Abstract
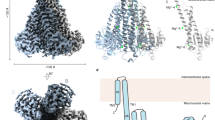
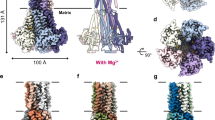
The human mitochondrial RNA splicing 2 protein (MRS2) has been implicated in Mg2+ transport across mitochondrial inner membranes, thus having an important role in Mg2+ homeostasis critical for mitochondrial integrity and function. However, the molecular mechanisms underlying its fundamental channel properties such as ion selectivity and regulation remain unclear. Here we present a structural and functional investigation of MRS2. Cryo-electron microscopy structures in various ionic conditions reveal a pentameric channel architecture and the molecular basis of ion permeation and potential regulation mechanisms. Electrophysiological analyses demonstrate that MRS2 is a Ca2+-regulated, nonselective channel permeable to Mg2+, Ca2+, Na+ and K+, which contrasts with its prokaryotic ortholog, CorA, operating as a Mg2+-gated Mg2+ channel. Moreover, a conserved arginine ring within the pore of MRS2 functions to restrict cation movements, thus preventing the channel from collapsing the proton motive force that drives mitochondrial adenosine triphosphate synthesis. Together, our results provide a molecular framework for further understanding MRS2 in mitochondrial function and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The raw images were deposited to the EM Public Image Archive with accession codes EMPIAR-12018, EMPIAR-12019 and EMPIAR-12035. The cryo-EM maps were deposited to EM Data Bank with accession codes EMD-41587, EMD-41588 and EMD-41589. Atomic coordinates were deposited to the PDB with accession codes 8TS1, 8TS2 and 8TS3. A previously published structure used for comparison was obtained from the PDB with accession code 4I0U. Previously published cryo-EM maps referred to are available from the EM Data Bank with accession codes EMD-41624, EMD-41628, EMD-35630 and EMD-35631. Source data are provided with this paper.
References
de Baaij, J. H., Hoenderop, J. G. & Bindels, R. J. Magnesium in man: implications for health and disease. Physiol. Rev. 95, 1–46 (2015).
Jin, F., Huang, Y. & Hattori, M. Recent advances in the structural biology of Mg(2+) channels and transporters. J. Mol. Biol. 434, 167729 (2022).
Quamme, G. A. Molecular identification of ancient and modern mammalian magnesium transporters. Am. J. Physiol. Cell Physiol. 298, C407–C429 (2010).
Wiesenberger, G., Waldherr, M. & Schweyen, R. J. The nuclear gene MRS2 is essential for the excision of group II introns from yeast mitochondrial transcripts in vivo. J. Biol. Chem. 267, 6963–6969 (1992).
Zsurka, G., Gregan, J. & Schweyen, R. J. The human mitochondrial Mrs2 protein functionally substitutes for its yeast homologue, a candidate magnesium transporter. Genomics 72, 158–168 (2001).
Bui, D. M., Gregan, J., Jarosch, E., Ragnini, A. & Schweyen, R. J. The bacterial magnesium transporter CorA can functionally substitute for its putative homologue Mrs2p in the yeast inner mitochondrial membrane. J. Biol. Chem. 274, 20438–20443 (1999).
Gregan, J., Kolisek, M. & Schweyen, R. J. Mitochondrial Mg(2+) homeostasis is critical for group II intron splicing in vivo. Genes Dev. 15, 2229–2237 (2001).
Kolisek, M. et al. Mrs2p is an essential component of the major electrophoretic Mg2+ influx system in mitochondria. EMBO J. 22, 1235–1244 (2003).
Schindl, R., Weghuber, J., Romanin, C. & Schweyen, R. J. Mrs2p forms a high conductance Mg2+ selective channel in mitochondria. Biophys. J. 93, 3872–3883 (2007).
Daw, C. C. et al. Lactate elicits ER-mitochondrial Mg2+ dynamics to integrate cellular metabolism. Cell 183, 474–489.e17 (2020).
Piskacek, M., Zotova, L., Zsurka, G. & Schweyen, R. J. Conditional knockdown of hMRS2 results in loss of mitochondrial Mg(2+) uptake and cell death. J. Cell. Mol. Med. 13, 693–700 (2009).
Kuramoto, T. et al. A mutation in the gene encoding mitochondrial Mg2+ channel MRS2 results in demyelination in the rat. PLoS Genet. 7, e1001262 (2011).
Madaris, T. R. et al. Limiting Mrs2-dependent mitochondrial Mg(2+) uptake induces metabolic programming in prolonged dietary stress. Cell Rep. 42, 112155 (2023).
Zhao, Y. et al. Differentially expressed gene profiles between multidrug resistant gastric adenocarcinoma cells and their parental cells. Cancer Lett. 185, 211–218 (2002).
Chen, Y. et al. Human mitochondrial Mrs2 protein promotes multidrug resistance in gastric cancer cells by regulating p27, cyclin D1 expression and cytochrome C release. Cancer Biol. Ther. 8, 607–614 (2009).
Yamanaka, R. et al. Mitochondrial Mg(2+) homeostasis decides cellular energy metabolism and vulnerability to stress. Sci. Rep. 6, 30027 (2016).
Franken, G. A. C., Huynen, M. A., Martinez-Cruz, L. A., Bindels, R. J. M. & de Baaij, J. H. F. Structural and functional comparison of magnesium transporters throughout evolution. Cell. Mol. Life Sci. 79, 418 (2022).
Lunin, V. V. et al. Crystal structure of the CorA Mg2+ transporter. Nature 440, 833–837 (2006).
Payandeh, J. & Pai, E. F. A structural basis for Mg2+ homeostasis and the CorA translocation cycle. EMBO J. 25, 3762–3773 (2006).
Eshaghi, S. et al. Crystal structure of a divalent metal ion transporter CorA at 2.9 angstrom resolution. Science 313, 354–357 (2006).
Guskov, A. et al. Structural insights into the mechanisms of Mg2+ uptake, transport, and gating by CorA. Proc. Natl Acad. Sci. USA 109, 18459–18464 (2012).
Dalmas, O. et al. A repulsion mechanism explains magnesium permeation and selectivity in CorA. Proc. Natl Acad. Sci. USA 111, 3002–3007 (2014).
Dalmas, O., Sompornpisut, P., Bezanilla, F. & Perozo, E. Molecular mechanism of Mg2+ -dependent gating in CorA. Nat. Commun. 5, 3590 (2014).
Matthies, D. et al. Cryo-EM structures of the magnesium channel CorA reveal symmetry break upon gating. Cell 164, 747–756 (2016).
Rangl, M., Schmandt, N., Perozo, E. & Scheuring, S. Real time dynamics of gating-related conformational changes in CorA. Elife 8, e47322 (2019).
Lee, A. et al. Ca2+/calmodulin binds to and modulates P/Q-type calcium channels. Nature 399, 155–159 (1999).
Imredy, J. P. & Yue, D. T. Mechanism of Ca(2+)-sensitive inactivation of L-type Ca2+ channels. Neuron 12, 1301–1318 (1994).
Diwan, J. J., Daze, M., Richardson, R. & Aronson, D. Kinetics of Mg2+ flux into rat liver mitochondria. Biochemistry 18, 2590–2595 (1979).
Brierley, G. P., Davis, M. & Jung, D. W. Respiration-dependent uptake and extrusion of Mg2+ by isolated heart mitochondria. Arch. Biochem. Biophys. 253, 322–332 (1987).
Bragadin, M., Pozzan, T. & Azzone, G. F. Kinetics of Ca2+ carrier in rat liver mitochondria. Biochemistry 18, 5972–5978 (1979).
Bernardi, P. Mitochondrial transport of cations: channels, exchangers, and permeability transition. Physiol. Rev. 79, 1127–1155 (1999).
Gati, C., Stetsenko, A., Slotboom, D. J., Scheres, S. H. W. & Guskov, A. The structural basis of proton driven zinc transport by ZntB. Nat. Commun. 8, 1313 (2017).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Chun, E. et al. Fusion partner toolchest for the stabilization and crystallization of G protein-coupled receptors. Structure 20, 967–976 (2012).
Ren, F. et al. X-ray structures of the high-affinity copper transporter Ctr1. Nat. Commun. 10, 1386 (2019).
Deng, Z. et al. Cryo-EM structure of a proton-activated chloride channel TMEM206. Sci. Adv. 7, eabe5983 (2021).
He, Z. et al. Structural and functional analysis of human pannexin 2 channel. Nat. Commun. 14, 1712 (2023).
Khan, M. B. et al. Structural and functional characterization of the N-terminal domain of the yeast Mg2+ channel Mrs2. Acta Crystallogr. D 69, 1653–1664 (2013).
Stetsenko, A., Stehantsev, P., Dranenko, N. O., Gelfand, M. S. & Guskov, A. Structural and biochemical characterization of a novel ZntB (CmaX) transporter protein from Pseudomonas aeruginosa. Int. J. Biol. Macromol. 184, 760–767 (2021).
Payandeh, J. et al. Probing structure–function relationships and gating mechanisms in the CorA Mg2+ transport system. J. Biol. Chem. 283, 11721–11733 (2008).
Nordin, N. et al. Exploring the structure and function of Thermotoga maritima CorA reveals the mechanism of gating and ion selectivity in Co2+/Mg2+ transport. Biochem. J. 451, 365–374 (2013).
Fan, M. et al. Structure and mechanism of the mitochondrial Ca(2+) uniporter holocomplex. Nature 582, 129–133 (2020).
Perry, S. W., Norman, J. P., Barbieri, J., Brown, E. B. & Gelbard, H. A. Mitochondrial membrane potential probes and the proton gradient: a practical usage guide. Biotechniques 50, 98–115 (2011).
Li, M. et al. Molecular basis of Mg(2+) permeation through the human mitochondrial Mrs2 channel. Nat. Commun. 14, 4713 (2023).
Lai, L. T. F., Balaraman, J., Zhou, F. & Matthies, D. Cryo-EM structures of human magnesium channel MRS2 reveal gating and regulatory mechanisms. Nat. Commun. 14, 7207 (2023).
Barish, M. E. A transient calcium-dependent chloride current in the immature Xenopus oocyte. J. Physiol. 342, 309–325 (1983).
Hartzell, C., Putzier, I. & Arreola, J. Calcium-activated chloride channels. Annu Rev. Physiol. 67, 719–758 (2005).
Miledi, R. & Parker, I. Chloride current induced by injection of calcium into Xenopus oocytes. J. Physiol. 357, 173–183 (1984).
Yao, Y. & Tsien, R. Y. Calcium current activated by depletion of calcium stores in Xenopus oocytes. J. Gen. Physiol. 109, 703–715 (1997).
White, M. M. & Aylwin, M. Niflumic and flufenamic acids are potent reversible blockers of Ca2(+)-activated Cl- channels in Xenopus oocytes. Mol. Pharmacol. 37, 720–724 (1990).
Jung, D. W., Chapman, C. J., Baysal, K., Pfeiffer, D. R. & Brierley, G. P. On the use of fluorescent probes to estimate free Mg2+ in the matrix of heart mitochondria. Arch. Biochem. Biophys. 332, 19–29 (1996).
Jung, D. W., Panzeter, E., Baysal, K. & Brierley, G. P. On the relationship between matrix free Mg2+ concentration and total Mg2+ in heart mitochondria. Biochim. Biophys. Acta 1320, 310–320 (1997).
Corkey, B. E., Duszynski, J., Rich, T. L., Matschinsky, B. & Williamson, J. R. Regulation of free and bound magnesium in rat hepatocytes and isolated mitochondria. J. Biol. Chem. 261, 2567–2574 (1986).
Rutter, G. A., Osbaldeston, N. J., McCormack, J. G. & Denton, R. M. Measurement of matrix free Mg2+ concentration in rat heart mitochondria by using entrapped fluorescent probes. Biochem. J. 271, 627–634 (1990).
Crompton, M., Capano, M. & Carafoli, E. Respiration-dependent efflux of magnesium ions from heart mitochondria. Biochem. J. 154, 735–742 (1976).
Akerman, K. E. Inhibition and stimulation of respiration-linked Mg2+ efflux in rat heart mitochondria. J. Bioenerg. Biomembr. 13, 133–139 (1981).
Pressman, B. C. Biological applications of ionophores. Annu. Rev. Biochem. 45, 501–530 (1976).
Benz, R. & McLaughlin, S. The molecular mechanism of action of the proton ionophore FCCP (carbonylcyanide p-trifluoromethoxyphenylhydrazone). Biophys. J. 41, 381–398 (1983).
Garg, V. et al. The mechanism of MICU-dependent gating of the mitochondrial Ca(2+)uniporter. Elife 10, e69312 (2021).
Murphy, E. & Eisner, D. A. Regulation of intracellular and mitochondrial sodium in health and disease. Circ. Res. 104, 292–303 (2009).
Diwan, J. J. Mitochondrial transport of K+ and Mg2+. Biochim. Biophys. Acta 895, 155–165 (1987).
Gunter, T. E., Buntinas, L., Sparagna, G., Eliseev, R. & Gunter, K. Mitochondrial calcium transport: mechanisms and functions. Cell Calcium 28, 285–296 (2000).
Nicholls, D. G. Mitochondria and calcium signaling. Cell Calcium 38, 311–317 (2005).
Rizzuto, R., De Stefani, D., Raffaello, A. & Mammucari, C. Mitochondria as sensors and regulators of calcium signalling. Nat. Rev. Mol. Cell Biol. 13, 566–578 (2012).
Garlid, K. D. & Paucek, P. Mitochondrial potassium transport: the K(+) cycle. Biochim. Biophys. Acta 1606, 23–41 (2003).
Kulawiak, B. & Szewczyk, A. Current challenges of mitochondrial potassium channel research. Front Physiol. 13, 907015 (2022).
Szabo, I. & Zoratti, M. Mitochondrial channels: ion fluxes and more. Physiol. Rev. 94, 519–608 (2014).
Clapham, D. E. Calcium signaling. Cell 131, 1047–1058 (2007).
Jegla, T. & Salkoff, L. A novel subunit for shal K+ channels radically alters activation and inactivation. J. Neurosci. 17, 32–44 (1997).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D 74, 519–530 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360, 376 (1996).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Morin, A. et al. Collaboration gets the most out of software. Elife 2, e01456 (2013).
Tsai, C. W. & Tsai, M. F. Electrical recordings of the mitochondrial calcium uniporter in Xenopus oocytes. J. Gen. Physiol. 150, 1035–1043 (2018).
Acknowledgements
This work was partly supported by National Institutes of Health grants R01NS109307 and R01GM143440 (to P.Y.) and R01GM144485 (to M.T.). We thank T.-Y. Liu and Y.-L. Huang for technical assistance. We thank the staff scientists at Washington University Center for Cellular Imaging for data acquisition. Some of this work was performed at the Simons EM Center at the New York Structural Biology Center, with major support from the Simons Foundation (SF349247).
Author information
Authors and Affiliations
Contributions
Z.H. performed protein biochemistry. Z.H., J.M. and J.Z. collected cryo-EM images. J.M., J.Z. and P.Y. conducted cryo-EM structure determination and analysis. Y.T. and C.T. performed electrophysiology experiments. M.T. and P.Y. supervised the project and wrote the paper. All authors edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Motoyuki Hattori and Weiwei Wang for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM data processing and validation.
a-c, Image processing and map validation of MRS2Mg (a), MRS2EDTA (b), and MRS2Ca (c). Representative micrographs and 2D classes are shown. The final 3D reconstructions are colored by local resolution. Also shown are Fourier shell correlations (FSC) and orientation distribution of particles used in the final reconstruction.
Extended Data Fig. 2 Cryo-EM density.
a, Ribbon representation of a single subunit of MRS2Mg and cryo-EM density. b, Segments of the final refined model of MRS2Mg and the corresponding cryo-EM densities.
Extended Data Fig. 3 Structural comparison of human MRS2 and TmCorA.
a-b, A single subunit of human MRS2 (a) and TmCorA (b, PDB ID: 4I0U). The N-terminal α/β domain is also highlighted for comparison. c, Divalent ion binding sites in human MRS2 (left panels) and TmCorA (right panels). Overlays of sites 1 and 2 are also shown (MRS2 in green and red; CorA in gray).
Extended Data Fig. 4 Sequence alignment of MRS2 channels.
Multiple protein sequences, including Homo sapiens MRS2 (hMRS2, NCBI sequence: NP_065713.1), Mus musculus MRS2 (mMRS2, NCBI sequence: NP_001013407.2), Rattus norvegicus MRS2 (rMRS2, NCBI sequence: NP_076491.1), Danio rerio MRS2 (zMRS2, NCBI sequence: XP_693621.5), Saccharomyces cerevisiae MRS2 (ScMRS2, NCBI sequence: NP_014979.1), Schizosaccharomyces pombe MRS2 (SpMRS2, NCBI sequence: NP_596358.1), Arabidopsis thaliana MRS2 (AtMRS2, NCBI sequence: AAM62917.1),and Thermotoga maritima CorA (TmCorA, NCBI sequence: WP_004081315.1). Secondary structure elements on the basis of hMRS2 are indicated above the sequences. Critical amino acids are highlighted.
Extended Data Fig. 5 Ion densities.
a-b, Cryo-EM densities in the three ion binding sites for Mg2+ (a) and Ca2+ (b). Mg2+ and Ca2+ are shown as magenta and yellow spheres, respectively. c, Densities near the assigned Cl− binding site from Li et al. (EMD-35630 and EMD-35631). d, Densities near the two additional Mg2+ binding sites in the pore from Lai et al. (EMD-41624).
Extended Data Fig. 7 Control experiments for MRS2 function.
a, MRS2-specific Mg2+ currents. The traces show that Xenopus oocytes without MRS2RS expression (uninjected) or expressing the transmembrane subunits of the mitochondrial calcium uniporter (hMEWT) exhibit no Mg2+ currents. b, The effect of BAPTA injection on MRS2RS Ca2+ currents. 5 nmol of BAPTA was injected into oocytes through a third electrode as indicated by arrows. The maximal Ca2+ current amplitudes before and 40 s after BAPTA injection were compared, showing smaller currents after BAPTA injection, as summarized in the paired dot plot. This mimics the effect of mutating site 3 to abolish divalent cation binding in MRS2’s matrix domain. Similar results were obtained when Ca2+ was applied 2.5 min after BAPTA injection. Statistical analyses were performed using paired two-tailed t-test. c, Isolation of Ca2+-activated Cl− currents (CACC). The I-V relationship of Mg2+-conducting TmCorA was obtained before and after adding 1 mM niflumic acid (NA) in the same oocyte. Recordings from 6 oocytes were summed to create the ensemble I-V curve. Subtracting the I-V curve with NA from the I-V curve without NA reveals the outwardly-rectifying CACC that reverses at -20 mV. d-e, The effect of NA on the Ca2+ currents from TmCorA (d) or MRS2RS (e). f, Recovery of MRS2 from inactivation. Switching the solution directly from Ca2+ to Na+ leads to slow Na+-current increase (first red bar). However, Na+ currents rise more rapidly following a 1-min washout (second red bar). In this trace, and in a subset ( ~ 20%) of our MRS2RS recordings, we observed that Ca2+ currents would inactivate following a double-exponential time course with some residual currents. The residual currents might reflect Ca2+ currents from native Ca2+ channels in Xenopus oocytes. The double-exponential Ca2+ current decay suggests that there might be multiple intermediate states in the inactivation process, but the molecular nature of these states remain unclear currently.
Extended Data Fig. 8 MRS2 function without R332S.
a-b, MRS2WT shows Ca2+ inactivation (a) and conducts Na+ currents that are not inhibited by Mg2+ (b). c-d, The effects of introducing quad mutations into MRS2WT, as shown in representative traces (c) and a bar chart (d). Statistical analyses were performed using two-tailed t-test. n.s.: no significance. The exact number of independent repeats was provided above each bar. Data were presented as mean ± S.E.M.
Supplementary information
Source data
Source Data Fig. 3 (download XLSX )
Numerical source data.
Source Data Fig. 4 (download XLSX )
Numerical source data.
Source Data Fig. 5 (download XLSX )
Numerical source data.
Source Data Extended Data Fig.7 (download XLSX )
Numerical source data.
Source Data Extended Data Fig.8 (download XLSX )
Numerical source data.
Source Data Fig.3 (download PDF )
Unprocessed western blots.
Source Data Fig.5 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, Z., Tu, YC., Tsai, CW. et al. Structure and function of the human mitochondrial MRS2 channel. Nat Struct Mol Biol 32, 459–468 (2025). https://doi.org/10.1038/s41594-024-01420-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41594-024-01420-5