Abstract
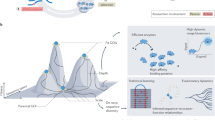
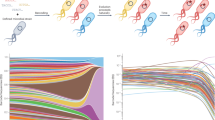
Evolution occurs when selective pressures from the environment shape inherited variation over time. Within the laboratory, evolution is commonly used to engineer proteins and RNA, but experimental constraints have limited the ability to reproducibly and reliably explore factors such as population diversity, the timing of environmental changes and chance on outcomes. We developed a robotic system termed phage- and robotics-assisted near-continuous evolution (PRANCE) to comprehensively explore biomolecular evolution by performing phage-assisted continuous evolution in high-throughput. PRANCE implements an automated feedback control system that adjusts the stringency of selection in response to real-time measurements of each molecular activity. In evolving three distinct types of biomolecule, we find that evolution is reproducibly altered by both random chance and the historical pattern of environmental changes. This work improves the reliability of protein engineering and enables the systematic analysis of the historical, environmental and random factors governing biomolecular evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data are publicly available at https://doi.org/10.17632/h9z94f9y6p.1 (https://data.mendeley.com/datasets/h9z94f9y6p/1). Source data are provided with this paper.
Code availability
All code will be made available via github, see https://github.com/dgretton/std-96-pace and https://github.com/dgretton/roboplaque.
References
Good, B. H., McDonald, M. J., Barrick, J. E., Lenski, R. E. & Desai, M. M. The dynamics of molecular evolution over 60,000 generations. Nature 551, 45–50 (2017).
Barrick, J. E. et al. Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature 461, 1243–1247 (2009).
Zhao, H., Giver, L., Shao, Z., Affholter, J. A. & Arnold, F. H. Molecular evolution by staggered extension process (StEP) in vitro recombination. Nat. Biotechnol. 16, 258–261 (1998).
Romero, P. A. & Arnold, F. H. Exploring protein fitness landscapes by directed evolution. Nat. Rev. Mol. Cell Biol. 10, 866–876 (2009).
Esvelt, K. M., Carlson, J. C. & Liu, D. R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Berman, C. M. et al. An adaptable platform for directed evolution in human cells. J. Am. Chem. Soc. 140, 18093–18103 (2018).
English, J. G. et al. VEGAS as a platform for facile directed evolution in mammalian cells. Cell https://doi.org/10.1016/j.cell.2019.05.051 (2019).
Crook, N. et al. In vivo continuous evolution of genes and pathways in yeast. Nat. Commun. 7, 13051 (2016).
Ravikumar, A., Arzumanyan, G. A., Obadi, M. K. A., Javanpour, A. A. & Liu, C. C. Scalable, continuous evolution of genes at mutation rates above genomic error thresholds. Cell 175, 1946–1957.e13 (2018).
Leconte, A. M. et al. A population-based experimental model for protein evolution: effects of mutation rate and selection stringency on evolutionary outcomes. Biochemistry 52, 1490–1499 (2013).
Dickinson, B. C., Leconte, A. M., Allen, B., Esvelt, K. M. & Liu, D. R. Experimental interrogation of the path dependence and stochasticity of protein evolution using phage-assisted continuous evolution. Proc. Natl Acad. Sci. USA 110, 9007–9012 (2013).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature https://doi.org/10.1038/nature26155 (2018).
Wong, B. G., Mancuso, C. P., Kiriakov, S., Bashor, C. J. & Khalil, A. S. Precise, automated control of conditions for high-throughput growth of yeast and bacteria with eVOLVER. Nat. Biotechnol. 36, 614–623 (2018).
Horinouchi, T., Minamoto, T., Suzuki, S., Shimizu, H. & Furusawa, C. Development of an automated culture system for laboratory evolution. J. Lab. Autom. 19, 478–482 (2014).
Zhong, Z. et al. Automated continuous evolution of proteins in vivo. ACS Synth. Biol. 9, 1270–1276 (2020).
Meyer, M. M. et al. Library analysis of SCHEMA-guided protein recombination. Protein Sci. 12, 1686–1693 (2003).
Crameri, A., Raillard, S. A., Bermudez, E. & Stemmer, W. P. DNA shuffling of a family of genes from diverse species accelerates directed evolution. Nature 391, 288–291 (1998).
Karanicolas, J. et al. A de novo protein binding pair by computational design and directed evolution. Mol. Cell 42, 250–260 (2011).
Amini, Z. N. & Müller, U. F. Low selection pressure aids the evolution of cooperative ribozyme mutations in cells. J. Biol. Chem. 288, 33096–33106 (2013).
Zinkus-Boltz, J., DeValk, C. & Dickinson, B. C. A phage-assisted continuous selection approach for deep mutational scanning of protein–protein interactions. ACS Chem. Biol. 14, 2757–2767 (2019).
Blount, Z. D., Lenski, R. E. & Losos, J. B. Contingency and determinism in evolution: replaying life’s tape. Science 362, eaam5979 (2018).
Chory, E. J., Gretton, D. W., DeBenedictis, E. A. & Esvelt, K. M. Enabling high-throughput biology with flexible open-source automation. Mol. Syst. Biol. 17, e9942 (2021).
Balagaddé, F. K., You, L., Hansen, C. L., Arnold, F. H. & Quake, S. R. Long-term monitoring of bacteria undergoing programmed population control in a microchemostat. Science 309, 137–140 (2005).
Badran, A. H. & Liu, D. R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015).
Polycarpo, C. et al. An aminoacyl-tRNA synthetase that specifically activates pyrrolysine. Proc. Natl Acad. Sci. USA 101, 12450–12454 (2004).
Bryson, D. I. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat. Chem. Biol. https://doi.org/10.1038/nchembio.2474 (2017).
Umehara, T. et al. N-acetyl lysyl-tRNA synthetases evolved by a CcdB-based selection possess N-acetyl lysine specificity in vitro and in vivo. FEBS Lett. 586, 729–733 (2012).
Chin, J. W. et al. Addition of p-azido-l-phenylalanine to the genetic code of Escherichia coli. J. Am. Chem. Soc. 124, 9026–9027 (2002).
Srinivasan, G., James, C. M. & Krzycki, J. A. Pyrrolysine encoded by UAG in Archaea: charging of a UAG-decoding specialized tRNA. Science 296, 1459–1462 (2002).
DeBenedictis, E. A., Carver, G. D., Chung, C. Z., Söll, D. & Badran, A. H. Multiplex suppression of four quadruplet codons via tRNA directed evolution. Nat. Commun. 12, 5706 (2021).
Magliery, T. J., Anderson, J. C. & Schultz, P. G. Expanding the genetic code: selection of efficient suppressors of four-base codons and identification of ‘shifty’ four-base codons with a library approach in Escherichia coli. J. Mol. Biol. 307, 755–769 (2001).
Anderson, J. C., Magliery, T. J. & Schultz, P. G. Exploring the limits of codon and anticodon size. Chem. Biol. 9, 237–244 (2002).
Wang, K., Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Evolved orthogonal ribosomes enhance the efficiency of synthetic genetic code expansion. Nat. Biotechnol. 25, 770–777 (2007).
DeBenedictis, E., Söll, D. & Esvelt, K. Measuring the tolerance of the genetic code to altered codon size. Preprint at bioRXiv https://doi.org/10.1101/2021.04.26.441066 (2021).
Nourmohammad, A. & Eksin, C. Optimal evolutionary control for artificial selection on molecular phenotypes. Phys. Rev. X 11, 011044 (2021).
Simutis, R. & Lübbert, A. Bioreactor control improves bioprocess performance. Biotechnol. J. 10, 1115–1130 (2015).
Hartl, R. F., Mehlmann, A. & Novak, A. Cycles of fear: periodic bloodsucking rates for vampires. J. Optim. Theory Appl. 75, 559–568 (1992).
Baym, M. et al. Spatiotemporal microbial evolution on antibiotic landscapes. Science 353, 1147–1151 (2016).
Wadman, M. United States rushes to fill void in viral sequencing. Science 371, 657–658 (2021).
Zhao, Y. & Yang, Y. Profiling metabolic states with genetically encoded fluorescent biosensors for NADH. Curr. Opin. Biotechnol. 31, 86–92 (2015).
Zhang, L. et al. Ratiometric fluorescent pH-sensitive polymers for high-throughput monitoring of extracellular pH. RSC Adv. 6, 46134–46142 (2016).
Zhujun, Z. & Seitz, W. R. A carbon dioxide sensor based on fluorescence. Anal. Chim. Acta 160, 305–309 (1984).
Cho, I., Jia, Z.-J. & Arnold, F. H. Site-selective enzymatic C‒H amidation for synthesis of diverse lactams. Science 364, 575–578 (2019).
Jiang, R. & Krzycki, J. A. PylSn and the homologous N-terminal domain of pyrrolysyl-tRNA synthetase bind the tRNA that is essential for the genetic encoding of pyrrolysine. J. Biol. Chem. 287, 32738–32746 (2012).
Suzuki, T. et al. Crystal structures reveal an elusive functional domain of pyrrolysyl-tRNA synthetase. Nat. Chem. Biol. 13, 1261–1266 (2017).
Cheetham, G. M. T., Jeruzalmi, D. & Steitz, T. A. Structural basis for initiation of transcription from an RNA polymerase–promoter complex. Nature 399, 80–83 (1999).
Delignette-Muller, M. L. et al. fitdistrplus: an R package for fitting distributions. J. Stat. Softw. 64, 1–34 (2015).
Dray, S. et al. The ade4 package: implementing the duality diagram for ecologists. J. Stat. Softw. 22, 1–20 (2007).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Yu, G. Using ggtree to visualize data on tree-like structures. Curr. Protoc. Bioinforma. 69, e96 (2020).
Noble, R. ggmuller: create muller plots of evolutionary dynamics (GitHub, 2019).
Acknowledgements
We acknowledge W. Consigli, W. Fu, A. Cuevas and others at Hamilton Robotics for their guidance and assistance. We thank K. Prather’s laboratory for equipment use and assistance. We thank E. Alley, S. Von Stetina and B. Thuronyi for their thoughtful comments on the paper. This work was supported by the MIT Media Laboratory, an Alfred P. Sloan Research Fellowship (to KME), gifts from the Open Philanthropy Project and the Reid Hoffman Foundation (to K.M.E.), and the National Institute of Digestive and Kidney Diseases (grant no. R00 DK102669-01 to KME). E.A.D. was supported by the National Institute for Allergy and Infectious Diseases (grant no. F31 AI145181-01). E.J.C. was supported by a Ruth L. Kirschstein NRSA fellowship from the National Cancer Institute (grant no. F32 CA247274-01).
Author information
Authors and Affiliations
Contributions
E.A.D. and K.M.E. conceived the study. E.A.D. and D.W.G. developed the platform with support from E.J.C. and advice from K.M.E. D.W.G. and S.G. developed the software with advice from E.A.D., E.J.C. and K.M.E. E.A.D., E.J.C. and D.W.G. designed the experiments with advice from K.M.E. E.A.D., E.J.C., D.W.G. and B.W. performed the experiments. E.J.C. analyzed and visualized data. E.J.C., E.A.D., B.W. and K.M.E. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
E.A.D. and K.M.E. have filed US Patent 16405380 on this work.
Peer review information
Nature Methods thanks Arjun Ravikumar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 PRANCE optimization.
(a) Robotic manipulations operate in a loop, which repeats every 30 minutes. (b) Culture source fluidics (media, turbidostat/static culture, waste) are peripherally separated from the robot. The maximum flow-through rate is determined by the frequency with which the robot exchanges liquid (operations per hour), as well as the fraction of the standing volume of the population that is exchanged during each operation (the bottlenecking fraction). There is a trade-off between the maximum flow rate and the extent of bottlenecking. (c) The number of populations that can be serviced assuming 2 robot operations per hour (ops/hr) impacts the experimenter-free/hands-off operation time of the robot. (d) Larger robot decks can fit more tip carriers, more tips, and therefore require less frequent servicing.
Extended Data Fig. 2 Relationship between real-time monitoring data and phage titer.
(a) Correlation between absorbance depression and luminescence for each evolving replicate. Kernel density estimates of the absorbance and luminescence data for the population are plotted on x and y axis, respectively. Luminescence data from Fig. 2b. (b) Comparison of real-time luminescence tracking (top) and corresponding phage titer as measured by traditional plaque assays (bottom). See Supplementary Table 1 for evolution construct details.
Extended Data Fig. 3 Reservoir diagrams.
Schematics of the 8-channel and 96-channel media reservoirs. These were printed on a Form 3 resin 3D printer. See the Supplementary Table 4 for .stl files for each.
Extended Data Fig. 4 Failure mode analysis.
Analysis of failure modes used to improve reliability and error handling.
Extended Data Fig. 5 Stochasticity of T7 RNAP evolution.
Validating T7 mutagenesis with cool PRANCE. MP-containing bacteria were provided with either 1) induction prior to cooling to 4 C, 2) given no inducer, or 3) induced on the robotic deck at 37 C and their luminescence was tracked for 30 hours to validate that mutagenesis behaved similarly to induction of cultures directly from a turbidostat (see Fig. 1d). (b) Real-time absorbance depression monitoring of 90 simultaneous directed evolution experiments with 6 no-phage controls, fit with a binomial regression of the total data. (c) Logistic regression of each luminescence trace during T7 RNAP evolution to bind the T3 promoter, used to calculate the average evolution times (Supplemental Methods). (d) Goodness-of-fit estimates of a logistic distribution of the total T7 evolution time data.
Extended Data Fig. 6 TAGA-qtRNA and AGGG-qtRNA PRANCE.
(a) Constructs for evolving TAGA-decoding qtRNAs. (b) Representative results for evolving TAGA-decoding qtRNAs. (c) Evolved qtRNAs exhibit increased ability to decode a TAGA quadruplet codon. Units are the percent luminescence when translating luxAB-357-TAGA in the presence of the qtRNA relative to expression of all-triplet-luxAB. (d) evolved genotypes E) Real-time absorbance and luminescence monitoring of qtRNA-encoding phage where either randomized or directed anticodons were used to evolve AGGG containing codons.
Extended Data Fig. 7 Edge effects.
Comparison between 96 replicates implemented in a densely packed 96-well plate (left) and 96 replicates split over two plates to reduce edge effects (right). Data plots the minimum time to phage detection via luminescence monitoring (below). Both plates above are normalized to their internal max value.
Extended Data Fig. 8 Tip contamination, sterilization and reuse.
To assess the maximum amount of possible cross-contamination, T7 RNAP-containing phage were inoculated into cultures containing pT7-psp-LuxAB bacteria in a grid-like pattern with 48 phage-containing wells and 48 no-phage-containing wells. PRANCE steps were performed with either (a) the 96-head channel or (b) the 8-channel pipettor. Use of the 96-well head gave less cross-contamination events, which we attributed to lower fly-over events. (c) To assess the impact of tip-sterilization and reuse in the optimized robotic method and configuration (Supplemental Video 1), robotic tips were submerged in either water or T7 RNAP-containing phage and then sterilized prior to being used to maintain high-throughput bacterial cultures containing pT7-psp-LuxAB22. Sterilized tips were also used to propagate bacteria inoculated with T7 RNAP phage as a positive control to ensure that bleach carryover did not affect phage propagation. No cross contamination was observed in the serialized tip condition over 12 hours, indicating that tips could be reused for a minimum of 12 hours without being replaced.
Extended Data Fig. 9 Kinetic Luminescence Activity Assays of −3, −5, and TP6 Promoter Variants.
To quantify the relative activities of each variant (independent of possible phage backbone mutations), phage from each replicate at time point t = 10 days were isolated and each T7 RNAP variant was cloned into an IPTG-inducible reporter construct. Subcloned variants were transformed into S2060 cells containing LuxAB driven by either the (a) −3 variant, (b) −5 variant, or (c) TP6 promoter and grown overnight to an OD > 1. Bacteria were then diluted 1:100 and grown to an OD of exactly 1.2 in DRM using a high-throughput robotic turbidostat method as described previously22. Once the bacteria reached an OD of 1.2 (approximately 2 hours), cells were induced with either 1 mM IPTG or [-] IPTG controls (n = 3 for each condition). Bacteria were autonomously maintained at an OD of 1.2 for the duration of the experiment and luminescence readings were taken once every 45 minutes for 10 hours. Fold change in luminescence (as shown in Fig. 6E), was calculated by averaging the luminescence in each turbidostat once the luminescence reached equilibrium (t > 8 hours) and then normalized to the average luminescence of the [-] IPTG controls within the same time window. WT T7 RNAP was used as a control for each respective promoter reporter construct (n = 6 for each WT control).
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1, legends for supplementary Tables 2–4.
Supplementary Video 1 (download MOV )
Tip sterilization robotic method.
Source data
Source Data Fig. 1 (download XLSX )
Annotated luminescence and absorbance data.
Source Data Fig. 2 (download XLSX )
Annotated luminescence and absorbance data, Inflection point analysis of evolution time, T7 RNAP sequence mutations.
Source Data Fig. 3 (download XLSX )
Annotated luminescence and absorbance data, PFU values for phage enrichment.
Source Data Fig. 4 (download XLSX )
Luminescence data as a percentage of WT, annotated luminescence and absorbance data.
Source Data Fig. 5 (download XLSX )
Luminescence data as a percentage of WT, next-generation sequencing data with annotation files.
Source Data Fig. 6 (download XLSX )
Annotated luminescence and absorbance data.
Source Data Extended Data Fig. 2 (download XLSX )
Annotated luminescence and absorbance data.
Source Data Extended Data Fig. 5 (download XLSX )
Annotated luminescence and absorbance data.
Source Data Extended Data Fig. 6 (download XLSX )
Annotated luminescence and absorbance data.
Source Data Extended Data Fig. 7 (download XLSX )
Annotated luminescence and absorbance data.
Source Data Extended Data Fig. 8 (download XLSX )
Annotated luminescence and absorbance data.
Source Data Extended Data Fig. 9 (download XLSX )
Annotated luminescence and absorbance data.
Rights and permissions
About this article
Cite this article
DeBenedictis, E.A., Chory, E.J., Gretton, D.W. et al. Systematic molecular evolution enables robust biomolecule discovery. Nat Methods 19, 55–64 (2022). https://doi.org/10.1038/s41592-021-01348-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-021-01348-4
This article is cited by
-
An industrial automated laboratory for programmable protein evolution
Nature Chemical Engineering (2025)
-
EvoAI enables extreme compression and reconstruction of the protein sequence space
Nature Methods (2025)
-
Continuous directed evolution of a compact CjCas9 variant with broad PAM compatibility
Nature Chemical Biology (2024)
-
Engineering is evolution: a perspective on design processes to engineer biology
Nature Communications (2024)
-
Printed polymer platform empowering machine-assisted chemical synthesis in stacked droplets
Nature Communications (2024)