Abstract
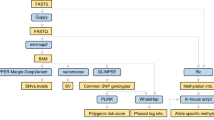
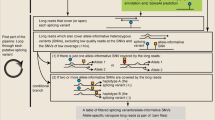
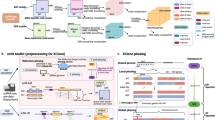
Long-read RNA sequencing is a powerful technology to link transcript structures to genetic variants, but this type of analysis is not often performed owing to the lack of end-user tools. Here we introduce longcallR for joint single-nucleotide polymorphism calling, haplotype phasing and allele-specific analysis, which achieves high accuracy on benchmark datasets. Applied to 202 human samples, longcallR identified 88 significant allele-specific splicing events per sample on average, of which 46% involved unannotated junctions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The raw reads for HG002a:MAS:cDNA are available at https://downloads.pacbcloud.com/public/dataset/Kinnex-full-length-RNA/DATA-Revio-HG002-1/. The raw reads for HG002b:MAS:cDNA, HG002c:MAS:cDNA, HG002d:MAS:cDNA, HG004:MAS:cDNA, HG005:MAS:cDNA, HG002a:ISO:cDNA, HG002b:ISO:cDNA, HG002c:ISO:cDNA, HG004:ISO:cDNA and HG005:ISO:cDNA can be accessed at https://ftp-trace.ncbi.nlm.nih.gov/giab/ftp/data_RNAseq/. The WTC11:ONT:cDNA reads can be found in ENCODE under ENCSR539ZXJ. The raw reads for HG002a:ONT:cDNA, HG002b:ONT:cDNA, HG004:ONT:cDNA and HG005:ONT:cDNA are available at https://s3.amazonaws.com/gtl-public-data/giab/bams/cDNA/05_09_23_R941_GIAB_cDNA_PCS111_NA26105_Guppy_6.4.6_sup.pass.fastq.gz.hg38.bam, https://s3.amazonaws.com/gtl-public-data/giab/bams/cDNA/05_09_23_R941_GIAB_cDNA_PCS111_NA27730_Guppy_6.4.6_sup.pass.fastq.gz.hg38.bam, https://s3.amazonaws.com/gtl-public-data/giab/bams/cDNA/05_09_23_R941_GIAB_cDNA_PCS111_NA24143_Guppy_6.4.6_sup.pass.fastq.gz.hg38.bam and https://s3.amazonaws.com/gtl-public-data/giab/bams/cDNA/05_09_23_R941_GIAB_cDNA_PCS111_NA24631_Guppy_6.4.6_sup.pass.fastq.gz.hg38.bam. The HG002:ONT:dRNA, HG004:ONT:dRNA and HG005:ONT:dRNA datasets, generated by the University of Hong Kong, are available from the NCBI under SRX26304755, SRX26304756 and SRX26304757. HPRC human samples are available at https://s3-us-west-2.amazonaws.com/human-pangenomics/index.html?prefix=working/HPRC/. For ground-truth DNA variants, the VCF files for HG002, HG004 and HG005 are available at GIAB v4.2.1. The VCF for WTC11 is provided by the Allen Institute at https://open.quiltdata.com/b/allencell/tree/aics/wtc11_short_read_genome_sequence/. The reference genome GRCh38 can be downloaded from NCBI. All generated SNP calls, ASJs and underlying data for the figures are available via Zenodo at https://doi.org/10.5281/zenodo.17842979 (ref. 27). Source data are provided with this paper.
Code availability
LongcallR is available via GitHub at https://github.com/huangnengCSU/longcallR. LongcallR-nn is available via GitHub at https://github.com/huangnengCSU/longcallR-nn. LongcallR scripts are available via GitHub at https://github.com/huangnengCSU/longcallR_scripts. The Nextflow of longcallR is available via GitHub at https://github.com/huangnengCSU/longcallR-nf.
References
Loving, R. K. et al. Long-read sequencing transcriptome quantification with lr-kallisto. PLoS Comput. Biol. 21, e101369 (2025).
Jousheghani, Z. Z. et al. Oarfish: enhanced probabilistic modeling leads to improved accuracy in long read transcriptome quantification. Bioinformatics 41, i304–i313 (2025).
Chen, Y. et al. Context-aware transcript quantification from long-read RNA-seq data with Bambu. Nat. Methods 20, 1187–1195 (2023).
Prjibelski, A. D. et al. Accurate isoform discovery with IsoQuant using long reads. Nat. Biotechnol. 41, 915–918 (2023).
Gao, Y. et al. ESPRESSO: robust discovery and quantification of transcript isoforms from error-prone long-read RNA-seq data. Sci. Adv. 9, eabq5072 (2023).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 278 (2019).
Pardo-Palacios, F. J. et al. SQANTI3: curation of long-read transcriptomes for accurate identification of known and novel isoforms. Nat. Methods 21, 793–797 (2024).
Quinones-Valdez, G., Amoah, K. & Xiao, X. Long-read RNA-seq demarcates cis- and trans-directed alternative RNA splicing. Nat. Commun. 16, 9603 (2025).
Tang, A. D. et al. Full-length transcript characterization of SF3B1 mutation in chronic lymphocytic leukemia reveals downregulation of retained introns. Nat. Commun. 11, 1438 (2020).
Tilgner, H., Grubert, F., Sharon, D. & Snyder, M. P. Defining a personal, allele-specific, and single-molecule long-read transcriptome. Proc. Natl Acad. Sci. USA 111, 9869–9874 (2014).
Tang, A. D. et al. Detecting haplotype-specific transcript variation in long reads with FLAIR2. Genome Biol. 25, 173 (2024).
Deonovic, B., Wang, Y., Weirather, J., Wang, X.-J. & Au, K. F. IDP-ASE: haplotyping and quantifying allele-specific expression at the gene and gene isoform level by hybrid sequencing. Nucleic Acids Res. 45, e32 (2017).
Edge, P. & Bansal, V. Longshot enables accurate variant calling in diploid genomes from single-molecule long read sequencing. Nat. Commun. 10, 4660 (2019).
Shafin, K. et al. Haplotype-aware variant calling with PEPPER-Margin-DeepVariant enables high accuracy in nanopore long-reads. Nat. Methods 18, 1322–1332 (2021).
Zheng, Z. et al. Symphonizing pileup and full-alignment for deep learning-based long-read variant calling. Nat. Comput. Sci. 2, 797–803 (2022).
Ahsan, M. U., Liu, Q., Fang, L. & Wang, K. NanoCaller for accurate detection of SNPs and indels in difficult-to-map regions from long-read sequencing by haplotype-aware deep neural networks. Genome Biol. 22, 261 (2021).
Huang, N. et al. NanoSNP: a progressive and haplotype-aware SNP caller on low-coverage nanopore sequencing data. Bioinformatics 39, btac824 (2023).
Wang, B. et al. Variant phasing and haplotypic expression from long-read sequencing in maize. Commun. Biol. 3, 78 (2020).
De Souza, V. B. C. et al. Transformation of alignment files improves performance of variant callers for long-read RNA sequencing data. Genome Biol. 24, 91 (2023).
Zheng, Z. et al. Clair3-RNA: a deep learning-based small variant caller for long-read RNA sequencing data. Nat. Commun. 16, 11553 (2025).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 770–778 (IEEE, 2016).
Zook, J. M. et al. An open resource for accurately benchmarking small variant and reference calls. Nat. Biotechnol. 37, 561–566 (2019).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Mansi, L. et al. REDIportal: millions of novel A-to-I RNA editing events from thousands of RNAseq experiments. Nucleic Acids Res. 49, D1012–D1019 (2021).
Lippert, R. et al. Algorithmic strategies for the single nucleotide polymorphism haplotype assembly problem. Brief Bioinform. 3, 23–31 (2002).
Krusche, P. et al. Best practices for benchmarking germline small-variant calls in human genomes. Nat. Biotechnol. 37, 555–560 (2019).
Huang, N. & Li, H. longcallR: SNP calling, haplotype phasing and allele-specific analysis with long RNA-seq reads. Zenodo https://doi.org/10.5281/zenodo.17842979 (2026).
Acknowledgements
This work is supported by National Institute of Health (grant nos. R01HG010040, R01HG014175, U24CA294203, U01HG013748 and U41HG010972 (to H.L.)). We thank the National Genome Research Institute for funding the following grants supporting the creation of the human pangenome reference: U41HG010972, U01HG010971, U01HG013760, U01HG013755, U01HG013748, U01HG013744 and R01HG011274, and the HPRC (BioProject ID: PRJNA730823).
Author information
Authors and Affiliations
Consortia
Contributions
N.H. and H.L. designed the algorithm, implemented longcallR and drafted the paper. N.H. conducted the performance evaluation of longcallR for SNP calling, haplotype phasing and allele-specific analysis. The HPRC provided the 202 MAS-seq human sample data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Andrey Prjibelski and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editors: Lei Tang and Lin Tang, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Accuracy of SNP calling and haplotype phasing.
A, precision and recall for SNP calling on PacBio datasets between longcallR (circles) and Clair3-RNA (triangles). Colors represent different samples. Genomic SNPs from GIAB are taken as the ground truth. B, precision and recall for SNP calling on Nanopore datasets. C, Phasing switch error rates between longcallR-phase (green) and WhatsHap (yellow) across PacBio and Nanopore datasets. Trio-based HG002 SNP phasing from GIAB is taken as the ground truth. Lower switch error rates indicate higher phasing accuracy. D, Phasing hamming error rates in haplotype phasing.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1–15, Figs. 1–12 and Notes 1–4.
Source data
Source Data Fig. 2 (download XLSX )
Upset plot source data, Venn plot source data, cumulative count of significant ASJ genes across HPRC samples, count of known and novel ASJ events across HPRC samples.
Source Data Extended Data Fig. 1 (download XLSX )
Precision and recall for SNP calling on PacBio datasets between longcallR and Clair3-RNA, precision and recall for SNP calling on Nanopore datasets, phasing switch error rates between longcallR-phase and WhatsHap across PacBio and Nanopore datasets, phasing hamming error rates between longcallR-phase and WhatsHap across PacBio and Nanopore datasets.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, N., Li, H. & Human Pangenome Reference Consortium. SNP calling, haplotype phasing and allele-specific analysis with long RNA-seq reads. Nat Methods (2026). https://doi.org/10.1038/s41592-026-03045-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41592-026-03045-6