Abstract
Biomolecular condensates regulate essential biological processes relevant to health and disease. However, the mechanisms driving pathogenic condensate formation and their therapeutic targeting have not been fully elucidated. In amyotrophic lateral sclerosis and frontotemporal dementia caused by C9orf72 GGGGCC repeat expansions (c9ALS/FTD), the expanded repeat RNA and repeat-associated non-AUG translation products are key pathogenic factors. Here, we show that the GGGGCC-repeat RNA and poly(GR) form cocondensates in vitro and in cellulo. The G-quadruplex and hairpin structures of GGGGCC-repeat RNA act as scaffolds to accelerate liquid-to-solid phase transition and aggregation of poly(GR), with the hairpin structure promoting amorphous solid-like condensates in vitro and reducing poly(GR) mobility. The cocondensation of GGGGCC-repeat RNA and poly(GR) exacerbates nucleolar stress and cellular toxicity. Targeting both G-quadruplex and hairpin structures of GGGGCC-repeat RNA with small molecules diminishes poly(GR) aggregation and ameliorates cellular dysfunction. These findings expand our understanding of poly(GR) aggregation in c9ALS/FTD, highlight the importance of RNA structure in regulating protein aggregation and suggest that targeting the RNA scaffold may expand the druggable space of pathogenic condensates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data in this study are provided in the main text and Supplementary Information. Source data are provided with this paper.
References
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Emmanouilidis, L. et al. A solid β-sheet structure is formed at the surface of FUS droplets during aging. Nat. Chem. Biol. 20, 1044–1052 (2024).
Shen, Y. et al. The liquid-to-solid transition of FUS is promoted by the condensate surface. Proc. Natl Acad. Sci. USA 120, e2301366120 (2023).
Soto, C. & Pritzkow, S. Protein misfolding, aggregation, and conformational strains in neurodegenerative diseases. Nat. Neurosci. 21, 1332–1340 (2018).
Ross, C. A. & Poirier, M. A. Protein aggregation and neurodegenerative disease. Nat. Med. 10, S10–S17 (2004).
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: mission possible? Nat. Rev. Drug Discov. 13, 828–851 (2014).
Dang, C. V., Reddy, E. P., Shokat, K. M. & Soucek, L. Drugging the ‘undruggable’ cancer targets. Nat. Rev. Cancer 17, 502–508 (2017).
Tesei, G. et al. Conformational ensembles of the human intrinsically disordered proteome. Nature 626, 897–904 (2024).
Mitrea, D. M., Mittasch, M., Gomes, B. F., Klein, I. A. & Murcko, M. A. Modulating biomolecular condensates: a novel approach to drug discovery. Nat. Rev. Drug Discov. 21, 841–862 (2022).
Childs-Disney, J. L. et al. Targeting RNA structures with small molecules. Nat. Rev. Drug Discov. 21, 736–762 (2022).
Wadsworth, G. M. et al. RNA-driven phase transitions in biomolecular condensates. Mol.Cell 84, 3692–3705 (2024).
Jain, A. & Vale, R. D. RNA phase transitions in repeat expansion disorders. Nature 546, 243–247 (2017).
Banerjee, P. R., Milin, A. N., Moosa, M. M., Onuchic, P. L. & Deniz, A. A. Reentrant phase transition drives dynamic substructure formation in ribonucleoprotein droplets. Angew. Chem. Int. Ed. Engl. 56, 11354–11359 (2017).
Sanchez de Groot, N. et al. RNA structure drives interaction with proteins. Nat. Commun. 10, 3246 (2019).
Oldani, E. G. et al. The effect of G-quadruplexes on TDP43 condensation, distribution, and toxicity. Structure 33, 1294–1303 (2025).
Matsuo, K. et al. RNA G-quadruplexes form scaffolds that promote neuropathological α-synuclein aggregation. Cell 187, 6835–6848 (2024).
Wan, L. et al. Small CAG repeat RNA forms a duplex structure with sticky ends that promote RNA condensation. J. Am. Chem. Soc. 147, 3813–3822 (2025).
Ding, M. et al. A biophysical basis for the spreading behavior and limited diffusion of Xist. Cell 188, 978–997 (2025).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9orf72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Balendra, R. & Isaacs, A. M. C9orf72-mediated ALS and FTD: multiple pathways to disease. Nat. Rev. Neurol. 14, 544–558 (2018).
Rodriguez, C. M. & Todd, P. K. New pathologic mechanisms in nucleotide repeat expansion disorders. Neurobiol. Dis. 130, 104515 (2019).
Zu, T. et al. RAN proteins and RNA foci from antisense transcripts in C9orf72 ALS and frontotemporal dementia. Proc. Natl Acad. Sci. USA 110, E4968–E4977 (2013).
Park, J. et al. Poly(GR) interacts with key stress granule factors promoting its assembly into cytoplasmic inclusions. Cell Rep. 42, 112822 (2023).
Zhang, Y. J. et al. Heterochromatin anomalies and double-stranded RNA accumulation underlie C9orf72 poly(PR) toxicity. Science 363, eaav2606 (2019).
Sirozh, O. et al. Nucleolar stress caused by arginine-rich peptides triggers a ribosomopathy and accelerates aging in mice. Mol. Cell 84, 1527–1540 (2024).
Lee, K. H. et al. C9orf72 dipeptide repeats impair the assembly, dynamics, and function of membrane-less organelles. Cell 167, 774–788 (2016).
Cook, C. N. et al. C9orf72 poly(GR) aggregation induces TDP-43 proteinopathy. Sci. Transl. Med. 12, eabb3774 (2020).
Wiedner, H. J. & Giudice, J. It’s not just a phase: function and characteristics of RNA-binding proteins in phase separation. Nat. Struct. Mol. Biol. 28, 465–473 (2021).
Su, Z. M. et al. Discovery of a biomarker and lead small molecules to target r(GGGGCC)-associated defects in c9FTD/ALS. Neuron 83, 1043–1050 (2014).
Boeynaems, S. et al. Spontaneous driving forces give rise to protein–RNA condensates with coexisting phases and complex material properties. Proc. Natl Acad. Sci. USA 116, 7889–7898 (2019).
Chen, X. J. et al. Visualizing RNA dynamics in live cells with bright and stable fluorescent RNAs. Nat. Biotechnol. 37, 1287–1293 (2019).
Woo, J. A. et al. Loss of function CHCHD10 mutations in cytoplasmic TDP-43 accumulation and synaptic integrity. Nat. Commun. 8, 15558 (2017).
Megat, S. et al. Integrative genetic analysis illuminates ALS heritability and identifies risk genes. Nat. Commun. 14, 342 (2023).
Kamei, N. et al. Insulin-inspired hippocampal neuron-targeting technology for protein drug delivery. Proc. Natl Acad. Sci. USA 121, e2407936121 (2024).
Morelli, C. et al. RNA modulates hnRNPA1A amyloid formation mediated by biomolecular condensates. Nat. Chem. 16, 1052–1061 (2024).
Mackenzie, I. R. et al. Quantitative analysis and clinico-pathological correlations of different dipeptide repeat protein pathologies in C9orf72 mutation carriers. Acta Neuropathol. 130, 845–861 (2015).
Ortega, J. A. et al. CLIP-Seq analysis enables the design of protective ribosomal RNA bait oligonucleotides against C9orf72 ALS/FTD poly-GR pathophysiology. Sci. Adv. 9, eadf7997 (2023).
Chen, Z. S. et al. Mutant GGGGCC RNA prevents YY1 from binding to Fuzzy promoter which stimulates Wnt/β-catenin pathway in C9ALS/FTD. Nat. Commun. 14, 8420 (2023).
Celona, B. et al. Zfp106 binds to G-quadruplex RNAs and inhibits RAN translation and formation of RNA foci caused by G4C2 repeats. Proc. Natl Acad. Sci. USA 121, e2220020121 (2024).
Wang, Y. et al. Structural investigation of pathogenic RFC1 AAGGG pentanucleotide repeats reveals a role of G-quadruplex in dysregulated gene expression in CANVAS. Nucleic Acids Res. 52, 2698–2710 (2024).
Ratmeyer, L., Vinayak, R., Zhong, Y. Y., Zon, G. & Wilson, W. D. Sequence specific thermodynamic and structural properties for DNA•RNA duplexes. Biochemistry 33, 5298–5304 (1994).
Wang, Z. F. et al. The hairpin form of r(G4C2)exp in c9ALS/FTD is repeat-associated non-ATG translated and a target for bioactive small molecules. Cell Chem. Biol. 26, 179–190 (2019).
Liang, T. et al. In situ formation of biomolecular condensates as intracellular drug reservoirs for augmenting chemotherapy. Nat. Biomed. Eng. 8, 1469–1482 (2024).
Iglesias-Artola, J. M. et al. Charge-density reduction promotes ribozyme activity in RNA–peptide coacervates via RNA fluidization and magnesium partitioning. Nat. Chem. 14, 407–416 (2022).
Visser, B. S., Lipinski, W. P. & Spruijt, E. The role of biomolecular condensates in protein aggregation. Nat. Rev. Chem. 8, 686–700 (2024).
Zhang, H. Y. et al. RNA controls polyQ protein phase transitions. Mol. Cell 60, 220–230 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Jawerth, L. et al. Protein condensates as aging Maxwell fluids. Science 370, 1317–1323 (2020).
Goldberg, D. C. et al. Manipulating adult neural stem and progenitor cells with G-quadruplex ligands. ACS Chem. Neurosci. 11, 1504–1518 (2020).
Asamitsu, S. et al. CGG repeat RNA G-quadruplexes interact with FMRpolyG to cause neuronal dysfunction in fragile X-related tremor/ataxia syndrome. Sci. Adv. 7, eabd9440 (2021).
Kwon, I. et al. Poly-dipeptides encoded by the C9orf72 repeats bind nucleoli, impede RNA biogenesis, and kill cells. Science 345, 1139–1145 (2014).
Mizielinska, S. et al. C9orf72 repeat expansions cause neurodegeneration in Drosophila through arginine-rich proteins. Science 345, 1192–1194 (2014).
Zhang, Y. J. et al. Poly(GR) impairs protein translation and stress granule dynamics in C9orf72 associated frontotemporal dementia and amyotrophic lateral sclerosis. Nat. Med. 24, 1136–1142 (2018).
Lin, Y. et al. Toxic PR poly-dipeptides encoded by the C9orf72 repeat expansion target LC domain polymers. Cell 167, 789–802 (2016).
Cooper-Knock, J. et al. Sequestration of multiple RNA recognition motif-containing proteins by C9orf72 repeat expansions. Brain 137, 2040–2051 (2014).
Das, M. R. et al. Repeat-associated non-AUG translation induces cytoplasmic aggregation of CAG repeat-containing RNAs. Proc. Natl Acad. Sci. USA 120, e2215071120 (2023).
Guo, W. et al. Tuning material states and functionalities of G-quadruplex-modulated RNA–peptide condensates. J. Am. Chem. Soc. 145, 2375–2385 (2023).
Raguseo, F. et al. The ALS/FTD-related C9orf72 hexanucleotide repeat expansion forms RNA condensates through multimolecular G-quadruplexes. Nat. Commun. 14, 8272 (2023).
Krainer, G. et al. Reentrant liquid condensate phase of proteins is stabilized by hydrophobic and non-ionic interactions. Nat. Commun. 12, 1085 (2021).
Liu, X. J. et al. G-quadruplex-induced liquid–liquid phase separation in biomimetic protocells. J. Am. Chem. Soc. 143, 11036–11043 (2021).
Wang, Y. et al. Mutual promotion of co-condensation of KRAS G-quadruplex and a well-folded protein HMGB1. Nucleic Acids Res. 52, 288–299 (2024).
Mahendran, T. S., Wadsworth, G. M., Singh, A., Gupta, R. & Banerjee, P. R. Homotypic RNA clustering accompanies a liquid-to-solid transition inside the core of multi-component biomolecular condensates. Nat. Chem. 17, 1236–1246 (2025).
Niedner-Boblenz, A. et al. Intrinsically disordered RNA-binding motifs cooperate to catalyze RNA folding and drive phase separation. Nucleic Acids Res. 52, 14205–14228 (2024).
Binas, O. et al. Structural basis for the recognition of transiently structured AU-rich elements by Roquin. Nucleic Acids Res. 48, 7385–7403 (2020).
Majumder, S., Coupe, S., Fakhri, N. & Jain, A. Sequence-encoded intermolecular base pairing modulates fluidity in DNA and RNA condensates. Nat. Commun. 16, 4258 (2025).
Dignon, G. L., Zheng, W., Best, R. B., Kim, Y. C. & Mittal, J. Relation between single-molecule properties and phase behavior of intrinsically disordered proteins. Proc. Natl Acad. Sci. USA 115, 9929–9934 (2018).
Poudyal, M. et al. Intermolecular interactions underlie protein/peptide phase separation irrespective of sequence and structure at crowded milieu. Nat. Commun. 14, 6199 (2023).
Linsenmeier, M. et al. Dynamic arrest and aging of biomolecular condensates are modulated by low-complexity domains, RNA and biochemical activity. Nat. Commun. 13, 3030 (2022).
Dall’Agnese, A. et al. Proteolethargy is a pathogenic mechanism in chronic disease. Cell 188, 207–221 (2025).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (22225402 to D.H., 22374132 to P.G., 22507123 to L.W. and 32341017 to D.H.), Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (JYB2025XDXM602 to D.H.), Natural Science Foundation of Zhejiang Province (QKHM25B0501 to P.G.) and Leading Innovative and Entrepreneur Team Introduction Program of Zhejiang Province (2024R01005 to D.H.). We acknowledge the support from the Scientific Experiment Center from Hangzhou Institute of Medicine, Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
P.G. and D.H. conceptualized and designed the work. Y.L. performed the majority of the experiments and data analysis with M.S. and L.W. as assistants. Y.L., P.G. and D.H. wrote the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Marco Di Antonio, Norifumi Shioda and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Dimitris Typas, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 GGGGCC repeat RNA and R-DPRs form condensates in vitro.
a, Bright-field microscopic images of (GR)8-r(G4C2)4 incubates and (PR)8-r(G4C2)4 incubates at various DPR concentrations and 0.025 μM r(G4C2)4 RNA. b, Bright-field microscopic images of (GR)8-r(G4C2)4 incubates and (PR)8-r(G4C2)4 incubates at various DPR concentrations and 25 μM r(G4C2)4 RNA. The buffer condition is 100 mM KCl, 1 mM MgCl2 and 10 mM Tris-HCl (pH 7). The incubation time is 30 min. The microscopic experiments were repeated at least three times with similar results.
Extended Data Fig. 2 Phase behaviors of five DPRs and GGGGCC repeat RNA.
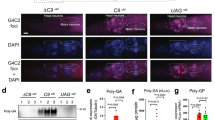
a, Microscopic images of (PA)8 and r(G4C2)4 incubates, (GA)8 and r(G4C2)4 incubates, and (GP)8 and r(G4C2)4 incubates, at 72/144/288 µM DPR and 2.5 µM RNA. The microscopic experiments were repeated at least three times with similar results. b, SPR sensorgrams show no binding of r(G4C2)4 for (PA)8, (GA)8 and (GP)8, respectively. c, Microscopic images of (GR)8 and rU24 incubates, and (PR)8 and rU24 incubates, at 36 µM DPR and 7.5/25/50 µM RNA. The microscopic experiments were repeated at least three times with similar results. The buffer condition in a-c is 1 mM MgCl2, 100 mM KCl, 10 mM Tris-HCl at pH 7. d, Microscopic images of live HEK293T cells transfected with (GR)29-r(G4C2)29 at 12 h (top) and 24 h (bottom). The microscopic experiments were repeated at least three times with similar results. e, Microscopic images of HEK293T cells transfected with (GR)29-r(G4C2)29 (top), (GR)29-r(GA4C)29 (middle), and (GP)29-r(G4C2)29 (bottom) at 24 h. The microscopic experiments were repeated at least three times with similar results. f, Quantification of DPR and RNA overlapping area per cell. Data are presented as mean ± s.d., n = 20 cells. Statistical analysis is performed using one-way ANOVA. ****P < 0.0001. g, Microscopic images of HT22 cells expressing (GR)29-r(G4C2)29 at 24 h after plasmid transfection. The microscopic experiments were repeated at least three times with similar results. Source data and exact P values are provided as a source data file.
Extended Data Fig. 3 Binding assays for five DPRs and r(G4C2)4 G4/hairpin.
a, SPR measuring the binding of five DPRs and r(G4C2)4 G4 in 100 mM K+. b, SPR measuring the binding of five DPRs and r(G4C2)4 hairpin in 5 mM Mg2+. The KD data are presented as mean ± s.d., n = 3 independent experiments. (PA)8, (GA)8 and (GP)8 have no measurable binding to r(G4C2)4 G4 and hairpin. Source data are provided as a source data file.
Extended Data Fig. 4 GGGGCC repeat RNA G4 and hairpin scaffolds regulate the morphology and fluidity of poly-GR condensates.
a, Bright-field microscopic images of (GR)8-G4 incubates (left) and (GR)8-HP incubates (right) at various (GR)8 and r(G4C2)4 RNA concentrations. The microscopic experiments were repeated at least three times with similar results. b, Phase diagrams of (GR)8-G4 incubates (left) and (GR)8-HP incubates (right) as shown in (a). c, Microscopic images show that the (GR)8-G4 condensates are spherical in shape and can fuse between droplets. The microscopic experiments were repeated at least three times with similar results. d, Fluorescence intensity of free NMM, NMM incubated with (GR)8-G4 condensates, and NMM incubated with (GR)8-HP condensates. e, Fluorescence intensity of free DOX, DOX incubated with (GR)8-G4 condensates, and DOX incubated with (GR)8-HP condensates. Source data are provided as a source data file.
Extended Data Fig. 5 GGGGCC repeat RNA G4 and hairpin spatially overlap with (GR)8 and reduce (GR)8 fluidity in the condensates.
a, The 3D reconstruction images of (GR)8-G4 condensates. b, The 3D reconstruction images of (GR)8-HP condensates. The (GR)8 concentration is 36 μM and the r(G4C2)4 RNA concentration is 2.5 μM. The microscopic experiments were repeated at least three times with similar results. c, The t1/2 of (GR)8 fluorescence recovery in the (GR)8-G4 condensates and (GR)8-HP condensates at 36 μM (GR)8 and 2.5/7.5 μM r(G4C2)4 RNA. The incubation time is 2 h. d, The t1/2 of (GR)8 fluorescence recovery in the (GR)8-G4 condensates and (GR)8-HP condensates at 36 μM (GR)8 and 2.5 μM r(G4C2)4 RNA, with incubation time of 2 h and 24 h. Data in c and d are presented as mean ± s.d., n = 3 condensates. The statistical analysis is performed using two-sided student’s t-test. *P < 0.05. Source data and exact P values are provided as a source data file.
Extended Data Fig. 6 Targeting GGGGCC repeat RNA G4 and hairpin structures with small molecules reduces cellular dysfunction.
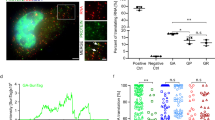
a, b, 1D 1H NMR spectra show binding of cPDS to r(G4C2)4 G4 (a), and binding of L4 to r(G4C2)4 hairpin (b). c-f, ITC thermograms show cPDS binding to r(G4C2)4 G4 (c) and r(G4C2)8 G4 (d), and L4 binding to r(G4C2)4 hairpin (e) and r(G4C2)8 hairpin (f). Data are presented as mean ± s.d., n = 3 independent experiments. g, h, Microscopic images show RNA foci in HEK293T cells expressing r(G4C2)29 only without treating ligand (-cPDS/L4), treated with 5 μM cPDS (+cPDS), 100 nM L4 (+L4), and 5 μM cPDS and 100 nM L4 (+cPDS/L4) (g), and quantification of number of RNA foci per cell by FISH (h). Data are presented as mean ± s.d., n = 30 cells. The microscopic experiments were repeated at least three times with similar results. i, Viability of HT22 cells without expressing (GR)29 and r(G4C2)29 (control), expressing r(G4C2)29 only, (GR)29 only, and (GR)29-r(G4C2)29. Data are presented as mean ± s.d., n = 3 biological replicates. j, Viability of HT22 cells expressing (GR)29-r(G4C2)29 without treating ligand (-cPDS/L4), treated with 5 μM cPDS (+cPDS), 100 nM L4 (+L4), and 5 μM cPDS and 100 nM L4 (+cPDS/L4). The viability is normalized to the control group. Data are presented as mean ± s.d., n = 3 biological replicates. Statistical analysis in h-j is performed using one-way ANOVA. ****P < 0.0001. ***P < 0.001. **P < 0.01. *P < 0.05. ns, not significant. Source data and exact P values are provided as a source data file.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–12 and Tables 1 and 2.
Supplementary Table 3 (download XLSX )
Light microscopy table.
Supplementary Data 1 (download XLSX )
Numerical source data for supplementary figures.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Y., Song, M., Wan, L. et al. Condensate protein aggregation in ALS/FTD is regulated by GGGGCC-repeat RNA scaffolds. Nat Struct Mol Biol (2026). https://doi.org/10.1038/s41594-026-01785-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41594-026-01785-9