Abstract
Proteostasis in mammalian oocytes is vital for successful reproduction. The cytoplasmic lattices (CPLs) of oocytes store essential maternal proteins for early embryo development. Here we show that PADI6, a core component of CPLs, forms a conserved ternary complex that we term MPU for maternal PADI6–UHRF1–UBE2D. The MPU complex regulates protein ubiquitination during oocyte maturation and early embryogenesis. We determined the cryo-electron microscopy structure of MPU and show that 86% (25/29) of clinically identified PADI6 missense variants disrupt MPU assembly, revealing a potential molecular mechanism linking dysregulation of ubiquitination on oocytes to abnormal embryonic development. Mechanistically, PADI6, with the assistance of UHRF1, sequesters UBE2D to prevent ubiquitin transfer from E2 to relevant substrate proteins, thereby suppressing the ubiquitination cascade. Therefore, our findings implicate PADI6 in the regulation of proteostasis by controlling the ubiquitination cascade, expanding our understanding of PADI6-dependent regulation of oocyte maturation and early embryogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The structural coordinates and factors were deposited to the Protein Data Bank under accession code 9LPK and the cryo-EM maps were deposited to the EM Data Bank under accession code EMD-63270. The MS proteomics data were deposited to the ProteomeXchange Consortium through the iProX partner repository60,61 with the dataset identifier PXD060230. Source data are provided with this paper.
References
Adam Bohnert, K. & Kenyon, C. A lysosomal switch triggers proteostasis renewal in the immortal C. elegans germ lineage. Nature 551, 629–633 (2017).
Zaffagnini, G. et al. Mouse oocytes sequester aggregated proteins in degradative super-organelles. Cell 187, 1109–1126 (2024).
Sala, A. J. & Morimoto, R. I. Protecting the future: balancing proteostasis for reproduction. Trends Cell Biol. 32, 202–215 (2022).
Jentoft, I. M. A. et al. Mammalian oocytes store proteins for the early embryo on cytoplasmic lattices. Cell 186, 5308–5327 (2023).
Rosswag de Souza, S., Böke, E. & Zaffagnini, G. Proteostasis in cellular dormancy: lessons from yeast to oocytes. Trends Biochem. Sci. 50, 646–662 (2025).
Zaffagnini, G., Solé, M., Duran, J. M., Polyzos, N. P. & Böke, E. The proteostatic landscape of healthy human oocytes. EMBO J. 44, 4611–4630 (2025).
Wang, S. et al. Proteome of mouse oocytes at different developmental stages. Proc. Natl Acad. Sci. USA 107, 17639–17644 (2010).
Liu, X. et al. UHRF1 targets DNMT1 for DNA methylation through cooperative binding of hemi-methylated DNA and methylated H3K9. Nat. Commun. 4, 1563 (2013).
Uemura, S. et al. UHRF1 is essential for proper cytoplasmic architecture and function of mouse oocytes and derived embryos. Life Sci. Alliance 6, e202301904 (2023).
Zhang, T., Liu, P., Yao, G., Zhang, X. & Cao, C. A complex heterozygous mutation in PADI6 causes early embryo arrest: a case report. Front. Genet. 13, 1104085 (2023).
Cao, G. et al. A novel homozygous variant in PADI6 is associate with human cleavage-stage embryonic arrest. Front. Genet. 14, 1243230 (2023).
Eggermann, T., Kadgien, G., Begemann, M. & Elbracht, M. Biallelic PADI6 variants cause multilocus imprinting disturbances and miscarriages in the same family. Eur. J. Hum. Genet. 29, 575–580 (2021).
Qian, J. et al. Biallelic PADI6 variants linking infertility, miscarriages, and hydatidiform moles. Eur. J. Hum. Genet. 26, 1007–1013 (2018).
Tannorella, P. et al. Germline variants in genes of the subcortical maternal complex and multilocus imprinting disturbance are associated with miscarriage/infertility or Beckwith–Wiedemann progeny. Clin. Epigenetics 14, 43 (2022).
Cubellis, M. V. et al. Loss-of-function maternal-effect mutations of PADI6 are associated with familial and sporadic Beckwith–Wiedemann syndrome with multi-locus imprinting disturbance. Clin. Epigenetics 12, 139 (2020).
Begemann, M. et al. Maternal variants in NLRP and other maternal effect proteins are associated with multilocus imprinting disturbance in offspring. J. Med. Genet. 55, 497–504 (2018).
Xu, Y. et al. Mutations in PADI6 cause female infertility characterized by early embryonic arrest. Am. J. Hum. Genet. 99, 744–752 (2016).
Zheng, W. et al. New biallelic mutations in PADI6 cause recurrent preimplantation embryonic arrest characterized by direct cleavage. J. Assist. Reprod. Genet. 37, 205–212 (2020).
Dong, J. et al. Novel biallelic mutations in PADI6 in patients with early embryonic arrest. J. Hum. Genet. 67, 285–293 (2022).
Xu, Y. et al. Novel homozygous PADI6 variants in infertile females with early embryonic arrest. Front. Cell Dev. Biol. 10, 819667 (2022).
Wang, X. et al. Novel mutations in genes encoding subcortical maternal complex proteins may cause human embryonic developmental arrest. Reprod. Biomed. Online 36, 698–704 (2018).
Rezaei, M. et al. Novel pathogenic variants in NLRP7, NLRP5, and PADI6 in patients with recurrent hydatidiform moles and reproductive failure. Clin. Genet. 99, 823–828 (2021).
Eggermann, T. et al. Trans-acting genetic variants causing multilocus imprinting disturbance (MLID): common mechanisms and consequences. Clin. Epigenetics 14, 41 (2022).
Maddirevula, S. et al. The human knockout phenotype of PADI6 is female sterility caused by cleavage failure of their fertilized eggs. Clin. Genet. 91, 344–345 (2017).
Liu, J. et al. Two novel mutations in PADI6 and TLE6 genes cause female infertility due to arrest in embryonic development. J. Assist. Reprod. Genet. 38, 1551–1559 (2021).
Zhang, M. et al. Case report: human early embryonic arrest in a consanguineous Chinese family caused by a novel missense variant of PADI6. QJM 116, 784–786 (2023).
Schwenk, F., Baron, U. & Rajewsky, K. A cre-transgenic mouse strain for the ubiquitous deletion of loxP-flanked gene segments including deletion in germ cells. Nucleic Acids Res. 23, 5080–5081 (1995).
Esposito, G. et al. Peptidylarginine deiminase (PAD) 6 is essential for oocyte cytoskeletal sheet formation and female fertility. Mol. Cell. Endocrinol. 273, 25–31 (2007).
Arita, K. et al. Structural basis for Ca2+-induced activation of human PAD4. Nat. Struct. Mol. Biol. 11, 777–783 (2004).
Williams, J. P. C. et al. Structural insight into the function of human peptidyl arginine deiminase 6. Comput. Struct. Biotechnol. J. 23, 3258–3269 (2024).
Ranaivoson, F. M. et al. Crystal structure of human peptidylarginine deiminase type VI (PAD6) provides insights into its inactivity. IUCrJ 11, 395–404 (2024).
Bertran, M. T. et al. A cyclic peptide toolkit reveals mechanistic principles of peptidylarginine deiminase IV regulation. Nat. Commun. 15, 9746 (2024).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Avvakumov, G. V. et al. Structural basis for recognition of hemi-methylated DNA by the SRA domain of human UHRF1. Nature 455, 822–825 (2008).
Williams, J. P. C. & Walport, L. J. PADI6: what we know about the elusive fifth member of the peptidyl arginine deiminase family. Philos. Trans. R. Soc. B 378, 20220242 (2023).
Chi, P. et al. Structural basis of the subcortical maternal complex and its implications in reproductive disorders. Nat. Struct. Mol. Biol. 31, 115–124 (2024).
Chi, P. et al. Cryo-EM structure of the human subcortical maternal complex and the associated discovery of infertility-associated variants. Nat. Struct. Mol. Biol. 31, 1798–1807 (2024).
Roman-Trufero, M. & Dillon, N. The UBE2D ubiquitin conjugating enzymes: potential regulatory hubs in development, disease and evolution. Front. Cell Dev. Biol. 10, 1058751 (2022).
Foster, B. M. et al. Critical role of the UBL domain in stimulating the E3 ubiquitin ligase activity of UHRF1 toward chromatin. Mol. Cell 72, 739–752 (2018).
Stewart, M. D., Ritterhoff, T., Klevit, R. E. & Brzovic, P. S. E2 enzymes: more than just middlemen. Cell Res. 26, 423–440 (2016).
Buetow, L. et al. Activation of a primed RING E3–E2–ubiquitin complex by non-covalent ubiquitin. Mol. Cell 58, 297–310 (2015).
Shen, D. et al. Novel cell- and tissue-based assays for detecting misfolded and aggregated protein accumulation within aggresomes and inclusion bodies. Cell Biochem. Biophys. 60, 173–185 (2011).
Gan, J., Leestemaker, Y., Sapmaz, A. & Ovaa, H. Highlighting the proteasome: using fluorescence to visualize proteasome activity and distribution. Front. Mol. Biosci. 6, 14 (2019).
Cecere, F., Argenziano, L., Pagano, A. & Riccio, A. New insights into oocyte cytoplasmic lattice-associated proteins. Trends Genet. 40, 880–890 (2024).
Giaccari, C. et al. A maternal-effect Padi6 variant causes nuclear and cytoplasmic abnormalities in oocytes, as well as failure of epigenetic reprogramming and zygotic genome activation in embryos. Genes Dev. 38, 131–150 (2024).
Li, Y. et al. Stella safeguards the oocyte methylome by preventing de novo methylation mediated by DNMT1. Nature 564, 136–140 (2018).
Meng, T. G. et al. NLRP14 safeguards calcium homeostasis via regulating the K27 ubiquitination of Nclx in oocyte-to-embryo transition. Adv. Sci. 10, e2301940 (2023).
Zhang, W. et al. NLRP14 deficiency causes female infertility with oocyte maturation defects and early embryonic arrest by impairing cytoplasmic UHRF1 abundance. Cell Rep. 42, 113531 (2023).
Yan, R. et al. Dynamics of DNA hydroxymethylation and methylation during mouse embryonic and germline development. Nat. Genet. 55, 130–143 (2023).
Han, Z. et al. The subcortical maternal complex modulates the cell cycle during early mammalian embryogenesis via 14-3-3. Nat. Commun. 15, 8887 (2024).
Xu, C. et al. The subcortical maternal complex safeguards mouse oocyte-to-embryo transition by preventing nuclear entry of SPIN1. Nat. Struct. Mol. Biol. 32, 1358–1372 (2025).
Zhang, G. et al. UBE2D3 functions in mouse oocyte meiotic maturation. FASEB J. 39, e70375 (2025).
Ratliff, M. et al. MIB-1 is required for spermatogenesis and facilitates LIN-12 and GLP-1 activity in Caenorhabditis elegans. Genetics 209, 173–193 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Jamali, K. et al. Automated model building and protein identification in cryo-EM maps. Nature 628, 450–457 (2024).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Bruderer, R. et al. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol. Cell. Proteomics 14, 1400–1410 (2015).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Acknowledgements
We thank BioRender.com for providing resources used to create Figs. 1a and 7 in this publication. We thank the members (D. Du, R. Wang, X. Wu and W. Li) of the MS Platform (Frontiers Science Center for Disease-related Molecular Network, West China Hospital) for data acquisition and analysis. We thank the SKLB West China Cryo-EM Center of Sichuan University Cryo-EM support. This study was supported by the Major Program of Shenzhen Bay Laboratory (C1012513007) and the National Natural Science Foundation of China (31971132 and 32171211 to D.D.; 82001540 to Y.L.). The funders had no role in study design, data collection and analysis, publication decisions or manuscript preparation.
Author information
Authors and Affiliations
Contributions
D.D. and S.G. supervised the project and edited the paper. Jinhong Li and Y.L. designed the study, conducted most of the experiments, analyzed the data, prepared the figures and wrote the paper. Z.X. and W.M. performed the MS experiments, analyzed the data and prepared the figures. Jinhong Li, P.C., Q.Q. and S.L. performed the construct design, protein purification, EM sample preparation, pulldown assays, co-IP, immunoblotting and data collection. Jinhong Li, P.C., Q.Q. and S.J. performed the ubiquitination activity assays, structure analysis and biochemical data analysis. Y.L. and Z.Z. collected oocytes and embryos and performed the immunofluorescence staining and immunoblotting. Jinhong Li and Jialu Li were responsible for the EM data collection, data processing and model building. Z.H. performed the IP. Q.L., J.C., X.W., L.G. and L.L. contributed to the data analysis and experiment design. W.H., L.D. and J.H. participated in the study design and supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Andrea Riccio and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Dimitris Typas, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The maternal effect factor PADI6 is essential for early embryonic development in mice.
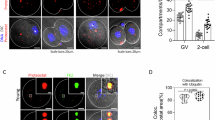
(a) Strategy for the generation of Padi6 knockout mice via the Cre-loxP system. (b) Western blotting showing PADI6 protein levels in GV oocytes from female Padi6Null, Padi6+/−, WT mice, with β-actin as the loading control. Three biologically repeated experiments were performed. (c) Immunofluorescent staining showing PADI6 protein levels in GV oocytes from female Padi6Null and WT mice. Three biologically repeated experiments were performed. Scale bar, 10 μm. (d) Fertility of female Padi6Null mice. Litter sizes of female Padi6+/− (cre, flox/+) and Padi6Null (cre, flox/flox) female mice after mating with Padi6+/+ (flox/flox), Padi6+/− (cre, flox/+) and Padi6Null (cre, flox/flox) males are presented as the means ± s.d. *Number of litters. †Number of females. (e) Representative images of WT and Padi6Null oocytes during oocyte maturation. Scale bars, 1 mm. (f) Percentages of oocytes that underwent GVBD and extruded their 1st polar body during in vitro maturation. WT (n = 122) and Padi6Null (n = 75) fully grown oocytes were collected after PMSG and HCG injection. (g) Representative images of preimplantation embryos derived from WT and Padi6Null female mice. Scale bars, 1 mm. (h) Development rates of 2-cell embryos and blastocysts after zygote collection from Padi6Null and WT mice. Number of zygotes: WT (n = 70), Padi6Null (n = 53). Rate of 2-cell/blastocyst = number of 2-cell/blastocyst/number of zygotes. In (f) and (h), n = number of oocytes analyzed, and data are presented as means ± s.d. P value were determined by unpaired, two-tailed t tests. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; n.s., no significance. WT and Padi6Null differed significantly in blastocyst development (P < 0.0001), whereas all other comparisons were not significant. Oocyte maturation potential: GVBD, WT v.s. Padi6Null, P = 0.3744; MII, WT v.s. Padi6Null, P = 0.3627. Embryo development potential:2PN, WT v.s. Padi6Null, both groups were zygotes, P = 1.0; 2-cell, WT v.s. Padi6Null, P = 0.1977; blastocyst, WT v.s. Padi6Null, P < 0.0001.
Extended Data Fig. 2 Comparative proteomic analysis of WT and Padi6Null oocytes.
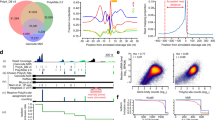
(a) Venn diagram illustrating the overlap and unique proteins identified in WT and Padi6Null oocytes. (b) Principal component analysis (PCA) of proteomic profiles demonstrates clear clustering of WT (blue) and Padi6Null (red) oocytes. (c) Bar plot illustrating the number of significantly upregulated (blue) and downregulated (red) proteins in Padi6Null oocytes compared to WT. Differential expression was assessed using an unpaired Student’s t-test (two-sided). Proteins with log2 (fold change) > 0.5 or log2 (fold change) < −0.5 and P < 0.05 were considered significantly upregulated or downregulated, respectively. (d) Volcano plot illustrating the differentially expressed proteins in Padi6Null compared to WT oocytes. Differential expression was assessed using an unpaired Student’s t-test (two-sided); proteins meeting log2 (fold change) > 0.5 or log2 (fold change) < −0.5 with P < 0.05 were highlighted as significantly upregulated (blue) or downregulated (red), respectively. Key proteins are labeled as indicated. (e-g) Venn diagrams showing the overlap of quantified proteins (e), downregulated proteins (f), and upregulated proteins (g) in Padi6Null oocytes between this study (MII stage) and Jentoft et al. (GV stage). (h) Scatter plot comparing fold changes of differentially expressed proteins from the MII stage in the GV stage. Proteins are categorized as showing consistent or inconsistent trends, and as significant or insignificant in the GV stage. (i, j) GO molecular function enrichment analysis of downregulated (i) and upregulated (j) proteins in Padi6Null oocytes at the MII stage (this study, red) and GV stage (Jentoft et al., blue). GO enrichment was performed using clusterProfiler::enrichGO (hypergeometric test), with P < 0.05 as the significance threshold. No adjustments for multiple comparisons were applied. (k) Heatmap showing the expression of shared downregulated proteins between MII- and GV-stage Padi6Null oocytes. (l) GO molecular function enrichment analysis of shared downregulated proteins between the MII and GV stages. GO enrichment was performed using clusterProfiler::enrichGO (hypergeometric test), with P < 0.05 as the significance threshold. No adjustments for multiple comparisons were applied.
Extended Data Fig. 3 In vitro reconstitution of human MPU from HEK293F cells.
(a) Representative SEC (Superose 6 Increase 10/300) of the human MPU complex purified from 293F cells. More than five biologically independent experiments were performed (n > 5). The column was calibrated with thyroglobulin (669 kDa), ferritin (440 kDa), and aldolase (158 kDa). The black box highlights the referenced protein from the bands of the MPU member, representing the relative mobility of endogenous UBE2D paralogs in HEK293F cells, as identified by mass spectrometry. The sequences of the unique peptides identified in the proteomic analysis are highlighted in red. The detailed information was described in Supplementary Table 3. (b) Protein identity matrix of UBE2D from humans and mice generated via the UniProt alignment tool. (c) In vitro reconstitution of human PADI6-UHRF1-UBE2D2. Purified Strep-hUHRF1 (SRA-RING construct) was incubated separately with His-hPADI6 and His-hUBE2D2, as well as in a combined mixture of all three proteins. The mixtures were pulled down using Strep resin. SDS-PAGE analysis of the eluates and input samples was performed to separate proteins, which were stained with Coomassie blue to assess their interactions. Three biologically repeated experiments were performed. (d) Flowchart of cryo-EM data processing for the complex of second peak contains nucleosomes. Details are described in the Methods section.
Extended Data Fig. 4 Flowchart of cryo-EM data processing for the human MPU complex.
(a) Representative micrograph of the cryo-EM data collection from 2,191 micrographs. (b) Workflow of the cryo-EM data processing. Details are described in the Methods section. (c) Representative micrograph of the 2D classification generated from the cryo-EM particles of human MPU complex. (d) Particle angular distribution for the map of the human MPU complex. (e) Gold-standard Fourier shell correlation (FSC) curve for the refined map of the human MPU complex.
Extended Data Fig. 5 Segmented map density of the human MPU.
The atomic structures of PADI6, UHRF1 and UBE2D3 were fitted into the cryo-EM map (white).
Extended Data Fig. 6 Comparisons of MPU members with their homologs.
(a) Alignment of hPADI4 (gray, PDB:1WDA) and hPADI6 (light sky blue and dodger blue, this study). In the middle of the panel, the surface representation of the hPADI4 active site cleft superimposed with a cartoon representation of hPADI6. The substrate BAG as part of the hPADI4 structure is displayed in yellow. D673 and N598 of hPADI6 are shown with potential hydrogen bonds represented by black dashed lines. The RMSD values for the N-terminal, middle, and C-terminal domains of PADI are 1.0 Å, 1.0 Å, and 1.2 Å, respectively. (b) Alignment of hPADI6 (gray, PDB:9FMN) and hPADI6 (light sky blue and dodger blue, this study). In the middle of the panel, a detailed view of the dimer interface of hPADI6 is presented. The residues participating in interactions and hydrogen bonds are indicated (represented by dashed lines). The RMSD values for the N-terminal, middle, and C-terminal domains of PADI are 1.0 Å, 0.8 Å, and 1.1 Å, respectively. (c) The electrostatic potential mapped onto the accessible surface area of human MPU. Regions of negative potential are colored red, those of positive potential are colored blue, and neutral regions are shown in gray. (d) Alignment of hUHRF1SRA-DNA (sage green and tan, PDB: 3CLZ) and hUHRF1SRA (gray, this study). RMSD values for hUHRF1SRA is 1.0 Å.
Extended Data Fig. 7 Evaluating the impact of the disease-associated missense variants on human MPU stability.
(a) The localization of pathogenic variants is indicated in the domains of hPADI6. The variants associated with infertility in the carrier women are represented in red, and the variants associated with MLID in the offspring are represented in medium purple, respectively. (b) Co-immunoprecipitation assay and the effects on MPU assembly of pathogenic variants of PADI6. The assays were classified into four groups, labeled cluster 1 (b), cluster 2 (c), cluster 3 (c), and cluster 4 (d) based on the phenotype and amino acid sequence of the pathogenic mutation. hPADI6 (WT or mutants) and hUHRF1 (SRA-RING construct) were co-expressed in HEK293F cells. Cells were then lysed, immunoprecipitated with anti-Flag affinity gel, and immunoblotted for Flag-tagged hUHRF1 and Strep-tagged hPADI6. IP, immunoprecipitation. The panel beside (b–d) shows statistical data for the ratio of intensity changes of IP blots, which was utilized to determine the impact on the stability of human MPU. Data represent three biologically repeated experiments. Bars represent the means ± s.d., and dots indicate individual data points. P values were obtained by unpaired, two-tailed Student’s t tests and are presented at the top of each column.
Extended Data Fig. 8 hPADI6 inhibits the protein ubiquitination in vitro.
Coomassie blue staining image of histone H3 ubiquitination assay mediated by hUHRF1 in the absence or presence of hPADI6 or pathogenic variants of hPADI6 (N598S, S336R, D349G, R352C, Q589K, P632A, and R682Q). Three biologically repeated experiments were performed.
Extended Data Fig. 9 Sequence alignment of PADIs.
Sequence alignment of human PADI homologs and mouse PADI6. The residues involved in the interactions among the subunits of human MPU were denoted with blue and red triangles. The residues involved in the dimer interface of hPADI6 were denoted with black triangles. Sequence alignment was performed using multalin (http://multalin.toulouse.inra.fr) and visualized by ESPript (https://espript.ibcp.fr).
Extended Data Fig. 10 Sequence alignment of the MPU members.
(a) Sequence alignment between human UHRF1 and mouse UHRF1. (b) Sequence alignment of human and mouse UBE2D homologs. The residues involved in the interactions among the subunits of human MPU are denoted with blue and red triangles. Sequence alignment was performed using multalin (http://multalin.toulouse.inra.fr) and visualized by ESPript (https://espript.ibcp.fr).
Supplementary information
Supplementary Tables 1–4 (download XLSX )
Supplementary Table 1: Quantitative proteomic analysis of WT or Padi6Null oocytes. Supplementary Table 2: Proteomic analysis of PADI6-interacting proteins. Supplementary Table 3: Identification of the MPU member UBE2D by MS. Supplementary Table 4: Light microscopy reporting table.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download PDF )
Unprocessed gels and western blots.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed gels.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Lu, Y., Xia, Z. et al. The maternal PADI6–UHRF1–UBE2D complex regulates ubiquitination during oocyte maturation and embryogenesis. Nat Struct Mol Biol 33, 512–524 (2026). https://doi.org/10.1038/s41594-026-01758-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41594-026-01758-y