Abstract
Neurons of distinct subtypes compartmentalize subtype-specific function in part by differentially localizing and translating specific RNAs, but underlying mechanisms are not understood. Here we investigate messenger RNA localization and stability within subtype-specific growth cones (GCs), leading tips of growing axons, of long-range projection neurons (PNs) of the developing cerebral cortex. Comparison of GC-localized transcriptomes between two subtypes of PNs (interhemispheric-callosal and corticothalamic) across developmental stages identified both distinct and shared subcellular machinery involved in distinct phases of growth, target innervation and synaptogenesis, and enrichment of genes associated with neurodevelopmental and neuropsychiatric disorders. Further, we investigated sequence elements in dynamically GC-localized mRNAs, identifying GC-enriched motifs in 3′ untranslated regions. For example, we identified that CPEB4, a translational regulator, regulates axonal branching and that RBMS1 functions dynamically in callosal circuit formation. This work offers generalizable insights for subcellular specialization in other polarized cells, toward elucidating neurodevelopmental and behavioral-cognitive disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
RNA-sequencing data and analysis results are available in Supplementary Tables 1–4. Source data are provided with this paper.
Code availability
The full RNA-sequencing data processing code is available at https://github.com/priyaveeraraghavan/amalgam.
References
Stoeckli, E. T. Understanding axon guidance: are we nearly there yet? Development 145, dev151415 (2018).
Maday, S., Twelvetrees, A. E., Moughamian, A. J. & Holzbaur, E. L. F. Axonal transport: cargo-specific mechanisms of motility and regulation. Neuron 84, 292–309 (2014).
Holt, C. E., Martin, K. C. & Schuman, E. M. Local translation in neurons: visualization and function. Nat. Struct. Mol. Biol. 26, 557–566 (2019).
Davis, L., Dou, P., DeWit, M. & Kater, S. B. Protein synthesis within neuronal growth cones. J. Neurosci. 12, 4867–4877 (1992).
Ming, G. et al. Adaptation in the chemotactic guidance of nerve growth cones. Nature 417, 411–418 (2002).
Lyles, V., Zhao, Y. & Martin, K. C. Synapse formation and mRNA localization in cultured aplysia. Neuron 49, 349–356 (2006).
Andreassi, C. et al. An NGF-responsive element targets myo-inositol monophosphatase-1 mRNA to sympathetic neuron axons. Nat. Neurosci. 13, 291–301 (2010).
Cox, L. J., Hengst, U., Gurskaya, N., Lukyanov, K. A. & Jaffrey, S. R. Intra-axonal translation and retrograde trafficking of CREB promotes neuronal survival. Nat. Cell Biol. 10, 149–159 (2008).
Aakalu, G., Smith, W. B., Nguyen, N., Jiang, C. & Schuman, E. M. Dynamic visualization of local protein synthesis in hippocampal. Neuron 30, 489–502 (2001).
Kang, H. & Schuman, E. M. A requirement for local protein synthesis in neurotrophin-induced hippocampal synaptic plasticity. Science 273, 1402–1406 (1996).
Ben-Yaakov, K. et al. Axonal transcription factors signal retrogradely in lesioned peripheral nerve. EMBO J. 31, 1350–1363 (2012).
Donnelly, C. J. et al. Limited availability of ZBP1 restricts axonal mRNA localization and nerve regeneration capacity. EMBO J. 30, 4665–4677 (2011).
Hanz, S. et al. Axoplasmic importins enable retrograde injury signaling in lesioned nerve. Neuron 40, 1095–1104 (2003).
Perlson, E. et al. Vimentin-dependent spatial translocation of an activated MAP kinase in injured nerve. Neuron 45, 715–726 (2005).
Verma, P. et al. Axonal protein synthesis and degradation are necessary for efficient growth cone regeneration. J. Neurosci. 25, 331–342 (2005).
Yudin, D. et al. Localized regulation of axonal RanGTPase controls retrograde injury signaling in peripheral nerve. Neuron 59, 241–252 (2008).
Rotem, N. et al. ALS along the axons—expression of coding and noncoding RNA differs in axons of ALS models. Sci. Rep. 7, 44500 (2017).
Zappulo, A. et al. RNA localization is a key determinant of neurite-enriched proteome. Nat. Commun. 8, 583 (2017).
Biever, A. et al. Monosomes actively translate synaptic mRNAs in neuronal processes. Science 367, eaay4991 (2020).
Glock, C. et al. The translatome of neuronal cell bodies, dendrites, and axons. Proc. Natl Acad. Sci. USA 118, e2113929118 (2021).
Hafner, A.-S., Donlin-Asp, P. G., Leitch, B., Herzog, E. & Schuman, E. M. Local protein synthesis is a ubiquitous feature of neuronal pre- and postsynaptic compartments. Science 364, eaau3644 (2019).
Ostroff, L. E. et al. Axon TRAP reveals learning-associated alterations in cortical axonal mRNAs in the lateral amygdala. eLife 8, e51607 (2019).
Zivraj, K. H. et al. Subcellular profiling reveals distinct and developmentally regulated repertoire of growth cone mRNAs. J. Neurosci. 30, 15464–15478 (2010).
Engmann, A. K. et al. Neuronal subtype-specific growth cone and soma purification from mouse and other mammalian CNS via fractionation and fluorescent sorting for subcellular analyses and spatial mapping of local transcriptomes and proteomes. Nat. Protoc. 17, 222–251 (2022).
Poulopoulos, A. et al. Subcellular transcriptomes and proteomes of developing axon projections in cerebral cortex. Nature 565, 356–360 (2019).
De León Reyes, N. S., Bragg-Gonzalo, L. & Nieto, M. Development and plasticity of the corpus callosum. Development 147, dev189738 (2020).
Fenlon, L. R., Suárez, R. & Richards, L. J. The anatomy, organisation and development of contralateral callosal projections of the mouse somatosensory cortex. Brain Neurosci. Adv. 1, 2398212817694888 (2017).
Greig, L. C., Woodworth, M. B., Greppi, C. & Macklis, J. D. Ctip1 controls acquisition of sensory area identity and establishment of sensory input fields in the developing neocortex. Neuron 90, 261–277 (2016).
Leyva-Díaz, E. & López-Bendito, G. In and out from the cortex: development of major forebrain connections. Neuroscience 254, 26–44 (2013).
Greig, L. C., Woodworth, M. B., Galazo, M. J., Padmanabhan, H. & Macklis, J. D. Molecular logic of neocortical projection neuron specification, development and diversity. Nat. Rev. Neurosci. 14, 755–769 (2013).
Paolino, A., Fenlon, L. R., Suárez, R. & Richards, L. J. Transcriptional control of long-range cortical projections. Curr. Opin. Neurobiol. 53, 57–65 (2018).
Paul, L. K., Corsello, C., Kennedy, D. P. & Adolphs, R. Agenesis of the corpus callosum and autism: a comprehensive comparison. Brain 137, 1813–1829 (2014).
Dienel, S. J., Schoonover, K. E. & Lewis, D. A. Cognitive dysfunction and prefrontal cortical circuit alterations in schizophrenia: developmental trajectories. Biol. Psychiatry 92, 450–459 (2022).
Fame, R. M., MacDonald, J. L. & Macklis, J. D. Development, specification, and diversity of callosal projection neurons. Trends Neurosci. 34, 41–50 (2011).
Galazo, M. J., Emsley, J. G. & Macklis, J. D. Corticothalamic projection neuron development beyond subtype specification: Fog2 and intersectional controls regulate intraclass neuronal diversity. Neuron 91, 90–106 (2016).
Galazo, M. J., Sweetser, D. A. & Macklis, J. D. Tle4 controls both developmental acquisition and early post-natal maturation of corticothalamic projection neuron identity. Cell Rep. 42, 112957 (2023).
Satterstrom, F. K. et al. Large-scale exome sequencing study implicates both developmental and functional changes in the neurobiology of autism. Cell 180, 568–584 (2020).
Banerjee-Basu, S. & Packer, A. SFARI Gene: an evolving database for the autism research community. Dis. Model. Mech. 3, 133–135 (2010).
Singh, T. et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature 604, 509–516 (2022).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Rottner, K., Stradal, T. E. B. & Chen, B. WAVE regulatory complex. Curr. Biol. 31, R512–R517 (2021).
Estrada-Bernal, A. et al. Functional complexity of the axonal growth cone: a proteomic analysis. PLoS ONE 7, e31858 (2012).
Nozumi, M. et al. Identification of functional marker proteins in the mammalian growth cone. Proc. Natl Acad. Sci. USA 106, 17211–17216 (2009).
Wu, Y. et al. Multi-trait analysis for genome-wide association study of five psychiatric disorders. Transl. Psychiatry 10, 209 (2020).
Sato, D. et al. SHANK1 deletions in males with autism spectrum disorder. Am. J. Hum. Genet. 90, 879–887 (2012).
Hake, L. E. & Richter, J. D. CPEB is a specificity factor that mediates cytoplasmic polyadenylation during Xenopus oocyte maturation. Cell 79, 617–627 (1994).
Piqué, M., López, J. M., Foissac, S., Guigó, R. & Méndez, R. A combinatorial code for CPE-mediated translational control. Cell 132, 434–448 (2008).
Wu, L. et al. CPEB-mediated cytoplasmic polyadenylation and the regulation of experience-dependent translation of α-CaMKII mRNA at synapses. Neuron 21, 1129–1139 (1998).
Poetz, F. et al. Control of immediate early gene expression by CPEB4-repressor complex-mediated mRNA degradation. Genome Biol. 23, 193 (2022).
Suñer, C. et al. Macrophage inflammation resolution requires CPEB4-directed offsetting of mRNA degradation. eLife 11, e75873 (2022).
Parras, A. et al. Autism-like phenotype and risk gene mRNA deadenylation by CPEB4 mis-splicing. Nature 560, 441–446 (2018).
Huang, Y.-S., Carson, J. H., Barbarese, E. & Richter, J. D. Facilitation of dendritic mRNA transport by CPEB. Genes Dev. 17, 638–653 (2003).
Mendonsa, S. et al. Massively parallel identification of mRNA localization elements in primary cortical neurons. Nat. Neurosci. 26, 394–405 (2023).
Kozlov, E., Shidlovskii, Y. V., Gilmutdinov, R., Schedl, P. & Zhukova, M. The role of CPEB family proteins in the nervous system function in the norm and pathology. Cell Biosci. 11, 64 (2021).
Stebbins-Boaz, B., Cao, Q., de Moor, C. H., Mendez, R. & Richter, J. D. Maskin is a CPEB-associated factor that transiently interacts with elF-4E. Mol. Cell 4, 1017–1027 (1999).
Kvainickas, A. et al. Cargo-selective SNX-BAR proteins mediate retromer trimer independent retrograde transport. J. Cell Biol. 216, 3677–3693 (2017).
Südhof, T. C. et al. Synapsins: mosaics of shared and individual domains in a family of synaptic vesicle phosphoproteins. Science 245, 1474–1480 (1989).
Ushkaryov, Y. A. & Südhof, T. C. Neurexin III α: extensive alternative splicing generates membrane-bound and soluble forms. Proc. Natl Acad. Sci. USA 90, 6410–6414 (1993).
Xie, X. et al. C2 domain-containing phosphoprotein CDP138 regulates GLUT4 insertion into the plasma membrane. Cell Metab. 14, 378–389 (2011).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Habib, K., Bishayee, K., Kang, J., Sadra, A. & Huh, S.-O. RNA binding protein RBMS1 enables neuronal differentiation and radial migration during neocortical development by binding and stabilizing the RNA message for Efr3a. Mol. Cells 45, 588–602 (2022).
Dankert, J. T., Wiesehöfer, M., Wach, S., Czyrnik, E. D. & Wennemuth, G. Loss of RBMS1 as a regulatory target of miR-106b influences cell growth, gap closing and colony forming in prostate carcinoma. Sci. Rep. 10, 18022 (2020).
Liu, M. et al. RBMS1 promotes gastric cancer metastasis through autocrine IL-6/JAK2/STAT3 signaling. Cell Death Dis. 13, 287 (2022).
Yu, J. et al. RBMS1 suppresses colon cancer metastasis through targeted stabilization of its mRNA regulon. Cancer Discov. 10, 1410–1423 (2020).
Mitchell, B. D. & Macklis, J. D. Large-scale maintenance of dual projections by callosal and frontal cortical projection neurons in adult mice. J. Comp. Neurol. 482, 17–32 (2005).
Sohur, U. S., Padmanabhan, H. K., Kotchetkov, I. S., Menezes, J. R. L. & Macklis, J. D. Anatomic and molecular development of corticostriatal projection neurons in mice. Cereb. Cortex 24, 293–303 (2014).
Cederquist, G. Y., Azim, E., Shnider, S. J., Padmanabhan, H. & Macklis, J. D. LMO4 establishes rostral motor cortex projection neuron subtype diversity. J. Neurosci. 33, 6321–6332 (2013).
Shigeoka, T. et al. Dynamic axonal translation in developing and mature visual circuits. Cell 166, 181–192 (2016).
Ayanlaja, A. A. et al. Distinct features of doublecortin as a marker of neuronal migration and its implications in cancer cell mobility. Front. Mol. Neurosci. 10, 199 (2017).
Arora, A. et al. High-throughput identification of RNA localization elements in neuronal cells. Nucleic Acids Res. 50, 10626–10642 (2022).
Dalla Costa, I. et al. The functional organization of axonal mRNA transport and translation. Nat. Rev. Neurosci. 22, 77–91 (2021).
Kislauskis, E. H., Zhu, X. & Singer, R. H. Sequences responsible for intracellular localization of β-actin messenger RNA also affect cell phenotype. J. Cell Biol. 127, 441–451 (1994).
Ross, A. F., Oleynikov, Y., Kislauskis, E. H., Taneja, K. L. & Singer, R. H. Characterization of a β-actin mRNA zipcode-binding protein. Mol. Cell. Biol. 17, 2158–2165 (1997).
Colak, D., Ji, S.-J., Porse, B. T. & Jaffrey, S. R. Regulation of axon guidance by compartmentalized nonsense-mediated mRNA decay. Cell 153, 1252–1265 (2013).
Loedige, I. et al. mRNA stability and m6A are major determinants of subcellular mRNA localization in neurons. Mol. Cell 83, 2709–2725 (2023).
Gardiner, A. S., Twiss, J. L. & Perrone-Bizzozero, N. I. Competing interactions of RNA-binding proteins, microRNAs, and their targets control neuronal development and function. Biomolecules 5, 2903–2918 (2015).
Veeraraghavan, P., Tillman, D. E., Ross, M. I. & Macklis, J. D. Developmental dynamics and axonal transport of cerebral cortex projection neuron mRNAs in vivo. Preprint at bioRxiv https://doi.org/10.1101/2025.04.23.650263 (2025).
Hilgers, V. et al. Neural-specific elongation of 3′ UTRs during Drosophila development. Proc. Natl Acad. Sci. USA 108, 15864–15869 (2011).
Miura, P., Shenker, S., Andreu-Agullo, C., Westholm, J. O. & Lai, E. C. Widespread and extensive lengthening of 3′ UTRs in the mammalian brain. Genome Res. 23, 812–825 (2013).
Balasanyan, V. & Arnold, D. B. Actin and myosin-dependent localization of mRNA to dendrites. PLoS ONE 9, e92349 (2014).
Blichenberg, A. et al. Identification of a cis-acting dendritic targeting element in MAP2 mRNAs. J. Neurosci. 19, 8818–8829 (1999).
Caceres, A., Banker, G., Steward, O., Binder, L. & Payne, M. MAP2 is localized to the dendrites of hippocampal neurons which develop in culture. Brain Res. 315, 314–318 (1984).
Chen, J., Kanai, Y., Cowan, N. J. & Hirokawa, N. Projection domains of MAP2 and tau determine spacings between microtubules in dendrites and axons. Nature 360, 674–677 (1992).
Dehmelt, L. & Halpain, S. The MAP2/tau family of microtubule-associated proteins. Genome Biol. 6, 204 (2005).
Garner, C. C., Tucker, R. P. & Matus, A. Selective localization of messenger RNA for cytoskeletal protein MAP2 in dendrites. Nature 336, 674–677 (1988).
Matus, A. Microtubule-associated proteins and neuronal morphogenesis. J. Cell Sci. Suppl. 15, 61–67 (1991).
Park, H. Y., Trcek, T., Wells, A. L., Chao, J. A. & Singer, R. H. An unbiased analysis method to quantify mRNA localization reveals its correlation with cell motility. Cell Rep. 1, 179–184 (2012).
Przyborski, S. A. & Cambray-Deakin, M. A. Developmental regulation of MAP2 variants during neuronal differentiation in vitro. Brain Res. Dev. Brain Res. 89, 187–201 (1995).
Turner-Bridger, B., Caterino, C. & Cioni, J.-M. Molecular mechanisms behind mRNA localization in axons. Open Biol. 10, 200177 (2020).
Saito, T. & Nakatsuji, N. Efficient gene transfer into the embryonic mouse brain using in vivo electroporation. Dev. Biol. 240, 237–246 (2001).
Pfenninger, K. H., Ellis, L., Johnson, M. P., Friedman, L. B. & Somlo, S. Nerve growth cones isolated from fetal rat brain: subcellular fractionation and characterization. Cell 35, 573–584 (1983).
Catapano, L. A., Arnold, M. W., Perez, F. A. & Macklis, J. D. Specific neurotrophic factors support the survival of cortical projection neurons at distinct stages of development. J. Neurosci. 21, 8863–8872 (2001).
Palmer, D. S. et al. Exome sequencing in bipolar disorder identifies AKAP11 as a risk gene shared with schizophrenia. Nat. Genet. 54, 541–547 (2022).
Chen, B. et al. The WAVE regulatory complex links diverse receptors to the actin cytoskeleton. Cell 156, 195–207 (2014).
Bailey, T. L. STREME: accurate and versatile sequence motif discovery. Bioinformatics 37, 2834–2840 (2021).
Krismer, K. et al. Transite: a computational motif-based analysis platform that identifies RNA-binding proteins modulating changes in gene expression. Cell Rep. 32, 108064 (2020).
Arshadi, C., Günther, U., Eddison, M., Harrington, K. I. S. & Ferreira, T. A. SNT: a unifying toolbox for quantification of neuronal anatomy. Nat. Methods 18, 374–377 (2021).
Acknowledgements
We thank J. LaVecchio and N. Kheradmand of the HSCRB-HSCI Flow Cytometry Core, the Bauer RNA Sequencing Core and Harvard Center for Biological Imaging for infrastructure and support, G. D. Wheeler for technical support and T. Sackton for statistical advice. This work was supported by grants to J.D.M. (Allen Distinguished Investigator Award ADI 11855 from the Paul G. Allen Frontiers Group, NIH Pioneer Award DP1 NS106665 and the Max and Anne Wien Professor of Life Sciences fund), with additional infrastructure support from NIH grants NS045523 and NS104055. A.K.E. was supported by a Jean-Jacques et Felicia Lopez-Loreta Foundation Award and a Swiss National Science Foundation Postdoctoral Fellowship. P.V. was partially supported by National Science Foundation predoctoral fellowship GRFP 280932, the NSF-Simons Center for Mathematical and Statistical Analysis of Biology at Harvard 1764269 and NIH Training Grant T32 GM007306-43. J.J.H. was partially supported by NIH NRSA F31 NS103262 and NIH Training Grant T32 GM007226. We dedicate this paper to the memory of John J. Hatch, an exceptional and boundlessly curious friend, student and colleague.
Author information
Authors and Affiliations
Contributions
J.J.H. and J.D.M. initiated the study. J.J.H. and A.K.E. constructed mouse lines. T.A. and P.V. designed constructs. P.V. and J.J.H. performed in utero electroporations. A.K.E. and J.J.H. performed GC preps and GC and soma sorts. Y.I. performed soma preps. P.V., A.K.E. and D.N. designed and executed cellular and histological validation studies. P.V. performed all bioinformatic analyses. A.K.E., P.V. and J.D.M. interpreted the data and iteratively designed further analyses and experiments. A.K.E., P.V. and J.D.M. drafted the paper. P.V., A.K.E. and J.D.M. edited the paper with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Subtype-specific labeling of CPN and CThPN.
(a) In utero electroporation at embryonic day E15 in mice specifically labels CPN of cortical layer II/III with a multi-cistronic DNA construct encoding membrane-targeted tdTomato (for GCs) and nuclear-localized GFP (for somata). GFP-positive nuclei of CPN are localized mainly to cortical layer II, with limited labeling present in layer III, and are colocalized with the superficial layer marker SatB2 (inset). The membrane-targeted tdTomato reveals the entire extent of CPN dendritic processes and axonal projections spanning all cortical layers, as well as across the corpus callosum. (b) Unilateral in utero electroporation at embryonic day E15 results in highly reproducible targeting of CPN in cortical layer II/III in the lateral somatosensory cortex. (c) Total number of labeled CPN in cortical layers II/III in a single 40 μm coronal brain section is comparable across mice, N = 4. Only very few labeled cells were detected in cortical layer IV. (d) Intersectional mouse genetics (Ntsr1::Cre+/-, Emx1::FlpO+/-, Ai65(TdTom)FRT/wt, flox/wt) specifically labels CThPN of cortical layer VI. Labeled CThPN projections transverse the internal capsule and innervate the thalamus. TdTom-positive CThPN are colocalized with CThPN marker TBR1 but are negative for superficial layer marker SatB2 (inset). (e) RNA expression levels (transcripts per million (TPM) visualized on log2 scale) of subtype marker genes. Data depicted was obtained by RNA sequencing of flow cytometry sorted CPN and CThPN somata at developmental ages P1 and P3. CPN have markedly higher expression levels in superficial layer markers28,30,31,34, whereas CThPN have appropriate enrichment for layer VI marker genes, as expected35,36. Genes common to both subtypes are expressed at similar levels.
Extended Data Fig. 2 Parallel isolation of subtype-specific somata and GCs from the mouse neocortex and RNA sequencing quality metrics.
(a) Workflow enabling parallel extraction and subtype-specific sorting of CPN- or CThPN-specific somata and GCs. RNA isolated from respective samples sorter-purified by flow cytometry is then used for subcellular mapping of subtype-specific transcriptomes. A detailed description of this protocol can be found in54. (b) Side scatter over forward scatter data for size-calibrated BD SORP beads run on a flow cytometer in fluorescent small particle sorting (FSPS) configuration. FSPS enables detection, analysis, and efficient sorting of particles in the 100nm-1 μm diameter range. (c) Analysis of CPN GCs overlaid on top of BD SORP beads depicted in detail in panel (b). CPN GCs have diameters of ~200-800 nm, predominantly ~300-500 nm. (d/e) FSPS enables purification of subtype-specifically labeled GCs of CPN (D) and CThPN (E) from respective GCF samples. (f) Hydrolysis protection assays are used to assess the presence of intact GC particles with intact membranes, thereby protecting their specific molecular contents from RNase or proteinase digestion in the absence of detergents. Schematic modified from24 (g) Western blot of representative proteinase protection assay of CPN GCF detected for multiple subcellular markers, as previously reported in24. Likely ambient contaminations like GM130 or Map2 are degraded in the absence of detergents (-/+), while GC markers such as GAP43 or Actin are preserved in the absence of detergents (-/+), and are only degraded once GC membranes are permeabilized by detergent (+/+). (h) Tape station results of representative RNase protection assay of CPN GC. When the sample is treated with RNase, RNA quality is preserved in the absence of detergents (-/+), and only upon permeabilization of membranes does pronounced RNA degradation occur (+/+). (i) Schematic of bioinformatics workflow to process transcriptomic data from reads to differential expression between samples. (j) Sequencing depth is >40M fragments for all libraries across subtypes and stages, as well as for background samples (GC fraction pre-sort, GCF). (k) Alignment of reads to contaminating features and the nuclear genome indicates that most reads, as expected, originate from the nuclear genome and that the sorted GC samples have a higher proportion of reads aligning to rRNAs. (l) Distribution of reads to genomic features shows that most reads align to UTRs and coding sequences (CDSs), and not genomic regions downstream of transcription end site (TES) or upstream of transcription start site (TSS), as expected; GC and GCF samples have higher representation in 3’UTRs. (m) The coverage over 3’ ends is higher in GC and GCF samples relative to somata. (n) A cutoff of 2 counts per million (CPM) was used for soma samples, since this is the inflection point between signal and noise CPM distributions. (o) A CPM cutoff for sorted GC and GCF samples was defined as the 95th quantile of the distribution of CPMs of non-expressed genes (dotted lines), which were determined by the soma cutoff in panel (f). (p) Heatmap showing correlation between normalized transcript counts of all pairs of samples.
Extended Data Fig. 3 Subcellular RNA correction to minimize effect of potential ambient contamination and developmental stage agnostic analysis.
(a) To minimize effects of potential ambient contamination in the GC preparation, transcriptomes obtained from sorted GC samples are corrected for their respective input material, that is RNA extracted from the unsorted GCF. (b/c) GC transcripts for CPN and CThPN are defined as those having a higher abundance in sorted GCs than in GCF. (d) Correction for input material results in exclusion of likely ambient contaminants, for example marker genes for various non-neuronal cell types (grey), while preserving a class of robustly GC transcripts (red). (e) GO term enrichment comparing GC genes to likely contaminants from the GCF. (f) Principal component analysis of CPN (light and dark red) and CThPN (light and dark blue) soma transcriptomes collected at developmental stage P1 (light colors) and P3 (dark colors). Data show striking separation according to subtype and additional distinction within each subtype cluster across developmental stages. (g) Heatmap of correlated soma samples for CPN (red) and CThPN (blue) at P1 (lighter color) and P3 (darker color). LUT reflects pairwise Euclidean distance between the log2 TPMs of all genes in a sample. (h) Principal component analysis of CPN (light and dark red) and CThPN (light and dark blue) GC transcriptomes collected at developmental stage P1 (light colors) and P3 (dark colors). Data show striking separation according to subtype and additional, but less distinct separation within each subtype cluster across developmental stages. (i) Representative images of coronal brain sections from intersectional transgenic mice (genotype indicated on top) showing CThPN having reached the thalamus at P1, innervating and branching into the different thalamic nuclei at P3 and having established circuitry at P7. (j) ASD and schizophrenia disease-associated genes are enriched (95% confidence interval) in soma- and GC-enriched transcriptomes. Both CPN and CThPN soma-localized and GC- localized transcriptomes show enrichment for ASD-associated genes, based on comparison with two independent data sets. Cross-referencing with two independent GWAS studies for schizophrenia additionally revealed that CThPN soma- and GC-localized transcriptomes are also enriched for schizophrenia-associated genes. There is no enrichment detected with genes associated with bipolar disorder in either CPN or CThPN transcriptomes, serving as a neuropsychiatric disease control, highlighting specificity in ASD and schizophrenia.
Extended Data Fig. 4 Expression and subcellular transcript localization of WRC core components in CPN and CThPN.
(a) Volcano plot indicating CPN-specific subcellular transcript localization of core components of the WRC as orange dots (FDR < 0.05). (b) Volcano plot indicating CThPN-specific subcellular transcript localization of core components of the WRC as blue dots (FDR < 0.05). (c) 95% confidence intervals of log2 fold change GC/GCF for each of the WRC-associated transcripts. (d) Transcript abundance (transcripts per million, TPM, visualized on log2 scale) of the non-WRC associated members of the Wasp gene family N-Wasp, Jmy, and Whamm in somata and GCs of CPN (pink/brown) and CThPN (light/dark blue). (e) Transcript abundance (TPM, visualized on log2 scale) of WRC core components in somata and GCs of CPN (pink/brown) and CThPN (light/dark blue). (f/g) Representative western blot images and quantification of triplicates for each protein’s abundance of WAVE paralogs WAVE1, WAVE2, WAVE3, in the post nuclear homogenate (PNH) and growth cone fraction (GCF) of CPN, all quantification normalized to bAct, p-values indicate test results. All three paralogs are enriched in the GCF when compared to the PNH. (h) Maximum intensity projection (10 μm) of representative RNAscope confocal image of Wave1, the respective positive control (posCtrl, targeting Ppib), or negative control (negCtrl, targeting DapB) in GFP-positive cultured CPN. Solid arrowheads indicate RNAscope puncta localized to proximal or more distal neurites. (i) Maximum intensity projection (10 μm) of representative RNAscope confocal image of the Wave paralogs Wave2 (magenta) and Wave3 (red) in GFP-positive cultured CPN. Solid arrowheads indicate Wave3 RNAscope puncta, while open arrow heads indicate Wave2 RNAscope puncta localized to proximal or more distal neurites. Similar distribution of puncta were observed for 6-7 neurons per condition. (j) Maximum intensity projection (10 μm) of representative confocal image cultured CPN neurons stained with antibody directed against WAVE1 and DAPI. The left panel shows the secondary antibody only control with no WAVE1 signal, the right panel shows a full ICC staining. Both images were acquired using identical microscope and image adjustment settings.
Extended Data Fig. 5 Expression and subcellular transcript localization of WIRS-containing receptors in CPN and CThPN.
(a) Volcano plot indicating CPN-specific subcellular transcript localization of WIRS-containing receptors as orange dots (FDR < 0.05). (b) Volcano plot indicating CThPN-specific subcellular transcript localization of WIRS-containing receptors as blue dots (FDR < 0.05). (c) 95% confidence intervals of log2 fold change GC/GCF for each of the WIRS-containing transcripts. (d) Differential expression levels (FDR < 0.05, dark grey) of WIRS-containing genes in CPN (orange) vs. CThPN (blue) somata. WIRS-containing genes that are not differentially expressed between subtypes are shown in green. (e) Maximum intensity projection (10 μm) of representative RNAscope confocal images of the WIRS receptors Pcdh17 (magenta) and Robo1 (red) in GFP-positive cultured CPN. Solid arrowhead indicates Robo1 RNAscope puncta localized to proximal (top panel) and distal (bottom panel) CPN neurites. Similar distribution of puncta were observed for 6-7 neurons per condition. (f) Transcript abundance (transcripts per million, TPM, visualized on log2 scale)) of examples of WIRS-containing receptors with GC-enriched transcript localization in GCs and somata of CPN (pink/brown) and CThPN (light/dark blue).
Extended Data Fig. 6 3’UTR isoform analysis identifies enrichment of poly-U motifs in GC transcriptomes over those of somata.
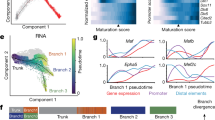
(a) Number of genes called by the gene-level approach for quantifying GC genes, last-250bp approach for quantifying GC 3’UTRs, or both. (b) MA plots depicting the log2 fold change and abundance of GC transcripts identified by the last-250bp approach. Each plot represents a subset of all transcripts: genes that are enriched in the last-250bp (purple), genes that were previously identified (black), isoforms of genes that were identified by both approaches (green), and as a control, isoforms of genes that were identified in neither approach (grey). Genes identified by only the gene-level approach (black) are generally of lower abundance. Last-250bp approach (purple) enables identification of GC-specific isoform variants of higher abundance genes. (c) GC 3’UTRs identified only in the last-250bp approach are on average longer (cdf – cumulative distribution function) (d) As an example, Mapt has one long 3’UTR enriched in GCs, but a shorter isoform not enriched in GCs, as seen by the read pileups; quantified in (e). (f) De novo motif analysis of GC-enriched 3’UTRs identifies multiple distinct versions of poly-U rich motifs preferentially detected in the GC compartment.
Extended Data Fig. 7 CPE location and spacing in GC 3’UTRs is consistent with baseline translational repression of targets.
(a) CPEBs bind to the cytoplasmic polyadenylation element (CPE) located in the 3‘-UTR of transcripts and complex with factors that control polyA tail length and translation. Cue-induced CPEB phosphorylation mediates polyA polymerase-mediated lengthening of the poly-A tail in the cytosol, enabling eiF4E interaction with PABP, and thereby initiation of translational activation of the respective transcripts [modified from54]. (b) Boxplots of expression levels in TPM (transcripts per million) of the three other Cpeb paralogs in CPN (pink/orange) and CThPN (light/dark blue) somata at P1 (brown, dark blue) and P3 (pink, light blue). (c) Representative western blot of protein samples obtained from whole forebrain (FB), micro-dissected cortex (Cx), and micro-dissected CPN axon bundles at corpus callosum (Ax), labeled for CPEB4, ACTIN, and TAU. Labeling for CPEB4 reveals 3 distinct bands, which are labeled as b1 (highest MW), b2 (middle MW), and b3 (lowest MW). Quantification of signal intensity in Cpeb4 b1-b3 from triplicates reveals relative enrichment of signal for micro-dissected axons to the middle band (b2). (d) Depending on the exact number and relative positioning of CPE motifs within the 3’-UTR, CPEBs can mediate either translational activation (as schematized in (A)) or translational repression [modified from54]. (e) Percent of transcripts per subcellular compartment and subtype (dark red: CPN soma, light red: CPN GC, dark blue: CThPN soma, light blue: CThPN GC) that have a CPE 6-25nt from the PAS (top panel) and density distribution of transcripts based on the distance from most distal CPE to the PAS (bottom panel), depicted on log2 scale. The region of 6-25nt distance to PAS is highlighted in light grey. (f) Top panels: Quantification of length (left), normalized number of contiguous binding sites (middle), and minimum distance between starts of CPE motifs (right) in 3’-UTRs enriched in GCs (orange, light blue) and somata (brown, dark blue) of CPN (orange/brown) and CThPN (light/dark blue), cdf – cumulative distribution function. Bottom panel: Density distribution of CPE motifs within 3’UTRs enriched in in GCs (orange, light blue) and somata (brown, dark blue) of CPN (orange/brown) and CThPN (light/dark blue). (g) Top panel: Barplot showing percent of transcripts per subcellular compartment and subtype (dark red: CPN soma, light red: CPN GC, dark blue: CThPN soma, light blue: CThPN GC) where the CPE(s) are likely activating or repressing (based on their location and distance relative to PAS). Bottom panel: Density distribution of transcripts per subcellular compartment and subtype (dark red: CPN soma, light red: CPN GC, dark blue: CThPN soma, light blue: CThPN GC) based on the distance between the two most distal CPEs. (h) Relative proportions of distinct variations of Cpeb4 motif, detected in 3’UTRs enriched in GCs (orange, light blue) and somata (brown, dark blue) of CThPN (light/dark blue) and CPN (orange/brown). (i/j) Efficiency of shRNA-mediated knockdown of Cpeb4. (i) Knockdown efficiency quantified via qPCR in transfected N2a cells and in in utero electroporated and FACS-purified CPN. (j) Knockdown efficiency assessed via immunohistochemistry in transfected N2a cells. (k) At P8, constitutive shRNA knockdown of Cpeb4 in CPN (red, n = 4, across two independent litters) by in utero electroporation does neither affect axon density in the corpus callosum nor fiber innervation in the contralateral gray matter compared to CPN treated with a scrambled shRNA control (scrb, teal, n = 6, across two independent litters). (l/m) Total number of labeled CPN in a single 40 μm coronal brain section of mice in utero electroporated with a scrambled control construct (ctrl) or an shRNA targeting Cpeb4 for knockdown (KD), tissue analyzed at P3 (upper panel) or P8 (lower panel). (n/o) Unilateral in utero electroporation at E15 with a scrambled control construct (ctrl, left panels, n = 3 for P3 and n=4 for P8), or shRNA targeting Cpeb4 (KD, right panels, n = 3 for P3 and n=6 for P8) results in reproducible targeting of CPN in cortical layer II/III in the lateral somatosensory cortex, tissue analyzed at P3 (upper panels) and P8 (lower panels).
Extended Data Fig. 8 Localized transcriptomes change dynamically from P1 to P3, as CPN switch from axon elongation to grey matter innervation and synapse formation.
(a) Somatosensory area CPN extend their axons across the midline around P1, innervate their homotopic target area in the contralateral cortex around P3, and collateralize within the cortical grey matter and likely start formation of synapses around P7. (b-e) Quantification of labeled axons/collaterals in progressively entered regions of interest (highlighted in schematic insets to the right, n = 3 for each developmental time), focusing on (b) CPN axon/collateral elongation in the subcortical white matter, (c) innervations into cortical grey matter (GM), (d) axon/collateral density in layer V or (e) in layers II/III at P1 (light pink), P3 (dark pink), and P7 (red). (f) Gene set enrichment analysis for CPN somata comparing P1 and P3. (g) Quadrant plot of differential gene expression by CPN at P3 vs. P1, and transcript localization by subcellular compartments: in somata (x axis) and GCs (y-axis). (h) Scaled transcript abundance of most significantly regulated CPNGC genes comparing P1 and P3, analyzed at gene level (left) and transcript level (right). (i) Median length of 3’UTR for transcripts detected in CPN and enriched at P1 or P3. Color ribbons indicate 50%, 75%, and 95% confidence intervals (CI). (j) Length of 3’UTRs of CPNGC genes increases from developmental stage P1 to P3.
Extended Data Fig. 9 The RBP Rbms1 increases transcript levels from P0 to P5, and is likely involved in localization and stabilization of CPNGC transcripts.
(a) Micro array data for RBPs Rbms1, Celf4, Pcbp3, and Tia1 at P0, P5, and P10, analyzed for CPN (red) and CThPN (blue), from Arlotta et al.55. (b) Boxplots highlighting RNA expression changes in TPM (transcripts per million) for the paralogs Rbms1, Rbms2, and Rbms3 in CPN somata at P1 (pink) and P3 (brown). (c) GO term enrichment for subset of CPNGC transcripts containing RBMS1 binding motifs at developmental stage P3. (d/e) Efficiency of shRNA-mediated knockdown of Rbms1, assessed via (D) qPCR in transfected N2a cells and in utero electroporated and FACS-purified CPN, as well as (E) via immunocytochemistry in transfected N2a cells. (f) Total number of labeled CPN in the cortex of a single 40 μm coronal brain section of mice electroporated with a scrambled control construct (ctrl) or a shRNA targeting Rbms1 (KD). (g) Unilateral in utero electroporation at E15 with a scrambled control construct (ctrl, left panel, n = 5), or shRNA targeting Rbms1 (KD, right panel, n = 4) results in comparable positioning of targeted CPN in cortical layer II/III in the lateral somatosensory cortex. The number of labeled CPN was higher in control mice, compared to KD mice.
Supplementary information
Supplementary Table 1 (download XLSX )
List of GO terms differentially enriched in either the soma or GC compartment for union of CPNs and CThPNs. The center statistics and P values come from an overrepresentation analysis (hypergeometric test) that accounts for hierarchical GO terms, calculated with the clusterProfiler tool in R. P values were adjusted with the Benjamini–Hochberg procedure.
Supplementary Table 2 (download XLSX )
Results of differential expression analysis at the gene and transcript levels, comparing expression across subtypes of cortical PNs and compartments and across developmental stages for CPNs. Statistics were estimated with a Wald test using the tool DESeq2. P values were adjusted with the Benjamini–Hochberg procedure. Tab names indicate <gene|transcript>_<positive_condition> versus <baseline_condition>.
Supplementary Table 3 (download XLSX )
Cross-referencing subcellular differential gene expression data of this report for CPNs and CThPNs with previously published data sets of genes associated with neurodevelopmental and neuropsychiatric disorders. Columns denote which study/census the gene was identified in (disease_group) and in which subcellular compartment(s) we identified mRNAs for this gene (express_group).
Supplementary Table 4 (download XLSX )
List of GO terms differentially enriched in GC compartments of distinct subtypes of cortical PNs. The center statistics and P values come from an overrepresentation analysis (hypergeometric test) that accounts for hierarchical GO terms, calculated with the clusterProfiler tool in R. P values were adjusted with the Benjamini–Hochberg procedure.
Source data
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Veeraraghavan, P., Engmann, A.K., Hatch, J.J. et al. Dynamic subtype- and context-specific subcellular RNA regulation in growth cones of developing neurons of the cerebral cortex. Nat Neurosci 29, 581–591 (2026). https://doi.org/10.1038/s41593-025-02173-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-025-02173-0