Abstract
The specificity of T cells is determined by T cell receptor (TCR) α and β chain sequences. While bulk TCR sequencing enables cost-effective repertoire profiling without chain pairing information, single-cell approaches provide paired data but are costly and limited in throughput. Here we present throughput-intensive rapid TCR library sequencing (TIRTL-seq), an experimental and computational methodology for paired TCR repertoire sequencing (TCR-seq). TIRTL-seq is based on the parallel generation of hundreds of TCR libraries in 384-well plates at less than US$200 per plate, allowing cohort-scale paired TCR-seq studies. We benchmarked TIRTL-seq against state-of-the-art bulk TCR-seq and 10x Genomics Chromium technologies on longitudinal samples and identified severe acute respiratory syndrome coronavirus 2- and Epstein–Barr virus-specific clonal expansions after infection with distinct dynamics. TIRTL-seq offers a universal protocol scalable from a single cell to millions of T cells per sample, simultaneously delivering both precise clonal frequency estimation and accurate TCR chain pairing, combining the strengths of bulk and single-cell TCR-seq.
Similar content being viewed by others
Main
In the adaptive immune system, αβ T cells are key players, recognizing antigens presented on major histocompatibility complex (MHC) molecules using T cell receptors (TCRs). The TCRα and TCRβ chains are formed in a stochastic V(D)J recombination process, creating a broad αβTCR repertoire. T cells recognizing a cognate peptide–MHC complex expand several thousand-fold during the immune response, but the peak frequency of individual clones in most cases is low (~1 in 10,000 (refs. 1,2)), with a combined frequency of all clones recognizing an immunodominant epitope of around 1%3. The great diversity of the TCR repertoire and low frequency of T cells of interest make high-throughput sequencing of TCR genes an essential tool to study T cell responses.
Bulk TCR repertoire sequencing (TCR-seq), introduced over 15 years ago4,5, can generate datasets with hundreds of thousands of unique T cell clones corresponding to millions of T cells. The major limitation of this method is that it does not provide paired αβTCR sequences. Most publicly available TCR-seq data comprise single-chain sequencing of TCRβ only, despite both TCRα and TCRβ making contact with the peptide–MHC complex and determining TCR specificity6,7. Although TCRs recognizing certain epitopes might have distinctive features in the TCRα or TCRβ chain8,9,10,11, both chains are needed to confirm specificity in vitro.
Concurrently with bulk sequencing, single-cell TCR-seq technologies were developed12,13,14 on the basis of single-cell sorting and multiplex amplification of TCR genes in individual wells of PCR plates. Although these methods provide chain pairings, they are limited in throughput to hundreds of cells and are time-consuming and expensive because of high reagent usage per well.
The development of droplet-based single-cell RNA sequencing (scRNA-seq) technologies increased the throughput and lowered per-cell costs for single-cell sequencing15,16, with the latest commercially available GEM-X kits from 10x Genomics17 capable of processing up to 20,000 cells in one reaction. Concurrent reaction miniaturization and transition to a 384-well-plate format have reduced the costs of plate-based scRNA-seq18,19. Combinatorial barcoding methods20 have further increased the throughput of scRNA-seq up to millions of cells. However, high reagent costs and tedious protocols limit the application of scRNA-seq to small numbers of samples.
To improve the throughput of paired αβTCR sequencing, a combinatorial approach (pairSEQ)21 has been suggested on the basis of the even distribution of T cells in a 96-well plate, bulk TCRα and TCRβ sequencing of each well, and reconstruction of αβ chain pairings by matching TCRα and TCRβ well-occurrence patterns. Alternative data analysis approaches for such pairSEQ experiments have also been suggested22,23. Despite the uniquely high throughput and yield (~36 million peripheral blood mononuclear cells (PBMCs) yielding ~212,000 unique αβTCRs), the protocol was not widely adopted, potentially owing to high reagent costs and computationally intensive algorithms for data analysis. Importantly, pairSEQ is also unable to pair high-frequency clonotypes, sampled in most or all wells, and low-frequency clonotypes, sampled in only a few wells.
An optimal TCR-seq technology should be affordable, able to process millions of T cells (deep), provide complete αβTCR sequences allowing for cloning and further screening (paired), and estimate frequencies of individual clones to identify clonal expansion (quantitative). Here, we present TIRTL-seq, an experimental and computational methodology satisfying these requirements.
Results
TIRTL-seq protocol development and optimization
Combinatorial TCR chain pairing relies on matching TCRα and TCRβ occurrence patterns across multiple replicates of T cell samples21,22,23. This method requires high sensitivity to detect transcripts from a single cell within a TCR clone. Furthermore, it must be robust enough to load thousands of T cells per replicate and process hundreds of replicates in a cost-effective way.
To achieve these goals, we developed an optimized protocol, TIRTL-seq, to generate bulk TCRα and TCRβ libraries from PBMCs or T cells in a 384-well-plate format (Fig. 1a). TIRTL-seq utilizes noncontact liquid dispensing with hydrophobic overlays for reaction miniaturization and simultaneous cell lysis and reverse transcription (RT) for complementary DNA synthesis to avoid multiple clean-up steps18. Targeted amplification of TCRα and TCRβ cDNA is carried out in a multiplex reaction using a set of V-segment and C-segment primers from ref. 21, which were further optimized to exclude pseudogenes and incorporate parts of Illumina Nextera adapter sequences. The C-segment reverse primers contain plate-specific barcodes to enable pooling and sequencing of multiple plates in a single sequencing run (Supplementary Data 1). In the second (indexing) PCR step, we introduce 384-well-specific unique dual indices (UDI) and full-length Illumina sequencing adapters. Final libraries are pooled, purified and size-selected using magnetic beads before sequencing on an Illumina platform. The entire protocol takes about 7 h, with less than 4 h hands-on time from cells to ready-to-sequence libraries and costs of about US$185 per 384-well plate for reagents and consumables. The default protocol requires a noncontact liquid dispenser and liquid handler to perform 384-to-384 plate transfer and could be implemented on a variety of automation platforms. We also developed a manual version of the protocol using larger volumes in a 96-well PCR plate, which requires no automation and costs about US$130 per plate (Methods).
a, Schematic of TIRTL-seq protocol. In brief, a cell suspension is distributed into 384-well plates containing an RT–lysis master mix under a hydrophobic overlay using noncontact liquid dispensers. After the RT reaction, PCR I master mix with V-segment and C-segment primers is dispensed into the same plate. The PCR I product is then diluted and transferred to the PCR II plate for indexing PCR with well-specific unique dual indices. The PCR II products are pooled by centrifugation, purified, size-selected using magnetic beads and sequenced on an Illumina platform. Total library preparation cost is listed for one 384-well plate. b,c, The sensitivity of single sorted T cells according to CellsDirect (b) and TIRTL-seq (Maxima H-based) (c). Green, both TCRα and TCRβ identified; orange, TCRα lost; yellow, TCRβ lost; blank, no cell present. Column 24 is a negative control (no cells sorted). d, Relative fraction of cells with both TCRα and TCRβ identified (green), lost TCRα chain (orange) and lost TCRβ chain (yellow) shown for Invitrogen CellsDirect RT protocol (left) and TIRTL-seq protocol (right). e, TIRTL-seq shows robustness to an increasing number of PBMCs. Number of unique clonotypes detected from each well (y axis) plotted for different numbers of cells per well (x axis) for the TIRTL-seq protocol (n = 32 wells per group), each box representing the interquartile range (IQR), the line inside the box indicating the median, whiskers showing ±1.5× IQR and overlaying points representing individual values. Panel a created with BioRender.com.
We tested the sensitivity of TIRTL-seq for detecting TCRα and TCRβ transcripts using single-sorted live CD3+ cells from healthy donor PBMCs. As efficient simultaneous cell lysis and RT is a crucial step in the protocol, we compared the commercially available Superscript IV-based CellsDirect kit with our Triton-X100/Maxima H-based cell lysis/RT reaction with the same amplification conditions for PCR I and PCR II. Using the TIRTL-seq RT protocol, we detected both TCRα and TCRβ transcripts in over 98% of wells containing single T cells, outperforming the CellsDirect kit at a fraction of the price (Fig. 1b–d). Next, we loaded 12,500–100,000 PBMCs per well of a 384-well plate to test the robustness of TIRTL-seq to increasing cell number. We found that the number of unique clonotypes identified increased monotonically with increasing numbers of cells for TCRβ (Fig. 1e) but plateaued at 25,000 cells per well for TCRα, potentially owing to a lower number of transcripts per cell24,25 (Fig. 1e).
Overall, the TIRTL-seq protocol proved robust and sensitive across a range of cell numbers up to 25,000 PBMCs per well for TCRα and TCRβ, corresponding to approximately 10 million PBMCs per 384-well plate, which is well within the expected counts from a standard blood draw of 1–10 ml.
TIRTL-seq identifies large numbers of αβTCR pairs
To predict αβTCR chain pairings, we applied the MAD-HYPE algorithm developed by the authors of ref. 23 based on the principle of combinatorial pairing by mapping the presence and absence patterns (originally proposed by ref. 21). In brief, for each possible TCRα/TCRβ pair, the algorithm calculates the number of wells in a plate where chains are detected simultaneously and the number of wells where each chain is found separately, and it then accepts or rejects the pair using a Bayesian statistical model. We reimplemented the algorithm in R and Python 3 to run on NVIDIA or Apple Silicon graphics processing units (GPUs) (Methods). Benchmarking our implementation of the MAD-HYPE algorithm achieved a runtime of 93 s for data from experiment 2 in ref. 21 when run on a single NVIDIA GPU A100 80 GB, in comparison with 24 h reported for the same dataset in ref. 23 on a 96-core central processing unit (CPU) cluster一an almost 1,000× increase in computation speed. Using this pairing approach, we identified 169,423 unique αβTCR pairs from TIRTL-seq data generated in a 384-well plate loaded with 10 million PBMCs. Using the manual 96-well protocol, we identified 88,142 paired TCRs from 10 million PBMCs per plate, suggesting loss of resolution from using fewer wells.
We then investigated how increasing the number of 384-well plates, each loaded with 10 million PBMCs from the same donor, would increase the number of identified αβTCR pairings. As we added plates, we tried three strategies to analyze the data: (1) calling pairs independently for each 384-well plate and combining results (‘more plates’), (2) combining wells from multiple plates into a single 384-well plate before analysis (‘more cells per well’) and (3) analyzing all wells collectively (‘plate with more wells’) (Fig. 2a). We found the last strategy to be the most effective: increasing the number of wells analyzed together improved the resolution of matching αβ chain patterns, resulting in higher statistical power. We identified 989,241 paired αβTCR clonotypes corresponding to more than 30 million T cells from six 384-well plates each loaded with 10 million PBMCs (60 million PBMCs in total). However, decreasing the number of wells per sample would accommodate T cells from multiple donors in the same plate to further reduce the cost for cohort studies. We tested this scenario by downsampling to 96 wells, which decreased the number of identified αβTCRs to 25,000 from approximately 2.5 million PBMCs (Fig. 2a).
a, TIRTL-seq scaling with different analysis methods. The total number of unique paired αβTCRs recovered (y axis) is plotted using three distinct analytical approaches: increasing the number of individual plates analyzed separately and concatenating the results (red), increasing the number of cells per well analyzed (blue) or increasing the number of total wells analyzed jointly (green). The dotted orange line shows the number of unique αβTCRs from a single 10x Genomics scTCR-seq reaction. b, Clone-size distribution of paired and unpaired (dark gray) clones by TIRTL-seq. Paired αβTCRs from a 10x Genomics scTCR-seq experiment are ordered by the number of cells with a clonotype (y axis, log scale), while the x axis shows the clone rank. Each color indicates overlap between 10x Genomics scTCR-seq and αβTCRs called by MAD-HYPE (orange), T-SHELL (green) or both (blue). Dark gray indicates lack of pairing in TIRTL-seq compared with 10X Genomics scTCR-seq. c, T-SHELL algorithm pairs TCR chain through frequency correlation. Top: correlation of relative per-well frequencies of largest TCRα no. 1 and top 5 largest TCRβs in the repertoire; red line shows linear fit. Bottom: Manhattan plot for unadjusted P values (two-sided Pearson correlation t-test) for pairing of TCRα no. 1 to 10,000 most abundant TCRβs. Dotted line shows P value cutoff after Bonferroni multiple testing adjustment. d, T cell expansion increases pairing efficiency. T cell clone frequency (y-axis) is plotted against rank (x-axis). The dotted line shows the minimal three-well occurrence threshold for αβ chain pairing by TIRTL-seq. An increased number of clones clearing the threshold after expansion (orange curve) results in more called pairs compared with before expansion (green curve). The inset shows clonal frequency distortion after antigen-independent T cell expansion. Clonal frequency before expansion (x axis) is plotted against clonal frequency after expansion (y axis). e, The fraction of TCRβs overlapping between 10X Genomics scTCR-seq (filtered or unfiltered for clones with >1 cell) and TIRTL-seq experiments (x-axis) with matching or mismatching TCRα for MAIT and non-MAIT clones. f, The fraction of clonotypes with a given chain (α or β) paired with one or more partner chains (TIRTL-seq data from one 384-well-plate experiment).
Extending αβTCR pairing to large and small clonal frequencies
To validate TCR chain pairings inferred by TIRTL-seq, we compared results with those from state-of-the-art single-cell TCR-seq (scTCR-seq) using the 10x Genomics 5′ GEM-X protocol. We generated 10x Genomics scTCR-seq data for 20,000 T cells from the same donor, resulting in 11,113 cells with paired αβTCR sequences, corresponding to 8,705 unique αβTCRs. We compared these pairs with those identified by MAD-HYPE analyses of TIRTL-seq data from one 384-well plate to measure concordance between methods. We found that, although 2,706 αβTCRs matched between technologies, no pairs for the abundant T cell clones present at more than five cells in the 10x Genomics experiment matched (Fig. 2b, orange and blue dots). Upon further investigation, we found that all these abundant clones were sampled in all or almost all wells of the 384-well plate, making them impossible to pair using the MAD-HYPE algorithm, as noted in the original MAD-HYPE algorithm and pairSEQ protocol21,23. In total, the top 74 pairs (more than 5 cells) identified by 10X Genomics, representing 1,944 cells out of the total 11,113, failed to be paired by MAD-HYPE.
To address this problem, we developed a new heuristic pairing algorithm called TCRαβ sequence highly efficient linkage learning (T-SHELL). The T-SHELL algorithm relies on the fact that clones present in all wells still have variability in the number of cells across wells. Thus, the relative frequency of both TCRα and TCRβ transcripts within a well should increase if more cells of a given clone are sampled in a given well. T-SHELL uses correlation between TCRα and TCRβ clonotype relative frequencies (measured as read fraction) across wells instead of presence and absence patterns for pairing. To demonstrate the principle of T-SHELL, we plotted the correlation between the largest TCRα clonotype (by average frequency across wells) and the top 5 largest TCRβ clonotypes from the 384-well experiment, as shown in Fig. 2a. The read fraction of TCRα no. 1 strongly correlated (Pearson r = 0.6) with TCRβ no. 4 (Fig. 2c, top), which remained the only TCRβ chain that had a significant correlation (P = 10−38) with TCRα no. 1 when we correlated the read fractions of TCRα no. 1 against the top 10,000 largest TCRβs (Fig. 2c, bottom). Corresponding 10x Genomics scTCR-seq data confirmed that TCRβ no. 4 is indeed the correct pair for TCRα no. 1, indicating that T-SHELL can accurately pair large clones. We subsequently applied T-SHELL to pair other TCRαβ chains found across the 384-well plate and found that the majority of abundant clones (73/74 with more than five cells per clone on Fig. 2b) were paired and matched the 10x Genomics scTCR-seq pairings (Fig. 2b, blue and green color). Out of 2,159 TCRαβ pairs identified by T-SHELL that matched the 10x Genomics scTCR-seq pairs with one to five cells per clone, 94% (2031/2159) were also identified by MAD-HYPE, suggesting good agreement between the two algorithms (Fig. 2b, blue). Thus, we developed a clonotype frequency-based algorithm capable of pairing TCR chains for abundant clones present in all wells. To validate T-SHELL in the 96-well format and calibrate P value thresholds across different clone sizes, we performed a series of experiments spiking in up to 10% of cells with a Jurkat T cell line with a known αβ pair (Extended Data Fig. 1). We found that T-SHELL correctly and unambiguously paired large spike-in clones, although tuning the adjusted P value threshold is necessary for 96-well experiments, as described in our 96-well protocol.
For small clones, which account for the majority of the peripheral blood TCR repertoire, TIRTL-seq pairing relies on occurrence-pattern matching. Consequently, a clone has to be present in at least 3/384 wells, and thus have at least three cells per clone in a sample to be paired by TIRTL-seq. To address this limitation and to increase the number of cells per clone, we cultured T cells in vitro for 1 week, which substantially expanded clones, and consequently increased the number of TCRαβ chain pairings by an order of magnitude from approximately 17,681 (unexpanded sample with the same initial cell count) to 313,106 pairs. However, T cell expansion distorted the clonal frequency hierarchy (Fig. 2d): None of the top 5 largest clonotypes before expansion were detected within the top 10 after expansion, and more than 10,000 clones increased or decreased in clonal frequency more than ten fold.
We then used both MAD-HYPE and T-SHELL algorithms to call pairs and compared TCRαβ pairings identified by either method with 10x Genomics scTCR-seq. For TCRβ clonotypes overlapping between 10x Genomics scTCR-seq and TIRTL-seq, 91.5% (2,886/3,155) for a single 384-well plate and 91.4% (3,858/4,221) for the six 384-well plate experiments had the same TCRα. Interestingly, many mismatches between TIRTL-seq and 10x Genomics scTCR-seq (15% of mismatches for the 6 × 384-plate experiment) corresponded to mucosal-associated invariant T (MAIT) cells, a specialized T cell subset featuring an invariant TCRα chain paired with various TCRβs, breaking the TCRαβ chain pattern matching in our algorithms26. In general, most mismatching αβTCRs were found in clones with just one cell in the 10x Genomics scTCR-seq data. The proportion of mismatches dramatically reduced from 8.5% to less than 2% (4/8 mismatches were MAIT) when we selected clones with more than one cell per clone in the 10x Genomics scTCR-seq dataset (Fig. 2e). To further compare the pairing precision of 10x Genomics and TIRTL-seq, we generated an additional dataset of individually sorted and sequenced T cells from the same donor. We obtained 568 individual cells with paired αβ TCRs corresponding to 495 clones. For TCRβ clonotypes overlapping between this benchmarking single-cell TCR-seq dataset and the 10x Genomics scTCR-seq data, 70/73 (96%) had the same TCRα. For TIRTL-seq, we found the same TCRα for 191/197 overlapping clonotypes (97%). We note that our metric of pairing precision has limitations, particularly that some mismatches might not be errors but rather distinct low-frequency clonotypes sharing the exact same TCRβ sampled at different time points. It has been previously observed that up to 30% of αβT cells express two TCRα chains27,28. We identified 27.5% of TCRβ with two TCRα partners (Fig. 2f), suggesting that TIRTL-seq reliably captures T cells expressing two functional TCRα chains. Thus, TIRTL-seq is as accurate for paired αβTCR sequencing as 10x Genomics scTCR-seq at a fraction of the price.
Longitudinal TCR repertoire profiling with TIRTL-seq
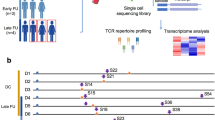
Longitudinal TCR-seq allows identification of TCR clonal expansion and contraction after an immune challenge in an antigen-agnostic manner. To test the performance of TIRTL-seq for longitudinal TCR repertoire profiling, we used three longitudinal samples collected from an individual infected with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) at −143 days (‘baseline’), 6 days (‘acute’) and 29 days (‘convalescent’) after a positive PCR test (Fig. 3a). On each PBMC sample, we performed immunomagnetic positive isolation of CD4+ and CD8+ T cells, which were distributed into two halves of a 384-well plate.
a, Longitudinal sampling of a donor with SARS-CoV-2 infection. scGEX, single-cell gene expression. b, Top: TIRTL-seq identifies expansions and contractions in the CD8+ T cell repertoire. Bottom: CD4+ T cell clonal frequencies from the same time points. The dashed diagonal indicates the line of equality. c, Colored dots show clonotypes matching known TCRs specific for A*02 YLQ (cyan) and B*07 SPR SARS-CoV-2 epitopes (green) and A*02 GLC (orange) and B*07 RAK EBV epitopes (red) on pairwise time point comparisons. The dashed diagonal indicates the line of equality. d, Cumulative frequency of CD8+ clones specific to A*02 YLQ (cyan) and B*07 SPR SARS-CoV-2 epitopes (green) and A*02 GLC (orange) and B*07 RAK EBV epitopes (red) across time points. conv, convalescent. Panel a created with BioRender.com.
To call expanded and contracted clonotypes from TIRTL-seq data, we calculated mean frequency and standard error of the mean (s.e.m.) for each TCRβ chain over wells. We call clones significantly expanded or contracted between time points if there is a log2 fold change (log2FC) >3 between average frequencies and the difference between average frequencies exceeds five s.e.m. intervals. We identified 980 CD8+ (Fig. 3b, top left) and 213 CD4+ clones (Fig. 3b, bottom left) that expanded when comparing acute and baseline time points. Conversely, 312 CD8+ (Fig. 3b, top right) and 134 CD4+ clones (Fig. 3b, bottom right) contracted when comparing acute and convalescent time points.
Paired αβTCR sequencing data generated by TIRTL-seq allowed us to identify TCRs with known specificities in our dataset by searching for highly similar TCRs with matching HLA-restriction in VDJdb29. We used TCRdist8 as a pairwise distance metric and found that many clones that expand from baseline to acute time points have highly similar sequences to known TCRs for HLA-B*07 SPRWYFYYL and HLA-A*02 YLQPRTFLL epitopes from SARS-CoV-2. As expected for SARS-CoV-2-specific T cell responses, these clones contracted from the acute to convalescent time points, indicating death of effector T cells after viral clearance. Interestingly, we also identified 91 expanding clonotypes matching to Epstein–Barr virus (EBV) epitopes (HLA-B*08 RAKFKQLL and HLA-A*02 GLCTLVAML), including the largest clone in the repertoire at acute and convalescent, but not baseline, time points (Fig. 3c). We cloned two representative TCRs per epitope into NFAT-GFP reporter Jurkat cell lines and confirmed their predicted peptide specificity in vitro (Extended Data Fig. 2). In stark contrast to SARS-CoV-2, EBV-specific clones, once established, remained stable from acute to convalescent time points, corresponding to novel clones at the very top of the TCR repertoire (Fig. 3d). We interpret these data to reflect two distinct infections (SARS-CoV-2 and EBV) occurring in this individual during the sample window, as almost none of the epitope-specific responses were detectable at baseline. We measured anti-EBV viral capsid antigen and anti-SARS-CoV-2 IgG in serum collected alongside the PBMC preparations and confirmed that this donor was seronegative at baseline and seropositive at acute and convalescent time points for both viruses, confirming the longitudinal TIRTL-seq results (Extended Data Fig. 3). These results indicate differential clonal dynamics between acute and chronic viral infections that might explain clonal dominance and the emergence of large stable clones over the lifespan.
Benchmarking TIRTL-seq against state-of-the-art single-cell and bulk TCR-seq approaches
We showed that, given enough cells, TIRTL-seq yields over 100 times more paired clonotypes (989,241 versus 8,705; Fig. 2a) at equal price to one Chromium GEM-X V(D)J assay reaction with αβ pairings matching between the approaches (Fig. 2e). However, in a more realistic scenario with cryopreserved PBMC samples from a standard blood draw, such large cell numbers are rarely available. To compare TIRTL-seq with state-of-the art approaches in real-world scenarios, we performed both 10x Genomics scTCR-seq (20,000 T cells per time point) and bulk 5′ rapid amplification of cloned/cDNA ends (RACE) TCRα and TCRβ sequencing on samples from the same individual infected with SARS-CoV-2 at the same time points as above (Fig. 3a) and compared the results with the longitudinal TIRTL-seq dataset described in the previous section.
We first compared the number of paired clones identified by 10x Genomics scTCR-seq and TIRTL-seq in each sample and assessed whether the clones were repeatedly detected across time points (Fig. 4a). Not only did TIRTL-seq identify approximately four times more paired TCRs at each time point (CD4+/CD8+ combined), but these TCRs were also more reproducible across samples. The vast majority (>95%) of paired TCRs identified by 10x Genomics scTCR-seq were found in only one cell, and >90% of paired clonotypes at any given time point were not detected in 10x Genomics scTCR-seq from other timepoints. TIRTL-seq requires T cell clones to be independently found in at least three wells to be paired, leading to preferential pairing of clones that are more likely to be detected again.
a, Number of paired TCR clonotypes identified in each sample by 10x Genomics on CD3+ cells and TIRTL-seq (split into CD8+ and CD4+ subsets). Color shows clonotypes identified only in one time point (orange), two time points (yellow) and three time points (green) by each method. b, Number of expanded and contracted clonotypes between time points identified with 10x Genomics scTCR-seq, bulk 5′RACE TCRβ-seq and TIRTL-seq between baseline/acute and acute/convalescent time point pairs. c, The number of expanded (from baseline to acute) clonotypes from an independent bulk TCRβ sequencing experiment paired using 10x Genomics scTCR-seq (blue) and TIRTL-seq (green). d, The overlap between expanded clonotypes between baseline and acute time point identified with bulk TCRβ sequencing (yellow) experiments or TIRTL-seq (green, CD4+ and CD8+ combined).
To assess quantitative accuracy, we compared TIRTL-seq clonal frequencies with those from 10x Genomics. Both the absolute frequencies of abundant clones (five or more cells in 10x) and their log2FC between time points correlated strongly between the methods (Extended Data Fig. 4a,b), supporting TIRTL-seq’s utility for tracking clonal dynamics quantitatively. We then compared each approach’s ability to identify expanded and contracted T cell clones between time points. To identify contracted and expanded clonotypes in 10x Genomics scTCR-seq datasets, we assumed the Poisson distribution for the number of cells with a given TCRβ and used the same significance and log2FC thresholds as for TIRTL-seq in Fig. 3b. While a small number (34) of strongly expanding clones between baseline and acute infection were captured by 10x Genomics scTCR-seq (Extended Data Fig. 5, left), the contraction wave between acute and convalescent was largely undetectable (Extended Data Fig. 5, right). This is primarily due to the majority of the participating clones having a peak frequency of 1/1,000, resulting in only a few cells in the 10x Genomics scTCR-seq reaction. By contrast, TIRTL-seq identified 1,193 expanding clones between the baseline and acute time points and 446 contracting clones between acute and convalescent time points, similar to the magnitude observed in 5′RACE bulk TCR-seq (Fig. 4b).
Bulk TCR sequencing is a state-of-the-art method for deep TCR-seq and is frequently used in combination with 10x Genomics scTCR-seq to simultaneously identify temporal changes and obtain paired TCR clonotypes30,31,32. Thus, we searched for the 1,511 TCRβ clonotypes expanding between baseline and acute time points identified by 5′RACE in the 10x Genomics scTCR-seq and TIRTL-seq results to find TCRα partner chains. We found 1,247 TCRα pairs for expanded TCRβs in TIRTL-seq, but only 521 pairs in the 10x Genomics scTCR-seq (Fig. 4c). The overlap between expanded clonotypes identified by TIRTL-seq and bulk 5′RACE TCRβ-seq was 947/1,193 (~80%) (Fig. 4d). Importantly, in TIRTL-seq, the threshold to call an expanded clonotype between time points is higher than the minimal pairing threshold, leading to the vast majority of expanding clones also being paired: 887/980 (91%) of CD8+ and 195/213 (92%) of CD4+ TCRβs. Overall we show that TIRTL-seq is efficient both in calling expansions and contractions and in identifying correct TCR partner chain pairings.
Discussion
Existing approaches for paired TCR sequencing are limited in throughput, expensive and difficult to implement. Integration of the combinatorial TCR chain pairing strategy suggested by Howie et al. with miniaturization techniques used in the Smart-seq3xpress method18 allowed us to transition from a 96- to a 384-well-plate format and decrease the price per well over 50 times from US$2,500 per 96-well pairSEQ experiment to US$185 per 384-well TIRTL-seq experiment. In addition, we reimplemented the MAD-HYPE framework23 for GPUs, making αβTCR pairing inference from experiments with thousands of wells feasible on consumer grade hardware. Furthermore, we developed a new algorithm (T-SHELL) on the basis of the correlation between the relative abundances of TCR αβ transcripts across wells to overcome the limitations of combinatorial TCR pairing for highly expanded clones sampled in all wells21,22,23. The result is a method that can process 10 million PBMCs for one tenth the cost of a standard 10x Genomics experiment, identifying >160,000 unique paired αβ TCRs from a single 384-well plate, albeit without accompanying gene expression data. Our method could be scaled even further by integrating the data across multiple 384-well plates per sample, as we show for the 60 million PBMC experiment, while the highest-throughput commercially available kit for paired TCR sequencing (ParseBio Evercode TCR Mega) is limited to 1 million cells.
Longitudinal tracking of TCR clones can provide biological insights into how the immune system reacts to an immune challenge. For example, here we show different trajectories for SARS-CoV-2- and EBV-reactive TCRs in patient samples before and after confirmed SARS-CoV-2 infection. EBV-reactive clones that were not observed at the baseline time point became the most expanded clones in the repertoire and remained stable at the convalescent time point. While the presence of herpesvirus-specific TCRs among the most abundant clones is consistent with previous reports33,34, here we serendipitously observed how this hierarchy is established in an adult. Further studies into mechanisms that allow certain clones to expand and persist over time will be useful for rational design of vaccine and T cell therapies. For functional validation of antigen-specific clones identified by TIRTL-seq, we used de novo gene synthesis of αβ TCRs followed by expression in a reporter cell line. Direct cloning from TIRTL-seq libraries is currently limited because the short amplicons include only the CDR3 and flanking FR3 regions, omitting most of the V segment. Although using a V-multiplex primer set targeting the FR1 region35 offers a potential solution, optimizing the protocol for efficient full-length TCR amplification may be necessary.
The clonal frequency distribution within TCR repertoires follows a power law. The largest T cell clone in the repertoire typically has a frequency ranging from 1% to 10%, whereas the frequency drops to about 0.1% for the tenth largest clone and about 0.01% for the hundredth largest clone (equal to 2 of 20,000 cells loaded into the state-of-the-art 10x Genomics GEM-X reaction). Such low counts make clonal frequency estimation and longitudinal tracking challenging with scTCR-seq. TIRTL-seq, however, allows us to not only determine TCR chain pairings, but also to quantitatively track the frequencies of those clones over time using each well within the plate as a biological replicate. Averaging of relative clonal frequencies across all wells results in a good clone size estimation. However, the V-region multiplex RT–PCR strategy without unique molecular identifiers (UMIs) used in TIRTL-seq is susceptible to amplification bias, so incorporating computational adjustments for primer amplification bias could improve the precision of clonal tracking36.
Our methodology permits chain pairing even for the largest TCR clones in a sample; however, an important limitation is its inability to pair TCRs found in only a few cells. For a 384-well-plate experiment, three cells per sample is the lowest possible frequency for TCR chain pairing. If pairings of small clones are of interest for a particular application, TIRTL-seq can be combined with single-cell sorting to produce paired TCRs for >98% of sorted cells, irrespective of clone size. However, in this scenario, the number of analyzed cells is limited by the number of wells in a plate. If each well contains a single cell, additional primers targeting transcripts of interest could be added to PCR I to determine cellular phenotype or genotype in addition to αβTCR sequence, similar to ref. 13. If sample size is limited to a few thousand cells, conventional scTCR-seq approaches such as 10x Genomics Chromium will probably be more efficient than TIRTL-seq. Alternatively, we show that T cell expansion before TIRTL-seq increases the number of cells per clone and thus pairing probability; this approach, however, disrupts clonal frequency estimates. Low frequency clones are still present in TIRTL-seq results as single TCRα and TCRβ chains in large numbers: for 160,000 paired clonotypes in a 384-well TIRTL-seq experiment from 10 million PBMCs we find approximately 5 million and 6 million unpaired TCRα and TCRβ, respectively. Unpaired TCR chains could be used to further refine TCR motifs identified in paired results or as input for any algorithms developed for single-chain bulk TCR-seq.
Several methods have been developed to determine TCR specificity from αβTCR sequence in vitro37,38,39,40,41,42,43, indicating that all information about TCR specificity is encoded in the paired TCR sequence and suggesting that this problem will eventually be solvable in silico44. We anticipate that affordable and high-throughput paired TCR sequencing techniques will be essential for diagnostics, vaccine and personalized therapy development.
Methods
Primary human cells
Healthy donor PBMCs were isolated from apheresis rings obtained from the St. Jude Blood Donor Center under Department of Pathology protocol BDC035. All apheresis rings were deidentified before release. Whole blood was collected from the apheresis ring and adjusted to a total volume of 30 ml with Dulbecco’s phosphate-buffered saline (DPBS; Gibco). PBMCs were separated by density gradient centrifugation using Lymphocyte Separation Medium (MP Biomedicals; 10 ml Lymphocyte Separation Medium to 30 ml whole blood, 600g at room temperature for 30 min, no centrifuge brake). The PBMC layer was then collected, washed with DPBS and subjected to red blood cell lysis using ACK lysis buffer to remove residual red blood cells. PBMCs were either used immediately for experiments or cryopreserved in Recovery Cell Culture Freezing Medium (Thermo Scientific).
SJTRC (NCT04362995) is a prospective, longitudinal cohort study involving adult employees (18 years and older) at St. Jude Children’s Research Hospital. The study received approval from the St. Jude Institutional Review Board. All participants provided written informed consent before enrollment and regularly completed questionnaires regarding demographics, medical history, treatment and symptoms if they tested positive for SARS-CoV-2 by PCR. Participants underwent weekly PCR screening for SARS-CoV-2 infection while on the St. Jude campus. Blood samples were collected in 8-ml cell preparation tubes, processed within 24 h into cellular and plasma components, aliquoted and then frozen for future analysis. For our analysis, we selected samples from a healthy 33-year-old female donor with naturally acquired mild SARS-CoV-2 infection and no prior history of SARS-CoV-2 infection or vaccination. Samples were collected 143 days before this donor’s first positive SARS-CoV-2 PCR test (‘baseline’ sample), 6 days after (‘acute’ sample) and 29 days after (‘convalescent’ sample).
Single-cell sorting
Frozen PBMCs were thawed in a water bath and resuspended in 5 ml of prewarmed complete Roswell Park Memorial Institute medium (RPMI 1640 (Gibco) supplemented with 10% fetal bovine serum (FBS) (Gibco), 2 mM L-glutamine (Gibco) and 100 U ml−1 penicillin–streptomycin (Gibco)). After centrifugation (500g, 5 min, room temperature), the supernatant was discarded and the cell pellet resuspended in 4 ml DPBS containing 0.1% bovine serum albumin (BSA) and 2 mM EDTA. A total of 10 µl aliquot of the cell suspension was used for cell counting with acridine orange and propidium iodide (AO/PI) staining reagent on CellDrop FL (Denovix). Following another centrifugation (500g, 5 min), the supernatant was discarded and the cell pellet was resuspended in 50 µl fluorescence-activated cell sorting buffer (0.5% BSA, 2 mM EDTA in DPBS) with 1 µl human TruStain Fc-block (1:50, Biolegend) and incubated for 15 min at 4 °C. The cells were then stained with 50 µl of a surface antibody cocktail containing Ghost Violet 510 viability dye (1:100, Tonbo Biosciences) and anti-human CD3-FITC (1:100, Biolegend, clone SK7) for 30 min at 4 °C. After washing with DPBS, the cells were filtered through a 50-µm filter into sorting tubes with DPBS. Live, single, CD3+ cells were sorted (one cell per well) using Sony SY3200 into 384-well plates containing cell lysis and RT mix (as described below). After sorting, the plates were immediately sealed and briefly centrifuged before proceeding with TIRTL-seq.
T cell expansion
Frozen PBMCs were thawed as described above. After centrifugation (500g, 5 min, room temperature), the supernatant was discarded and the cell pellet resuspended in 5 ml RP10 medium (RPMI 1640 (Gibco) supplemented with 10% human AB serum (Gemini BioProducts), 2 mM L-glutamine (Gibco) and 100 U ml−1 penicillin–streptomycin (Gibco)). Cells were counted as described above. Following another centrifugation (500g, 5 min), the supernatant was discarded and the cell pellet was resuspended to 1 million cells per ml in RP10 medium supplemented with 50 ng ml−1 anti-human CD3 antibody (Miltenyi, clone OKT3), 3000 IU ml−1 recombinant human IL-2 (Stemcell Technologies) and 15 ng ml−1 recombinant human IL-15 (PeproTech). A total of 100 µl cell suspension (100,000 PBMCs) per well was plated in a 96-well round-bottom plate. Cells were incubated at 37 °C with 5% CO2 for 7 days.
CD4 and CD8 selection
Frozen PBMCs were thawed as described above. After centrifugation (500g, 5 min, room temperature), the supernatant was discarded and the cell pellet was resuspended in 1 ml Dynabeads Buffer (0.1% BSA, 2 mM EDTA in DPBS). CD8+ and CD4+ T cells were sequentially positively isolated using Dynabeads CD8 and CD4 Positive Isolation Kits (Thermo Scientific), respectively, according to the manufacturer’s protocol. In brief, for CD8 selection, 25 µl CD8 Dynabeads were washed with 1 ml Dynabeads Buffer, then resuspended with PBMCs in 1 ml Dynabeads Buffer. PBMCs and beads were incubated with slow continuous rotation for 20 min at 4 °C. After incubation, the cell–bead suspension was placed on a magnet for 2 min. Supernatant containing the CD8− fraction was then carefully removed and used for CD4 T cell isolation. The bead-bound CD8+ fraction was washed once with DPBS, then resuspended in 100 µl complete RPMI medium and placed on ice during CD4 selection. For CD4 selection, 25 µl CD4 Dynabeads were washed with 1 ml Dynabeads Buffer, then resuspended with the CD8− supernatant. CD8− cells and beads were incubated with slow continuous rotation for 20 min at 4 °C. After incubation, the cell–bead suspension was placed on a magnet for 2 min, after which CD4− supernatant was carefully removed. The bead-bound CD4+ fraction was washed once with DPBS, then resuspended in 100 µl complete RPMI medium and placed on ice. Both bead-bound CD4+ and CD8+ populations were then detached from beads by adding 10 µl CD4 or CD8 DETACHaBEAD reagent to the appropriate tube and then incubating with slow continuous rotation for 45 min at room temperature. After incubation, tubes were placed on a magnet for 1 min. Supernatants containing isolated CD4+ and CD8+ T cells were carefully removed and placed in new tubes, and residual cells were collected by washing the beads with 500 µl complete RPMI twice and collecting supernatant. Isolated CD4+ and CD8+ T cells were then washed twice with DPBS before proceeding with TIRTL-seq.
TIRTL-seq library preparation
Thawed PBMCs were centrifuged (500g, 5 min), and the cell pellet was resuspended in 5 ml DPBS (Gibco). Cells were counted as described above. Following another centrifugation (500g, 5 min), the supernatant was discarded and the cell pellet was resuspended in DPBS (Gibco) to the desired final volume and kept on ice until use.
RT and PCR I were performed in 384-well plates pre-loaded with 3 µl Vapor-Lock (Qiagen). Cell lysis buffer and reverse transcriptase mix (0.4 µl per well) containing 0.1% TritonX-100 (Sigma-Aldrich), 0.5 mM deoxyribonucleotide triphosphate (dNTP; Thermo Scientific), 2.5 ng µl−1 Random Hexamer Primer (Thermo Scientific), 2.5 µM Oligo(dT) (Thermo Scientific), 1× Maxima H RT buffer (Thermo Scientific), 5 mM dithiothreitol (Thermo Scientific), 1 U µl−1 RNasin ribonuclease Inhibitor (Promega) and 4 U µl−1 Maxima H Minus Reverse Transcriptase (Thermo Scientific) was dispensed into 384-well plates followed by cell suspension (0.250 µl per well) using I-DOT Mini (Dispendix). The plate was pulse centrifuged to ensure that the lysis buffer containing RT master mix and the cell suspension were merged underneath the Vapor-Lock overlay. RT was performed by incubating the plate at 42 °C for 5 min, 25 °C for 10 min, 50 °C for 60 min and 94 °C for 5 min.
For multiplex PCR I, an equimolar mix at 1.125 µM each of 88 V-segment forward primers (see the Supplementary information for the complete sequence) containing Illumina Nextera reverse adapter sequence was prepared. For reverse primers, an equimolar mix at 10 µM each of TRBC and TRAC C-segment-specific primers with 6- or 8-nt barcodes for experimental identification and Illumina Nextera forward adapter sequence (see the Supplementary information for the complete sequence) were prepared. Multiplex PCR I master mix was prepared by mixing 0.45 µl of V-segment forward primer mix, 0.2 µl C-segment reverse primer mix with experiment-specific barcodes and 1.25 µl KAPA2G Fast Multiplex Mix (Roche) with nuclease-free water (Thermo Scientific) to a final volume of 2 µl. PCR I mix (2 µl per well) was dispensed directly into the RT plate using a MANTIS Liquid Dispenser (Formulatrix) to perform targeted amplification of TCRα and TCRβ cDNA. Following the addition of PCR I mix, the plate was centrifuged to ensure the PCR mix was merged underneath the Vapor-Lock overlay. The plate was incubated for 3 min at 95 °C for initial denaturation, followed by 20 cycles of 15 s at 95 °C, 30 s at 59 °C and 1 min at 72 °C. Final elongation was performed for 5 min at 72 °C. Pre- and post-PCR I steps were performed in different rooms to avoid cross-contamination.
For indexing PCR II, 1 µM each of forward and reverse primers containing full-length Illumina adapter sequences and 384-well-specific UDI barcodes (see the Supplementary information for the complete sequence) were prepared and stored at −20 °C. PCR II mix was prepared by mixing 1 µl 5× Q5 Reaction Buffer (NEB), 0.1 µl dNTP Mix (10 mM each, Thermo Scientific), 0.05 µl Q5 Hot Start High-Fidelity DNA Polymerase (NEB) and nuclease-free water (Thermo Scientific) to a final volume of 3 µl. PCR II mix (3 µl per well) was transferred to a new 384-well plate containing Vapor-Lock overlay and centrifuged briefly. A total of 1 µl of a mix of Illumina UDI primers with well-specific barcodes (1 µM each) was stamped into a PCR II plate using Viaflo 384 (Integra Biosciences). The PCR I products were diluted ~1:10 by adding 18 µl Ultrapure Distilled Water using a Welljet dispenser (Integra Biosciences). Thereafter, 1 µl of the diluted PCR I products were transferred into a PCR II plate using Viaflo 384 (Integra Biosciences). The plate was spun down briefly and incubated at 98 °C for 30 s, followed by 15 cycles of 10 s at 98 °C, 10 s at 58 °C and 50 s at 72 °C. Final elongation was performed for 2 min at 72 °C. The libraries were pooled in the VBLOK200 reservoir (ClickBio) by spinning the PCR II plate upside down and purified using 1X AMPure XP beads (Beckman Coulter). Final libraries were sequenced on an Illumina NovaSeq platform at 300,000 reads per well.
Manual 96-well-plate TIRTL-seq
For the manual version of the protocol, the following changes were made: All steps were carried out in 96-well plates with no Vapor-Lock, all pipetting steps were carried out using standard 8- or 12-well multichannel pipettes, volumes for RT and PCR I were increased 4× (a final volume of 2.6 µl and 10.6 µl per well, respectively) and volumes for PCR II were increased 2× (a final volume of 10 µl per well). Libraries were sequenced on an Illumina NovaSeq platform at 2 million reads per well. The 96-well format accommodates a similar total cell input per plate as the 384-well format owing to larger well volumes and offers slightly lower costs per plate. However, the reduced number of wells (96 versus 384) leads to lower statistical power for both pairing algorithms, resulting in fewer unique TCR pairs identified for a given cell input (Fig. 2a) and requiring adjustment of the T-SHELL P value threshold from 10−10 to 10−3 for optimal performance (Extended Data Fig. 1d,e). The 96-well protocol is validated for manual multichannel pipetting and is adaptable to standard tip-based liquid handling robots.
Jurkat spike-in
Jurkat (clone E6-1) cells were obtained from the American Type Culture Collection (ATCC) and cultured in complete RPMI medium (RPMI 1640 (Gibco) supplemented with 10% FBS (Gibco), 2 mM L-glutamine (Gibco) and 100 U ml−1 penicillin–streptomycin (Gibco)). For spike-in experiments, Jurkats were washed with Dynabeads Buffer (0.1% BSA, 2 mM EDTA in DPBS) and stained with Ghost Violet 510 viability dye (1:100, Tonbo Biosciences). Defined numbers of live Jurkats were then sorted using a CytoFLEX SRT cell sorter (Beckman Coulter) to achieve three different final spike-in concentrations: 10% Jurkats (that is, 100,000 Jurkats per 1 × 106 CD8+ T cells), 1% Jurkats (that is, 10,000 Jurkats per 1 × 106 CD8+ T cells) and 0.01% Jurkats (that is, 100 Jurkats per 1 × 106 CD8+ T cells). Meanwhile, PBMCs were thawed, counted and CD4+/CD8+ selected as described above. CD8+ T cells were then added to the sorted Jurkats to achieve final Jurkat concentrations of 10%, 1% and 0.01%. Specifically, as shown in Extended Data Fig. 1a, 1.25 × 106 CD8+ T cells were added to 125,000 Jurkats for the 10% partition; 1.25 × 106 CD8+ T cells were added to 12,500 Jurkats for the 1% partition; and 2.5 × 106 CD8+ T cells were added to 250 Jurkats for the 0.01% partition, for a total of approximately 5 × 106 total cells in the 384-well plate. As shown in Extended Data Fig. 1b, 5 × 106 CD8+ T cells were added to 500,000 Jurkats to achieve a final concentration of 10%, with total cell numbers approximately equal to the full multi-partition 384-well plate (Extended Data Fig. 1a). As shown in Extended Data Fig. 1c, 1.25 × 106 CD8+ T cells were added to 125,000 Jurkats to achieve a final concentration of 10%, with total cell numbers equal to the 10% partition in the multi-partition 384-well plate (Extended Data Fig. 1a). Each cell mixture was centrifuged (500g, 5 min) and resuspended in DPBS to the target volume for dispensing for 384-well or manual 96-well TIRTL-seq, which were then completed as described above.
Bulk 5′RACE
Frozen PBMCs were thawed as described above. After centrifugation (500g, 5 min, room temperature), the supernatant was discarded and the cell pellet resuspended in 5 ml complete RPMI for counting on a Vi-Cell XR automatic cell counter (Beckman Coulter). After counting, the cell suspension was split equally into two replicate aliquots, each containing approximately 5 million viable PBMCs. Following another centrifugation (500g, 5 min), the supernatant was discarded and the cell pellets were resuspended in 1 ml Trizol (Thermo Scientific) each. RNA was isolated from each replicate sample according to the manufacturer’s protocol and quantified by Qubit RNA High Sensitivity Assay (Thermo Scientific). cDNA synthesis was carried out as described previously45. In brief, 250 ng RNA and a cDNA synthesis mix containing 1X First Strand Buffer (Takara Bio), 2 mM dithiothreitol (Thermo Scientific), 1 mM of each dNTP (Thermo Scientific), 1 μM each of human TCRα and TCRβ RT UMI primers, 1 μM 5′ template-switch adapter, 10 U µl−1 Smartscribe Reverse Transcriptase (Takara Bio) and 0.4 U µl−1 RNasin (Promega) were mixed together, followed by incubation at 42 °C for 60 min. After cDNA synthesis, Uracil-DNA Glycosylase (NEB) was added at 0.25 U µl−1, and samples were incubated for an additional 40 min at 37 °C. cDNA was purified using 1X AMPure XP beads (Beckman Coulter).
PCR I reaction mix was prepared by mixing purified cDNA with 1X Q5 polymerase buffer (NEB), 0.2 mM of each dNTP (Thermo Scientific), 0.2 μM M1ss forward primer, 0.2 μM each of human TCRα and human TCRβ UMI first round primers, 0.02 U µl−1 Q5 Hot Start Polymerase (NEB) and nuclease-free water to a total volume of 50 µl per reaction. PCR I was performed by incubating at 98 °C for 30 s, followed by 20 cycles of 98 °C for 10 s, 55 °C for 10 s and 72 °C for 50 s. Final elongation was performed at 72 °C for 2 min. Pre- and post-PCR workflows were performed in separate rooms to prevent cross-contamination.
For PCR II, each sample was split and TCRα and TCRβ samples were processed separately. PCR II master mix was prepared by mixing 1× Q5 polymerase buffer (NEB), 0.2 mM of each dNTP, 0.02 U µl−1 Q5 polymerase and nuclease-free water to a final volume of 50 µl per reaction. Two tubes (one TCRα and one TCRβ) were prepared for each sample, and 2 µl unpurified PCR I product, 0.2 μM H1s primer and 0.2 μM human acj_i (TCRα) or bcj_i (TCRβ) were added to each sample. PCR II was performed by incubating at 98 °C for 30 s, followed by 16 cycles of 98 °C for 10 s, 58 °C for 10 s and 72 °C for 50 s. Final elongation was performed at 72 °C for 2 min. PCR II product was purified using 0.8× AMPure XP beads (Beckman Coulter).
10x Genomics scTCR-seq
PBMCs were stained as described in the ‘Single-cell sorting’ section. Live, single CD3+ cells were sorted on a Beckman Coulter Cytoflex SRT sorter, counted on a hemocytometer, adjusted to 1,400 live cells per µl in DPBS and loaded into 10x Genomics Chromium GEM-X 5′ reaction for target recovery of 20,000 cells. scTCR libraries were prepared by following the manufacturer’s protocol for Chromium GEM-X Single Cell 5′ Reagent Kits v3 (CG000733, Rev A) and sequenced on an Illumina NovaSeq platform.
Raw data processing
Raw fastq files from each well were processed with the mixcr version 4.6.046 package using the ‘analyze’ tool. Optional switches were set to analyze the ‘generic-amplicon’ preset with ‘floating-left-alignment-boundary’, ‘floating-right-alignment-boundary’ for C segment and ‘split-by-sample’. All other parameters were set to default. ‘Tag-pattern’ switch was used to match and split the samples on the basis of plate barcodes incorporated during the PCR I step. Processing steps included demultiplexing raw reads by plate barcode; aligning V, D and J segments; assembling identical reads into clonotypes; and frequency-based error-correction.
Statistics and reproducibility
Sample size in terms of number of cells per well was determined on the basis of the yield from the test experiment (Fig. 1e). To select the number of replicate wells per sample, we used guidelines from ref. 21 and used 96 wells or more for each sample; see Fig. 2a for yield scaling with an increasing sample size. No donors were excluded from the analysis. Particular replicate wells of 384-well plates were excluded if they did not pass our quality control criteria (see below). The experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment.
scTCR-seq with TIRTL-seq
To call functional TCR chains from single-cell sorted T cells and filter sequencing errors from each well, we selected functional (no frameshifts and no stop codons inside CDR3) TCR chains with >50 reads and >10% of reads in a well corresponding to these clonotypes.
TCR pairing with MAD-HYPE
Owing to variability in cell dispensing and library preparation, some wells produce substantially fewer clonotypes than average. To avoid confounding the downstream analysis, we excluded wells with <50% of the average number of unique TCRα clonotypes across the plate.
We reimplemented the MAD-HYPE algorithm23 in R and Python 3. In brief, for the given experimental design (number of wells and number of cells per well) and TCRα and TCRβ overlap statistics (wij, the number of wells in which TCRα and TCRβ are found together; wi, the number of wells with TCRα only; and wj, the number of wells with TCRβ only), MAD-HYPE outputs a posterior probability log ratio for the hypothesis that TCRα and TCRβ are paired versus unpaired. To call TCRαβ chain pairs, we calculated this ratio for all the possible TCRαβ pairs and filtered all pairs with a score above 0.1. To speed up the computations, we first computed a look-up table containing threshold values for maximal chain loss: for each overlap wij and for each loss of TCRβ wi, what is the maximum loss of TCRα wj such that the paired versus unpaired hypothesis probability log ratio is still above the 0.1 threshold.
To efficiently calculate wij, wi and wj for all the possible TCRαβ pairs, we used GPU and cupy47 or MLX frameworks48, although for the vast majority of 384-well-plate TIRTL-seq experiments, a single central processing unit core is enough to analyze the results with our implementation.
T-SHELL heuristic algorithm for TCRα/TCRβ pairing
If the clone is large and present in all (or almost all) wells, there should still be Poissonian variation in the exact number of cells per well, influencing the clonal frequency of the respective TCRα and TCRβ transcripts (fraction of reads corresponding to this clone of all reads) measured in each well.
To pair the αβ TCR chains found in all the wells, we calculated the Pearson correlation coefficient between per well clonal frequencies (defined as the sum of reads corresponding to a given unique α or β CDR3nt sequence divided by the total number of sequencing reads per well) for each possible TCRα/TCRβ pair. We next computed the P value for H0: r = 0 (using Student’s t-distribution with n − 2 degrees of freedom, where n is the number of wells). After that, for each TCRα, we ordered P values and divide each by the third-smallest P value to get an adjusted P value, reflecting our belief that each TCRα is paired to two TCRβ sequences at most. We then put a 10−10 threshold on the adjusted P value to call chain pairs for 384- and 192-well experiments and a 10−3 threshold for 96-well experiments.
Identification of expanded and contracted clones
To identify expanded and contracted clones between time points with TIRTL-seq, we first averaged relative frequency (defined as the sum of reads mapping to a given CDR3β divided by the sum of reads mapping to all CDR3βs) across all wells in a plate, including wells in which the clone was not present. We filtered out clones found in fewer than five wells at either time point to limit the number of comparisons. For each clonotype, we calculated the standard error of the mean at both time points and called clones significantly expanding or contracting if log2FC between the average frequencies at two time points was >3 and if 2.5 s.e.m. intervals did not overlap. If the clone had a very low frequency at one of the time points and was found in fewer than three wells, we assumed that s.e.m. was twice the s.e.m. of the clones found in three wells at this time point, and a pseudocount of 10−6 was added to all clonal frequencies to obtain reasonable fold change estimates.
To identify expanded and contracted clones from bulk TCRβ repertoire replicates, we used the edgeR package49, as described previously2, using Padj < 0.001 and log2FC < 2 thresholds.
TCR specificity matching to VDJdb
To identify clusters of highly similar clones matching previously reported TCR specificities, we coclustered paired data from the acute time point with VDJdb and identified tightly interconnected clusters, including both TCRs with previously described specificity and αβ TCRs from our experiment. First, we filtered VDJdb (accessed on 6 August 2024) for paired human TCRs with reported HLA-restriction matching HLA class I alleles of our donor (A*02, A*03, B*07, B*08 and C*07). We excluded A*02 GILGFVFTL and A*03-restricted KLGGALQAK epitopes to avoid spurious matching. We then merged the cleaned-up VDJdb dataset with paired TCRs detected on acute time points and used TCRdist < 90 to connect similar clones with edges. To separate tightly interconnected clusters of highly similar TCRs, we excluded 1% of nodes with the highest betweenness values from the network. We then assigned specificity to TCR clusters if more than 40% of the nodes from the cluster were from given epitope-specific TCRs from VDJdb.
TCR cloning and screening
TCR cloning and screening was performed as previously described10,50. In brief, to generate artificial antigen-presenting cells, gBlock gene fragments encoding HLA-A*02:01, HLA-B*08:01 and HLA-B*07:02 were synthesized by GenScript and cloned into a pLVX-EF1α-IRES-Puro lentiviral vector (Clontech). Lentivirus was produced by transfecting HEK 293 T cells (ATCC CRL-3216) with the HLA-containing vector, psPAX2 and pMD2.G plasmids (Addgene) at a ratio of 4:3:1 using the Lipofectamine 3000 kit (Thermo Fisher Scientific). Viral supernatant was filtered, concentrated and used to transduce K562 cells (ATCC CCL-243), followed by selection with puromycin for 1 week. Surface HLA expression was confirmed by flow cytometry.
Representative TCRαβ pairs for each epitope were selected (Supplementary Data 2), modified with murine constant regions and linked to mCherry via 2A sites. This construct was cloned into a pLVX-EF1α-IRES-Puro vector. Lentivirus was generated by transfecting HEK 293 T cells (ATCC CRL-3216) with the TCR-mCherry vector, psPAX2 and pMD2.G at a ratio of 4:3:1 using the Lipofectamine 3000 kit (Thermo Fisher Scientific). 2D3 Jurkat 76.7 cells expressing human CD8+ and NFAT-GFP51 (a kind gift from Fumihiro Fujiki) were transduced and selected with puromycin for 1 week.
Resulting TCR-transgenic Jurkat cells (105) were cocultured with 105 K562 aAPC cells with a single predicted HLA in a round-bottom 96-well cell culture plate in 100 µl RPMI 1640 media (Gibco) supplemented with 10% FBS, 1% penicillin–streptomycin and 1% L-glutamine and pulsed with predicted peptide at a final concentration of 10 µM. Predicted HLA-B*07:02-SPRWYFYYL-specific TCRs were additionally tested against LPRWYFYYL, an epitope variant found in HKU1 and OC43 common cold coronaviruses. Fraction of GFP+ cells among mCherry+ cells was measured on BD FACSymphony A3 flow cytometer.
Anti-EBV and anti-SARS-CoV-2 enzyme-linked immunosorbent assay
Anti-SARS-CoV-2 IgG analysis was performed as previously described52. In brief, 384-well microtiter plates were coated overnight at 4 °C, with recombinant SARS-CoV-2 Spike (Sino Biological) diluted in PBS at 2 µg ml−1. Plates were washed three times the next day with PBS with Tween 20 (PBS-T; 0.1% Tween 20) before being blocked with 3% Omniblok nonfat milk (AmericanBio) in PBS-T for 1 h. Plates were washed as before, then incubated with the serum samples diluted 1:50 in 1% milk in PBS-T for 90 min at room temperature. Before dilution, serum samples were thawed and heat-inactivated at 56 °C for 15 min. The plates were washed as before and incubated for 30 min at room temperature with anti-human IgG secondary antibody (Invitrogen) at a 1:10,000 dilution in 1% milk in PBS-T. After plates were washed as before, they were incubated with SIGMAFAST OPD (Sigma-Aldrich) for 8 min in the dark at room temperature. To stop the chemiluminescence reaction, 3 N HCl was added to the wells of the plate. The plates were then read at 490 nm on a microplate reader. Binding antibody units per ml (BAU per ml) were determined by comparing the optical densities (ODs) of the target samples with those of blank wells and samples calibrated to World Health Organization standard samples (NIBSC) set at 1,000 BAU per ml. A cutoff value of 90.9 BAU per ml was used to determine anti-SARS-CoV-2 Spike IgG positivity. This value has been used by others when measuring these antibody levels using the same World Health Organization control53 and was slightly higher than the values of other baseline serum samples from SJTRC participants. For anti-EBV viral capsid antigen IgG analysis, serum samples were prepared and measured according to the manufacturer’s instructions (Abcam ab108730). Standard units were calculated using the provided control sample OD measured at 450 nm. The ODs of the control samples and blanks were within the manufacturer’s criteria for a valid assay run.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
TCR sequencing data are available via Zenodo at https://doi.org/10.5281/zenodo.14010377 (ref. 54). Raw sequencing data are available via the Sequence Read Archive (SRA) (PRJNA1256512).
Code availability
Code is available via GitHub at https://github.com/pogorely/TIRTL.
References
DeWitt, W. S. et al. Dynamics of the cytotoxic T cell response to a model of acute viral infection. J. Virol. 89, 4517–4526 (2015).
Pogorelyy, M. V. et al. Precise tracking of vaccine-responding T cell clones reveals convergent and personalized response in identical twins. Proc. Natl Acad. Sci. USA 115, 12704–12709 (2018).
Akondy, R. S. et al. The yellow fever virus vaccine induces a broad and polyfunctional human memory CD8+ T cell response. J. Immunol. 1950 183, 7919–7930 (2009).
Robins, H. S. et al. Comprehensive assessment of T-cell receptor β-chain diversity in αβ T cells. Blood 114, 4099–4107 (2009).
Wang, C. et al. High throughput sequencing reveals a complex pattern of dynamic interrelationships among human T cell subsets. Proc. Natl Acad. Sci. USA 107, 1518–1523 (2010).
Garcia, K. C. & Adams, E. J. How the T cell receptor sees antigen—a structural view. Cell 122, 333–336 (2005).
Krogsgaard, M. & Davis, M. M. How T cells ‘see’ antigen. Nat. Immunol. 6, 239–245 (2005).
Dash, P. et al. Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 547, 89–93 (2017).
Glanville, J. et al. Identifying specificity groups in the T cell receptor repertoire. Nature 547, 94–98 (2017).
Mudd, P. A. et al. SARS-CoV-2 mRNA vaccination elicits a robust and persistent T follicular helper cell response in humans. Cell 185, 603–613 (2022).
Meysman, P. et al. Benchmarking solutions to the T-cell receptor epitope prediction problem: IMMREP22 workshop report. ImmunoInformatics 9, 100024 (2023).
Dash, P. et al. Paired analysis of TCRα and TCRβ chains at the single-cell level in mice. J. Clin. Invest. 121, 288–295 (2011).
Han, A., Glanville, J., Hansmann, L. & Davis, M. M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Ludwig, J., Huber, A.-K., Bartsch, I., Busse, C. E. & Wardemann, H. High-throughput single-cell sequencing of paired TCRα and TCRβ genes for the direct expression-cloning and functional analysis of murine T-cell receptors. Eur. J. Immunol. 49, 1269–1277 (2019).
McDaniel, J. R., DeKosky, B. J., Tanno, H., Ellington, A. D. & Georgiou, G. Ultra-high-throughput sequencing of the immune receptor repertoire from millions of lymphocytes. Nat. Protoc. 11, 429–442 (2016).
Spindler, M. J. et al. Massively parallel interrogation and mining of natively paired human TCRαβ repertoires. Nat. Biotechnol. 38, 609–619 (2020).
Ortolano, N. The neXt generation of single cell RNA-seq: an introduction to GEM-X technology. 10x Genomics https://www.10xgenomics.com/blog/the-next-generation-of-single-cell-rna-seq-an-introduction-to-gem-x-technology (2024).
Hagemann-Jensen, M., Ziegenhain, C. & Sandberg, R. Scalable single-cell RNA sequencing from full transcripts with Smart-seq3xpress. Nat. Biotechnol. 40, 1452–1457 (2022).
Hahaut, V. et al. Fast and highly sensitive full-length single-cell RNA sequencing using FLASH-seq. Nat. Biotechnol. 40, 1447–1451 (2022).
Rosenberg, A. B. et al. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360, 176–182 (2018).
Howie, B. et al. High-throughput pairing of T cell receptor α and β sequences. Sci. Transl. Med. 7, 301ra131 (2015).
Lee, E. S., Thomas, P. G., Mold, J. E. & Yates, A. J. Identifying T cell receptors from high-throughput sequencing: dealing with promiscuity in TCRα and TCRβ pairing. PLoS Comput. Biol. 13, e1005313 (2017).
Holec, P. V., Berleant, J., Bathe, M. & Birnbaum, M. E. A Bayesian framework for high-throughput T cell receptor pairing. Bioinformatics 35, 1318–1325 (2019).
Genolet, R. et al. TCR sequencing and cloning methods for repertoire analysis and isolation of tumor-reactive TCRs. Cell Rep. Methods 3, 100459 (2023).
Ma, K.-Y. et al. Immune repertoire sequencing using molecular identifiers enables accurate clonality discovery and clone size quantification. Front. Immunol. 9, 33 (2018).
Garner, L. C., Klenerman, P. & Provine, N. M. Insights Into mucosal-associated invariant T cell biology from studies of invariant natural killer T cells. Front. Immunol. 9, 1478 (2018).
Dupic, T., Marcou, Q., Walczak, A. M. & Mora, T. Genesis of the αβ T-cell receptor. PLoS Comput. Biol. 15, e1006874 (2019).
Padovan, E. et al. Expression of two T cell receptor α chains: dual receptor T cells. Science 262, 422–424 (1993).
Goncharov, M. et al. VDJdb in the pandemic era: a compendium of T cell receptors specific for SARS-CoV-2. Nat. Methods 19, 1017–1019 (2022).
Minervina, A. A. et al. Primary and secondary anti-viral response captured by the dynamics and phenotype of individual T cell clones. eLife 9, e53704 (2020).
Rojas, L. A. et al. Personalized RNA neoantigen vaccines stimulate T cells in pancreatic cancer. Nature 618, 144–150 (2023).
Ford, E. S. et al. Repeated mRNA vaccination sequentially boosts SARS-CoV-2-specific CD8+ T cells in persons with previous COVID-19. Nat. Immunol. 25, 166–177 (2024).
Lossius, A. et al. High-throughput sequencing of TCR repertoires in multiple sclerosis reveals intrathecal enrichment of EBV-reactive CD8+ T cells. Eur. J. Immunol. 44, 3439–3452 (2014).
Huth, A., Liang, X., Krebs, S., Blum, H. & Moosmann, A. Antigen-specific TCR signatures of cytomegalovirus infection. J. Immunol. 202, 979–990 (2019).
Wahl, I. et al. An efficient single-cell based method for linking human T cell phenotype to T cell receptor sequence and specificity. Eur. J. Immunol. 52, 237–246 (2022).
Carlson, C. S. et al. Using synthetic templates to design an unbiased multiplex PCR assay. Nat. Commun. 4, 2680 (2013).
Birnbaum, M. E. et al. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157, 1073–1087 (2014).
Kula, T. et al. T-Scan: a genome-wide method for the systematic discovery of T cell epitopes. Cell 178, 1016–1028 (2019).
Kisielow, J., Obermair, F.-J. & Kopf, M. Deciphering CD4+ T cell specificity using novel MHC–TCR chimeric receptors. Nat. Immunol. 20, 652–662 (2019).
Joglekar, A. V. et al. T cell antigen discovery via signaling and antigen-presenting bifunctional receptors. Nat. Methods 16, 191–198 (2019).
Dobson, C. S. et al. Antigen identification and high-throughput interaction mapping by reprogramming viral entry. Nat. Methods 19, 449–460 (2022).
Zdinak, P. M. et al. De novo identification of CD4+ T cell epitopes. Nat. Methods 21, 846–856 (2024).
Kohlgruber, A. C. et al. High-throughput discovery of MHC class I- and II-restricted T cell epitopes using synthetic cellular circuits. Nat. Biotechnol. 43, 623–634 (2025).
Hudson, D., Fernandes, R. A., Basham, M., Ogg, G. & Koohy, H. Can we predict T cell specificity with digital biology and machine learning?. Nat. Rev. Immunol. 23, 511–521 (2023).
Egorov, E. S. et al. Quantitative profiling of immune repertoires for minor lymphocyte counts using unique molecular identifiers. J. Immunol. 194, 6155–6163 (2015).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Okuta, R., Unno, Y., Nishino, D., Hido, S. & Loomis, C. CuPy: a NumPy-compatible library for NVIDIA GPU calculations. In Proc. Workshop on Machine Learning Systems (LearningSys) in 31st Annual Conference on Neural Information Processing Systems (NIPS) http://learningsys.org/nips17/assets/papers/paper_16.pdf (2017).
Hannun, A., Digani, J., Katharopoulos, A. & Collobert, R. mlx. GitHub https://github.com/ml-explore (2023).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Minervina, A. A. et al. SARS-CoV-2 antigen exposure history shapes phenotypes and specificity of memory CD8+ T cells. Nat. Immunol. 23, 781–790 (2022).
Morimoto, S. et al. Establishment of a novel platform cell line for efficient and precise evaluation of T cell receptor functional avidity. Oncotarget 9, 34132–34141 (2018).
Lin, C.-Y. et al. Pre-existing humoral immunity to human common cold coronaviruses negatively impacts the protective SARS-CoV-2 antibody response. Cell Host Microbe 30, 83–96 (2022).
Winichakoon, P. et al. Diagnostic performance between in-house and commercial SARS-CoV-2 serological immunoassays including binding-specific antibody and surrogate virus neutralization test (sVNT). Sci. Rep. 13, 34 (2023).
Pogorelyy, M. V. et al. TCR sequencing data. Zenodo https://doi.org/10.5281/zenodo.14010377 (2025).
Acknowledgements
This work was supported by grant nos. R01 AI136514, U01 AI144616 and P01 AI165077 from the St. Jude Center for Influenza Research and Response (SJCEIRR) contract 75N93021C00016, the Center for Influenza Vaccine Research in High Risk Populations (CIVR-HRP) contract 75N93019C00052, the TIRTL Blue-sky Initiative and American Lebanese Syrian Associated Charities (ALSAC) at St. Jude. D.C.B. was supported through The American Association of Immunologists Intersect Fellowship Program for Computational Scientists and Immunologists. The funders had no role in the study design, data collection and interpretation, or decision to submit the work for publication. We thank the Hartwell Center Team at St. Jude and, in particular S. Olsen, G. Neale and D. Darnell for their help with high-throughput sequencing; G. Lennon at the St. Jude Immunology Flow Core for help with cell sorting; A. Walczak, T. Mora, P. Bradley, K. Mayer-Blackwell and A. Fiore-Gartland for their valuable discussions of data analysis methods and A. Green for her advice on serology assays.
Author information
Authors and Affiliations
Consortia
Contributions
Conceptualization: M.V.P., A.M.K. and P.G.T. Methods development: M.V.P., A.M.K., S.A. and B.S. Investigation: M.V.P., A.M.K., A.A.M., S.A., B.S., D.C.B. and Z.B.S. Formal analysis: M.V.P., B.S. and D.C.B. Resources: Z.B.S. and K.V. Data and sample curation: M.V.P., A.M.K., Z.B.S. and the SJTRC Study Group consortium. Visualization: M.V.P., A.M.K. and A.A.M. Writing, original draft: M.V.P. Writing, review and editing: all authors. Funding acquisition: P.G.T. Supervision: M.V.P. and P.G.T.
Corresponding authors
Ethics declarations
Competing interests
P.G.T. is on the Scientific Advisory Board of Immunoscape and Shennon Bio, has received research support and personal fees from Elevate Bio, and consulted for 10X Genomics, Illumina, Pfizer, Cytoagents, Sanofi, Merck and JNJ. P.G.T., M.V.P., A.M.K. and A.A.M. have patents related to TCR amplification, cloning and/or applications thereof. The other authors declare no competing interests.
Peer review
Peer review information
Peer reviewer reports are available. Nature Methods thanks Michael Birnbaum and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Madhura Mukhopadhyay, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Benchmarking of TIRTL-seq with Jurkat spike-in.
a-c. Plate maps showing the number of unique TCRbeta clonotypes detected in each well. Plates a and b were processed with automated 384-well protocol, plate c was processed with manual 96-well protocol without non-contact liquid handling. Row I in the 384-well plates (a,b) and well H12 in the 96-well plate (c) were intentionally left empty (except well I13, which was loaded with Jurkat cells only). d. Distributions of adjusted p-values for clones that were independently paired using 10x Genomics scTCR-seq. Vertical dashed lines indicate the p-value obtained for the Jurkat spike-in pair in each corresponding plate. e. Comparison of the number of pairs identified in the 96-well spike-in experiments by T-SHELL, MADHYPE, or both algorithms, using two different T-SHELL adjusted p-value thresholds (10−3 and 10−10).
Extended Data Fig. 2 Functional validation of paired virus-specific TCRs identified by TIRTL-seq.
a. Gating strategy and representative flow plots for in vitro validation experiments. b. In vitro validation of predicted TCR specificity. Bars show average percentage of GFP+ cells out of TCR-transgenic Jurkat cells (mCherry + ), dots show replicates (n = 2).
Extended Data Fig. 3 Anti-EBV VCA (left) and Anti-SARS-CoV-2 Spike IgG levels across time points.
Lowest dashed line shows seronegativity cut-offs.
Extended Data Fig. 4 Clone size estimation accuracy with TIRTL-seq.
a. Correlation (Spearman’s ρ = 0.71) between clonal frequencies estimated by 10x Genomics scTCR-seq and TIRTL-seq for the most abundant clones on acute timepoint (defined as those with ≥5 cells detected in the 10x data). TIRTL-seq CD8+ frequencies were multiplied by 0.32 (the frequency of CD8+ cells among CD3+ cells for this donor) to estimate frequency relative to total CD3+ cells, comparable to the 10x scTCR-seq measurement. (b) Comparison of log2 fold change (log2FC) between acute and convalescent time points for large clones (defined as those with ≥ 5 cells detected in 10x at either of the compared time points). Log2FC calculated from 10x scTCR-seq (x-axis) is plotted against log2FC calculated from an independent TIRTL-seq experiment on isolated CD8+ cells (y-axis). The dashed diagonal indicates the line of equality. Pearson’s correlation coefficient (R) and p-value are shown. Colors indicate clone status determined by the longitudinal TIRTL-seq analysis shown in Fig. 3b: orange for expanding clones, green for contracting clones, grey for stable clones.
Extended Data Fig. 5 Longitudinal clonal tracking with 10x Genomics Chromium scTCR-seq.
10x Genomics scTCR-seq identifies expansions and contractions in the CD3 + T cell repertoire. Each dot represents TCRβ clonotype, frequency at two timepoints is plotted in log-scale. Orange and green color show significantly expanding and contracting clones, respectively.
Supplementary information
Supplementary Data 1 (download XLS )
List of oligonucleotides.
Supplementary Data 2 (download XLS )
TCR sequences used for cloning.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pogorelyy, M.V., Kirk, A.M., Adhikari, S. et al. TIRTL-seq: deep, quantitative and affordable paired TCR repertoire sequencing. Nat Methods 23, 56–64 (2026). https://doi.org/10.1038/s41592-025-02907-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-025-02907-9
This article is cited by
-
TIRTL-seq: heroes with T-SHELL
Nature Methods (2026)
-
A therapeutic peptide vaccine for fibrolamellar hepatocellular carcinoma: a phase 1 trial
Nature Medicine (2025)