Abstract
The intrinsic organization underlying the central cognitive role of the prefrontal cortex (PFC) is poorly understood. We approached organization by profiling the activity and spatial location of >24,000 neurons recorded in awake mice. High-resolution activity maps of the PFC did not align with cytoarchitecturally defined subregions. Instead, spontaneous activity and tuning to choice during a behavioral task were both related to intra-PFC hierarchy, suggesting that connectivity, rather than cytoarchitecture, shapes the PFC’s activity landscape. Low-rate, regular spontaneous firing was a hallmark of both the PFC and high hierarchy. Surprisingly, choice tuning was overrepresented in units displaying high spontaneous firing rates, linking connectivity-based hierarchy to distinct functional properties in separate neuronal populations. Our data-driven approach provides a scalable roadmap to explore functional organizations in diverse brain regions and species, opening avenues to obtain an integrated view of activity, structure and function in the brain.
Similar content being viewed by others
Main
The PFC integrates information from all over the brain and is crucial for emotional and cognitive functions1,2. While mapping of gene expression3,4,5, cytoarchitecture6,7,8 and connectivity9,10,11,12,13 provides insights into the PFC’s infrastructure, a functional description of the PFC remains elusive14. It is unclear to what extent subregions of the PFC hold functional specializations14,15, and the existing structural descriptions of the PFC are yet to be integrated with the neuronal activities underlying this region’s information processing16. Furthermore, it is an open question whether the PFC’s neuronal activity is distinctive from other cortical regions and what kind of activity patterns would support PFC-specific functional demands.
In the current study, we approach the organization of the PFC at the level of single-neuron activity. A neuron’s firing pattern is a reflection of intrinsic biophysical properties and the neuron’s embedding in a particular network17,18,19. The activity profile of neurons should, therefore, inform about the functional properties of brain regions. In line with this, it has been shown that brain regions differ in activity patterns18,20,21,22 and that there is a correlation between connectivity-based hierarchy and firing patterns23,24,25. Yet, while cytoarchitecture is used to spatially define regions in the brain—and to sample and summarize activity—the relevance of cytoarchitectural delineations to function remains unclear, especially in case of the PFC, whose functions are considered to be distributed rather than localized26.
Here, we recorded single-neuron activity using high-density probes (Neuropixels) in awake mice and evaluated neuronal firing patterns at multiple spatial scales. Specifically, we analyzed spontaneous activity patterns, tone response profiles and tuning to aspects of goal-directed behavior. We addressed the general relationship between single-unit activity and brain anatomy, as well as the PFC’s distinctive activity characteristics and internal structure. The approach presented is a roadmap, applicable across brain regions and species.
Capturing the diversity of spontaneous firing patterns
We recorded the firing activity of 24,248 single units from awake, head-fixed mice (dataset KI; Fig. 1a–c). Overall, 52% of the units originated from the PFC and 48% from other brain regions (3 cortical, 10 subcortical; Fig. 1d and Extended Data Fig. 1a,b). In the PFC, we included 11 subregions9,14: secondary motor area (MOs), anterior cingulate area—dorsal and ventral parts (ACAd and ACAv, respectively), prelimbic area (PL), infralimbic area (ILA), orbital area—medial, lateral and ventrolateral parts (ORBm, ORBvl and ORBl, respectively), agranular insular area—dorsal and ventral parts (AId and AIv, respectively) and frontal pole (FRP). As spontaneous activity, we extracted 3-s epochs preceding the onset of auditory stimuli (Fig. 1c; ~227 epochs per recording session). Epochs contaminated by ‘sleep-like’ periods27 were excluded due to their fundamentally different, nonstationary firing characteristics (Extended Data Fig. 1c–e and Methods). While generally stationary for individual units, activity patterns varied considerably across units (Fig. 1d). To capture this diversity, we characterized each unit by three metrics: firing rate, burstiness and memory. Burstiness reflects the variability of a unit’s inter-spike intervals (ISIs), while memory is defined as the correlation coefficient between subsequent ISIs (Fig. 1e)28. Note that burstiness solely describes the distribution of ISI durations, while memory reflects the temporal ordering of ISIs. Memory and burstiness are mathematically independent of each other and together comprehensively capture two key aspects of firing patterns: sequential structure and regularity. These two metrics thus offer a robust characterization that encompasses the information provided by other commonly used interval metrics, such as CV2 and ‘local variation’ (LvR)20. The memory metric is related to neuronal timescales that recent studies23,24 derived from firing autocorrelations. However, the complexity of the autocorrelations at the single-neuron level disagrees with parametrization into a timescale through exponential fitting17. In contrast, the memory metric used here avoids the assumptions underlying timescales and describes the sequential structure of firing in a less biased way. Each metric was computed per 3-s epoch and averaged across epochs, thus summarizing each unit’s firing pattern into three metrics (Fig. 1e,f and Extended Data Fig. 1f–i).
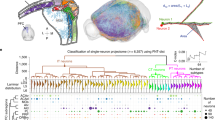
a, Experimental design; acute Neuropixels recordings in awake, head-fixed mice. Pure tones were presented during the sessions. b, Left, Coronal brain section with a Neuropixels probe track labeled by 1,1′-dioctadecyl-3,3,3′3′-tetramethylindocarbocyanine perchlorate (CM-DiI; red). Nuclei counterstained with DAPI (cyan). Scale bar, 1 mm. Right, Corresponding plate in the Allen mouse brain reference atlas (CCFv3, plate no. 576989109) with color coding of the recorded brain regions. c, Spike raster plot (top) of the 617 single units recorded with the probe shown in b. Bottom, Electromyogram (EMG). Turquoise shading indicates a 3-s epoch of spontaneous activity; orange shading indicates tone presentation (200 ms); black vertical line denotes tone onset. d, Top, Anatomical three-dimensional (3D) localization of all probe tracts (n = 99, black lines) in distinct subregions of the PFC (colored) and other brain regions. Bottom, Example spike raster plots (100 consecutive epochs aligned to sound onset at 0) of four single units (1–4), recorded in layer 5 and 6 (L5, L6). e, Schematic illustrating the three firing metrics used to characterize spontaneous firing patterns. In these examples, each epoch (box) holds the same number of action potentials (n = 30; vertical bars), that is, the firing rate is the same in all epochs. Top row, three distinct examples of burstiness: regular, Poissonian (random) and bursty, with constant neutral memory (M ~ 0). Bottom row, three distinct examples of memory: anti, neutral and high, with constant Poissonian burstiness (B ~ 0). Adapted with permission from ref. 28, European Physical Society. f, The three firing metrics used to characterize spontaneous activity of all recorded units: firing rate (log10FR, top), burstiness (middle) and memory (bottom). The gradient bars outline the respective metric range. The mean metrics across spontaneous epochs of the four units (1–4) in d are indicated, with highlighting of the metric combination of unit 2 (red). g, Examples of individual (n = 100, gray) and mean (black/purple) waveforms of a nw (top) and a ww (bottom) unit. Double arrows indicate peak-to-trough durations. h, Distribution of peak-to-trough durations of all mean waveforms (n = 24,248 single units; bin size = 1 ms). Peak-to-trough duration < 0.38 ms = nw, n = 4,200 units; >0.43 ms = ww, n = 19,186 units. Units with intermediate peak-to-trough duration were not classified (n = 862 units) and excluded from further analysis. i, Distributions of the three firing metrics: log10FR, burstiness and memory for nw (purple) and ww (black) units. Statistics used were mixed-effect regression, with two-sided P values corrected for multiple comparisons as described in the Methods.
Spike waveforms have been widely used to distinguish two neuron types, wide-width (ww) and narrow-width (nw) units (Fig. 1g), identified as putative excitatory and putative inhibitory neurons, respectively29. As expected, spike widths were bimodally distributed, allowing us to distinguish ww (n = 19,186; 81.1%) and nw (n = 4,200; 17.7%) units (Fig. 1h). nw neurons are expected to fire at high rates29,30, and indeed, nw units displayed significantly higher firing rates, along with higher burstiness and memory compared to ww units (Fig. 1i). To adequately analyze differences within each unit type, we processed ww and nw units separately. In the following section, we focus on ww units.
Brain regions have distinct firing repertoires
To obtain an overview of naturally occurring firing patterns, we trained a self-organizing map (SOM) on the three metrics extracted from the ww units. A SOM consists of a grid of nodes (here visualized as hexagons), where each node represents a combination of metrics; similar nodes are neighbors on the map (Extended Data Fig. 2a). The SOM displayed perpendicular gradients of burstiness and firing rate, showing that these two metrics vary independently of each other. In contrast, high memory patches coincided with medium to low burstiness, indicating that high memory and high burstiness are mutually exclusive (Fig. 2a). Thus, burstiness and memory were empirically dependent, albeit mathematically independent. Note that linear correlation did not reveal this empirical dependence between burstiness and memory (Extended Data Fig. 1f,g).
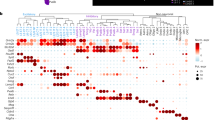
a, The SOM’s component planes for the three firing metrics. Each component plane consists of a hexagonal grid of nodes and displays the respective metric value per node in color. Together, the component planes visualize the feature landscape of the SOM. Contours (black/purple) delineate the unit categories defined in c. The original metric value ranges are displayed; for this, we reverted the standardization applied to each metric before SOM calculation (Methods). b, Count of PFC ww units assigned to each SOM node. Contours (purple) delineate the unit categories defined in c. c, Top, Partitioning of the SOM nodes into eight ww unit categories using hierarchical clustering. Bottom, Count of ww units per unit category. d, Summarizing the characteristics of each unit category. Median (dot) and 10th to 90th percentile (vertical line) of metrics across units assigned to each category. Circles below (‘summary’) further summarize the metric composition of each category: color indicates the median metric value based on a; radii are scaled linearly per metric across the eight categories, ranging from a fixed minimum radius reflecting the lowest median metric value to a fixed maximum radius reflecting the highest median metric value, to facilitate comparison between categories. e, Stability of ww unit categories across time. Top, Spontaneous 3-s epochs were allocated to blocks (~50 epochs per block) and each unit’s category was calculated per block. Bottom, Stability (fraction of units retaining their category) from one block to the next (black) compared to stability expected from marginal distributions (gray). f, Coincidence coefficient matrix quantifying the relative transition probabilities between unit categories (e, top) from one block to the next: −1, zero transitions; 0, random; 1, maximal possible number of transitions as derived from the marginal distributions (Methods). g, Right, Category enrichment profiles (Methods and Extended Data Fig. 3e) of brain (sub)regions. PFC subregions are in bold. d, deep layers. Nonsignificant E-scores are whitened (Supplementary Table 3). Left, hierarchical tree derived from the enrichment profiles. h, Graph representation of the data in g. Brain (sub)regions (dots) are arranged according to the first and second uniform manifold approximation and projection (UMAP) dimension of their enrichment profiles; line width scales with cosine similarity between category enrichment profiles of (sub)regions (only shown for similarities > 0.1). PFC subregions are in bold. i, Comparison of the enrichment of category 1 units in cortical (sub)regions between dataset KI (black) and dataset IBL Passive (gray). Dots and crosses indicate significant and nonsignificant enrichment, respectively. PFC subregions are in bold. Statistics used were Pearson correlation and the two-sided P value; n = 14. j, Matrix of Pearson correlation coefficients indicating similarity between ww and nw category enrichments across n = 9 brain regions in dataset KI. Only brain regions with sufficient nw units were included (Extended Data Fig. 5g). Data: dataset KI, ww units, all brain (sub)regions and layers, n = 19,186 units; a,c–f, dataset KI, PFC ww units, all layers, n = 10,413 units; b, dataset KI, ww units, all brain (sub)regions, for cortex restricted to deep layers (L5–6), n = 18,056 units; g,h, ww units, cortical (sub)regions, deep layers (L5–6), n = 9,715 units from dataset KI, n = 5,783 units from dataset IBL Passive; i, dataset KI, all brain (sub)regions, for cortex restricted to deep layers (L5–6), n = 3,984 nw units, n = 9,542 ww units; j.
Each unit was assigned to the node of the SOM that matched its metrics best. Every node represented units from numerous recordings, confirming that inter-animal variability did not drive the SOM’s feature landscape (Extended Data Fig. 2b). While PFC units constituted 54% of the ww dataset used to create the SOM, units from other brain regions were represented similarly well by the SOM (Extended Data Fig. 2c,d). Overall, PFC units were widely distributed on the SOM, with many units matching best to regular firing (that is, low burstiness) and avoiding anti-memory territories (Fig. 2b), while units from other brain regions populated different territories (Extended Data Fig. 2e). Notably, occupancy of SOM territories varied across PFC subregions, indicating spatial differentiation of firing repertoires within the PFC (Extended Data Fig. 2f).
Stable classification of single-unit firing patterns
To enable a statistical analysis of the spatial distribution of firing patterns, we categorized firing patterns by hierarchically clustering the SOM’s nodes—and, implicitly, the matched units (Fig. 2c and Extended Data Fig. 3a,b). Units within a category displayed similar firing statistics (Fig. 2d). To test whether units maintained stable firing properties over time, we allocated the spontaneous epochs into equally sized blocks and determined each unit’s category in the respective block. The category label (1–8) assigned to a unit per block matched well with the unit’s category obtained across all epochs (Extended Data Fig. 3c). Consistently, around 57% of units remained in the category they had in the previous block, which is considerably above the 18% expected from chance (marginal distributions; Fig. 2e). Splitting data into blocks reduces the sampling of spikes, deteriorating metric estimation and consequently also categorization (clustering) of units. This should disproportionately affect low-rate units, leading to underestimation of the stability for units with low firing rate in particular. Indeed, the block-wise assignment of categories to units was substantially less stable for low-rate categories (1–3, 7 and 8) compared to high-rate categories (Fig. 2f and Extended Data Fig. 3d): low-rate, regular-firing categories (1–3) mostly transitioned among themselves, as did the low-rate, bursty categories (7 and 8), suggesting the existence of statistically similar category groups.
Enrichment of low-rate, regular-firing units in the PFC
To quantify differences in firing patterns across various spatial entities—brain (sub)regions, layers and regions of interest (ROIs)—we defined a variable, the E-score, expressing for any selection of units the enrichment/depletion of a given unit category (1–8) relative to a chance distribution (Extended Data Fig. 3e). E-score analysis of ww unit activity revealed substantial enrichment in bursty units in the superficial cortical layers (L2/3), while deep cortical layers (L5–6) were enriched in regular-firing units (Extended Data Fig. 3f,g and Supplementary Tables 1 and 2). Given the anatomical organization of the mouse PFC and the experimental constraints, 95% of cortical units were recorded from deep layers. Therefore, the subsequent analyses include only the deep-layer (L5–6) units from cortical regions as well as all units from non-cortical regions.
The majority of the cytoarchitectural PFC subregions, that is, ILA, PL, ACAd, ACAv, ORBm and ORBvl, were generally enriched in low-rate, regular-firing units (categories 1–3), while holding subregion-specific differentiations (Fig. 2g). Interestingly, ILA and PL were both heavily and significantly enriched in units of the most regular-firing category 1 (Supplementary Table 3). The ORBl, MOs and AIv, in contrast, were significantly enriched in bursty category 8 units. Thus, the subregions ILA, PL, ACAd/ACAv and ORBm/ORBvl formed an activity-defined prefrontal entity, excluding ORBl, MOs and AIv (Fig. 2h). Other brain regions like thalamus (TH) and hippocampal formation (HPF) were heavily and significantly enriched in bursty units (categories 6–8), revealing a PFC-specific signature of low-rate, regular-firing units. Characterizing brain regions by firing properties requires reproducibility and robustness across datasets. Repeating the analyses with spontaneous activity obtained from the International Brain Laboratory (dataset IBL Passive31) revealed a high degree of similarity between the regional enrichment profiles of the two datasets (Fig. 2i and Extended Data Fig. 4).
To analyze nw unit activity, we trained a separate SOM and then repeated categorizations and comparisons of brain regions (Extended Data Fig. 5). ILA, PL and ACAd/v were significantly enriched in high-rate, regular-firing nw units with high memory (Extended Data Fig. 5g and Supplementary Table 4). ORBl was consistently and significantly enriched in high-rate, bursty units. Considering both nw and ww unit activity, we found regular-firing patterns to be the hallmark of the medial wall PFC subregions (ILA, PL and ACAd/ACAv). Relating nw and ww category enrichment profiles (Fig. 2j), we observed that brain (sub)regions enriched in low-rate, regular-firing ww units were enriched in high-rate, regular-firing nw units. Similarly, regions enriched in low-rate, bursty ww units were enriched in high-rate, bursty nw units. This could be interpreted as high-rate, putative inhibitory nw units contributing to the low-rate characteristic of ww units and implies that the PFC operates in an inhibition-dominated regime, which could be conducive to fine-tuned processing required for complex cognitive processes.
Enrichment in units displaying regular spontaneous firing reflects high cortical hierarchy
Having found differences between brain regions in terms of firing patterns, we next asked whether these differences are linked to the hierarchical organization of brain regions. The Allen Mouse Brain Connectivity Atlas describes the hierarchy of mouse cortical regions based on input/output connectivity motifs occurring between cortical and thalamic regions9. To investigate the relationship between the spontaneous unit categories (1–8) and cortical hierarchy, we correlated the E-scores of cortical (sub)regions with their respective cortical hierarchy score established by Harris et al.9. We found a positive correlation between cortical hierarchy and enrichment in low-rate, regular-firing units (categories 1–3). This positive correlation was not driven by a continuous relationship but due to a bimodal distribution of hierarchical scores between the low-hierarchy sensory regions versus the high-hierarchy PFC subregions (Fig. 3a and Extended Data Fig. 6a). To sample a wider range of cortical regions and hierarchical scores, we repeated the analysis with the dataset IBL Passive and found positive correlations between cortical hierarchy and the PFC’s signature categories 1–3 (Fig. 3b and Extended Data Fig. 6b). We identified even stronger, negative correlations between cortical hierarchy and enrichment in bursty, low-memory units (categories 6–8; Fig. 3c and Extended Data Fig. 6a,b). Importantly, when we limited the correlation analysis to PFC subregions, we found no direct relationship between cortical hierarchy and enrichment/depletion in unit categories (Supplementary Table 5), suggesting that firing patterns did not reflect cortical hierarchy at the level of cytoarchitectural subregions.
a,b, Correlation between enrichment in unit category 2 and cortical hierarchy score for dataset KI (a) and dataset IBL Passive (b). One dot/square per cortical (sub)region (dataset KI: n = 17 (sub)regions; dataset IBL: n = 32 (sub)regions). PFC subregions are in bold. Gray line indicates least-squares regression. Statistics used were Pearson correlation and two-sided P values. c, Pearson correlation coefficients between cortical hierarchy score and enrichment in the different unit categories for dataset KI (dark colors) and dataset IBL Passive (light colors); *P < 0.05, **P < 0.01, ***P < 0.001. Exact P values (two sided) are reported in a, b and Extended Data Fig. 6. Data: dataset KI, ww units, cortical (sub)regions, deep layers (L5–6), n = 10,898 units; a,c, dataset IBL Passive, ww units, cortical (sub)regions, deep layers (L5–6), n = 7,168 units; b,c, Cortical hierarchy scores from Harris et al.9.
Toward an activity-defined map of the PFC
Given the accumulating evidence for the incongruence between cytoarchitecture and connectivity in the mouse PFC14, demarcating and analyzing activity profiles based on cytoarchitecture could be called into question. Therefore, we parcellated the PFC subregions into smaller ROIs (dataROIs, n = 42), each containing a similar number of deep-layer ww units (~200; Fig. 4a,b and Extended Data Fig. 7a) and calculated E-scores per dataROI. Low-rate, regular-firing units (categories 1 and 3) were significantly enriched in dataROIs located in ILA and PL, while low-rate, bursty units (categories 7 and 8) were enriched in dataROIs located in the MOs and the ORBl (Fig. 4c,d, Extended Data Fig. 7b,c and Supplementary Table 6). Clustering dataROIs based on category enrichment profiles enabled us to define and spatially outline activity-based modules (Extended Data Fig. 7c,d). This revealed a homogeneous enrichment profile in ILA, creating an ILA-specific activity module that bordered a module with similar firing patterns in the adjacent parts of PL and ACAd (Fig. 4e). Other activity modules stretched over cytoarchitectural boundaries and occurred in multiple places. Thus, several cytoarchitectural subregions lacked spatially homogeneous activity signatures. The strong enrichment of low-rate, bursty units (categories 7 and 8) characterizing ORBl (Fig. 2g) could now, due to the improved spatial resolution, be pinpointed to a dataROI in the most ventral portion of this subregion. The similarity in the activity patterns in ILA and the adjacent portions of PL is consistent with the proposition of a ventromedial subdivision of the PFC defined by input and output connectivity12,14. We could not, however, identify specific activity signatures for the proposed dorsomedial and ventrolateral subdivisions14. A confound could be that our focus on deep layers primarily captures output activity, while the connectivity-based subdivisions of the PFC also consider the superficial input layers. Conversely, our relatively coarse sampling of the extensive territory of the dorsomedial (MOs and ACAd/ACAv) and the ventrolateral regions (ORBl and ORBvl) precludes discovery of small, homogeneous activity modules.
a, Flatmap of the mouse PFC with the anatomical location of all recording sites (black dots, from n = 99 Neuropixels probes) of dataset KI. Colors indicate cytoarchitectural PFC subregions. b, PFC subregions (black outlines, colors as in a) parcellated into 42 ROIs (dataROIs, white outlines) each holding ~200 ww units (Extended Data Fig. 7a) and identified by an ID number. Box is an enlargement of the corresponding box on the flatmap. c, Enrichment in category 1 ww units—reflecting low-rate, regular spontaneous firing. d, Enrichment in category 8 ww units—reflecting low-rate, bursty spontaneous firing. e, Activity modules of the PFC. Modules were obtained by clustering dataROIs based on their category enrichment profiles (Extended Data Fig. 7b–d). f, Large flatmap, the mouse PFC parcellated into 60 ROIs (GaoROIs, white outlines) of approximately the same volume (mean ± s.d.: 0.288 ± 0.070 mm3) and identified by an ID number. Each GaoROI is colored according to the PFC subregion where most of the units were located (colors as in a). GaoROIs with fewer than 20 units (white) were not analyzed. Small flatmap shows GaoROIs colored according to intra-PFC hierarchy score. g, Correlation between enrichment in unit category 1 and intra-PFC hierarchy score. One dot per GaoROI, colored as in f (n = 30 ROIs). Gray line indicates least-squares regression. Statistics used were Pearson correlation and a two-sided P value. h, Pearson correlation coefficients between intra-PFC hierarchy score and enrichment in the different unit categories across GaoROIs; *P < 0.05, **P < 0.01. P values are two sided and reported as exact values in g and Extended Data Fig. 7j–p. Data: dataset KI, PFC ww units, deep layers (L5–6), n = 10,413 units; PFC hierarchy scores in f–h from Gao et al.10.
Recently, two studies from Gao et al.10,11 used single-neuron connectivity tracing to describe the mouse intra-PFC connectivity in detail and provided a map of the hierarchical organization of the PFC. Gao et al. parcellated the PFC into 60 ROIs of similar volume (GaoROIs) and found no apparent correspondence between connectivity-based hierarchy and cytoarchitecture. Using GaoROIs (Fig. 4f), we corroborated the category enrichment profiles and the modules of spontaneous activity obtained with dataROIs (Extended Data Fig. 7e–h and Supplementary Table 7). To determine how spontaneous firing patterns relate to the detailed intra-PFC hierarchy proposed by Gao et al., we correlated the intra-PFC hierarchy score with the E-scores across GaoROIs (Fig. 4g,h and Extended Data Fig. 7i–p). Enrichment in low-rate, regular-firing units (category 1) was positively correlated with intra-PFC hierarchy (Fig. 4g), while enrichment in high-rate, bursty units (category 6) was negatively correlated to intra-PFC hierarchy (Extended Data Fig. 7n). Specifically, GaoROIs (19, 20, 41) located in PL displayed exceptionally high hierarchy scores and the strongest enrichment in unit category 1, while the GaoROI (23) located in the ventral part of ORBl featured the lowest hierarchy score and most prominent depletion in unit category 1 (Fig. 4g). Unfortunately, no hierarchy scores were available for ROIs located in ILA, where we had observed a particularly strong enrichment in unit category 1 (Fig. 4c). Overall, our results demonstrate that the positive correlation between enrichment in regular-firing unit categories (1–3) and connectivity-based hierarchy observed across cortical regions (Fig. 3) is replicated within the PFC across ROIs, indicating a general, scale-invariant link between enrichment in regular-firing units and hierarchy. Yet, as mentioned, the category enrichment profiles of cytoarchitecturally defined PFC subregions did not show an obvious relationship to cortical hierarchy (Fig. 3a, Extended Data Fig. 6 and Supplementary Table 5). A partitioning scheme transcending the traditionally defined subregions might, therefore, better capture both activity and connectivity-based hierarchy in the PFC.
Tone response maps do not align with cytoarchitecture
We addressed how sensory responses in the PFC map to cytoarchitecture by analyzing PFC unit activity related to presentation of 10-kHz tones (200 ms, 5–10-s random interstimulus intervals; dataset KI). To characterize tone responses, we extracted peri-stimulus time histograms (PSTHs) of ww units in deep layers (n = 15,352 units; Fig. 5a). Each PSTH was normalized to the pre-stimulus period. The resulting response traces were clustered based on their temporal profile into tone response categories 1–8, labeled according to their average peak delay. Tone response categories differed in response onset time, amplitude and persistence (Fig. 5b and Extended Data Fig. 8a–c). Most units displayed flat or negative response traces (category 8). Units belonging to tone response categories 1–4 exhibited well-defined peaks and consistent response peak onsets. In contrast, units belonging to tone response categories 5–7 had variable peak onset latencies and persistent responses.
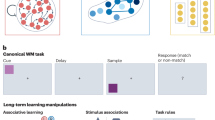
a, Spike raster plot (top) showing the response of an L5 ACAd ww unit to a 10-kHz tone (200 ms; gray horizontal bar) and normalized PSTH (bottom). Red vertical line indicates tone onset. b, Eight tone response categories of ww units. Each line represents a normalized PSTH averaged across the units assigned to a tone response category. Gray shading: tone presentation (200 ms). c, PFC flatmap with dataROIs (black outlines) colored according to enrichment in ww units of tone response category 1. d, Tone response modules of the PFC. Modules were obtained by clustering dataROIs based on their category enrichment profiles (Extended Data Fig. 8e,f). Black outlines indicate PFC subregions. e, Enrichment in tone-responsive ww units, that is, units significantly changing their firing rate in response to tone presentation (Methods; ZETA test32). f, Structure of the IBL task. Tuning to three task variables was analyzed for each unit: side of visual stimulus (left versus right; mustard), choice (clockwise (CW) versus counterclockwise (CCW) wheel turn; turquoise) and feedback (reward versus white noise; burgundy). Analyzed time windows (colored horizontal bars) were aligned to trial events (vertical lines). Adapted from ref. 31 under a Creative Commons license CC BY 4.0. g, Small flatmap shows PFC subregions (black outlines; colors as in Fig. 4a) parcellated into 73 ROIs (IBL ROIs; white outlines) each holding ~200 ww units (Extended Data Fig. 9c). Large flatmaps show enrichment in units significantly tuned to visual stimulus side (left), choice (middle) and feedback (right). h, Correlation between enrichment in choice-tuned units and intra-PFC hierarchy score. One dot per GaoROI (N = 39 ROIs). Each GaoROI is colored according to the PFC subregion where most of the units were located (Extended Data Fig. 9d). Gray line indicates least-squares regression. Statistics used were Pearson correlation, and two-sided P value. i, Cosine similarity between enrichment maps of spontaneous activity (Fig. 4c,d and Extended Data Fig. 7b) and enrichment maps of tone responsiveness (e) and task tuning (g). Cosine similarity values range from −1 (inversely proportional) to 1 (proportionally identical); 0 indicates no relationship (orthogonality). j, Mutual enrichment matrix quantifying the co-occurrence of spontaneous and tone response properties within single units. k, Mutual enrichment matrix quantifying the co-occurrence of spontaneous and task-tuning properties within single units. Data: dataset KI, PFC ww units with tone responses, deep layers (L5–6), n = 7,184 units; b–e, dataset IBL, PFC ww units, deep layers (L5–6), n = 16,148 units; g,h, dataset KI, PFC ww units with tone responses and spontaneous activity, deep layers (L5–6), n = 6,681; j, dataset IBL, PFC ww units with both task and spontaneous activity, deep layers (L5–6), n = 1,854; k, Intra-PFC hierarchy scores in h from Gao et al.10.
To map the different tone responses onto the PFC space, we calculated the tone response enrichment profiles for dataROIs. Units displaying the earliest responses (category 1) were enriched in anterior parts of ACAd and ORBvl and in parts of PL (Fig. 5c). Neither this response map nor the maps of the other response types (categories 2–8) obeyed cytoarchitectural boundaries (Extended Data Fig. 8d). Clustering the dataROIs based on their enrichment profiles into modules corroborated tone response heterogeneity within PFC subregions (Fig. 5d and Extended Data Fig. 8e,f). Furthermore, classifying units as either tone-responsive or nonresponsive (Zenith of event-based time-locked anomalies (ZETA) test32; Methods) confirmed that tone-responsive units were enriched in spatial patches distributed across the PFC (Fig. 5e).
Mapping task variables during goal-directed behavior
Previous research has attributed cognitive processes to specific PFC subregions14. Our findings, however, show that spontaneous and tone response activity patterns defy these cytoarchitecturally defined subregions. To investigate how processes involved in goal-directed behavior map onto PFC space, we analyzed single-unit activity from the IBL decision-making task (dataset IBL), which includes sensory integration, decision-making and action execution31. Briefly, mice viewed a stimulus patch appearing on either the left or the right side of a screen and should turn a steering wheel to center the patch (left stimulus indicates a clockwise turn; right stimulus indicates a counterclockwise turn). Correct turns were rewarded with water, while incorrect turns were punished with white noise. Like the original study31, we focused on three task variables: (1) stimulus (0–100 ms after visual stimulus onset), (2) choice (100–0 ms before first wheel movement) and (3) feedback (0–200 ms after reward/noise delivery; Fig. 5f). Using a statistical procedure to isolate the individual task variables and account for spurious correlations31,33 (Methods), we identified units as ‘tuned’ if their firing rate significantly differed between stimuli (left-versus-right stimulus position), choices (leading to clockwise-versus-counterclockwise wheel turns; Extended Data Fig. 9a) or feedback type (reward versus noise delivery).
Partitioning the PFC into ROIs each containing ~200 units from dataset IBL (IBL ROIs; Extended Data Fig. 9b,c), we analyzed enrichment of tuned units in IBL ROIs. Only 105 of 16,148 PFC units were tuned to the position of the visual stimulus, with a single IBL ROI in the MOs showing strong enrichment (Fig. 5g). Choice-tuned units (n = 1,109 units) were enriched in a spatially cohesive territory spanning central MOs, the FRP and the anterior parts of ACAd, ORBvl and PL; ILA, ORBm, ORBl and ACAv were consistently depleted in choice-tuned units (Fig. 5g). Feedback-tuned units (n = 2,868 units) were enriched in AId, FRP and anterior parts of MOs, ORBvl and PL, while they were depleted in ILA, ORBm and ACAd/ACAv. Overall, visually tuned IBL ROIs were embedded in the choice-tuned territory, which partially overlapped with the feedback-tuned territory. These overlaps could be interpreted as a spatial evolution of tuning following the temporal progression34,35,36 from stimulus to choice to action.
To determine the relation of tone responsiveness and task variables to connectivity-based hierarchy, we correlated enrichment in tuned and tone-responsive units with the intra-PFC hierarchy score across GaoROIs. Enrichment in choice-tuned units correlated strongly with intra-PFC hierarchy score (R = 0.48, P = 0.002; Fig. 5h and Extended Data Fig. 9d), whereas correlations were weak for visual tuning and absent for feedback tuning and tone responsiveness (Extended Data Fig. 9e–g).
Relationship between spontaneous and evoked activity patterns at spatial and single-unit level
We hypothesized that spontaneous firing properties reflect processing properties of local networks and thus evaluated the similarity of maps of tone response and task tuning to maps of spontaneous activity. To compare maps with distinct ROI partitioning (dataROIs for tone response and spontaneous activity; IBL ROIs for task variables), we rasterized the PFC flatmap into 311,563 pixels. For any given map, each pixel was assigned the E-score of the ROI covering the pixel, resulting in a vector with 311,563 E-score entries. The similarity between two maps was quantified as the cosine similarity between their E-score vectors. Although no pairs of activity maps showed strong overall congruence, several maps exhibited partial spatial overlap (Fig. 5i): the map of choice-tuned units (Fig. 5g) was most similar to the map of low-rate, regular-firing units (spontaneous category 1; Fig. 4c). This spatial similarity aligns with our finding that both enrichment in choice-tuned units (Fig. 5h) and enrichment in low-rate, regular-firing units (Fig. 4g) correlated positively with intra-PFC hierarchy. The map of feedback-tuned units was most similar to the map of high-rate, bursty units (spontaneous category 6), while the map of overall tone responsiveness (Fig. 5e) was most similar to the map of low-rate, bursty units (spontaneous category 8; Fig. 4d). We next established whether the observed spatial similarities between maps reflect associations between spontaneous firing properties and tone response/tuning properties at the single-unit level. We expected, for instance, that tone responsiveness is overrepresented in units with low-rate, bursty spontaneous firing. Surprisingly, we found that tone-responsive units (tone response categories 1–7) were more likely than expected to have high spontaneous firing rates (spontaneous categories 4–6; Fig. 5j). Conversely, units lacking clear positive tone responses (tone response category 8) tended to have low spontaneous firing rates (spontaneous categories 1–3, 7 and 8). Similarly, analysis of goal-directed task variables revealed that units with choice tuning, as well as units displaying feedback tuning, were associated with high-rate, Poissonian firing (spontaneous category 4; Fig. 5k).
Discussion
In this study, we demonstrate that enrichment in units with low-rate, regular spontaneous firing is a hallmark of the PFC (Fig. 2) and a correlate of both cortical (Fig. 3) and intra-PFC (Fig. 4) hierarchy. These central findings are in line with research tying regular-firing patterns to computational processes supporting cognition, such as sustained input integration and stable representations37,38. Mechanistically, regular firing aligns with known PFC properties, including a high level of recurrent connectivity and ion channels with slow kinetics5,39,40. In apparent contradiction to this, we found that task-tuned units did not display low-rate, regular firing (Fig. 5). An explanation for this could be that regular-firing units support the emergence of task tuning in high-rate units within local networks.
Our results differ from Mochizuki et al.20, who reported near-random (Poisson) firing in the PFC. This is likely due to methodological differences, as Mochizuki et al. focused on in-task, nonstationary activity, analyzed very few units in rodents, pooled units across layers and waveform types and used different delineations of brain regions. In contrast, our analyses specifically focused on ww units in deep layers. Importantly, our enrichment score (E-score) reflects relative deviation from the sampled population; given the overrepresentation of PFC units in the dataset KI, we may even have underestimated how prominent the PFC’s regular-firing signature actually is.
The consistent correlations between connectivity-based hierarchy and spontaneous activity across both dataset KI and dataset IBL Passive affirm a robust relationship between cortical hierarchy and spontaneous firing repertoire (Fig. 3). This activity–hierarchy correlation is in line with studies on mice, macaques and humans showing that spontaneous neuronal timescales correlate with connectivity-based hierarchy. In contrast, Siegle et al.23 found no such correlation in the mouse visual system, potentially because (i) their application of the timescale metric to single-unit activity might have been inadequate to capture the complexity of single-unit autocorrelations17, and (ii) spontaneous activity–hierarchy correlations may be absent within the mouse visual system.
Contributing to the ongoing efforts to map the mouse brain3,4,5,9,10,11,12,13, we here provide detailed activity-based maps of the PFC. While we observed that one spontaneous activity module aligned with the cytoarchitecturally defined ILA subregion, other modules did not correspond to single subregions. Likewise, maps of spontaneous activity patterns, tone responses and task variables did not align with PFC subregions. Feedback-tuned units were particularly enriched in the anterolateral part of MOs. This is consistent with previous findings that feedback tuning is mostly carried by licking in the IBL task31 and that the anterolateral part of MOs is implicated in licking37.
Choice-tuned units were enriched in a spatially cohesive, hitherto undefined territory, covering central MOs and anterior parts of several PFC subregions. Interestingly, choice tuning was, despite spatially overlapping with low-rate, regular spontaneous firing, overrepresented specifically in units with high spontaneous firing rates. This indicates that choice tuning and low-rate, regular firing—the two hallmarks of high hierarchy we identified here—are found in separate neuronal populations that overlap in space. As intra-PFC hierarchy is derived from intrinsic input and output connectivity10, our results thus suggest that connectivity, rather than cytoarchitecture, shapes the PFC’s activity landscape. The discovery that a high spontaneous firing rate was associated not only with choice tuning, but also with responsiveness to tones and feedback tuning at the single-unit level, further suggests that spontaneous high firing rates, implying high excitability, predispose units to engage in sensory and cognitive processing.
Overall, our results highlight how distinct aspects of neuronal activity—spontaneous firing patterns, sensory responses and task-related tuning—contribute unique and complementary insights into the functional organization of the PFC. By linking spontaneous activity to connectivity-based hierarchy and uncovering how specific firing properties predispose neurons to engage in tone responsiveness and task tuning, we provide a multifaceted perspective on how network organization supports function in this region. Importantly, these findings challenge the traditional emphasis on cytoarchitecture, instead pointing to intrinsic connectivity as a primary organizing principle. Moving forward, expanding the range of tasks, stimuli and behavioral contexts will be essential to piece together the functional organization of the PFC. Our data-driven framework offers a scalable path forward, with broad applicability to other brain regions and species.
Methods
Animals
All procedures and experiments on animals were performed according to the guidelines of the Stockholm Municipal Committee for animal experiments and the Karolinska Institutet in Sweden (approval numbers 7362-2019 and 1535-2024). Adult wild-type mice (C57BL/6J, Charles River; 27 male and 39 female) aged 3–6 months were used. Animals were group-housed, up to five per cage, in a temperature-controlled (23 °C) and humidity-controlled (55%) environment in standard cages on a 12:12-h reversed light/dark cycle with ad libitum access to food and water, unless placed on a water restriction schedule. All water-restricted mice were restricted to 85–90% of their initial body weight by administering 1 ml of water per day.
Surgery
Adult mice were anesthetized with isoflurane (3% for induction, then 1–2%). Buprenorphine (0.1 mg per kg body weight, subcutaneous (s.c.)), carprofen (5 mg per kg body weight, s.c.) and lidocaine (4 mg per kg body weight, s.c.) were administered. The body temperature was maintained at 37 °C by a heating pad. An ocular ointment (Viscotears, Alcon) was applied over the eyes. The head of the mouse was fixed in a stereotaxic apparatus (Kopf). Lidocaine (2%) was injected locally before skin incision. The skin overlying the cortex was removed; the skull was first cleaned with chlorhexidine and then gently scratched with a scalpel blade. A thin layer of glue was applied on the exposed skull. A lightweight metal head-post was fixed with lightcuring dental adhesive (OptiBond FL, Kerr) and cement (Tetric EvoFlow, Invoclar Vivadent). For extracellular recordings, a chamber was constructed by building a wall with dental cement along the coronal suture and the front of the skull. Brain regions were targeted using stereotaxic coordinates. After the surgery, the animal was returned to its home cage, and carprofen (5 mg per kg body weight, s.c.) was provided for postoperative pain relief 24 h following surgery.
Habituation and behavioral settings
Following surgery recovery, mice were handled and progressively habituated to the head-fixation procedure over a period of 3 to 4 days by increasing the head-restriction time from 15 min to 1 h. After habituation, the mice were engaged in distinct behavioral settings (see Supplementary Methods for details). For each behavioral setting, auditory stimuli (10-kHz pure tones) were delivered through earphones. Importantly, although the mice were engaged in various behavioral tasks, firing metric distributions were not significantly different across behavioral settings. For all behavioral settings, delivery of the auditory stimuli was controlled with custom-written computer routines using a National Instruments board (PCI-6221) interfaced through MATLAB (MathWorks).
Head-fixed recordings
Animal preparation
For acute recordings, two to four small craniotomies (300–500 μm in diameter) were opened a few hours (>3 h) before the experiment to access the pre-marked targeted entry points (PFC: +2.20 to +1.60 mm AP, ±0.3 to ±1.5 mm ML; AUD: −3.20 mm AP, ±4.20 mm ML; CA1: −1.50 mm AP, ±1.6 mm ML; MOp: +1.90 mm to +1.50 mm AP, ±2.50 mm ML; SSp: −3.10 mm AP, ±2.80 mm ML). The mice were anesthetized with isoflurane (3% for induction, then 1–2%). Buprenorphine (0.1 mg per kg body weight, s.c.), carprofen (5 mg per kg body weight, s.c.) and lidocaine (4 mg per kg body weight, s.c.) were administered. The open craniotomy was covered with silicone sealant (Kwik-Cast, WPI), and the mouse was returned to its home cage for recovery.
Probe preparation
The probes were coated with CM-DiI (Thermo Fisher), a fixable lipophilic dye for post hoc recovery of the recording location. The coating was achieved by holding a drop of CM-DiI at the end of a micropipette and repeatedly painting the probe shank with the drop, letting it dry, after which the probe appeared pink.
Probe insertion
First, the reference electrode was connected to a silver wire positioned over the pia in a craniotomy using a micromanipulator. Then, the probe(s) was (were) lowered gradually (speed ~20 μ s−1) with a micromanipulator (uMp-4, Sensapex), using 0° to 20° angles, until the tip reached a depth of ~3,800–4,200 μm under the surface of the pia. The probe(s) was (were) allowed to sit in the brain for 20–30 min before the recordings started. A total of 99 probes were lowered with a maximum of 3 probes inserted simultaneously.
Data acquisition
Extracellular potentials were recorded using Neuropixels probes (phase 3B Option 1, IMEC) with 383 recording sites along a single shank covering 3,800 μm in depth or with Neuronexus probes (A1x32-Poly2-10mm-50s-177) with 32 recording sites along a single 750-μm shank. The spike-band data were digitized with a sampling frequency of 30 kHz with a gain of 500 while the local field potential (LFP) band data were digitized with a sampling frequency of 2.5 kHz with a gain of 250. The digitized signal was transferred to our data acquisition system (a PXIe acquisition module PXI-Express chassis: PXIe-1071 and MXI-Express interface: PCIe-8381 and PXIe-8381, National Instruments for Neuropixels recordings or an Open Ephys acquisition board for Neuronexus probes), written to disk using SpikeGLX (B. Karsh, Janelia) for Neuropixels probes or Open Ephys GUI for Neuronexus probes, and stored on a local server for later analyses. The action potential signals (‘spike band’) were filtered between 0.3 Hz and 10 kHz and amplified. The LFP signals (‘LFP band’) were filtered between 0.5 Hz and 500 Hz.
Probe cleaning
After recording, probes were cleaned for 30 min with a fresh Tergazyme solution (Sigma-Aldrich) and rinsed with distilled water overnight.
Perfusion
At the end of the last recording session, each mouse was deeply anesthetized with pentobarbital (60 mg per kg body weight; intraperitoneal), and then transcardially perfused with 4% paraformaldehyde. The brain was removed and post-fixed in 4% paraformaldehyde in 1× phosphate buffer.
Tissue clearing and Neuropixels probe track reconstruction
Brain sections were cut on a vibratome at a thickness of 400 μm (Leica VT1000, Leica Microsystems). Slices were repeatedly washed in phosphate buffer and cleared using ‘CUBIC reagent 1’ (25 wt% urea, 25 wt% N,N,N’,N’-tetrakis(2-hydroxypropyl) ethylenediamine, and 15 wt% polyethylene glycol mono-p-isooctylphenyl ether/Triton X-100) for two days. After repeated washing in phosphate buffer, slices were incubated with DAPI (1:50,000 dilution) for one day at room temperature. Slices were then re-washed in phosphate buffer and submerged in ‘CUBIC reagent 2’ (50 wt% sucrose, 25 wt% urea, 10 wt% 2,20,20’-nitrilotriethanol, and 0.1% vol/vol% Triton X-100) for further clearing. Slices were mounted on customized 400-μm-thick slides using CUBIC reagent 2 solution and covered with 1.5-mm cover glasses. Blue and red channels were imaged at 4× magnification using a Zeiss 800 or 880 confocal microscope and exported via ZEN black (2.1 SP3 v14.0). For each brain section, 6 to 7 z-stacks spaced by 50 μm were obtained and downsampled to 10-μm resolution. The z-stacks containing the probe’s red fluorescent signal (DiI) were registered in the Allen Institute Common Coordinate Framework (CCFv3) and the probe position was estimated using the accompanying ‘SHARP-Track’ pipeline (https://github.com/cortex-lab/allenCCF; Supplementary Fig. 1). The electrode locations along the probe were transformed into CCFv3 space based on the orientation and position of the probe track. A unit’s location was assigned based on the location of the electrode where the unit had the highest waveform amplitude.
Extraction of single-unit activity
Preprocessing
The high-pass-filtered spike-band data were preprocessed using common-average referencing: the channel’s median was subtracted to remove baseline offset fluctuations, then the median across channels was also subtracted from each channel to remove artifacts.
Semiautomatic spike sorting
The data were spike sorted with Kilosort2.0 (https://github.com/MouseLand/Kilosort/releases/tag/v2.0/). Clusters of waveforms were manually curated using the phy2 GUI (https://github.com/cortex-lab/phy/). During manual curation, clusters of waveforms showing near-zero amplitudes, non-physiological waveforms, inconsistent waveform shapes and/or refractory period violations were discarded. The remaining units were compared with spatially neighboring clusters. Units showing similar waveforms, clear common refractory periods and putative drift patterns were subjected to a merge attempt: if the cluster resulting from merging was showing consistent waveforms and a clear refractory period in its autocorrelogram, the merged cluster was used. Clusters were split when the principal features of waveforms indicated distinct clusters, and two or more groups of waveforms could be clearly identified. Double-counted spikes were removed.
Unit quality control
Automatic spike sorting (Kilosort2.0) followed by manual curation in Phy2 yielded an initial dataset of 34,642 units. We first excluded 24 recordings containing prominent LFP artifacts, reducing the dataset to 29,782 units. Next, we applied the following inclusion criteria based on Siegle et al.23: (1) ISI violation ratio < 0.1 and (2) presence of a plausible spike waveform. Presence ratio > 0 and amplitude cutoff < 0.1 were also checked and enforced (Supplementary Fig. 2). In addition, units were required to have a sufficient number of ISIs (≥4) in at least 12 separate 3-s epochs to allow reliable estimation of the memory metric. Applying these criteria resulted in a dataset KI of 24,248 high-quality units (Supplementary Fig. 2).
Temporal drift correction and removal of spikes during saturation epochs
Spike times were corrected for temporal drift during the recording time (~10 ms h−1) relative to a clock signal registered independently by the PXIe acquisition module and the PCI-6221 card logging the behavioral signals. First, the temporal drift between the two devices was measured for each recording. Second, a linear regression was applied to correct the behavioral timestamps relative to the spike times. Spikes within an interval from 1 s before saturation onset to 1 s after offset were removed. Spike times were saved in Neurodata Without Borders (NWB) files for subsequent analysis.
EMG extraction
In Neuropixels recordings, we defined the EMG from the high-frequency (1–10 kHz) muscular tone (neck muscles), picked up by the reference electrode (situated, for example, over the visual cortex) with zero time lag30,38,39. EMG signals were extracted as the band-pass filtered (ellipsoid band-pass filter with a lower and upper cutoff frequency of 1 kHz and 10 kHz, respectively) median signal across all the Neuropixels probe active channels (common noise source). EMG signals were downsampled to 1 kHz and stored in NWB files for further analysis.
Statistics and reproducibility
Pairwise correlations were quantified with the Pearson correlation coefficient (R). Significance of correlations was tested under the null hypothesis of zero correlation, assuming bivariate normality. All P values reported are two sided. Effects of neuron type on spontaneous firing metrics and differences in response onset time across unit categories were assessed using linear mixed-effect models. Enrichment was assessed using custom shuffling statistics. Responsiveness of single units was assessed using a ZETA test32. Details concerning linear mixed-effect models, enrichment statistics and the ZETA test are provided in ‘Data analysis’. No statistical method was used to predetermine sample size. Data were not anonymized for analysis. Recording sessions were excluded from any further analysis if (1) the presence of slow waves in the LFP and up-and-down states in cortical activity indicated that the session was dominated by sleep-like states, (2) the abundance of artifacts in the LFP made such assessment impossible, or (3) experiments involved licking and it was not possible to reliably detect licking from LFP artifacts or the piezo element. Following these criteria, 24 recordings were excluded in total.
Data analysis
Unless stated otherwise, continuous variables are reported as the median and interquartile range (IQR, 25th–75th percentiles) in the following.
Detection and exclusion of licking periods
In 18 of the 99 recording sessions in the dataset KI, mice could lick to obtain water rewards following the tone stimuli. Licking requires motor action, which could influence firing patterns. Licking also occurred outside the reward window and could thus affect not only tone response epochs but also spontaneous epochs. To improve comparability of patterns across sessions, licking periods were detected and both spontaneous and tone response epochs overlapping with licking periods were excluded. If available, licking was detected using a piezoelectric sensor. Otherwise, licking was detected as artifacts in LFPs (LFPs downsampled to 500 Hz with a polyphase filter) using independent component analysis. Lick periods invariably presented as a single independent component displaying a characteristic, rhythmic, saw-tooth like pattern with close to uniform weight contribution across the electrodes of a probe, that is, the lick artifacts were similarly strong across electrodes. Periods of lick artifacts in the lick component were detected using a semiautomatic procedure, which involved manually setting an amplitude threshold for each recording, detecting threshold crossings and manual curation that allowed deletion of false positives and inclusion of false negative detections, respectively.
Detection of sleep-like periods
Sleep-like periods27 were detected per recording session, as illustrated in Extended Data Fig. 1c. A mean firing rate vector across all cortical spikes (10 ms bin width) was computed and smoothed with a Savitzky–Golay filter (11 pts window width; order, 3) to obtain the mean smooth firing rate vector FR(mean). Two thresholds were defined, ϴgmean, the geometric mean of FR(mean) and ϴtrough = 0.2 × ϴgmean. Periods when FR(mean) was below ϴtrough for longer than 5 ms were detected as ‘troughs’. To obtain ‘off periods’, troughs were extended forward and backward in time until FR(mean) reached above ϴgmean. Off periods formed the basis of sleep-like activity. Successive off periods closer than 1.5 s were merged. Around merged off periods and individually occurring off periods 1 s and 0.3 s, respectively, were included into the sleep-like periods. All sleep-like periods were reviewed visually and consistently coincided with a flat EMG, indicating absence of movement, and large-amplitude, irregular, slow-wave activity in the LFP typical of non-rapid eye movement sleep and drowsiness27. Detection of sleep-like periods was only performed for the dataset KI, as sessions of this dataset consistently featured a sufficient number of cortical units to detect collective off periods. Sleep-like episodes occupied 6% (IQR, 0–18%) of the time in recording sessions.
Epoch selection
Dataset KI, spontaneous epochs
In the dataset KI, spontaneous epochs were selected as time windows from 3 s before tone onset until tone onset. All epochs containing licking (see ‘Detection and exclusion of licking periods’) and where saturation occurred in the spike band were excluded. Sleep-like epochs were defined as all episodes overlapping with sleep-like periods (see ‘Detection of sleep-like periods’). Sleep-like epochs are only featured in Extended Data Fig. 1c–e. All other analyses of spontaneous activity are based on ‘spontaneous active’ epochs occurring with at least 1 s of temporal distance to sleep-like episodes. Recording sessions contained 227 (IQR, 172–284) spontaneous active epochs and 19 (IQR, 1–84) sleep-like epochs.
Dataset IBL Passive, spontaneous epochs
The dataset IBL Passive is a subset of the dataset IBL and comprises 303 recordings that contain, in addition to task activity, spontaneous activity (IBL Neuropixels Brainwide Map31, accessed February 2024 from https://registry.opendata.aws/ibl-brain-wide-map/). A 5-min block of spontaneous activity was recorded after about 68 min (IQR, 58–80 min), toward the end of a session. We obtained spontaneous epochs by splitting the 5-min block into 99 epochs lasting 3 s each.
Robustness of results for shorter spontaneous epoch durations
We selected a pre-stimulus epoch of 3 s, as the inter-tone intervals varied randomly between 5 s and 10 s, and tone-evoked responses were observed to dissipate within 2 s after tone onset. Analyses based on shorter pre-stimulus epochs of 2 s or 1 s were also performed, but fewer units could be included due to insufficient data (number of spikes) for reliably estimating activity patterns in these shorter epochs. However, when using shorter epochs, results remained qualitatively consistent, validating the robustness of our findings.
Dataset KI, tone response epochs
In the dataset KI, epochs used to compute tone response traces were selected as time windows starting 2 s before tone onset until 0.7 s after tone onset. All epochs containing licking, sleep-like periods, and where saturation occurred in the spike band were excluded. Recording sessions contained 188 (IQR: 86–228) tone response epochs. The number of tone response epochs per recording was lower than the number of spontaneous active epochs per recording because licking disproportionately occurred during tone response time windows, which coincided with reward presentation in some recordings (see ‘Detection and exclusion of licking periods’).
Waveform classification
A maximum of 2,000 randomly selected 2.8-ms waveforms per unit were extracted from the spike band. A total of 82 sample points (–1.4 ms and +1.4 ms) around each spike time provided by Kilosort2 were collected per waveform. The mean waveforms per unit were obtained by averaging across all collected waveforms per unit. Each mean waveform was converted from 16-bit analog values (i) to voltage values (V) according to equation (1):
With Vmax = 0.6 V, Imax = 512 bit and gain = 500, where the factor (Vmax) ÷ (Imax × gain) is the least significant byte. The mean waveforms were then interpolated by a factor of 1,000 (one-dimensional linear interpolation) and baseline corrected. A custom script was used to detect and compute the main peaks, amplitude values, the polarity of the waveform, the main slopes and the amplitude ratio of the interpolated mean waveform per unit. Only the peak-to-trough duration was used for waveform classification. The valley observed in the distribution of peak-to-trough durations (Fig. 1h) was used to label units as nw units (peak-to-trough duration < 0.38 ms), ww units (peak-to-trough duration > 0.43 ms) and unclassified units (0.38 ms ≤ peak-to-trough duration ≤ 0.43 ms) in accordance with previous studies30.
Extraction of spontaneous activity metrics
Spontaneous activity metrics were extracted from spontaneous epochs (see ‘Epoch selection’). For each unit, the three metrics characterizing firing rate, burstiness and memory were first calculated per epoch. The firing rate metric log10FR was computed as the decadic logarithm of the number of spikes in an epoch divided by epoch duration (3 s). The burstiness and memory metrics were based on ISIs and defined according to ref. 28. In brief, burstiness ‘B’ can be understood as the coefficient of variation of ISIs normalized to a range between –1 (completely regular; s.d.(ISI)«mean(ISI)) and 1 (maximally bursty; s.d.(ISI)»mean(ISI)) and was calculated according to equation (2):
with mean(ISI) and s.d.(ISI) being the mean and standard deviation of ISIs, respectively. Memory ‘M’ was defined as the Pearson correlation coefficient (PCC) between successive ISIs as given by equation (3):
Burstiness and memory were only computed in epochs with at least six spikes available (corresponding to at least four ISIs). Units with fewer than 12 epochs with at least six spikes were excluded from further analyses. To characterize a unit’s firing pattern, each of the three metrics was averaged across epochs. Note that calculating burstiness and memory per epoch before averaging has the advantage that the distribution-based burstiness and the sequence-based memory can both be faithfully computed. The metric LvR proposed by Shinomoto et al.40 was designed to mitigate vulnerability of ISI-based irregularity metrics to slow fluctuations in firing rates when computing ISI-based metrics over longer stretches of time. Yet, given the brief and stationary (stimulation free) character of the spontaneous epochs in the dataset KI, burstiness and memory are a more informative variable choice than LvR as these metrics allow to address the distributional and sequential character of bursting independently of each other.
Extraction and post-processing of tone responses
Tone responses were extracted from tone response epochs, which spanned an interval from 2 s before the stimulus (‘prestim-window’) to 0.7 s after the stimulus (‘poststim-window’; see ‘Epoch selection’).
Extraction of PSTHs
For each unit, spikes times relative to tone onset were collected from all tone response epochs to construct a firing rate PSTH (0.5 ms bin width, Fig. 5a). The PSTH was smoothed by convolving it with a Gaussian kernel (10 ms standard deviation of the kernel; 80 ms width of convolution window). Units with an overall average firing rate below 0.1 Hz across all tone response epochs or an average firing rate below 0.1 Hz across all prestim-windows were excluded from further analyses. To obtain a trace reflecting relative stimulus-induced rate changes, each unit’s smoothed PSTH was z-scored to its prestim-window through subtracting the mean and dividing by the standard deviation across the prestim-window. Tone response traces were defined as the smoothed, z-scored PSTHs extending from stimulus onset to 0.65 s after onset.
Dimensionality reduction of tone response traces
To facilitate later classification of tone responses (see ‘SOM analysis’ and ‘Hierarchical clustering analyses’ below), the dimensionality of the response traces was reduced using principal component analysis (PCA). Tone response traces from all units (both nw and ww units pooled) were collected into a data matrix (dimensions: (number of units, number of response trace time points)). Each time point in the data matrix was mean centered. PCA was performed on the time points. The top eight principal components, explaining together over 95% of the variance, were retained. The dot product of a unit’s original tone response trace and the principal components resulted in eight scores summarizing the tone response trace of a unit.
Task-tuning analyses of the dataset IBL
We closely followed the methods of single-unit analyses used in International Brain Laboratory31, where a comprehensive description of the experiment and its statistical evaluation can be found. For our analyses, which were limited to PFC ww units from deep layers, the dataset IBL provided 16,148 units from 66 recordings in 29 mice.
Task variables and block structure
We assessed tuning to three task variables: visual stimulus, choice and feedback. Each task variable could adopt two alternative task values (visual stimulus: left versus right, choice: counterclockwise versus clockwise, feedback: reward versus noise). Depending on whether the visual stimulus was shown on the left or right, mice had to perform a counterclockwise or clockwise turn of a wheel to obtain a reward. Only trials where the first wheel movement occurred within 0.08 s to 2 s after stimulus onset were included. The task was structured into blocks. The first block had a 50:50 distribution of left/right stimuli, while subsequent blocks alternated between left-biased and right-biased distributions (80:20). Each block contained between 20 and 100 trials.
Combined-condition Mann–Whitney U test
The aim was to determine, for each unit and task variable, whether the firing rates were significantly higher for one task value as compared to the other, while controlling for the influence of other task variables and spurious correlations due to changes of a unit’s firing rate during the experiment41. Firing rates were computed in the analysis windows shown in Fig. 5f. To assess, for example, tuning to choice, firing rates between trials with counterclockwise versus clockwise turns were compared. These comparisons were made within unique condition combinations, defined by the same visual stimulus side and block identity. For each combination, trials were ranked by firing rate and a Mann–Whitney U statistic comparing the two task values (clockwise versus counterclockwise) was computed. Then a combined U statistic across all unique condition combinations was calculated. Correspondingly, when evaluating visual stimulus tuning and feedback tuning the choice value was kept fixed.
Significance testing via shuffling
To assess significance, the combined U statistic was compared to a null distribution of 2,000 surrogate U statistics generated by shuffling the task values within each condition combination. A unit was considered significantly tuned to a task variable if both the combined-condition U test yielded a P value < 0.05, and a standard (unconditioned) Mann–Whitney U test yielded a P value < 0.001.
Enrichment analysis of task tuning
Enrichment flatmaps, quantifying the enrichment in significantly tuned units in IBL ROIs, were computed for each task variable (see ‘PFC flatmap projection and parcellation’ and ‘Enrichment statistics’). Note that the dataset IBL Passive provided only 1,854 units (24 recordings; 18 mice) in the PFC, which we deemed insufficient for a robust ROI-based enrichment analysis. Yet, the mutual enrichment between spontaneous activity and task variables shown in Fig. 5k is of necessity based on the dataset IBL Passive, for which spontaneous activity was available along with task data.
SOM analysis
A SOM is an unsupervised machine learning algorithm42 used here to summarize a set of n-dimensional feature vectors into a two-dimensional grid of nodes. Each node represents an n-dimensional prototype vector (visualized as a hexagon). After training the SOM, similar prototype vectors become neighbors on the SOM grid. Separate SOMs were constructed for the following data selections of the dataset KI: (i) spontaneous activity of ww units, (ii) spontaneous activity of nw units (iii) and tone responses of ww units.
Input features
To obtain input for training the SOM, each unit j was represented by a feature vector. For spontaneous activity (i and ii), the feature vector xj consisted of the three spontaneous firing metrics: xj = [log10FR, burstiness, memory]; see ‘Extraction of spontaneous activity metrics’. For tone response activity (iii), the feature vector comprised eight principal component scores (PCSs) summarizing the tone response trace of each unit: xj = [PCSj,1,PCSj,2,…PCSj,8]; see ‘Extraction and post-processing of tone responses’. Feature vectors were collected from all units in the respective data selection. Each feature was standardized by subtracting the mean and dividing by the standard deviation across all included units.
SOM architecture and initialization
The number of nodes in the SOM was manually determined to balance a detailed representation of the input space (requiring more nodes) with preservation of neighborhood relationships and a compact visualization (requiring fewer nodes). The shape (x–y dimensions) of the SOM and initialization of nodes were determined using the linear initialization method suggested by Kohonen43. This involved performing a PCA on the input feature vectors and setting the height and width of the SOM proportional to the ratio of the two largest eigenvalues.
SOM training
The SOM was trained using the batch training method and the neighborhood functions provided by Kind and Brunner44. In brief, each feature vector (summarizing a unit’s characteristics) was first assigned to the SOM node with the closest prototype vector in Euclidean space, referred to as the best matching node (BMN). The prototype vectors of the BMNs and their neighboring nodes were then updated to more closely resemble their assigned input feature vectors, using a Gaussian neighborhood function. Over the 200 training iterations, the neighborhood radius was gradually reduced, leading to more localized and subtle updates to the prototype vectors. After the SOM was trained, each unit was assigned to (that is, represented by) its BMN.
Projection of the dataset IBL Passive on SOMs of the dataset KI
IBL data were projected on the SOM trained on the respective subset of dataset KI by extracting the same features from units of the dataset IBL Passive and standardizing them to the mean and standard deviation of the features of the respective subset of dataset KI. As for the dataset KI, a BMN on the SOM derived from the dataset KI was then assigned to each IBL unit.
Advantages of SOMs
The SOM here serves as the first step in a two-step clustering procedure (with the second step detailed in ‘Hierarchical clustering analyses’). Each prototype vector can be viewed as representing a cluster of similar units. This coarse graining into prototype vectors makes subsequent clustering less sensitive to outliers. Additionally, SOMs are well suited for heuristic approaches like ours, as they offer a compact visualization of similarity structures and can accommodate and elucidate nonlinear relationships between features (metrics). Furthermore, new data can be readily projected on existing SOMs (as done here for the dataset IBL Passive), which increases reproducibility and comparability of results across datasets.
Hierarchical clustering analyses
Hierarchical trees were computed for three types of data: (i) SOM nodes (for example, see Fig. 2c and a detailed illustration in Extended Data Fig. 3b), (ii) PFC flatmap ROIs (for example, Fig. 4e) and (iii) enrichment matrices (for example, Fig. 2g), using Ward’s agglomerative hierarchical clustering algorithm45.
Feature selection and standardization
Before clustering, all features were standardized by subtracting the mean and dividing by the standard deviation across samples. The features used for constructing the hierarchical tree varied by type of data; SOM nodes were represented by prototype vectors (‘SOM analysis’) and PFC flatmap ROIs and enrichment matrices by enrichment profiles (‘Enrichment statistics’). In the case of enrichment matrices, the hierarchical tree was used solely to order the matrix rows for better visualization.
Clustering
For SOM nodes and PFC flatmap ROIs, the hierarchical tree was cut at a specific level to define clusters representing categories of spontaneous and tone response activity (for example, Figs. 2c and Fig. 5b) or modules of spontaneous and tone response activity (for example, Figs. 4e and Fig. 5d), respectively.
Categorization of unit activity
Specifically, in the case of spontaneous/response categories obtained by clustering the SOM nodes, units inherited their category from their BMN. Repeating analyses using various numbers of unit activity categories, that is, SOM clusters (ww: 5–10 categories; nw: 4–8 categories), led to qualitatively similar results, validating the robustness of our approach.
Determining a suitable number of clusters
The number of clusters (that is implicitly the threshold for cutting the hierarchical tree) was manually determined according to the Thorndike criterion46 and the Dunn index47 (for example, as shown in Extended Data Fig. 3a), aiming for a balance between cluster compactness and separation and for summarizing the data with the smallest number of clusters that aligned with both criteria.
Stability analyses
To assess the validity of unit categories, we tested how stable unit categories were when subdividing data into blocks and evaluating unit categories per block. Obtaining spontaneous unit categories per block: Obtaining the categories of a unit per block involved three steps: (i) allocating epochs to each block, (ii) calculating spontaneous metrics per block, and (iii) categorizing the unit in each block. (i) In experimental settings where there was an inherent block structure, spontaneous epochs belonging to a certain inherent block were used. Experimentally defined blocks contained 50 or 100 epochs. Blocks of 100 epochs were split in two. In experimental settings without block structure, epochs were allocated to blocks of around 50 epochs. There were 5 (IQR: 3–5) blocks available per recording. Epochs with licking or sleep-like activity (see ‘Epoch selection’) were discarded from each block. For each unit, blocks with fewer than 12 epochs with at least six spikes were excluded from further analyses. Per unit, blocks analyzed contained 36 (IQR: 23–49) epochs for ww units and 43 (IQR: 29–51) epochs for nw units, respectively. (ii) For each unit, metrics were extracted per epoch and then averaged per block, as detailed in ‘Extraction of spontaneous activity metrics’. (iii) For each unit, the three spontaneous metrics (features) describing a block were standardized to the mean and standard deviation of the respective original dataset used to compute the reference SOM (‘SOM analysis’). Block-wise ww unit data were standardized to and projected on the SOM derived from the original (full epoch) ww dataset. Block-wise nw unit data were standardized to and projected on the SOM derived from the original (full epoch) nw dataset. After standardizing, the BMN of each unit was identified on the respective SOM. The block-specific category of a unit was inherited from the BMN (‘Hierarchical clustering analyses’).
Assessment of overall stability per transition
For each unit u and transition t from one block to the next, we calculated a transition matrix Mu,t (dimensions: k × k; where k is the number of categories available for a dataset, that is k = 8 for ww units, and k = 5 for the nw units). This binary matrix contains exactly one nonzero entry at the row corresponding to the category the unit had in the current block and the column corresponding to the category the unit transitioned to in the next block. For each transition, the unit-wise transition matrices Mu,t were summed across all units, resulting in a k × k transition matrix Mt for each transition t (t = 1 corresponds to the transition from block 1 to block 2, t = 2 corresponds to the transition from block 2 to block 3, and so on). Each entry mi,j,t represents the number of times units of a certain category i in a block transitioned to a certain category j in the following block during a transition t. We defined the stability value per transition as the sum of the diagonal elements in Mt divided by the sum of all the entries in Mt. The stability value thus expresses the fraction of units that retained their category in a transition. In Fig. 2e, the stability value is contrasted with the chance stability expected from the marginal distributions of Mt.
Assessment of stability per category
To assess how stable individual categories were and which categories preferentially transitioned to which other categories, we summed transition matrices Mt across transitions to obtain an overall k × k transition count matrix M (Extended Data Fig. 3d) and analyzed various matrices derived from M. The transition probability matrix P was obtained by dividing the entries of a row i in M by the sum of the row entries, such that each row sums to 1 (Extended Data Fig. 3d, third panel, rows and columns reversed for visualization). Here, each entry pi,j gives the probability of transitioning to category j in the next block given category i was assigned in the current block. Evaluating stability from P has the disadvantage that categories more abundantly present in the dataset will appear more stable purely by chance. Therefore, we also considered the observed-to-expected ratio matrix O (Extended Data Fig. 3d, last panel). For this, we first defined an expected transition count matrix E derived from the marginal distributions of M (Extended Data Fig. 3d, second panel). An entry ei,j in E is defined as (Ri × Cj)/n, with Ri being the row total count of category i, Cj the column total of category j, and n the grand total of counts of M. O was obtained by dividing M by E. Each entry oi,j gives thus the ratio of the observed count and the expected count. This display accentuates the stability of overall small categories yet underemphasizes the stability of large categories.
Coincidence coefficient
To overcome the interpretative limitations of P and O, we developed the coincidence coefficient matrix X (Fig. 2f). We defined the coincidence coefficient xi,j as given by equation (4):
The coincidence coefficient ranges from –1 to 1. A value of –1 indicates that a transition never happens, 0 indicates that the transition happens as often as expected by chance, and 1 indicates that the transition happens as often as possible (relative to the size of the smallest category considered for the transition, that is min(Ri,Cj)).
Enrichment statistics
The need to control for sampling differences between groups (for example, groups of animals undergoing different behavioral settings) necessitated a statistical approach beyond fractional composition to determine whether a certain attribute c (for example, an activity-based unit category) is disproportionately frequent or rare in a statistical population r (for example, all units in a certain brain region). To avoid biases introduced by group-specific sampling, we developed an enrichment statistic that uses group-level shuffling (see Extended Data Fig. 3e for an illustration of the procedure described here).
Obtaining the coincidence matrix X
For each group g counting how often an attribute c appears within a population r generates a group-specific coincidence count matrix Xg (dimensions: n × m; where n is the number of populations and m is the number of attributes). Each entry xr,c,g represents the number of times c occurred in r for group g. Summing these matrices across all k groups produces the overall n × m coincidence matrix X, with xr,c = xr,c,1 + xr,c,2 + …. + xr,c,k.
Obtaining the surrogate matrix S
To estimate the coincidences expected by chance while controlling for group sampling biases, r–c combinations are shuffled (dissociated and randomly re-associated) within each group, generating a surrogate dataset j. The shuffling is repeated 1,000 times, creating 1,000 surrogate datasets. For each surrogate dataset, the process of obtaining X (described above) is repeated, leading to a group-specific surrogate coincidence matrix Sg,j and through summing over groups to an overall matrix Sj. Ultimately, all 1,000 Sj are combined into an n × m × 1,000 surrogate matrix S.
Calculating the E-score
The E-score er,c for each r–c combination is calculated by standardizing the original coincidence count xr,c to the distribution of the 1,000 surrogate counts in sr,c,1:1,000, as given by equation (5):
E-scores for all r–c combinations are collected in the n × m enrichment matrix E.
Significance of enrichment
The significance of an E-score (P value, Pr,c) is obtained from the fraction of surrogate counts sr,c,1:1,000 falling below (fr,c,−) or exceeding (fr,c,+) the original coincidence count xr,c: Pr,c = 1 – max(fr,c,−,fr,c,+).
Group-bias control used for specific types of data
Category enrichment statistics of (sub)regions and PFC flatmap ROIs (for example, Fig. 2g and Extended Data Fig. 7c), as well as the mutual enrichment of spontaneous activity and tone response categories at single-unit level (Fig. 5j), were group-controlled for behavioral settings. The layer-based category enrichment statistic (Extended Data Fig. 3f) was controlled for combinations of behavioral settings and brain (sub)regions, that is, each unique combination of a sub(region) and a behavioral setting constituted a separate group g.
Mutual enrichment
To evaluate mutual enrichment between spontaneous activity categories and tone response categories (Fig. 5j), as well as between spontaneous activity categories and task variables (Fig. 5k), each spontaneous activity category was treated as the statistical population r, and each tone response category or task variable as the attribute c.
PFC flatmap projection and parcellation
For a compact 2D representation of the PFC, we used the flatmap kindly provided by Gao et al.10. Each unit was positioned on the flatmap by projecting the 3D stereotactic coordinates (AP, ML, DL) from the unit’s recording site (where the unit’s waveform exhibited the highest amplitude) into the corresponding (u,v) coordinates of the flatmap.
Parcellation into ROIs of similar unit count
To generate statistically comparable ROIs that respected subregion boundaries, we parcellated cytoarchitecturally defined subregions of the PFC flatmap into ROIs (dataROIs for dataset KI, IBL ROIs for dataset IBL), each containing a similar number of units (dataset KI, Extended Data Fig. 7a; dataset IBL, Extended Data Fig. 9c). This parcellation was based on deep-layer units, aiming for approximately 200 units per ROI. The subregion FRP, which contained fewer than 200 units in total, was not subdivided further. All other subregions were subdivided as follows: The number of ROIs n for each subregion was determined by n = ((unit count in a subregion)/200). The (u,v) coordinates of the units contained in the subregion were then clustered into n groups using a constrained k-means algorithm48 with a target range of 160–240 units per ROI as the size constraint for each cluster. The subregion was then parcellated using a Voronoi diagram49 based on the cluster centers, such that a ROI was defined as the space closest to a common cluster center.
ZETA test
To test for responsiveness of single units to tones and estimate the onset of tone responses, we used the parameter-free ZETA test32. We analyzed the 700-ms tone response time windows following tone onset and obtained a two-sided P value and the response onset time (the time after which half of the response peak is reached) for each unit. Units with a P value < 0.01 were considered responsive to tones.
Linear mixed-effect regression models
To estimate the effect of neuron type on each spontaneous firing metric (log10FR, burstiness and memory), we fitted linear mixed-effect (LME) regression models with unit type (categorical variable: ww or nw) as a fixed effect and mouse as random effect50, as shown in equation (6):
Significance of the effects and interactions was assessed with post hoc F tests51 using the Kenward–Roger method for degree-of-freedom adjustment52 and Tukey’s method for multiple-comparison correction. We also used LME analysis to estimate the effect of epoch type (categorical variable ‘active’ or ‘sleep-like’) on each firing metric with the model in equation (7):
Correlation between firing metrics
To estimate the correlation between firing metrics, we fitted LME models for each pair of metrics (Metric1 and Metric2) with mouse as random effect as given by equation (8):
The correlation between any two variables is reported as the marginal coefficient of determination53.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data from the 99 recorded Neuropixels sessions used to generate all main and supplementary figures is available for download in NWB format on the DANDI Archive via https://dandiarchive.org/dandiset/000473/ and https://dandiarchive.org/dandiset/001260/. Source data are provided with this paper.
Code availability
The code used to preprocess the data and generate all manuscript figures is available at GitHub via https://github.com/hejDMC/pfcmap/ and Zenodo via https://doi.org/10.5281/zenodo.17559037 (ref. 54).
The following open-source software and toolboxes were used: Python: matplotlib, netgraph, numpy, scipy, shapely, sklearn, SOMz; Julia: LightXML, MixedModels, Statistics, StatsBase, StatsModels, StatsPlots; R: emmeans, lme4, MuMIn, pbkrtest.
References
Fuster, J. M. The Prefrontal Cortex (Elsevier, 2015).
Carlen, M. What constitutes the prefrontal cortex?. Science 358, 478–482 (2017).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Ortiz, C. et al. Molecular atlas of the adult mouse brain. Sci. Adv. 6, eabb3446 (2020).
Bhattacherjee, A. et al. Spatial transcriptomics reveals the distinct organization of mouse prefrontal cortex and neuronal subtypes regulating chronic pain. Nat. Neurosci. 26, 1880–1893 (2023).
Brodmann, K. Vergleichende Lokalisationslehre der Grosshirnrinde in ihren Prinzipien dargestellt auf Grund des Zellenbaues (Barth, 1909).
Franklin, K. B. J. & Praxinos, G. The Mouse Brain in Stereotaxic Coordinates (Academic Press, 2007).
Van De Werd, H. J. J. M. & Uylings, H. B. M. Comparison of (stereotactic) parcellations in mouse prefrontal cortex. Brain Struct. Funct. 219, 433–459 (2014).
Harris, J. A. et al. Hierarchical organization of cortical and thalamic connectivity. Nature 575, 195–202 (2019).
Gao, L. et al. Single-neuron projectome of mouse prefrontal cortex. Nat. Neurosci. 25, 515–529 (2022).
Gao, L. et al. Single-neuron analysis of dendrites and axons reveals the network organization in mouse prefrontal cortex. Nat. Neurosci. 26, 1111–1126 (2023).
Zingg, B. et al. Neural networks of the mouse neocortex. Cell 156, 1096–1111 (2014).
Ährlund-Richter, S. et al. A whole-brain atlas of monosynaptic input targeting four different cell types in the medial prefrontal cortex of the mouse. Nat. Neurosci. 22, 657–668 (2019).
Le Merre, P., Ährlund-Richter, S. & Carlén, M. The mouse prefrontal cortex: unity in diversity. Neuron 109, 1925–1944 (2021).
Wilson, C. R. E., Gaffan, D., Browning, P. G. F. & Baxter, M. G. Functional localization within the prefrontal cortex: missing the forest for the trees?. Trends Neurosci. 33, 533–540 (2010).
Christensen, A. J., Ott, T. & Kepecs, A. Cognition and the single neuron: How cell types construct the dynamic computations of frontal cortex. Curr. Opin. Neurobiol. 77, 102630 (2022).
Swindale, N. V., Spacek, M. A., Krause, M. & Mitelut, C. Spontaneous activity in cortical neurons is stereotyped and non-Poisson. Cereb. Cortex 33, 6508–6525 (2023).
Maimon, G. & Assad, J. A. Beyond Poisson: increased spike-time regularity across primate parietal cortex. Neuron 62, 426–440 (2009).
Wang, X. -J. Macroscopic gradients of synaptic excitation and inhibition in the neocortex. Nat. Rev. Neurosci. 21, 169–178 (2020).
Mochizuki, Y. et al. Similarity in neuronal firing regimes across mammalian species. J. Neurosci. 36, 5736–5747 (2016).
Tolossa, G. B., Schneider, A. M., Dyer, E. L. & Hengen, K. B. A conserved code for anatomy: Neurons throughout the brain embed robust signatures of their anatomical location into spike trains. eLife https://doi.org/10.7554/elife.101506 (2024).
Yu, H. et al. In vivo cell-type and brain region classification via multimodal contrastive learning. Preprint at bioRxiv (2024) https://doi.org/10.1101/2024.11.05.622159 (2024).
Siegle, J. H. et al. Survey of spiking in the mouse visual system reveals functional hierarchy. Nature 592, 86–92 (2021).
Murray, J. D. et al. A hierarchy of intrinsic timescales across primate cortex. Nat. Neurosci. 17, 1661–1663 (2014).
Zeisler, Z. R., Love, M., Rutishauser, U., Stoll, F. M. & Rudebeck, P. H. Consistent hierarchies of single-neuron timescales in mice, macaques and humans. J. Neurosci. 45, 19e2155242025 (2025).
Fuster, J. M. The prefrontal cortex–an update: time is of the essence. Neuron 30, 319–333 (2001).
Vyazovskiy, V. V. et al. Local sleep in awake rats. Nature 472, 443–447 (2011).
Goh, K. -I. & Barabási, A. -L. Burstiness and memory in complex systems. EPL 81, 48002 (2008).
Petersen, P. C., Siegle, J. H., Steinmetz, N. A., Mahallati, S. & Buzsáki, G. CellExplorer: a framework for visualizing and characterizing single neurons. Neuron 109, 3594–3608 (2021).
Senzai, Y., Fernandez-Ruiz, A. & Buzsáki, G. Layer-specific physiological features and interlaminar interactions in the primary visual cortex of the mouse. Neuron 101, 500–513 (2019).
International Brain Laboratory et al. A brain-wide map of neural activity during complex behaviour. Nature 645, 177–191 (2025).
Montijn, J. S. et al. A parameter-free statistical test for neuronal responsiveness. eLife 10, e71969 (2021).
Steinmetz, N. A., Zatka-Haas, P., Carandini, M. & Harris, K. D. Distributed coding of choice, action and engagement across the mouse brain. Nature 576, 266–273 (2019).
Gallero-Salas, Y. et al. Sensory and behavioral components of neocortical signal flow in discrimination tasks with short-term memory. Neuron 109, 135–148 (2021).
Esmaeili, V. et al. Rapid suppression and sustained activation of distinct cortical regions for a delayed sensory-triggered motor response. Neuron 109, 2183–2201 (2021).
Oryshchuk, A. et al. Distributed and specific encoding of sensory, motor, and decision information in the mouse neocortex during goal-directed behavior. Cell Rep. 43, 113618 (2024).
Chen, S. et al. Brain-wide neural activity underlying memory-guided movement. Cell 187, 676–691 (2024).
Schomburg, E. W. et al. Theta phase segregation of input-specific gamma patterns in entorhinal-Hippocampal networks. Neuron 84, 470–485 (2014).
Osanai, H., Yamamoto, J. & Kitamura, T. Extracting electromyographic signals from multi-channel LFPs using independent component analysis without direct muscular recording. Cell Rep. Methods 3, 100482 (2023).
Shinomoto, S. et al. Relating neuronal firing patterns to functional differentiation of cerebral cortex. PLOS Comput. Biol. 5, e1000433 (2009).
Harris, K. D. Nonsense correlations in neuroscience. Preprint at bioRxiv https://doi.org/10.1101/2020.11.29.402719 (2020).
Kohonen, T. Essentials of the self-organizing map. Neural Netw. 37, 52–65 (2013).
Kohonen, T. Self-Organizing Maps. Vol. 30 (Springer, 2001).
Kind, M. C. & Brunner, R. J. SOMz: photometric redshift PDFs with self organizing maps and random atlas. Preprint at https://doi.org/10.48550/arXiv.1312.5753 (2013).
Ward, J. H. Jr. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 58, 236–244 (1963).
Thorndike, R. L. Who belongs in the family?. Psychometrika 18, 267–276 (1953).
Dunn, J. C. Well-separated clusters and optimal fuzzy partitions. J. Cybernetics 4, 95–104 (1974).
Levy-Kramer, J. k-means-constrained. https://github.com/joshlk/k-means-constrained (2018).
Gillies, S. Shapely: manipulation and analysis of geometric objects. https://github.com/shapely/shapely (2007).
Yu, Z. et al. Beyond t-test and ANOVA: applications of mixed-effects models for more rigorous statistical analysis in neuroscience research. Neuron 110, 21–35 (2022).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest package: tests in linear mixed effects models. J. Stat. Softw. 82, 1–26 (2017).
Halekoh, U. & Højsgaard, S. A Kenward-Roger approximation and parametric bootstrap methods for tests in linear mixed models – the R package pbkrtest. J. Stat. Softw. 59, 1–32 (2014).
Burnham, K. P & Anderson, D. R. Model Selection and Multimodel Inference (Springer, 2004). https://doi.org/10.1007/b97636
Le Merre, P., Heining, K., & Carlén, M. A prefrontal cortex map based on single neuron activity. Zenodo https://doi.org/10.5281/zenodo.17559037 (2025).
Acknowledgements
We thank L. Gao and J. Yan for sharing the code for projecting 3D coordinates into flatmap space and intra-PFC hierarchy score data; G. Chapuis, O. Winter and other IBL members for their technical help and early access to the dataset IBL Passive. A. Wolthon for technical help in handling our mouse colony. This work was supported by the Wallenberg Scholar program (Knut and Alice Wallenberg Foundation), a KAW project grant (Knut and Alice Wallenberg Foundation), the WennerGren foundation, Hjärnfonden, a NARSAD Young Investigator Grant (Brain & Behavior Research Foundation) and a Stratneuro Postdoctoral grant. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Funding
Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Contributions
Conceptualization: P.L.M., K.H. and M.C. Methodology: P.L.M. and K.H. Investigation: P.L.M., M.S., F.J., E.M., N.G., R.Y., H.P. and F.W. Data analysis: P.L.M. and K.H. Project design and supervision: M.C. Visualization: P.L.M., K.H. and M.C. Writing: P.L.M., K.H. and M.C. All authors discussed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Adam Kepecs and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The dataset KI, with description of data selection and correlations between spontaneous firing metrics.
a, b Count, per brain region (a) and per PFC subregion (b), of all high-quality, manually sorted, single units included in the dataset KI. c, Excluding ‘sleep-like’ periods. Gray horizontal bars indicate 3 s epochs of spontaneous activity directly preceding tone presentation (200 ms, orange bar); black vertical line: tone onset. The left side exemplifies spontaneous epochs included in analysis; the right side exemplifies spontaneous sleep-like epochs excluded from analysis. Any 3 s epoch of spontaneous activity temporally overlapping with a sleep-like period was excluded. Sleep-like periods were identified by the presence of ‘off’ periods (blue vertical bars). Off periods were defined as drops in FR(mean) below the 20% threshold and extended forward and backward to when FR(mean) reached above its geometric mean. Off periods are accompanied by high-amplitude LFP waves, and sleep-like periods in general coincide with large-amplitude, highly, variable LFP. Time periods neighboring off periods (light blue shadings) were excluded as well (Methods). Top to bottom, Spike raster plot of the single units recorded with the probe whose location is shown in Fig. 1b. FR(mean): the smoothed firing rate averaged across cortical units (bin = 10 ms). Blue solid horizontal line: the geometric mean of FR(mean); blue dashed horizontal line: a threshold corresponding to 20% of the geometric mean of FR(mean). Colored traces of LFPs from the recording site situated centrally in the respective PFC subregion. Concurrent electromyogram (EMG). d, Distributions of the firing metrics log10FR, burstiness, and memory for ww units in included (black) and sleep-like excluded (light blue) 3 s epochs of spontaneous activity. Statistics: mixed-effect regression, two-sided p-values corrected for multiple comparisons as described in Methods. e, Same as d for nw units. f, Correlation between firing rate, burstiness, and memory across all ww units. Marginal coefficient of determination (R2m) values are shown above the diagonal, the respective p-values are shown below the diagonal. n.s.: non-significant, *** p < 0.001, mixed-effect regression. g, Two-dimensional histogram showing, across all ww units, the relationship between firing rate and burstiness (left), firing rate and memory (middle), and memory and burstiness (right). h, i, Same as f and g for nw units. *** p < 0.001, mixed-effect regression. Data: dataset KI, all units across brain (sub)regions and layers, n = 24,248 units; a dataset KI, all units across subregions and layers of the PFC, n = 12,674 units; b dataset KI, ww units, all brain (sub)regions and layers, n = 19,186 units with sufficient number of included 3 s epochs; d black, f, g dataset KI, ww units, all brain (sub)regions and layers, n = 11,599 units with sufficient number of excluded 3 s epochs; d blue dataset KI, nw units, all brain (sub)regions and layers, n = 4,200 units with sufficient number of included 3 s epochs; e black, h, i dataset KI, nw units, all brain (sub)regions and layers, n = 2,549 units with sufficient number of excluded 3 s epochs; e blue.
Extended Data Fig. 2 SOM validation and brain (sub)region specific occupancy of SOM territories.
a, U-matrix visualizing the distance between neighboring nodes of the SOM trained on ww units (Fig. 2a). In the U-matrix, an additional interpolating node is placed between each pair of neighboring (original) nodes. The interpolating node displays in color the Euclidean distance between the pair’s standardized metric vectors. The original SOM nodes are colored based on the average Euclidean distance value of all their surrounding interpolating nodes. b, Diversity of units assigned to each SOM node in terms of recording probe. Each pie chart (one pie chart/SOM node) reflects the range of different recording probes contributing units assigned to the node. A piece of the pie reflects the fraction of ww units assigned to the respective node that originate from a specific probe. The dataset KI holds 99 recording probes in total. c, Quantization error (QError) indicating how well ww units of different PFC subregions were represented on the SOM. The QError is defined as the Euclidean distance between a unit’s metrics vector and the metrics vector of the SOM node the unit is assigned to. A lower QError indicates better representation. Dots: median, vertical lines: 25th to 75th percentiles. d, Same as c but QErrors for the PFC overall (black) and other brain regions (gray). e, Count of ww units (all layers) assigned to each SOM node for different brain regions (for PFC see Fig. 2b). Orange contours delineate the unit categories defined in Fig. 2c. f, Count of ww units in deep layers (L5–6) assigned to each SOM node for PFC subregions. Data: dataset KI, ww units, brain (sub)regions, all layers, n = 19,186 units; a, b, d, e. dataset KI, ww units, all PFC subregions and layers, n = 10,413 units; c. dataset KI, PFC ww units, deep layers (L5–6), n = 9,319 units; f.
Extended Data Fig. 3 Classification of firing patterns, enrichment statistics, and layer-resolved enrichment profiles.
a, Criteria for determining a suitable number of clusters for partitioning the SOM trained on ww units (Fig. 2a,c). Red dashed line indicates the eight clusters used. Top, Euclidean distance in standardized metric space between the last pair of clusters joined as a function of the number of clusters (categories) defined when hierarchically clustering SOM nodes (read graph from right to left). Lower values indicate more homogenous clusters47. Bottom, Dunn index as a function of the number of clusters. The Dunn index is the ratio between minimal between-cluster distance and maximal within-cluster distance, with a higher Dunn index implying more compact and well-separated clusters48. b, Hierarchical clustering (ward linkage) of SOM nodes. Left, dendrogram displaying the hierarchical relationship of the eight unit categories partitioning the SOM in Fig. 2c (same coloring). Right, partitioning of the SOM when opting for four to seven clusters. Coloring follows the respective main branch on the left. c, Percentage distribution of the ww unit category labels computed per block (obtained when splitting unit data into blocks, Methods) for each ww unit category (row). Each row sums close to 100%, as percentages were rounded off. Number of blocks per unit category are indicated on the right. d, Statistics of ww unit category transitions from one block to the next. From left to right: count matrix with color indicting how many units that had a certain category in a block (x-axis) transitioned to a certain category in the subsequent block (y-axis); expected count matrix showing the transition counts expected from the marginal distributions of the count matrix (Methods); probability matrix showing the probability of a unit transitioning to a certain category given its category in the current block, with columns summing to 1; observed-to-expected ratio matrix showing the ratio between the observed and expected count for each transition. e, Statistical procedure for calculating enrichment scores (E-scores). From top to bottom: Schematic representation of two experimental groups (X1 and X2) with differential sampling of single-unit activity in two brain regions, rA and rB, respectively. Vertical bars: recording probes, gray = region rA, black = region rB; colored circles = three unit categories (red/blue/yellow). Coincidences of brain regions (r) and unit categories (c) are counted within each group (X1 and X2) and summed to generate the overall coincidence count matrix across groups (X). To estimate the coincidences expected by chance while controlling for group effects, r and c are dissociated and randomly re-associated for each group, resulting in group-specific surrogate coincidence matrices (S1,j and S2,j). These surrogate matrices are then summed to obtain an across-group surrogate matrix (Sj). The random dissociation, re-association, and coincidence counting process is repeated 1,000 times to generate an overall surrogate matrix, S, with dimensions [n regions] x [n activity categories] x 1,000 (bottom left). To calculate the E-score, er,c, for a specific activity category c in a given region r (shown here for region rB and category c3, bottom middle), the original coincidence count xr,c (yellow vertical line) is standardized to the distribution of the 1,000 surrogate counts in sr,c (yellow shaded area) by subtracting the mean (sr,c) and dividing by the standard deviation (σ) of sr,c. E-scores for all combinations of r and c are collected in the enrichment matrix E. Note that r and c can represent any attribute (for example, r could also refer to layer or experimental group), and that ‘group’ in this schematic denotes the attribute being controlled for (for example, ‘group’ could also refer to brain region, when controlling for brain regions, as in f). f, Category enrichment profiles of different cortical layers. Enrichment profiles were obtained statistically controlling for brain regions (e and Methods). Rows are sorted according to hierarchical clustering (ward linkage) of enrichment profiles. Non-significant E-scores are whitened. Circles below summarize the metric composition of each category as detailed in Fig. 2d. g, Right, Layer-specific category enrichment profiles of cortical (sub)regions. Bold: PFC subregions. Non-significant E-scores are whitened. Left, hierarchical tree (ward) derived from the enrichment profiles. Data: dataset KI, ww units, all brain (sub)regions and layers, n = 19,186 units; a–d, f, g.
Extended Data Fig. 4 Comparison of category enrichment profiles between dataset KI and dataset IBL Passive.
a, b Count of all high-quality, manually sorted, single units included in the dataset IBL Passive per brain region (a) and per PFC subregion (b). c, Count of ww units in deep layers (L5–6) per cortical (sub)region in dataset KI (black) and dataset IBL Passive (gray). d, Comparison of the enrichment of ww unit category 2 units in cortical (sub)regions between dataset KI (black) and dataset IBL Passive (gray). Dots and crosses indicate significant and non-significant enrichment, respectively. Bold: PFC subregions. |d: deep layers. Statistics: Pearson correlation, two-sided p-value, n = 14. e–j, Same as d, but for ww unit categories 3–8. k, Count of nw units in deep layers (L5–6) per cortical (sub)region in dataset KI (black) and dataset IBL Passive (gray). l–p, Same as d, but for nw units (nw unit categories 1–5), n = 9 (sub)regions. Data: dataset IBL Passive, all units, all brain (sub)regions and layers, n = 21,119 units; a. dataset IBL Passive, all units, all subregions and layers of the PFC, n = 3,053 units; b. dataset KI, ww units, cortical (sub)regions (matching dataset IBL Passive), deep layers, n = 9,603 units; c–j. dataset IBL Passive, ww units, cortical (sub)regions (matching dataset KI), deep layers, n = 3,010 units; c–j. dataset KI, nw units, cortical (sub)regions (matching dataset IBL Passive), deep layers, n = 1,637 units; k–p. dataset IBL Passive, nw units, cortical (sub)regions (matching dataset KI), deep layers, n = 759 units; k–p.
Extended Data Fig. 5 Spontaneous activity patterns in nw units: Characterization and analysis of category enrichment profiles in brain regions and PFC subregions.
a, The component planes of the SOM trained on the three firing metrics of nw units. Each component plane consists of a hexagonal grid of nodes and displays the respective metric value per node in color. Together, the component planes visualize the feature landscape of the SOM. Contours (black/purple) delineate the unit categories defined in c. The original metric value ranges are displayed; for this, we reverted the standardization applied to each metric before SOM calculation (Methods). b, Count of PFC nw units assigned to each SOM node. Contours (purple) delineate the unit categories defined in c. c, Top, Partitioning of the SOM nodes into five unit categories using hierarchical clustering. Bottom, count of nw units per unit category. d, Summarizing the characteristics of each unit category. Median (dot) and 10th to 90th percentile (vertical line) of metrics across units assigned to each category. Circles below (‘summary’) further summarize the metric composition of each category: color indicates the median metric value based on a; radii are scaled linearly per metric across the five categories, ranging from a fixed minimum radius reflecting the lowest median metric value to a fixed maximum radius reflecting the highest median metric value, to facilitate comparison between categories. e, Stability of nw unit categories across time. Top, spontaneous 3-s epochs were allocated to blocks ( ~ 50 epochs/block) and each unit’s category was calculated per block. Bottom, Stability (fraction of units retaining their category) from one block to the next (black) compared to stability expected from marginal distributions (gray). f, Quantification of the transitions between nw unit categories across all blocks shown as a coincidence coefficient matrix: -1: zero transitions; 0: random; 1: maximal possible number of transitions as derived from the marginal distributions (Methods). g, Right, Category enrichment profiles (Methods, Extended Data Fig. 3e) of brain (sub)regions. Bold: PFC subregions. |d: deep layers. Non-significant E-scores are whitened (Extended Data Table 4). Left, hierarchical tree (ward) derived from the enrichment profiles. h, Graph representation of the data in g. Nodes representing brain (sub)regions are arranged according to the first and second UMAP dimension of their enrichment profiles; line width scales with cosine similarity between category enrichment profiles of (sub)regions (only shown for similarities > 0.1). Bold: PFC subregions. |d: deep layers. Data: dataset KI, nw units, all brain (sub)regions and layers, n = 4,200 units; a, c–f. dataset KI, PFC nw units, all layers, n = 1,911 units; b. dataset KI, nw units, all brain (sub)regions, for cortex restricted to deep layers (L5–6), n = 3,984 units; g,h.
Extended Data Fig. 6 Spontaneous firing patterns reflect connectivity-based hierarchy of cortical regions.
a, Correlation between enrichment in unit categories and cortical hierarchy score for dataset KI. One dot/square per cortical (sub)region. Bold: PFC subregions. Gray line: least-squares regression. The pink frame delimits nw unit data. Statistics: Pearson correlation, two-sided p-values; n = 17 (ww), n = 13 (nw). b, Same as a, but for dataset IBL Passive. n = 32 (ww), n = 21 (nw). Data: dataset KI, cortical (sub)regions, deep layers (L5–6), n = 10,898 ww units, n = 1,826 nw units; a. dataset IBL Passive, cortical (sub)regions, deep layers (L5–6), n = 7,168 ww units, n = 1,026 nw units; b. . Cortical hierarchy scores from Harris et al.9.
Extended Data Fig. 7 ROI-based spontaneous activity mapping of the PFC and correlation with intra-PFC hierarchy.
a, Flatmap of the PFC with subregions (black outlines) parcellated into 42 regions of interest (dataROIs, white outlines) and identified by an ID number. Box: enlargement of the corresponding box on the flatmap. The color gradient visualizes unit count (ww units, L5–6) per ROI. b, PFC flatmaps with dataROIs (black outlines) colored according to enrichment in ww units of category 2–7. c, Clustering of PFC dataROIs into activity modules based on their category enrichment profiles. Top to bottom, hierarchical tree derived from enrichment profiles; dataROI ID numbers; PFC subregion identity of the dataROIs (color-coded as in Fig. 4b); activity modules (A–E); category enrichment profile of dataROIs. d, Criteria for determining a suitable number of clusters when partitioning the PFC flatmap into modules based on the dataROI’s category enrichment profiles (c). Red dashed line: the five clusters (modules) used. Top, Euclidean distance in standardized metric space between the last pair of clusters joined as a function of the number of modules defined during hierarchical clustering (read graph from right to left). Lower values indicate more homogenous clusters47. Bottom, Dunn index as a function of the number of clusters. The Dunn index is the ratio between minimal between-cluster distance and maximal within-cluster distance, with a higher Dunn index implying more compact and well-separated clusters48. e, Flatmap of the PFC colored according to the count of ww units (deep layers, L5–6) per GaoROI (gray outlines). Each GaoROI is identified by an ID number. GaoROIs with fewer than 20 units (white) were not included in analyses. f, Clustering of PFC GaoROIs into activity modules based on their category enrichment profiles. Top to bottom, hierarchical tree derived from enrichment profiles; GaoROI ID numbers; PFC subregion identity of the GaoROIs (color-coded as in Fig. 4f); activity modules (A–E); category enrichment profiles of GaoROIs. g, Same as d, but for clustering of GaoROIs. h, PFC flatmap with GaoROIs colored according to activity module. i, Data availability for correlations of E-scores with intra-PFC hierarchy using the GaoROI parcellation of the PFC (Fig. 4g and j–p). GaoROIs with both intra-PFC hierarchy score and category enrichment profile ( ≥ 20 units available) are colored according to the PFC subregion (Fig. 4a) where most of the units were located. Gray: no intra-PFC hierarchy score and too low (n < 20) unit count; white: intra-PFC hierarchy score but too low (n < 20) unit count; dark gray: sufficient number of units (n ≥ 20) but no intra-PFC hierarchy score. j–p, Correlation between enrichment in unit categories (2–8) and intra-PFC hierarchy score. One dot per GaoROI, colors as in i. Gray line: least-squares regression. Statistics: Pearson correlation, two-sided p-values, n = 30. Data: dataset KI, PFC ww units, deep layers (L5–6), n = 9,284 units. Intra-PFC hierarchy scores from Gao et al.10.
Extended Data Fig. 8 Tone responses in the PFC: characterization of categories, enrichment maps, and tone-response modules.
a, Response onset time (ZETA test38) per tone response category of PFC ww units in deep layers. The whisker plot displays the median (center line), interquartile range (25th to 75th percentiles), and the minimum and maximum non-outlier values (whiskers). Outliers, defined as values exceeding 1.5 times the interquartile range, are not shown. Only units that responded significantly according to the ZETA test were analyzed in terms of tone response onset time; numbers on top indicate the fraction of ZETA-significant units per category. b, Statistical assessment of the difference in response onset time between tone response categories of ZETA-significant units. The matrix shows p-values derived from a mixed-effect regression model (Methods). P-values are visualized on a log10 color scale and are to be interpreted from column to row (red: significantly increased latency, blue: significantly decreased latency, gray: no significant difference). c, Normalized PSTHs of single units arranged according to tone response category (1–8). Gray horizontal bar: tone presentation (200 ms); black vertical lines: tone onset/offset. Black dots: response onset time (ZETA test39, significant units only, p < 0.01). d, PFC flatmaps with dataROIs (black outlines) colored according to enrichment in ww units of tone response category 2–8. e, Criteria for determining a suitable number of dataROI clusters to partition the PFC flatmap into modules based on tone response category enrichment profiles of dataROIs (d and Fig. 5c). Red dashed line: the four clusters (modules) used. Left, Euclidean distance in standardized metric space between the last pair of clusters joined as a function of the number of clusters (modules) defined when hierarchically clustering the enrichment profiles of dataROIs (read graph from right to left). Lower values indicate more homogenous clusters47. Right, Dunn index as a function of the number of clusters. The Dunn index is the ratio between minimal between-cluster distance and maximal within-cluster distance, with a higher Dunn index implying more compact and well-separated clusters48. f, Clustering of PFC dataROIs into tone response modules based on their tone response category enrichment profiles. Top to bottom, hierarchical tree derived from dataROI enrichment profiles; dataROI ID numbers; PFC subregion identity of the dataROIs (colored as in Fig. 4b); tone response modules; tone response category enrichment profiles of dataROIs. Data: dataset KI, PFC ww units, deep layers (L5–6), with tone response activity available, n = 7,184 units.
Extended Data Fig. 9 The IBL’s goal-directed behavior dataset and correlation of task variables and tone responsiveness to intra-PFC hierarchy.
a, Spike raster plot of an example unit, aligned to movement onset (gray line). Turquoise bar: time window in which choice tuning was evaluated; mustard ticks: onset of visual stimulus. Top, trials with a clockwise (CW) choice. Bottom, trials with a counterclockwise (CCW) choice. b, Flatmap of the mouse PFC with the anatomical location of all recording sites (black dots, n = 66 Neuropixels probes) of dataset IBL with in-task data available. Colors: cytoarchitectural PFC subregions. c, Flatmap of the PFC with the subregions (black outlines) parcellated into IBL ROIs (white outlines). The color gradient visualizes unit count (ww units, L5–6) per ROI. d, Data availability for correlations of task-tuning E-scores with intra-PFC hierarchy using the GaoROI parcellation of the PFC (Fig. 5h and f, g). GaoROIs with both intra-PFC hierarchy score and category enrichment profile ( ≥ 20 units available) are colored according to the PFC subregion (Fig. 4a) where most of the units were located. White: intra-PFC hierarchy score but too low (n < 20) unit count; dark gray: sufficient number of units (n ≥ 20) but no intra-PFC hierarchy score. e, Correlation between enrichment in tone-responsive units and intra-PFC hierarchy score. One dot per GaoROI, colored as in Extended Data Fig. 7i. Gray line: least-squares regression. Statistics: Pearson correlation, two-sided p-value, n = 29. f, Correlation between enrichment in units tuned to the visual stimulus and intra-PFC hierarchy score. One dot per GaoROI, colored as in d. Gray line: least-squares regression. Statistics: Pearson correlation, two-sided p-value, n = 39. g, Same as f, but for units tuned to feedback. Data: dataset IBL, PFC ww units, deep layers (L5–6), n = 16,148 units; b–d, f, g. dataset KI, PFC ww units with tone responses, deep layers (L5–6), n = 7,184 units; e. Intra-PFC hierarchy scores from Gao et al.10.
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Figs. 1 and 2 and Tables 1–7.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download TIFF )
Original unmodified confocal image.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Le Merre, P., Heining, K., Slashcheva, M. et al. A prefrontal cortex map based on single-neuron activity. Nat Neurosci 29, 673–681 (2026). https://doi.org/10.1038/s41593-025-02190-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-025-02190-z