Abstract
Advances in genomics have identified thousands of risk genes impacting human health and diseases, but the functions of these genes and their mechanistic contribution to disease are often unclear. Moving beyond identification to actionable biological pathways requires dissecting risk gene function and cell type-specific action in intact tissues. This gap can in part be addressed by in vivo Perturb-seq, a method that combines state-of-the-art gene editing tools for programmable perturbation of genes with high-content, high-resolution single-cell genomic assays as phenotypic readouts. Here we describe a detailed protocol to perform massively parallel in vivo Perturb-seq using several versatile adeno-associated virus (AAV) vectors and provide guidance for conducting successful downstream analyses. Expertise in mouse work, AAV production and single-cell genomics is required. We discuss key parameters for designing in vivo Perturb-seq experiments across diverse biological questions and contexts. We further detail the step-by-step procedure, from designing a perturbation library to producing and administering AAV, highlighting where quality control checks can offer critical go–no-go points for this time- and cost-expensive method. Finally, we discuss data analysis options and available software. In vivo Perturb-seq has the potential to greatly accelerate functional genomics studies in mammalian systems, and this protocol will help others adopt it to answer a broad array of biological questions. From guide RNA design to tissue collection and data collection, this protocol is expected to take 9–15 weeks to complete, followed by data analysis.
Key points
-
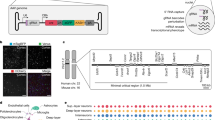
Versatile adeno-associated virus (AAV) vectors are combined with a transposon system to achieve high-level fast-onset expression of guide RNA perturbation libraries in live animals.
-
This protocol outlines strategies for designing the perturbation library, producing and administering AAV and performing downstream single-cell RNA sequencing and computational analyses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data included in this protocol are from the key reference publication11. The raw data generated in that key reference study are available on Mendeley Data (https://doi.org/10.17632/hvb39r62xw.1), NCBI Gene Expression Omnibus (GEO: GSE249416) and the Broad single cell portal (https://singlecell.broadinstitute.org/single_cell/study/SCP2443).
Code availability
The analysis pipeline used in the key reference publication is deposited in the GitHub repository (https://github.com/jinlabneurogenomics). The Python scripts referenced in this protocol are available in the Supplementary Information (Supplementary Softwares 1 and 2).
References
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882.e21 (2016).
Dixit, A. et al. Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016).
Jaitin, D. A. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-seq. Cell 167, 1883–1896.e15 (2016).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Schraivogel, D., Steinmetz, L. M. & Parts, L. Pooled genome-scale CRISPR screens in single cells. Annu. Rev. Genet. 57, 223–244 (2023).
Jin, X. et al. In vivo Perturb-Seq reveals neuronal and glial abnormalities associated with autism risk genes. Science 370, eaaz6063 (2020).
Kuzmin, D. A. et al. The clinical landscape for AAV gene therapies. Nat. Rev. Drug Discov. 20, 173–174 (2021).
Higashikawa, F. & Chang, L. Kinetic analyses of stability of simple and complex retroviral vectors. Virology 280, 124–131 (2001).
Lang, J. F., Toulmin, S. A., Brida, K. L., Eisenlohr, L. C. & Davidson, B. L. Standard screening methods underreport AAV-mediated transduction and gene editing. Nat. Commun. 10, 3415 (2019).
Zheng, X. et al. Massively parallel in vivo Perturb-seq reveals cell-type-specific transcriptional networks in cortical development. Cell 187, 3236–3248 (2024).
Santinha, A. J. et al. Transcriptional linkage analysis with in vivo AAV-Perturb-seq. Nature 622, 367–375 (2023).
Kutner, R. H., Zhang, X. Y. & Reiser, J. Production, concentration and titration of pseudotyped HIV-1-based lentiviral vectors. Nat. Protoc. 4, 495–505 (2009).
Brown, L. Y., Dong, W. & Kantor, B. An improved protocol for the production of lentiviral vectors. STAR Protoc. 1, 100152 (2020).
Challis, R. C. et al. Systemic AAV vectors for widespread and targeted gene delivery in rodents. Nat. Protoc. 14, 379–414 (2019).
Binan, L. et al. Simultaneous CRISPR screening and spatial transcriptomics reveals intracellular, intercellular, and functional transcriptional circuits. Preprint at bioRxiv https://doi.org/10.1101/2023.11.30.569494 (2023).
Gu, J. et al. Mapping multimodal phenotypes to perturbations in cells and tissue with CRISPRmap. Nat. Biotechnol. https://doi.org/10.1038/s41587-024-02386-x (2024).
Kudo, T. et al. Multiplexed, image-based pooled screens in primary cells and tissues with PerturbView. Nat. Biotechnol. https://doi.org/10.1038/s41587-024-02391-0 (2024).
Rood, J. E., Hupalowska, A. & Regev, A. Toward a foundation model of causal cell and tissue biology with a Perturbation Cell and Tissue Atlas. Cell 187, 4520–4545 (2024).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Joung, J. et al. Genome-scale CRISPR–Cas9 knockout and transcriptional activation screening. Nat. Protoc. 12, 828–863 (2017).
Yin, J. A. et al. Arrayed CRISPR libraries for the genome-wide activation, deletion and silencing of human protein-coding genes. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-024-01278-4 (2024).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Morris, J. A. et al. Discovery of target genes and pathways at GWAS loci by pooled single-cell CRISPR screens. Science 380, eadh7699 (2023).
Barrangou, R. & Doudna, J. A. Applications of CRISPR technologies in research and beyond. Nat. Biotechnol. 34, 933–941 (2016).
Nunez, J. K. et al. Genome-wide programmable transcriptional memory by CRISPR-based epigenome editing. Cell 184, 2503–2519.e17 (2021).
Huang, Y. H. et al. DNA epigenome editing using CRISPR–Cas SunTag-directed DNMT3A. Genome Biol. 18, 176 (2017).
Konstantakos, V., Nentidis, A., Krithara, A. & Paliouras, G. CRISPR–Cas9 gRNA efficiency prediction: an overview of predictive tools and the role of deep learning. Nucleic Acids Res. 50, 3616–3637 (2022).
Jensen, K. T. et al. Chromatin accessibility and guide sequence secondary structure affect CRISPR–Cas9 gene editing efficiency. FEBS Lett. 591, 1892–1901 (2017).
Bock, C. et al. High-content CRISPR screening. Nat. Rev. Methods Primers 2, 9 (2022).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Port, F. & Bullock, S. L. Augmenting CRISPR applications in Drosophila with tRNA-flanked sgRNAs. Nat. Methods 13, 852–854 (2016).
Xie, K., Minkenberg, B. & Yang, Y. Boosting CRISPR–Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl Acad. Sci. USA 112, 3570–3575 (2015).
Vidigal, J. & Ventura, A. Rapid and efficient one-step generation of paired gRNA CRISPR–Cas9 libraries. Nat. Commun. 6, 8083 (2015).
Xie, S., Cooley, A., Armendariz, D., Zhou, P. & Hon, G. C. Frequent sgRNA–barcode recombination in single-cell perturbation assays. PLoS ONE 13, e0198635 (2018).
Sanson, K. R. et al. Optimized libraries for CRISPR–Cas9 genetic screens with multiple modalities. Nat. Commun. 9, 5416 (2018).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 2, W171–W174 (2019).
Heigwer, F., Kerr, G. & Boutros, M. E-CRISP: fast CRISPR target site identification. Nat. Methods 11, 122–123 (2014).
Hanna, R. E. & Doench, J. G. Design and analysis of CRISPR–Cas experiments. Nat. Biotechnol. 38, 813–823 (2020).
Platt, R. J. et al. CRISPR–Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Wangensteen, K. J. et al. Combinatorial genetics in liver repopulation and carcinogenesis with a in vivo CRISPR activation platform. Hepatology 68, 663–676 (2018).
Zhou, H. et al. In vivo simultaneous transcriptional activation of multiple genes in the brain using CRISPR–dCas9-activator transgenic mice. Nat. Neurosci. 21, 440–446 (2018).
Li, K. et al. Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing. Nat. Commun. 11, 485 (2020).
Gemberling, M. P. et al. Transgenic mice for in vivo epigenome editing with CRISPR-based systems. Nat. Methods 8, 965–974 (2021).
Dow, L. E. et al. Apc restoration promotes cellular differentiation and reestablishes crypt homeostasis in colorectal cancer. Cell 161, P1539–P1552 (2015).
Katigbak, A., Robert, F., Paquet, M. & Pelletier, J. Inducible genome editing with conditional CRISPR–Cas9 mice. G3 8, 1627–1635 (2018).
Challis, R. C. et al. Adeno-associated virus toolkit to target diverse brain cells. Annu. Rev. Neurosci. 45, 447–469 (2022).
Chen, X. et al. Engineered AAVs for non-invasive gene delivery to rodent and non-human primate nervous systems. Neuron 110, 2242–2257.e6 (2022).
Swiech, L. et al. In vivo interrogation of gene function in the mammalian brain using CRISPR–Cas9. Nat. Biotechnol. 33, 102–106 (2015).
Bedbrook, C. N., Deverman, B. & Gradinaru, V. Viral strategies for targeting the central and peripheral nervous systems. Annu. Rev. Neurosci. 8, 323–348 (2018).
Issa, S. S., Shaimardanova, A. A., Solovyeva, V. V. & Rizvanov, A. A. Various AAV serotypes and their applications in gene therapy: an overview. Cells 12, 785 (2023).
Li, C. & Samulski, R. J. Engineering adeno-associated virus vectors for gene therapy. Nat. Rev. Genet. 21, 255–272 (2020).
Deverman, B. E. et al. Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain. Nat. Biotechnol. 34, 204–209 (2016).
Ojala, D. S. et al. In vivo selection of a computationally designed SCHEMA AAV library yields a novel variant for infection of adult neural stem cells in the SVZ. Mol. Ther. 26, 304–319 (2018).
Foust, K. et al. Intravascular AAV9 preferentially targets neonatal neurons and adult astrocytes. Nat. Biotechnol. 27, 59–65 (2009).
Brommel, C. M., Cooney, A. L. & Sinn, P. L. Adeno-associated virus-based gene therapy for lifelong correction of genetic disease. Hum. Gene Ther. 31, 985–995 (2020).
Replogle, J. M. et al. Combinatorial single-cell CRISPR screens by direct guide RNA capture and targeted sequencing. Nat. Biotechnol. 38, 954–961 (2020).
Feldman, D. et al. Optical pooled screens in human cells. Cell 179, 787–799.e17 (2019).
Replogle, J. M. et al. Mapping information-rich genotype-phenotype landscapes with genome-scale Perturb-seq. Cell 185, 2559–2575.e28 (2022).
Dvoretskova, E. et al. Spatial enhancer activation influences inhibitory neuron identity during mouse embryonic development. Nat. Neurosci. 27, 862–872 (2024).
Li, C. et al. Single-cell brain organoid screening identifies developmental defects in autism. Nature 621, 373–380 (2023).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Tanay, A. & Regev, A. Scaling single-cell genomics from phenomenology to mechanism. Nature 18, 331–338 (2017).
Vaishnav, E. D. et al. The evolution, evolvability and engineering of gene regulatory DNA. Nature 603, 455–463 (2022).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. eLife 5, e19760 (2016).
Morgens, D. W. et al. Genome-scale measurement of off-target activity using Cas9 toxicity in high-throughput screens. Nat. Commun. 8, 15178 (2017).
Gu, S. S. et al. Therapeutically increasing MHC-I expression potentiates immune checkpoint blockade. Cancer Discov. 11, 1524–1541 (2021).
Lawson, K. A. et al. Functional genomic landscape of cancer-intrinsic evasion of killing by T cells. Nature 586, 120–126 (2020).
Ma, H. et al. Pol III promoters to express small RNAs: delineation of transcription initiation. Mol. Ther. Nucleic Acids 3, e161 (2014).
Mullally, G. et al. 5′ Modifications to CRISPR–Cas9 gRNA can change the dynamics and size of R-loops and inhibit DNA cleavage. Nucleic Acids Res. 48, 6811–6823 (2020).
Saito, T. In vivo electroporation in the embryonic mouse central nervous system. Nat. Protoc. 1, 1552–1558 (2006).
Langford, D. J. et al. Coding of facial expressions of pain in the laboratory mouse. Nat. Methods 7, 447–449 (2010).
Yao, Z. et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317–332 (2023).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, P329–P337 (2019).
Lopez, R. et al. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Phillip, B. N. & Jeffrey, W. M. Model-based dimensionality reduction for single-cell RNA-seq using generalized bilinear models. Preprint at bioRxiv https://doi.org/10.1101/2023.04.21.537881 (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Antonsson, E. S. & Páll, M. Batch correction methods used in single cell RNA-sequencing analyses are often poorly calibrated. Preprint at bioRxiv https://doi.org/10.1101/2024.03.19.585562 (2024).
Wang, Q. et al. The allen mouse brain common coordinate framework: a 3D reference atlas. Cell 181, 936–953.e20 (2020).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Phipson, B. et al. propeller: testing for differences in cell type proportions in single cell data. Bioinformatics 38, 4720–4726 (2022).
Fleck, J. S. et al. Inferring and perturbing cell fate regulomes in human brain organoids. Nature 621, 365–372 (2022).
Renz, P. F. et al. In vivo single-cell CRISPR uncovers distinct TNF programmes in tumour evolution. Nature 632, 419–428 (2024).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Rubin, A. J. et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell 176, 361–376.e17 (2019).
Liscovitch-Brauer, N. et al. Profiling the genetic determinants of chromatin accessibility with scalable single-cell CRISPR screens. Nat. Biotechnol. 39, 1270–1277 (2021).
Ursu, O. et al. Massively parallel phenotyping of coding variants in cancer with Perturb-seq. Nat. Biotechnol. 40, 896–905 (2022).
Geiger-Schuller, K. et al. Systematically characterizing the roles of E3-ligase family members in inflammatory responses with massively parallel Perturb-seq. Preprint at bioRxiv https://doi.org/10.1101/2023.01.23.525198 (2023).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Acknowledgements
We thank B. Wu for advice on the scRNA-seq experiments; the Department of Animal Resources, Genomics Core and Flow Cytometry Core facilities at Scripps Research for technical assistance; N. Huynh, S. Simmons, B. Wu, J. Li, A. Selstad and E. Petty for comments on the manuscript; and all members of the Jin lab for their help and support. X.Z. and P.C.T. were supported by the Dorris Scholar Award. X.Z. was supported by Frank J. Dixon Graduate Fellowship. C.M.W. was supported by Skaggs-Oxford Fellowship and The Schimmel Family Endowed Fellowship. X.J. and this work were supported by the Simons Foundation for Autism Research Initiative Collaboration on Sex Differences (SFARI 736613), National Institute of Health (R01HG012819, R01MH137042), Impetus grant, One Mind Rising Star Award, Klingenstein-Simons Fellowship Award, G. Harold and Leila Y. Mathers Foundation, Larry L. Hillblom Foundation, Scripps Collaborative Innovative Fund, Chan Zuckerberg Initiative, Conrad Prebys Foundation, Pew Charitable Trusts, McKnight Foundation, Astera Institute and James Fickel.
Author information
Authors and Affiliations
Contributions
X.Z., P.C.T., C.M.W. and X.J. designed and performed the experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
X.J. and X.Z. are co-inventors on in vivo AAV-based Perturb-seq and CRISPR inventions filed by Scripps Research relating to the work in this protocol. P.C.T. and C.M.W. declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key reference
Zheng, X. et al. Cell 187, 3236–3248.e21 (2024): https://doi.org/10.1016/j.cell.2024.04.050
Extended data
Extended Data Fig. 1 FACS analysis and gating strategy for identifying BFP+ perturbed cells.
Dissociated cortical cell suspension from P7 mouse brains was stained with Vybrant™ DyeCycle™ Ruby and analyzed by FACS. First, cells are identified based on size (forward scatter, FSC-A) versus granularity (side scatter, SSC-A). Single cells were isolated by excluding doublets using SSC-A versus SSC-H. Then, a Vybrant (cell-permeable stain) versus FSC-A gate was used to remove cytosolic debris (Vybrant−). Alternatively, a viability dye (e.g., Sytox™ Red) can be used to exclude dead cells (SytoxRed+, not shown). Finally, the perturbed BFP+ population is defined by GFP (expressed constitutively in a Cas9 transgenic mouse line) versus mtagBFP (expressed in transduced cells) plot.
Supplementary information
Supplementary Table 1 (download XLSX )
AAV titration calculator.
Supplementary Software 1 and 2 (download ZIP )
2 Python scripts for generate gRNA oligo and map gRNA in fastq files separately.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zheng, X., Thompson, P.C., White, C.M. et al. Massively parallel in vivo Perturb-seq screening. Nat Protoc 20, 1733–1767 (2025). https://doi.org/10.1038/s41596-024-01119-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41596-024-01119-3
This article is cited by
-
Cutting-edge technologies in neural regeneration
Cell Regeneration (2025)
-
Single-cell CRISPR screening in mouse brain
Nature Protocols (2025)
-
Harnessing LLMs to decode genetic perturbations
Nature Computational Science (2025)