., M.E.R. and A.A.B. were involved in the acquisition of the data, including conducting the research and performing the experiments. B.M.C., H.I.B., G.T.R., V.D., C.E.B., C.D.W. and D.B.O. were involved in the interpretation of the data. B.M.C. and C.E.B. were involved in the drafting of the initial version of the manuscript. All authors reviewed the draft critically for important intellectual content and approved the manuscript before it was submitted by the corresponding authors.
Corresponding authors
Ethics declarations
Competing interests
B.M.C., R.R.M., C.W.B., K.Y., P.G.N., J.S., L.Y., R.E.P., E.B.C., J.L.H. and D.B.O. are employees of Merck and declare owning stock in the company. The other authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Kevin Pethe, John Rubinstein and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Lia Parkin, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Hydroxylation site of MK-7762 by upregulation of the dioxygenase encoded by Rv3161c.
Left triple quad mass spectrometric fragmentation of MK-7762 showing the difluoro aromatic nucleus release as the Collison Energy is increased. Center showing the fragmentation pattern of the mono-oxygenated analog +16. Right shows the di-hydroxylated analog likewise produces a fragment with both hydroxyl groups on the same ring.
Extended Data Fig. 2 Quantitative PCR testing the expression level of Rv3161c in the wild type and three in vitro selected mutants resistant to MK-7762.
RNA was isolated from all four strains early in log phase either with or without treatment with 2 µM MK-7762 for 2 h. The data represent two biological replicates of this.
Extended Data Fig. 3 Cryoelectron microscopy workflow for stalled mycobacterial ribosomes with MK-7762.
Workflow showing the data processing for structure determination from 7,925 movies.
Extended Data Fig. 4 Cryoelectron microscopy resolution statistics.
A. GSFSC resolution after masking and correction. B. Number of images processed by elevation and orientation. C. Local resolution map across the structure. D. Local density at the binding site.
Extended Data Fig. 5 Mitotoxicity of MK-7762 and two reduced analogs showing the sulfone is less toxic than either the sulfoxide or the thioether.
The assay was repeated three times on different days and the values and error bars shown represent the mean and the standard error of the mean for these.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–9 and Supplementary Notes (synthetic procedures, spectral data and mass spectrometry).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Crowley, B.M., Boshoff, H.I., Boving, A. et al. Discovery and development of a new oxazolidinone with reduced toxicity for the treatment of tuberculosis. Nat Med 32, 553–560 (2026). https://doi.org/10.1038/s41591-025-04164-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41591-025-04164-x