Abstract
A tool to map changes in synaptic strength during a defined time window could provide powerful insights into the mechanisms of learning and memory. Here we developed a technique, Extracellular Protein Surface Labeling in Neurons (EPSILON), to map α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptor (AMPAR) exocytosis in vivo by sequential pulse-chase labeling of surface AMPARs with membrane-impermeable dyes. This approach yields synaptic-resolution maps of AMPAR exocytosis, a proxy for synaptic potentiation, in genetically targeted neurons during memory formation. In mice undergoing contextual fear conditioning, we investigated the relationship between synapse-level AMPAR exocytosis in CA1 pyramidal neurons and cell-level expression of the immediate early gene product cFos, a frequently used marker of engram neurons. We observed a strong correlation between AMPAR exocytosis and cFos expression, suggesting a synaptic mechanism for the association of cFos expression with memory engrams. The EPSILON technique is a useful tool for mapping synaptic plasticity and may be extended to investigate trafficking of other transmembrane proteins.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data, including images of dye-stained HT-GluA1-expressing neurons and patch-clamp recordings, are available from the corresponding author upon request.
Code availability
Computer codes for data analysis are available from the corresponding author upon request.
References
Martin, S. J., Grimwood, P. D. & Morris, R. G. Synaptic plasticity and memory: an evaluation of the hypothesis. Annu. Rev. Neurosci. 23, 649–711 (2000).
Wu, Q.-L., Gao, Y., Li, J.-T., Ma, W.-Y. & Chen, N.-H. The role of AMPARs composition and trafficking in synaptic plasticity and diseases. Cell. Mol. Neurobiol. 42, 2489–2504 (2022).
Patterson, M. A., Szatmari, E. M. & Yasuda, R. AMPA receptors are exocytosed in stimulated spines and adjacent dendrites in a Ras-ERK–dependent manner during long-term potentiation. Proc. Natl Acad. Sci. USA 107, 15951–15956 (2010).
Choquet, D. & Opazo, P. The role of AMPAR lateral diffusion in memory. Semin. Cell Dev. Biol. 125, 76–83 (2022).
Zhang, Y., Cudmore, R. H., Lin, D.-T., Linden, D. J. & Huganir, R. L. Visualization of NMDA receptor–dependent AMPA receptor synaptic plasticity in vivo. Nat. Neurosci. 18, 402–407 (2015).
Kim, J. et al. mGRASP enables mapping mammalian synaptic connectivity with light microscopy. Nat. Methods 9, 96–102 (2012).
Choi, J.-H. et al. Interregional synaptic maps among engram cells underlie memory formation. Science 360, 430–435 (2018).
Dore, K. et al. SYNPLA, a method to identify synapses displaying plasticity after learning. Proc. Natl Acad. Sci. USA 117, 3214–3219 (2020).
Getz, A. M. et al. High-resolution imaging and manipulation of endogenous AMPA receptor surface mobility during synaptic plasticity and learning. Sci. Adv. 8, eabm5298 (2022).
Bulovaite, E. et al. A brain atlas of synapse protein lifetime across the mouse lifespan. Neuron 110, 4057–4073 (2022).
Mohar, B. et al. DELTA: a method for brain-wide measurement of synaptic protein turnover reveals localized plasticity during learning. Nat. Neurosci. https://doi.org/10.1038/s41593-025-01923-4 (2025).
Kastellakis, G. & Poirazi, P. Synaptic clustering and memory formation. Front. Mol. Neurosci. 12, 300 (2019).
Watson, J. F., Ho, H. & Greger, I. H. Synaptic transmission and plasticity require AMPA receptor anchoring via its N-terminal domain. eLife 6, e23024 (2017).
Bensussen, S. et al. A viral toolbox of genetically encoded fluorescent synaptic tags. iScience 23, 101330 (2020).
Xie, L. et al. A dynamic interplay of enhancer elements regulates Klf4 expression in naïve pluripotency. Genes Dev. 31, 1795–1808 (2017).
Kleinjan, M. S. et al. Dually innervated dendritic spines develop in the absence of excitatory activity and resist plasticity through tonic inhibitory crosstalk. Neuron 111, 362–371 (2023).
Wong, V. C. et al. Plasticity-induced actin polymerization in the dendritic shaft regulates intracellular AMPA receptor trafficking. eLife 13, e80622 (2024).
Lu, W. et al. Subunit composition of synaptic AMPA receptors revealed by a single-cell genetic approach. Neuron 62, 254–268 (2009).
Dörrbaum, A. R., Kochen, L., Langer, J. D. & Schuman, E. M. Local and global influences on protein turnover in neurons and glia. eLife 7, e34202 (2018).
Trachtenberg, J. T. et al. Long-term in vivo imaging of experience-dependent synaptic plasticity in adult cortex. Nature 420, 788–794 (2002).
Barrientos, S. A. & Tiznado, V. Hippocampal CA1 subregion as a context decoder. J. Neurosci. 36, 6602–6604 (2016).
Ramirez, S. et al. Creating a false memory in the hippocampus. Science 341, 387–391 (2013).
Josselyn, S. A. & Tonegawa, S. Memory engrams: recalling the past and imagining the future. Science 367, eaaw4325 (2020).
Hebb, D. O. The Organization of Behavior: A Neuropsychological Theory (Psychology Press, 2005).
Han, D. H., Park, P., Choi, D. I., Bliss, T. V. & Kaang, B.-K. The essence of the engram: cellular or synaptic? Semin. Cell Dev. Biol. 125, 122–135 (2022).
Curzon, P., Rustay, N. R. & Browman, K. E. in Methods of Behavior Analysis in Neuroscience Ch. 2 (CRC/Taylor & Francis, 2009).
Park, K. & Chung, C. Systemic cellular activation mapping of an extinction-impaired animal model. Front. Cell. Neurosci. 13, 99 (2019).
Magee, J. C. & Grienberger, C. Synaptic plasticity forms and functions. Annu. Rev. Neurosci. 43, 95–117 (2020).
Arai, A., Black, J. & Lynch, G. Origins of the variations in long‐term potentiation between synapses in the basal versus apical dendrites of hippocampal neurons. Hippocampus 4, 1–9 (1994).
Branco, T. & Häusser, M. The single dendritic branch as a fundamental functional unit in the nervous system. Curr. Opin. Neurobiol. 20, 494–502 (2010).
Landau, A. T. et al. Dendritic branch structure compartmentalizes voltage-dependent calcium influx in cortical layer 2/3 pyramidal cells. eLife 11, e76993 (2022).
Makino, H. & Malinow, R. Compartmentalized versus global synaptic plasticity on dendrites controlled by experience. Neuron 72, 1001–1011 (2011).
Bloss, E. B. et al. Single excitatory axons form clustered synapses onto CA1 pyramidal cell dendrites. Nat. Neurosci. 21, 353–363 (2018).
Murakoshi, H., Wang, H. & Yasuda, R. Local, persistent activation of Rho GTPases during plasticity of single dendritic spines. Nature 472, 100–104 (2011).
Routtenberg, A. & Rekart, J. L. Post-translational protein modification as the substrate for long-lasting memory. Trends Neurosci. 28, 12–19 (2005).
Park, P. et al. The role of calcium-permeable AMPARs in long-term potentiation at principal neurons in the rodent hippocampus. Front. Synaptic Neurosci. 10, 42 (2018).
Plitt, M. H., Kaganovsky, K., Südhof, T. C. & Giocomo, L. M. Hippocampal place code plasticity in CA1 requires postsynaptic membrane fusion. Preprint at bioRxiv https://doi.org/10.1101/2023.11.20.567978 (2023).
Shi, S. et al. Syntaxin-3 is dispensable for basal neurotransmission and synaptic plasticity in postsynaptic hippocampal CA1 neurons. Sci. Rep. 10, 709 (2020).
Huang, M. et al. Neuronal SNAP-23 is critical for synaptic plasticity and spatial memory independently of NMDA receptor regulation. iScience 26, 106664 (2023).
Graves, A. R. et al. Visualizing synaptic plasticity in vivo by large-scale imaging of endogenous AMPA receptors. eLife 10, e66809 (2021).
Xu, Y. K. T. et al. Cross-modality supervised image restoration enables nanoscale tracking of synaptic plasticity in living mice. Nat. Methods 20, 935–944 (2023).
Wang, C. et al. Multiplexed aberration measurement for deep tissue imaging in vivo. Nat. Methods 11, 1037–1040 (2014).
Juczewski, K., Koussa, J. A., Kesner, A. J., Lee, J. O. & Lovinger, D. M. Stress and behavioral correlates in the head-fixed method: stress measurements, habituation dynamics, locomotion, and motor-skill learning in mice. Sci. Rep. 10, 12245 (2020).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Tian, H. et al. Video-based pooled screening yields improved far-red genetically encoded voltage indicators. Nat. Methods 20, 1082–1094 (2023).
Lin, D. et al. Time-tagged ticker tapes for intracellular recordings. Nat. Biotechnol. 41, 631–639 (2023).
Manders, E., Verbeek, F. & Aten, J. Measurement of co‐localization of objects in dual‐colour confocal images. J. Microsc. 169, 375–382 (1993).
Goldey, G. J. et al. Removable cranial windows for long-term imaging in awake mice. Nat. Protoc. 9, 2515–2538 (2014).
Adam, Y. et al. Voltage imaging and optogenetics reveal behaviour-dependent changes in hippocampal dynamics. Nature 569, 413–417 (2019).
Holtmaat, A., Wilbrecht, L., Knott, G. W., Welker, E. & Svoboda, K. Experience-dependent and cell-type-specific spine growth in the neocortex. Nature 441, 979–983 (2006).
Seaton, G. et al. Dual-component structural plasticity mediated by αCaMKII autophosphorylation on basal dendrites of cortical layer 2/3 neurones. J. Neurosci. 40, 2228–2245 (2020).
Acknowledgements
We thank A. Preecha and S. Begum for technical assistance. We thank R. Huganir for sharing plasmids. We thank M. Tadross, R. Yasuda, S. Grant, B. Mohar, N. Spruston, K. Svoboda and B. Sabatini for helpful discussions. This work was supported by grants from Schmidt Futures and the Gordon and Betty Moore Foundation and by NIH grant no. 1-R21-EY033669. J.D.W.-C. is a Merck Awardee of the Life Sciences Research Foundation. J.B.G. and L.D.L. are supported by the Howard Hughes Medical Institute. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the paper.
Author information
Authors and Affiliations
Contributions
A.E.C. and D.K. conceived the project and designed the experiments. D.K. conducted the experiments and analyzed the data. P.P. performed acute brain slice patch-clamp measurements. X.L. contributed to molecular cloning. J.D.W.-C., H.T. and E.M.M. provided advice and technical support. L.D.L. and J.B.G. synthesized and provided the dyes used in this study. D.K. and A.E.C. wrote the paper with input from all authors. A.E.C. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 In vitro characterization of EPSILON.
(a) Selective labeling of surface HT-GluA1 with membrane-impermeable HTL dye. Left: Cultured neuron expressing HT-GluA1 stained with membrane-impermeable AF488-HTL while alive. Right: Another HT-GluA1 expressing cultured neuron stained with the same dye after fixation to permeabilize the cell membrane. Scale bars 10 μm. (b) Manders’ overlap coefficient between HT-GluA1 and PSD95 (PSD95.FingR-eGFP). n = 13 dendritic segments from 7 neurons. Error bars show mean ± s.e.m. (c) Confocal images of a fixed cultured neuron expressing (top) JF549-labeled HT-GluA1 stained with (middle) anti-GluA1-CTD (c-terminal domain) antibody. (Bottom) Merge. Scale bar, 1 μm. (d) Relationship between HT-GluA1 intensity and GluA1 antibody intensity among spines. n = 32 spines from 5 neurons. R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. (e) GluA1 expression level in HT-GluA1 +/− neurons stained with anti-GluA1-CTD antibody (n = 11 dendritic segments from 5 neurons for each group). Two-sided Wilcoxon rank-sum test. Error bars show mean ± s.e.m. (f) Confocal images of fixed cultured neurons showing labeling of HT-GluA1 with AF488-HTL (100 nM, cyan) or JF549i-HTL (1 μM, orange) or AF647-HTL (1 μM, red) at different times after dye addition. Scale bars: 100 μm. (g-i) Fluorescence vs. dye incubation time for cultured neurons expressing HT-GluA1 and treated with (g) AF488-HTL (100 nM) or (h) JF549i-HTL (1 μM) or (i) AF647-HTL (1 μM). (n = 5 cells for each timepoint). Data are represented as mean ± SD. Red: fitted curve. (j) Surface GluA1 was saturated by labeling with 100 nM AF488-HTL for 5 minutes, followed immediately by chase-dye labeling (1 μM of JF549i-HTL for 30 s). (k) Confocal images of fixed cultured neurons after saturation with AF488-HTL (left) with and (middle) without JF549i-HTL chase. (Right) JF549i-HTL only without AF488-HTL. Scale bars: 50 μm. (l) JF549i-to-AF488 intensity ratios (n = 16 cells from 3 cultures for Dye 2 +; n = 16 cells from 3 cultures for Dye 2 -; n = 12 cells from 3 cultures for Dye 2 only). Error bars show mean ± s.e.m. Two-sided Student’s t-test. (m) Surface GluA1 was saturated by labeling with 1 μM AF647-HTL for 1 minute, followed immediately by chase-dye labeling (1 μM of JF549i-HTL for 30 s). (n) Confocal images of fixed cultured neurons (left) with and (middle) without JF549i-HTL after saturation with AF647-HTL. (Right) JF549i-HTL only without AF647-HTL. Scale bars: 50 μm. (o) JF549i-to-AF647 intensity ratios (n = 5 cells from 2 cultures for each group). Error bars show mean ± s.e.m. Two-sided Student’s t-test. (p) Experimental timeline of surface HT-GluA1 turnover measurement with multi-color labeling. (q) Confocal images of fixed cultured neurons showing replacement of old (AF488, cyan) with new (JF549i, orange) AMPARs. Scale bars: 200 μm. Turnover occurred faster in soma and perisomatic neurites than in distal neurites. (r) Normalized dye intensity ratio at 5, 20, 30, 60, and 120-min (n = 5 cells for each timepoint). Normalized dye intensity ratio: 2nd dye intensity / (1st dye intensity + 2nd dye intensity). Data are represented as mean ± SD. Schematics in j, m and p created using BioRender.com.
Extended Data Fig. 2 Effects of MNI-glutamate uncaging and cLTP on AMPAR exocytosis.
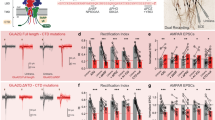
(a) Time-lapse images of the stimulated spines expressing GPI-eGFP +/− HT-GluA1 expression. Scale bars, 10 μm. (b) Time course for spine size increases +/− HT-GluA1 expression (HT-GluA1 (+): n = 13 spines from 3 neurons; HT-GluA1 (−): n = 14 spines from 3 neurons). Error bars show mean ± s.e.m. (c) Confocal images of spines expressing GPI-eGFP (green) and HT-GluA1 labeled with Dye 1 (cyan, AF647) and Dye 2 (orange, JF549i), before and after glutamate photo-uncaging. Controls had no glutamate uncaging. Scale bars, 1 μm. (d) Dye 1 intensity change for control and uncaged spine groups. Control: n = 10 spines; Uncaged: n = 20 spines. Two-sided Wilcoxon rank-sum test. Error bars show mean ± s.e.m. (e) Correlation between the change in Dye 1 intensity and the change in spine size for spines from (d). R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. (f) Scatterplots of spine fluorescence intensities in the two color channels for (left) control, (middle) cTLP-treated, and (right) TeTX-LC co-expressing neurons. Same data as in Fig. 1m, with vertical scale adjusted to include all spines. Control: n = 32,828 spines from 6 cultures; cLTP: n = 47,090 spines from 6 cultures; TeTX-LC: n = 17,231 spines from 6 cultures. Dye 2 intensity thresholds indicated with dashed lines (see Methods for calculation of threshold). Replicate dishes represented by different shape symbols. (g) The extent of AMPAR exocytosis in each spine measured by the fluorescence of Dye 2 above threshold. Spines with Dye 2 intensity higher than the threshold were included for analysis (control: 1,563 ± 112 counts, mean ± s.e.m., n = 195 spines, 54 neurons, 6 cultures, cLTP: 2,398 ± 74.9 counts, mean ± s.e.m., n = 1,024 spines, 44 neurons, 6 cultures; TeTX-LC: 868.7 ± 147 counts, mean ± s.e.m., n = 35 spines, 45 neurons, 6 cultures). Replicate dishes represented by different symbols. Box plot shows extrema, 25th and 75th percentiles and median. Two-sided Wilcoxon rank-sum test.
Extended Data Fig. 3 Validation of EPSILON in mouse brain.
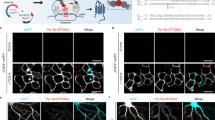
HT-GluA1 stained with AF647-HTL in (a) barrel cortex (b) CA1 pyramidal cells. Each set of micrographs represents a single experiment. Scale bars 50 μm. (c) Postsynaptic trafficking of HT-GluA1 in layer 2/3 pyramidal neurons in barrel cortex. (Left) HT-GluA1 stained with JF549i-HTL. (Middle) PSD95 (PSD95.FingR-eGFP) from same region of interest. (Right) Merge. Scale bars, 5 μm. (d), (e) Dendritic segments expressing (d) HT-GluA1 only or (E) PSD95.FingR-eGFP only. Scale bars, 2 μm. (f) Manders’ overlap coefficient between HT-GluA1 and PSD95 (PSD95.FingR-eGFP). n = 9 dendritic segments from 5 neurons. Error bars show mean ± s.e.m. (g) Correlation between PSD95.FingR-eGFP intensity and HT-GluA1 intensity on individual spines (n = 112 spines from 8 dendritic segments from 8 neurons). R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. HT-GluA1 expressing layer 2/3 pyramidal neurons in barrel cortex in acute brain slices were identified by staining with 1uM JFX608-HTL. (h) Representative patch-clamp recordings in acute brain slice. (i) Measurements of electrophysiological properties of neurons with or without HT-GluA1 expression. Membrane resistance: 204 ± 16 MΩ vs. 212 ± 12 MΩ, P = 0.69; membrane capacitance: 47 ± 5 pF vs. 43 ± 3 pF, P = 0.50; resting potential: −68.1 ± 1.4 mV vs. −69.2 ± 1.1 mV, P = 0.53; and rheobase 97 ± 9.8 pA vs. 94 ± 15 pA, P = 0.86 (n = 13 neurons for each group). Error bars show mean ± s.e.m. n.s. not significant, two-sided Student’s t-test. Spine density on dendritic segments from neurons +/− HT-GluA1 in (j) layer 2/3 pyramidal neurons in barrel cortex and (k) pyramidal neurons in CA1 (HT-GluA1 (+) in barrel cortex: n = 11 dendritic segments from 10 neurons; HT-GluA1 (−) in barrel cortex: n = 11 dendritic segments from 11 neurons; HT-GluA1 (+) in CA1: n = 8 dendritic segments from 5 neurons; HT-GluA1 (−) in CA1: n = 8 dendritic segments from 4 neurons). Two-sided Wilcoxon rank-sum test. Error bars show mean ± s.e.m. (l) Confocal images of HT-GluA1 stained with JF549-HTL (orange), co-expressed GPI-eGFP (green), and immunostained for GluA1 (red). Selected regions for quantifying GluA1 expression levels in neurons with HT-GluA1 (red) or without HT-GluA1 (yellow) are indicated on the GPI-eGFP panels. HT-GluA1 (+) and (−) regions were selected to be within 100 μm of each other, considering possible heterogenous immunostaining. Scale bars, 10 μm. (m) GluA1 expression level in HT-GluA1 +/− neurons (n = 10 neurons from 3 animals for each group). Two-sided Wilcoxon rank-sum test. Error bars show mean ± s.e.m.
Extended Data Fig. 4 HaloTag dye labeling in live mouse brain.
(a) Relative coordinates from bregma and lambda for intracortical dye injections to cover layer 2/3 pyramidal neurons in left barrel cortex. (b) Relative coordinates from bregma and lambda for intrahippocampal dye injections to cover CA1 pyramidal neurons in left hippocampus. (c) Immunostaining of the astrocyte marker GFAP after dye injections. (Top) GFAP expression in the (top left) surgery hemisphere, (top middle) intentionally injured hemisphere, and (top right) control hemisphere without dye injection. (Bottom) HT-GluA1 stained with AF647-HTL in the same regions of interest as the upper panels. 1 μM of AF647-HTL was injected at three different sites as shown in (a) (80 nL at each depth) three times (20 minutes between the 1st and 2nd injections; 90 minutes between the 2nd and 3rd injections). Some of the injection paths are indicated by white dashed lines on the GFAP panels. These injections were performed 7 days after the installation of the injection port (see Methods). Each set of micrographs represents a single experiment from at least three repeated experiments. Scale bar, 1 mm. (d) Saturated labeling of surface GluA1 with Dye 1 blocked chase-dye labeling with Dye 2. Surface GluA1 was saturated by labeling with (left) 1 μM JF549i-HTL for 20 minutes in layer 2/3 barrel cortex, (middle) 1 μM AF647-HTL for 20 minutes in layer 2/3 barrel cortex, (right) 1 μM AF647-HTL for 20 minutes in CA1. After Dye 1 labeling, 10 μM of Dye 2 was immediately injected to the same injection sites (Methods). Confocal images show Dye 1 (green) and Dye 2 (magenta) labeled dendrites (top) with Dye 2, (middle) without Dye 2, and (bottom) Dye 2 only. Scale bars, 5 μm. (e) Dye 2-to-Dye 1 intensity ratios for the injection conditions from (d). n = 21 dendritic segments from 3 neurons, 20 dendritic segments from 3 neurons, 15 dendritic segments from 3 neurons for Dye 2 (+), Dye 2 (−), Dye 1 only groups in JF549i/AF488 in barrel cortex, respectively; n = 21 dendritic segments from 3 neurons for each group in AF647/JF549i in barrel cortex; n = 21 dendritic segments from 3 neurons for each group in CA1. Error bars show mean ± s.e.m. Two-sided Student’s t-test. (f) Background fluorescence analysis in mouse barrel cortex layer 2/3. Left: Spherical regions of interest (yellow) used for background fluorescence measurement on top of Fig. 2g. Right: Scatterplots of spine (blue) and background (red) fluorescence intensities (n = 53,457 spines and 747 background regions). (g) Background fluorescence analysis in mouse CA1. Left: Spherical regions of interest (yellow) used for background fluorescence measurement on top of Fig. 3c. Right: Scatterplots of spine (blue) and background (red) fluorescence intensities (n = 5,314 spines and 2,836 background regions). Scale bars: 200 μm. Schematics in a and b created using BioRender.com.
Extended Data Fig. 5 Validation of contextual fear conditioning protocol.
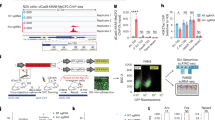
(a) The CFC system was validated by measuring the percent of time mouse spent freezing (that is immobile) 24 hours after conditioning. The percentage of freezing was measured for the mice that underwent full conditioning (shock) and for the mice exposed to the identical context but that did not receive an aversive stimulus (no shock). Error bars represent mean ± s.e.m. (n = 5 mice for each group). Two-sided Wilcoxon rank-sum test. Representative images from CA1 from mice that (b) were not exposed to any context (that is euthanized after 3 hours of anesthesia), (c) were not exposed to the novel context (stayed in their home cages, home cage control), (d) were exposed to the novel context but did not receive an aversive foot shock (context-only), and (e) underwent CFC. The images are shown in the same contrast scale and are maximum-intensity projections of z-stacks. These images are representative of the dataset used in Fig. 3m. Scale bars: 200 μm.
Extended Data Fig. 6 Relationships among cFos expression, AMPAR exocytosis levels, and percentage of potentiated spines.
(a) Relation of the percentage of potentiated spines to the mean HT-GluA1 exocytosis among potentiated spines (distance of Dye 2 signal above threshold, averaged over above-threshold spines), for (left) CFC and (right) context-only control. (b) Relation between the mean HT-GluA1 exocytosis among potentiated spines and the corresponding cFos intensity. CFC: n = 19 cells from 4 animals; context-only: n = 22 cells from 6 animals. R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. Distinct animals represented by different shape symbols. (c) Relation between the percentage of potentiated spine and the z-coordinate of the center of soma of the corresponding neuron for (left) CFC, (middle) context-only control, and (right) home cage control. (d) Relation between cFos intensity and the z-coordinate of the center of soma of the corresponding neuron for (left) CFC, (middle) context-only control, and (right) home cage control. (e) Relation between the total number of identified spines and the z-coordinate of the center of soma of the corresponding neuron for (left) CFC, (middle) context-only control, and (right) home cage control. CFC: n = 19 cells from 4 animals; context-only: n = 23 cells from 6 animals; home cage control: n = 12 cells from 3 animals. R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. Distinct animals represented by different shape symbols. (f) Relation between the total number of identified spines and the cFos intensity of the corresponding neuron for (left) CFC, (middle) context-only control, and (right) home cage control. CFC: n = 19 cells from 4 animals; context-only: n = 23 cells from 6 animals; home cage control: n = 12 cells from 3 animals. R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. Distinct animals represented by different shape symbols.
Extended Data Fig. 7 Subcellular distribution of potentiated spines.
(a) Representative neuron with dendrites segmented and colored by their branch order. (b) Same neuron with spines registered to the nearest segmented dendrites. Scale bars: 100 μm. (c) Distribution of potentiated spines as a function of projected distance from the stratum pyramidale. (Top) Overlapped images of all identified spines (gray) and spines with high HT-GluA1 exocytosis (red) from the context-only control group. (Bottom) Total number of identified spines (grey), number of spines with high HT-GluA1 exocytosis (red), and fraction of potentiated spines (blue) plotted against the projected distance from the stratum pyramidale (x-axis from the top panel). Error bars represent count ± square root(count). (d) Schematic diagram showing the contour distance of a neuron’s spine from its soma. This is the distance along the dendrite backbone. (e, f) Same data as in Fig. 4a and (c), but plotted vs. contour distance instead of x. (g) Schematic drawing of a neuron with dendrites numbered by their branch order. (h), (i) Total number of identified spines (left), number of potentiated spines (middle), fraction of potentiated spines (right) vs. corresponding dendrites’ branch order from (top) apical dendrites or (bottom) basal dendrites in (h) CFC group and (i) context-only controls. CFC: n = 19 neurons; context-only: n = 23 neurons. Data are represented as mean ± s.e.m. The right panels are truncated due to the division by small numbers of spines on branch orders greater than 10. Schematics in d and g created using BioRender.com.
Extended Data Fig. 8 Comparison of AMPAR exocytosis in basal vs. apical dendrites.
Fraction of potentiated spines for each neuron vs. cFos intensity for (a) CFC (n = 19 neurons) and (b) context-only control (n = 23 neurons). Spine fraction was separately evaluated for basal and apical dendrites. (c) Normalized basal to apical ratio of percent potentiated spines. Normalized ratio: basal percent potentiated spines / (basal percent potentiated spines + apical percent potentiated spines). 13 of 19 neurons in the CFC group and 17 of 23 neurons in the context-only group had higher basal than apical fraction of potentiated spines. There was no significant difference between the CFC and context-only groups in the ratio of basal to apical potentiated spines (CFC: 0.61 ± 0.06, mean ± s.e.m.; context-only: 0.70 ± 0.07, mean ± s.e.m.). Two-sided Wilcoxon rank-sum test. (d) Basal to apical ratio of percent potentiated spines vs. cFos intensity. The basal to apical ratio was not correlated with the corresponding cFos levels. R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. Distinct mice represented by different shape symbols. Mean contour distance from soma to potentiated spines vs. corresponding cFos intensity for (e) CFC group and (f) context-only control. The mean distance was separately evaluated for spines in basal and apical dendrites. R, Pearson’s linear correlation coefficient, P value from two-sided Student’s t-test. Distinct mice represented by different shape symbols. (g) Mean contour distance from soma to potentiated spines in CFC and context-only group. Data are represented as mean ± s.e.m. Two-sided Wilcoxon rank-sum test. CFC: n = 19 neurons; context-only: n = 22 neurons.
Extended Data Fig. 9 Clustering of potentiated spines in context-only control.
(a) Density profile of potentiated spines as a function of distance from the nearest potentiated spine. The single-cell profiles are plotted with light colors. Random: simulation where the same number of potentiated spines are distributed randomly and independently among all detected spines. Bold and dashed lines represent mean ± s.e.m. (b) Fraction of potentiated spine clusters of different sizes from the context-only group (n = 22 neurons). The single-cell profiles are plotted with light colors. Bold and dashed lines represent mean ± s.e.m. Random defined as in (a).
Extended Data Fig. 10 Possible extensions of the EPSILON technique.
(a,b) Monitoring spine dynamics in vivo. (a) Experimental setup for intracortical injection of membrane-impermeable AF488-HTL dye into mouse barrel cortex expressing HT-GluA1, followed by in vivo 2-photon imaging. (b) Representative in vivo 2-photon images of layer 2/3 pyramidal neuron apical tuft dendrites and spines stained with AF488-HTL. This set of micrographs represents a single experiment. Scale bars, 5 μm. (c) Detecting new spine formation. New spines are expected to be labeled only with Dye 2, not Dye 1. Locations of the spines labeled with Dye 2 only are indicated with red dots. Neuron from Fig. 3c. This micrograph represents a single experiment. See Methods for the detection of Dye 2 only spines. Scale bar, 200 μm. Schematic in a created using BioRender.com.
Supplementary information
Supplementary Information (download PDF )
Image processing and data analysis.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, D., Park, P., Li, X. et al. EPSILON: a method for pulse-chase labeling to probe synaptic AMPAR exocytosis during memory formation. Nat Neurosci 28, 1099–1107 (2025). https://doi.org/10.1038/s41593-025-01922-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-025-01922-5